Structural Analysis of Jumbo Coliphage phAPEC6
Abstract
1. Introduction
2. Results and Discussion
2.1. Coliphage phAPEC6 is A Jumbo Phage
2.2. Most of the phAPEC6 Genome Encodes for Unknown Proteins
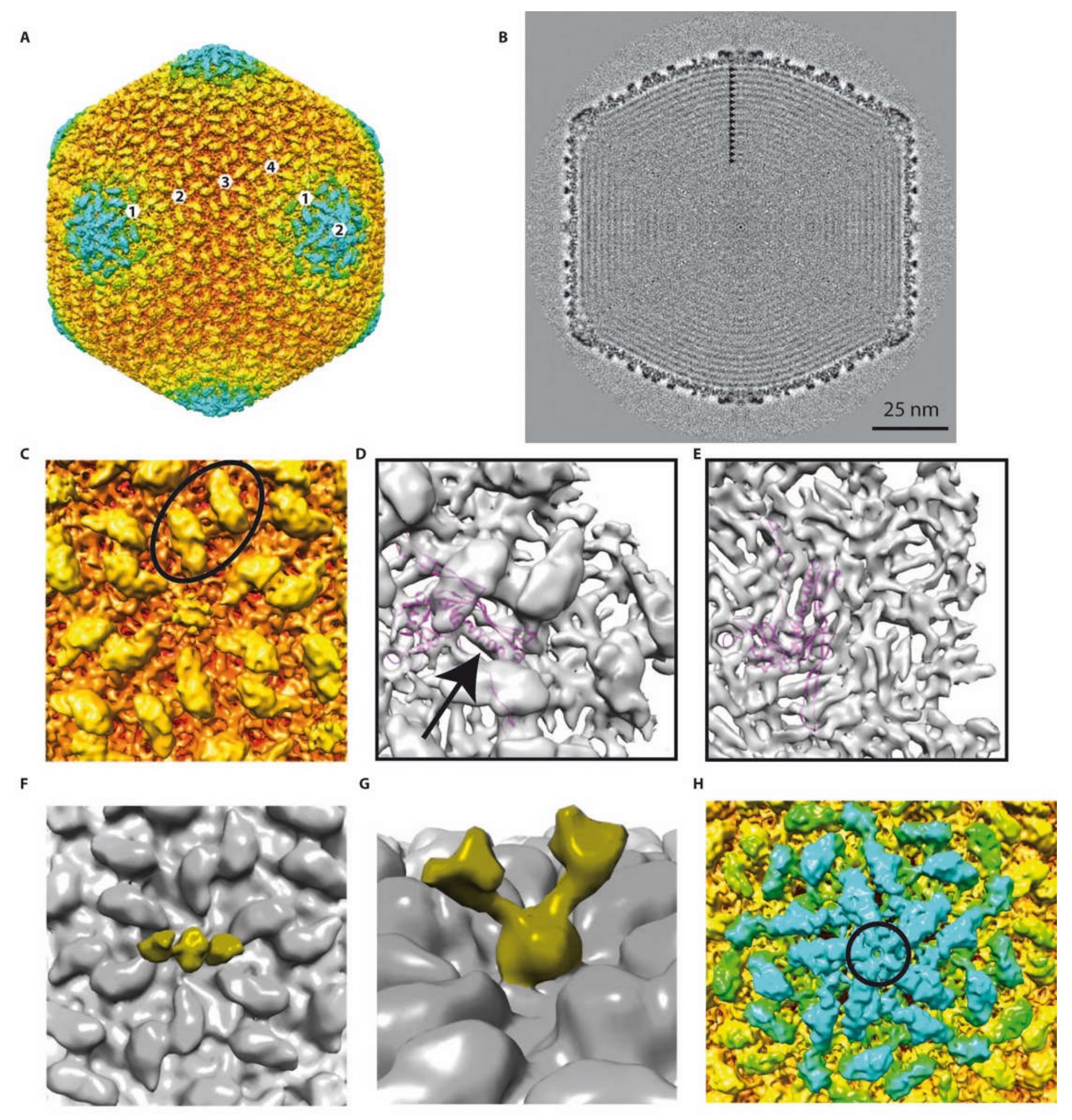
2.3. 3D Reconstruction of the phAPEC6 Capsid
2.4. 3D Reconstruction of the Entire Phage Tail
3. Materials and Methods
3.1. Molecular Analysis of phAPEC6
3.2. Electron Microscopy
3.3. Phage Head Reconstruction
3.4. 3D Reconstruction of phAPEC6 Full Tail
Author Contributions
Funding
Conflicts of Interest
Data Availability
References
- Hendrix, R.W. Jumbo bacteriophages. Curr. Top. Microbiol. Immunol. 2009, 328, 229–240. [Google Scholar] [PubMed]
- Van Etten, J.L.; Lane, L.C.; Dunigan, D.D. DNA viruses: The really big ones (giruses). Annu. Rev. Microbiol. 2010, 64, 83–99. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Gao, M. Jumbo bacteriophages: An overview. Front. Microbiol. 2017, 8, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Fokine, A.; Kostyuchenko, V.A.; Efimov, A.V.; Kurochkina, L.P.; Sykilinda, N.N.; Robben, J.; Volckaert, G.; Hoenger, A.; Chipman, P.R.; Battisti, A.J.; et al. A three-dimensional cryo-electron microscopy structure of the bacteriophage phiKZ head. J. Mol. Biol. 2005, 352, 117–124. [Google Scholar] [CrossRef]
- Thomas, J.A.; Hardies, S.C.; Rolando, M.; Hayes, S.J.; Lieman, K.; Carroll, C.A.; Weintraub, S.T.; Serwer, P. Complete genomic sequence and mass spectrometric analysis of highly diverse, atypical Bacillus thuringiensis phage 0305 phi 8-36. Virology 2007, 368, 405–421. [Google Scholar] [CrossRef]
- Thomas, J.A.; Weintraub, S.T.; Hakala, K.; Serwer, P.; Hardies, S.C. Proteome of the large Pseudomonas Myovirus 201 phi 2-1. Mol. Cell. Proteomics 2010, 9, 940–951. [Google Scholar] [CrossRef]
- Lecoutere, E.; Ceyssens, P.-J.; Miroshnikov, K.A.; Mesyanzhinov, V.V.; Krylov, V.N.; Noben, J.-P.; Robben, J.; Hertveldt, K.; Volckaert, G.; Lavigne, R. Identification and comparative analysis of the structural proteomes of phiKZ and EL, two giant Pseudomonas aeruginosa bacteriophages. Proteomics 2009, 9, 3215–3219. [Google Scholar] [CrossRef]
- Hua, J.; Huet, A.; Lopez, C.A.; Toropova, K.; Pope, W.H.; Duda, R.L.; Hendrix, R.W.; Conway, J.F. Capsids and genomes of jumbo-sized bacteriophages reveal the evolutionary reach of the HK97 fold. MBio 2017, 8, 1–15. [Google Scholar] [CrossRef]
