Genetic Dissection of Alzheimer’s Disease Using Drosophila Models
Abstract
1. Introduction
1.1. Genetics of Alzheimer’s Disease
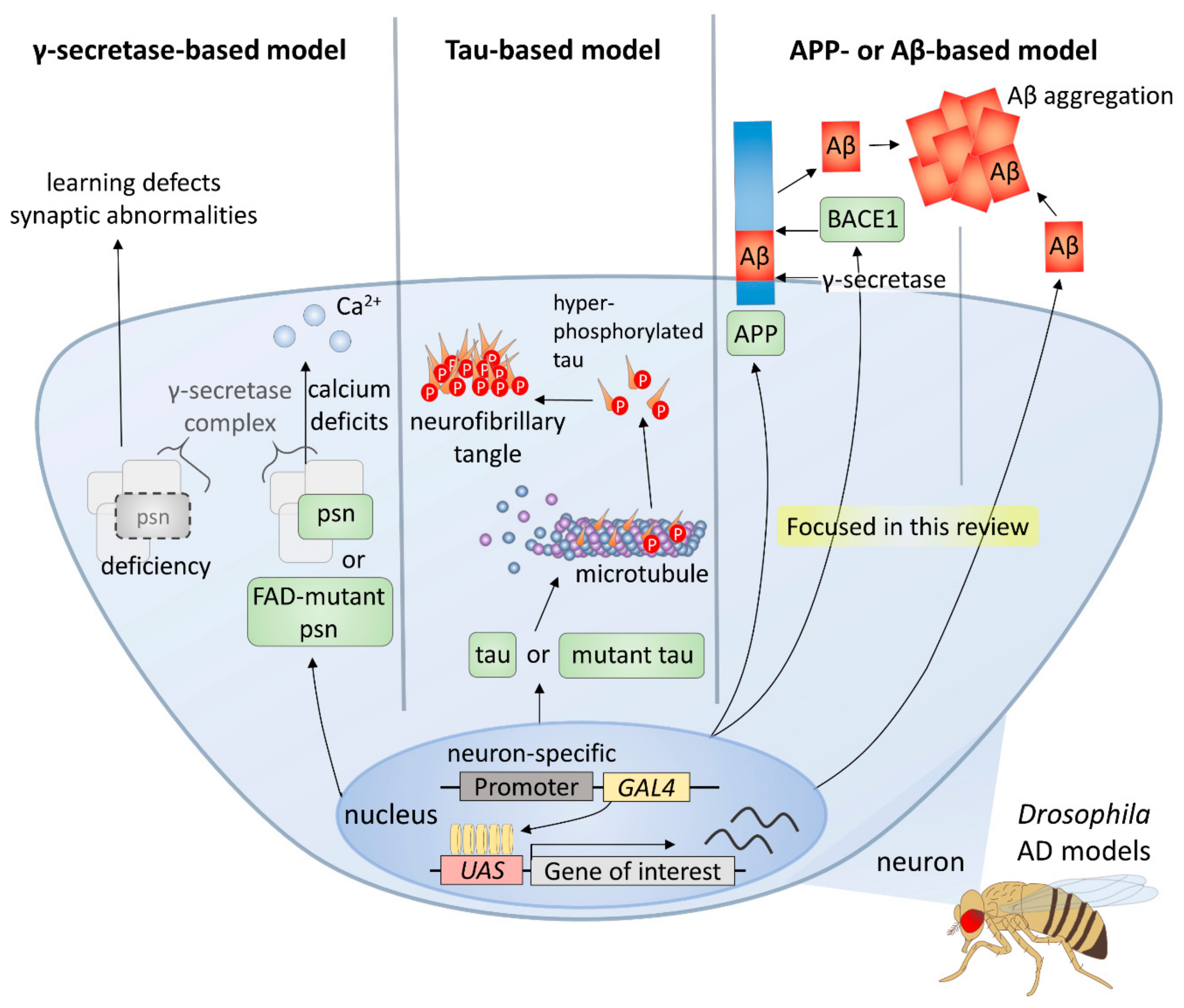
1.2. Drosophila Models of Alzheimer’s Disease
2. AD-Related Mechanisms and Genetic Modifiers Identified Using the Drosophila Model
2.1. Amyloid Beta Accumulation
2.1.1. Soluble Aβ Oligomer Toxicity and Aggregation
2.1.2. Aβ Degradation
2.1.3. Intraneuronal Accumulation of Aβ
2.2. Amyloid-Mediated Tauopathy
2.3. Modifiers Related to Stress-Responsive Pathways
2.3.1. Oxidative Stress
2.3.2. Endoplasmic Reticulum Stress
2.4. Modifiers Involved in ERK Pathway or Cell Cycle
2.4.1. EGFR/ERK Signaling
2.4.2. Cell Cycle
2.5. Modifiers Related to Apoptosis
2.6. Modifiers Related to Epigenetic Regulation
2.7. Modifiers Related With Synaptic Abnormalities
2.7.1. Small GTPases
2.7.2. Impaired Axonal Transport
2.7.3. Synaptic Proteins
2.8. Mitochondrial Dysfunction or Mislocation
2.9. Modifiers Related with Inflammation
2.9.1. Phagocytic Receptor Draper
2.9.2. TREM2
2.9.3. Toll and IMD Pathways
3. Conclusions and Perspectives
Author Contributions
Acknowledgments
Conflicts of Interest
Abbreviations
| AD | Alzheimer’s disease |
| Aβ | Amyloid β |
| APLP1 | Amyloid precursor like protein 1 |
| APP | Amyloid precursor protein |
| ATF6 | Activating transcription factor 6 |
| B4GALT6 | β1,4-galactosyltransferases |
| BHQ | Bromo, HAT and poly glutamine stretch |
| CA | Constitutive active |
| CAT | Catalase |
| CCR | Cell cycle reentry |
| CBP | CREB-binding protein |
| Ctr1B | Copper transporter 1B |
| dFOXO | Drosophila forkhead box subgroup O |
| DIAP1 | Death-associated inhibitor of apoptosis 1 |
| Drp1 | Dynamin-related protein 1 |
| EGFR | Epidermal growth factor receptor |
| EOAD | Early onset AD |
| ER | Endoplasmic reticulum |
| ERC2 | ELKS/RAB6-interacting/CAST family member 2 |
| ERK | Extracellular signal-regulated kinase |
| Fer1HCH | Ferritin 1 heavy chain homologue |
| Fer2LCH | Ferritin 2 light chain homologue |
| GSK3β | Glycogen synthase kinase 3β |
| GWAS | Genome-wide association studies |
| HAT | Histone acetyltransferases |
| HDAC | Histone deacetylases |
| hep | hemipterous |
| HPD | 4-hydroxyphenylpyruvate dioxygenase |
| Hsp70 | Heat shock protein 70 |
| IDE | Insulin degradation enzyme |
| IMD | Immune deficiency |
| JNK | c-jun N-terminal kinase |
| LOAD | Late onset AD |
| MARK | Microtubule affinity regulating kinase |
| MAPKs | Mitogen-activated protein kinases |
| NEP | Neprilysin |
| NFTs | Neurofibrillary tangles |
| PERK | Protein kinase R-like endoplasmic reticulum kinase |
| PGRP-SC1b | Peptidogylcan recognition protein SC1b |
| PLD3 | Phospholipase D3 |
| PRCC | Proline rich mitotic checkpoint control factor |
| psn | presenilin |
| RBO | Rolling blackout |
| SAT1 | α2,3-sialyltransferase |
| Ser | Serine |
| SH2B1 | Src homology 2B1 |
| SOD | Superoxide dismutase |
| sra | sarah |
| Thr | Threonine |
| TLRs | Toll-like receptors |
| TREM2 | Triggering receptor expressed on myeloid cell 2 |
| Trx80 | Thioredoxin-80 |
| TSPO | Translocator protein 18 kDa |
| TTC7 | Tetratricopeptide repeat domain 7 |
| XBP1 | X-box binding protein 1 |
| XBP1-S | spliced XBP1 |
| Zip1 | zinc/iron regulated transporter-related protein 1 |
References
- Alzheimer’s Disease International. Executive summary. In World Alzheimer Report 2019: Attitudes to Dementia; Alzheimer’s Disease International (ADI): London, UK, 2019; pp. 1–160. [Google Scholar]
- Cummings, J.L. Alzheimer’s disease. N. Engl. J. Med. 2004, 351, 56–67. [Google Scholar] [CrossRef]
- Ballard, C.; Gauthier, S.; Corbett, A.; Brayne, C.; Aarsland, D.; Jones, E. Alzheimer’s disease. Lancet 2011, 377, 1019–1031. [Google Scholar] [CrossRef]
- Hardy, J.A.; Higgins, G.A. Alzheimer’s disease: The amyloid cascade hypothesis. Science 1992, 256, 184–186. [Google Scholar] [CrossRef] [PubMed]
- Mudher, A.; Lovestone, S. Alzheimer’s disease–do tauists and baptists finally shake hands? Trends Neurosci. 2002, 25, 22–26. [Google Scholar] [CrossRef]
- Haass, C.; Selkoe, D.J. Soluble protein oligomers in neurodegeneration: Lessons from the Alzheimer’s amyloid β-peptide. Nat. Rev. Mol. Cell Biol. 2007, 8, 101. [Google Scholar] [CrossRef] [PubMed]
- Wisniewski, K.; Wisniewski, H.; Wen, G. Occurrence of neuropathological changes and dementia of Alzheimer’s disease in Down’s syndrome. Ann. Neurol. 1985, 17, 278–282. [Google Scholar] [CrossRef]
- Mullan, M.; Crawford, F.; Axelman, K.; Houlden, H.; Lilius, L.; Winblad, B.; Lannfelt, L. A pathogenic mutation for probable Alzheimer’s disease in the APP gene at the N–terminus of β–amyloid. Nat. Genet. 1992, 1, 345. [Google Scholar] [CrossRef]
- Hardy, J. Amyloid, the presenilins and Alzheimer’s disease. Trends Neurosci. 1997, 20, 154–159. [Google Scholar] [CrossRef]
- Ballatore, C.; Lee, V.M.-Y.; Trojanowski, J.Q. Tau-mediated neurodegeneration in Alzheimer’s disease and related disorders. Nat. Rev. Neurosci. 2007, 8, 663. [Google Scholar] [CrossRef]
- Mehta, D.; Jackson, R.; Paul, G.; Shi, J.; Sabbagh, M. Why do trials for Alzheimer’s disease drugs keep failing? A discontinued drug perspective for 2010-2015. Expert Opin. Investig. Drugs 2017, 26, 735–739. [Google Scholar] [CrossRef]
- Barnes, D.E.; Yaffe, K. The projected effect of risk factor reduction on Alzheimer’s disease prevalence. Lancet Neurol. 2011, 10, 819–828. [Google Scholar] [CrossRef]
- Bettens, K.; Sleegers, K.; Van Broeckhoven, C. Genetic insights in Alzheimer’s disease. Lancet Neurol. 2013, 12, 92–104. [Google Scholar] [CrossRef]
- Gatz, M.; Reynolds, C.A.; Fratiglioni, L.; Johansson, B.; Mortimer, J.A.; Berg, S.; Fiske, A.; Pedersen, N.L. Role of genes and environments for explaining Alzheimer disease. Arch. Gen. Psychiatry 2006, 63, 168–174. [Google Scholar] [CrossRef] [PubMed]
- Van Dam, D.; De Deyn, P.P. Animal models in the drug discovery pipeline for Alzheimer’s disease. Br. J. Pharmacol. 2011, 164, 1285–1300. [Google Scholar] [CrossRef]
- Tan, F.H.P.; Azzam, G. Drosophila melanogaster: Deciphering Alzheimer’s Disease. Malays. J. Med. Sci. 2017, 24, 6–20. [Google Scholar] [CrossRef]
- Yeates, C.J.; Sarkar, A.; Kango-Singh, M.; Singh, A. Unraveling Alzheimer’s Disease Using Drosophila. In Insights into Human Neurodegeneration: Lessons Learnt from Drosophila; Mutsuddi, M., Mukherjee, A., Eds.; Springer: Singapore, 2019; pp. 251–277. [Google Scholar]
- Guo, Y.; Livne-Bar, I.; Zhou, L.; Boulianne, G.L. Drosophila presenilin is required for neuronal differentiation and affects notch subcellular localization and signaling. J. Neurosci. 1999, 19, 8435–8442. [Google Scholar] [CrossRef]
- Ye, Y.; Fortini, M.E. Apoptotic activities of wild-type and Alzheimer’s disease-related mutant presenilins in Drosophila melanogaster. J. Cell Biol. 1999, 146, 1351–1364. [Google Scholar] [CrossRef]
- Michno, K.; Knight, D.; Campusano, J.M.; van de Hoef, D.; Boulianne, G.L. Intracellular calcium deficits in Drosophila cholinergic neurons expressing wild type or FAD-mutant presenilin. PLoS ONE 2009, 4, e6904. [Google Scholar] [CrossRef]
- Knight, D.; Iliadi, K.; Charlton, M.P.; Atwood, H.L.; Boulianne, G.L. Presynaptic plasticity and associative learning are impaired in a Drosophila presenilin null mutant. Dev. Neurobiol. 2007, 67, 1598–1613. [Google Scholar] [CrossRef]
- Wittmann, C.W.; Wszolek, M.F.; Shulman, J.M.; Salvaterra, P.M.; Lewis, J.; Hutton, M.; Feany, M.B. Tauopathy in Drosophila: Neurodegeneration without neurofibrillary tangles. Science 2001, 293, 711–714. [Google Scholar] [CrossRef]
- Jackson, G.R.; Wiedau-Pazos, M.; Sang, T.K.; Wagle, N.; Brown, C.A.; Massachi, S.; Geschwind, D.H. Human wild-type tau interacts with wingless pathway components and produces neurofibrillary pathology in Drosophila. Neuron 2002, 34, 509–519. [Google Scholar] [CrossRef]
- Shulman, J.M.; Feany, M.B. Genetic modifiers of tauopathy in Drosophila. Genetics 2003, 165, 1233–1242. [Google Scholar] [PubMed]
- Folwell, J.; Cowan, C.M.; Ubhi, K.K.; Shiabh, H.; Newman, T.A.; Shepherd, D.; Mudher, A. Aβ exacerbates the neuronal dysfunction caused by human tau expression in a Drosophila model of Alzheimer’s disease. Exp. Neurol. 2010, 223, 401–409. [Google Scholar] [CrossRef]
- Tanzi, R.E.; Bertram, L. Twenty years of the Alzheimer’s disease amyloid hypothesis: A genetic perspective. Cell 2005, 120, 545–555. [Google Scholar] [CrossRef] [PubMed]
- Greeve, I.; Kretzschmar, D.; Tschape, J.A.; Beyn, A.; Brellinger, C.; Schweizer, M.; Nitsch, R.M.; Reifegerste, R. Age-dependent neurodegeneration and Alzheimer-amyloid plaque formation in transgenic Drosophila. J. Neurosci. 2004, 24, 3899–3906. [Google Scholar] [CrossRef]
