Abstract
Kernel hardness is a key trait of wheat seeds, largely controlled by two tightly linked genes Puroindoline a and b (Pina and Pinb). Genes homologous to Pinb, namely Pinb2, have been studied. Whether these genes contribute to kernel hardness and other important seed traits remains inconclusive. Using the high-quality bread wheat reference genome, we show that PINB2 are encoded by three homoeologous loci Pinb2 not syntenic to the Hardness locus, with Pinb2-7A locus containing three tandem copies. PINB2 proteins have several features conserved for the Pin/Pinb2 phylogenetic cluster but lack a structural basis of significant impact on kernel hardness. Pinb2 are seed-specifically expressed with varied expression levels between the homoeologous copies and among wheat varieties. Using the high-quality genome information, we developed new Pinb2 allele specific markers and demonstrated their usefulness by 1) identifying new Pinb2 alleles in Triticeae species; and 2) performing an association analysis of Pinb2 with kernel hardness. The association result suggests that Pinb2 genes may have no substantial contribution to kernel hardness. Our results provide new insights into Pinb2 evolution and expression and the new allele-specific markers are useful to further explore Pinb2’s contribution to seed traits in wheat.
1. Introduction
Wheat (Triticum aestivum L.) is one of the major stable crops on Earth, feeding around 40% of the world population. Kernel hardness directly affects a set of physical and chemical properties of wheat seeds, such as water absorption, starch damage and flour particles [1,2]. Kernel hardness is one of the key traits in wheat seeds, largely determining milling quality and influencing the end-use qualities [3,4]. The major genetic determinant of wheat kernel hardness is the Hardness locus (Ha) on chromosome 5DS, harboring three closely linked genes: Grain Softness Protein-1 (GSP-1), Puroindoline a (Pina) and Puroindoline b (Pinb). The starch granule-associated protein, Friabilin, determines kernel hardness and is composed of PINA and PINB, encoded by the Pina and Pinb genes, respectively [5,6,7]. Genetic studies and transgenic studies show that wildtype PINA and PINB proteins confers a soft kernel phenotype [8,9,10,11,12,13,14,15,16]. Certain Pina or Pinb mutations and their combinations lead to hard kernel phenotypes, with slight variation in kernel hardness between the alleles of Pina and/or Pinb [17]. As the major causal genes of kernel hardness, the allelic diversity of Pina/Pinb has been extensively investigated in a wide range of wheat germplasm, revealing 26 alleles of Pina and 33 of Pinb, as well as a few double null alleles [18,19,20,21,22,23,24,25]. Genotype–phenotype association studies show that the diverse Pin alleles are related to the phenotypic variations in kernel hardness.
Due to the functional importance of PINs in wheat, efforts have been made to search for genes that are potentially homologous to Puroindolines using the wheat Expressed Sequence Tag (EST) database, resulting in the identification of several transcripts known as Pinb-like genes (also known as Pinb-2v) [26,27]. Due to their high sequence similarity to Pinb (~60%), the molecular characterization of these Pinb-like genes and genotype–phenotype correlations have drawn research attentions. Physical mapping of the Pinb-2v genes, using several wheat genetic stocks, proves that Pinb-2v1 is located on chromosome 7D, and Pinb-2v2 and Pinb-2v3 are allelic and located on 7B, with Pinb-2v4 or Pinb-2v5 located on 7A [27,28,29,30], indicating that these Pinb-2v genes might be homoeologous.
Mining for Pinb-2v genotypic diversity revealed 23 variants, suggesting less sequence diversity compared to Pina or Pinb [29,31]. With the development of Pinb-2v genotyping primers specific for Pinb-2v1, 2v2, 2v3, and 2v4 genotype–phenotype association has been studied in several collections of wheat varieties, from which different results were reported [28,29,32,33]. Chen et al. found that Pinb-2v3 was associated with preferable grain traits and higher kernel hardness compared to those possessing Pinb-2v2 in soft wheat varieties [26]. The results of Pinb-2v’s impacts on kernel hardness have been supported by association analysis using other wheat populations [34]. Another association mapping study that considered population structure and kinship showed that Pinb-2v variants were associated with semolina extraction but not kernel hardness [32]. By contrast, a genotype–phenotype study surveying representative U.S. wheat accessions did not support an ascertained role of Pinb-2v in kernel hardness [33]. An explanation for this could be that the expression levels of Pinb-2v variants (2v1, 2v2, 2v3 and 2v4) were much lower compared to that of Pinb, while varied expression levels were also observed among the Pinb-2v variants [35].
It is still inconclusive whether the Pinb-2v loci or particular alleles could influence kernel hardness or other kernel traits, even at a lesser extent than Pina/Pinb. Nevertheless, the prevalent Pinb-2v variants identified thus far have limited sequence polymorphisms within open reading frames (ORFs). The Pinb-2v genotyping primers were specific for variants 2v1, 2v2, 2v3 and 2v4, with Pinb-2v2 and Pinb-2v3 being the most frequently genotyped. Previously, the genomic resources were unavailable to address these critical issues of Pinb-2v variants. More recently, owing to the advance in new genomics technologies, such as NRGene deNovoMagic, PacBio single-molecule sequencing, and optical mapping, the contiguous and well-ordered genome assemblies with high-quality annotations are now available in several Triticeae species, including bread wheat (Triticum aestivum), wild emmer wheat (Triticum turgidum spp. dicoccoides), the progenitor of the wheat D genome, Aegilops tauschii, and that of the wheat A genome Triticum urartu [36,37,38,39,40]. These advanced genomics technologies have also substantially contributed to several unambiguously reconstructed, contiguous monocot genomes (maize, broomcorn millet, sugarcane, etc.), as well as to the chromosomal regions with tandem gene clusters and extensive haplotype variations in the gene family [41,42,43,44]. With the bread wheat genome (International Wheat Genome Sequencing Consortium (IWGSC) RefSeq v1.0 for the cultivar Chinese Spring (CS42)), we show that the Pinb-2v genes are encoded by five gene models at three homoeologous loci, Pinb2-7A, Pinb2-7B and Pinb2-7D, with the Pinb2-7A locus containing three tandemly duplicated copies, Pinb2-7A1, Pinb2-7A2, and Pinb2-7A3. We propose the new nomenclature of Pinb-2v for Pinb2, so as to be consistent with the IWGSC RefSeq v1.0 annotation (Table S1) [36]. We establish a series of new PCR-based genotyping primers for Pinb2 genes. Genome-wide analysis of Pinb2 genes and applications of the Pinb2 genotyping method provide new insights into this gene family.
2. Results
2.1. Genomic and Phylogenetic Analyses of Pinb2 Genes
The DNA and predicted protein sequences of known Pinb2 variants were used to identify gene models that could be Pinb-like genes, annotated in the IWGSC bread wheat genome RefSeq v1.0. Five gene models were identified, two on the B and D genomes (TraesCS7B02G431200 and TraesCS7D02G504800, respectively), with the other three on the A genome in a tandemly manner (TraesCS7A02G514400, TraesCS7A02G514500 and TraesCS7A02G514505). Sequence alignments confirmed their high sequence similarity with the kernel hardness-related genes Pinb and Pina, as well as the gene encoding GSP-1 on the Ha locus (Figures S1 and S2).
We further expanded our BLAST search to several recently published reference genomes of the Triticeae species and identified the Pinb2 homolog, AET0Gv20021600, in the Ae. tauschii genome of accession AL8/78; TuG1812G0700005557.01 in the T. urartu genome of accession G1812; and TRIDC7BG068420.1 in the B genome of wild emmer wheat accession Zavitan; as well as a tandem cluster of three Pinb2 homologous sequences in the A genome of wild emmer wheat, which lack annotations as gene models. The nomenclature of Pinb2 alleles and the corresponding variants, reported previously, are shown in Table S1 with the suffixes 7A, 7B, and 7D used to indicate the chromosomal locations. The gene models corresponding to Pinb2 genes from the Triticeae species are listed in Table S2.
Due to the high sequence similarity between Pinb and Pinb2, it has long been considered that Pinb2 might be associated with kernel hardness. Therefore, we sought to address its relationship with Pinb by using synteny alignment. The gene order at the Ha locus in the IWGSC RefSeq v1.0 assembly of the bread wheat genome is consistent with previous analyses of the Bacterial Artificial Chromosome (BAC) library containing the Ha locus. The BGGP gene, encoding b-1-3-galactosyl-O-glycosyl-glycoprotein, and the GSP-1 gene, encoding Grain Softness Protein-1, are located at the 5′ end of Ha (these genes indicated as ‘BG’ and ‘Gs’, respectively, in Figure 1, Table S3) [45,46,47].

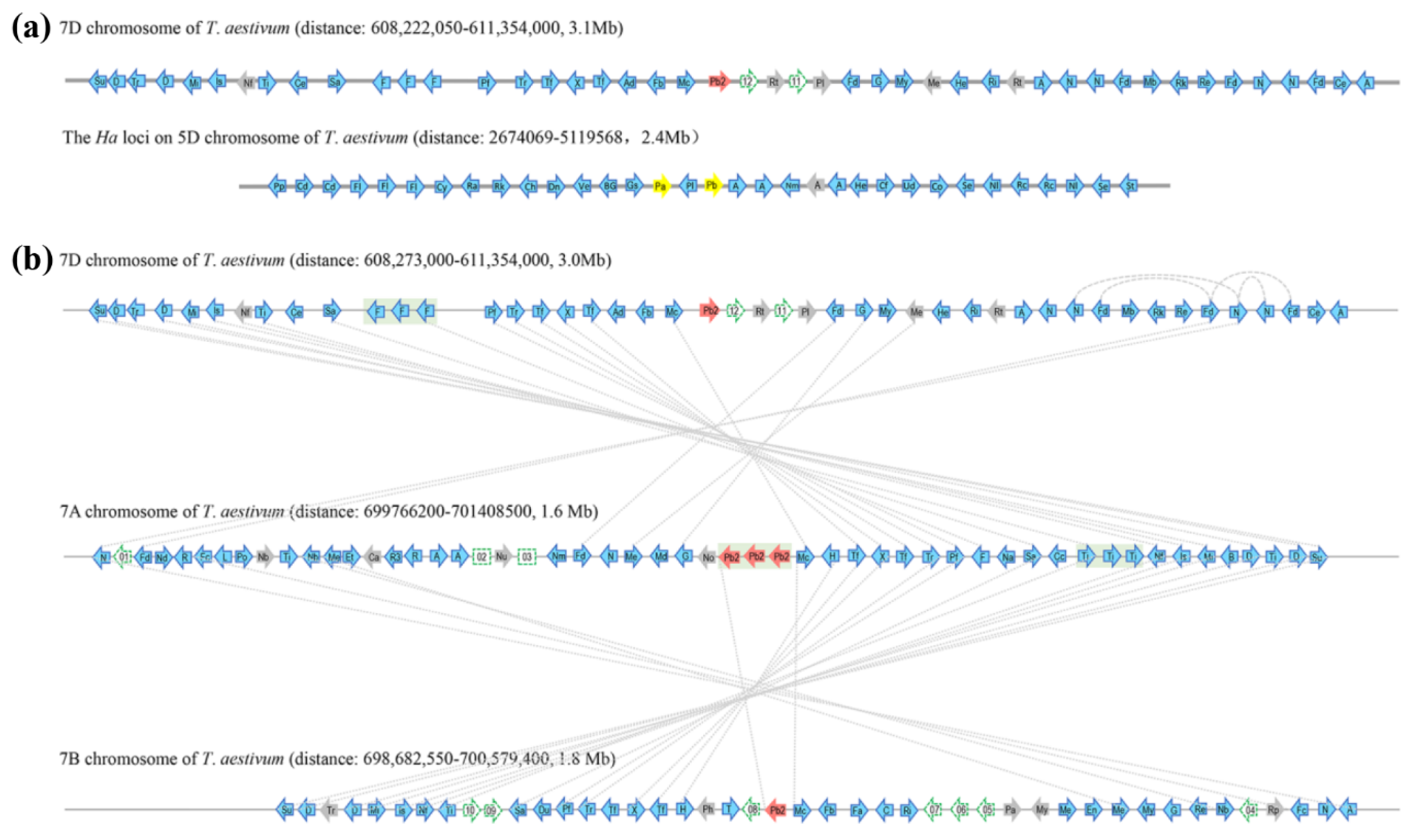
Figure 1.