- Day, A.; Ahn, J.; Salmond, G.P.C. Jumbo bacteriophages are represented within an increasing diversity of environmental viruses infecting the emerging phytopathogen, Dickeya solani. Front. Microbiol. 2018, 9, 2169. [Google Scholar] [CrossRef]
- Saad, A.M.; Soliman, A.M.; Kawasaki, T.; Fujie, M.; Nariya, H.; Shimamoto, T.; Yamada, T. Systemic method to isolate large bacteriophages for use in biocontrol of a wide-range of pathogenic bacteria. J. Biosci. Bioeng. 2018, 127, 73–78. [Google Scholar] [CrossRef]
- Sharma, R.; Pielstick, B.A.; Bell, K.A.; Nieman, T.B.; Stubbs, O.A.; Yeates, E.L.; Baltrus, D.A.; Grose, J.H. A novel, highly related jumbo family of bacteriophages that were isolated against Erwinia. Front. Microbiol. 2019, 10, 1533. [Google Scholar] [CrossRef] [PubMed]
- Baker, T.S.; Olson, N.H.; Fuller, S.D. Adding the third dimension to virus life cycles: Three-dimensional reconstruction of icosahedral viruses from cryo-electron micrographs. Microbiol. Mol. Biol. Rev. 1999, 63, 862–922. [Google Scholar] [CrossRef] [PubMed]
- Ackermann, H.W.; Nguyen, T.M. Sewage coliphages studied by electron microscopy. Appl. Environ. Microbiol. 1983, 45, 1049–1059. [Google Scholar] [CrossRef]
- Drulis-Kawa, Z.; Olszak, T.; Danis, K.; Majkowska-Skrobek, G.; Ackermann, H.-W. A giant Pseudomonas phage from Poland. Arch. Virol. 2014, 159, 567–572. [Google Scholar] [CrossRef] [PubMed]
- Šimoliunas, E.; Kaliniene, L.; Truncaite, L.; Zajančkauskaite, A.; Staniulis, J.; Kaupinis, A.; Ger, M.; Valius, M.; Meškys, R. Klebsiella phage vB_KleM-RaK2—A giant singleton virus of the family Myoviridae. PLoS ONE 2013, 8, e60717. [Google Scholar] [CrossRef] [PubMed]
- Moran, N.A. Microbial minimalism: Genome reduction in bacterial pathogens. Cell 2002, 108, 583–586. [Google Scholar] [CrossRef]
- Kim, M.S.; Hong, S.S.; Park, K.; Myung, H. Genomic analysis of bacteriophage PBECO4 infecting Escherichia coli O157:H7. Arch. Virol. 2013, 158, 2399–2403. [Google Scholar] [CrossRef]
- Amitsur, M.; Levitz, R.; Kaufmann, G. Bacteriophage T4 anticodon nuclease, polynucleotide kinase and RNA ligase reprocess the host lysine tRNA. EMBO J. 1987, 6, 2499–2503. [Google Scholar] [CrossRef]
- Cornelissen, A.; Ceyssens, P.-J.J.; Krylov, V.N.; Noben, J.-P.P.; Volckaert, G.; Lavigne, R. Identification of EPS-degrading activity within the tail spikes of the novel Pseudomonas putida phage AF. Virology 2012, 434, 251–256. [Google Scholar] [CrossRef]