- Carmine-Simmen, K.; Proctor, T.; Tschape, J.; Poeck, B.; Triphan, T.; Strauss, R.; Kretzschmar, D. Neurotoxic effects induced by the Drosophila amyloid-beta peptide suggest a conserved toxic function. Neurobiol. Dis. 2009, 33, 274–281. [Google Scholar] [CrossRef]
- Chakraborty, R.; Vepuri, V.; Mhatre, S.D.; Paddock, B.E.; Miller, S.; Michelson, S.J.; Delvadia, R.; Desai, A.; Vinokur, M.; Melicharek, D.J. Characterization of a Drosophila Alzheimer’s disease model: Pharmacological rescue of cognitive defects. PLoS ONE 2011, 6, e20799. [Google Scholar] [CrossRef]
- Stokin, G.B.; Almenar-Queralt, A.; Gunawardena, S.; Rodrigues, E.M.; Falzone, T.; Kim, J.; Lillo, C.; Mount, S.L.; Roberts, E.A.; McGowan, E.; et al. Amyloid precursor protein-induced axonopathies are independent of amyloid-beta peptides. Hum. Mol. Genet. 2008, 17, 3474–3486. [Google Scholar] [CrossRef]
- Muller, T.; Meyer, H.E.; Egensperger, R.; Marcus, K. The amyloid precursor protein intracellular domain (AICD) as modulator of gene expression, apoptosis, and cytoskeletal dynamics-relevance for Alzheimer’s disease. Prog. Neurobiol. 2008, 85, 393–406. [Google Scholar] [CrossRef]
- Finelli, A.; Kelkar, A.; Song, H.J.; Yang, H.; Konsolaki, M. A model for studying Alzheimer’s Abeta42-induced toxicity in Drosophila melanogaster. Mol. Cell. Neurosci. 2004, 26, 365–375. [Google Scholar] [CrossRef]
- Crowther, D.C.; Kinghorn, K.J.; Miranda, E.; Page, R.; Curry, J.A.; Duthie, F.A.; Gubb, D.C.; Lomas, D.A. Intraneuronal Abeta, non-amyloid aggregates and neurodegeneration in a Drosophila model of Alzheimer’s disease. Neuroscience 2005, 132, 123–135. [Google Scholar] [CrossRef] [PubMed]
- Iijima, K.; Iijima-Ando, K. Drosophila models of Alzheimer’s amyloidosis: The challenge of dissecting the complex mechanisms of toxicity of amyloid-β 42. J. Alzheimers Dis. 2008, 15, 523–540. [Google Scholar] [CrossRef] [PubMed]
- Casas-Tinto, S.; Zhang, Y.; Sanchez-Garcia, J.; Gomez-Velazquez, M.; Rincon-Limas, D.E.; Fernandez-Funez, P. The ER stress factor XBP1s prevents amyloid-beta neurotoxicity. Hum. Mol. Genet. 2011, 20, 2144–2160. [Google Scholar] [CrossRef] [PubMed]
- Jeon, Y.; Lee, S.; Shin, M.; Lee, J.H.; Suh, Y.S.; Hwang, S.; Yun, H.S.; Cho, K.S. Phenotypic differences between Drosophila Alzheimer’s disease models expressing human Aβ42 in the developing eye and brain. Anim. Cells Syst. 2017, 21, 160–168. [Google Scholar] [CrossRef]
- Iijima, K.; Chiang, H.C.; Hearn, S.A.; Hakker, I.; Gatt, A.; Shenton, C.; Granger, L.; Leung, A.; Iijima-Ando, K.; Zhong, Y. Abeta42 mutants with different aggregation profiles induce distinct pathologies in Drosophila. PLoS ONE 2008, 3, e1703. [Google Scholar] [CrossRef]
- Tare, M.; Modi, R.M.; Nainaparampil, J.J.; Puli, O.R.; Bedi, S.; Fernandez-Funez, P.; Kango-Singh, M.; Singh, A. Activation of JNK signaling mediates amyloid-ss-dependent cell death. PLoS ONE 2011, 6, e24361. [Google Scholar] [CrossRef]
- Moran, M.T.; Tare, M.; Kango-Singh, M.; Singh, A. Homeotic Gene teashirt (tsh) has a neuroprotective function in amyloid-beta 42 mediated neurodegeneration. PLoS ONE 2013, 8, e80829. [Google Scholar] [CrossRef]
- Steffensmeier, A.M.; Tare, M.; Puli, O.R.; Modi, R.; Nainaparampil, J.; Kango-Singh, M.; Singh, A. Novel neuroprotective function of apical-basal polarity gene crumbs in amyloid beta 42 (aβ42) mediated neurodegeneration. PLoS ONE 2013, 8, e78717. [Google Scholar] [CrossRef]
- Jucker, M.; Walker, L.C. Self-propagation of pathogenic protein aggregates in neurodegenerative diseases. Nature 2013, 501, 45–51. [Google Scholar] [CrossRef]
- Jaunmuktane, Z.; Mead, S.; Ellis, M.; Wadsworth, J.D.; Nicoll, A.J.; Kenny, J.; Launchbury, F.; Linehan, J.; Richard-Loendt, A.; Walker, A.S. Evidence for human transmission of amyloid-β pathology and cerebral amyloid angiopathy. Nature 2015, 525, 247. [Google Scholar] [CrossRef]
- Eisele, Y.S. From Soluble A β to Progressive A β Aggregation: Could prion-like templated misfolding play a role? Brain Pathol. 2013, 23, 333–341. [Google Scholar] [CrossRef]
- Sowade, R.F.; Jahn, T.R. Seed-induced acceleration of amyloid-β mediated neurotoxicity in vivo. Nat. Commun. 2017, 8, 512. [Google Scholar] [CrossRef]
- Pitschke, M.; Prior, R.; Haupt, M.; Riesner, D. Detection of single amyloid β-protein aggregates in the cerebrospinal fluid of Alzheimer’s patients by fluorescence correlation spectroscopy. Nat. Med. 1998, 4, 832. [Google Scholar] [CrossRef]
- McLean, C.A.; Cherny, R.A.; Fraser, F.W.; Fuller, S.J.; Smith, M.J.; Vbeyreuther, K.; Bush, A.I.; Masters, C.L. Soluble pool of Aβ amyloid as a determinant of severity of neurodegeneration in Alzheimer’s disease. Ann. Neurol. 1999, 46, 860–866. [Google Scholar] [CrossRef]
- Näslund, J.; Haroutunian, V.; Mohs, R.; Davis, K.L.; Davies, P.; Greengard, P.; Buxbaum, J.D. Correlation between elevated levels of amyloid β-peptide in the brain and cognitive decline. JAMA 2000, 283, 1571–1577. [Google Scholar] [CrossRef]
- Hardy, J.; Selkoe, D.J. The amyloid hypothesis of Alzheimer’s disease: Progress and problems on the road to therapeutics. Science 2002, 297, 353–356. [Google Scholar] [CrossRef]
- Kumar, A.; Srivastava, S.; Tripathi, S.; Singh, S.K.; Srikrishna, S.; Sharma, A. Molecular insight into amyloid oligomer destabilizing mechanism of flavonoid derivative 2-(4‘-benzyloxyphenyl)-3-hydroxy-chromen-4-one through docking and molecular dynamics simulations. J. Biomol. Struct. Dyn. 2016, 34, 1252–1263. [Google Scholar] [CrossRef]
- Nerelius, C.; Sandegren, A.; Sargsyan, H.; Raunak, R.; Leijonmarck, H.; Chatterjee, U.; Fisahn, A.; Imarisio, S.; Lomas, D.; Crowther, D. α-Helix targeting reduces amyloid-β peptide toxicity. Proc. Natl. Acad. Sci. USA 2009, 106, 9191–9196. [Google Scholar] [CrossRef]
- Kinghorn, K.J.; Crowther, D.C.; Sharp, L.K.; Nerelius, C.; Davis, R.L.; Chang, H.T.; Green, C.; Gubb, D.C.; Johansson, J.; Lomas, D.A. Neuroserpin binds Aβ and is a neuroprotective component of amyloid plaques in Alzheimer disease. J. Biol. Chem. 2006, 281, 29268–29277. [Google Scholar] [CrossRef]
- Martín-Peña, A.; Rincón-Limas, D.E.; Fernandez-Fúnez, P. Engineered Hsp70 chaperones prevent Aβ42-induced memory impairments in a Drosophila model of Alzheimer’s disease. Sci. Rep. 2018, 8, 9915. [Google Scholar] [CrossRef]
- Fernandez-Funez, P.; Sanchez-Garcia, J.; de Mena, L.; Zhang, Y.; Levites, Y.; Khare, S.; Golde, T.E.; Rincon-Limas, D.E. Holdase activity of secreted Hsp70 masks amyloid-beta42 neurotoxicity in Drosophila. Proc. Natl. Acad. Sci. USA 2016, 113, E5212–E5221. [Google Scholar] [CrossRef]
- Younan, N.D.; Chen, K.-F.; Rose, R.-S.; Crowther, D.C.; Viles, J.H. Prion protein stabilizes amyloid-β (Aβ) oligomers and enhances Aβ neurotoxicity in a Drosophila model of Alzheimer’s disease. J. Biol. Chem. 2018, 293, 13090–13099. [Google Scholar] [CrossRef]
- Yamasaki, Y.; Tsuda, L.; Suzuki, A.; Yanagisawa, K. Induction of ganglioside synthesis in Drosophila brain accelerates assembly of amyloid β protein. Sci. Rep. 2018, 8, 8345. [Google Scholar] [CrossRef]
- Iwata, N.; Tsubuki, S.; Takaki, Y.; Watanabe, K.; Sekiguchi, M.; Hosoki, E.; Kawashima-Morishima, M.; Lee, H.-J.; Hama, E.; Sekine-Aizawa, Y. Identification of the major Aβ 1–42-degrading catabolic pathway in brain parenchyma: Suppression leads to biochemical and pathological deposition. Nat. Med. 2000, 6, 143. [Google Scholar] [CrossRef]
- Iijima-Ando, K.; Hearn, S.A.; Granger, L.; Shenton, C.; Gatt, A.; Chiang, H.-C.; Hakker, I.; Zhong, Y.; Iijima, K. Overexpression of neprilysin reduces alzheimer amyloid-β42 (Aβ42)-induced neuron loss and intraneuronal Aβ42 deposits but causes a reduction in cAMP-responsive element-binding protein-mediated transcription, age-dependent axon pathology, and premature death in Drosophila. J. Biol. Chem. 2008, 283, 19066–19076. [Google Scholar]
- Qiu, W.Q.; Walsh, D.M.; Ye, Z.; Vekrellis, K.; Zhang, J.; Podlisny, M.B.; Rosner, M.R.; Safavi, A.; Hersh, L.B.; Selkoe, D.J. Insulin-degrading enzyme regulates extracellular levels of amyloid β-protein by degradation. J. Biol. Chem. 1998, 273, 32730–32738. [Google Scholar] [CrossRef]
- Tsuda, M.; Kobayashi, T.; Matsuo, T.; Aigaki, T. Insulin-degrading enzyme antagonizes insulin-dependent tissue growth and Aβ-induced neurotoxicity in Drosophila. FEBS Lett. 2010, 584, 2916–2920. [Google Scholar] [CrossRef]
- Shen, Y.; Xia, Y.; Meng, S.; Lim, N.K.; Wang, W.; Huang, F. SH2B1 is involved in the accumulation of amyloid-β 42 in Alzheimer’s disease. J. Alzheimers Dis. 2017, 55, 835–847. [Google Scholar] [CrossRef]
- O’Neill, C. PI3-kinase/Akt/mTOR signaling: Impaired on/off switches in aging, cognitive decline and Alzheimer’s disease. Exp. Gerontol. 2013, 48, 647–653. [Google Scholar] [CrossRef]
