A Novel Ruthenium Based Coordination Compound Against Pathogenic Bacteria
Abstract
1. Introduction
2. Results
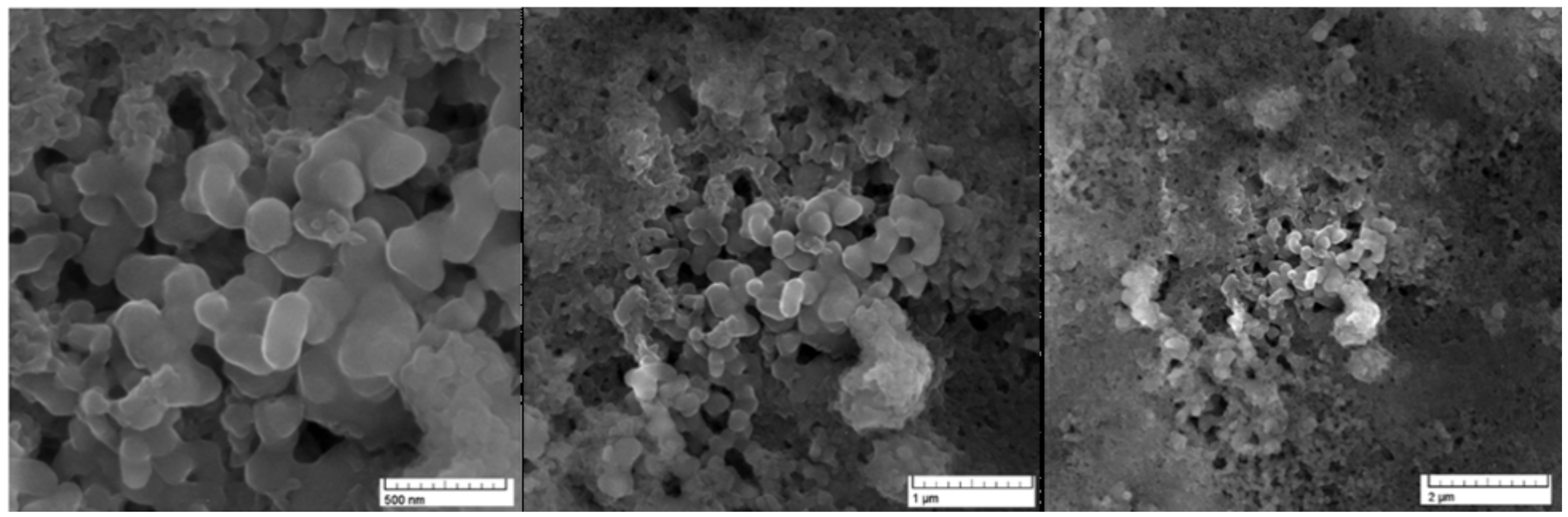
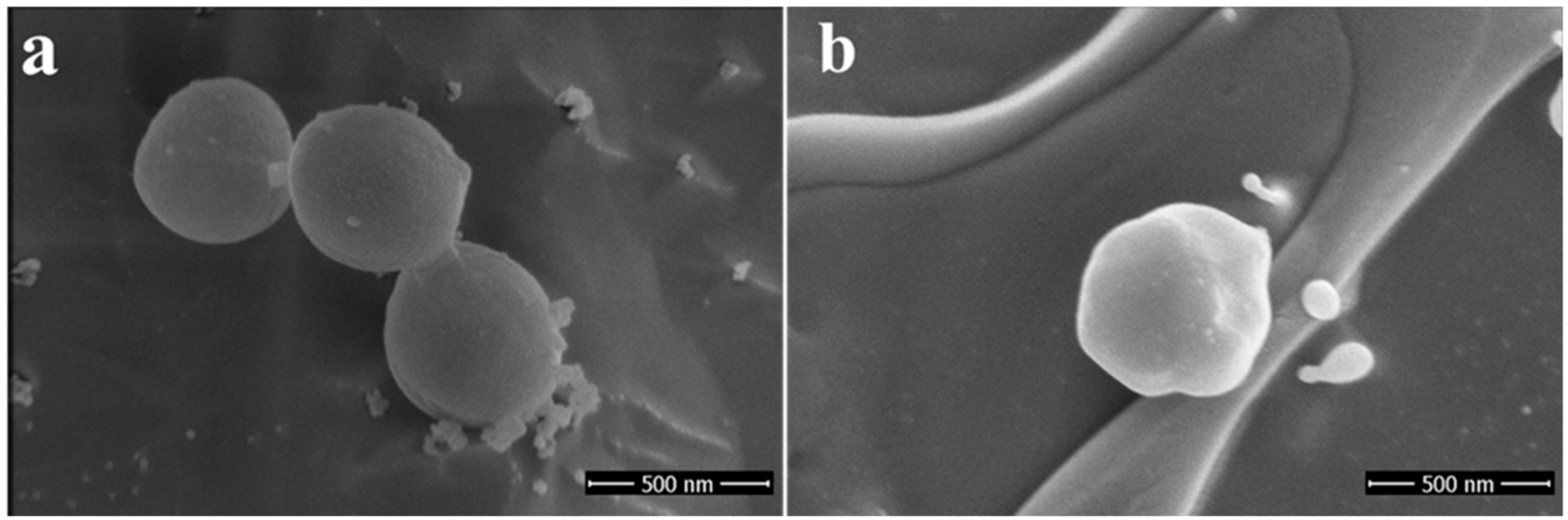
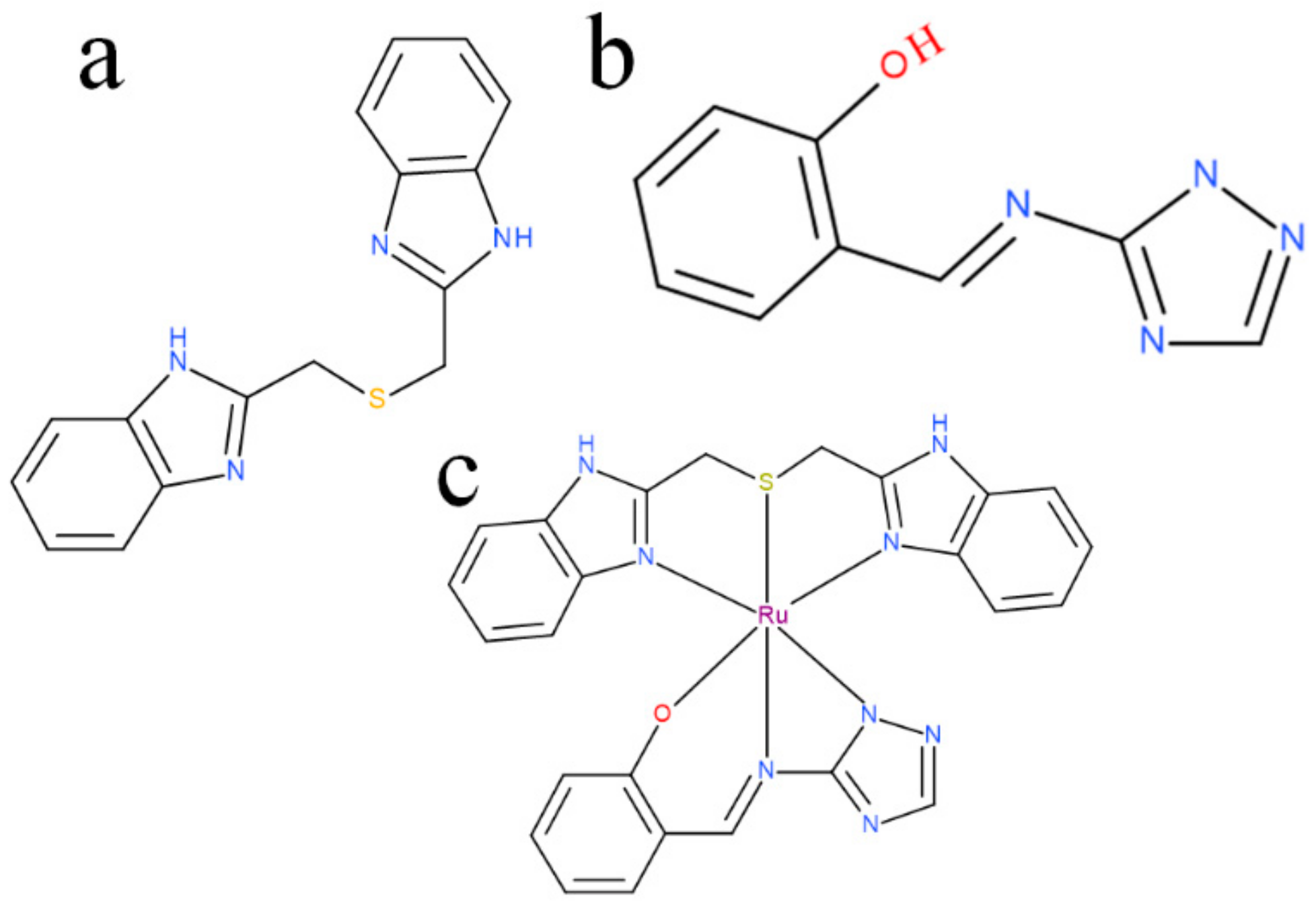
2.1. Chemical Synthesis and Electron Microscopy and Energy-Dispersive X-ray Spectroscopy (EDS) Confirmation
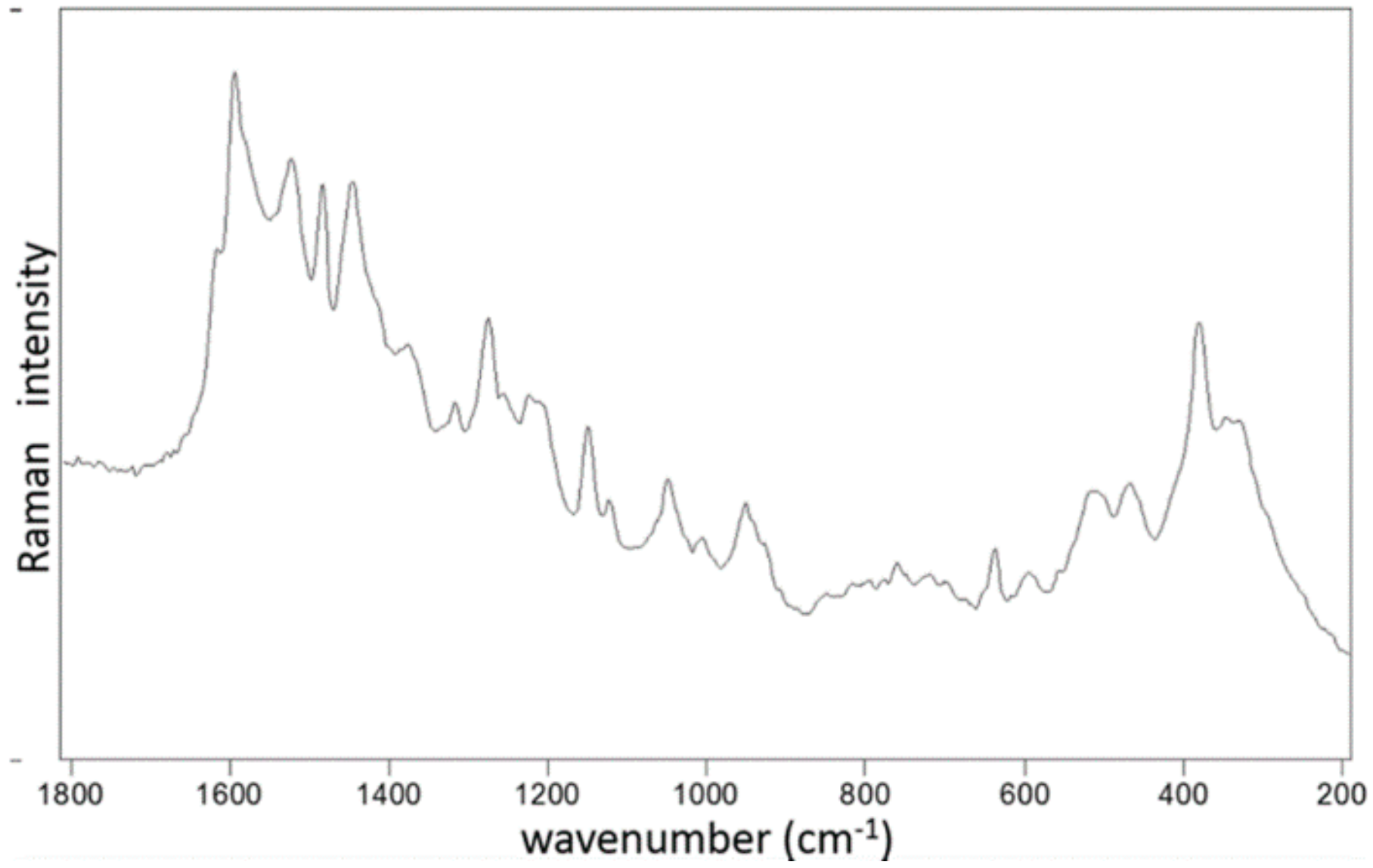
2.2. Characterization of the Ru Complexes by Raman Spectroscopy
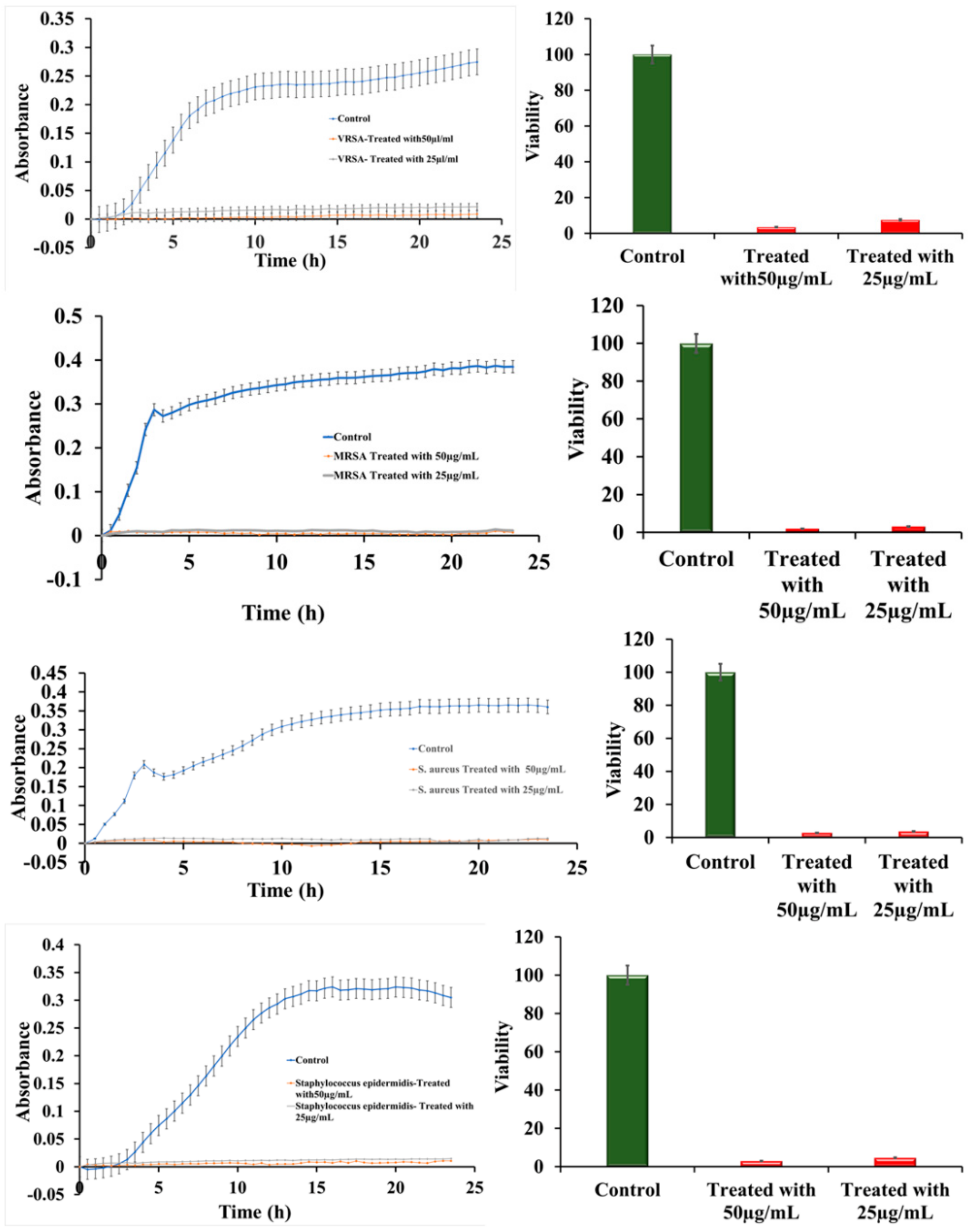
2.3. Growth Curve Snalysis
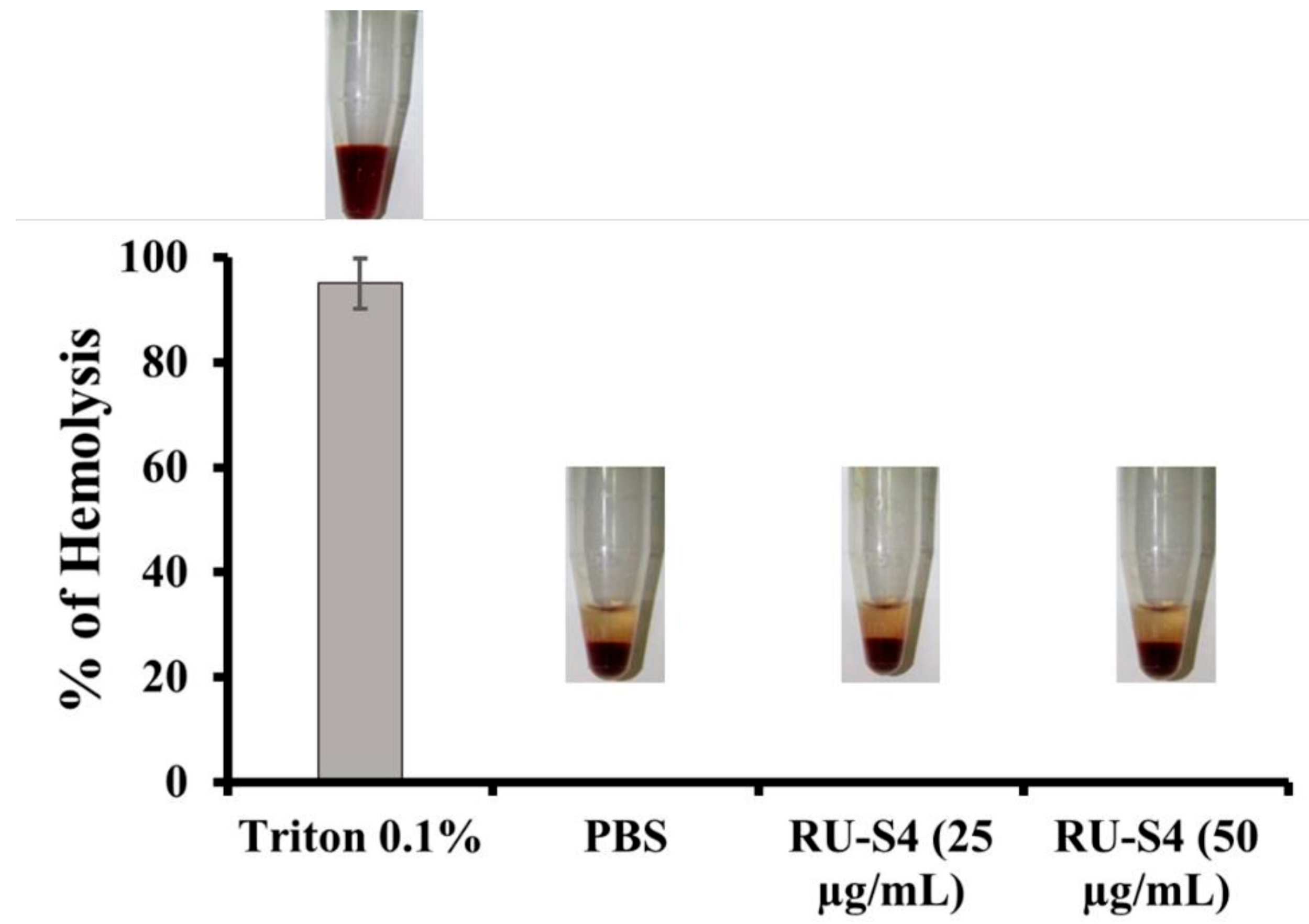
2.4. Hemolytic Assay
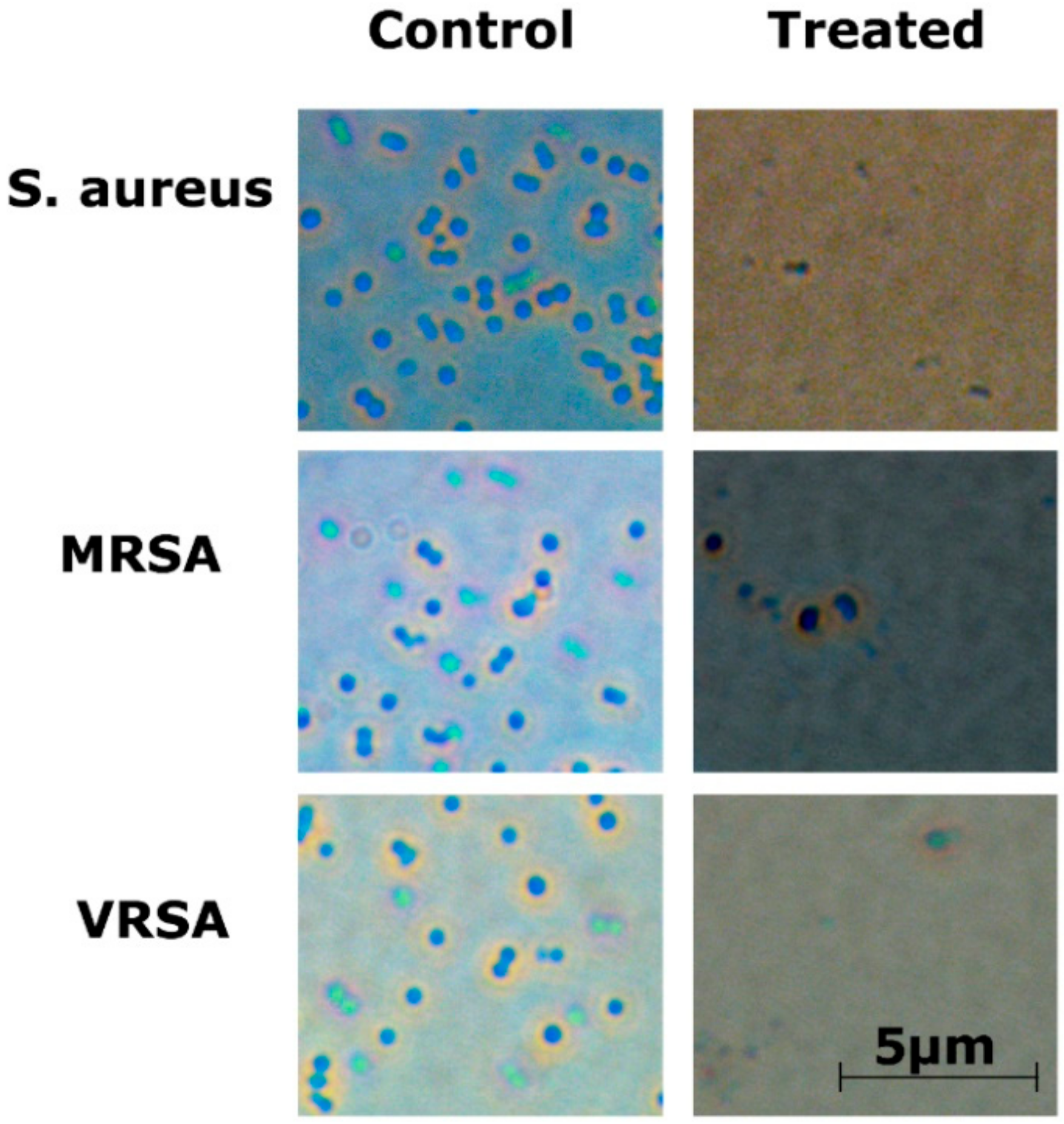
2.5. Optical Microscopy
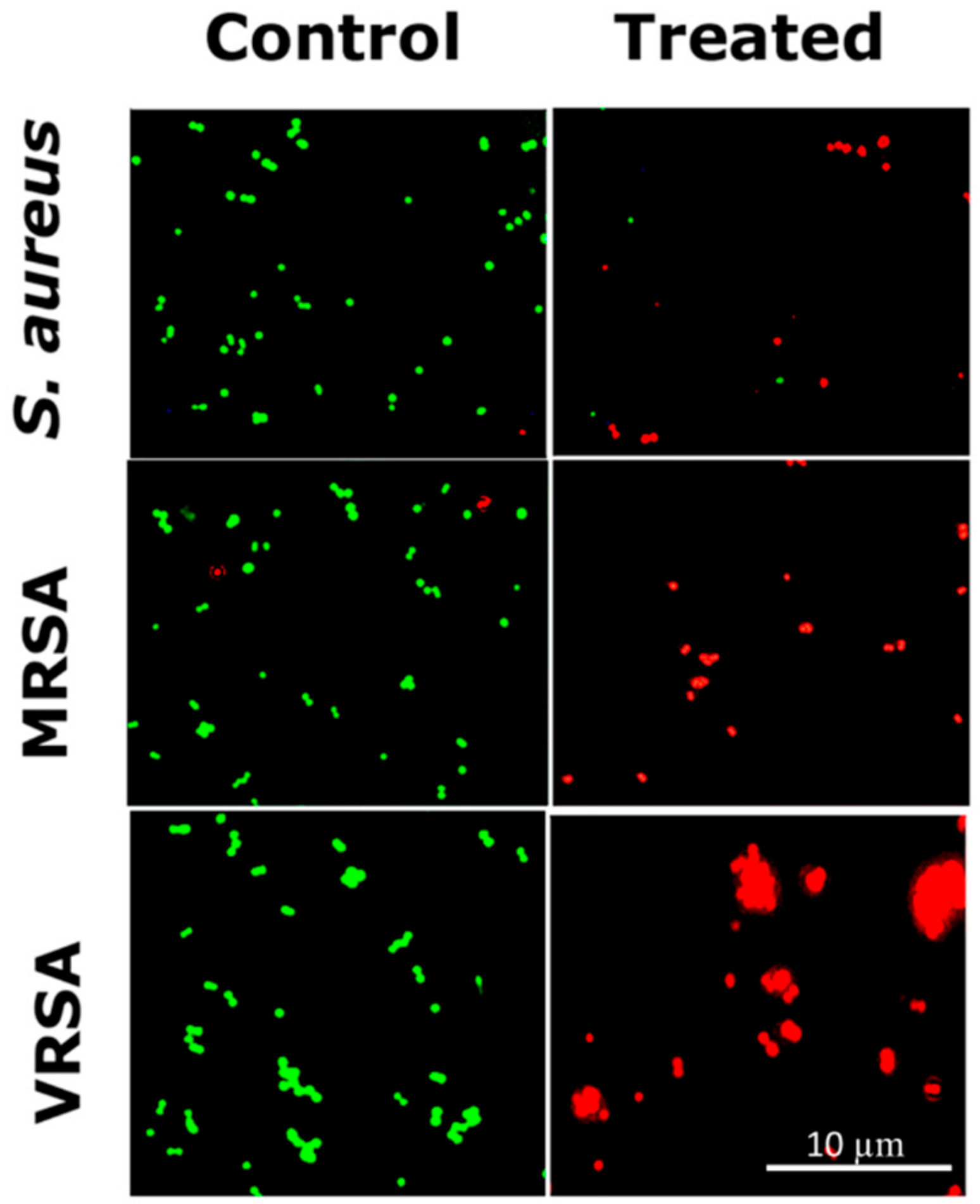
2.6. LIVE/DEAD Cell Imaging by Fluorescence Microscopy
2.7. Cryo-SEM Microscopy Imaging for Bacterial Cells Treatment with RU-S4
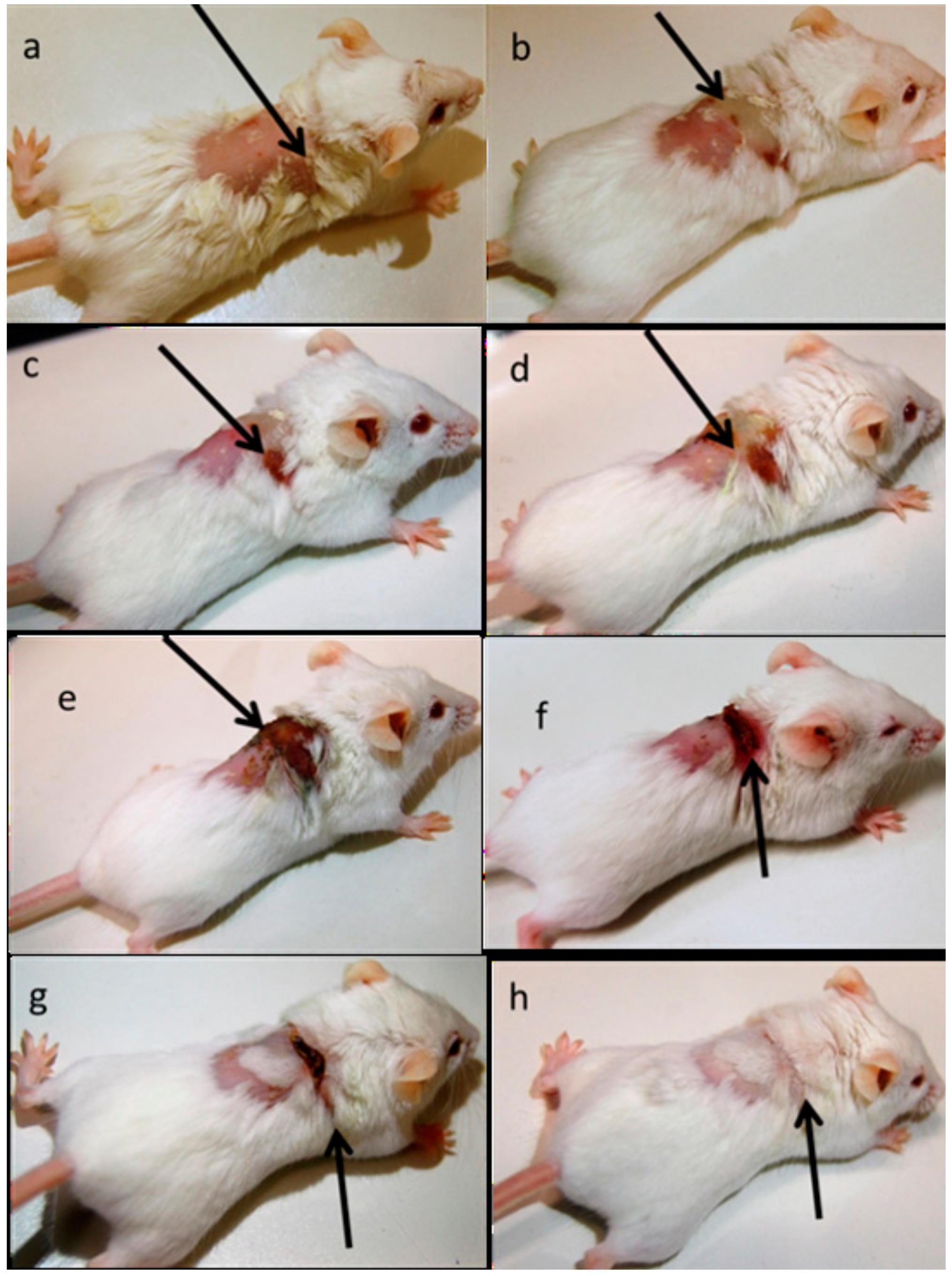
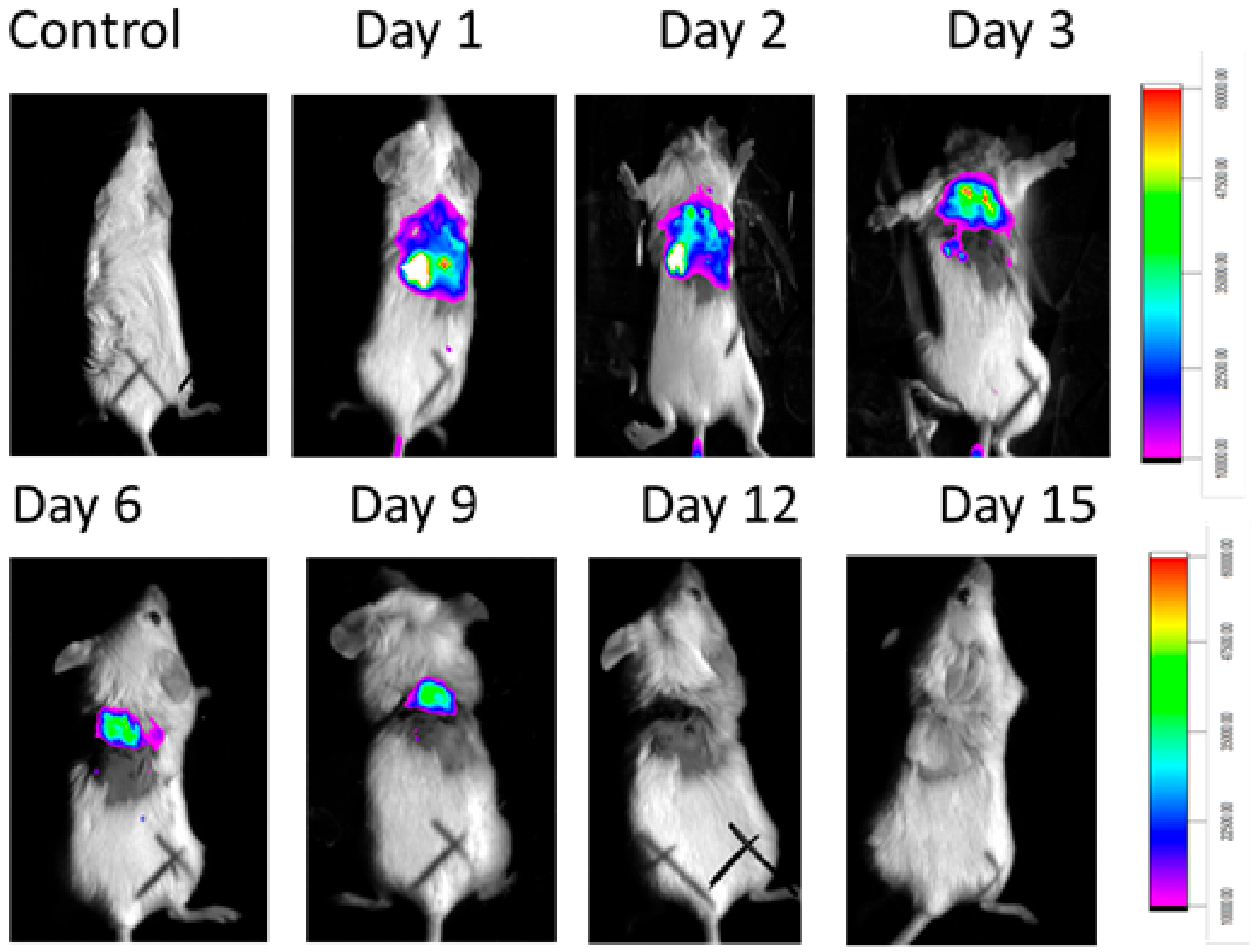
2.8. In Vivo Animal Model
3. Discussion
4. Materials and Methods
4.1. Synthesis of Ruthenium-Schiff Base-Benzimidazole Complex (RU-S4)
4.2. Electron Microscopy Imaging and Energy-dispersive X-ray Spectroscopy (EDS)
4.3. Raman Spectroscopy
4.4. Bacterial cultures
4.5. Hospital Samples
4.6. Bactericidal Effect of Ru-S4 and Growth Curve Analysis
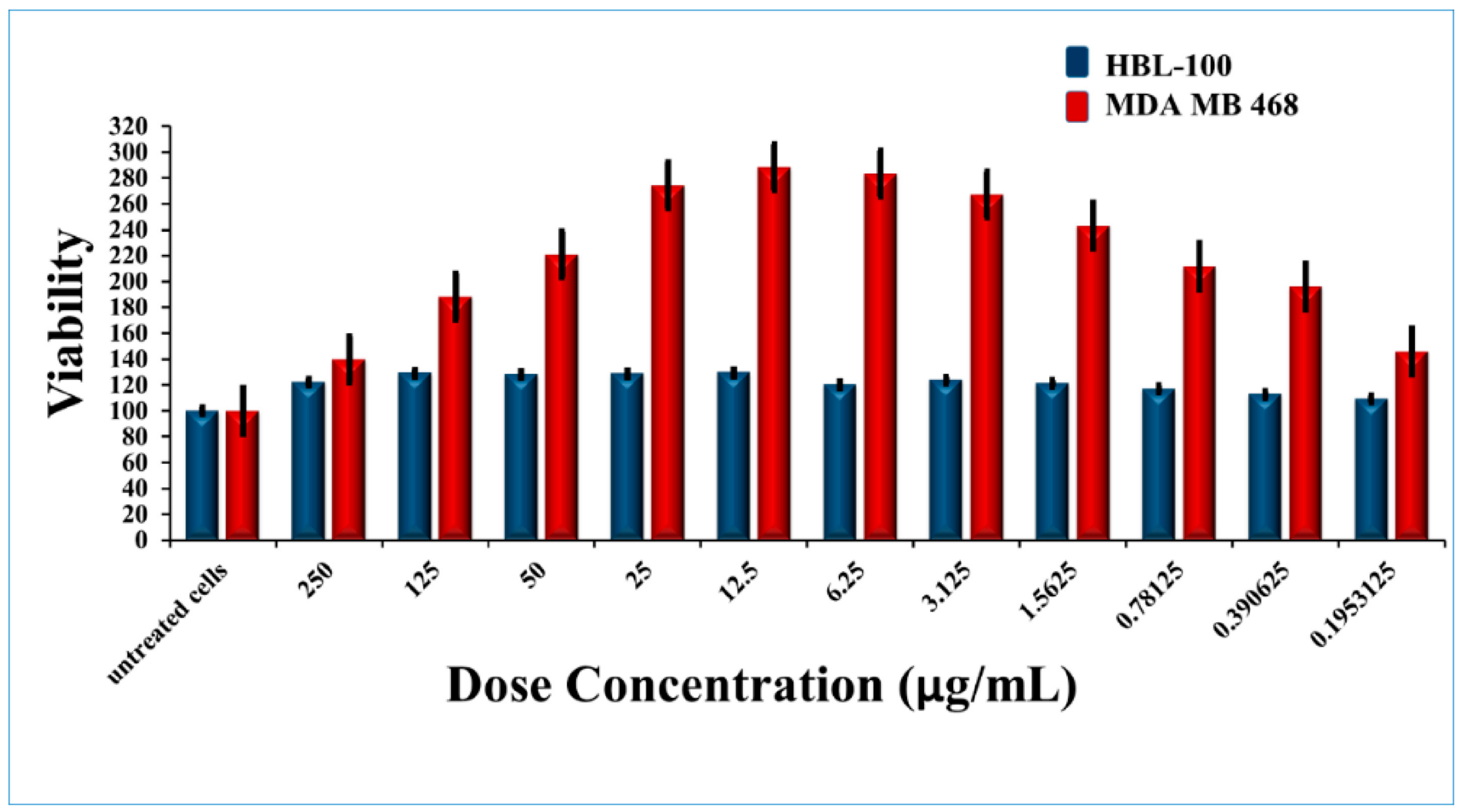
4.7. In Vitro Cytotoxicity Assay
4.8. Heamolytic Assay
4.9. Optical Microscopy and Fluorescence Microscopy Imaging of RU-S4 Treated Bacteria
4.10. Electron Microscopy Imaging of Bacteria with RU-S4 Compound
4.11. In vivo Infection Model Preparation
4.12. In Vivo Infection Treatment
4.13. In Vivo Imaging
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Boucher, H.; Miller, L.G.; Razonable, R.R. Serious Infections Caused by Methicillin-Resistant Staphylococcus aureus. Clin. Infect. Dis. 2010, 51 (Suppl. S2), S183–S197. [Google Scholar] [CrossRef] [PubMed]
- Loomba, P.; Taneja, J.; Mishra, B. Methicillin and vancomycin resistant S. aureus in hospitalized patients. J. Glob. Infect. Dis. 2010, 2, 275–283. [Google Scholar] [CrossRef]
- Tacconelli, E.; Carrara, E.; Savoldi, A.; Harbarth, S.; Mendelson, M.; Monnet, D.L.; Pulcini, C.; Kahlmeter, G.; Kluytmans, J.; Carmeli, Y.; et al. Discovery, research, and development of new antibiotics: The WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 2018, 18, 318–327. [Google Scholar] [CrossRef]
