Identification and Pilot Evaluation of Salivary Peptides from Anopheles albimanus as Biomarkers for Bite Exposure and Malaria Infection in Colombia
Abstract
1. Introduction
2. Results
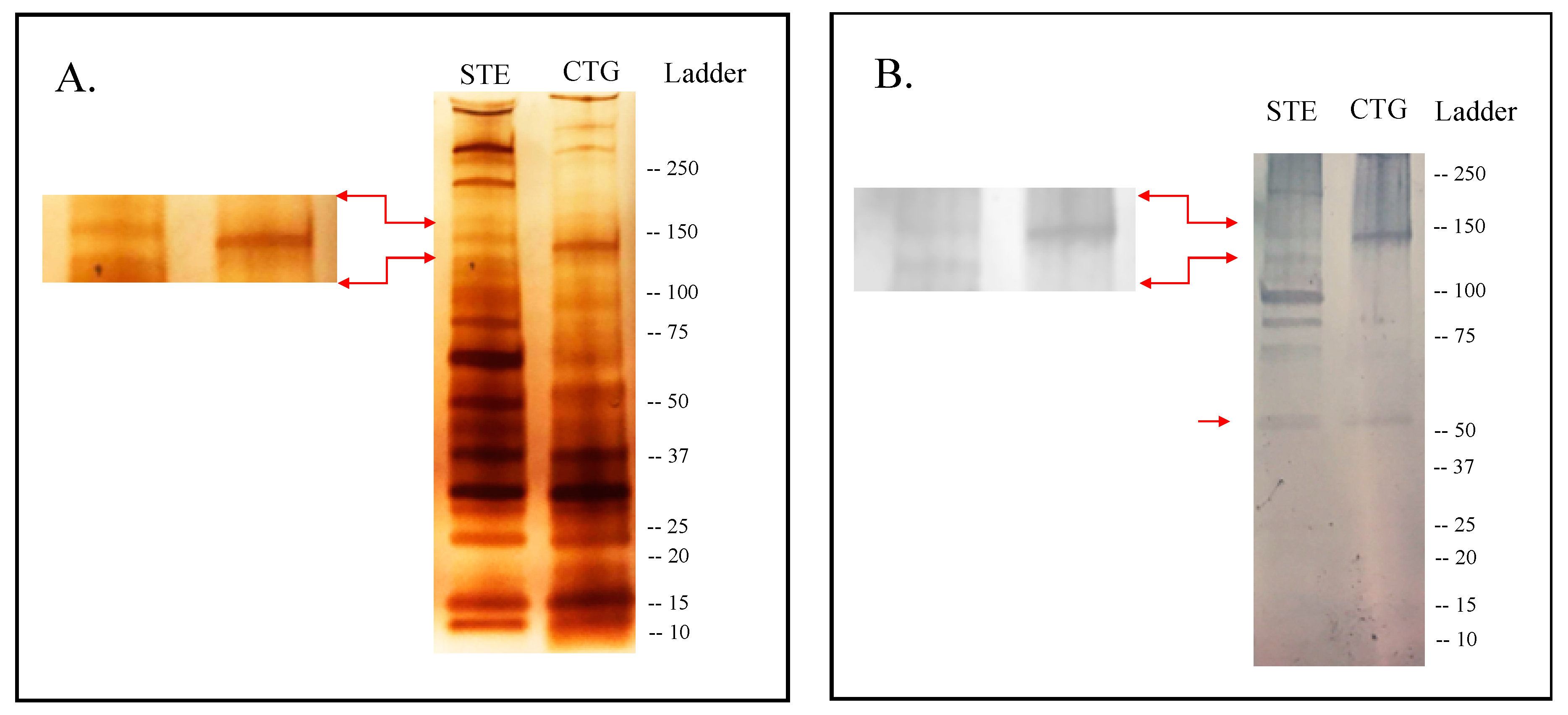
2.1. Immunogenic Candidates Identified in the Sialome of An. Albimanus
