Overexpression of Tamarix hispida ThTrx5 Confers Salt Tolerance to Arabidopsis by Activating Stress Response Signals
Abstract
1. Introduction
2. Results
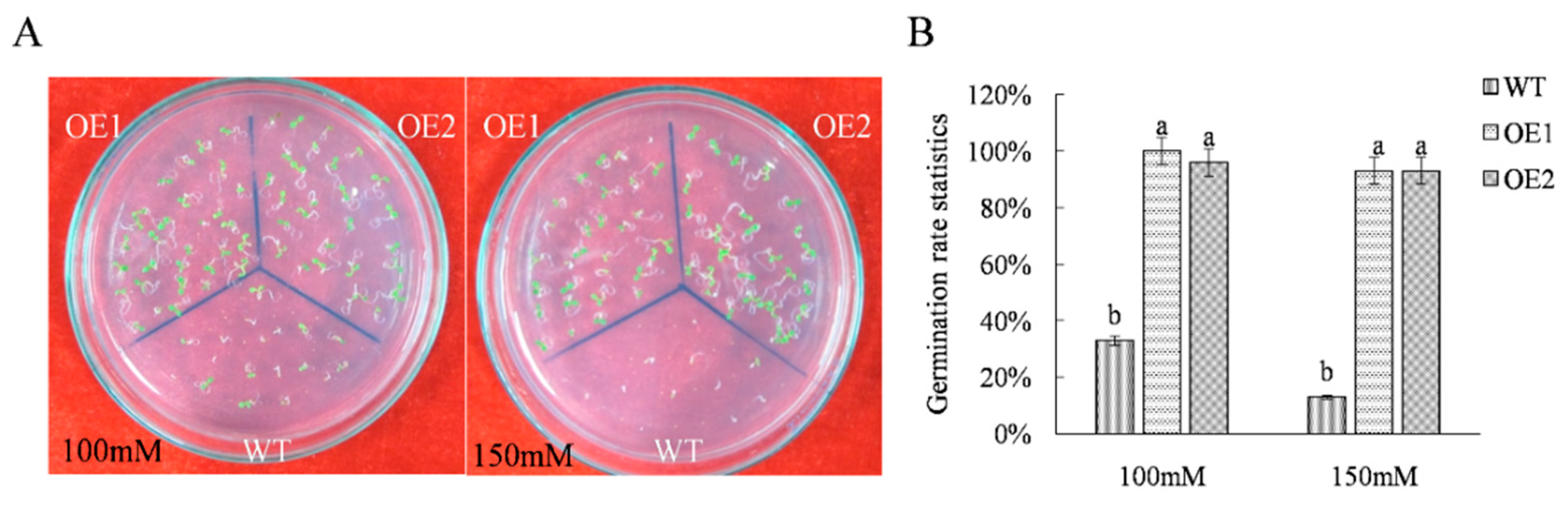
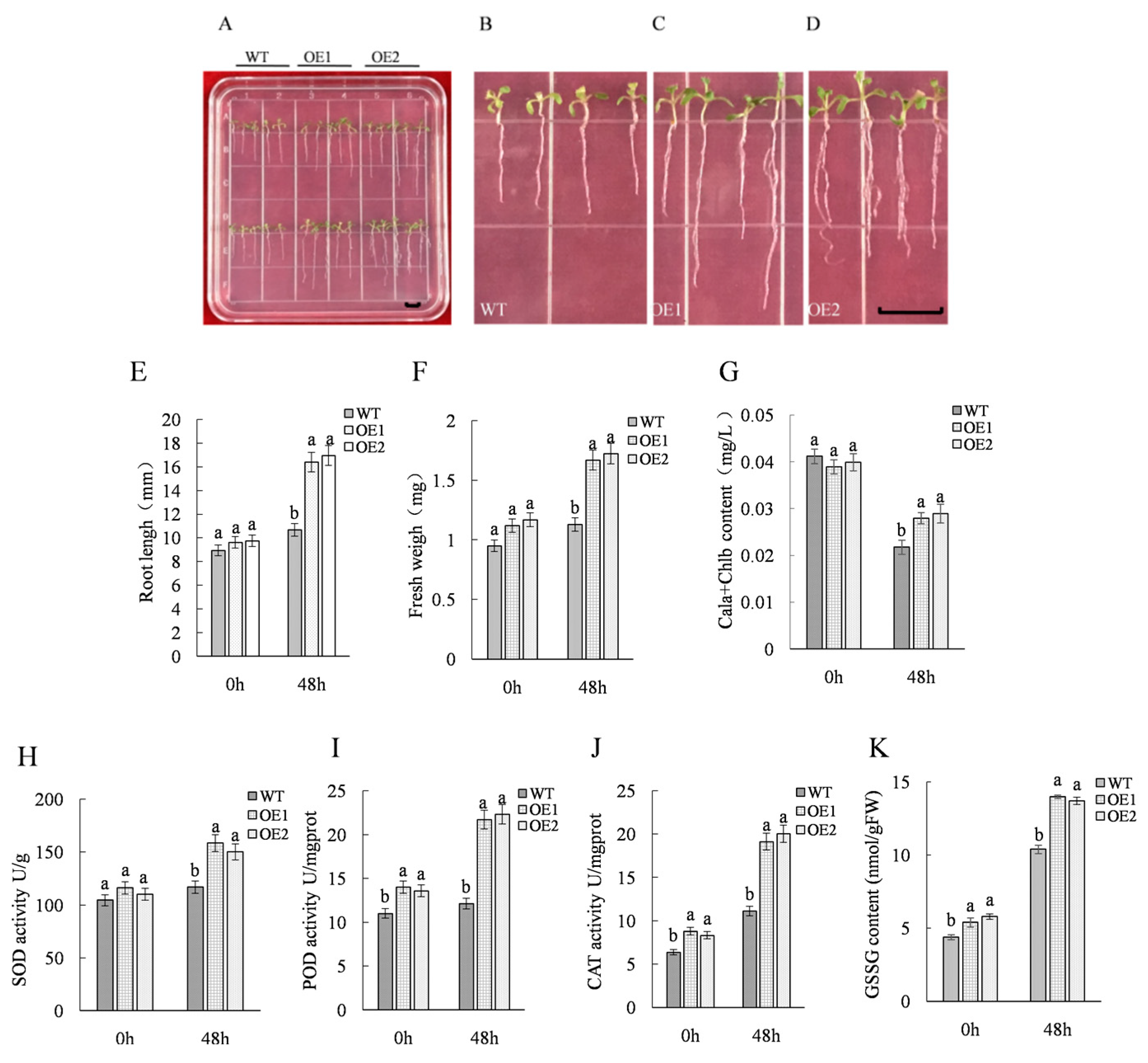
2.1. Acquisition and Salt Resistance Analysis of ThTrx5-Overexpressing Transgenic Lines
2.2. Screening of Interacting Proteins
Pull Down
2.3. Transcriptome Analysis
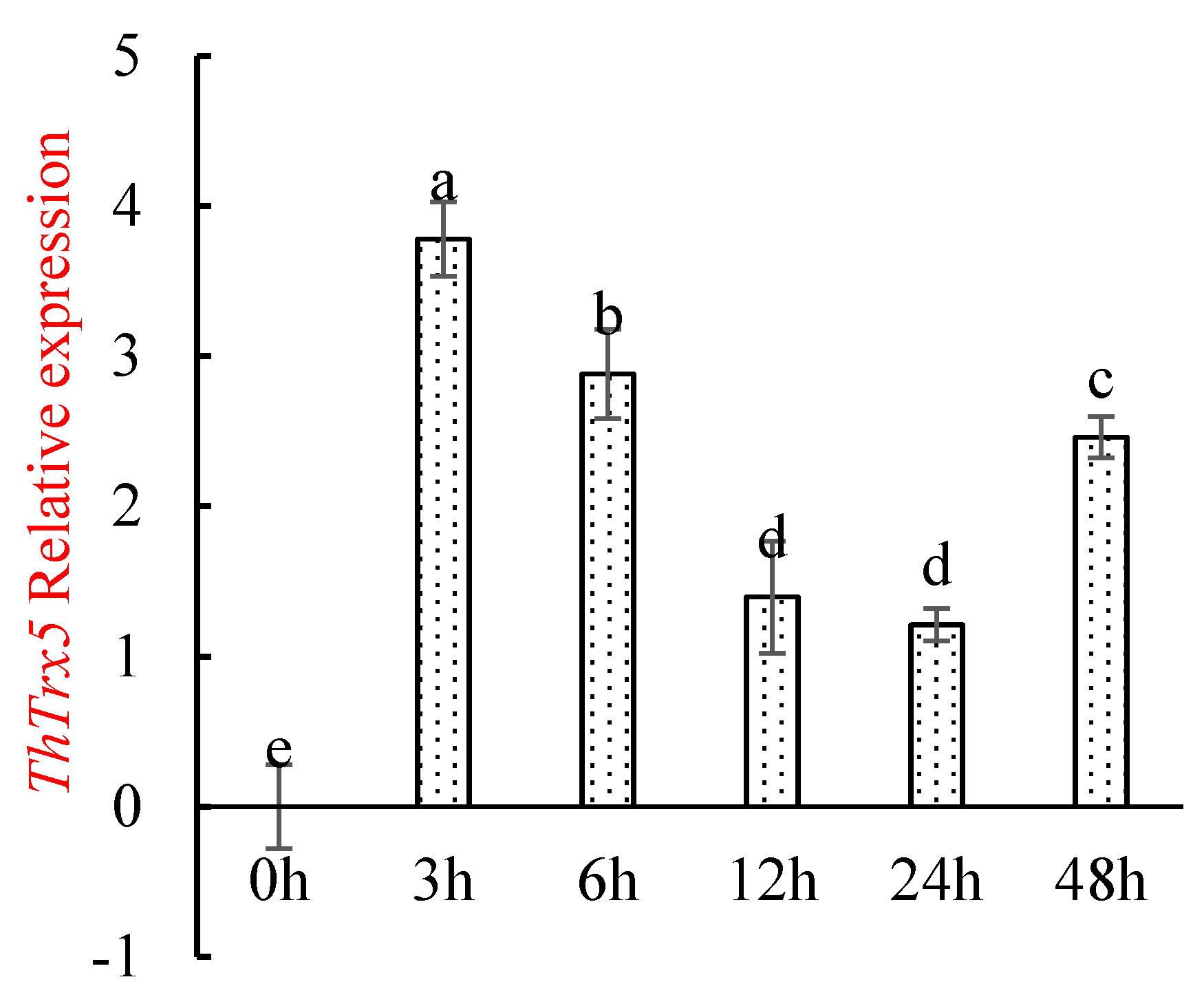
2.3.1. Changes in ThTrx5 Gene Expression in Tamarix Hispida under NaCl Stress
2.3.2. Illumina Sequencing
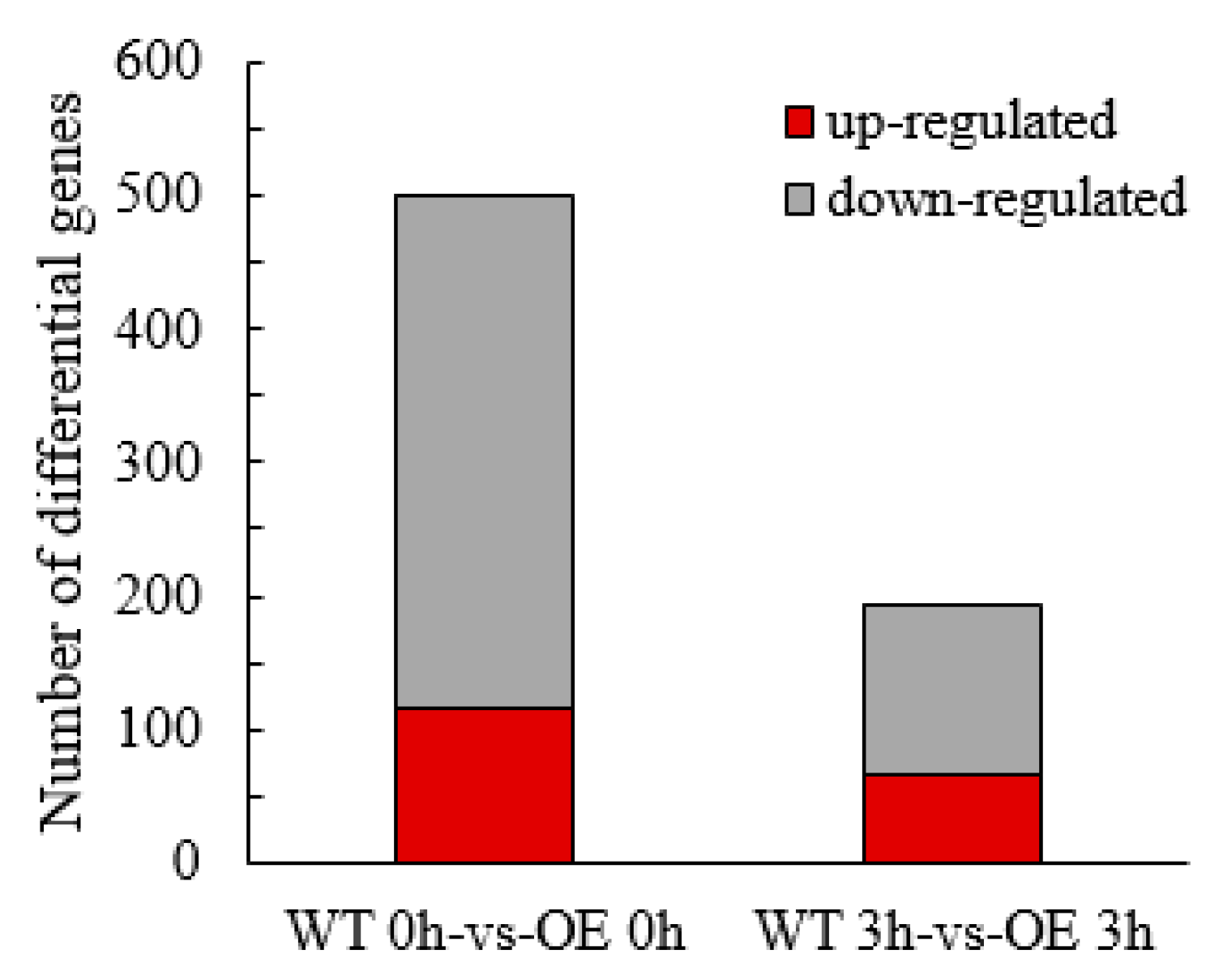
2.3.3. Screening and Analysis of Differentially Expressed Genes
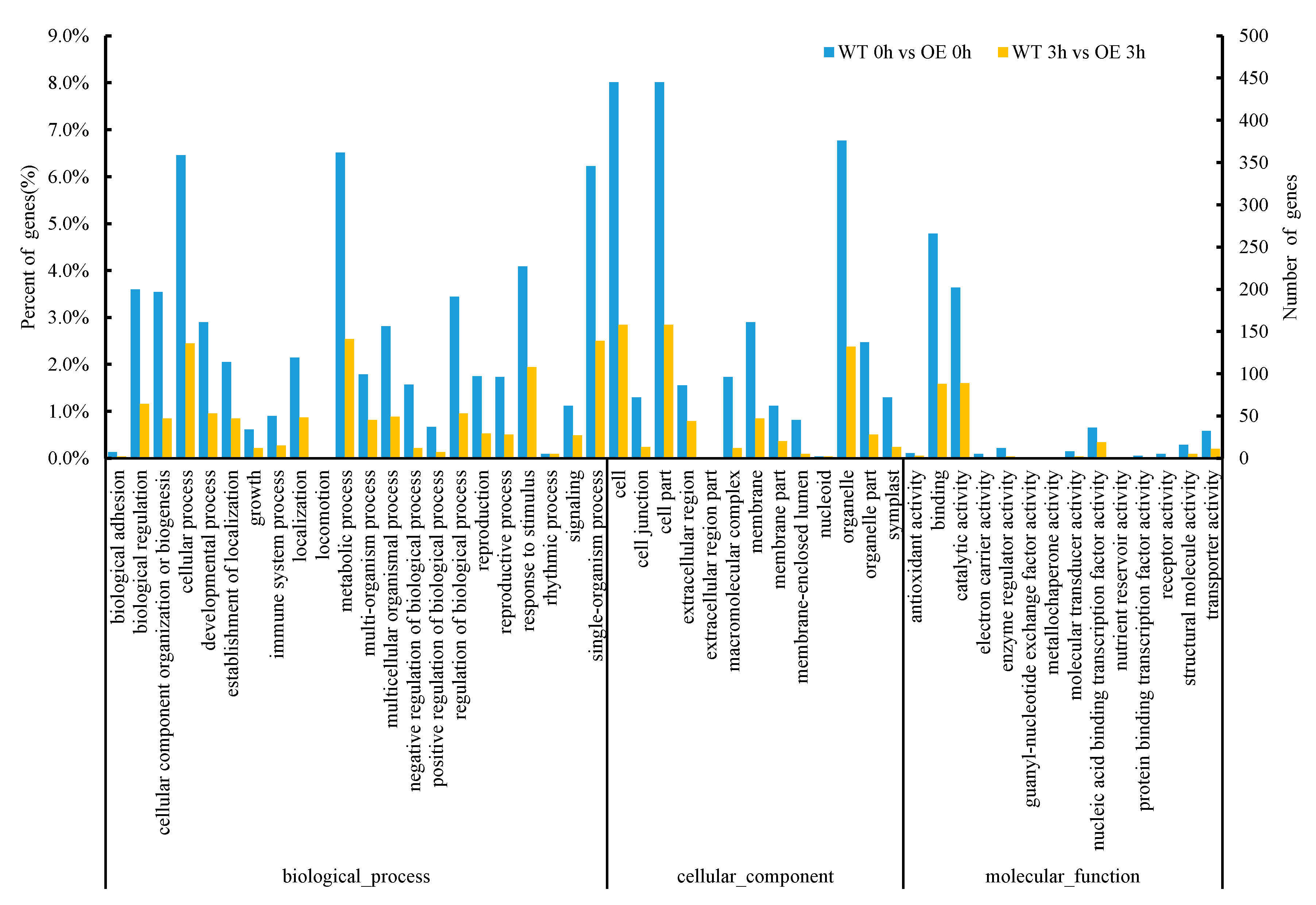
2.3.4. Analysis of GO (Gene Ontology) Enrichment of DEGs
2.3.5. Analysis of Transcription Factor (TF) Families Represented by DEGs Identified during Salt Stress Exposure
2.3.6. Analysis of DEGs Involved in Hormone Synthesis and Signal Transduction Pathways Triggered by Salt Stress Exposure
2.3.7. Analysis of DEGs Involved in Redox-Based Reaction Processes upon Salt Stress Exposure
2.3.8. qRT-PCR Analysis
3. Discussion
3.1. Overexpressing ThTRX5 Improves Salt Resistance of Arabidopsis Plants
3.2. Arabidopsis Proteins Interacting with ThTRX5 Participate with Stress Resistance