- Boland, B.; Kumar, A.; Lee, S.; Platt, F.M.; Wegiel, J.; Yu, W.H.; Nixon, R.A. Autophagy induction and autophagosome clearance in neurons: Relationship to autophagic pathology in Alzheimer’s disease. J. Neurosci. 2008, 28, 6926–6937. [Google Scholar] [CrossRef]
- Chiang, H.-C.; Wang, L.; Xie, Z.; Yau, A.; Zhong, Y. PI3 kinase signaling is involved in Aβ-induced memory loss in Drosophila. Proc. Natl. Acad. Sci. USA 2010, 107, 7060–7065. [Google Scholar] [CrossRef]
- Gerenu, G.; Persson, T.; Goikolea, J.; Calvo-Garrido, J.; Loera-Valencia, R.; Pottmeier, P.; Santiago, C.; Poska, H.; Presto, J.; Cedazo-Minguez, A. Thioredoxin-80 protects against amyloid-beta pathology through autophagic-lysosomal pathway regulation. Mol. Psychiatry 2019, 1–14. [Google Scholar] [CrossRef]
- Demirev, A.V.; Song, H.-L.; Cho, M.-H.; Cho, K.; Peak, J.-J.; Yoo, H.J.; Kim, D.-H.; Yoon, S.-Y. V232M substitution restricts a distinct O-glycosylation of PLD3 and its neuroprotective function. Neurobiol. Dis. 2019, 129, 182–194. [Google Scholar] [CrossRef]
- LaFerla, F.M.; Green, K.N.; Oddo, S. Intracellular amyloid-β in Alzheimer’s disease. Nat. Rev. Neurosci. 2007, 8, 499. [Google Scholar] [CrossRef]
- Gouras, G.K.; Tampellini, D.; Takahashi, R.H.; Capetillo-Zarate, E. Intraneuronal β-amyloid accumulation and synapse pathology in Alzheimer’s disease. Acta Neuropathol. 2010, 119, 523–541. [Google Scholar] [CrossRef]
- Hong, S.; Ostaszewski, B.L.; Yang, T.; O’Malley, T.T.; Jin, M.; Yanagisawa, K.; Li, S.; Bartels, T.; Selkoe, D.J. Soluble Aβ oligomers are rapidly sequestered from brain ISF in vivo and bind GM1 ganglioside on cellular membranes. Neuron 2014, 82, 308–319. [Google Scholar] [CrossRef]
- McLaurin, J.; Chakrabartty, A. Characterization of the interactions of Alzheimer β-amyloid peptides with phospholipid membranes. Eur. J. Biochem. 1997, 245, 355–363. [Google Scholar] [CrossRef]
- McIntire, L.B.J.; Berman, D.E.; Myaeng, J.; Staniszewski, A.; Arancio, O.; Di Paolo, G.; Kim, T.-W. Reduction of synaptojanin 1 ameliorates synaptic and behavioral impairments in a mouse model of Alzheimer’s disease. J. Neurosci. 2012, 32, 15271–15276. [Google Scholar] [CrossRef]
- Zhu, L.; Zhong, M.; Zhao, J.; Rhee, H.; Caesar, I.; Knight, E.M.; Volpicelli-Daley, L.; Bustos, V.; Netzer, W.; Liu, L. Reduction of synaptojanin 1 accelerates Aβ clearance and attenuates cognitive deterioration in an Alzheimer mouse model. J. Biol. Chem. 2013, 288, 32050–32063. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, W.-A.; Jiang, L.-X.; Liu, H.-Y.; Zhang, B.-Z.; Lim, N.; Li, Q.-Y.; Huang, F.-D. Downregulation of RBO-PI4KIIIα facilitates Aβ42 secretion and ameliorates neural deficits in Aβ42-expressing Drosophila. J. Neurosci. 2017, 37, 4928–4941. [Google Scholar] [CrossRef]
- Sun, M.; Zhao, Y.; Han, M.; Zhang, B.; Zhang, X.; Zhang, Q.; Lim, N.K.-H.; Wang, W.-A.; Huang, F.-D. TTC7 and Hyccin regulate neuronal Aβ 42 accumulation and its associated neural deficits in Aβ 42-expressing Drosophila. J. Alzheimers Dis. 2018, 65, 1–10. [Google Scholar] [CrossRef]
- Belfiori-Carrasco, L.F.; Marcora, M.S.; Bocai, N.I.; Ceriani, M.F.; Morelli, L.; Castaño, E.M. A novel genetic screen identifies modifiers of age-dependent amyloid β toxicity in the Drosophila brain. Front. Aging Neurosci. 2017, 9, 61. [Google Scholar] [CrossRef][Green Version]
- Heidary, G.; Fortini, M.E. Identification and characterization of the Drosophila tau homolog. Mech. Dev. 2001, 108, 171–178. [Google Scholar] [CrossRef]
- Fulga, T.A.; Elson-Schwab, I.; Khurana, V.; Steinhilb, M.L.; Spires, T.L.; Hyman, B.T.; Feany, M.B. Abnormal bundling and accumulation of F-actin mediates tau-induced neuronal degeneration in vivo. Nat. Cell Biol. 2007, 9, 139. [Google Scholar] [CrossRef]
- Iijima, K.; Gatt, A.; Iijima-Ando, K. Tau Ser262 phosphorylation is critical for Aβ42-induced tau toxicity in a transgenic Drosophila model of Alzheimer’s disease. Hum. Mol. Genet. 2010, 19, 2947–2957. [Google Scholar] [CrossRef]
- Ando, K.; Maruko-Otake, A.; Ohtake, Y.; Hayashishita, M.; Sekiya, M.; Iijima, K.M. Stabilization of microtubule-unbound tau via tau phosphorylation at Ser262/356 by Par-1/MARK contributes to augmentation of AD-related phosphorylation and Aβ42-induced tau toxicity. PLoS Genet. 2016, 12, e1005917. [Google Scholar] [CrossRef]
- Götz, J.; Chen, F.; Van Dorpe, J.; Nitsch, R. Formation of neurofibrillary tangles in P301L tau transgenic mice induced by Aβ42 fibrils. Science 2001, 293, 1491–1495. [Google Scholar] [CrossRef]
- Lewis, J.; Dickson, D.W.; Lin, W.-L.; Chisholm, L.; Corral, A.; Jones, G.; Yen, S.-H.; Sahara, N.; Skipper, L.; Yager, D. Enhanced neurofibrillary degeneration in transgenic mice expressing mutant tau and APP. Science 2001, 293, 1487–1491. [Google Scholar] [CrossRef]
- Pennanen, L.; Götz, J. Different tau epitopes define Aβ 42-mediated tau insolubility. Biochem. Biophys. Res. Commun. 2005, 337, 1097–1101. [Google Scholar] [CrossRef]
- Oddo, S.; Caccamo, A.; Cheng, D.; Jouleh, B.; Torp, R.; LaFerla, F.M. Genetically augmenting tau levels does not modulate the onset or progression of Aβ pathology in transgenic mice. J. Neurochem. 2007, 102, 1053–1063. [Google Scholar] [CrossRef]
- Roberson, E.D.; Scearce-Levie, K.; Palop, J.J.; Yan, F.; Cheng, I.H.; Wu, T.; Gerstein, H.; Yu, G.Q.; Mucke, L. Reducing endogenous tau ameliorates amyloid beta-induced deficits in an Alzheimer’s disease mouse model. Science 2007, 316, 750–754. [Google Scholar] [CrossRef]
- Hu, S.; Begum, A.N.; Jones, M.R.; Oh, M.S.; Beech, W.K.; Beech, B.H.; Yang, F.; Chen, P.; Ubeda, O.J.; Kim, P.C.; et al. GSK3 inhibitors show benefits in an Alzheimer’s disease (AD) model of neurodegeneration but adverse effects in control animals. Neurobiol. Dis. 2009, 33, 193–206. [Google Scholar] [CrossRef]
- Takashima, A.; Murayama, M.; Murayama, O.; Kohno, T.; Honda, T.; Yasutake, K.; Nihonmatsu, N.; Mercken, M.; Yamaguchi, H.; Sugihara, S.; et al. Presenilin 1 associates with glycogen synthase kinase-3beta and its substrate tau. Proc. Natl. Acad. Sci. USA 1998, 95, 9637–9641. [Google Scholar] [CrossRef]
- Song, M.S.; Rauw, G.; Baker, G.B.; Kar, S. Memantine protects rat cortical cultured neurons against beta-amyloid-induced toxicity by attenuating tau phosphorylation. Eur. J. Neurosci. 2008, 28, 1989–2002. [Google Scholar] [CrossRef]
- Yang, T.; Knowles, J.K.; Lu, Q.; Zhang, H.; Arancio, O.; Moore, L.A.; Chang, T.; Wang, Q.; Andreasson, K.; Rajadas, J.; et al. Small molecule, non-peptide p75 ligands inhibit Abeta-induced neurodegeneration and synaptic impairment. PLoS ONE 2008, 3, e3604. [Google Scholar] [CrossRef]
- Noh, M.Y.; Koh, S.H.; Kim, Y.; Kim, H.Y.; Cho, G.W.; Kim, S.H. Neuroprotective effects of donepezil through inhibition of GSK-3 activity in amyloid-beta-induced neuronal cell death. J. Neurochem. 2009, 108, 1116–1125. [Google Scholar] [CrossRef]
- Drewes, G.; Trinczek, B.; Illenberger, S.; Biernat, J.; Schmitt-Ulms, G.; Meyer, H.E.; Mandelkow, E.M.; Mandelkow, E. Microtubule-associated protein/microtubule affinity-regulating kinase (p110mark). A novel protein kinase that regulates tau-microtubule interactions and dynamic instability by phosphorylation at the Alzheimer-specific site serine 262. J. Biol. Chem. 1995, 270, 7679–7688. [Google Scholar] [CrossRef]
- Drewes, G.; Ebneth, A.; Preuss, U.; Mandelkow, E.M.; Mandelkow, E. MARK, a novel family of protein kinases that phosphorylate microtubule-associated proteins and trigger microtubule disruption. Cell 1997, 89, 297–308. [Google Scholar] [CrossRef]
- Hasegawa, M.; Morishima-Kawashima, M.; Takio, K.; Suzuki, M.; Titani, K.; Ihara, Y. Protein sequence and mass spectrometric analyses of tau in the Alzheimer’s disease brain. J. Biol. Chem. 1992, 267, 17047–17054. [Google Scholar]
- Hanger, D.P.; Brion, J.P.; Gallo, J.M.; Cairns, N.J.; Luthert, P.J.; Anderton, B.H. Tau in Alzheimer’s disease and Down’s syndrome is insoluble and abnormally phosphorylated. Biochem. J. 1991, 275 Pt 1, 99–104. [Google Scholar] [CrossRef]
- Hanger, D.P.; Betts, J.C.; Loviny, T.L.; Blackstock, W.P.; Anderton, B.H. New phosphorylation sites identified in hyperphosphorylated tau (paired helical filament-tau) from Alzheimer’s disease brain using nanoelectrospray mass spectrometry. J. Neurochem. 1998, 71, 2465–2476. [Google Scholar] [CrossRef]
- Morishima-Kawashima, M.; Hasegawa, M.; Takio, K.; Suzuki, M.; Yoshida, H.; Titani, K.; Ihara, Y. Proline-directed and non-proline-directed phosphorylation of PHF-tau. J. Biol. Chem. 1995, 270, 823–829. [Google Scholar] [CrossRef]