2.2. Selection of Candidate Biomarkers
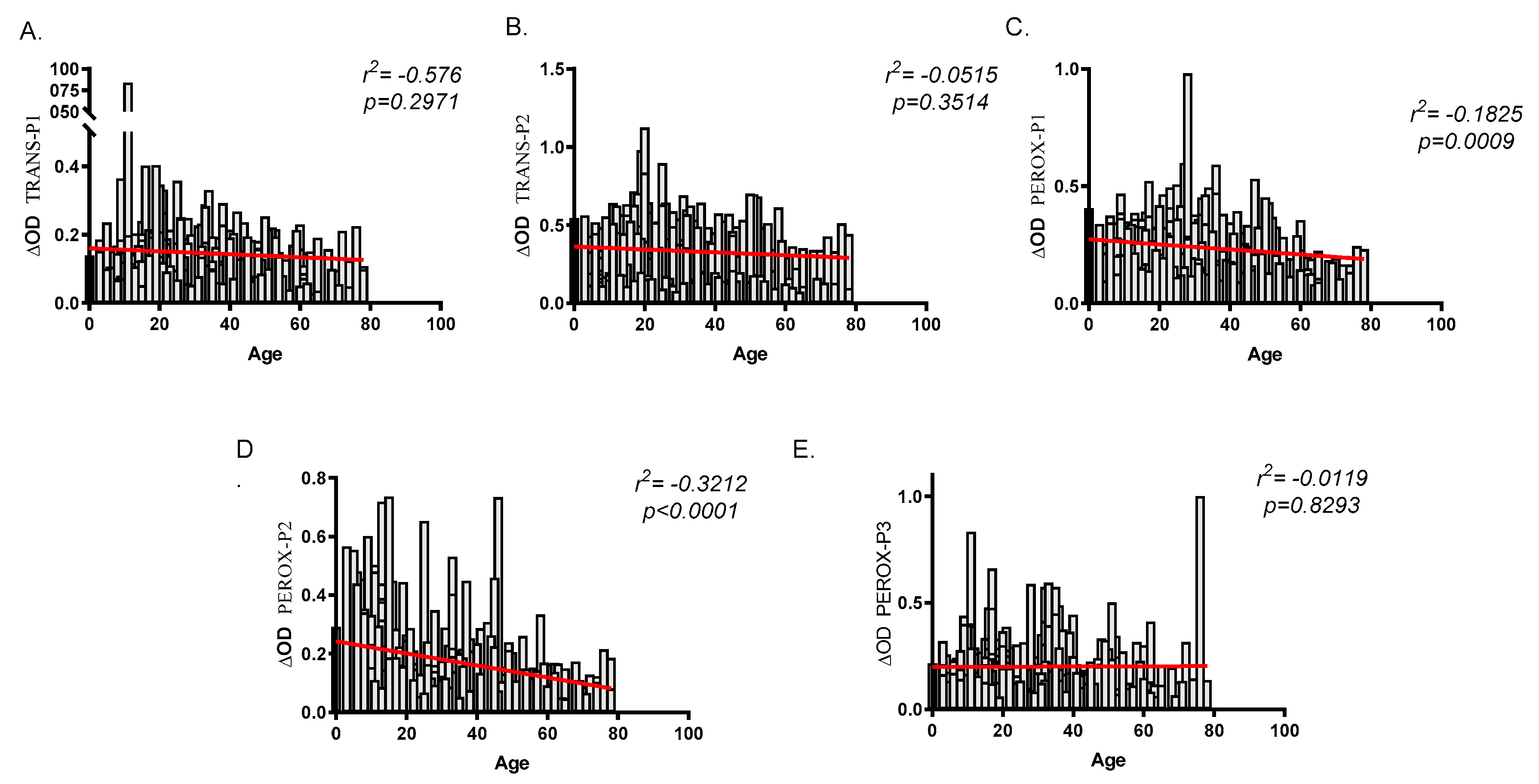
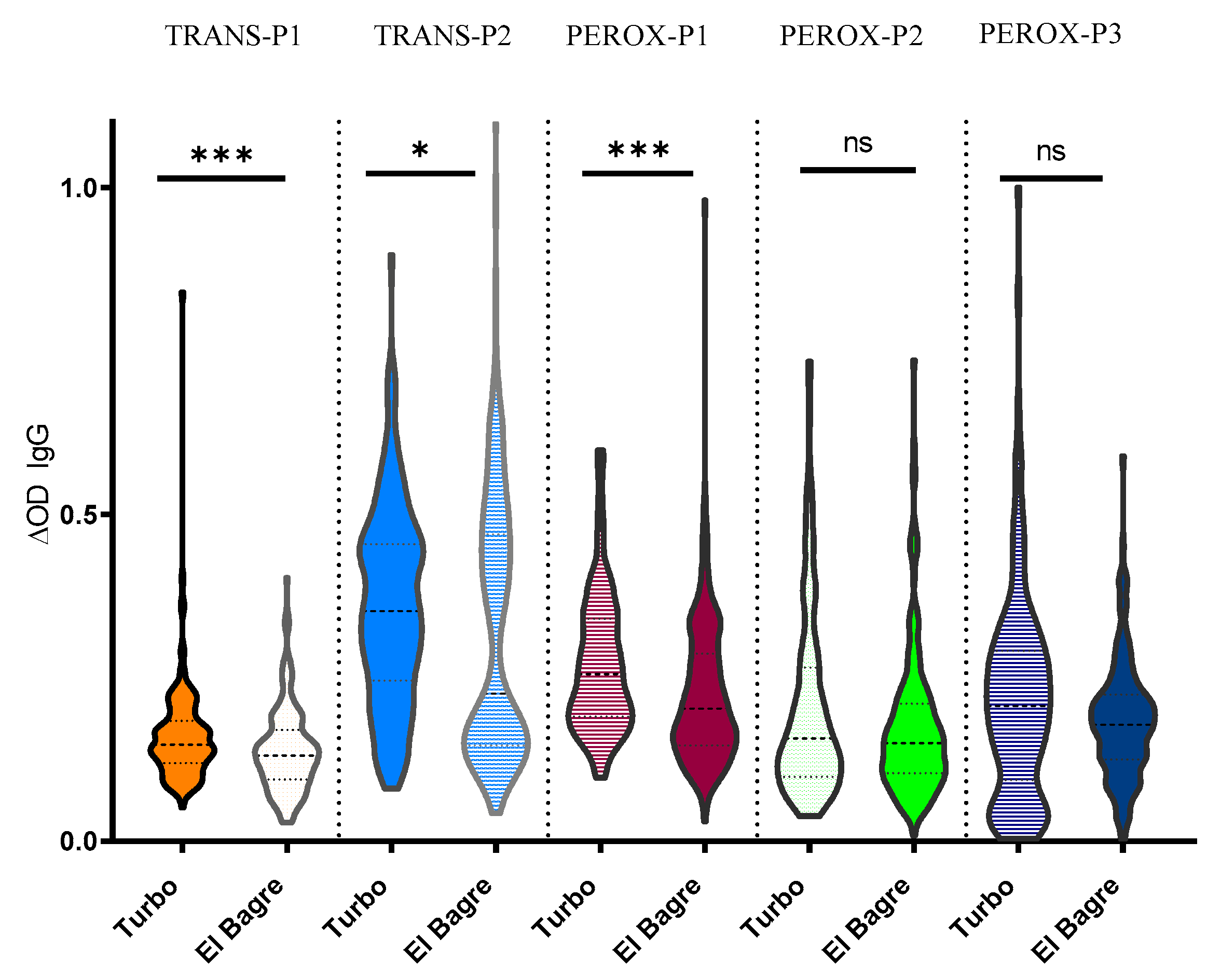
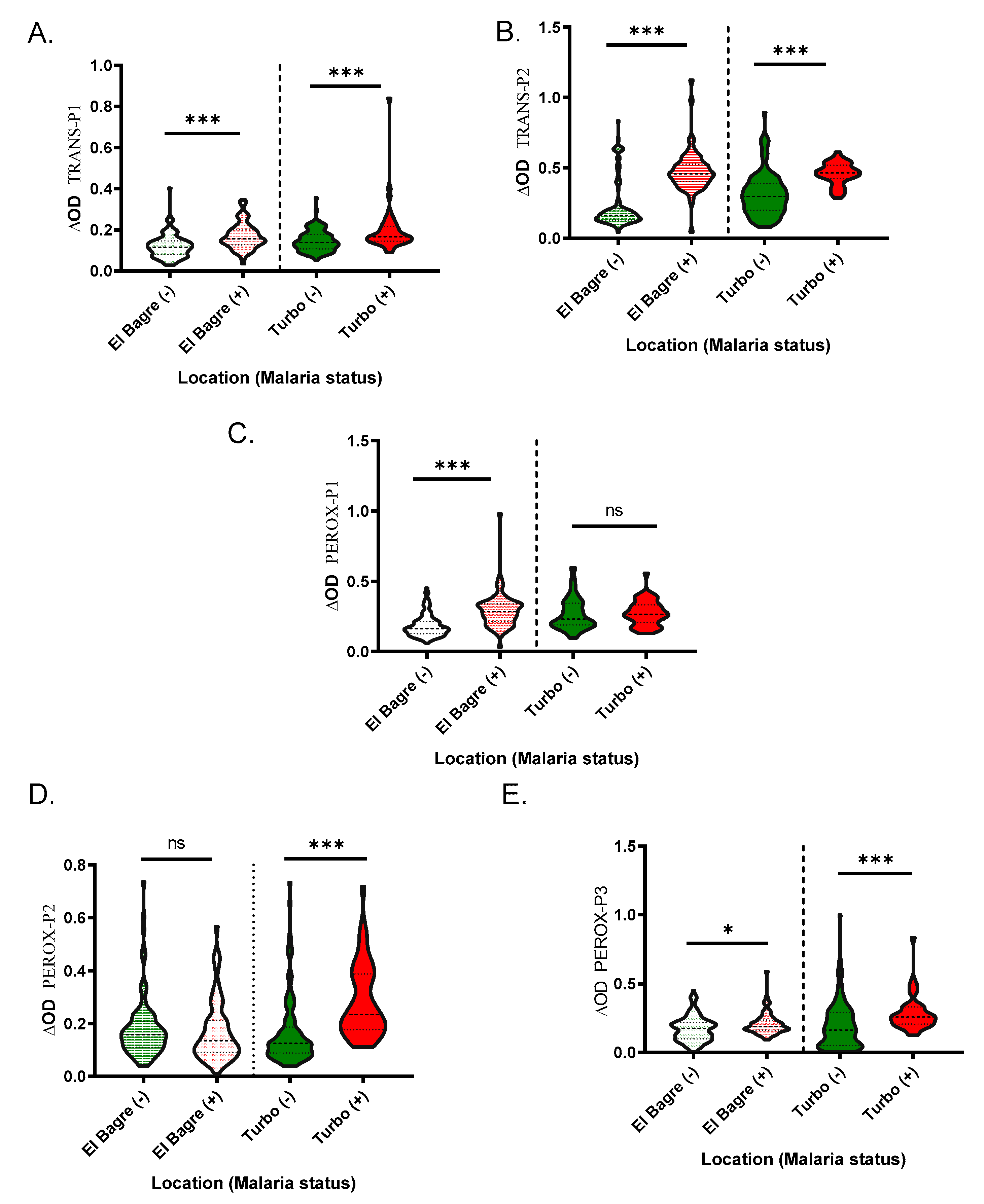
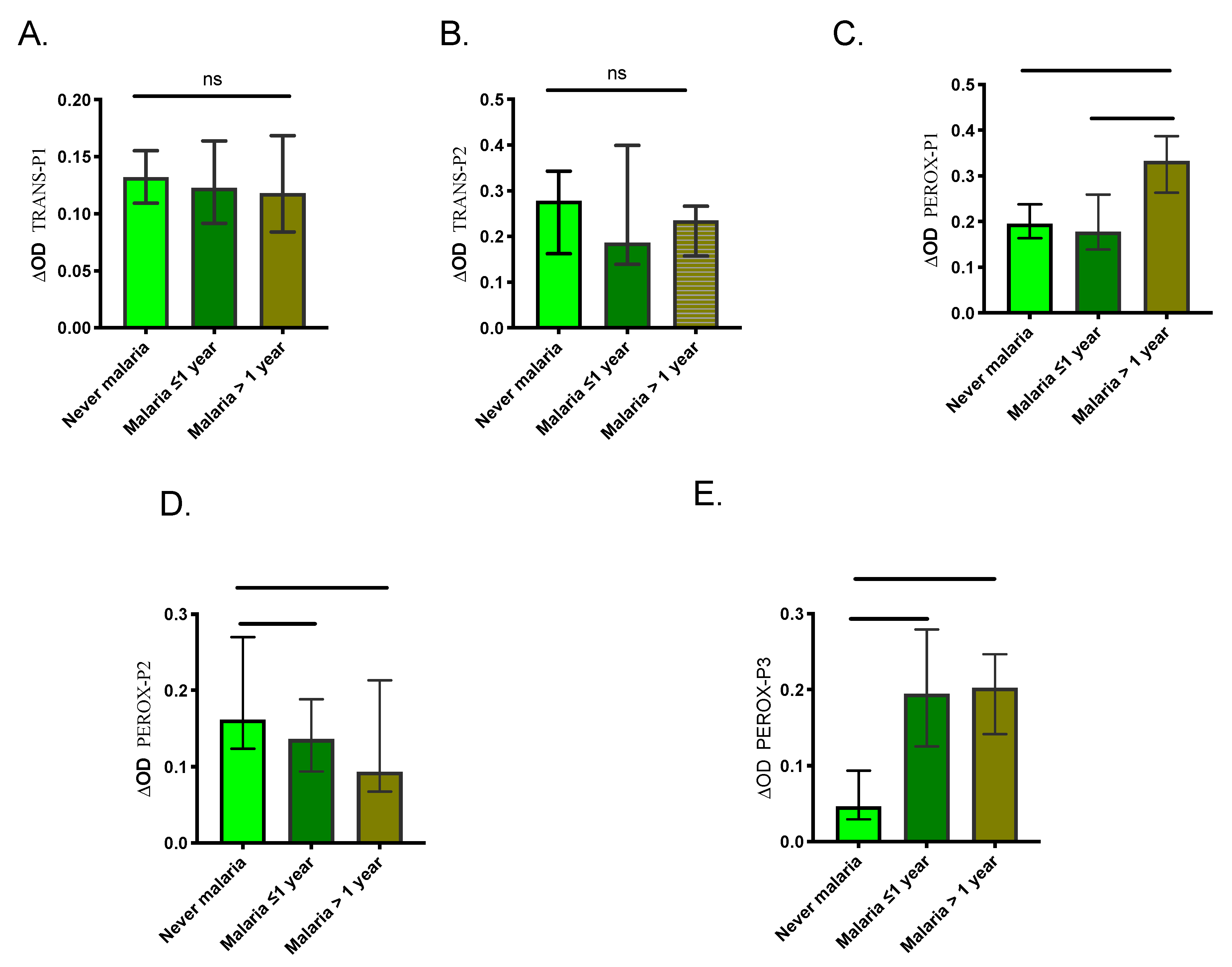
2.3. Human Antibody Responses Specific to An. albimanus Salivary Peptides
3. Discussion
4. Materials and Methods
4.1. Study Site and Serum Samples for Validation of Biomarkers
4.2. Mosquito Rearing and Salivary Gland Extraction
4.3. SDS and Western Blot
4.4. Peptide Design and Selection
4.5. Human IgG Detection by Indirect ELISA
4.6. Statistical Analyses
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- World Health Organization. World Malaria Report 2018. Available online: http://www.who.int/iris/handle/10665/275867 (accessed on 3 May 2019).
