Pyramiding Bacterial Blight Resistance Genes in Tainung82 for Broad-Spectrum Resistance Using Marker-Assisted Selection
Abstract
1. Introduction
2. Results
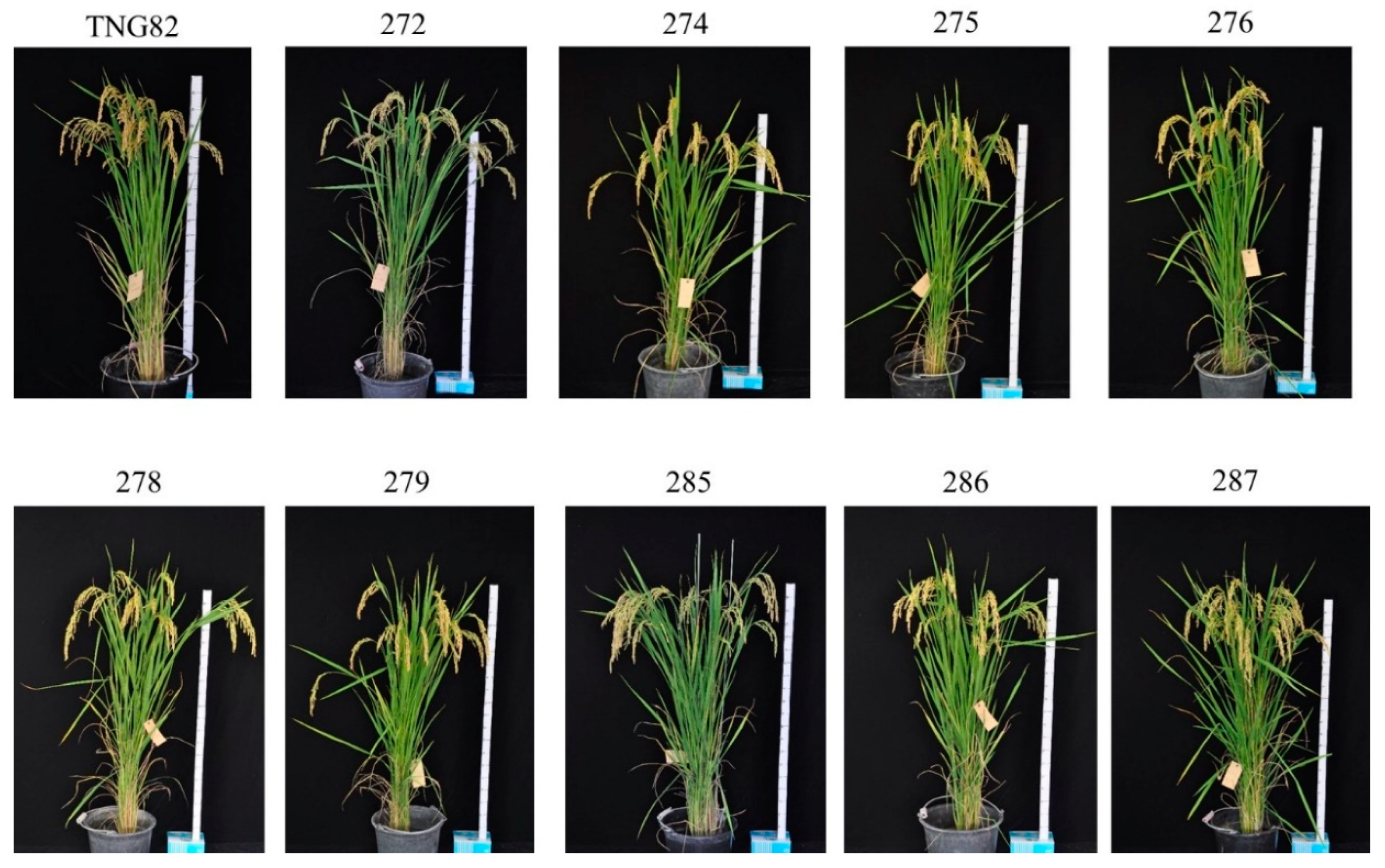
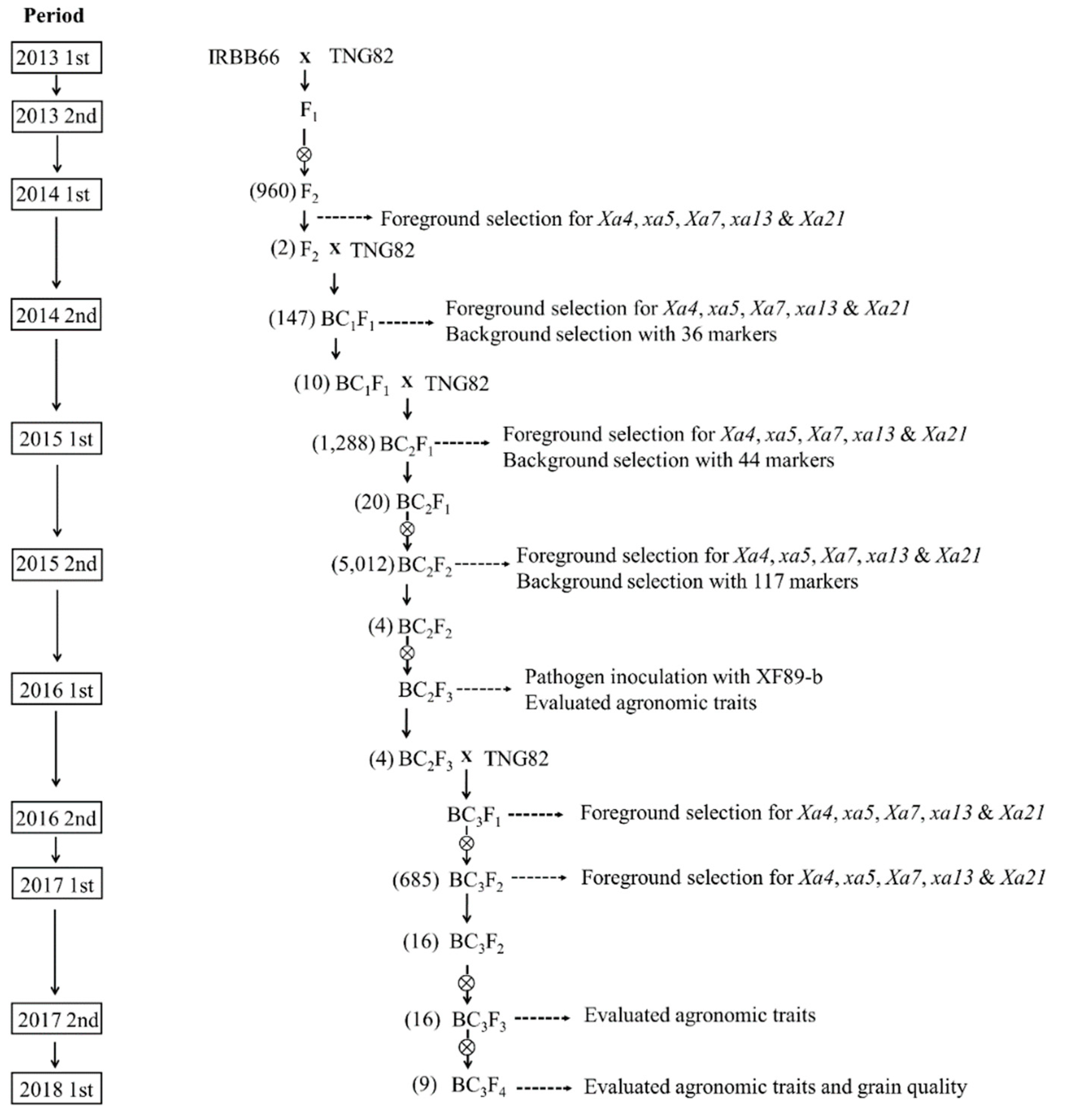
2.1. Development of BC3F4 Pyramided Lines Using Marker-Assisted Breeding
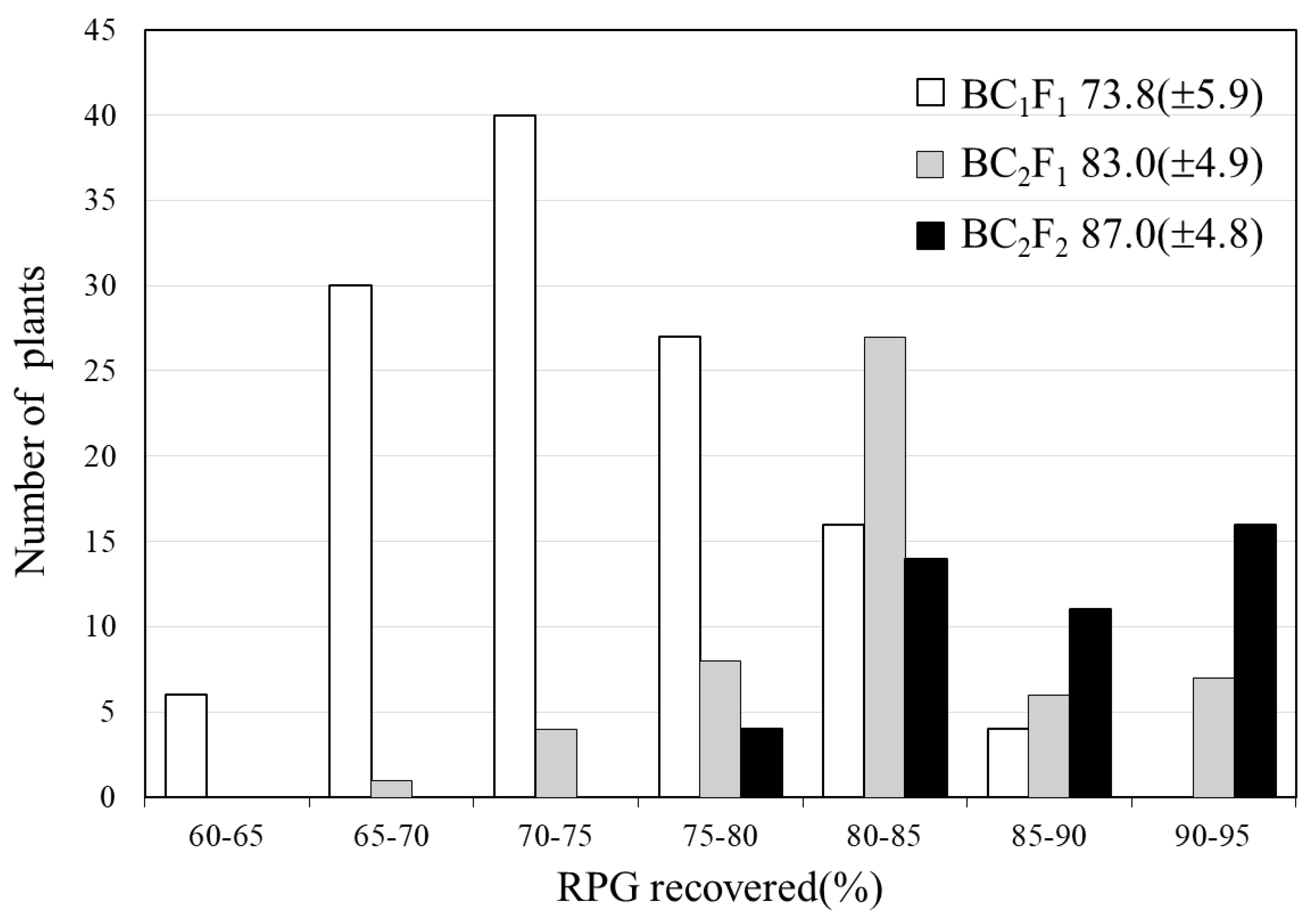
2.2. Development of BC3F4 Pyramided Lines Using Marker-Assisted Breeding
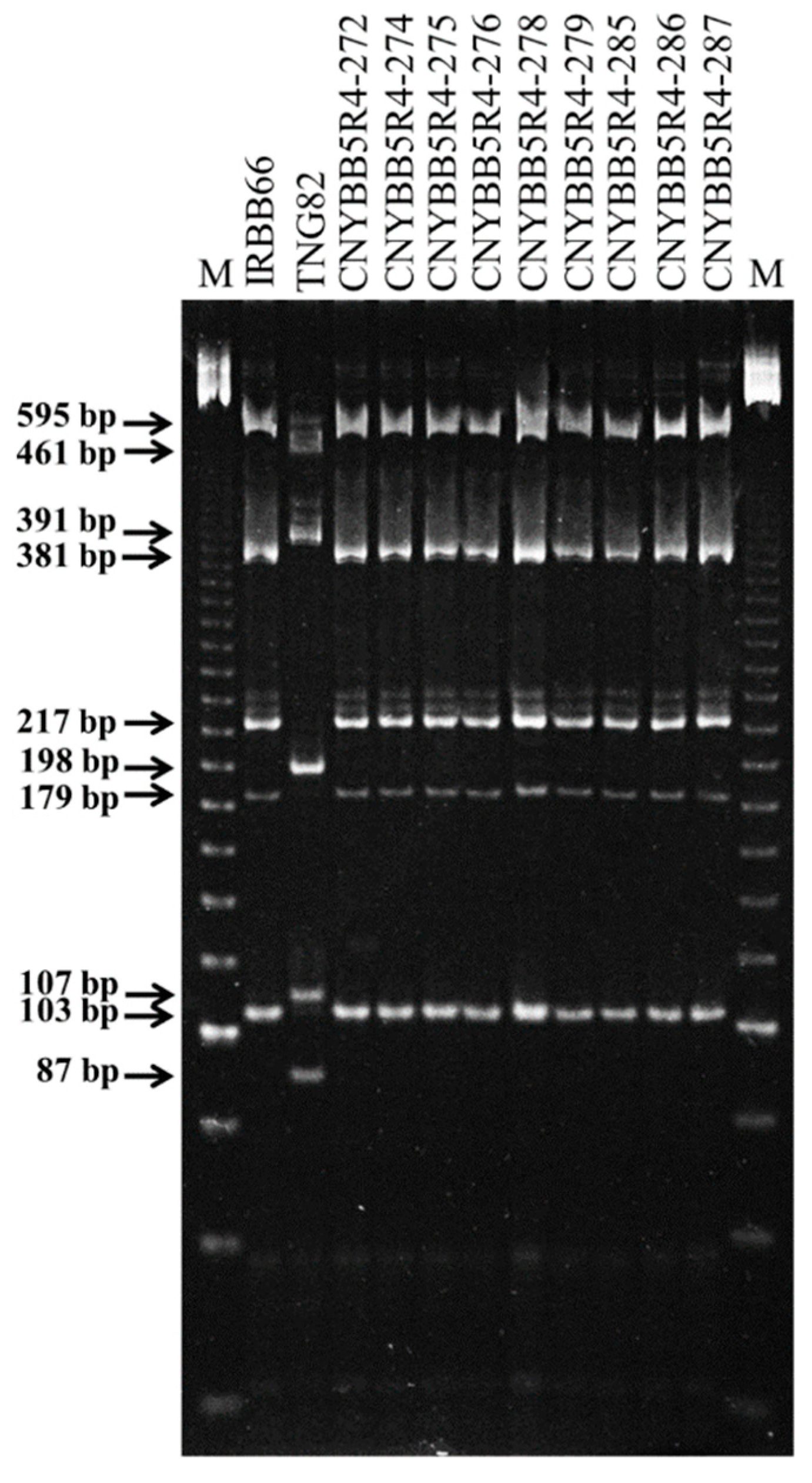
2.3. Development of BC3F4 Pyramided Lines Using Marker-Assisted Breeding
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Evaluation of Bacterial Blight Resistance
4.3. Evaluation of Agronomic Traits
4.4. DNA Isolation and PCR Amplification
4.5. Marker Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- FAO. Global Agriculture Towards 2050; FAO: Rome, Italy, 2009. [Google Scholar]
- Ray, D.K.; Mueller, N.D.; West, P.C.; Foley, J.A. Yield trends are insufficient to double global crop production by 2050. PLoS ONE 2013, 8, e66428. [Google Scholar] [CrossRef] [PubMed]
- Khush, G.S.; Mackill, D.J.; Sidhu, G.S. Breeding rice for resistance to bacterial leaf blight. In IRRI Bacterial Blight of Rice; IRRI: Manila, Philippines, 1989; pp. 207–217. [Google Scholar]
- Singh, G.P.; Singh, S.R.; Singh, R.V.; Singh, R.M. Variation and qualitative losses caused by bacterial blight in different rice varieties. Indian Phytopathol. 1997, 30, 180–185. [Google Scholar]
- Perez, L.M.; Redoña, E.D.; Mendioro, M.S.; Vera Cruz, C.M.; Leung, H. Introgression of Xa4, Xa7 and Xa21 for resistance to bacterial blight in thermosensitive genetic male sterile rice (Oryza sativa L.) for the development of two-line hybrids. Euphytica 2008, 164, 627–636. [Google Scholar] [CrossRef]
- Pradhan, S.K.; Nayak, D.K.; Mohanty, S.; Behera, L.; Barik, S.R.; Pandit, E.; Lenka, S.; Anandan, A. Pyramiding of three bacterial blight resistance genes for broad-spectrum resistance in deepwater rice variety, Jalmagna. Rice 2015, 8, 51. [Google Scholar] [CrossRef]
- Prahalada, G.D.; Ramkumar, G.; Hechanova, S.L.; Vinarao, R.; Jena, K.K. Exploring key blast and bacterial blight resistance genes in genetically diverse rice accessions through molecular and phenotypic evaluation. Crop Sci. 2017, 57, 1881–1892. [Google Scholar] [CrossRef]