3.3. ThTrx5 Participates in the Redox Process, TF Expression, and Hormone Signal Transduction
3.4. ThTrx5 Regulates Biological and Metabolic Pathways via Protein Interactions
4. Materials and Methods
4.1. Materials
4.1.1. Plant Materials and Growth Conditions
4.1.2. Processing Method
4.2. Gene Cloning and Vector Construction
4.2.1. ThTrx5 Gene Cloning and Expression Vector Construction
4.2.2. Construction of Fusion Protein Expression Vector
4.2.3. qPT-PCR
4.2.4. Data Analysis
4.3. Preparation of Engineered Bacteria and Arabidopsis Transformation
4.4. Screening of Interacting Proteins
4.4.1. Induction of Protein Expression
4.4.2. Pull-Down
4.4.3. Mass Spectrometry
4.5. Transcriptome Analysis
4.5.1. RNA Extraction and Differential Expression Library Construction
4.5.2. De Novo Assembly and Functional Annotation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| BLAST | the Basic Local Alignment Search Tool |
| DEGs | differentially expressed genes |
| GO | Gene ontology |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| SwissProt | Swiss-prot sequence data bases |
| ORFs | Open reading frames |
References
- Wei, L.; Shu, H.X.; Kun, L.P.; Qin, Z.H.; Chao, W.T. Research Progress in Plant Thioredoxin. J. Guizhou Agric. Sci. 2006, 34, 129–131. [Google Scholar]