- Rodriguez, J.C.; Uribe, G.A.; Araujo, R.M.; Narvaez, P.C.; Valencia, S.H. Epidemiology and control of malaria in Colombia. Memórias Inst. Oswaldo Cruz 2011, 106, 114–122. [Google Scholar] [CrossRef]
- Feged-Rivadeneira, A.; Angel, A.; Gonzalez-Casabianca, F.; Rivera, C. Malaria intensity in Colombia by regions and populations. PLoS ONE 2018, 13, e0203673. [Google Scholar] [CrossRef]
- Montoya-Lerma, J.; Solarte, Y.A.; Giraldo-Calderon, G.I.; Quinones, M.L.; Ruiz-Lopez, F.; Wilkerson, R.C.; Gonzalez, R. Malaria vector species in Colombia: A review. Memórias Inst. Oswaldo Cruz 2011, 106, 223–238. [Google Scholar] [CrossRef] [PubMed]
- Naranjo-Diaz, N.; Rosero, D.A.; Rua-Uribe, G.; Luckhart, S.; Correa, M.M. Abundance, behavior and entomological inoculation rates of anthropophilic anophelines from a primary Colombian malaria endemic area. Parasit Vectors 2013, 6, 61. [Google Scholar] [CrossRef]
- Gutierrez, L.A.; Naranjo, N.; Jaramillo, L.M.; Muskus, C.; Luckhart, S.; Conn, J.E.; Correa, M.M. Natural infectivity of Anopheles species from the Pacific and Atlantic Regions of Colombia. Acta Trop. 2008, 107, 99–105. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, L.A.; Gonzalez, J.J.; Gomez, G.F.; Castro, M.I.; Rosero, D.A.; Luckhart, S.; Conn, J.E.; Correa, M.M. Species composition and natural infectivity of anthropophilic Anopheles (Diptera: Culicidae) in the states of Cordoba and Antioquia, Northwestern Colombia. Memórias Inst. Oswaldo Cruz 2009, 104, 1117–1124. [Google Scholar] [CrossRef] [PubMed]
- Beier, J.C.; Killeen, G.F.; Githure, J.I. Short report: Entomologic inoculation rates and Plasmodium falciparum malaria prevalence in Africa. Am. J. Trop. Med. Hyg. 1999, 61, 109–113. [Google Scholar] [CrossRef] [PubMed]
- Drakeley, C.J.; Corran, P.H.; Coleman, P.G.; Tongren, J.E.; McDonald, S.L.; Carneiro, I.; Malima, R.; Lusingu, J.; Manjurano, A.; Nkya, W.M.; et al. Estimating medium- and long-term trends in malaria transmission by using serological markers of malaria exposure. Proc. Natl. Acad. Sci. USA 2005, 102, 5108–5113. [Google Scholar] [CrossRef] [PubMed]
- Bousema, T.; Okell, L.; Felger, I.; Drakeley, C. Asymptomatic malaria infections: Detectability, transmissibility and public health relevance. Nat. Rev. Microbiol. 2014, 12, 833–840. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, J.M.; Francischetti, I.M. Role of arthropod saliva in blood feeding: Sialome and post-sialome perspectives. Ann. Rev. Entomol. 2003, 48, 73–88. [Google Scholar] [CrossRef]
- Orlandi-Pradines, E.; Almeras, L.; Denis de Senneville, L.; Barbe, S.; Remoue, F.; Villard, C.; Cornelie, S.; Penhoat, K.; Pascual, A.; Bourgouin, C.; et al. Antibody response against saliva antigens of Anopheles gambiae and Aedes aegypti in travellers in tropical Africa. Microbes Infect. 2007, 9, 1454–1462. [Google Scholar] [CrossRef] [PubMed]
- Londono-Renteria, B.L.; Eisele, T.P.; Keating, J.; James, M.A.; Wesson, D.M. Antibody response against Anopheles albimanus (Diptera: Culicidae) salivary protein as a measure of mosquito bite exposure in Haiti. J. Med. Entomol. 2010, 47, 1156–1163. [Google Scholar] [CrossRef] [PubMed]
- Remoue, F.; Cisse, B.; Ba, F.; Sokhna, C.; Herve, J.P.; Boulanger, D.; Simondon, F. Evaluation of the antibody response to Anopheles salivary antigens as a potential marker of risk of malaria. Trans. R. Soc. Trop. Med. Hyg. 2006, 100, 363–370. [Google Scholar] [CrossRef] [PubMed]
- Cardenas, J.C.; Drame, P.M.; Luque-Burgos, K.A.; Berrio, J.D.; Entrena-Mutis, E.; Gonzalez, M.U.; Carvajal, D.J.; Gutierrez-Silva, L.Y.; Cardenas, L.D.; Colpitts, T.M.; et al. IgG1 and IgG4 antibodies against Aedes aegypti salivary proteins and risk for dengue infections. PLoS ONE 2019, 14, e0208455. [Google Scholar] [CrossRef]