- Foster, T.J. Antibiotic resistance in Staphylococcus aureus. Current status and future prospects. FEMS Microbiol. Rev. 2017, 41, 430–449. [Google Scholar] [CrossRef] [PubMed]
- Boneca, I.G.; Chiosis, G. Vancomycin resistance: Occurrence, mechanisms and strategies to combat it. Expert Opin. Ther. Targets 2003, 7, 311–328. [Google Scholar] [CrossRef] [PubMed]
- Kugelberg, E.; Norström, T.; Petersen, T.K.; Duvold, T.; Andersson, D.I.; Hughes, D. Establishment of a Superficial Skin Infection Model in Mice by Using Staphylococcus aureus and Streptococcus pyogenes. Antimicrob. Agents Chemother. 2005, 49, 3435. [Google Scholar] [CrossRef] [PubMed]
- Kopel, P.; Wawrzak, D.; Langer, V.; Cihalova, K.; Chudobova, D.; Vesely, R.; Adam, V.; Kizek, R. Biological Activity and Molecular Structures of Bis(benzimidazole) and Trithiocyanurate Complexes. Molecules 2015, 20, 10360–10376. [Google Scholar] [CrossRef]
- Mazumdar, A.; Haddad, Y.; Milosavljevic, V.; Michalkova, H.; Guran, R.; Bhowmick, S.; Moulick, A. Peptide-Carbon Quantum Dots conjugate, Derived from Human Retinoic Acid Receptor Responder Protein 2, against Antibiotic-Resistant Gram Positive and Gram Negative Pathogenic Bacteria. Nanomaterials 2020, 10, 325. [Google Scholar] [CrossRef]
- Jayabalakrishnan, C.; Natarajan, K. Ruthenium(II) carbonyl complexes with tridentate Schiff bases and their antibacterial activity. Transit. Met. Chem. 2002, 27, 75–79. [Google Scholar] [CrossRef]
- Li, F.; Collins, J.G.; Keene, F.R. Ruthenium complexes as antimicrobial agents. Chem. Soc. Rev. 2015, 44, 2529–2542. [Google Scholar] [CrossRef]
- Mårtensson, A.K.F.; Bergentall, M.; Tremaroli, V.; Lincoln, P. Diastereomeric bactericidal effect of Ru(phenanthroline)(2) dipyridophenazine. Chirality 2016, 28, 713–720. [Google Scholar] [CrossRef] [PubMed]
- Selwin Joseyphus, R.; Sivasankaran Nair, M. Synthesis, characterization and antimicrobial activity of transition metal complexes with the Schiff base derived from imidazole-2-carboxaldehyde and glycylglycine. J. Coord. Chem. 2009, 62, 319–327. [Google Scholar] [CrossRef]