Syntenic alignments of bread wheat chromosomal segments that contains Pinb2-7D locus at 7DL and Ha locus at 5DS (a), and those that contains Pinb2-7D, Pinb2-7A, Pinb2-B at 7DL, 7AL and 7BL, respectively (b). Wheat chromosomes are represented as grey horizontal lines, whereas genes along the chromosome are represented as arrowheads. Pina-D1 and Pinb-D1 are represented as yellow arrowheads. Pinb2 are represented as red arrowheads, while the other protein-coding genes with high confidence annotations in the IWGSC RefSeq v1.0 bread wheat genome are represented as blue arrowheads. Homoeologous gene pairs are indicated as dotted lines connecting arrowheads. Tandem duplication of the three Pinb2-7A genes is shaded in the grey box. To simplify visualization, only high-confidence protein-coding genes are shown with their orders and orientations along the chromosomal segments, shown as the same in wheat genome assembly. The intergenic regions are not in proportion to the wheat genome assembly. For visualization, only abbreviations of the genes are labeled on the arrowheads, with their full names provided in Table S3.
Two clusters of ATPase genes with a Nodulin gene in between are located at the 3′ downstream of Pina and Pinb (these genes are indicated as ‘A’ and ‘Nm’, respectively, in Figure 1). Comparison of the genes flanking Pina/Pinb and Pinb2 failed to identify clear synteny between the Ha and the Pinb2-7D locus (Figure 1, Table S3). We then analyzed the homologous regions containing the Pinb2 genes at 7AL, 7BL, and 7DL. Alignment of the Pinb2 flanking genes revealed a clear collinearity between the segments containing Pinb2-7A, Pinb2-7B, and Pinb2-7D (Figure 1, Table S3). Interestingly, three copies of Pinb2-7A with their flanking sequences were arranged in a cluster, suggesting their origin from tandem duplication.
Next, we investigated the collinearity of Pinb2-containing segments among the sequenced Triticeae species. A comparison of the orthologous segments on chromosome 7A showed collinearity at this region between bread wheat, wild emmer wheat, and T. urartu, while only bread and wild emmer wheats have three copies of Pinb2-7A (Figure S3a, Table S3). Comparison of the segments on chromosome 7B, between bread and wild emmer wheats, showed a good collinearity between the regions (Figure S3b, Table S3). Furthermore, TaPinb2-7D1 is contained within a segment that is collinear with an orthologous region of Ae. tauschii containing AtPinb2-7D1 (Figure S3c, Table S3). The Pinb2-7A cluster is only detected in polyploidy wheat species, but not in the A-genome and D-genome donor species, T.urartu and Ae. tauschii, suggesting that the tandem duplication of Pinb2-7A could emerge after the polyploidization of wheat.
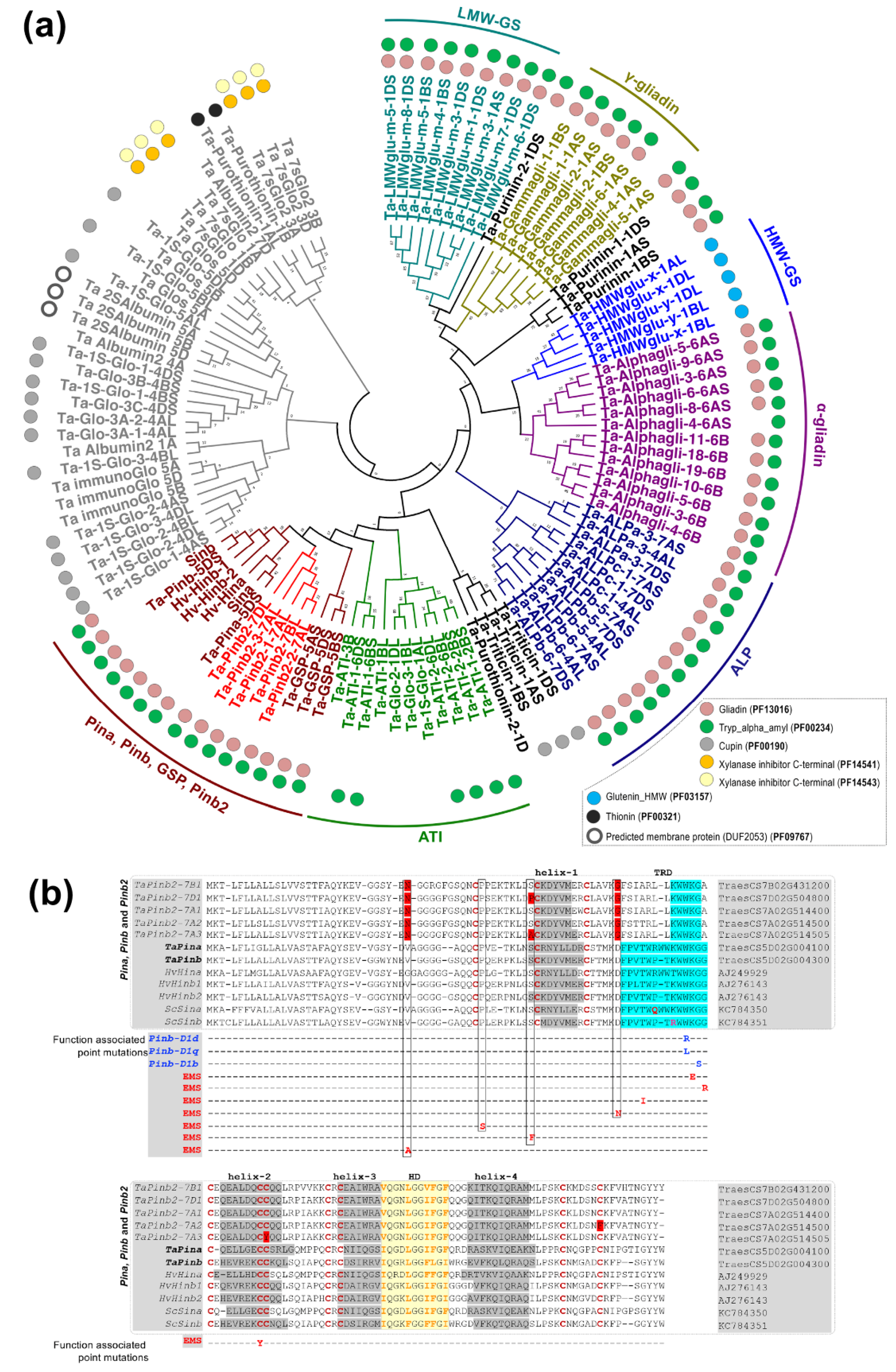
While there is a lack of synteny between the genomic segments harboring Pinb2 and the Pina/Pinb genes, the Pinb2 genes indeed have a high sequence similarity with Pinb, indicating that Pinb2 might have the sequence basis for kernel hardness or interactions with other seed proteins. Therefore, we analyzed its phylogeny together with other seed proteins, many of which belong to the prolamin superfamily [48]. The phylogenetic results separated wheat globulins and albumins from those in the prolamin superfamily (Figure 2a, Table S4). PINB2 proteins were clustered together with PINA, PINB and GSP. The puroindoline cluster was grouped together with a cluster largely consisting of α-amylase inhibitors (ATIs). As the repetitive sequences exist in many proteins of the prolamin superfamily and could be problematic for sequence alignment or annotation, we therefore used the domain signatures and conserved cysteine residue patterns to help define the phylogenetic clusters of wheat seed proteins. The results showed that almost all the phylogenetically analyzed prolamin proteins, including PINB2, have Gliadin and Tryp_alpha_amyl domains (PF13016 and PF00234, respectively) except that the α-amylase inhibitors only contain domain PF00234 and high-molecular-weight glutenin subunits (HMW-GSs) contain the HMW domain (PF03157). In addition, the conserved cysteine-rich patterns and number of cysteine residuals can be used as characteristics for defining groups of storage proteins within the prolamin superfamily [43]. For example, HMW-glutenin x- and y-type subunits contain four to six cysteines, while α-gliadin and γ-gliadin have six to eight and eight to eleven cysteines, respectively (Figure S4). Type a-, b- and c- avenin-like proteins (ALPs) have 14, 18–20, and 10–12 cysteine residues, respectively, and some ALPs have effects on dough quality, likely due to the number of cysteine residues for interaction with wheat storage proteins (Figure S4) [49,50,51,52]. Here, PINB2 identified in the bread wheat genome of CS shares the same cysteine-rich backbone consisting of 10 residues as PINA, PINB, and GSP (Figure S4).
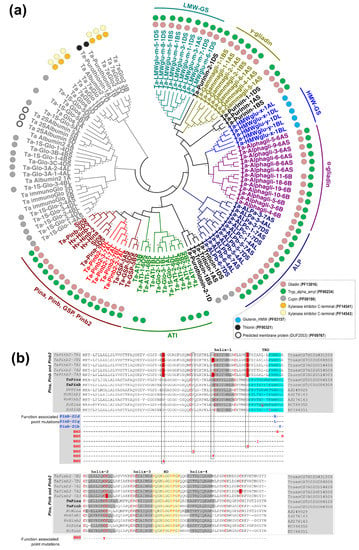
Figure 2.
Analyses of phylogeny and protein sequence alignment for PINB2, together with other wheat seed proteins. (a) Maximum likelihood (ML) phylogeny of HMW-GS, LMW-GS, α-gliadin, γ-gliadin, purinin, purothionin, avenin-like proteins (ALPs), PINA, PINB and GSP from the bread wheat CS and the homologous proteins of PINA, PINB and GSP from triticale and barley. Results for protein domain analysis using profile hidden Markov Models (HMMER) are shown as color-coded circles. (b) Alignment of the amino acid sequences between PINA, PINB, GSP, PINB2, from wheat, barley and triticale, highlights that PINB2 proteins share the cysteine residue backbone, five α-helixes and hydrophobic domain with PINs but lack TRD domain. PINB2 proteins also contain some changes in the conserved amino acids that are important for PINs’ function in kernel hardness (those amino acids shaded in red for PINB2 proteins). α-helixes, hydrophobic domain and TRD are shaded in grey, yellow and blue, respectively. The cysteine residues are shown in red.