- Song, W.Y.; Wang, G.L.; Chen, L.L.; Kim, H.S.; Pi, L.Y.; Holsten, T.; Gardner, J.; Wang, B.; Zhai, W.X.; Zhu, L.H.; et al. A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science 1995, 270, 1804–1806. [Google Scholar] [CrossRef]
- Sun, X.; Yang, Z.; Wang, S.; Zhang, Q. Identification of a 47-kb DNA fragment containingXa4, a locus for bacterial blight resistance in rice. Theor. Appl. Genet. 2003, 106, 683–687. [Google Scholar] [CrossRef]
- Yap, R.; Hsu, Y.-C.; Wu, Y.P.; Lin, Y.R.; Kuo, C.W. Multiplex PCR genotyping for five bacterial blight resistance genes applied to marker-assisted selection in rice (Oryza sativa). Plant Breed. 2016, 135, 309–317. [Google Scholar] [CrossRef]
- Iyer, A.S.; McCouch, S.R. The rice bacterial blight resistance gene xa5 encodes a novel form of disease resistance. Mol. Plant Microbe Interact. 2004, 17, 1348–1354. [Google Scholar] [CrossRef]
- Wang, Y.H.; Liu, S.J.; Ji, S.L.; Zhang, W.W.; Wang, C.M.; Jiang, L.; Wan, J.M. Fine mapping and marker-assisted selection (MAS) of a low glutelin content gene in rice. Cell Res. 2005, 15, 622–630. [Google Scholar] [CrossRef]
- Chu, Z.; Fu, B.; Yang, H.; Xu, C.; Li, Z.; Sanchez, A.; Park, Y.J.; Bennetzen, J.L.; Zhang, Q.; Wang, S. Targetingxa13, a recessive gene for bacterial blight resistance in rice. Theor. Appl. Genet. 2006, 112, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Peng, H.; Chen, Z.; Fang, Z.; Zhou, J.; Xia, Z.; Gao, L.; Chen, L.; Li, L.; Li, T.; Zhai, W.; et al. Rice Xa21 primed genes and pathways that are critical for combating bacterial blight infection. Sci. Rep. 2015, 5, 12165. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Sidhu, J.S.; Huang, N.; Vikal, Y.; Li, Z.; Brar, D.S.; Dhaliwal, H.S.; Khush, G.S. Pyramiding three bacterial blight resistance genes (Xa5, Xa13 and Xa21) using marker-assisted selection into indica rice cultivar PR106. Theor. Appl. Genet. 2001, 102, 1011–1015. [Google Scholar] [CrossRef]
- Suh, J.P.; Jeung, J.U.; Noh, T.H.; Cho, Y.C.; Park, S.H.; Park, H.S.; Shin, M.S.; Kim, C.K.; Jena, K.K. Development of breeding lines with three pyramided resistance genes that confer broad-spectrum bacterial blight resistance and their molecular analysis in rice. Rice 2013, 6, 5. [Google Scholar] [CrossRef] [PubMed]