- Ranjith, P.K.; Rajeesh, P.; Haridas, K.R.; Susanta, N.K.; Guru Row, T.N.; Rishikesan, R.; Suchetha Kumari, N. Design and synthesis of positional isomers of 5 and 6-bromo-1-[(phenyl)sulfonyl]-2-[(4-nitrophenoxy)methyl]-1H-benzimidazoles as possible antimicrobial and antitubercular agents. Bioorg. Med. Chem. Lett. 2013, 23, 5228–5234. [Google Scholar] [CrossRef] [PubMed]
- Gullapelli, K.; Brahmeshwari, G.; Ravichander, M.; Kusuma, U. Synthesis, antibacterial and molecular docking studies of new benzimidazole derivatives. Egypt. J. Basic Appl. Sci. 2017, 4, 303–309. [Google Scholar] [CrossRef]
- Hameed, P.S.; Raichurkar, A.; Madhavapeddi, P.; Menasinakai, S.; Sharma, S.; Kaur, P.; Nandishaiah, R.; Panduga, V.; Reddy, J.; Sambandamurthy, V.K.; et al. Benzimidazoles: Novel Mycobacterial Gyrase Inhibitors from Scaffold Morphing. ACS Med. Chem. Lett. 2014, 5, 820–825. [Google Scholar] [CrossRef]
- Suwaiyan, A.; Zwarich, R.; Baig, N. Infrared and Raman spectra of benzimidazole. J. Raman Spectrosc. 1990, 21, 243–249. [Google Scholar] [CrossRef]
- Wang, G.; Harrison, A.; Li, X.; Whittaker, G.; Shi, J.; Wang, X.; Yang, H.; Cao, P.; Zhang, Z. Study of the adsorption of benzimidazole and 2-mercaptobenzothiazole on an iron surface by confocal micro-Raman spectroscopy. J. Raman Spectrosc. 2004, 35, 1016–1022. [Google Scholar] [CrossRef]
- Sondi, I.; Salopek-Sondi, B. Silver nanoparticles as antimicrobial agent: A case study on E. coli as a model for Gram-negative bacteria. J. Colloid Interface Sci. 2004, 275, 177–182. [Google Scholar] [CrossRef]
- Van Sorge, N.M.; Beasley, F.C.; Gusarov, I.; Gonzalez, D.J.; Blickwede, M.V.K.; Anik, S.; Borkowski, A.W.; Dorrestein, P.C.; Nudler, E.; Nizet, V. Methicillin-resistant Staphylococcus aureus bacterial nitric-oxide Synthase affects antibiotic sensitivity and skin abscess development. J. Biol. Chem. 2013, 288, 6417–6426. [Google Scholar] [CrossRef]
- Li, R.C.; Nix, D.E.; Schentag, J.J. New turbidimetric assay for quantitation of viable bacterial densities. Antimicrob. Agents Chemother. 1993, 37, 371–374. [Google Scholar] [CrossRef]
- Sur, V.P.; Mazumdar, A.; Kopel, P.; Moulick, A. Novel Ruthenium coordinate compound combined with Schiff base and benzimidazole as a potent antibacterial agent against VRSA and MRSA. MendelNet 2019, 26, 654–658. [Google Scholar]