- Sperber, B.R.; Leight, S.; Goedert, M.; Lee, V.M. Glycogen synthase kinase-3 beta phosphorylates tau protein at multiple sites in intact cells. Neurosci. Lett. 1995, 197, 149–153. [Google Scholar] [CrossRef]
- Singh, T.J.; Haque, N.; Grundke-Iqbal, I.; Iqbal, K. Rapid Alzheimer-like phosphorylation of tau by the synergistic actions of non-proline-dependent protein kinases and GSK-3. FEBS Lett. 1995, 358, 267–272. [Google Scholar] [CrossRef]
- Singh, T.J.; Wang, J.Z.; Novak, M.; Kontzekova, E.; Grundke-Iqbal, I.; Iqbal, K. Calcium/calmodulin-dependent protein kinase II phosphorylates tau at Ser-262 but only partially inhibits its binding to microtubules. FEBS Lett. 1996, 387, 145–148. [Google Scholar] [CrossRef]
- Sengupta, A.; Kabat, J.; Novak, M.; Wu, Q.; Grundke-Iqbal, I.; Iqbal, K. Phosphorylation of tau at both Thr 231 and Ser 262 is required for maximal inhibition of its binding to microtubules. Arch. Biochem. Biophys. 1998, 357, 299–309. [Google Scholar] [CrossRef]
- Nishimura, I.; Yang, Y.; Lu, B. PAR-1 kinase plays an initiator role in a temporally ordered phosphorylation process that confers tau toxicity in Drosophila. Cell 2004, 116, 671–682. [Google Scholar] [CrossRef]
- Kosmidis, S.; Grammenoudi, S.; Papanikolopoulou, K.; Skoulakis, E.M. Differential effects of Tau on the integrity and function of neurons essential for learning in Drosophila. J. Neurosci. 2010, 30, 464–477. [Google Scholar] [CrossRef]
- Leroy, K.; Ando, K.; Laporte, V.; Dedecker, R.; Suain, V.; Authelet, M.; Heraud, C.; Pierrot, N.; Yilmaz, Z.; Octave, J.N.; et al. Lack of tau proteins rescues neuronal cell death and decreases amyloidogenic processing of APP in APP/PS1 mice. Am. J. Pathol. 2012, 181, 1928–1940. [Google Scholar] [CrossRef]
- Sofola, O.; Kerr, F.; Rogers, I.; Killick, R.; Augustin, H.; Gandy, C.; Allen, M.J.; Hardy, J.; Lovestone, S.; Partridge, L. Inhibition of GSK-3 ameliorates Abeta pathology in an adult-onset Drosophila model of Alzheimer’s disease. PLoS Genet. 2010, 6, e1001087. [Google Scholar] [CrossRef]
- Burnouf, S.; Gronke, S.; Augustin, H.; Dols, J.; Gorsky, M.K.; Werner, J.; Kerr, F.; Alic, N.; Martinez, P.; Partridge, L. Deletion of endogenous Tau proteins is not detrimental in Drosophila. Sci. Rep. 2016, 6, 23102. [Google Scholar] [CrossRef]
- Klein, J.A.; Ackerman, S.L. Oxidative stress, cell cycle, and neurodegeneration. J. Clin. Investig. 2003, 111, 785–793. [Google Scholar] [CrossRef]
- Moreira, P.I.; Santos, M.S.; Oliveira, C.R.; Shenk, J.C.; Nunomura, A.; Smith, M.A.; Zhu, X.; Perry, G. Alzheimer disease and the role of free radicals in the pathogenesis of the disease. CNS Neurol. Disord. Drug Targets 2008, 7, 3–10. [Google Scholar]
- Eckert, A.; Schmitt, K.; Gotz, J. Mitochondrial dysfunction—The beginning of the end in Alzheimer’s disease? Separate and synergistic modes of tau and amyloid-beta toxicity. Alzheimers Res. Ther. 2011, 3, 15. [Google Scholar] [CrossRef]
- Rival, T.; Page, R.M.; Chandraratna, D.S.; Sendall, T.J.; Ryder, E.; Liu, B.; Lewis, H.; Rosahl, T.; Hider, R.; Camargo, L.M.; et al. Fenton chemistry and oxidative stress mediate the toxicity of the beta-amyloid peptide in a Drosophila model of Alzheimer’s disease. Eur. J. Neurosci. 2009, 29, 1335–1347. [Google Scholar] [CrossRef]
- Ott, S.; Dziadulewicz, N.; Crowther, D.C. Iron is a specific cofactor for distinct oxidation- and aggregation-dependent Abeta toxicity mechanisms in a Drosophila model. Dis. Model. Mech. 2015, 8, 657–667. [Google Scholar] [CrossRef]
- Liu, B.; Moloney, A.; Meehan, S.; Morris, K.; Thomas, S.E.; Serpell, L.C.; Hider, R.; Marciniak, S.J.; Lomas, D.A.; Crowther, D.C. Iron promotes the toxicity of amyloid beta peptide by impeding its ordered aggregation. J. Biol. Chem. 2011, 286, 4248–4256. [Google Scholar] [CrossRef]
- Dias-Santagata, D.; Fulga, T.A.; Duttaroy, A.; Feany, M.B. Oxidative stress mediates tau-induced neurodegeneration in Drosophila. J. Clin. Investig. 2007, 117, 236–245. [Google Scholar] [CrossRef]
- Lee, S.; Bang, S.M.; Hong, Y.K.; Lee, J.H.; Jeong, H.; Park, S.H.; Liu, Q.F.; Lee, I.S.; Cho, K.S. The calcineurin inhibitor Sarah (Nebula) exacerbates Abeta42 phenotypes in a Drosophila model of Alzheimer’s disease. Dis. Model. Mech. 2016, 9, 295–306. [Google Scholar] [CrossRef]
- Favrin, G.; Bean, D.M.; Bilsland, E.; Boyer, H.; Fischer, B.E.; Russell, S.; Crowther, D.C.; Baylis, H.A.; Oliver, S.G.; Giannakou, M.E. Identification of novel modifiers of Abeta toxicity by transcriptomic analysis in the fruitfly. Sci. Rep. 2013, 3, 3512. [Google Scholar] [CrossRef]
- Muller, F.L.; Lustgarten, M.S.; Jang, Y.; Richardson, A.; Van Remmen, H. Trends in oxidative aging theories. Free Radic. Biol. Med. 2007, 43, 477–503. [Google Scholar] [CrossRef]
- Jung, I.; Kim, T.Y.; Kim-Ha, J. Identification of Drosophila SOD3 and its protective role against phototoxic damage to cells. FEBS Lett. 2011, 585, 1973–1978. [Google Scholar] [CrossRef]
- Bush, A.I. The metallobiology of Alzheimer’s disease. Trends Neurosci. 2003, 26, 207–214. [Google Scholar] [CrossRef]
- Lang, M.; Wang, L.; Fan, Q.; Xiao, G.; Wang, X.; Zhong, Y.; Zhou, B. Genetic inhibition of solute-linked carrier 39 family transporter 1 ameliorates abeta pathology in a Drosophila model of Alzheimer’s disease. PLoS Genet. 2012, 8, e1002683. [Google Scholar] [CrossRef]
- Lang, M.; Fan, Q.; Wang, L.; Zheng, Y.; Xiao, G.; Wang, X.; Wang, W.; Zhong, Y.; Zhou, B. Inhibition of human high-affinity copper importer Ctr1 orthologous in the nervous system of Drosophila ameliorates Abeta42-induced Alzheimer’s disease-like symptoms. Neurobiol. Aging 2013, 34, 2604–2612. [Google Scholar] [CrossRef]
- Lindholm, D.; Wootz, H.; Korhonen, L. ER stress and neurodegenerative diseases. Cell Death Differ. 2006, 13, 385. [Google Scholar] [CrossRef]
- Hitomi, J.; Katayama, T.; Eguchi, Y.; Kudo, T.; Taniguchi, M.; Koyama, Y.; Manabe, T.; Yamagishi, S.; Bando, Y.; Imaizumi, K.; et al. Involvement of caspase-4 in endoplasmic reticulum stress-induced apoptosis and Abeta-induced cell death. J. Cell Biol. 2004, 165, 347–356. [Google Scholar] [CrossRef]
- Costa, R.O.; Ferreiro, E.; Cardoso, S.M.; Oliveira, C.R.; Pereira, C.M. ER stress-mediated apoptotic pathway induced by Abeta peptide requires the presence of functional mitochondria. J. Alzheimers Dis. 2010, 20, 625–636. [Google Scholar] [CrossRef]
- Costa, R.O.; Ferreiro, E.; Oliveira, C.R.; Pereira, C.M. Inhibition of mitochondrial cytochrome c oxidase potentiates Abeta-induced ER stress and cell death in cortical neurons. Mol. Cell. Neurosci. 2013, 52, 1–8. [Google Scholar] [CrossRef]
- Kang, E.B.; Kwon, I.S.; Koo, J.H.; Kim, E.J.; Kim, C.H.; Lee, J.; Yang, C.H.; Lee, Y.I.; Cho, I.H.; Cho, J.Y. Treadmill exercise represses neuronal cell death and inflammation during Abeta-induced ER stress by regulating unfolded protein response in aged presenilin 2 mutant mice. Apoptosis 2013, 18, 1332–1347. [Google Scholar] [CrossRef]
- Ron, D.; Walter, P. Signal integration in the endoplasmic reticulum unfolded protein response. Nat. Rev. Mol. Cell Biol. 2007, 8, 519–529. [Google Scholar] [CrossRef]
- Marcora, M.S.; Belfiori-Carrasco, L.F.; Bocai, N.I.; Morelli, L.; Castano, E.M. Amyloid-beta42 clearance and neuroprotection mediated by X-box binding protein 1 signaling decline with aging in the Drosophila brain. Neurobiol. Aging 2017, 60, 57–70. [Google Scholar] [CrossRef]
- Perry, G.; Roder, H.; Nunomura, A.; Takeda, A.; Friedlich, A.L.; Zhu, X.; Raina, A.K.; Holbrook, N.; Siedlak, S.L.; Harris, P.L.; et al. Activation of neuronal extracellular receptor kinase (ERK) in Alzheimer disease links oxidative stress to abnormal phosphorylation. Neuroreport 1999, 10, 2411–2415. [Google Scholar] [CrossRef]
- Zhu, X.; Castellani, R.J.; Takeda, A.; Nunomura, A.; Atwood, C.S.; Perry, G.; Smith, M.A. Differential activation of neuronal ERK, JNK/SAPK and p38 in Alzheimer disease: The ‘two hit’ hypothesis. Mech. Ageing Dev. 2001, 123, 39–46. [Google Scholar] [CrossRef]
- Dineley, K.T.; Westerman, M.; Bui, D.; Bell, K.; Ashe, K.H.; Sweatt, J.D. Beta-amyloid activates the mitogen-activated protein kinase cascade via hippocampal alpha7 nicotinic acetylcholine receptors: In vitro and in vivo mechanisms related to Alzheimer’s disease. J. Neurosci. 2001, 21, 4125–4133. [Google Scholar] [CrossRef]
- Ma, Q.L.; Harris-White, M.E.; Ubeda, O.J.; Simmons, M.; Beech, W.; Lim, G.P.; Teter, B.; Frautschy, S.A.; Cole, G.M. Evidence of Abeta- and transgene-dependent defects in ERK-CREB signaling in Alzheimer’s models. J. Neurochem. 2007, 103, 1594–1607. [Google Scholar] [CrossRef]
- Zhu, X.; Lee, H.-G.; Raina, A.K.; Perry, G.; Smith, M.A. The role of mitogen-activated protein kinase pathways in Alzheimer’s disease. Neurosignals 2002, 11, 270–281. [Google Scholar] [CrossRef]