- Meyer, Y.; Buchanan, B.B.; Vignols, F.; Reichheld, J.-P. Thioredoxins and Glutaredoxins: Unifying Elements in Redox Biology. Annu. Rev. Genet. 2009, 43, 335–367. [Google Scholar] [CrossRef] [PubMed]
- Michelet, L.; Zaffagnini, M.; Morisse, S.; Sparla, F.; Pérez-Pérez, M.E.; Francia, F.; Danon, A.; Marchand, C.H.; Fermani, S.; Trost, P.; et al. Redox regulation of the Calvin–Benson cycle: Something old, something new. Front. Plant Sci. 2013, 4, 470. [Google Scholar] [CrossRef]
- Buchanan, B.B. The Path to Thioredoxin and Redox Regulation in Chloroplasts. Annu. Rev. Plant Biol. 2016, 67, 1–24. [Google Scholar] [CrossRef]
- Shaykholeslam Esfahani, E.; Shahpiri, A. Thioredoxin h isoforms from rice are differentially reduced by NADPH/thioredoxin or GSH/glutaredoxin systems. Int. J. Biol. Macromol. 2015, 74, 243–248. [Google Scholar] [CrossRef]
- Gelhaye, E.; Rouhier, N.; Navrot, N.; Jacquot, J.-P. The plant thioredoxin system. Cell. Mol. Life Sci. 2005, 62, 24–35. [Google Scholar] [CrossRef]
- Mouaheb, N.; Thomas, M.; Verdoucq, L.; Monfort, P.; Meyer, Y. In vivo functional discrimination between plant thioredoxins by heterologous expression in the yeast Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 1998, 95, 3312–3317. [Google Scholar] [CrossRef]