- Ramalingam, J.; Savitha, P.; Alagarasan, G.; Saraswathi, R.; Chandrababu, R. Functional marker assisted improvement of stable cytoplasmic male sterile lines of rice for bacterial blight resistance. Front. Plant Sci. 2017, 8, 1131. [Google Scholar] [CrossRef]
- Ruengphayak, S.; Chaichumpoo, E.; Phromphan, S.; Kamolsukyunyong, W.; Sukhaket, W.; Phuvanartnarubal, E.; Korinsak, S.; Korinsak, S.; Vanavichit, A. Pseudo-backcrossing design for rapidly pyramiding multiple traits into a preferential rice variety. Rice 2015, 8, 7. [Google Scholar] [CrossRef]
- Guvvala, L.; Koradi, P.; Shenoy, V.; Marella, L. Making an Indian traditional rice variety Mahsuri, bacterial blight resistant using marker-assisted selection. J. Crop Sci. Biotech. 2013, 16, 111–121. [Google Scholar] [CrossRef]
- Huang, N.; Angeles, E.R.; Domingo, J.; Magpantay, G.; Singh, S.; Zhang, G.; Kumaravadivel, N.; Bennett, J.; Khush, G.S. Pyramiding of bacterial blight resistance genes in rice: Marker-assisted selection using RFLP and PCR. Theor. Appl. Genet. 1997, 95, 313–320. [Google Scholar] [CrossRef]
- Crossa, J.; Perez-Rodriguez, P.; Cuevas, J.; Montesinos-Lopez, O.; Jarquin, D.; de Los Campos, G.; Burgueno, J.; Gonzalez-Camacho, J.M.; Perez-Elizalde, S.; Beyene, Y.; et al. Genomic selection in plant breeding: Methods, models, and perspectives. Trends Plant Sci. 2017, 22, 961–975. [Google Scholar] [CrossRef]
- Mehta, S.; Singh, B.; Dhakate, P.; Rahman, M.; Islam, M.A. Rice, Marker-Assisted Breeding, and Disease Resistance. In Disease Resistance in Crop Plants: Molecular, Genetic and Genomic Perspectives; Wani, S.H., Ed.; Springer International Publishing: Cham, Switzerland, 2019; pp. 83–111. [Google Scholar]
- Abhilash Kumar, V.; Balachiranjeevi, C.H.; Bhaskar Naik, S.; Rambabu, R.; Rekha, G.; Harika, G.; Hajira, S.K.; Pranathi, K.; Anila, M.; Kousik, M.; et al. Development of gene-pyramid lines of the elite restorer line, RPHR-1005 possessing durable bacterial blight and blast resistance. Front Plant Sci. 2016, 7, 1195. [Google Scholar] [CrossRef]
- Abhilash Kumar, V.; Balachiranjeevi, C.H.; Bhaskar Naik, S.; Rekha, G.; Rambabu, R.; Harika, G.; Pranathi, K.; Hajira, S.K.; Anila, M.; Kousik, M.; et al. Marker-assisted pyramiding of bacterial blight and gall midge resistance genes into RPHR-1005, the restorer line of the popular rice hybrid DRRH-3. Mol. Breed. 2017, 37, 86. [Google Scholar] [CrossRef]