- Krylov, V.N.; Dela Cruz, D.M.; Hertveldt, K.; Ackermann, H.-W. “phiKZ-like viruses”, a proposed new genus of myovirus bacteriophages. Arch. Virol. 2007, 152, 1955–1959. [Google Scholar] [CrossRef]
- Effantin, G.; Hamasaki, R.; Kawasaki, T.; Bacia, M.; Moriscot, C.; Weissenhorn, W.; Yamada, T.; Schoehn, G. Cryo-electron microscopy three-dimensional structure of the jumbo phage ΦRSL1 infecting the phytopathogen Ralstonia solanacearum. Structure 2013, 21, 298–305. [Google Scholar] [CrossRef] [PubMed]
- Helgstrand, C.; Wikoff, W.R.; Duda, R.L.; Hendrix, R.W.; Johnson, J.E.; Liljas, L. The refined structure of a protein catenane: The HK97 bacteriophage capsid at 3.44Å resolution. J. Mol. Biol. 2003, 334, 885–899. [Google Scholar] [CrossRef] [PubMed]
- Falco, S.C.; Laan, K.V.; Rothman-Denes, L.B. Virion-associated RNA polymerase required for bacteriophage N4 development. Proc. Natl. Acad. Sci. USA 1977, 74, 520–523. [Google Scholar] [CrossRef] [PubMed]
- Yamada, T.; Satoh, S.; Ishikawa, H.; Fujiwara, A.; Kawasaki, T.; Fujie, M.; Ogata, H. A jumbo phage infecting the phytopathogen Ralstonia solanacearum defines a new lineage of the Myoviridae family. Virology 2010, 398, 135–147. [Google Scholar] [CrossRef] [PubMed]
- Alberts, B.M.; Frey, L. T4 bacteriophage gene 32: A structural protein in the replication and recombination of DNA. Nature 1970, 227, 1313–1318. [Google Scholar] [CrossRef]
- Fokine, A.; Chipman, P.R.; Leiman, P.G.; Mesyanzhinov, V.V.; Rao, V.B.; Rossmann, M.G. Molecular architecture of the prolate head of bacteriophage T4. Proc. Natl. Acad. Sci. USA 2004, 101, 6003–6008. [Google Scholar] [CrossRef]
- Krylov, V.N.; Smirnova, T.A.; Minenkova, I.B.; Plotnikova, T.G.; Zhazikov, I.Z.; Khrenova, E.A. Pseudomonas bacteriophage phiKZ contains an inner body in its capsid. Can. J. Microbiol. 1984, 30, 758–762. [Google Scholar] [CrossRef]
- Thomas, J.A.; Rolando, M.R.; Carroll, C.A.; Shen, P.S.; Belnap, D.M.; Weintraub, S.T.; Serwer, P.; Hardies, S.C. Characterization of Pseudomonas chlororaphis myovirus 201-phi2-1 via genomic sequencing, mass spectrometry, and electron microscopy. Virology 2008, 376, 330–338. [Google Scholar] [CrossRef]
- Wu, W.; Thomas, J.A.; Cheng, N.; Black, L.W.; Steven, A.C. Bubblegrams reveal the inner body of bacteriophage phiKZ. Science 2012, 335, 182. [Google Scholar] [CrossRef]