- Murata, Y.; Pei, Z.M.; Mori, I.C.; Schroeder, J. Abscisic Acid Activation of Plasma Membrane Ca2+ Channels in Guard Cells Requires Cytosolic NAD(P)H and Is Differentially Disrupted Upstream and Downstream of Reactive Oxygen Species Production in abi1-1 and abi2-1 Protein Phosphatase 2C Mutants. Plant Cell 2001, 13, 2513–2523. [Google Scholar] [CrossRef]
- Gilbert, H.F. Molecular and cellular aspects of thiol-disulfide exchange. Adv. Enzymol. Relat. Areas Mol. Boil. 1990, 63, 69–172. [Google Scholar]
- Jaeger, T.; Flohé, L. The thiol-based redox networks of pathogens: Unexploited targets in the search for new drugs. BioFactors 2006, 27, 109–120. [Google Scholar] [CrossRef]
- Laloi, C.; Rayapuram, N.; Chartier, Y.; Grienenberger, J.-M.; Bonnard, G.; Meyer, Y. Identification and characterization of a mitochondrial thioredoxin system in plants. Proc. Natl. Acad. Sci. USA 2001, 98, 14144–14149. [Google Scholar] [CrossRef]
- Wang, P.; Liu, J.; Liu, B.; Feng, D.; Da, Q.; Shu, S.; Su, J.; Zhang, Y.; Wang, J.; Wang, H.-B. Evidence for a Role of Chloroplastic m-Type Thioredoxins in the Biogenesis of Photosystem II in Arabidopsis. Plant Physiol. 2013, 163, 1710–1728. [Google Scholar] [CrossRef] [PubMed]