- Yugander, A.; Sundaram, R.M.; Singh, K.; Senguttuvel, P.; Ladhalakshmi, D.; Kemparaju, K.B.; Madhav, M.S.; Prasad, M.S.; Hariprasad, A.S.; Laha, G.S. Improved versions of rice maintainer line, APMS 6B, possessing two resistance genes, Xa21 and Xa38, exhibit high level of resistance to bacterial blight disease. Mol. Breed. 2018, 38, 100. [Google Scholar] [CrossRef]
- Bharathkumar, S.; Paulraj, R.S.D.; Brindha, P.V.; Kavitha, S.; Gnanamanickam, S.S. Improvement of bacterial blight resistance in rice cultivars Jyothi and IR50 via marker-assisted backcross breeding. J. Crop Improv. 2008, 21, 101–116. [Google Scholar] [CrossRef]
- Joseph, M.; Krishnan, S.; Sharma, R.; Singh, V.P.; Singh, A.; Singh, N.; Mohapatra, T. Combining bacterial blight resistance and Basmati quality characteristics by phenotypic and molecular marker-assisted selection in rice. Molecular Breed. 2004, 13, 377–387. [Google Scholar] [CrossRef]
- Luo, Y.; Sangha, J.S.; Wang, S.; Li, Z.; Yang, J.; Yin, Z. Marker-assisted breeding of Xa4, Xa21 and Xa27 in the restorer lines of hybrid rice for broad-spectrum and enhanced disease resistance to bacterial blight. Molecular Breed. 2012, 30, 1601–1610. [Google Scholar] [CrossRef]
- Lalitha Devi, G.; Koradi, P.; Shenoy, V.; Shanti, L. Improvement of resistance to bacterial blight through marker assisted backcross breeding and field validation in rice (Oryza sativa). Res. J. Biol. 2013, 1, 52–66. [Google Scholar]
- Shanti, M.L.; Shenoy, V.V.; Lalitha Devi, G.; Mohan Kumar, V.; Premalatha, P.; Naveen Kumar, G.; Shashidhar, H.E.; Zehr, U.B.; Freeman, W.H. Marker-assisted breeding for resistance to bacterial leaf blight in popular cultivar and parental lines of hybrid rice. J. Plant Pathol. 2010, 92, 495–501. [Google Scholar]
- Huang, B.; Xu, J.Y.; Hou, M.S.; Ali, J.; Mou, T.M. Introgression of bacterial blight resistance genes Xa7, Xa21, Xa22 and Xa23 into hybrid rice restorer lines by molecular marker-assisted selection. Euphytica 2012, 187, 449–459. [Google Scholar] [CrossRef]
- Hasan, M.M.; Rafii, M.Y.; Ismail, M.R.; Mahmood, M.; Rahim, H.A.; Alam, M.A.; Ashkani, S.; Malek, M.A.; Latif, M.A. Marker-assisted backcrossing: A useful method for rice improvement. Biotechnol. Biotechnol. Equip. 2015, 29, 237–254. [Google Scholar] [CrossRef]
- Jiang, G.L. Molecular Markers and Marker–Assisted Breeding in Plants. In Plant Breeding from Laboratories to Fields; Andersen, S.B., Ed.; IntechOpen: Croatia, 2013; pp. 45–83. [Google Scholar]