- Chang, J.T.; Schmid, M.F.; Haase-Pettingell, C.; Weigele, P.R.; King, J.A.; Chiu, W. Visualizing the structural changes of bacteriophage Epsilon15 and its Salmonella host during infection. J. Mol. Biol. 2010, 402, 731–740. [Google Scholar] [CrossRef]
- Thomas, J.A.; Weintraub, S.T.; Wu, W.; Winkler, D.C.; Cheng, N.; Steven, A.C.; Black, L.W. Extensive proteolysis of head and inner body proteins by a morphogenetic protease in the giant Pseudomonas aeruginosa phage phiKZ. Mol. Microbiol. 2012, 84, 324–339. [Google Scholar] [CrossRef] [PubMed]
- Ohtsubo, M.; Kai, R.; Furuno, N.; Sekiguchi, T.; Sekiguchi, M.; Hayashida, H.; Kuma, K.; Miyata, T.; Fukushige, S.; Murotsu, T. Isolation and characterization of the active cDNA of the human cell cycle gene (RCC1) involved in the regulation of onset of chromosome condensation. Genes Dev. 1987, 1, 585–593. [Google Scholar] [CrossRef] [PubMed]
- McNulty, R.; Cardone, G.; Gilcrease, E.B.; Baker, T.S.; Casjens, S.R.; Johnson, J.E. Cryo-EM elucidation of the structure of bacteriophage P22 virions after genome release. Biophys. J. 2018, 114, 1295–1301. [Google Scholar] [CrossRef] [PubMed]
- Buttimer, C.; Hendrix, H.; Oliveira, H.; Casey, A.; Neve, H.; McAuliffe, O.; Paul Ross, R.; Hill, C.; Noben, J.P.; O’Mahony, J.; et al. Things are getting hairy: Enterobacteria bacteriophage vB_PcaM_CBB. Front. Microbiol. 2017, 8, 1–16. [Google Scholar] [CrossRef]
- Briers, Y.; Walmagh, M.; Puyenbroeck, V. Van; Cornelissen, A.; Cenens, W.; Aertsen, A.; Oliveira, H. Engineered endolysin based “ artilysins ” to combat multidrug resistant Gram-negative pathogens. MBio 2014, 5, 1–10. [Google Scholar] [CrossRef]
- De Smet, J.; Hendrix, H.; Blasdel, B.G.; Danis-Wlodarczyk, K.; Lavigne, R. Pseudomonas predators: Understanding and exploiting phage-host interactions. Nat. Rev. Microbiol. 2017, 15, 517–530. [Google Scholar] [CrossRef]
- Tsonos, J.; Oosterik, L.H.; Tuntufye, H.N.; Klumpp, J.; Butaye, P.; De Greve, H.; Hernalsteens, J.P.; Lavigne, R.; Goddeeris, B.M. A cocktail of in vitro efficient phages is not a guarantee for in vivo therapeutic results against avian colibacillosis. Vet. Microbiol. 2014, 171, 470–479. [Google Scholar] [CrossRef]
- Seemann, T. Shovill: Faster SPAdes assembly of Illumina reads. Available online: https://github.com/tseemann/shovill (accessed on 15 March 2020).