- Andrade, B.B.; Rocha, B.C.; Reis-Filho, A.; Camargo, L.M.; Tadei, W.P.; Moreira, L.A.; Barral, A.; Barral-Netto, M. Anti-Anopheles darlingi saliva antibodies as marker of Plasmodium vivax infection and clinical immunity in the Brazilian Amazon. Malar. J. 2009, 8, 121. [Google Scholar] [CrossRef] [PubMed]
- Arca, B.; Lombardo, F.; Struchiner, C.J.; Ribeiro, J.M. Anopheline salivary protein genes and gene families: An evolutionary overview after the whole genome sequence of sixteen Anopheles species. BMC Genom. 2017, 18, 153. [Google Scholar] [CrossRef]
- Rizzo, C.; Ronca, R.; Fiorentino, G.; Mangano, V.D.; Sirima, S.B.; Nebie, I.; Petrarca, V.; Modiano, D.; Arca, B. Wide cross-reactivity between Anopheles gambiae and Anopheles funestus SG6 salivary proteins supports exploitation of gSG6 as a marker of human exposure to major malaria vectors in tropical Africa. Malar. J. 2011, 10, 206. [Google Scholar] [CrossRef]
- Stone, W.; Bousema, T.; Jones, S.; Gesase, S.; Hashim, R.; Gosling, R.; Carneiro, I.; Chandramohan, D.; Theander, T.; Ronca, R.; et al. IgG responses to Anopheles gambiae salivary antigen gSG6 detect variation in exposure to malaria vectors and disease risk. PLoS ONE 2012, 7, e40170. [Google Scholar] [CrossRef]
- Poinsignon, A.; Cornelie, S.; Ba, F.; Boulanger, D.; Sow, C.; Rossignol, M.; Sokhna, C.; Cisse, B.; Simondon, F.; Remoue, F. Human IgG response to a salivary peptide, gSG6-P1, as a new immuno-epidemiological tool for evaluating low-level exposure to Anopheles bites. Malar. J. 2009, 8, 198. [Google Scholar] [CrossRef]
- Coutinho-Abreu, I.V.; Guimaraes-Costa, A.B.; Valenzuela, J.G. Impact of Insect Salivary Proteins in Blood Feeding, Host Immunity, Disease, and in the Development of Biomarkers for Vector Exposure. Curr. Opin. Insect Sci. 2015, 10, 98–103. [Google Scholar] [CrossRef]
- Olano, V.A.; Carrillo, M.P.; de la Vega, P.; Espinal, C.A. Vector competence of Cartagena strain of Anopheles albimanus for Plasmodium falciparum and P. vivax. Trans. R. Soc. Trop. Med. Hyg. 1985, 79, 685–686. [Google Scholar] [CrossRef]
- Papa, F.; Windbichler, N.; Waterhouse, R.M.; Cagnetti, A.; D’Amato, R.; Persampieri, T.; Lawniczak, M.K.N.; Nolan, T.; Papathanos, P.A. Rapid evolution of female-biased genes among four species of Anopheles malaria mosquitoes. Genome Res. 2017, 27, 1536–1548. [Google Scholar] [CrossRef] [PubMed]
- Champagne, D.E.; Smartt, C.T.; Ribeiro, J.M.; James, A.A. The salivary gland-specific apyrase of the mosquito Aedes aegypti is a member of the 5′-nucleotidase family. Proc. Natl. Acad. Sci. USA 1995, 92, 694–698. [Google Scholar] [CrossRef]
- Choumet, V.; Carmi-Leroy, A.; Laurent, C.; Lenormand, P.; Rousselle, J.C.; Namane, A.; Roth, C.; Brey, P.T. The salivary glands and saliva of Anopheles gambiae as an essential step in the Plasmodium life cycle: A global proteomic study. Proteomics 2007, 7, 3384–3394. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, J.M.; Valenzuela, J.G. Purification and cloning of the salivary peroxidase/catechol oxidase of the mosquito Anopheles albimanus. J. Exp. Biol. 1999, 202, 809–816. [Google Scholar]
- Gutierrez, L.A.; Gomez, G.F.; Gonzalez, J.J.; Castro, M.I.; Luckhart, S.; Conn, J.E.; Correa, M.M. Microgeographic genetic variation of the malaria vector Anopheles darlingi root (Diptera: Culicidae) from Cordoba and Antioquia, Colombia. Am. J. Trop. Med. Hyg. 2010, 83, 38–47. [Google Scholar] [CrossRef]