- Rajpurohit, D.; Kumar, R.; Kumar, M.; Paul, P.; Awasthi, A.; Osman Basha, P.; Puri, A.; Jhang, T.; Singh, K.; Dhaliwal, H.S. Pyramiding of two bacterial blight resistance and a semidwarfing gene in Type 3 Basmati using marker-assisted selection. Euphytica 2011, 178, 111–126. [Google Scholar] [CrossRef]
- Sundaram, R.M.; Vishnupriya, M.R.; Biradar, S.K.; Laha, G.S.; Reddy, G.A.; Rani, N.S.; Sarma, N.P.; Sonti, R.V. Marker assisted introgression of bacterial blight resistance in Samba Mahsuri, an elite indica rice variety. Euphytica 2008, 160, 411–422. [Google Scholar] [CrossRef]
- Balachiranjeevi, C.; Bhaskar, N.S.; Abhilash, V.; Akanksha, S.; Viraktamath, B.C.; Madhav, M.S.; Hariprasad, A.S.; Laha, G.S.; Prasad, M.S.; Balachandran, S.; et al. Marker-assisted introgression of bacterial blight and blast resistance into DRR17B, an elite, fine-grain type maintainer line of rice. Mol. Breed. 2015, 35, 151. [Google Scholar] [CrossRef]
- Singh, A.; Singh, V.K.; Singh, S.P.; Pandian, R.T.P.; Ellur, R.K.; Singh, D.; Bhowmick, P.K.; Gopala Krishnan, S.; Nagarajan, M.; Vinod, K.K.; et al. Molecular breeding for the development of multiple disease resistance in Basmati rice. AoB Plants 2012, 2012. [Google Scholar] [CrossRef] [PubMed]
- Cobb, J.N.; Biswas, P.S.; Platten, J.D. Back to the future: Revisiting MAS as a tool for modern plant breeding. Theor. Appl. Genet. 2019, 132, 647–667. [Google Scholar] [CrossRef]
- Tseng, H.Y.; Lin, D.G.; Hsieh, H.Y.; Tseng, Y.J.; Tseng, W.B.; Chen, C.W.; Wang, C.S. Genetic analysis and molecular mapping of QTLs associated with resistance to bacterial blight in a rice mutant, SA0423. Euphytica 2015, 205, 231–241. [Google Scholar] [CrossRef]
- Wang, C.S.; Lin, D.G. The Application of Genomic Approaches in Studying a Bacterial Blight–Resistant Mutant in Rice. In Advances in International Rice Research; Li, J.Q., Li, J., Eds.; IntechOpen: London, UK, 2017. [Google Scholar]
- Kauffman, H.E.; Reddy, A.P.K.; Hsieh, S.P.Y.; Merca, S.D. An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae. Plant Dis. Rep. 1973, 57, 537–541. [Google Scholar]
- Kado, C.I.; Heskett, M.G. Selective media for isolation of Agrobacterium, Corynebacterium, Erwinia, Psedumonoas and Xanthomonas. Phytopathology 1970, 60, 969–976. [Google Scholar] [CrossRef]