- Cheung, E.C.; Slack, R.S. Emerging role for ERK as a key regulator of neuronal apoptosis. Sci. STKE. 2004, 2004, pe45. [Google Scholar] [CrossRef]
- Park, S.H.; Lee, S.; Hong, Y.K.; Hwang, S.; Lee, J.H.; Bang, S.M.; Kim, Y.K.; Koo, B.S.; Lee, I.S.; Cho, K.S. Suppressive effects of SuHeXiang Wan on amyloid-beta42-induced extracellular signal-regulated kinase hyperactivation and glial cell proliferation in a transgenic Drosophila model of Alzheimer’s disease. Biol. Pharm. Bull. 2013, 36, 390–398. [Google Scholar] [CrossRef]
- Xia, Z.; Dickens, M.; Raingeaud, J.; Davis, R.J.; Greenberg, M.E. Opposing effects of ERK and JNK-p38 MAP kinases on apoptosis. Science 1995, 270, 1326–1331. [Google Scholar] [CrossRef]
- Cobb, M.H. MAP kinase pathways. Prog. Biophys. Mol. Biol. 1999, 71, 479–500. [Google Scholar] [CrossRef]
- Liu, Q.F.; Jeong, H.; Lee, J.H.; Hong, Y.K.; Oh, Y.; Kim, Y.M.; Suh, Y.S.; Bang, S.; Yun, H.S.; Lee, K.; et al. Coriandrum sativum suppresses Abeta42-Induced ROS increases, glial cell proliferation, and ERK activation. Am. J. Chin. Med. 2016, 44, 1325–1347. [Google Scholar] [CrossRef]
- Liu, Q.F.; Jeon, Y.; Sung, Y.W.; Lee, J.H.; Jeong, H.; Kim, Y.M.; Yun, H.S.; Chin, Y.W.; Jeon, S.; Cho, K.S.; et al. Nardostachys jatamansi ethanol extract ameliorates Abeta42 cytotoxicity. Biol. Pharm. Bull. 2018, 41, 470–477. [Google Scholar] [CrossRef]
- Rahn, T.; Leippe, M.; Roeder, T.; Fedders, H. EGFR signaling in the brain is necessary for olfactory learning in Drosophila larvae. Learn. Mem. 2013, 20, 194–200. [Google Scholar] [CrossRef][Green Version]
- Wang, L.; Chiang, H.C.; Wu, W.; Liang, B.; Xie, Z.; Yao, X.; Ma, W.; Du, S.; Zhong, Y. Epidermal growth factor receptor is a preferred target for treating amyloid-beta-induced memory loss. Proc. Natl. Acad. Sci. USA 2012, 109, 16743–16748. [Google Scholar] [CrossRef]
- Herrup, K. Post-mitotic role of the cell cycle machinery. Curr. Opin. Cell Biol. 2013, 25, 711–716. [Google Scholar] [CrossRef]
- Lee, H.G.; Casadesus, G.; Zhu, X.; Castellani, R.J.; McShea, A.; Perry, G.; Petersen, R.B.; Bajic, V.; Smith, M.A. Cell cycle re-entry mediated neurodegeneration and its treatment role in the pathogenesis of Alzheimer’s disease. Neurochem. Int. 2009, 54, 84–88. [Google Scholar] [CrossRef]
- Crews, L.; Rockenstein, E.; Masliah, E. APP transgenic modeling of Alzheimer’s disease: Mechanisms of neurodegeneration and aberrant neurogenesis. Brain Struct. Funct. 2010, 214, 111–126. [Google Scholar] [CrossRef]
- Varvel, N.H.; Bhaskar, K.; Patil, A.R.; Pimplikar, S.W.; Herrup, K.; Lamb, B.T. Abeta oligomers induce neuronal cell cycle events in Alzheimer’s disease. J. Neurosci. 2008, 28, 10786–10793. [Google Scholar] [CrossRef]
- Seward, M.E.; Swanson, E.; Norambuena, A.; Reimann, A.; Cochran, J.N.; Li, R.; Roberson, E.D.; Bloom, G.S. Amyloid-beta signals through tau to drive ectopic neuronal cell cycle re-entry in Alzheimer’s disease. J. Cell Sci. 2013, 126, 1278–1286. [Google Scholar] [CrossRef]
- Kong, Y.; Li, K.; Fu, T.; Wan, C.; Zhang, D.; Song, H.; Zhang, Y.; Liu, N.; Gan, Z.; Yuan, L. Quercetin ameliorates Abeta toxicity in Drosophila AD model by modulating cell cycle-related protein expression. Oncotarget 2016, 7, 67716–67731. [Google Scholar] [CrossRef] [PubMed]
- Peng, F.; Zhao, Y.; Huang, X.; Chen, C.; Sun, L.; Zhuang, L.; Xue, L. Loss of Polo ameliorates APP-induced Alzheimer’s disease-like symptoms in Drosophila. Sci. Rep. 2015, 5, 16816. [Google Scholar] [CrossRef] [PubMed]
- Alberi, L.; Hoey, S.E.; Brai, E.; Scotti, A.L.; Marathe, S. Notch signaling in the brain: In good and bad times. Ageing Res. Rev. 2013, 12, 801–814. [Google Scholar] [CrossRef] [PubMed]
- Marathe, S.; Liu, S.; Brai, E.; Kaczarowski, M.; Alberi, L. Notch signaling in response to excitotoxicity induces neurodegeneration via erroneous cell cycle reentry. Cell Death Differ. 2015, 22, 1775–1784. [Google Scholar] [CrossRef]
- Kong, Y.; Wu, J.; Zhang, D.; Wan, C.; Yuan, L. The Role of miR-124 in Drosophila Alzheimer’s Disease Model by Targeting Delta in Notch Signaling Pathway. Curr. Mol. Med. 2015, 15, 980–989. [Google Scholar] [CrossRef]
- Roth, K.A. Caspases, apoptosis, and Alzheimer disease: Causation, correlation, and confusion. J. Neuropathol. Exp. Neurol. 2001, 60, 829–838. [Google Scholar] [CrossRef]
- Wu, S.C.; Cao, Z.S.; Chang, K.M.; Juang, J.L. Intestinal microbial dysbiosis aggravates the progression of Alzheimer’s disease in Drosophila. Nat. Commun. 2017, 8, 24. [Google Scholar] [CrossRef]
- Hong, Y.K.; Lee, S.; Park, S.H.; Lee, J.H.; Han, S.Y.; Kim, S.T.; Kim, Y.K.; Jeon, S.; Koo, B.S.; Cho, K.S. Inhibition of JNK/dFOXO pathway and caspases rescues neurological impairments in Drosophila Alzheimer’s disease model. Biochem. Biophys. Res. Commun. 2012, 419, 49–53. [Google Scholar] [CrossRef]
- Hawkins, C.J.; Yoo, S.J.; Peterson, E.P.; Wang, S.L.; Vernooy, S.Y.; Hay, B.A. The Drosophila caspase DRONC cleaves following glutamate or aspartate and is regulated by DIAP1, HID, and GRIM. J. Biol. Chem. 2000, 275, 27084–27093. [Google Scholar]
- Meier, P.; Silke, J.; Leevers, S.J.; Evan, G.I. The Drosophila caspase DRONC is regulated by DIAP1. EMBO J. 2000, 19, 598–611. [Google Scholar] [CrossRef]
- Yu, S.Y.; Yoo, S.J.; Yang, L.; Zapata, C.; Srinivasan, A.; Hay, B.A.; Baker, N.E. A pathway of signals regulating effector and initiator caspases in the developing Drosophila eye. Development 2002, 129, 3269–3278. [Google Scholar] [PubMed]
- Lin, R.; Angelin, A.; Da Settimo, F.; Martini, C.; Taliani, S.; Zhu, S.; Wallace, D.C. Genetic analysis of dTSPO, an outer mitochondrial membrane protein, reveals its functions in apoptosis, longevity, and Ab42-induced neurodegeneration. Aging Cell 2014, 13, 507–518. [Google Scholar] [CrossRef] [PubMed]
- Goedert, M.; Hasegawa, M.; Jakes, R.; Lawler, S.; Cuenda, A.; Cohen, P. Phosphorylation of microtubule-associated protein tau by stress-activated protein kinases. FEBS Lett. 1997, 409, 57–62. [Google Scholar] [CrossRef]
- Reynolds, C.H.; Betts, J.C.; Blackstock, W.P.; Nebreda, A.R.; Anderton, B.H. Phosphorylation sites on tau identified by nanoelectrospray mass spectrometry: Differences in vitro between the mitogen-activated protein kinases ERK2, c-Jun N-terminal kinase and P38, and glycogen synthase kinase-3beta. J. Neurochem. 2000, 74, 1587–1595. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Rottkamp, C.A.; Boux, H.; Takeda, A.; Perry, G.; Smith, M.A. Activation of p38 kinase links tau phosphorylation, oxidative stress, and cell cycle-related events in Alzheimer disease. J. Neuropathol. Exp. Neurol. 2000, 59, 880–888. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Raina, A.K.; Rottkamp, C.A.; Aliev, G.; Perry, G.; Boux, H.; Smith, M.A. Activation and redistribution of c-jun N-terminal kinase/stress activated protein kinase in degenerating neurons in Alzheimer’s disease. J. Neurochem. 2001, 76, 435–441. [Google Scholar] [CrossRef]
- Song, Q.; Feng, G.; Huang, Z.; Chen, X.; Chen, Z.; Ping, Y. Aberrant axonal arborization of PDF neurons induced by Abeta42-mediated JNK activation underlies sleep disturbance in an Alzheimer’s model. Mol. Neurobiol. 2017, 54, 6317–6328. [Google Scholar] [CrossRef]
- Hong, Y.K.; Park, S.H.; Lee, S.; Hwang, S.; Lee, M.J.; Kim, D.; Lee, J.H.; Han, S.Y.; Kim, S.T.; Kim, Y.K.; et al. Neuroprotective effect of SuHeXiang Wan in Drosophila models of Alzheimer’s disease. J. Ethnopharmacol. 2011, 134, 1028–1032. [Google Scholar] [CrossRef]
- Wang, X.; Ma, Y.; Zhao, Y.; Chen, Y.; Hu, Y.; Chen, C.; Shao, Y.; Xue, L. APLP1 promotes dFoxO-dependent cell death in Drosophila. Apoptosis 2015, 20, 778–786. [Google Scholar] [CrossRef]
- Coelho, D.S.; Schwartz, S.; Merino, M.M.; Hauert, B.; Topfel, B.; Tieche, C.; Rhiner, C.; Moreno, E. Culling less fit neurons protects against amyloid-β-induced brain damage and cognitive and motor decline. Cell Rep. 2018, 25, 3661–3673. [Google Scholar] [CrossRef]
- Lord, J.; Cruchaga, C. The epigenetic landscape of Alzheimer’s disease. Nat. Neurosci. 2014, 17, 1138–1140. [Google Scholar] [CrossRef] [PubMed]
- Govindarajan, N.; Rao, P.; Burkhardt, S.; Sananbenesi, F.; Schlüter, O.M.; Bradke, F.; Lu, J.; Fischer, A. Reducing HDAC6 ameliorates cognitive deficits in a mouse model for Alzheimer’s disease. EMBO Mol. Med. 2013, 5, 52–63. [Google Scholar] [CrossRef] [PubMed]
- Ding, H.; Dolan, P.J.; Johnson, G.V. Histone deacetylase 6 interacts with the microtubule-associated protein tau. J. Neurochem. 2008, 106, 2119–2130. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.; Choi, H.; Jung, E.S.; Lee, W.; Oh, S.; Jeon, N.L.; Mook-Jung, I. HDAC6 inhibitor blocks amyloid beta-induced impairment of mitochondrial transport in hippocampal neurons. PLoS ONE 2012, 7, e42983. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; Zhao, K.; Wu, J.; Xu, Z.; Jin, S.; Zhang, Y.Q. HDAC6 mutations rescue human tau-induced microtubule defects in Drosophila. Proc. Natl. Acad. Sci. USA 2013, 110, 4604–4609. [Google Scholar] [CrossRef]