- Stasiak-Różańska, L.; Błażejak, S.; Gientka, I. Effect of glycerol and dihydroxyacetone concentrations in the culture medium on the growth of acetic acid bacteria Gluconobacter oxydans ATCC 621. Eur. Food Res. Technol. 2014, 239, 453–461. [Google Scholar] [CrossRef]
- Hogenkamp, A.; Herías, M.V.; Tooten, P.C.J.; Veldhuizen, E.J.A.; Haagsman, H.P. Effects of surfactant protein D on growth, adhesion and epithelial invasion of intestinal Gram-negative bacteria. Mol. Immunol. 2007, 44, 3517–3527. [Google Scholar] [CrossRef] [PubMed]
- Greenwood, D.; Bidgood, K.; Turner, M. A comparison of the responses of staphylococci and streptococci to teicoplanin and vancomycin. J. Antimicrob. Chemother. 1987, 20, 155–164. [Google Scholar] [CrossRef] [PubMed]
- Jelinkova, P.; Splichal, Z.; Jimenez, A.M.J.; Haddad, Y.; Mazumdar, A.; Sur, V.P.; Milosavljevic, V.; Kopel, P.; Buchtelova, H.; Guran, R.; et al. Novel vancomycin-peptide conjugate as potent antibacterial agent against vancomycin-resistant Staphylococcus aureus. Infect. Drug Resist. 2018, 11, 1807–1817. [Google Scholar] [CrossRef]
- Heger, Z.; Merlos Rodrigo, M.A.; Michalek, P.; Polanska, H.; Masarik, M.; Vit, V.; Plevova, M.; Pacik, D.; Eckschlager, T.; Stiborova, M.; et al. Sarcosine Up-Regulates Expression of Genes Involved in Cell Cycle Progression of Metastatic Models of Prostate Cancer. PLoS ONE 2016, 11, e0165830. [Google Scholar] [CrossRef]
- Sur, V.P.; Kominkova, M.; Buchtova, Z.; Dolezelikova, K.; Zitka, O.; Moulick, A. CdSe QD Biosynthesis in Yeast Using Tryptone-Enriched Media and Their Conjugation with a Peptide Hecate for Bacterial Detection and Killing. Nanomaterials 2019, 9, 1463. [Google Scholar] [CrossRef]
- Wu, Y.; Liang, J.; Rensing, K.; Chou, T.-M.; Libera, M. Extracellular Matrix Reorganization during Cryo Preparation for Scanning Electron Microscope Imaging of Staphylococcus aureus Biofilms. Microsc. Microanal. 2014, 20, 1348–1355. [Google Scholar] [CrossRef]
- Goszczynska, A.; Kwiecien, H.; Fijalkowski, K. Synthesis and antibacterial activity of Schiff bases and amines derived from alkyl 2-(2-formyl-4-nitrophenoxy) alkanoates. Med. Chem. Res. 2015, 24, 3561–3577. [Google Scholar] [CrossRef]