- IRRI. Standard Evaluation System; In International Rice Research Institute: Manila, Philippines, 1996; p. 52. [Google Scholar]
- Latif, M.A.; Badsha, M.A.; Tajul, M.I.; Kabir, M.S.; Rafii, M.Y.; Mia, M.A.T. Dentification of genotypes resistant to blast, bacterial leaf blight, sheath blight and tungro and efficacy of seed treating fungicides against blast disease of rice. Sci. Res. Essays 2011, 6, 2804–2811. [Google Scholar]
- Hsu, Y.C.; Tseng, M.C.; Wu, Y.P.; Lin, M.Y.; Wei, F.J.; Hwu, K.K.; Hsing, Y.I.; Lin, Y.R. Genetic factors responsible for eating and cooking qualities of rice grains in a recombinant inbred population of an inter-subspecific cross. Mol. Breed. 2014, 34, 655–673. [Google Scholar] [CrossRef]
- Van Berloo, R. GGT: Software for the display of graphical genotypes. J. Hered. 1999, 90, 328–330. [Google Scholar] [CrossRef]
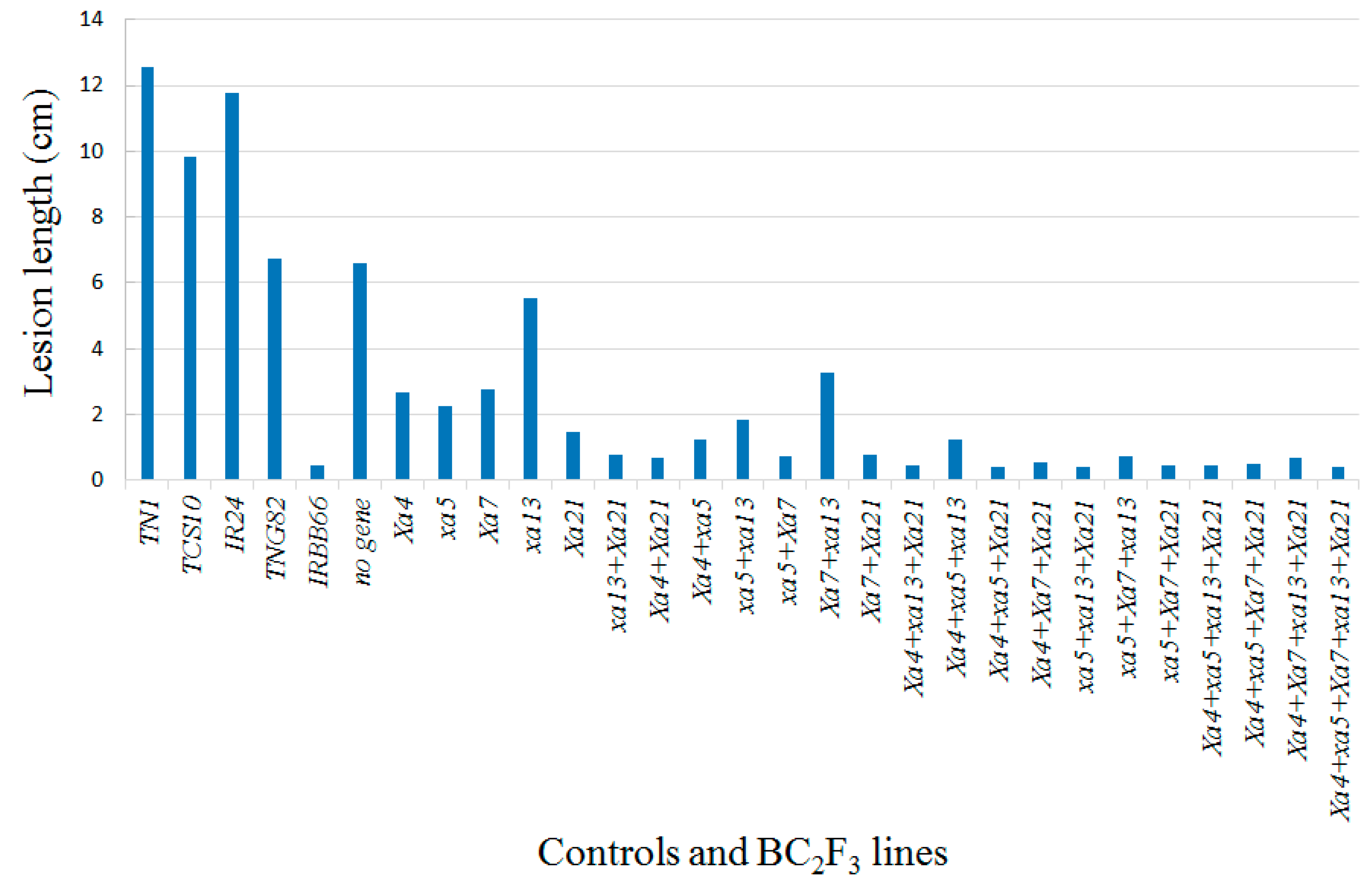
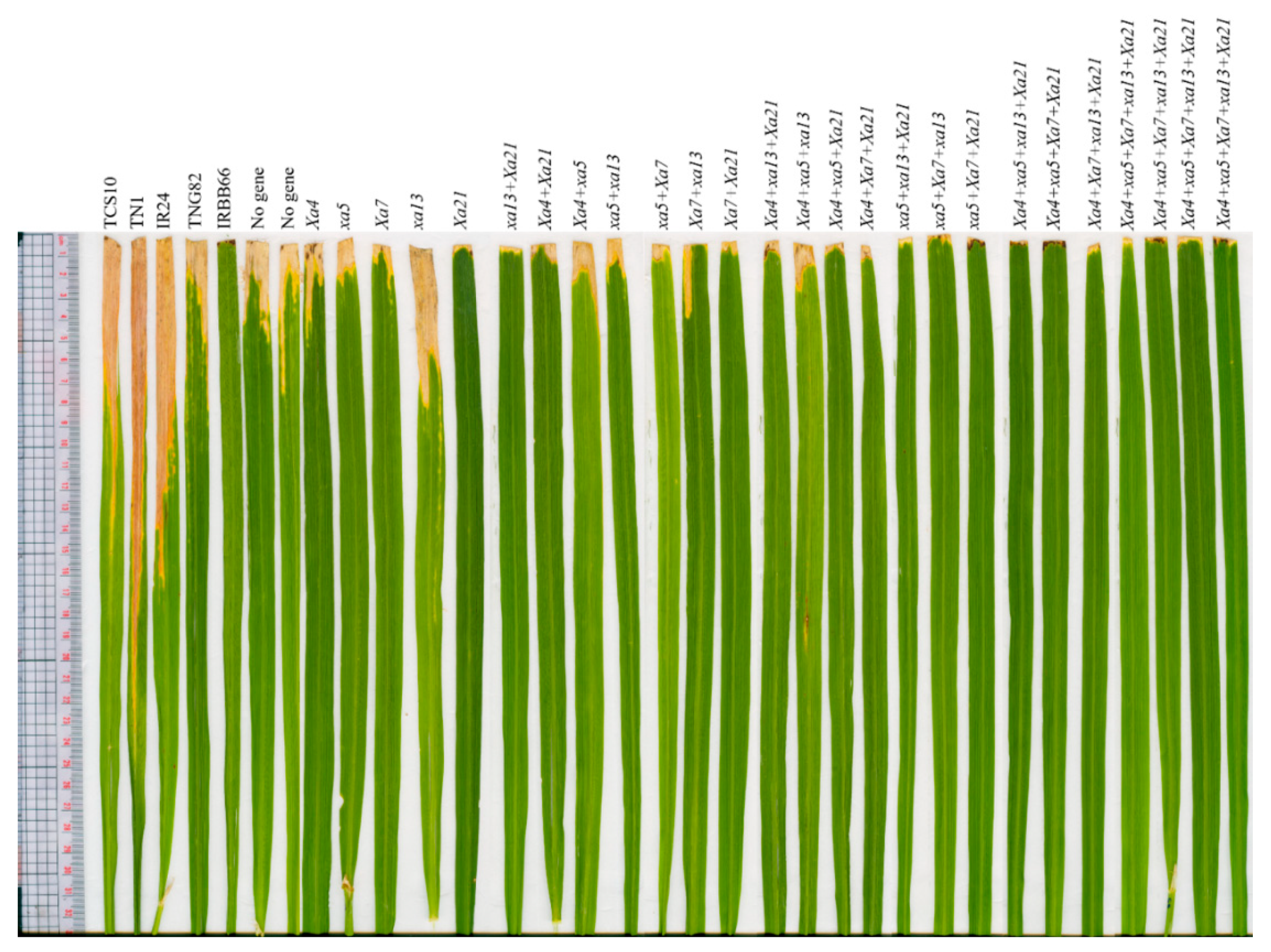

| No. | Lines | Genotypes | Lesion Length † (cm) | Resistance Scale ¥ |
|---|---|---|---|---|
| 1 | TN1 | 12.56 ± 2.98 a | S | |
| 2 | TCS10 | 9.85 ± 2.07 b | MS | |
| 3 | IR24 | 11.75 ± 1.80 a | S | |
| 4 | TNG82 | 6.75 ± 2.54 c,d | MS | |
| 5 | IRBB66 | Xa4 + xa5 + Xa7 + xa13 + Xa21 | 0.43 ± 0.70 i | HR |
| 6 | CNYBB0R01 | Without resistance gene | 7.30 ± 1.50 d | MS |
| 7 | CNYBB0R02 | Without resistance gene | 5.91 ± 2.04 c | MR |
| 8 | CNYBB1R01 | Xa4 | 2.67 ± 1.00 e,f | R |
| 9 | CNYBB1R02 | xa5 | 2.27 ± 0.73 e,f,g | R |
| 10 | CNYBB1R03 | Xa7 | 2.77 ± 1.04 e,f | R |
| 11 | CNYBB1R04 | xa13 | 5.55 ± 2.28 d | MR |
| 12 | CNYBB1R05 | Xa21 | 1.48 ± 1.48 f,g,h,i | R |
| 13 | CNYBB2R03 | xa13 + Xa21 | 0.77 ± 0.45 h,i | HR |
| 14 | CNYBB2R04 | Xa4 + Xa21 | 0.68 ± 0.24 h,i | HR |
| 15 | CNYBB2R05 | Xa4 + xa5 | 1.24 ± 0.98 g,h,i | R |
| 16 | CNYBB2R06 | xa5 + xa13 | 1.82 ± 0.54 f,g,h | R |
| 17 | CNYBB2R01 | xa5 + Xa7 | 0.75 ± 0.33 h,i | HR |
| 18 | CNYBB2R07 | Xa7 + xa13 | 3.25 ± 0.73 e | MR |
| 19 | CNYBB2R02 | Xa7 + Xa21 | 0.76 ± 0.19 h,i | HR |
| 20 | CNYBB3R03 | Xa4 + xa13 + Xa21 | 0.45 ± 0.17 h,i | HR |
| 21 | CNYBB3R04 | Xa4 + xa5 + xa13 | 1.25 ± 0.74 g,h,i | R |
| 22 | CNYBB3R05 | Xa4 + xa5 + Xa21 | 0.46 ± 0.17 h,i | HR |
| 23 | CNYBB3R01 | Xa4 + Xa7 + Xa21 | 0.56 ± 0.16 h,i | HR |
| 24 | CNYBB3R06 | xa5 + xa13 + Xa21 | 0.44 ± 0.08 h,i | HR |