We performed a detailed sequence comparison between Pinb2 and Pina, Pinb, puroindoline-like genes in barley and hexaploid triticale (Hordoindolines, Hin, and Secaloindolines, Sin), as well as those Pinb alleles that naturally exist or are mutagenized by ethylmethanesulfonate (EMS) with functional changes (Figure 2b) [53,54,55,56,57,58]. The puroindoline-like genes in barley include Hina, Hinb1 and Hinb2 with Hinbs, but not Hina associated with grain hardness, while the puroindoline-like genes in hexaploid triticale (AABBRR), Sina and Sinb, don’t affect grain hardness [53,56,57,58]. The point mutations identified in these natural and EMS-mutagenized alleles of Pinb have been reported to be associated with kernel hardness and were summarized in our previous study [4]. The results showed that PINB2s encoded by all three homoeologous genes had the four helixes and a hydrophobic domain (HD), as with PINA/PINB and their homologs in barley and triticale, but lacked a functional tryptophan-rich domain (TRD), which is key for interactions with polar lipids on starch granule surfaces [59]. Additionally, PINB2s had distinct amino acid residues at several conserved locations that were proved important for PINB’s function in kernel hardness by genetic analyses [4,54,55]. Despite these important sequence differences compared to PINs, PINB2s have the conserved HD domain which could be involved in protein–protein interactions in Puroindolines (Figure 2b) [59,60]. In summary, analyses of PINB2s’s sequence signatures (domains, cysteine residue patterns, etc.) and its phylogeny emphasize the sequence and structural similarities between PINB2 and PINA/PINB but also point out the sequence distinction, suggesting one of the probable reasons for PINB2 not being associated with obvious kernel hardness differences.
2.2. Expression Levels of Pinb2
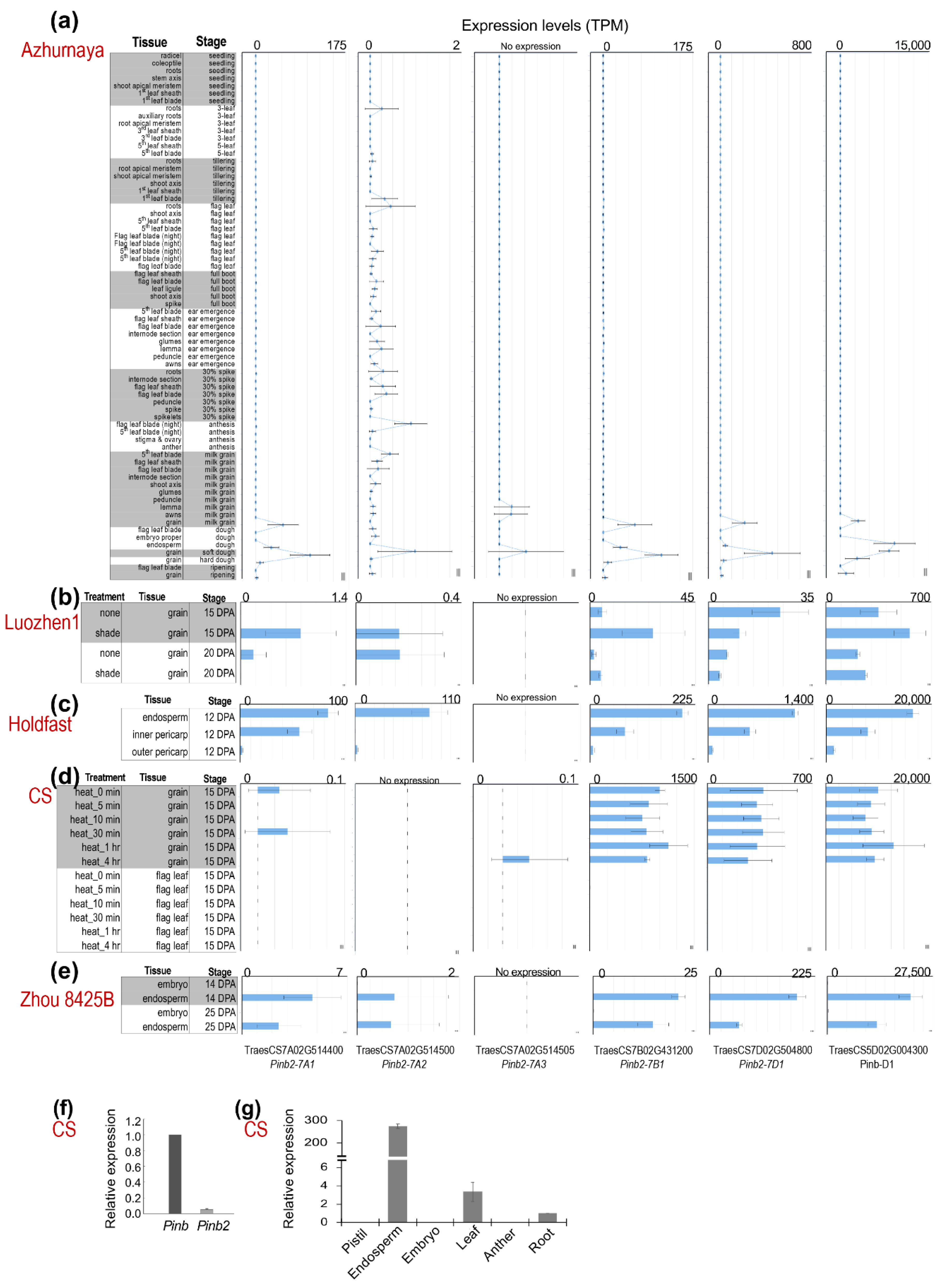
High expression levels of Pina and Pinb during seed development are important for their functions in kernel hardness. We analyzed the Pinb2 expression dynamics in wheat using the many publicly available wheat RNA-seq datasets. The RNA-seq datasets from five different wheat varieties showed that Pinb2 genes were expressed at much lower levels compared to Pinb (Figure 3; RNA-seq datasets, listed in Table S5 and briefly described in Materials and Methods). For example, Pinb was expressed at ~20,000 transcripts per million (TPM) in CS, while Pinb2-7B1 and Pinb2-7D1 were expressed at less than 1500 TPM (Figure 3d). Here, to visualize the expression levels of Pinb2 genes across different RNA-seq experiments or libraries, TPM was used as the expression measure, since it has been suggested to be more consistent and accurate in reflecting the quantitative differences of genes across wheat RNA-seq samples with the consideration that Pinb2 genes are comprised of a single-exon and short [61,62]. The expression levels of Pinb2-7A1, -7A2, -7A3, -7B1 and -7D1 differed from each other, with Pinb2-7D1 being the one with a relatively higher expression, followed by Pinb2-7B1. Pinb2-7A3 was expressed at a low level, while Pinb2-7A1 and Pinb2-7A2 were expressed at extremely low levels around the RNA-seq expression threshold (>=0.5 TPM), except for their expression in cv. Holdfast (Figure 3c). Previous RNA-seq data of the wheat variety Azhurnaya covers a wide range of tissues and developmental stages, representing a comprehensive wheat expression atlas [62]. Azhurnaya RNA-seq data showed that Pinb2 genes were highly expressed in several samples of developing seeds, particularly during soft and hard dough stages, with Pinb2-7D1 most highly expressed followed by Pinb2-7B1 and Pinb2-7A1 (Figure 3a). The results also exhibited a very low expression of Pinb2-7A2 in leaf and root tissues. These RNA-seq data highlights that the expression levels of Pinb2 genes vary between loci (Figure 3). To explore the Pinb2 expression across wheat varieties, we visualized the Pinb2 expression levels together with a reference gene, TaActin (TraesCS1D02G274400; Figure S5) [63]. The reference gene TaActin showed relative stable expression for the seed samples within each variety except for cv. Holdfast (Figure S5a). To take into account for potential RNA-seq batch effects between varieties, the TPM expression values of Pinb2 genes were normalized to TaActin. The results showed that Pinb2-7D1 was highly expressed in Holdfast, followed by that in CS, then Azhuranaya and Zhou 8425B, while Pinb2-7B1 was highly expressed in CS, followed by that in Holdfast, then Azhurnaya and Zhou 8425B (Figure S5b). Both TPM expression and relative expression data suggest that the expression levels of Pinb2 genes vary between varieties. Such varied expression levels of Pinb2 indicates potential regulatory differences between the loci and varieties, consistent with the much lower sequence similarities in the promoter regions than in the ORFs (Figure S1). Quantitative RT-PCR results validated the lower expression levels of Pinb2 in comparison with Pinb (Figure 3f, Table S6). Our qRT-PCR results also confirmed that Pinb2 genes were specifically expressed in the developing endosperm of CS (Figure 3g) [35].
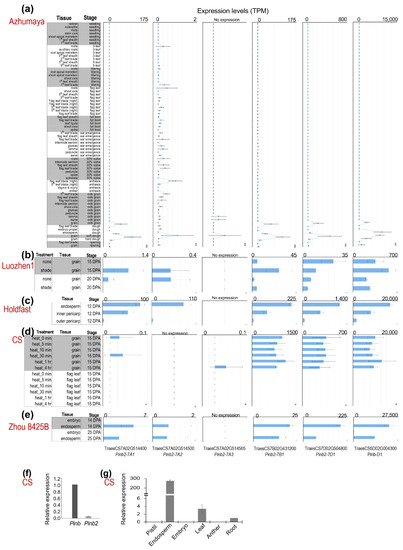
Figure 3.
Expression patterns of Pinb2 genes. Expression levels of Pinb2 genes and Pinb-D1 from five publicly available RNA-seq datasets and shown in bar charts (a–e). The varieties used for RNA-seq analyses are indicated in red, while the tissues, developmental stages and conditions for each RNA-seq experiments are labeled on the left of each panel. Expression levels are shown using TPM and are not normalized across datasets. (f) The relative expression levels of Pinb and Pinb2 genes were quantified using qRT-PCR at 28 Days Post Anthesis (DPA) in CS seeds. (g) Expression levels of Pinb2 genes in different tissues from CS proved Pinb2 genes are expressed specific in seeds. Detailed information of the RNA-seq datasets used are given in Table S5.
2.3. Genotyping of Pinb2
PCR methods to amplify the sequences of Pinb-like variants in wheat have been established, and these contribute to our understanding of Pinb-like variants [26,27,28,29]. Particularly, primers of PCR or derived cleaved amplified polymorphic sequence (dCAPS) assays detecting certain Pinb2-7B1 or Pinb2-7D1 variants were reported and have been widely used [32,33,34]. The use of Pinb2 PCR markers has significantly helped to identify new variants, to study genetic diversity and to facilitate association analysis of Pinb2 with seed traits, including kernel hardness.