- Hui-Yu, L.; Kun, N.; Xin, S.; Zi-Jia, W.; Gui-Feng, L.; Jing, J.; Rui-Ping, Z.; Kai-Long, L. Molecular Cloning and Expression Analysis of Nine ThTrx Genes in Tamarix hispida. Plant Mol. Boil. Rep. 2013, 31, 917–924. [Google Scholar] [CrossRef]
- Mangelsen, E.; Kilian, J.; Berendzen, K.W.; Kolukisaoglu, Ü.H.; Harter, K.; Jansson, C.; Wanke, D. Phylogenetic and comparative gene expression analysis of barley (Hordeum vulgare) WRKY transcription factor family reveals putatively retained functions between monocots and dicots. BMC Genom. 2008, 9, 194. [Google Scholar] [CrossRef]
- Kim, Y.-C.; Masutani, H.; Yamaguchi, Y.; Itoh, K.; Yamamoto, M.; Yodoi, J. Hemin-induced Activation of the Thioredoxin Gene by Nrf2. J. Boil. Chem. 2001, 276, 18399–18406. [Google Scholar] [CrossRef]
- Kanzok, S.M.; Fechner, A.; Bauer, H.; Ulschmid, J.K.; Müller, H.M.; Botella-Munoz, J.; Schneuwly, S.; Schirmer, R.H.; Becker, K. Substitution of the Thioredoxin System for Glutathione Reduetase in Drosophila melanogaster. Science 2001, 291, 643–645. [Google Scholar] [CrossRef]
- Foyer, C.H.; Noctor, G. Managing the cellular redox hub in photosynthetic organismspce. Plant Cell Environ. 2012, 35, 199–201. [Google Scholar] [CrossRef]