| 25 | CNYBB3R07 | xa5 + Xa7 + xa13 | 0.71 ± 0.45 h,i | HR |
| 26 | CNYBB3R02 | xa5 + Xa7 + Xa21 | 0.44 ± 0.12 h,i | HR |
| 27 | CNYBB4R03 | Xa4 + xa5 + xa13 + Xa21 | 0.43 ± 0.12 i | HR |
| 28 | CNYBB4R01 | Xa4 + xa5 + Xa7 + Xa21 | 0.48 ± 0.17 h,i | HR |
| 29 | CNYBB4R02 | Xa4 + Xa7 + xa13 + Xa21 | 0.68 ± 0.50 h,i | HR |
| 30 | CNYBB5R01 | Xa4 + xa5 + Xa7 + xa13 + Xa21 | 0.37 ± 0.13 i | HR |
| 31 | CNYBB5R02 | Xa4 + xa5 + Xa7 + xa13 + Xa21 | 0.38 ± 0.12 i | HR |
| 32 | CNYBB5R03 | Xa4 + xa5 + Xa7 + xa13 + Xa21 | 0.46 ± 0.14 h,i | HR |
| 33 | CNYBB5R04 | Xa4 + xa5 + Xa7 + xa13 + Xa21 | 0.38 ± 0.12 i | HR |
| Pyramided Lines | Plant Height (cm) (n = 20) | Days to 50% Flowering (n = 20) | Panicle Length (cm) (n = 20) | Panicles/Plant (n = 20) | Panicle Weight (g) (n = 20) | No. of Grains/Panicle (n = 20) | 1000-Seed Weight (g) (n = 20) | Single Plant Yield (g) (n = 20) |
|---|---|---|---|---|---|---|---|---|
| TNG82 | 105.4 ± 1.1 | 90 | 20.3 ± 0.2 | 14 ± 0.7 | 3.3 ± 0.1 | 102 ± 2.5 | 30.8 ± 0.1 | 36.8 ± 0.1 |
| IRBB66 | 90.2 ± 1.4 | 95 | 22.6 ± 0.1 | 17 ± 0.5 | 3.7 ± 0.1 | 108 ± 1.0 | 24.2 ± 0.1 | 30.1 ± 0.7 |
| CNYBB5R4-272 | 120.4 ± 1.3 | 92 | 22.8 ± 1.8 | 16 ± 0.8 | 3.6 ± 0.4 | 122 ± 16.7 | 26.0 ± 0.1 | 36.3 ± 7.3 |
| CNYBB5R4-274 | 115.9 ± 2.7 | 91 | 21.4 ± 1.6 | 15 ± 0.5 | 2.8 ± 0.1 | 95 ± 8.2 | 28.4 ± 0.4 | 32.6 ± 4.4 |
| CNYBB5R4-275 | 116.3 ± 3.7 | 91 | 20.1 ± 0.6 | 16 ± 2.1 | 3.2 ± 0.3 | 113 ± 17.8 | 25.2 ± 0.4 | 41.2 ± 3.3 |
| CNYBB5R4-276 | 116.3 ± 3.2 | 91 | 20.4 ± 0.7 | 16 ± 0.5 | 2.8 ± 0.2 | 100 ± 9.9 | 26.4 ± 0.4 | 43.7 ± 0.9 |
| CNYBB5R4-278 | 115.8 ± 2.4 | 91 | 21.7 ± 0.2 | 12 ± 0.8 | 3.9 ± 0.3 | 135 ± 10.3 | 26.2 ± 0.1 | 44.5 ± 1.3 |
| CNYBB5R4-279 | 112.7 ± 4.2 | 91 | 18.3 ± 0.1 | 15 ± 1.7 | 3.1 ± 0.1 | 120 ± 4.7 | 25.2 ± 0.2 | 39.2 ± 3.0 |
| CNYBB5R4-285 | 121.8 ± 3.0 | 92 | 21.6 ± 0.1 | 14 ± 0.8 | 3.1 ± 0.1 | 106 ± 0.8 | 28.2 ± 0.2 | 38.0 ± 0.3 |
| CNYBB5R4-286 | 113.7 ± 6.8 | 91 | 20.2 ± 1.0 | 15 ± 1.0 | 2.9 ± 0.2 | 100 ± 12.1 | 27.6 ± 0.1 | 35.9 ± 2.6 |
| CNYBB5R4-287 | 115.3 ± 6.6 | 91 | 21.7 ± 0.6 | 14 ± 0.2 | 2.8 ± 0.4 | 103 ± 16.5 | 26.6 ± 1.6 | 37.1 ± 8.1 |
| LSD (p = 0.05) | 9.2 | 0.6 | 2.6 | 3.1 | 0.7 | 24.0 | 1.7 | 12.1 |
| Pyramided Lines | Palatability (n = 20) | Protein (n = 20) | Amylose (n = 20) | Brown Rice (%) (n = 20) | Head Rice (%) (n = 20) | Total Milled Rice (%) (n = 20) |