The previous Pinb2 PCR primers target within the ORF regions, where the sequence variation is much lower compared to the flanking regions, where very few SNPs can be used to distinguish certain variants (Figure S1). With the high-quality reference genomes of several Triticeae species, we compared the sequences of Pinb2 homoeologous genes and designed several new primer pairs for amplifying and detecting Pinb2. These PCR primers target Pinb2-flanking regions to ensure PCR specificity (Table S7).
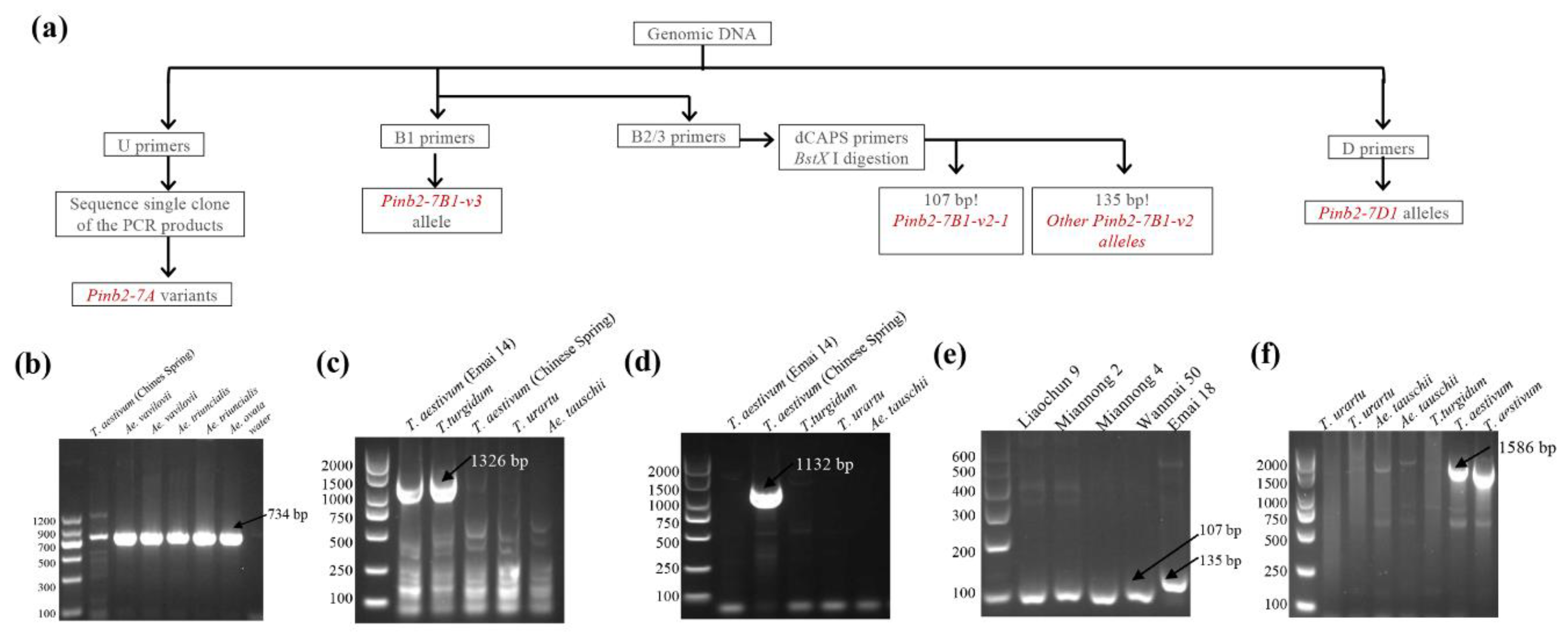
Among the newly designed Pinb2 primers, primer pair D can be used to amplify Pinb2-7D1, and primer pair U can be used to amplify all three homoeologous Pinb2 genes (Figure 4f). Primer pair B1 is designed to specifically amplify the Pinb2-7B1-v3 allele as a ~1326 bp PCR product (Figure 4c). Nested primer pairs B2 and B3 can be used to detect Pinb2-7B1-v2 alleles [64]. Further, a pair of dCAPS primers (primer pair C) was designed specifically for the Pinb2-7B1-v2-1 allele. BstX I digestion, after amplification by primer pair C, distinguishes the Pinb2-7B1-v2-1 allele (107-bp product) from the Pinb2-7B1-v2 allele (135-bp product), which only has an SNP difference (Figure 4e). Therefore, primer pairs B1, B2/B3 and the dCAPS primers form a sequential pipeline for detecting the alleles of Pinb2-7B1.
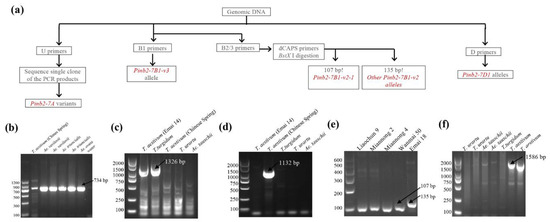
Figure 4.
Genotyping of Pinb2 genes using the new PCR method. (a) Diagram describing the workflow of set of PCR primers to genotype Pinb2-7A, Pinb2-7B and Pinb2-7D loci. (b) Primer pair U non-specifically amplified Pinb2 genes encoded by Pinb2-7A, Pinb2-7B and Pinb2-7D loci from several Triticeae species. Sizes of DNA markers used are labeled. Specific PCR products are pointed out by black arrowheads. (c) PCR primer pair B1 specifically detected the Pinb2-7B1-v3 allele in bread and durum wheats (T. aestivum and T. turgidum spp. durum). (d) Nested PCR primer sets B2/B3 specifically detected the Pinb2-7B1-v2 allele in bread wheat (T. aestivum) Chinese Spring. (e) The dCAPS primers-derived PCR amplification and subsequent BstX I digestion distinguished the Pinb2-7B1-v2-1 allele (107 bp) from the other Pinb2-7B1-v2 alleles (135 bp). (f) PCR primer pair D specifically amplified the Pinb2-7D1 gene in bread wheat.
To validate the primers’ specificity, several accessions of bread wheat (T. aestivum), durum wheat (T. turgidum spp. durum), Ae. tauschii and T. urartu were used for PCR. Indeed, primer pair B1 amplified the Pinb2-7B1 genes from bread wheat Emai 14 and durum wheat Ofanto but did not detect the Pinb2-7B1-v2 allele from CS (Figure 4b,c). The sequencing results of purified PCR products showed that the Pinb-2v on chromosome 7B of Emai14 and T. durum are Pinb-B2v3b and Pinb-B2v3a, respectively. The dCAPS primer pair followed by enzyme digestion specifically distinguished the Pinb2-7B1-v2-1 allele (with a 107-bp product) from other Pinb2-7B1 alleles (with a 135-bp product, Figure 4e). Additionally, primer pair D only detected the Pinb2-7D1 genes in bread wheat as designed (Figure 4f), while primer pair U had amplification for all the accessions used (Figure 4b). Overall, these PCR validation results demonstrated the feasibility of our Pinb2 genotyping pipeline, as illustrated in Figure 4a.
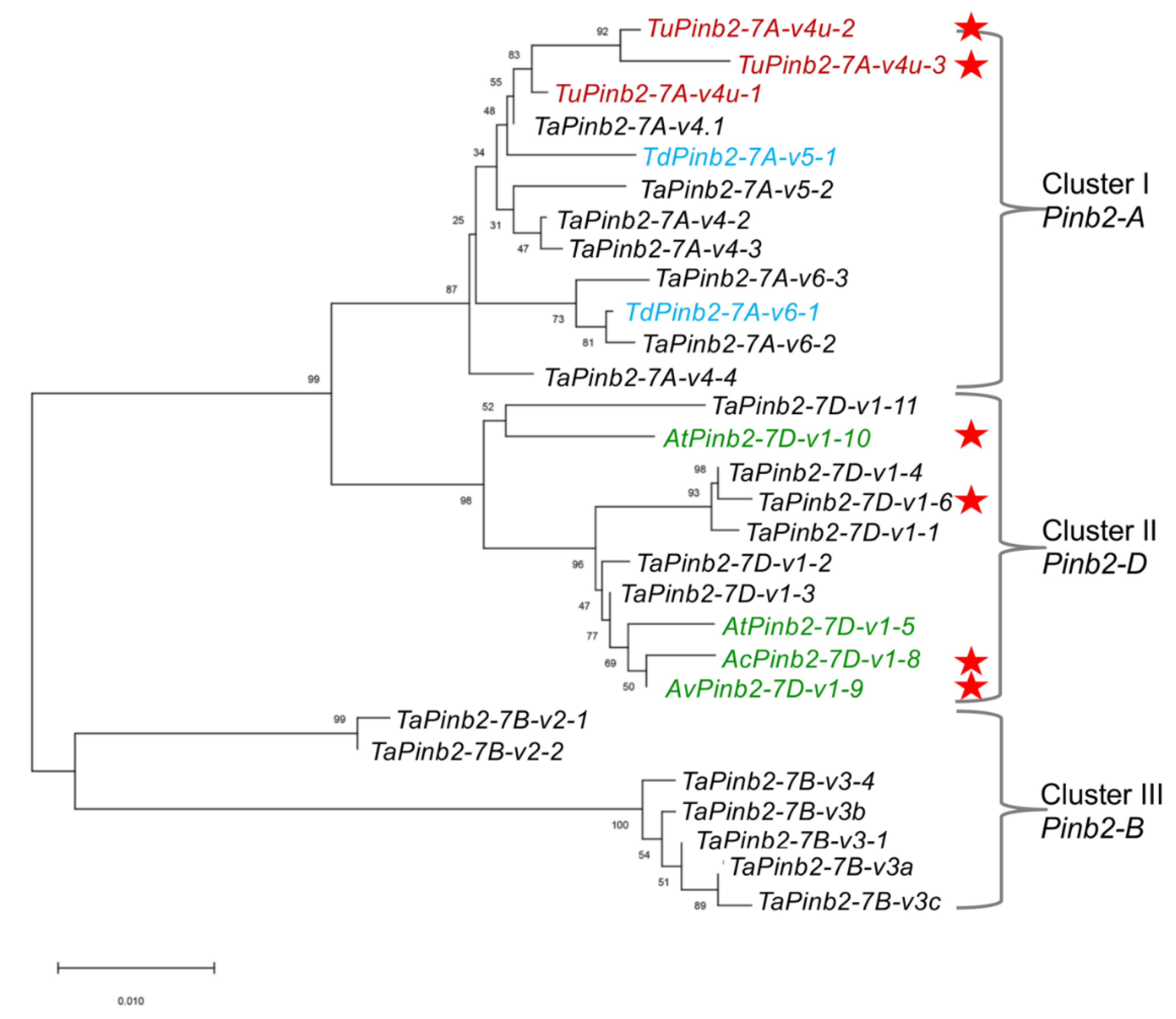
We then sought to show the usefulness of our Pinb2 genotyping primers with two examples. First, we explored the diversity of Pinb2 genes using 70 selected bread wheat accessions and several accessions of Ae. tauschii, Aegilops vavilovii, Aegilops triuncialis, Aegilops ovata and T. urartu. A new Pinb2-7D1 allele was amplified from the bread wheat varieties Zhengmai 101, Wanke 06229, Jimai 107, and Laoqimai, designated as Pinb2-7D1-v1-6. Four new alleles of Pinb2-7D1 were obtained from Ae. cylindrical, Ae. vavilovii, Ae. triuncialis and Ae. geniculate, designated as Pinb2-7D1-v1-8, Pinb2-7D1-v1-9, Pinb2-7D1-v1-10 and Pinb2-7D1-v1-11, respectively (Figure 4b). We sequenced the five new Pinb2 alleles, Pinb2-7D1-v1-6, Pinb2-7D1-v1-8, Pinb2-7D1-v1-9, Pinb2-7D1-v1-10, and Pinb2-7D1-v1-11. Sequence alignment and phylogenetic analysis of the new Pinb2 alleles with the previously identified Pinb2 sequences showed that the Pinb2 sequences are clustered according to the A, B and D genomes (Figure 5 and Figure S6).