- Pile, L.; Lee, F.-H.; Wassarman, D. The histone deacetylase inhibitor trichostatin A influences the development of Drosophila melanogaster. Cell. Mol. Life. Sci. 2001, 58, 1715–1718. [Google Scholar] [CrossRef]
- Jeong, M.R.; Hashimoto, R.; Senatorov, V.V.; Fujimaki, K.; Ren, M.; Lee, M.S.; Chuang, D.-M. Valproic acid, a mood stabilizer and anticonvulsant, protects rat cerebral cortical neurons from spontaneous cell death: A role of histone deacetylase inhibition. FEBS Lett. 2003, 542, 74–78. [Google Scholar] [CrossRef]
- Cao, X.; Südhof, T.C. A transcriptively active complex of APP with Fe65 and histone acetyltransferase Tip60. Science 2001, 293, 115–120. [Google Scholar] [CrossRef]
- Baek, S.H.; Ohgi, K.A.; Rose, D.W.; Koo, E.H.; Glass, C.K.; Rosenfeld, M.G. Exchange of N-CoR corepressor and Tip60 coactivator complexes links gene expression by NF-κB and β-amyloid precursor protein. Cell 2002, 110, 55–67. [Google Scholar] [CrossRef]
- Panikker, P.; Xu, S.-J.; Zhang, H.; Sarthi, J.; Beaver, M.; Sheth, A.; Akhter, S.; Elefant, F. Restoring Tip60 HAT/HDAC2 balance in the neurodegenerative brain relieves epigenetic transcriptional repression and reinstates cognition. J. Neurosci. 2018, 38, 4569–4583. [Google Scholar] [CrossRef]
- Pirooznia, S.K.; Sarthi, J.; Johnson, A.A.; Toth, M.S.; Chiu, K.; Koduri, S.; Elefant, F. Tip60 HAT activity mediates APP induced lethality and apoptotic cell death in the CNS of a Drosophila Alzheimer’s disease model. PLoS ONE 2012, 7, e41776. [Google Scholar] [CrossRef] [PubMed]
- Cutler, T.; Sarkar, A.; Moran, M.; Steffensmeier, A.; Puli, O.R.; Mancini, G.; Tare, M.; Gogia, N.; Singh, A. Drosophila eye model to study neuroprotective role of CREB binding protein (CBP) in Alzheimer’s disease. PLoS ONE 2015, 10, e0137691. [Google Scholar] [CrossRef] [PubMed]
- Selkoe, D.J. Alzheimer’s disease is a synaptic failure. Science 2002, 298, 789–791. [Google Scholar] [CrossRef] [PubMed]
- Terry, R.D.; Masliah, E.; Salmon, D.P.; Butters, N.; DeTeresa, R.; Hill, R.; Hansen, L.A.; Katzman, R. Physical basis of cognitive alterations in Alzheimer’s disease: Synapse loss is the major correlate of cognitive impairment. Ann. Neurol. 1991, 30, 572–580. [Google Scholar] [CrossRef] [PubMed]
- Hsia, A.Y.; Masliah, E.; McConlogue, L.; Yu, G.-Q.; Tatsuno, G.; Hu, K.; Kholodenko, D.; Malenka, R.C.; Nicoll, R.A.; Mucke, L. Plaque-independent disruption of neural circuits in Alzheimer’s disease mouse models. Proc. Natl. Acad. Sci. USA 1999, 96, 3228–3233. [Google Scholar] [CrossRef] [PubMed]
- Lue, L.-F.; Kuo, Y.-M.; Roher, A.E.; Brachova, L.; Shen, Y.; Sue, L.; Beach, T.; Kurth, J.H.; Rydel, R.E.; Rogers, J. Soluble amyloid β peptide concentration as a predictor of synaptic change in Alzheimer’s disease. Am. J. Pathol. 1999, 155, 853–862. [Google Scholar] [CrossRef]
- Koistinaho, M.; Ort, M.; Cimadevilla, J.M.; Vondrous, R.; Cordell, B.; Koistinaho, J.; Bures, J.; Higgins, L.S. Specific spatial learning deficits become severe with age in β-amyloid precursor protein transgenic mice that harbor diffuse β-amyloid deposits but do not form plaques. Proc. Natl. Acad. Sci. USA 2001, 98, 14675–14680. [Google Scholar] [CrossRef]
- Kamenetz, F.; Tomita, T.; Hsieh, H.; Seabrook, G.; Borchelt, D.; Iwatsubo, T.; Sisodia, S.; Malinow, R. APP processing and synaptic function. Neuron 2003, 37, 925–937. [Google Scholar] [CrossRef]
- Townsend, M.; Shankar, G.M.; Mehta, T.; Walsh, D.M.; Selkoe, D.J. Effects of secreted oligomers of amyloid β-protein on hippocampal synaptic plasticity: A potent role for trimers. J. Physiol. 2006, 572, 477–492. [Google Scholar] [CrossRef]
- Moechars, D.; Dewachter, I.; Lorent, K.; Reversé, D.; Baekelandt, V.; Naidu, A.; Tesseur, I.; Spittaels, K.; Van Den Haute, C.; Checler, F. Early phenotypic changes in transgenic mice that overexpress different mutants of amyloid precursor protein in brain. J. Biol. Chem. 1999, 274, 6483–6492. [Google Scholar] [CrossRef]
- Chapman, P.F.; White, G.L.; Jones, M.W.; Cooper-Blacketer, D.; Marshall, V.J.; Irizarry, M.; Younkin, L.; Good, M.A.; Bliss, T.; Hyman, B.T. Impaired synaptic plasticity and learning in aged amyloid precursor protein transgenic mice. Nat. Neurosci. 1999, 2, 271. [Google Scholar] [CrossRef] [PubMed]
- Fitzjohn, S.M.; Morton, R.A.; Kuenzi, F.; Rosahl, T.W.; Shearman, M.; Lewis, H.; Smith, D.; Reynolds, D.S.; Davies, C.H.; Collingridge, G.L. Age-related impairment of synaptic transmission but normal long-term potentiation in transgenic mice that overexpress the human APP695SWE mutant form of amyloid precursor protein. J. Neurosci. 2001, 21, 4691–4698. [Google Scholar] [CrossRef] [PubMed]
- Fang, L.; Duan, J.; Ran, D.; Fan, Z.; Yan, Y.; Huang, N.; Gu, H.; Zhu, Y. Amyloid-β depresses excitatory cholinergic synaptic transmission in Drosophila. Neurosci. Bull. 2012, 28, 585–594. [Google Scholar] [CrossRef] [PubMed]
- Iijima, K.; Liu, H.-P.; Chiang, A.-S.; Hearn, S.A.; Konsolaki, M.; Zhong, Y. Dissecting the pathological effects of human Aβ40 and Aβ42 in Drosophila: A potential model for Alzheimer’s disease. Proc. Natl. Acad. Sci. USA 2004, 101, 6623–6628. [Google Scholar] [CrossRef]
- Klyubin, I.; Walsh, D.M.; Lemere, C.A.; Cullen, W.K.; Shankar, G.M.; Betts, V.; Spooner, E.T.; Jiang, L.; Anwyl, R.; Selkoe, D.J.; et al. Amyloid beta protein immunotherapy neutralizes Abeta oligomers that disrupt synaptic plasticity in vivo. Nat. Med. 2005, 11, 556–561. [Google Scholar] [CrossRef]
- Newey, S.E.; Velamoor, V.; Govek, E.E.; Van Aelst, L. Rho GTPases, dendritic structure, and mental retardation. J. Neurobiol. 2005, 64, 58–74. [Google Scholar] [CrossRef]
- Lee, T.; Winter, C.; Marticke, S.S.; Lee, A.; Luo, L. Essential roles of Drosophila RhoA in the regulation of neuroblast proliferation and dendritic but not axonal morphogenesis. Neuron 2000, 25, 307–316. [Google Scholar] [CrossRef]
- Nakayama, A.Y.; Harms, M.B.; Luo, L. Small GTPases Rac and Rho in the maintenance of dendritic spines and branches in hippocampal pyramidal neurons. J. Neurosci. 2000, 20, 5329–5338. [Google Scholar] [CrossRef]
- Huesa, G.; Baltrons, M.A.; Gómez-Ramos, P.; Morán, A.; García, A.; Hidalgo, J.; Francés, S.; Santpere, G.; Ferrer, I.; Galea, E. Altered distribution of RhoA in Alzheimer’s disease and AβPP overexpressing mice. J. Alzheimers Dis. 2010, 19, 37–56. [Google Scholar] [CrossRef]
- Cook, M.; Mani, P.; Wentzell, J.S.; Kretzschmar, D. Increased RhoA prenylation in the loechrig (loe) mutant leads to progressive neurodegeneration. PLoS ONE 2012, 7, e44440. [Google Scholar] [CrossRef]
- Wu, W.; Du, S.; Shi, W.; Liu, Y.; Hu, Y.; Xie, Z.; Yao, X.; Liu, Z.; Ma, W.; Xu, L. Inhibition of Rac1-dependent forgetting alleviates memory deficits in animal models of Alzheimer’s disease. Protein Cell. 2019, 10, 745–759. [Google Scholar] [CrossRef] [PubMed]
- Morfini, G.A.; Burns, M.; Binder, L.I.; Kanaan, N.M.; LaPointe, N.; Bosco, D.A.; Brown, R.H.; Brown, H.; Tiwari, A.; Hayward, L. Axonal transport defects in neurodegenerative diseases. J. Neurosci. 2009, 29, 12776–12786. [Google Scholar] [CrossRef] [PubMed]
- Gunawardena, S.; Goldstein, L.S. Disruption of axonal transport and neuronal viability by amyloid precursor protein mutations in Drosophila. Neuron 2001, 32, 389–401. [Google Scholar] [CrossRef]
- Stokin, G.B.; Lillo, C.; Falzone, T.L.; Brusch, R.G.; Rockenstein, E.; Mount, S.L.; Raman, R.; Davies, P.; Masliah, E.; Williams, D.S. Axonopathy and transport deficits early in the pathogenesis of Alzheimer’s disease. Science 2005, 307, 1282–1288. [Google Scholar] [CrossRef]
- Shaw, J.L.; Chang, K.T. Nebula/DSCR1 upregulation delays neurodegeneration and protects against APP-induced axonal transport defects by restoring calcineurin and GSK-3β signaling. PLoS Genet. 2013, 9, e1003792. [Google Scholar] [CrossRef]
- Rusu, P.; Jansen, A.; Soba, P.; Kirsch, J.; Löwer, A.; Merdes, G.; Kuan, Y.H.; Jung, A.; Beyreuther, K.; Kjaerulff, O. Axonal accumulation of synaptic markers in APP transgenic Drosophila depends on the NPTY motif and is paralleled by defects in synaptic plasticity. Eur. J. Neurosci. 2007, 25, 1079–1086. [Google Scholar] [CrossRef]
- Yao, P.J.; Zhu, M.; Pyun, E.I.; Brooks, A.I.; Therianos, S.; Meyers, V.E.; Coleman, P.D. Defects in expression of genes related to synaptic vesicle traffickingin frontal cortex of Alzheimer’s disease. Neurobiol. Dis. 2003, 12, 97–109. [Google Scholar] [CrossRef]