|---|---|---|---|---|---|---|
| TNG82 | 76.3 ± 3.2 | 6.0 ± 0.5 | 16.0 ± 0.1 | 77.6 ± 0.2 | 55.0 ± 0.1 | 61.5 ± 0.7 |
| IRBB66 | 60.3 ± 1.1 | 8.7 ± 0.3 | 18.1 ± 0.1 | 72.3 ± 0.1 | 42.3 ± 0.1 | 45.7 ± 0.4 |
| CNYBB5R4-272 | 69.0 ± 2.1 | 7.4 ± 0.4 | 14.5 ± 0.9 | 74.1 ± 0.5 | 50.5 ± 0.7 | 57.8 ± 0.8 |
| CNYBB5R4-274 | 70.0 ± 3.5 | 7.3 ± 0.7 | 15.3 ± 0.2 | 72.8 ± 0.6 | 31.2 ± 1.8 | 48.6 ± 0.1 |
| CNYBB5R4-275 | 69.8 ± 1.1 | 7.2 ± 0.2 | 16.0 ± 0.4 | 75.1 ± 2.3 | 53.8 ± 1.5 | 58.7 ± 1.3 |
| CNYBB5R4-276 | 74.5 ± 2.8 | 6.4 ± 0.5 | 15.4 ± 0.1 | 77.2 ± 0.9 | 50.1 ± 2.4 | 59.0 ± 1.0 |
| CNYBB5R4-278 | 72.3 ± 1.8 | 6.7 ± 0.3 | 15.6 ± 0.2 | 77.5 ± 0.7 | 55.8 ± 0.3 | 62.1 ± 0.1 |
| CNYBB5R4-279 | 72.3 ± 0.4 | 6.8 ± 0.1 | 15.3 ± 0.2 | 77.2 ± 1.8 | 49.3 ± 2.6 | 58.1 ± 3.6 |
| CNYBB5R4-285 | 72.0 ± 1.4 | 7.0 ± 0.7 | 15.2 ± 0.1 | 77.0 ± 1.4 | 37.0 ± 1.9 | 54.4 ± 2.2 |
| CNYBB5R4-286 | 74.3 ± 1.4 | 6.4 ± 0.3 | 15.3 ± 0.2 | 79.3 ± 0.5 | 48.4 ± 2.4 | 61.2 ± 0.1 |
| CNYBB5R4-287 | 74.3 ± 0.4 | 6.8 ± 0.1 | 15.4 ± 0.1 | 75.5 ± 0.5 | 42.0 ± 9.0 | 54.7 ± 0.6 |
| LSD (p = 0.05) | 4.6 | 0.9 | 1.0 | 3.5 | 9.8 | 4.5 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hsu, Y.-C.; Chiu, C.-H.; Yap, R.; Tseng, Y.-C.; Wu, Y.-P. Pyramiding Bacterial Blight Resistance Genes in Tainung82 for Broad-Spectrum Resistance Using Marker-Assisted Selection. Int. J. Mol. Sci. 2020, 21, 1281. https://doi.org/10.3390/ijms21041281
Hsu Y-C, Chiu C-H, Yap R, Tseng Y-C, Wu Y-P. Pyramiding Bacterial Blight Resistance Genes in Tainung82 for Broad-Spectrum Resistance Using Marker-Assisted Selection. International Journal of Molecular Sciences. 2020; 21(4):1281. https://doi.org/10.3390/ijms21041281
Chicago/Turabian StyleHsu, Yu-Chia, Chih-Hao Chiu, Ruishen Yap, Yu-Chien Tseng, and Yong-Pei Wu. 2020. "Pyramiding Bacterial Blight Resistance Genes in Tainung82 for Broad-Spectrum Resistance Using Marker-Assisted Selection" International Journal of Molecular Sciences 21, no. 4: 1281. https://doi.org/10.3390/ijms21041281
APA StyleHsu, Y.-C., Chiu, C.-H., Yap, R., Tseng, Y.-C., & Wu, Y.-P. (2020). Pyramiding Bacterial Blight Resistance Genes in Tainung82 for Broad-Spectrum Resistance Using Marker-Assisted Selection. International Journal of Molecular Sciences, 21(4), 1281. https://doi.org/10.3390/ijms21041281