- Rodrigues, P.T.; Valdivia, H.O.; de Oliveira, T.C.; Alves, J.M.P.; Duarte, A.; Cerutti-Junior, C.; Buery, J.C.; Brito, C.F.A.; de Souza, J.C., Jr.; Hirano, Z.M.B.; et al. Human migration and the spread of malaria parasites to the New World. Sci. Rep. 2018, 8, 1993. [Google Scholar] [CrossRef]
- Hemingway, J.; Shretta, R.; Wells, T.N.; Bell, D.; Djimde, A.A.; Achee, N.; Qi, G. Tools and Strategies for Malaria Control and Elimination: What Do We Need to Achieve a Grand Convergence in Malaria? PLoS Biol. 2016, 14, e1002380. [Google Scholar] [CrossRef]
- Maldonado-Ruiz, L.P.; Montenegro-Cadena, L.; Blattner, B.; Menghwar, S.; Zurek, L.; Londono-Renteria, B. Differential Tick Salivary Protein Profiles and Human Immune Responses to Lone Star Ticks (Amblyomma americanum) From the Wild vs. a Laboratory Colony. Front. Immunol. 2019, 10, 1996. [Google Scholar] [CrossRef]
- Ben Hadj Ahmed, S.; Chelbi, I.; Kaabi, B.; Cherni, S.; Derbali, M.; Zhioua, E. Differences in the salivary effects of wild-caught versus colonized Phlebotomus papatasi (Diptera: Psychodidae) on the development of zoonotic cutaneous leishmaniasis in BALB/c mice. J. Med. Entomol. 2010, 47, 74–79. [Google Scholar] [CrossRef][Green Version]
- Fontaine, A.; Diouf, I.; Bakkali, N.; Misse, D.; Pages, F.; Fusai, T.; Rogier, C.; Almeras, L. Implication of haematophagous arthropod salivary proteins in host-vector interactions. Parasit Vectors 2011, 4, 187. [Google Scholar] [CrossRef] [PubMed]
- Carlson, J.C.; Dyer, L.A.; Omlin, F.X.; Beier, J.C. Diversity cascades and malaria vectors. J. Med. Entomol. 2009, 46, 460–464. [Google Scholar] [CrossRef] [PubMed]
- Marie, A.; Ronca, R.; Poinsignon, A.; Lombardo, F.; Drame, P.M.; Cornelie, S.; Besnard, P.; Le Mire, J.; Fiorentino, G.; Fortes, F.; et al. The Anopheles gambiae cE5 salivary protein: A sensitive biomarker to evaluate the efficacy of insecticide-treated nets in malaria vector control. Microbes Infect. 2015, 17, 409–416. [Google Scholar] [CrossRef]
- Yoshiga, T.; Hernandez, V.P.; Fallon, A.M.; Law, J.H. Mosquito transferrin, an acute-phase protein that is up-regulated upon infection. Proc. Natl. Acad. Sci. USA 1997, 94, 12337–12342. [Google Scholar] [CrossRef] [PubMed]
- Rawal, R.; Vijay, S.; Kadian, K.; Adak, T.; Pande, V.; Sharma, A. Comparative proteomics of salivary glands of Anopheles culicifacies mosquitoes using tandem mass tag (TMT) mass spectrometry. J. Vector Borne Dis. 2018, 55, 98–110. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, M.H.; Hernandez-Hernandez Fde, L. Insect-malaria parasites interactions: The salivary gland. Insect Biochem. Mol. Biol. 2004, 34, 615–624. [Google Scholar] [CrossRef]
- Wells, M.B.; Andrew, D.J. Anopheles Salivary Gland Architecture Shapes Plasmodium Sporozoite Availability for Transmission. mBio 2019, 10. [Google Scholar] [CrossRef]
- Das, S.; Radtke, A.; Choi, Y.J.; Mendes, A.M.; Valenzuela, J.G.; Dimopoulos, G. Transcriptomic and functional analysis of the Anopheles gambiae salivary gland in relation to blood feeding. BMC Genom. 2010, 11, 566. [Google Scholar] [CrossRef]
- Gutierrez, L.A.; Naranjo, N.J.; Cienfuegos, A.V.; Muskus, C.E.; Luckhart, S.; Conn, J.E.; Correa, M.M. Population structure analyses and demographic history of the malaria vector Anopheles albimanus from the Caribbean and the Pacific regions of Colombia. Malar. J. 2009, 8, 259. [Google Scholar] [CrossRef] [PubMed]
- DSSA. Dirección Seccional de Salud de Antioquia. Eventos de Interés en Salud Pública Por Subregiones y Municipios. Antioquia 2007–2017. Available online: http://www.dssa.gov.co/index.php/estadisticas/eventos-en-salud-publica/item/71-eventos-de-interes-en-salud-publica-por-subregiones-y-municipios-antioquia-2007-2012 (accessed on 15 May 2019).