- Bello, B.; Zhang, X.; Liu, C.; Yang, Z.; Yang, Z.; Wang, Q.; Zhao, G.; Li, F. Cloning of Gossypium hirsutum Sucrose Non-Fermenting 1-Related Protein Kinase 2 Gene (GhSnRK2) and Its Overexpression in Transgenic Arabidopsis Escalates Drought and Low Temperature Tolerance. PLoS ONE 2014, 9, e112269. [Google Scholar] [CrossRef]
- Baier, M. Protective Function of Chloroplast 2-Cysteine Peroxiredoxin in Photosynthesis. Evidence from Transgenic Arabidopsis. Plant Physiol. 1999, 119, 1407–1414. [Google Scholar] [CrossRef][Green Version]
- Baier, M.; Noctor, G.; Foyer, C.H.; Dietz, K.-J. Antisense suppression of 2-Cys peroxiredoxin in Arabidopsis specifically enhances the activities and expression of enzymes associated with ascorbate metabolism but not glutathione metabolism. Plant Physiol. 2000, 124, 823–832. [Google Scholar] [CrossRef]
- Baier, M.; Dietz, K.-J. The plant 2-Cys peroxiredoxin BAS1 is a nuclear-encoded chloroplast protein: Its expressional regulation, phylogenetic origin, and implications for its specific physiological function in plants. Plant J. 1997, 12, 179–190. [Google Scholar] [CrossRef]
- König, J.; Baier, M.; Horling, F.; Kahmann, U.; Harris, G.; Schürmann, P.; Dietz, K.-J. The plant-specific function of 2-Cys peroxiredoxin-mediated detoxification of peroxides in the redox-hierarchy of photosynthetic electron flux. Proc. Natl. Acad. Sci. USA 2002, 99, 5738–5743. [Google Scholar] [CrossRef]