- Kittel, R.J.; Wichmann, C.; Rasse, T.M.; Fouquet, W.; Schmidt, M.; Schmid, A.; Wagh, D.A.; Pawlu, C.; Kellner, R.R.; Willig, K.I. Bruchpilot promotes active zone assembly, Ca2+ channel clustering, and vesicle release. Science 2006, 312, 1051–1054. [Google Scholar] [CrossRef]
- Mhatre, S.D.; Satyasi, V.; Killen, M.; Paddock, B.E.; Moir, R.D.; Saunders, A.J.; Marenda, D.R. Synaptic abnormalities in a Drosophila model of Alzheimer’s disease. Dis. Model. Mech. 2014, 7, 373–385. [Google Scholar] [CrossRef]
- Huang, J.-K.; Ma, P.-L.; Ji, S.-Y.; Zhao, X.-L.; Tan, J.-X.; Sun, X.-J.; Huang, F.-D. Age-dependent alterations in the presynaptic active zone in a Drosophila model of Alzheimer’s disease. Neurobiol. Dis. 2013, 51, 161–167. [Google Scholar] [CrossRef]
- Thomas, U.; Ebitsch, S.; Gorczyca, M.; Koh, Y.; Hough, C.; Woods, D.; Gundelfinger, E.; Budnik, V. Synaptic targeting and localization of discs-large is a stepwise process controlled by different domains of the protein. Curr. Biol. 2000, 10, 1108–1117. [Google Scholar] [CrossRef]
- Chen, K.; Featherstone, D.E. Discs-large (DLG) is clustered by presynaptic innervation and regulates postsynaptic glutamate receptor subunit composition in Drosophila. BMC Biol. 2005, 3, 1. [Google Scholar] [CrossRef] [PubMed]
- Furotani, K.; Kamimura, K.; Yajima, T.; Nakayama, M.; Enomoto, R.; Tamura, T.; Okazawa, H.; Sone, M. Suppression of the synaptic localization of a subset of proteins including APP partially ameliorates phenotypes of the Drosophila Alzheimer’s disease model. PLoS ONE 2018, 13, e0204048. [Google Scholar] [CrossRef] [PubMed]
- Verri, M.; Pastoris, O.; Dossena, M.; Aquilani, R.; Guerriero, F.; Cuzzoni, G.; Venturini, L.; Ricevuti, G.; Bongiorno, A.I. Mitochondrial alterations, oxidative stress and neuroinflammation in Alzheimer’s disease. Int. J. Immunopathol. Pharmacol. 2012, 25, 345–353. [Google Scholar] [CrossRef]
- Lv, F.; Yang, X.; Cui, C.; Su, C. Exogenous expression of Drp1 plays neuroprotective roles in the Alzheimer’s disease in the Abeta42 transgenic Drosophila model. PLoS ONE 2017, 12, e0176183. [Google Scholar] [CrossRef] [PubMed]
- Verstreken, P.; Ly, C.V.; Venken, K.J.; Koh, T.-W.; Zhou, Y.; Bellen, H.J. Synaptic mitochondria are critical for mobilization of reserve pool vesicles at Drosophila neuromuscular junctions. Neuron 2005, 47, 365–378. [Google Scholar] [CrossRef]
- Iijima-Ando, K.; Hearn, S.A.; Shenton, C.; Gatt, A.; Zhao, L.; Iijima, K. Mitochondrial mislocalization underlies Aβ42-induced neuronal dysfunction in a Drosophila model of Alzheimer’s disease. PLoS ONE 2009, 4, e8310. [Google Scholar] [CrossRef]
- Akiyama, H.; Barger, S.; Barnum, S.; Bradt, B.; Bauer, J.; Cole, G.M.; Cooper, N.R.; Eikelenboom, P.; Emmerling, M.; Fiebich, B.L. Inflammation and Alzheimer’s disease. Neurobiol. Aging 2000, 21, 383–421. [Google Scholar] [CrossRef]
- Zhang, B.; Gaiteri, C.; Bodea, L.-G.; Wang, Z.; McElwee, J.; Podtelezhnikov, A.A.; Zhang, C.; Xie, T.; Tran, L.; Dobrin, R. Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell 2013, 153, 707–720. [Google Scholar] [CrossRef]
- Wyss-Coray, T.; Mucke, L. Inflammation in neurodegenerative disease—A double-edged sword. Neuron 2002, 35, 419–432. [Google Scholar] [CrossRef]
- Ransohoff, R.M.; Brown, M.A. Innate immunity in the central nervous system. J. Clin. Investig. 2012, 122, 1164–1171. [Google Scholar] [CrossRef]
- Petersen, A.J.; Rimkus, S.A.; Wassarman, D.A. ATM kinase inhibition in glial cells activates the innate immune response and causes neurodegeneration in Drosophila. Proc. Natl. Acad. Sci. USA 2012, 109, E656–E664. [Google Scholar] [CrossRef]
- Cao, Y.; Chtarbanova, S.; Petersen, A.J.; Ganetzky, B. Dnr1 mutations cause neurodegeneration in Drosophila by activating the innate immune response in the brain. Proc. Natl. Acad. Sci. USA 2013, 110, E1752–E1760. [Google Scholar] [CrossRef]
- Lee, C.D.; Landreth, G.E. The role of microglia in amyloid clearance from the AD brain. J. Neural Transm. 2010, 117, 949–960. [Google Scholar] [CrossRef]
- Coutinho-Budd, J.; Freeman, M.R. Probing the enigma: Unraveling glial cell biology in invertebrates. Curr. Opin. Neurobiol. 2013, 23, 1073–1079. [Google Scholar] [CrossRef]
- Ray, A.; Speese, S.D.; Logan, M.A. Glial draper rescues Aβ toxicity in a Drosophila model of Alzheimer’s disease. J. Neurosci. 2017, 37, 11881–11893. [Google Scholar] [CrossRef]
- Purice, M.D.; Speese, S.D.; Logan, M.A. Delayed glial clearance of degenerating axons in aged Drosophila is due to reduced PI3K/Draper activity. Nat. Commun. 2016, 7, 12871. [Google Scholar] [CrossRef]
- Ulland, T.K.; Colonna, M. TREM2—A key player in microglial biology and Alzheimer disease. Nat. Rev. Neurol. 2018, 14, 667–675. [Google Scholar] [CrossRef]
- Tahara, K.; Kim, H.-D.; Jin, J.-J.; Maxwell, J.A.; Li, L.; Fukuchi, K.-I. Role of toll-like receptor signalling in Aβ uptake and clearance. Brain 2006, 129, 3006–3019. [Google Scholar] [CrossRef]
- Cameron, B.; Landreth, G.E. Inflammation, microglia, and Alzheimer’s disease. Neurobiol. Dis. 2010, 37, 503–509. [Google Scholar] [CrossRef]
- Sessa, G.; Podini, P.; Mariani, M.; Meroni, A.; Spreafico, R.; Sinigaglia, F.; Colonna, M.; Panina, P.; Meldolesi, J. Distribution and signaling of TREM2/DAP12, the receptor system mutated in human polycystic lipomembraneous osteodysplasia with sclerosing leukoencephalopathy dementia. Eur. J. Neurosci. 2004, 20, 2617–2628. [Google Scholar] [CrossRef] [PubMed]
- Jiang, T.; Yu, J.-T.; Zhu, X.-C.; Tan, L. TREM2 in Alzheimer’s disease. Mol. Neurobiol. 2013, 48, 180–185. [Google Scholar] [CrossRef] [PubMed]
- Guerreiro, R.; Wojtas, A.; Bras, J.; Carrasquillo, M.; Rogaeva, E.; Majounie, E.; Cruchaga, C.; Sassi, C.; Kauwe, J.S.; Younkin, S. TREM2 variants in Alzheimer’s disease. N. Engl. J. Med. 2013, 368, 117–127. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Cella, M.; Mallinson, K.; Ulrich, J.D.; Young, K.L.; Robinette, M.L.; Gilfillan, S.; Krishnan, G.M.; Sudhakar, S.; Zinselmeyer, B.H. TREM2 lipid sensing sustains the microglial response in an Alzheimer’s disease model. Cell 2015, 160, 1061–1071. [Google Scholar] [CrossRef] [PubMed]
- Sekiya, M.; Wang, M.; Fujisaki, N.; Sakakibara, Y.; Quan, X.; Ehrlich, M.E.; De Jager, P.L.; Bennett, D.A.; Schadt, E.E.; Gandy, S. Integrated biology approach reveals molecular and pathological interactions among Alzheimer’s Aβ42, Tau, TREM2, and TYROBP in Drosophila models. Genome Med. 2018, 10, 26. [Google Scholar] [CrossRef]
- Khush, R.S.; Leulier, F.; Lemaitre, B. Drosophila immunity: Two paths to NF-kappaB. Trends Immunol. 2001, 22, 260–264. [Google Scholar] [CrossRef]
- Richard, K.L.; Filali, M.; Préfontaine, P.; Rivest, S. Toll-like receptor 2 acts as a natural innate immune receptor to clear amyloid β1–42 and delay the cognitive decline in a mouse model of Alzheimer’s disease. J. Neurosci. 2008, 28, 5784–5793. [Google Scholar] [CrossRef]
- Scholtzova, H.; Kascsak, R.J.; Bates, K.A.; Boutajangout, A.; Kerr, D.J.; Meeker, H.C.; Mehta, P.D.; Spinner, D.S.; Wisniewski, T. Induction of toll-like receptor 9 signaling as a method for ameliorating Alzheimer’s disease-related pathology. J. Neurosci. 2009, 29, 1846–1854. [Google Scholar] [CrossRef]
- Walker, D.G.; Tang, T.M.; Lue, L.-F. Increased expression of toll-like receptor 3, an anti-viral signaling molecule, and related genes in Alzheimer’s disease brains. Exp. Neurol. 2018, 309, 91–106. [Google Scholar] [CrossRef]
- Tan, L.; Schedl, P.; Song, H.-J.; Garza, D.; Konsolaki, M. The Toll→ NFκB signaling pathway mediates the neuropathological effects of the human Alzheimer’s Aβ42 polypeptide in Drosophila. PLoS ONE 2008, 3, e3966. [Google Scholar] [CrossRef]
- Li, Y.; Sibon, O.; Dijkers, P. Inhibition of NF-κB in astrocytes is sufficient to delay neurodegeneration induced by proteotoxicity in neurons. J. Neuroinflammation 2018, 15, 261. [Google Scholar] [CrossRef] [PubMed]
- Bertram, L.; Tanzi, R.E. Alzheimer disease risk genes: 29 and counting. Nat. Rev. Neurol. 2019, 15, 191. [Google Scholar] [CrossRef] [PubMed]
- Jansen, I.E.; Savage, J.E.; Watanabe, K.; Bryois, J.; Williams, D.M.; Steinberg, S.; Sealock, J.; Karlsson, I.K.; Hägg, S.; Athanasiu, L. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 2019, 51, 404–413. [Google Scholar] [CrossRef]
- Ridge, P.G.; Hoyt, K.B.; Boehme, K.; Mukherjee, S.; Crane, P.K.; Haines, J.L.; Mayeux, R.; Farrer, L.A.; Pericak-Vance, M.A.; Schellenberg, G.D. Assessment of the genetic variance of late-onset Alzheimer’s disease. Neurobiol. Aging 2016, 41, e13–e200. [Google Scholar] [CrossRef]
- Hall, A.M.; Roberson, E.D. Mouse models of Alzheimer’s disease. Brain Res. Bull. 2012, 88, 3–12. [Google Scholar] [CrossRef]