- Adriaenssens, E.M.; Ceyssens, P.J.; Dunon, V.; Ackermann, H.W.; Van Vaerenbergh, J.; Maes, M.; De Proft, M.; Lavigne, R. Bacteriophages LIMElight and LIMEzero of Pantoea agglomerans, belonging to the “phiKMV-like viruses”. Appl. Environ. Microbiol. 2011, 77, 3443–3450. [Google Scholar] [CrossRef]
- Merrill, B.D.; Ward, A.T.; Grose, J.H.; Hope, S. Software-based analysis of bacteriophage genomes, physical ends, and packaging strategies. BMC Genomics 2016, 17, 1–16. [Google Scholar] [CrossRef]
- Abbasifar, R.; Griffiths, M.W.; Sabour, P.M.; Ackermann, H.W.; Vandersteegen, K.; Lavigne, R.; Noben, J.P.; Alanis Villa, A.; Abbasifar, A.; Nash, J.H.E.; et al. Supersize me: Cronobacter sakazakii phage GAP32. Virology 2014, 460–461, 138–146. [Google Scholar] [CrossRef]
- Shevchenko, A.; Tomas, H.; Havli, J.; Olsen, J.V.; Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 2007, 1, 2856–2860. [Google Scholar] [CrossRef]
- Ceyssens, P.-J.; Minakhin, L.; Van den Bossche, A.; Yakunina, M.; Klimuk, E.; Blasdel, B.; De Smet, J.; Noben, J.-P.; Blasi, U.; Severinov, K.; et al. Development of giant bacteriophage phiKZ is independent of the host transcription apparatus. J. Virol. 2014, 88, 10501–10510. [Google Scholar] [CrossRef] [PubMed]
- Mas, G.; Guan, J.-Y.; Crublet, E.; Debled, E.C.; Moriscot, C.; Gans, P.; Schoehn, G.; Macek, P.; Schanda, P.; Boisbouvier, J. Structural investigation of a chaperonin in action reveals how nucleotide binding regulates the functional cycle. Sci. Adv. 2018, 4, eaau4196. [Google Scholar] [CrossRef] [PubMed]
- Fuller, S.D.; Butcher, S.J.; Cheng, R.H.; Baker, T.S. Three-dimensional reconstruction of icosahedral particles—The uncommon line. J. Struct. Biol. 1996, 116, 48–55. [Google Scholar] [CrossRef] [PubMed]
- Conway, J.F.; Steven, A.C. From cryoelectron micrographs to subnanometer resolution. J. Struct. Biol. 1999, 118, 106–118. [Google Scholar] [CrossRef] [PubMed]
- Van Heel, M. Angular reconstitution: A posteriori assignment of projection directions for 3D reconstruction. Ultramicroscopy 1987, 21, 111–123. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera: A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Frank, J.; Radermacher, M.; Penczek, P.; Zhu, J.; Li, Y.; Ladjadj, M.; Leith, A. SPIDER and WEB: Processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 1996, 116, 190–199. [Google Scholar] [CrossRef]




| Protein | Function | MW (kDa) | Unique Peptide Count | Sequence Coverage (%) |
|---|---|---|---|---|
| gp003 | DNA condensation protein | 41.39 | 4 | 11.6 |
| gp005 | DNA condensation protein | 40.25 | 2 | 5.7 |
| gp006 | hypothetical protein | 41.36 | 1 | 3.3 |
| gp007 | DNA condensation protein | 30.06 | 3 | 17.5 |
| gp009 | DNA condensation protein | 39.30 | 3 | 10.8 |
| gp010 | hypothetical protein | 41.39 | 1 | 3.1 |
| gp035c | chromosome segregation protein | 46.39 | 8 | 28.8 |
| gp037c | hypothetical protein | 14.83 | 5 | 48.0 |
| gp077 | hypothetical protein | 42.88 | 10 | 37.8 |
| gp078c | hypothetical protein | 29.73 | 4 | 21.4 |
| gp149c | hypothetical protein | 25.61 | 4 | 29.2 |
| gp150c | hypothetical protein | 21.33 | 5 | 34.8 |
| gp158c | hypothetical protein | 90.30 | 9 | 17.1 |
| gp159c | topoisomerase II small subunit | 54.54 | 2 | 8.1 |
| gp161c | hypothetical protein | 28.98 | 6 | 28.9 |
| gp162c | hypothetical protein | 60.05 | 10 | 25.4 |
| gp167c | hypothetical protein | 20.66 | 4 | 23.4 |
| gp179c | hypothetical protein | 21.48 | 2 | 15.0 |
| gp182c | hypothetical protein | 12.33 | 1 | 11.7 |
| gp192c | hypothetical protein | 36.90 | 7 | 32.0 |