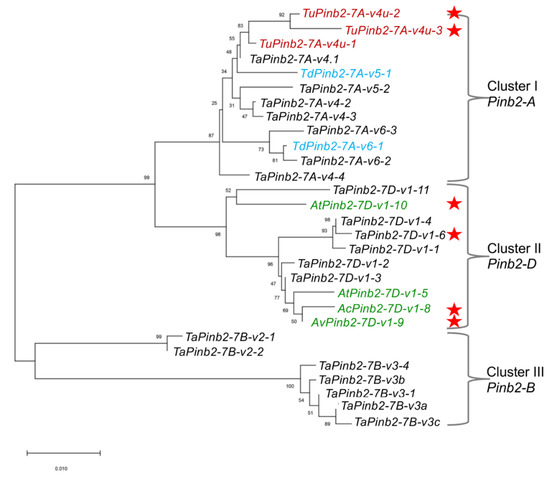
Figure 5.
Phylogenetic tree of Pinb2 variants highlights that these sequences are clustered according to the loci where they are located. Phylogeny of Pinb2 alleles was constructed using the neighbor-joining method with 1000-time bootstrap and the substitution model of Kimura 2. Different Pinb2 alleles from different Triticeae species in this study are labeled in different colors. The Pinb2 alleles from durum wheats were labeled in blue color, bread wheat in black, T. urartu in red, Ae. tauschii in green. The new alleles identified in this study are indicated using red stars.
2.4. Association of Pina, Pinb and Pinb2 with Wheat Kernel Hardness
As another application of our Pinb2 genotyping primers, we genotyped Pinb2-7B1 and Pinb2-7D1 loci for 70 Chinese bread wheat varieties, of which the kernel hardness phenotypes and Pina/Pinb genotypes were reported previously [4]. The 70 varieties were selected based on their highly correlated kernel hardness phenotypes between two field seasons (2016–2017 and 2017–2018) and their uniform Pina-D1a genotype, so as to simplify the genotype–phenotype association analysis. The genotypes of Pinb2-7B1 and Pinb2-7D1 for these 70 varieties are given in Table S8, with the Pin and Pinb2 frequencies shown in Table 1.

Table 1.
The distribution and frequency of Pina, Pinb and Pinb2 for the 70 Chinese wheat varieties.
For the Pinb2-7B1 locus, four alleles (Pinb2-7B1-v2-1, Pinb2-7B1-v3a, Pinb2-7B1-v3b and Pinb2-7B1-v3c) were detected and Pinb2-7B1-v3b was the most prevalent allele (50%) followed by Pinb2-7B1-v2-1. For the Pinb2-7D1 locus, three alleles (Pinb2-7D1-v1-3, Pinb2-7D1-v1-4, and Pinb2-7D1-v1-6) were identified, with Pinb2-7D1-v1-4 being the most predominant allele. While all of the 70 varieties uniformly contained the Pina-D1a allele, these varieties have three Pinb alleles, Pinb-D1a, Pinb-D1b, and Pinb-D1p, of which the latter two are causal for the hard-kernel phenotype due to a point mutation and a lack of PINB protein, respectively (summarized by Li et al.) [4].
Among the 70 varieties, six had a medium-hard kernel (hardness index (HI) between 40 and 60) in one season but were detected as hard kernel (HI > 60) in the other season. These varieties were then classified as mixed kernel-hardness varieties (likely containing multiple Pin-D1 haplotypes) [65,66,67] and excluded from further analysis. A Chi-square independence test showed that Pinb-D1 was significantly associated with kernel hardness in both seasons (P = 3.5 × 10−11), and Pinb2-7B1 was not correlated to kernel hardness (Table 2). Chi-square results didn’t support a dependency between Pinb2-7D1 and kernel hardness although a provisional significance was calculated (P = 0.0503). The results here highlight the usefulness of the new Pinb2 genotyping method and its contribution to association studies of Pinb2. Whether Pinb2-7D1 could play a minor role in kernel hardness, at least in some varieties under particular conditions, requires more large-scale, robust genetic analyses.

Table 2.
Chi-square analysis of the association between Pinb, Pinb2-7B1 or Pinb2-7D1 loci with kernel hardness.
3. Discussion
Pinb2 is a group of important genes with several sequence features similar to the kernel hardness-determinant gene Puroindolines, though some previous results of association and expression do not support Pinb2 as playing a major role in controlling wheat kernel hardness. Nevertheless, some genotype–phenotype analyses have indicated that Pinb2 could be associated with kernel traits [26,34]. Recently, new insights into the functions of Puroindolines have emerged, including their abilities to interact with wheat gluten proteins and lipids, possibly through hydrophobic interactions and/or disulfide bonds besides the known TRD–polarlipid interaction [68,69,70]. PINB2 maintains the same cysteine-residue backbone as PIN (Figure 2 and Figure S4). Taken together, these pieces of information justify the need to explore the sequences and function of Pinb2 in Triticeae species, as recently reported for PINs, which was previously mainly reliant on PCR-based genotyping using the limited sequence information of the Pinb2 ORFs. To facilitate Pinb2 analysis, we used the high-quality reference genomes of T. aestivum and other closely related Triticeae species to compare genomic synteny between Pin and Pinb2, phylogenetically position Pinb2 among major groups of wheat seed proteins, and develop the new PCR primers for Pinb2 genes/alleles. Similar types of studies, such as the large-scale identification of immunoresponsive allergens from wheat seed proteins and a genome-wide study of avenin-like proteins or MADS-box genes, were only made possible recently, thanks to the high-quality Triticeae genomes [48,51,71].
Using the high-quality wheat genome, we show that Pinb2 loci are not colinear with the Hardness locus on chromosome 5DS, where Pina and Pinb reside. While there is a lack of synteny between the genomic segments, PINB2 proteins are, indeed, phylogenetically clustered with PINs as well as other ATIs and share the conserved cysteine residue backbone, helixes, and hydrophobic domain with PINs, suggesting an evolutionary relationship between these proteins. Sequence analysis highlights the lack of a functional TRD region and several important amino acids in PINB2, which are required for determining kernel hardness. The contrasting results between the synteny and phylogeny and sequence analyses allow us to speculate the possibility that Pinb2 or Puroindolines might emerge in the progenitor of diploid Triticeae species, and a duplicated copy of Pinb2 might reinsert into the Ha locus at chromosome 5DS, or vice versa, followed by an independent evolution for each loci as the divergence of diploid Triticeae species and polyploidization to form Triticum aestivum. This hypothesis would explain the syntenic alignment results and the emergence of unique TRD in Puroindolines. The evolutionary aspect of Pinb2 and Pin went beyond the scope of our study and requires additional evolutionary and bioinformatics analyses using much broader genomics information, such as those from barley and rye.
Previous studies on the association of Pinb2-7B with kernel hardness drew somewhat different results and some results have indicated potential minor effects of certain Pinb2-7B alleles on kernel hardness [26,28,32,33,34]. Pinb2 has also been reported to be associated with other kernel traits in wheat [26,32]. In the present study, we designed the new Pinb2 PCR markers using the high-quality genome assemblies of bread wheat and several Triticeae species and proved the PCR markers to be useful. However, we acknowledged that the Pinb2 PCR primers might not be perfect. For example, there are a few polymorphisms in the reverse U primer (Figure S1), although it did not affect the annealing of the primers and successful amplification using the wheat materials (Figure 4). Future work will be needed to improve the Pinb2 genotyping primers toward a higher sensitivity and a broader adaptability for more wheat varieties and Triticeae species. With the new Pinb2 genotyping primers described here, we expanded the genotyping ability of certain Pinb2-7B alleles to both three homoeologous Pinb2 loci. The result of the Chi-square test didn’t support the dependency of Pinb2-7D or Pinb2-7B on kernel hardness, although the association results of Pinb2-7D approached significance (P = 0.0503). As the major purpose of the present study is to conduct a comprehensive analysis of Pinb2 genes using the high-quality wheat genome and to proof-of-concept the new Pinb2 genotyping primers, rather than to draw definite solid conclusions for the association between Pinb2 and kernel hardness, our interpretation comes with a caveat based on the possibility that the number of varieties may not be sufficient or the varieties used here may not represent broad enough genetic diversity to draw definite solid conclusions for genotype–phenotype association analysis. The marginal significance between Pinb2-7D and kernel hardness highlights the necessity of performing further genotype–phenotype association studies for the Pinb2 genes and to expand such appropriate studies to a larger and more representative collection of wheat accessions using the Pinb2 genotyping primers reported here. The wheat accessions used previously are different between studies, also making it difficult to directly compare the results. Importantly, PINB2 proteins possibly hold the structure and/or sequence basis for interacting with gluten proteins or lipids [68,69,70,72].
More recently, studies provide evidence of PIN–gluten protein aggregation, likely through hydrophobic interactions, particularly the evidence for PIN–gliadin interaction affecting gliadins’ aggregative properties [69,70]. Considering the variations in expression levels between Pinb2 homoeologous copies observed in the RNA-seq data, it may be possible that certain PINB2 proteins with high expression levels could exert similar properties as PINs, interacting with gluten proteins or lipids through hydrophobic interactions. Thus, our results emphasize future research directions to address the issue whether PINB2 proteins could interact with gluten proteins and/or lipids as PINs, and whether such interactions would exert some effects on kernel hardness or other kernel traits.
4. Materials and Methods
4.1. Plant Material
A collection of seventy bread wheat varieties from the wheat-producing regions of the Yellow and Huai Valleys and Yangtze Valley of China, provided by the Institute of Crop Breeding and Cultivation of the Hubei Agricultural Academy of Science, and the Institute of Crop Sciences, Chinese Academy of Agricultural Sciences, was used for genotyping Pinb2 genes and for interrogating their association with kernel hardness using the new genotyping primers developed in the present study (Table S8). The kernel hardness phenotypes of these varieties, collected in the 2016–2017 and 2017–2018 field seasons, have been reported previously [4]. In both seasons, they were planted in the experimental field of Huazhong University of Science and Technology with a randomized complete block design consisting of five 200 cm long rows per accession.