- Cummings, J.L.; Morstorf, T.; Zhong, K. Alzheimer’s disease drug-development pipeline: Few candidates, frequent failures. Alzheimers Res. Ther. 2014, 6, 37. [Google Scholar] [CrossRef] [PubMed]
- Abbott, A.; Dolgin, E. Failed Alzheimer’s trial does not kill leading theory of disease. Nature 2016, 540, 15. [Google Scholar] [CrossRef]
- Sabbagh, M.N.; Hendrix, S.; Harrison, J.E. FDA position statement “Early Alzheimer’s disease: Developing drugs for treatment, Guidance for Industry”. Alzheimer’s Dement. Transl. Res. Clin. Interv. 2019, 5, 13–19. [Google Scholar]
- Wang, X.; Perumalsamy, H.; Kwon, H.W.; Na, Y.-E.; Ahn, Y.-J. Effects and possible mechanisms of action of acacetin on the behavior and eye morphology of Drosophila models of Alzheimer’s disease. Sci. Rep. 2015, 5, 16127. [Google Scholar] [CrossRef]
- Lee, B.I.; Lee, S.; Suh, Y.S.; Lee, J.S.; Kim, A.k.; Kwon, O.Y.; Yu, K.; Park, C.B. Photoexcited porphyrins as a strong suppressor of β-Amyloid aggregation and synaptic toxicity. Angew. Chem. Int. Ed. 2015, 54, 11472–11476. [Google Scholar] [CrossRef]
- Lee, B.I.; Suh, Y.S.; Chung, Y.J.; Yu, K.; Park, C.B. Shedding light on Alzheimer’s β-amyloidosis: Photosensitized methylene blue inhibits self-assembly of β-amyloid peptides and disintegrates their aggregates. Sci. Rep. 2017, 7, 7523. [Google Scholar] [CrossRef]
- Deshpande, P.; Gogia, N.; Singh, A. Exploring the efficacy of natural products in alleviating Alzheimer’s disease. Neural. Regen. Res. 2019, 14, 1321. [Google Scholar] [PubMed]
- Lee, S.; Bang, S.M.; Lee, J.W.; Cho, K.S. Evaluation of traditional medicines for neurodegenerative diseases using Drosophila models. Evid. Based Complement. Altern. Med. 2014, 2014. [Google Scholar] [CrossRef] [PubMed]
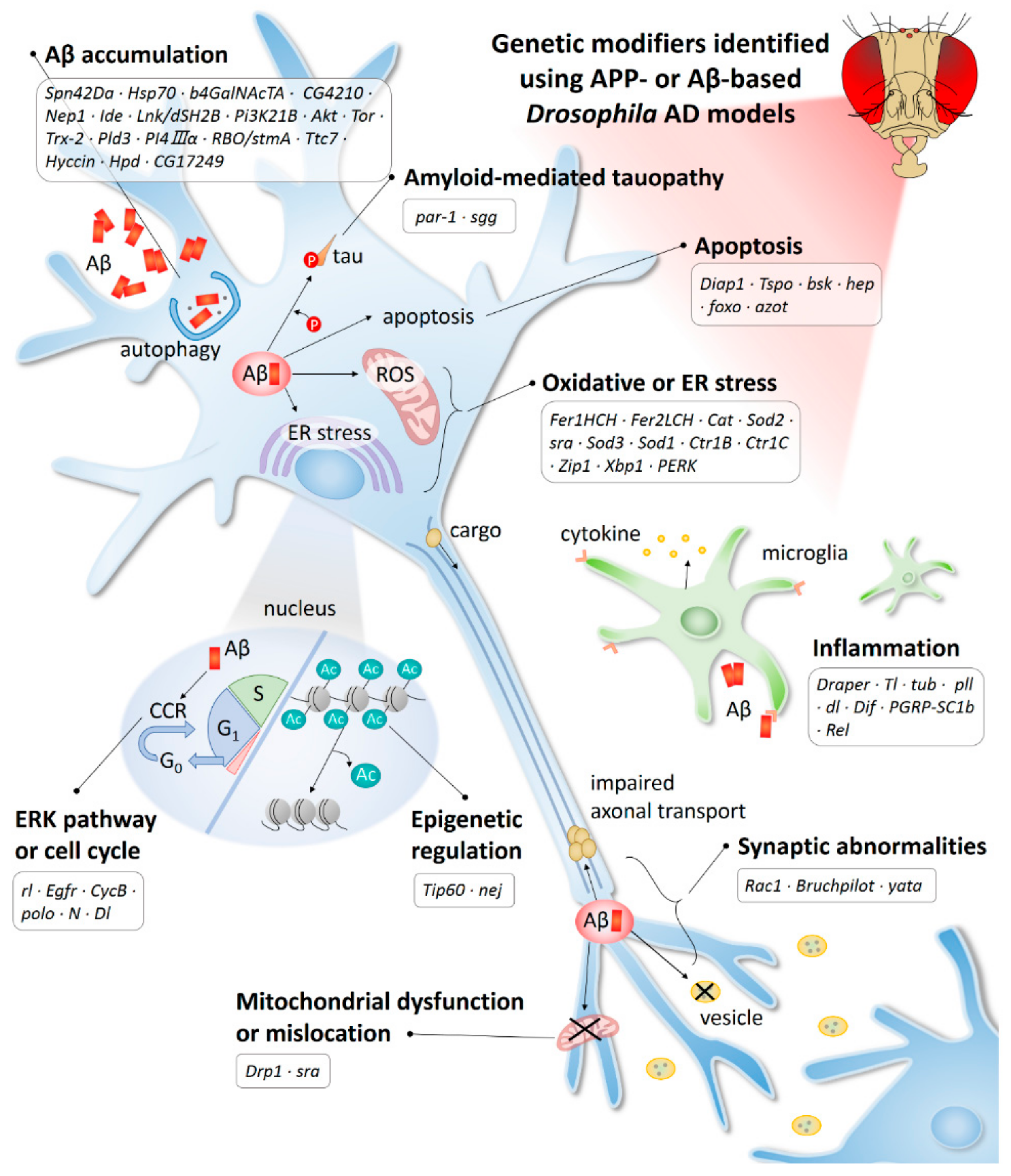
| Pathway | Drosophila Genes | Human Genes | Protein Type and Function | Effect | References |
|---|---|---|---|---|---|
| Aβ production and aggregation | Spn42Da | SERPINI | Serine protease inhibitor that interacts with tissue plasminogen activator | S 1 | [51] |
| Hsp70 | HSPA1A | Central component of the cellular network of molecular chaperones and folding catalysts | S | [52,53] | |
| b4GalNAcTA | B4GALT6 | β-1,4-galactosyltransferase | E 2 | [55] | |
| CG4210 | SAT1 | α-2,3-sialyltransferase | E | [55] | |
| Nep1 | MME | Membrane metalloendopeptidase Aβ degrading enzyme | S | [57] | |
| Ide | IDE | Zinc metallopeptidase that degrades insulin | S | [59] | |
| Lnk/dSH2B | SH2B1 | SH2-domain containing mediator important for insulin receptor signaling | S | [60] | |
| Pi3K21B | PIK3R3 | Regulatory subunit of PI3K that phosphorylates phosphatidylinositol | E | [63] | |
| Akt | AKT | AKT serine/threonine kinase | E | [63] | |
| Tor | MTOR | Phosphatidylinositol kinase-related kinase | E | [63] | |
| Trx-2 | TXN | Thioredoxin-1 that is involved in many redox reaction | S | [64] | |
| Pld3 | PLD3 | Enzyme that catalyzes the hydrolysis of membrane phospholipids | S | [65] | |
| PI4KIIIα | PI4KA | Lipid kinase that synthesizes phosphatidylinositol 4-phosphate from phosphatidylinositol | E | [72] | |
| RBO/stmA | EFR3B | Component of complex required to localize PI4K to the plasma membrane | E | [72] | |
| Ttc7 | TTC7B | Component of complex required to localize PI4K to the plasma membrane | E | [73] | |
| Hyccin | FAM126A | Component of complex required to localize PI4K to the plasma membrane | E | [73] | |
| Hpd | HPD | 4-hydroxyphenylpyruvate dioxygenase that catalyzes the conversion of 4-hydroxyphenyl-pyruvate to homogentisate | S | [74] | |
| CG17249 | PRCC | Protein that may play a role in pre-mRNA splicing | S | [74] | |
| Tauopathy | par-1 | MARK | Serine/threonine kinase that plays critical roles in cell polarity and microtubule dynamics | E | [99] |
| sgg | GSK3β | Serine/threonine kinase that plays multiple roles in various signaling pathways | E | [99] | |
| Oxidative stress | Fer1HCH | FTH1 | Subunit of Ferritin, an iron-storage protein | S | [107,108,109] |
| Fer2LCH | FTMT | Subunit of mitochondrial Ferritin, an iron-storage protein | S | [107,108,109] | |
| Cat | CAT | Enzyme that catalyzes decomposition of hydrogen peroxide to water and oxygen | S | [107] | |
| Sod2 | SOD2 | Mitochondrial Mn-dependent superoxide dismutase | S | [107] [110] | |
| sra | RCAN1 | Inhibitor of calcineurin | E | [111] | |
| Sod3 | SOD3 | Extracellular Cu/Zn-dependent superoxide dismutase | E | [112] | |
| Sod1 | SOD1 | Cytoplasmic Cu/Zn-dependent superoxide dismutase | E | [107] | |
| Ctr1B | SLC31A1 | Copper importer | E | [117] | |
| Ctr1C | SLC31A1 | Copper importer | E | [117] | |
| Zip1 | SLC39A3 | Zinc importer | E | [116] | |
| ER stress | Xbp1 | XBP1 | Transcriptional factor that mediates the unfolded protein response | S | [35,124] |
| PERK | EIF2AK3 | ER transmembrane kinase that phosphorylates eukaryotic translation-initiation factor 2-alpha during ER stress | S | [124] | |
| EGFR/ERK pathway | rl | MAPK1 | Serine/threonine kinase and core component of the EGFR/MAPK pathway | E | [131] |
| Egfr | EGFR | Membrane-localized tyrosine kinase receptor for epidermal growth factor | E | [137] | |
| Cell cycle | CycB | CCNB1 | G2/mitotic-specific protein which is a member of the cyclin family | E | [143] |
| polo | PLK1 | Serine/threonine kinase involved in mitosis | E | [144] | |
| N | NOTCH1 | Transmembrane receptor for Notch signaling | E | [147] | |
| Dl | DLL1 | Transmembrane ligand for Notch signaling | E | [147] | |
| Apoptosis | Diap1 | BIRC2 | E3 ligase with inhibitory activity on caspase | S E | [150] [162] |
| Tspo | TSPO | Outer mitochondrial membrane protein related to steroid and heme biosynthesis, apoptosis, protein import, cell proliferation, and differentiation | E | [154] | |
| bsk | MAPK8 | Serine/threonine kinase that phosphorylates the Jra transcription factor | E | [38] [150] | |
| hep | MAP2K7 | Serine/threonine kinase involved in the JNK pathway by phosphorylating JNK | E | [160] | |
| foxo | FOXO | Forkhead family of transcription factor regulated by various signaling pathway | E | [150] [161] | |
| azot | CALM1 | EF-hand calcium binding protein act as fitness checkpoint | S | [162] | |
| Epigenetic regulation | Tip60 | KAT5 | Histone acetyltransferase | S | [172] |
| nej | CBP | Histone acetyltransferase | S | [174] | |
| Synaptic abnormalities | Rac1 | RAC1 | GTPase which belongs to the RAS superfamily of small GTP-binding proteins | E | [193] |
| Bruchpilot | ERC2 | Cytoskeletal protein critical for structural integrity of electron-dense projection at pre-active zones | S | [202] | |
| yata | SCYL1 | Transcriptional regulator belonging to the SCY1-like family of kinase-like proteins | E | [205] | |
| Mitochondrial dysfunction or mislocation | Drp1 | DNM1L | Dynamin related protein that regulates mitochondrial fission | S | [207] |
| sra | RCAN1 | Inhibitor of calcineurin | S E | [197] [111] | |
| Inflammation and innate immune system | Draper | MEFG10 | Multiple EGF like domains 10, which encodes a phagocytic receptor of glia | S | [218] |
| Tl | TLR | Toll-like receptor that promotes NF-kB like transcription factors | E | [232] | |
| tub | IRAK1 | Downstream component of the Toll pathway | E | [232] | |
| pll | IRAK4 | Downstream component of the Toll pathway | E | [232] | |
| dl | RELA | Transcription factor regulated by the Toll pathway | E | [232] | |
| Dif | RELB | Transcription factor regulated by the Toll pathway | E | [232] | |
| PGRP-SC1b | PGLYRP1 | Peptidoglycan recognition protein SC1b which is a negative regulator of Imd pathway | E | [112] | |
| Rel | NFKB1 | Subunit 1 of nuclear factor kappa B, which is a main regulatory gene in the Imd pathway | E | [233] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jeon, Y.; Lee, J.H.; Choi, B.; Won, S.-Y.; Cho, K.S. Genetic Dissection of Alzheimer’s Disease Using Drosophila Models. Int. J. Mol. Sci. 2020, 21, 884. https://doi.org/10.3390/ijms21030884
Jeon Y, Lee JH, Choi B, Won S-Y, Cho KS. Genetic Dissection of Alzheimer’s Disease Using Drosophila Models. International Journal of Molecular Sciences. 2020; 21(3):884. https://doi.org/10.3390/ijms21030884
Chicago/Turabian StyleJeon, Youngjae, Jae Ha Lee, Byoungyun Choi, So-Yoon Won, and Kyoung Sang Cho. 2020. "Genetic Dissection of Alzheimer’s Disease Using Drosophila Models" International Journal of Molecular Sciences 21, no. 3: 884. https://doi.org/10.3390/ijms21030884
APA StyleJeon, Y., Lee, J. H., Choi, B., Won, S.-Y., & Cho, K. S. (2020). Genetic Dissection of Alzheimer’s Disease Using Drosophila Models. International Journal of Molecular Sciences, 21(3), 884. https://doi.org/10.3390/ijms21030884