| gp203c | RecA-like recombination protein | 41.58 | 2 | 6.7 |
| gp204c | single-stranded DNA binding protein | 38.89 | 4 | 17.1 |
| gp205c | hypothetical protein | 48.48 | 2 | 8.4 |
| gp210 | hypothetical protein | 20.92 | 6 | 69.8 |
| gp211 | hemagglutinin repeat-containing protein | 24.30 | 13 | 69.9 |
| gp215c | hypothetical protein | 26.56 | 5 | 26.3 |
| gp225c | major head protein | 42.18 | 14 | 74.9 |
| gp226c | scaffolding protein | 42.21 | 2 | 4.9 |
| gp227c | prohead core scaffold and protease | 23.01 | 2 | 15.0 |
| gp228c | hypothetical protein | 31.12 | 5 | 16.7 |
| gp230c | portal vertex protein of head | 64.56 | 9 | 21.1 |
| gp231c | hypothetical protein | 79.72 | 1 | 1.4 |
| gp238 | hypothetical protein | 27.50 | 2 | 12.8 |
| gp239 | hypothetical protein | 12.37 | 1 | 8.7 |
| gp240 | hypothetical protein | 85.40 | 5 | 10.0 |
| gp241 | lysozyme family protein | 95.90 | 7 | 9.9 |
| gp244 | baseplate wedge | 131.18 | 21 | 24.2 |
| gp245 | hypothetical protein | 381.88 | 17 | 7.6 |
| gp248 | proximal tail protein | 51.67 | 4 | 13.9 |
| gp250c | hypothetical protein | 40.33 | 9 | 40.2 |
| gp251c | hypothetical protein | 21.95 | 2 | 13.4 |
| gp253 | hypothetical protein | 40.99 | 14 | 65.6 |
| gp256 | tail fiber protein | 180.38 | 15 | 15.9 |
| gp259c | thymidylate synthase | 35.51 | 3 | 10.7 |
| gp262c | hypothetical protein | 25.02 | 1 | 6.4 |
| gp265c | ATPase | 49.52 | 6 | 19.0 |
| gp267c | hypothetical protein | 35.96 | 1 | 4.9 |
| gp268c | hypothetical protein | 51.12 | 6 | 17.8 |
| gp270c | hypothetical protein | 16.20 | 4 | 41.1 |
| gp273c | hypothetical protein | 40.42 | 7 | 29.9 |
| gp274c | hypothetical protein | 28.76 | 3 | 16.3 |
| gp275c | hypothetical protein | 25.07 | 2 | 9.2 |
| gp277c | tail sheath monomer | 97.05 | 36 | 68.0 |
| gp280c | neck protein | 29.93 | 4 | 20.5 |
| gp306c | hypothetical protein | 16.90 | 6 | 65.8 |
| gp314c | tail fiber protein | 105.34 | 8 | 13.0 |
| gp315c | hypothetical protein | 4.66 | 2 | 69.0 |
| gp323c | hypothetical protein | 98.62 | 4 | 5.6 |
| gp341c | PhoH-like protein | 53.21 | 8 | 30.9 |
| gp347 | hypothetical protein | 14.10 | 8 | 75.2 |
| gp549 | DNA condensation protein | 41.77 | 4 | 13.2 |
| gp551 | DNA condensation protein | 41.23 | 5 | 14.2 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wagemans, J.; Tsonos, J.; Holtappels, D.; Fortuna, K.; Hernalsteens, J.-P.; De Greve, H.; Estrozi, L.F.; Bacia-Verloop, M.; Moriscot, C.; Noben, J.-P.; et al. Structural Analysis of Jumbo Coliphage phAPEC6. Int. J. Mol. Sci. 2020, 21, 3119. https://doi.org/10.3390/ijms21093119
Wagemans J, Tsonos J, Holtappels D, Fortuna K, Hernalsteens J-P, De Greve H, Estrozi LF, Bacia-Verloop M, Moriscot C, Noben J-P, et al. Structural Analysis of Jumbo Coliphage phAPEC6. International Journal of Molecular Sciences. 2020; 21(9):3119. https://doi.org/10.3390/ijms21093119
Chicago/Turabian StyleWagemans, Jeroen, Jessica Tsonos, Dominique Holtappels, Kiandro Fortuna, Jean-Pierre Hernalsteens, Henri De Greve, Leandro F. Estrozi, Maria Bacia-Verloop, Christine Moriscot, Jean-Paul Noben, and et al. 2020. "Structural Analysis of Jumbo Coliphage phAPEC6" International Journal of Molecular Sciences 21, no. 9: 3119. https://doi.org/10.3390/ijms21093119
APA StyleWagemans, J., Tsonos, J., Holtappels, D., Fortuna, K., Hernalsteens, J.-P., De Greve, H., Estrozi, L. F., Bacia-Verloop, M., Moriscot, C., Noben, J.-P., Schoehn, G., & Lavigne, R. (2020). Structural Analysis of Jumbo Coliphage phAPEC6. International Journal of Molecular Sciences, 21(9), 3119. https://doi.org/10.3390/ijms21093119