Several Triticum and Aegilops species were used to validate the PCR specificity and to explore the genetic diversity of Pinb2 genes; they include Aegilops tauschii (donor of the bread wheat A genome, AA), Ae. vaviolovii, Ae. triuncialis, Ae. ovata, T. urartu (donor of the bread wheat D genome, DD; accessions G1937 and G1906) and tetraploid Triticum turgidum spp. durum variety Ofanto (AABB). G1937 and G1906 were obtained from USDA-GRIN.
4.2. Phenotyping of Kernel Hardness
The kernel hardness index (HI) was determined with the Single Kernel Characterization System (SKCS) 4100 (Perten Instruments North America Inc., Springfield, IL, USA), following the American Association for Cereal Chemists (AACC) approved method as previously described [73]. Briefly, 200 kernels harvested from each replicate field plot were used for measuring HI. The wheat varieties were planted and measured in the 2016–2017 and 2017–2018 seasons, with two replicated plots for the 2016–2017 season and three replicated plots for the 2017–2018 season. These varieties were classified into soft-, medium-, and hard-kernels based on the HI value: varieties with HI less than 40 were the soft-kernel type, while those with HI greater than 60 were the hard-kernel type, with those with a HI value between 40 to 60 were the medium-kernel type.
4.3. Identification and Genotyping of Puroindoline-D1 and Puroindoline b-2 Variants
The genotyping of Pin a and b was described previously [4]. Six Pinb2 variants reported previously, Pinb-2v1, Pinb-2v2, Pinb-2v3, Pinb-2v4, Pinb-2v5, and Pinb-2v6, were used to identify the gene models encoding Pinb2 genes. Then, the coding sequences of Pinb2 gene models and predicted protein sequences were obtained from the Gramene and Triticeae Multi-omics Center Website, the latter of which has been established by Shandong Agricultural University (http://202.194.139.32/blast/viroblast.php). The coding and flanking sequences of Pinb2 genes were aligned and used to design primers specific to particular loci or variants. Specific primers were designed successfully in the light of variant- or loci-specific regions based on alignment (Figure S1, Table S7). Particularly, the conserved regions of the Pinb2 variants allowed us to design the universal primer pair U. A pair of dCAPS primers was newly designed to identify allelic variation in Pinb-B2v2-1 on chromosome 7B. Based on a special, single-nucleotide polymorphism of Pinb-B2v2-1, dCAPS Finder v2.0 was used to design dCAPS primer pair C for its use together with restriction enzyme BstX I (Fermentas, Waltham, MA, USA) [74]. Using the new primer pairs described here, a pipeline for detecting known Pinb2 genes and identifying new Pinb2 variants at all three homoeologous loci will be possible (details in Results, Figure 4).
Genomic DNA samples from three wheat seedlings of each variety were extracted with the Cetyltrimethylammonium Ammonium Bromide (CTAB) method and used for Pina/Pinb and Pinb2 genotyping [73]. PCR amplification was done in a 25 μL reaction volume containing 100 ng of genomic DNA, 0.2 μM primers, and 12.5 µL 2×Es Taq MasterMix (CWBIO, Beijing, China) using the the PCR program as follows: 95 °C for 3 min, 34 cycles of 95 °C for 30 s, 55–62 °C 30 s and 72 °C for 30 s, with a final extension of 5 min in a Bio-Rad-T100 thermal cycler. The PCR products were separated by 1.5% (w/v) agarose-gel electrophoresis and visualized under UV light after ethidium bromide staining. A sequencing service for the purified PCR products (provided by AuGCT company, Beijing, China) was used to validate the Pina/Pinb and Pinb-2v genotypes and to obtain the sequences of the newly identified alleles of Pinb2 (Table 3).

Table 3.
The information of new Pinb2 alleles identified in this study.
4.4. Synteny Analysis of the Genomic Segments Containing Pinb2 Genes
The gene order and annotation from several high-quality Triticeae species were used for syntenic analysis, including T. aestivum IWSGC RefSeq v1.0, the T. urartu genome, Ae. tauschii genome, and wild emmer wheat (T. turgidum spp. dicoccoides) genome [36,37,38,39]. High-confidence protein-coding genes flanking Pina/Pinb genes at 5DS, and those flanking Pinb2 at 7AL, 7BL and 7DL (within ~2–3 Mb), were retrieved from the genomes. The orthologous gene pairs were determined by using reciprocal BLAST (10e-5, the score is greater than 70%, and the matching base is longer than 100 bp) to compare genes between any two of the genomes. Annotations of the genes and their abbreviated names shown in Table S3.
4.5. Quantitative Real-Time Reverse Transcription PCR (qRT-PCR)
Quantitative PCR primers were designed to target the gene-specific coding regions for Pina, Pinb and Pinb2, respectively, according to the sequence alignment by DNAMAN v6.0. Actin (TraesCS4B01G050600.1) was used as the internal reference gene for qPCR. Due to the high sequence identity within the Pinb2 coding region, qRT-PCR primers for quantifying Pinb2 were only able to detect the overall expression levels of all expressed copies at the three loci (Table S6). The total RNA was extracted from different tissues of the wheat cultivar Chinse Spring [75]. The first-strand cDNA was generated from 1 μg RNA using a FastKing RT kit and gDNase (Tiangen, China) and qRT-PCR was conducted using AceQ qPCR SYBR Green Master Mix (Vazyme, Nanjing, China). The data were evaluated between the three replicates by using the relative quantification method (2−ΔΔCt).
4.6. Analyses of the Phylogeny and Protein Features for Wheat Seed Proteins
The predicted protein sequences from the bread wheat reference genome were used for phylogenetic analysis together with the predicted protein sequences of Puroindoline homologs in triticale and barley (namely, Sina and Sinb from hexaploid triticale, and Hina, Hinb-1 and Hinb-2 from barley, Hordeum vulgare) (Table S4). Domain analysis using profile hidden Markov Models (HMMER) was performed for the predicted protein sequences of Pinb2 and other seed proteins to identify several protein domain signatures, for instance the Tryp_alpha_amyl domain (PF00234), HMW-GS domain (PF03157) and gliadin domain (PF13016) [76]. Annotations of wheat seed proteins were reported previously by Juhasz et al. [43] and avenin-like proteins (ALP) were annotated by Zhang et al. [51]. The presence of conserved cysteine residual patterns was used as a feature for the subfamily assignment of the prolamin superfamily [47,48]. The phylogenetic analysis followed a detailed protocol [77]. The protein sequences were aligned using MUltiple Sequence Comparison by Log-Expectation (MUSCLE), guided by an unweighted pair-group method with arithmetic means (UPGMA) tree. The phylogenetic tree of the wheat seed proteins was constructed using the maximum likelihood method provided in the MEGA7 software with a 500-time bootstrap using a Jones–Taylor–Thornton (JTT) model [78].
4.7. Statistical Analysis
The association of Pinb and Pinb2 loci with kernel hardness phenotypes was determined by a Chi-square test of independence.
5. Conclusions
In conclusion, Pinb2 genes consist of five copies rather than three homoeologous genes due to the additional tandem duplicated copies at Pinb2-7A. Based on the synteny and phylogeny analyses, Pinb2 genes likely preserve the sequence features for interacting with gluten proteins through hydrophobic connections but lack the basis for determining kernel hardness, such as the TRD domain. These results are in line with the association analysis results: Pinb2 genes do not exert major impacts on kernel hardness as Pina/Pinb. Leveraging the high-quality reference genome of bread wheat, we develop new Pinb2 genotyping primers and demonstrate their application in identifying new Pinb2 alleles and in facilitating association studies. The present study exemplifies an application of the high-quality Triticeae genomic resources, and the results implicate the areas for further study to unveil Pinb2’s function and its potential use in genetic engineering.
Supplementary Materials
Supplementary materials can be found at https://www.mdpi.com/1422-0067/21/4/1304/s1.
Author Contributions
G.H., G.Y., X.L. and Y.L. conceived and designed the experiments; X.L., F.S. and X.Y. performed the experiments; X.L., F.S., X.Y. and Y.L. analyzed the data; G.H. and G.Y. contributed reagents/materials/analysis tools. X.L., Y.L. drafted the manuscript. G.Y. and G.H. finalized the manuscript. funding acquisition, G.H. and G.Y. All authors have read and agreed to the published version of the manuscript.
Funding
The work was funded by National Genetically Modified New Varieties of Major Projects of China (2016ZX08010004-004), the National Natural Science Foundation of China (No. 31771418, 31570261) and Key Project of Hubei Province (2017AHB041).