- Singh, B.; Bobogare, A.; Cox-Singh, J.; Snounou, G.; Abdullah, M.S.; Rahman, H.A. A genus—And species-specific nested polymerase chain reaction malaria detection assay for epidemiologic studies. Am. J. Trop. Med. Hyg. 1999, 60, 687–692. [Google Scholar] [CrossRef] [PubMed]
- Burtnick, M.N.; Brett, P.J.; DeShazer, D. Proteomic analysis of the Burkholderia pseudomallei type II secretome reveals hydrolytic enzymes, novel proteins, and the deubiquitinase TssM. Infect. Immun. 2014, 82, 3214–3226. [Google Scholar] [CrossRef]
- Hurst, K.E.; Lawrence, K.A.; Essman, M.T.; Walton, Z.J.; Leddy, L.R.; Thaxton, J.E. Endoplasmic Reticulum Stress Contributes to Mitochondrial Exhaustion of CD8(+) T Cells. Cancer Immunol. Res. 2019, 7, 476–486. [Google Scholar] [CrossRef] [PubMed]
- Bronisz, A.; Wang, Y.; Nowicki, M.O.; Peruzzi, P.; Ansari, K.; Ogawa, D.; Balaj, L.; De Rienzo, G.; Mineo, M.; Nakano, I.; et al. Extracellular vesicles modulate the glioblastoma microenvironment via a tumor suppression signaling network directed by miR-1. Cancer Res. 2014, 74, 738–750. [Google Scholar] [CrossRef] [PubMed]
- Londono-Renteria, B.; Drame, P.M.; Weitzel, T.; Rosas, R.; Gripping, C.; Cardenas, J.C.; Alvares, M.; Wesson, D.M.; Poinsignon, A.; Remoue, F.; et al. An. gambiae gSG6-P1 evaluation as a proxy for human-vector contact in the Americas: A pilot study. Parasit Vectors 2015, 8, 533. [Google Scholar] [CrossRef] [PubMed]
| Protein ID | Description | CTG | STE | Length | Molecular Weight (Da) |
|---|---|---|---|---|---|
| A0A182FAJ2 | Transferrin | YES | YES | 532 | 59.390 |
| A0A182FH19 | Uncharacterized protein | YES | NO | 561 | 60.603 |
| A0A182FTN8 | Uncharacterized protein | YES | YES | 568 | 63.192 |
| A0A182FP42 | Uncharacterized protein | YES | YES | 573 | 63.502 |
| Q9XYP9 | Salivary peroxidase | NO | YES | 591 | 65.445 |
| A0A1Y9G8H0 | Uncharacterized protein | YES | YES | 592 | 65.508 |
| A0A1Y9G8K4 | Uncharacterized protein | YES | YES | 593 | 66.000 |
| A0A1Y9G9T7 | Uncharacterized protein | YES | NO | 589 | 66.231 |
| A0A1Y9G9L7 | Uncharacterized protein | YES | NO | 589 | 66.339 |
| Protein ID | Description | Length | Mass | Peptide Name | Peptide Sequence | Amino-Acid Position |
|---|---|---|---|---|---|---|
| A0A182FAJ2 | Transferrin | 532 | 59.390 | TRANS-P1 | YSPNADIDGLMKKRYSNL | 185–202 |
| TRANS_P2 | SYLCEDGTTRPVSDQNVC | 271–288 | ||||
| A0A1Y9G8H0/Q9XYP9 | Salivary peroxidase | 592 | 65.508 | PEROX-P1 | RTITDCDADPSSCSNSKKAE | 162–181 |
| PEROX-P2 | MKVETRDGSDWPPRNPNAST | 214–233 | ||||
| 591 | 65.445 | PEROX-P3 | QRARDHGLPSYNSFREKCGL | 434–453 |
| Status | Turbo | El Bagre | Total | ||
|---|---|---|---|---|---|
| Females | Males | Females | Males | ||
| Malaria + | 15 | 29 | 23 | 45 | 112 |
| Malaria − | 63 | 44 | 67 | 50 | 224 |
| Total | 78 | 73 | 90 | 95 | 336 * |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Londono-Renteria, B.; Drame, P.M.; Montiel, J.; Vasquez, A.M.; Tobón-Castaño, A.; Taylor, M.; Vizcaino, L.; Lenhart, A.E. Identification and Pilot Evaluation of Salivary Peptides from Anopheles albimanus as Biomarkers for Bite Exposure and Malaria Infection in Colombia. Int. J. Mol. Sci. 2020, 21, 691. https://doi.org/10.3390/ijms21030691
Londono-Renteria B, Drame PM, Montiel J, Vasquez AM, Tobón-Castaño A, Taylor M, Vizcaino L, Lenhart AE. Identification and Pilot Evaluation of Salivary Peptides from Anopheles albimanus as Biomarkers for Bite Exposure and Malaria Infection in Colombia. International Journal of Molecular Sciences. 2020; 21(3):691. https://doi.org/10.3390/ijms21030691
Chicago/Turabian StyleLondono-Renteria, Berlin, Papa M. Drame, Jehidys Montiel, Ana M. Vasquez, Alberto Tobón-Castaño, Marissa Taylor, Lucrecia Vizcaino, and Audrey E. Lenhart. 2020. "Identification and Pilot Evaluation of Salivary Peptides from Anopheles albimanus as Biomarkers for Bite Exposure and Malaria Infection in Colombia" International Journal of Molecular Sciences 21, no. 3: 691. https://doi.org/10.3390/ijms21030691
APA StyleLondono-Renteria, B., Drame, P. M., Montiel, J., Vasquez, A. M., Tobón-Castaño, A., Taylor, M., Vizcaino, L., & Lenhart, A. E. (2020). Identification and Pilot Evaluation of Salivary Peptides from Anopheles albimanus as Biomarkers for Bite Exposure and Malaria Infection in Colombia. International Journal of Molecular Sciences, 21(3), 691. https://doi.org/10.3390/ijms21030691