- Thangadurai, T.D.; Ihm, S.-K. Novel bidentate ruthenium(III) Schiff base complexes: Synthetic, spectral, electrochemical, catalytic and antimicrobial studies. Transit. Met. Chem. 2004, 29, 189–195. [Google Scholar] [CrossRef]
- Li, X.; Gorle, A.K.; Ainsworth, T.D.; Heimann, K.; Woodward, C.E.; Grant Collins, J.; Richard Keene, F. RNA and DNA binding of inert oligonuclear ruthenium(ii) complexes in live eukaryotic cells. Dalton Trans. 2015, 44, 3594–3603. [Google Scholar] [CrossRef] [PubMed]
- Shoemaker, M.; Cohen, I.; Campbell, M. Reduction of MTT by aqueous herbal extracts in the absence of cells. J. Ethnopharmacol. 2004, 93, 381–384. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.V.; Scottwell, S.Ø.; Waugh, E.; McAdam, C.J.; Hanton, L.R.; Brooks, H.J.L.; Crowley, J.D. Antimicrobial Properties of Tris(homoleptic) Ruthenium(II) 2-Pyridyl-1,2,3-triazole “Click” Complexes against Pathogenic Bacteria, Including Methicillin-Resistant Staphylococcus aureus (MRSA). Inorg. Chem. 2016, 55, 9767–9777. [Google Scholar] [CrossRef] [PubMed]
- Mulyana, Y.; Weber, D.K.; Buck, D.P.; Motti, C.A.; Collins, J.G.; Keene, F.R. Oligonuclear polypyridylruthenium(ii) complexes incorporating flexible polar and non-polar bridges: Synthesis, DNA-binding and cytotoxicity. Dalton Trans. 2011, 40, 1510–1523. [Google Scholar] [CrossRef] [PubMed]
- Bolhuis, A.; Hand, L.; Marshall, J.E.; Richards, A.D.; Rodger, A.; Aldrich-Wright, J. Antimicrobial activity of ruthenium-based intercalators. Eur. J. Pharm. Sci. 2011, 42, 313–317. [Google Scholar] [CrossRef]
- Hegerova, D.; Vesely, R.; Cihalova, K.; Kopel, P.; Milosavljevic, V.; Heger, Z.; Hynek, D.; Guran, R.; Vaculovicova, M.; Sedlacek, P.; et al. Antimicrobial Agent Based on Selenium Nanoparticles and Carboxymethyl Cellulose for the Treatment of Bacterial Infections. J. Biomed. Nanotechnol. 2017, 13, 767–777. [Google Scholar] [CrossRef]
- Manicone, A.M.; McGuire, J.K. Matrix metalloproteinases as modulators of inflammation. Semin. Cell Dev. Biol. 2008, 19, 34–41. [Google Scholar] [CrossRef]
- Caley, M.P.; Martins, V.L.C.; O’Toole, E.A. Metalloproteinases and Wound Healing. Adv. Wound Care 2015, 4, 225–234. [Google Scholar] [CrossRef]