Acknowledgments
The authors thank the anonymous editors and reviewers whose constructive comments were helpful in improving the quality of this work.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
| ATI | α-amylase inhibitor |
| ALP | avenin-like protein |
| BAC | bacterial artificial chromosome |
| BGGP | β-1-3- galactosyl- O-glycosyl-glycoprotein |
| CS | Chinese Spring |
| dCAPS | derived cleaved amplified polymorphic sequence |
| GSP | grain softness protein |
| Ha | Hardness |
| HD | hydropbic domain |
| HMW-GS | High-Molecular-Weight Glutenin Subunits |
| IWGSC | International Wheat Genome Sequencing Consortium |
| LTP | Lipid transfer proteins |
| ORF | open reading frame |
| PIN | puroindoline |
| TPM | Transcripts Per Million |
| TRD | tryptophan-rich domain |
| UPGMA | unweighted pair group method using arithmetic average |
References
- Martin, J.M.; Frohberg, R.C.; Morris, C.F.; Talbert, L.E.; Giroux, M.J. Milling and bread baking traits associated with puroindoline sequence type in hard red spring wheat. Crop Sci. 2001, 41, 228–234. [Google Scholar] [CrossRef]
- Martin, J.M.; Meyer, F.D.; Morris, C.F.; Giroux, M.J. Pilot scale milling characteristics of transgenic isolines of a hard wheat over-expressing Puroindolines. Crop Sci. 2007, 47, 497–506. [Google Scholar] [CrossRef]
- Li, Y.; Mao, X.; Wang, Q.; Zhang, J.R.; Li, X.Y.; Ma, F.Y.; Sun, F.S.; Chang, J.L.; Chen, M.J.; Wang, Y.S.; et al. Overexpression of Puroindoline a gene in transgenic durum wheat (Triticum turgidum ssp. durum) leads to a medium–hard kernel texture. Mol. Breed. 2014, 33, 545–554. [Google Scholar]
- Li, X.; Li, Y.; Zhang, M.; Yu, X.; Hu, R.; Chang, J.; Yang, G.; Wang, Y.; He, G. Diversity of Puroindoline genes and their association with kernel hardness in Chinese wheat cultivars and landraces. Mol. Breed. 2019, 39, 61. [Google Scholar] [CrossRef]
- Morris, C.F. Puroindolines: The molecular genetic basis of wheat grain hardness. Plant Mol. Biol. 2002, 48, 633–647. [Google Scholar] [CrossRef]
- Bhave, M.; Morris, C.F. Molecular genetics of Puroindolines and related genes: Allelic diversity in wheat and other grasses. Plant Mol. Biol. 2008, 66, 205–219. [Google Scholar] [CrossRef]
- Bhave, M.; Morris, C.F. Molecular genetics of Puroindolines and related genes: Regulation of expression, membrane binding properties and applications. Plant Mol. Biol. 2008, 66, 221–231. [Google Scholar] [CrossRef]
- Beecher, B.; Bettge, A.; Smidansky, E.; Giroux, M. Expression of wild-type pinB sequence in transgenic wheat complements a hard phenotype. Theor. Appl. Genet. 2002, 105, 870–877. [Google Scholar] [CrossRef]
- Giroux, M.J.; Morris, C.F. A glycine to serine change in puroindoline b is associated with wheat grain hardness and low levels of starch-surface friabilin. Theor. Appl. Genet. 1997, 95, 857–864. [Google Scholar] [CrossRef]
- Hogg, A.C.; Sripo, T.; Beecher, B.; Martin, J.M.; Giroux, M.J. Wheat puroindolines interact to form friabilin and control wheat grain hardness. Theor. Appl. Genet. 2004, 108, 1089–1097. [Google Scholar] [CrossRef]
- Krishnamurthy, K.; Giroux, M.J. Expression of wheat puroindoline genes in transgenic rice enhances grain softness. Nat. Biotechnol. 2001, 19, 162–166. [Google Scholar] [CrossRef] [PubMed]
- Martin, J.M.; Meyer, F.D.; Smidansky, E.D.; Wanjugi, H.; Blechl, A.E.; Giroux, M.J. Complementation of the pina (null) allele with the wild type Pina sequence restores a soft phenotype in transgenic wheat. Theor. Appl. Genet. 2006, 113, 1563–1570. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Martin, J.M.; Beecher, B.; Morris, C.F.; Hannah, L.C.; Giroux, M.J. Seed-specific expression of the wheat puroindoline genes improves maize wet milling yields. Plant Biotechnol. J. 2009, 7, 733–743. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Li, Y.; Sun, F.; Li, X.; Wang, P.; Yang, G.; He, G. Expression of Puroindoline a in durum wheat affects milling and pasting properties. Front. Plant Sci. 2019, 10, 482. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Li, Y.; Sun, F.; Li, X.; Wang, P.; Chang, J.; Wang, Y.; Yang, G.; He, G. Co-expression of high-molecular-weight glutenin subunit 1Ax1 and Puroindoline a (Pina) genes in transgenic durum wheat (Triticum turgidum ssp. durum) improves milling and pasting quality. BMC Plant Biol. 2019, 19, 126. [Google Scholar]
- Rai, A.; Mahendru-Singh, A.; Raghunandan, K.; Kumar, T.; Sharma, P.; Ahlawat, A.; Singh, S.; Ganiewala, D.; Shukla, R.; Sivasamy, M. Marker-assisted transfer of PinaD1a gene to develop soft grain wheat cultivars. 3 Biotech. 2019, 9, 183. [Google Scholar] [CrossRef]
- Ma, X.; Xue, H.; Sun, J.; Sajjad, M.; Wang, J.; Yang, W.; Li, X.; Zhang, A.; Liu, D. Transformation of Pinb-D1x to soft wheat produces hard wheat kernel texture. J. Cereal Sci. 2020, 91, 102889. [Google Scholar] [CrossRef]
- Ali, I.; Sardar, Z.; Rasheed, A.; Mahmood, T. Molecular characterization of the puroindoline-a and b alleles in synthetic hexaploid wheats and in silico functional and structural insights into Pina-D1. J. Theor. Biol. 2015, 376, 1–7. [Google Scholar] [CrossRef]
- Chen, F.; He, Z.H.; Xia, X.C.; Xia, L.Q.; Zhang, X.Y.; Lillemo, M.; Morris, C.F. Molecular and biochemical characterization of puroindoline a and b alleles in Chinese landraces and historical cultivars. Theor. Appl. Genet. 2006, 112, 400–409. [Google Scholar] [CrossRef]
- Kumar, R.; Arora, S.; Singh, K.; Garg, M. Puroindoline allelic diversity in Indian wheat germplasm and identification of new allelic variants. Breed. Sci. 2015, 65, 319–326. [Google Scholar] [CrossRef][Green Version]
- Ma, X.; Sajjad, M.; Wang, J.; Yang, W.; Sun, J.; Li, X.; Zhang, A.; Liu, D. Diversity, distribution of Puroindoline genes and their effect on kernel hardness in a diverse panel of Chinese wheat germplasm. BMC Plant Biol. 2017, 17, 158. [Google Scholar] [CrossRef]
- Wang, J.; Sun, J.Z.; Liu, D.C.; Yang, W.L.; Wang, D.W.; Tong, Y.P.; Zhang, A.M. Analysis of Pina and Pinb alleles in the microcore collections of Chinese wheat germplasm by Ecotilling and identification of a novel Pinb allele. J. Cereal Sci. 2008, 48, 836–842. [Google Scholar] [CrossRef]
- Ayala, M.; Guzmán, C.; Alvarez, J.; Peña, R. Characterization of genetic diversity of puroindoline genes in Mexican wheat landraces. Euphytica 2012, 190, 53–63. [Google Scholar] [CrossRef]
- Klimushina, M.; Divashuk, M.; Mokhammed, T.; Semenov, O.; Karlov, G. Analysis of allelic state of genes responsible for baking properties in allocytoplasmic wheat hybrids. Russ. J. Genet. 2013, 49, 530–538. [Google Scholar] [CrossRef]
- Morris, C.; Kiszona, A.; Peden, G. Registration of extra-hard kernel near-isogenic hexaploid wheat genetic stocks lacking puroindoline genes. J. Plant Regist. 2020, 2. [Google Scholar] [CrossRef]
- Chen, F.; Zhang, F.Y.; Cheng, X.Y.; Craig, M.; Xu, H.X.; Dong, Z.D.; Zhan, K.H.; Cui, D.Q. Association of Puroindoline b-B2 variants with grain traits, yield components and flag leaf size in bread wheat (Triticum aestivum L.) varieties of the Yellow and Huai Valleys of China. J. Cereal Sci. 2010, 52, 247–253. [Google Scholar] [CrossRef]
- Wilkinson, M.; Wan, Y.; Tosi, P.; Leverington, M.; Snape, J.; Mirchell, A.C.R.; Shewry, P.R. Identification and genetic mapping of variant forms of puroindoline b expressed in developing wheat grain. J. Cereal Sci. 2008, 48, 722–728. [Google Scholar] [CrossRef]
- Chen, F.; Beecher, B.S.; Morris, C.F. Physical mapping and a new variant of Puroindoline b-2 genes in wheat. Theor. Appl. Genet. 2010, 120, 745–751. [Google Scholar] [CrossRef]
- Chen, F.; Xu, H.X.; Zhang, F.Y.; Xia, X.C.; He, Z.H.; Wang, D.W.; Dong, Z.D.; Zhan, K.H.; Cheng, X.Y.; Cui, D.Q. Physical mapping of puroindoline b-2 genes and molecular characterization of a novel variant in durum wheat (Triticum turgidum L.). Mol. Breed. 2011, 28, 153–161. [Google Scholar] [CrossRef]
- Geng, H.; Beecher, B.S.; He, Z.H.; Morris, C.F. Physical Mapping of Puroindoline b-2 Genes in Wheat using ‘Chinese Spring’ Chromosome Group 7 Deletion Lines. Crop Sci. 2012, 52, 2674–2678. [Google Scholar] [CrossRef]
- Ramalingam, A.; Palombo, E.A.; Bhave, M. The Pinb-2 genes in wheat comprise a multigene family with great sequence diversity and important variants. J. Cereal Sci. 2012, 56, 171–180. [Google Scholar] [CrossRef]
- Mohler, V.; Schmolke, M.; Paladey, E.; Seling, S.; Hartl, L. Association analysis of Puroindoline-D1 and Puroindoline b-2 loci with 13 quality traits in European winter wheat (Triticum aestivum L.). J. Cereal Sci. 2012, 56, 623–628. [Google Scholar] [CrossRef]
- Geng, H.; Beecher, B.S.; He, Z.; Kiszonas, A.M.; Morris, C.F. Prevalence of Puroindoline D1 and Puroindoline b-2 variants in U.S. Pacific Northwest wheat breeding germplasm pools, and their association with kernel texture. Theor. Appl. Genet. 2012, 124, 1259–1269. [Google Scholar] [CrossRef]
- Chen, F.; Zhang, F.; Li, H.; Morris, C.F.; Cao, Y.; Shang, X.; Cui, D. Allelic variation and distribution independence of Puroindoline b-B2 variants and their association with grain texture in wheat. Mol. Breed. 2013, 32, 399–409. [Google Scholar] [CrossRef]
- Giroux, M.J.; Kim, K.H.; Hogg, A.C.; Martin, J.M.; Beecher, B. The Puroindoline b-2 variants are expressed at low levels relative to the Puroindoline D1 genes in wheat seeds. Crop Sci. 2013, 53, 833–841. [Google Scholar] [CrossRef]
- International Wheat Genome Sequencing Consortium (IWGSC). Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 2018, 361, eaar7191. [Google Scholar] [CrossRef]
- Avni, R.; Nave, M.; Barad, O.; Baruch, K.; Twardziok, S.O.; Gundlach, H.; Hale, I.; Mascher, M.; Spannagl, M.; Wiebe, K.; et al. Wild emmer genome architecture and diversity elucidate wheat evolution and domestication. Science 2017, 357, 93–97. [Google Scholar] [CrossRef]
- Ling, H.; Ma, B.; Shi, X.; Liu, H.; Dong, L.; Sun, H.; Cao, Y.; Gao, Q.; Zheng, S.; Li, Y.; et al. Genome sequence of the progenitor of wheat A subgenome Triticum Urartu. Nature 2018, 557, 424–428. [Google Scholar] [CrossRef]
- Luo, M.C.; Gu, Y.; Puiu, D.; Wang, H.; Twardziok, S.O.; Deal, K.R.; Huo, N.; Zhu, T.; Wang, L.; Wang, Y.; et al. Genome sequence of the progenitor of the wheat D genome Aegilops tauschii. Nature 2017, 551, 498–502. [Google Scholar] [CrossRef]
- Zhao, G.; Zou, C.; Li, K.; Wang, K.; Li, T.; Gao, L.; Zhang, X.; Wang, H.; Yang, Z.; Liu, X.; et al. The Aegilops tauschii genome reveals multiple impacts of transposons. Nat. Plants 2017, 3, 946–955. [Google Scholar] [CrossRef]
- Dong, J.; Feng, Y.; Kumar, D.; Zhang, W.; Zhu, T.; Luo, M.C.; Messing, J. Analysis of tandem gene copies in maize chromosomal regions reconstructed from long sequence reads. Proc. Natl. Acad. Sci. USA 2016, 113, 7949–7956. [Google Scholar] [CrossRef]
- Li, C.; Xiang, X.; Huang, Y.; Zhou, Y.; An, D.; Dong, J.; Zhao, C.; Liu, H.; Li, Y.; Wang, Q.; et al. Long-read sequencing reveals genomic structural variations that underlie creation of quality protein maize. Nat. Commun. 2020, 11, 17. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, X.; Tang, H.; Zhang, Q.; Hua, X.; Ma, X.; Zhu, F.; Jones, T.; Zhu, X.; Bowers, J.; et al. Allele-defined genome of the autopolyploid sugarcane Saccharum spontaneum L. Nat. Genet. 2018, 50, 1565–1573. [Google Scholar] [CrossRef]
- Shi, J.; Ma, X.; Zhang, J.; Zhou, Y.; Liu, M.; Huang, L.; Sun, S.; Zhang, X.; Gao, X.; Zhan, W.; et al. Chromosome conformation capture resolved near complete genome assembly of broomcorn millet. Nat. Commun. 2019, 10, 464. [Google Scholar] [CrossRef]
- Chantret, N.; Salse, J.; Sabot, F.; Rahman, S.; Bellec, A.; Laubin, B.; Dubois, I.; Dossat, C.; Sourdile, P.; Philippe, J.; et al. Molecular basis of evolutionary events that shaped the hardness locus in diploid and polyploid wheat species (Triticum and Aegilops). Plant Cell 2005, 17, 1033–1045. [Google Scholar] [CrossRef]
- Li, W.; Huang, L.; Gill, B.S. Recurrent deletions of Puroindoline genes at the grain hardness locus in four independent lineages of polyploid wheat. Plant Physiol. 2008, 146, 200–212. [Google Scholar] [CrossRef]
- Charles, M.; Tang, H.; Belcram, H.; Paterson, A.; Gornick, P.; Chalhoub, B. Sixty million years in evolution of soft grain trait in grasses: Emergence of the softness locus in the common ancestor of pooideae and ehrhartoideae, after their divergence from panicoideae. Mol. Biol. Evol. 2009, 26, 1651–1661. [Google Scholar] [CrossRef]
- Juhasz, A.; Belova, T.; Florides, C.G.; Maulis, C.; Fischer, I.; Gell, G.; Birinyi, Z.; Ong, J.; Keeble-Gagnere, G.; Maharajan, A.; et al. Genome mapping of seed-borne allergens and immunoresponsive proteins in wheat. Sci. Adv. 2018, 4, eaar8602. [Google Scholar] [CrossRef]
- Ma, F.; Li, M.; Yu, L.; Li, Y.; Liu, Y.; Li, T.; Liu, W.; Wang, H.; Zheng, Q.; Li, K.; et al. Transformation of common wheat (Triticum aestivum L.) with avenin-like b gene improves flour mixing properties. Mol. Breed. 2013, 32, 853–865. [Google Scholar] [CrossRef]
- Ma, F.; Li, M.; Li, T.; Liu, W.; Liu, Y.; Li, Y.; Hu, W.; Zheng, Q.; Wang, Y.; He, G. Overexpression of avenin-like b proteins in bread wheat (Triticum aestivum L.) improves dough mixing properties by their incorporation into glutenin polymers. PLoS ONE 2013, 8, e66758. [Google Scholar] [CrossRef]
- Zhang, Y.; Hu, X.; Islam, S.; She, M.; Peng, Y.; Yu, Z.; Wylie, S.; Juhasz, A.; Dowla, M.; Yang, R.; et al. New insights into the evolution of wheat avenin-like proteins in wild emmer wheat (Triticum dicoccoides). Proc. Natl. Acad. Sci. USA 2018, 115, 13312–13317. [Google Scholar] [CrossRef]
- Wang, Y.; Li, M.; Guan, Y.; Li, L.; Sun, F.; Han, J.; Chang, J.; Chen, M.; Yang, G.; He, G. Effects of an additional cysteine residue of avenin-like b protein by site-directed mutagenesis on dough properties in wheat (Triticum aestivum L.). J. Agric. Food Chem. 2019, 67, 8557–8572. [Google Scholar] [CrossRef]
- Beecher, B.; Bowman, J.; Martin, J.M.; Bettge, A.D.; Morris, C.F.; Blake, T.K.; Giroux, M.J. Hordoindolines are associated with a major endosperm-texture QTL in Barley (Hordeum vulgare). Genome 2002, 45, 584–591. [Google Scholar] [CrossRef]
- Feiz, L.; Beecher, B.S.; Martin, J.M.; Giroux, M.J. In planta mutagenesis determines the functional regions of the wheat puroindoline proteins. Genetics 2009, 183, 853–860. [Google Scholar] [CrossRef][Green Version]
- Feiz, L.; Martin, J.M.; Giroux, M.J. Creation and functional analysis of new Puroindoline alleles in Triticum aestivum. Theor. Appl. Genet. 2009, 118, 247–257. [Google Scholar] [CrossRef]
- Takahashi, A.; Ikeda, T.M.; Takayama, T.; Yanagisawa, T. A barley Hordoindoline mutation resulted in an increase in grain hardness. Theor. Appl. Genet. 2010, 120, 519–526. [Google Scholar] [CrossRef]
- Gasparis, S.; Orczyk, W.; Nadolska-Orczyk, A. Sina and Sinb genes in triticale do not determine grain hardness contrary to their orthologs Pina and Pinb in wheat. BMC Plant Biol. 2013, 13, 190. [Google Scholar] [CrossRef]
- Shabrangy, A.; Roustan, V.; Reipert, S.; Weidinger, M.; Roustan, P.J.; Stoger, E.; Weckwerth, W.; Ibl, V. Using RT-qPCR, proteomics, and microscopy to unravel the spatio-temporal expression and subcellular localization of hordoindolines across development in barley endosperm. Front. Plant Sci. 2018, 9, 775. [Google Scholar] [CrossRef]
- Alfred, R.L.; Palombo, E.A.; Panozzo, J.F.; Bhave, M. The cooperative interaction of puroindolines in wheat grain texture may involve the hydrophobic domain. J. Cereal Sci. 2014, 60, 323–330. [Google Scholar] [CrossRef]
- Geneix, N.; Dalgalarrondo, M.; Bakan, B.; Rolland-Sabate, A.; Elmorjani, K.; Marion, D. A single amino acid substitution in puroindoline b impacts its self-assembly and the formation of heteromeric assemblies with puroindoline a. J. Cereal Sci. 2015, 64, 116–125. [Google Scholar] [CrossRef]
- Borrill, P.; Ramirez-Gonzalez, R.H.; Uauy, C. expVIP: A customizable RNA-seq data analysis and visualization platform. Plant Physiol. 2016, 170, 2172–2186. [Google Scholar] [CrossRef]
- Ramirez-Gonzalez, R.H.; Borrill, P.; Lang, D.; Harrington, S.A.; Brinton, J.; Venturini, L.; Davey, M.; Jacobs, J.; van Ex, F.; Pasha, A.; et al. The transcriptional landscape of polyploid wheat. Science 2018, 361, eaar6089. [Google Scholar] [CrossRef]
- Yu, X.; Wang, T.; Zhu, M.; Zhang, L.; Zhang, F.; Jing, E.; Ren, Y.; Wang, Z.; Xin, Z.; Lin, T. Transcriptome and physiological analyses for revealing genes involved in wheat response to endoplasmic reticulum stress. BMC Plant Biol. 2019, 19, 193. [Google Scholar] [CrossRef]
- Geng, H.; Beecher, B.; Pumphrey, M.; He, Z.; Morris, C.F. Segregation analysis indicates that Puroindoline b-2 variants 2 and 3 are allelic in Triticum aestivum and that a revision to Puroindoline b-2 gene symbolization is indicated. J. Cereal Sci. 2013, 57, 61–66. [Google Scholar] [CrossRef]
- Chen, F.; He, Z.H.; Xia, X.C.; Lillemo, M.; Morris, C.F. A new puroindoline b mutation presented in Chinese winter wheat cultivar Jingdong 11. J. Cereal Sci. 2005, 42, 267–269. [Google Scholar] [CrossRef]
- Lillemo, M.; Chen, F.; Xia, X.C.; William, M.; Peña, R.J.; Trethowan, R.; He, Z.H. Puroindoline grain hardness alleles in CIMMYT bread wheat germplasm. J. Cereal Sci. 2006, 44, 86–92. [Google Scholar] [CrossRef]
- Morris, C.F.; Lillemo, M.; Simeone, M.C.; Giroux, M.J.; Babb, S.L.; Kidwell, K.K. Prevalence of puroindoline grain hardness genotypes among historically significant North American spring and winter wheats. Crop Sci. 2001, 41, 218–228. [Google Scholar] [CrossRef]
- Sanders, M.R.; Clifton, L.A.; Frazier, R.A.; Green, R.J. Role of lipid composition on the interaction between a tryptophan-rich protein and model bacterial membranes. Langmuir 2016, 32, 2050–2057. [Google Scholar] [CrossRef]
- Quayson, E.T.; Marti, A.; Morris, C.F.; Marengo, M.; Bonomi, F.; Seetharaman, K.; Iametti, S. Structural consequences of the interaction of puroindolines with gluten proteins. Food Chem. 2018, 253, 255–261. [Google Scholar] [CrossRef]
- Geneix, N.; Dlgalarrondo, M.; Tassy, C.; Nadaud, I.; Barret, P.; Bakan, B.; Elmorjani, K.; Marion, D. Relationships between puroindoline-prolamin interactions and wheat grain hardness. bioRxv 2019. [Google Scholar] [CrossRef]
- Schilling, S.; Kennedy, A.; Pan, S.; Jermiin, L.S.; Melzer, R. Genome-wide analysis of MIKC-type MADS-box genes in wheat: Pervasive duplications, functional conservation and putative neofunctionalization. New Phytol. 2020, 225, 511–529. [Google Scholar] [CrossRef]
- Qin, H.; Ma, D.; Huang, X.; Zhang, J.; Sun, W.; Hou, G.; Wang, C.; Guo, T. Accumulation of glycolipids in wheat grain and their role in hardness during grain development. Crop J. 2019, 7, 19–29. [Google Scholar] [CrossRef]
- Li, Y.; Wang, Q.; Li, Y.; Xiao, X.; Sun, S.; Wang, C.; Hu, W.; Feng, J.; Chang, L.; Chen, J.; et al. Coexpression of the high molecular weight glutenin subunit 1Ax1 and puroindoline improves dough mixing properties in durum wheat (Triticum turgidum L. ssp. durum). PLoS ONE 2012, 7, e50057. [Google Scholar] [CrossRef]
- Neff, M.M.; Turk, E.; Kalishman, M. Web-based primer design for single nucleotide polymorphism analysis. Trends Genet. 2002, 18, 613–615. [Google Scholar] [CrossRef]
- Sun, F.; Liu, X.; Wei, Q.; Liu, J.; Yang, T.; Jia, L.; Wang, Y.; Yang, G.; He, G. Functional characterization of TaFUSCA3, a B3-superfamily transcription factor gene in the wheat. Front. Plant Sci. 2017, 8, 1133. [Google Scholar] [CrossRef]
- Potter, S.C.; Luciani, A.; Eddy, Y.; Park, S.R.; Lopez, R.; Finn, R.D. HMMER web server: 2018 update. Nucleic Acids Res. 2018, Web Server Issue 46. W200–W204. [Google Scholar] [CrossRef]
- Hall, B.G. Building phylogenetic trees from molecular data with MEGA. Mol. Biol. Evol. 2013, 30, 1229–1235. [Google Scholar] [CrossRef]
- Kumar, S.; Stechelr, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).