- Küster, T.; Hermann, C.; Hemphill, A.; Gottstein, B.; Spiliotis, M. Subcutaneous Infection Model Facilitates Treatment Assessment of Secondary Alveolar Echinococcosis in Mice. PLoS Negl. Trop. Dis. 2013, 7, e2235. [Google Scholar] [CrossRef]
- Chen, H.Y.; Zhang, M.; Li, B.W.; Chen, D.; Dong, X.Y.; Wang, Y.H.; Gu, Y.Q. Versatile antimicrobial peptide-based ZnO quantum dots for in vivo bacteria diagnosis and treatment with high specificity. Biomaterials 2015, 53, 532–544. [Google Scholar] [CrossRef]
- Joseyphus, R.S.; Nair, M.S. Antibacterial and Antifungal Studies on Some Schiff Base Complexes of Zinc(II). Mycobiology 2008, 36, 93–98. [Google Scholar] [CrossRef] [PubMed]
- Jayaseelan, P.; Prasad, S.; Vedanayaki, S.; Rajavel, R. Synthesis, characterization, anti-microbial, DNA binding and cleavage studies of Schiff base metal complexes. Arab. J. Chem. 2016, 9, S668–S677. [Google Scholar] [CrossRef]
- Yang, Y.; Liao, G.; Fu, C. Recent Advances on Octahedral Polypyridyl Ruthenium(II) Complexes as Antimicrobial Agents. Polymers 2018, 10, 650. [Google Scholar] [CrossRef] [PubMed]
- Clarke, M.J.; Zhu, F.; Frasca, D.R. Non-Platinum Chemotherapeutic Metallopharmaceuticals. Chem. Rev. 1999, 99, 2511–2534. [Google Scholar] [CrossRef] [PubMed]
- Weder, J.E.; Dillon, C.T.; Hambley, T.W.; Kennedy, B.J.; Lay, P.A.; Biffin, J.R.; Regtop, H.L.; Davies, N.M. Copper complexes of non-steroidal anti-inflammatory drugs: An opportunity yet to be realized. Coord. Chem. Rev. 2002, 232, 95–126. [Google Scholar] [CrossRef]
- Bistrović, A.; Krstulović, L.; Stolić, I.; Drenjančević, D.; Talapko, J.; Taylor, M.C.; Kelly, J.M.; Bajić, M.; Raić-Malić, S. Synthesis, anti-bacterial and anti-protozoal activities of amidinobenzimidazole derivatives and their interactions with DNA and RNA. J. Enzym. Inhib. Med. Chem. 2018, 33, 1323–1334. [Google Scholar] [CrossRef]
- Spunda, R.; Hruby, J.; Mericka, P.; Mlcek, M.; Pecha, O.; Splith, K.; Schmelzle, M.; Krenzien, F.; Lindner, J.; Matia, I.; et al. Immunosuppressive protocols with tacrolimus after cryopreserved aortal allotransplantation in rats. PLoS ONE 2018, 13, e0201984. [Google Scholar] [CrossRef]
- Gargiulo, S.; Greco, A.; Gramanzini, M.; Esposito, S.; Affuso, A.; Brunetti, A.; Vesce, G. Mice Anesthesia, Analgesia, and Care, Part I: Anesthetic Considerations in Preclinical Research. ILAR J. 2012, 53, E55–E69. [Google Scholar] [CrossRef]
- Malachowa, N.; Kobayashi, S.D.; Braughton, K.R.; DeLeo, F.R. Mouse Model of Staphylococcus aureus Skin Infection. In Mouse Models of Innate Immunity: Methods and Protocols; Allen, I.C., Ed.; Humana Press: Totowa, NJ, USA, 2013; pp. 109–116. [Google Scholar]
- Pletzer, D.; Mansour, S.C.; Wuerth, K.; Rahanjam, N.; Hancock, R.E.W. New Mouse Model for Chronic Infections by Gram-Negative Bacteria Enabling the Study of Anti-Infective Efficacy and Host-Microbe Interactions. MBio 2017, 8, e00140-17. [Google Scholar] [CrossRef]
- Hussain, S.; Joo, J.; Kang, J.; Kim, B.; Braun, G.B.; She, Z.-G.; Kim, D.; Mann, A.P.; Mölder, T.; Teesalu, T.; et al. Antibiotic-loaded nanoparticles targeted to the site of infection enhance antibacterial efficacy. Nat. Biomed. Eng. 2018, 2, 95–103. [Google Scholar] [CrossRef]
- Mölne, L.; Tarkowski, A. An Experimental Model of Cutaneous Infection Induced by Superantigen-Producing Staphylococcus aureus. J. Investig. Dermatol. 2000, 114, 1120–1125. [Google Scholar] [CrossRef] [PubMed]
- Sykes, E.A.; Dai, Q.; Tsoi, K.M.; Hwang, D.M.; Chan, W.C.W. Nanoparticle exposure in animals can be visualized in the skin and analysed via skin biopsy. Nat. Commun. 2014, 5, 3796. [Google Scholar] [CrossRef] [PubMed]
- Koch, M.A. Chapter 18—Experimental Modeling and Research Methodology. In The Laboratory Rat, 2nd ed.; Suckow, M.A., Weisbroth, S.H., Franklin, C.L., Eds.; Academic Press: Burlington, NJ, USA, 2006; pp. 587–625. [Google Scholar]
- Sevgi, M.; Toklu, A.; Vecchio, D.; Hamblin, M.R. Topical antimicrobials for burn infections—An update. Recent Pat. Anti-Infect. Drug Discov. 2013, 8, 161–197. [Google Scholar] [CrossRef] [PubMed]
- O’Toole, M.G.; Henderson, R.M.; Soucy, P.A.; Fasciotto, B.H.; Hoblitzell, P.J.; Keynton, R.S.; Ehringer, W.D.; Gobin, A.S. Curcumin Encapsulation in Submicrometer Spray-Dried Chitosan/Tween 20 Particles. Biomacromolecules 2012, 13, 2309–2314. [Google Scholar] [CrossRef]
- Blazkova, I.; Viet Nguyen, H.; Kominkova, M.; Konecna, R.; Chudobova, D.; Krejcova, L.; Kopel, P.; Hynek, D.; Zitka, O.; Beklova, M.; et al. Fullerene as a transporter for doxorubicin investigated by analytical methods and in vivo imaging. Electrophoresis 2013, 35, 1040–1049. [Google Scholar] [CrossRef]
- Merritt, J.; Senpuku, H.; Kreth, J. Let there be bioluminescence: Development of a biophotonic imaging platform for in situ analyses of oral biofilms in animal models. Environ. Microbiol. 2015, 18, 174–190. [Google Scholar] [CrossRef]
- Moulick, A.; Blazkova, I.; Milosavljevic, V.; Fohlerova, Z.; Hubalek, J.; Kopel, P.; Vaculovicova, M.; Adam, V.; Kizek, R. Application of CdTe/ZnSe Quantum Dots in In Vitro Imaging of Chicken Tissue and Embryo. Photochem. Photobiol. 2014, 91, 417–423. [Google Scholar] [CrossRef]


| Animal Set | Purpose |
|---|---|
| First set | Infected model with drug administration |
| Second set | Infected model without drug administration |
| Third set | Uninfected control |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sur, V.P.; Mazumdar, A.; Kopel, P.; Mukherjee, S.; Vítek, P.; Michalkova, H.; Vaculovičová, M.; Moulick, A. A Novel Ruthenium Based Coordination Compound Against Pathogenic Bacteria. Int. J. Mol. Sci. 2020, 21, 2656. https://doi.org/10.3390/ijms21072656
Sur VP, Mazumdar A, Kopel P, Mukherjee S, Vítek P, Michalkova H, Vaculovičová M, Moulick A. A Novel Ruthenium Based Coordination Compound Against Pathogenic Bacteria. International Journal of Molecular Sciences. 2020; 21(7):2656. https://doi.org/10.3390/ijms21072656
Chicago/Turabian StyleSur, Vishma Pratap, Aninda Mazumdar, Pavel Kopel, Soumajit Mukherjee, Petr Vítek, Hana Michalkova, Markéta Vaculovičová, and Amitava Moulick. 2020. "A Novel Ruthenium Based Coordination Compound Against Pathogenic Bacteria" International Journal of Molecular Sciences 21, no. 7: 2656. https://doi.org/10.3390/ijms21072656
APA StyleSur, V. P., Mazumdar, A., Kopel, P., Mukherjee, S., Vítek, P., Michalkova, H., Vaculovičová, M., & Moulick, A. (2020). A Novel Ruthenium Based Coordination Compound Against Pathogenic Bacteria. International Journal of Molecular Sciences, 21(7), 2656. https://doi.org/10.3390/ijms21072656