- König, J.; Lotte, K.; Plessow, R.; Brockhinke, A.; Baier, M.; Dietz, K.-J. Reaction mechanism of the 2-Cys peroxiredoxin: Role of the C-terminus and the quarternary structure. Biol. Chem. 2003, 278, 24409–24420. [Google Scholar] [CrossRef]
- Baier, M.; Stroher, E.; Karl-Josef, D. The Acceptor Availability at Photosystem I and ABA Control Nuclear Expression of 2-Cys Peroxiredoxin-A in Arabidopsis thaliana. Plant Cell Physiol. 2004, 45, 997. [Google Scholar] [CrossRef]
- Nonn, L.; Berggren, M.; Powis, G. Increased Expression of Mitochondrial Peroxiredoxin-3 (Thioredoxin Peroxidase-2) Protects Cancer Cells against Hypoxia and Drug-Induced Hydrogen Peroxide-Dependent Apoptosis11CA52995 and CA772049. Mol. Cancer Res. 2003, 1, 682–689. [Google Scholar]
- Tartaglia, L.A.; Storz, G.; Brodsky, M.H.; Lai, A.; Ames, B.N. Alkyl hydroperoxide reductase from Salmonella typhimurium. Sequence and homology to thioredoxin reductase and other flavoprotein disulfide oxidoreductases. J. Boil. Chem. 1990, 265, 10535–10540. [Google Scholar]
- Chae, H.Z.; Chung, S.J.; Rhee, S.G. Thioredoxin-Dependent Peroxide Reductase from Yeast. J. Boil. Chem. 1994, 269, 27670–27678. [Google Scholar]
- Pillai, M.A.; Lihuang, Z.; Akiyama, T. Molecular cloning, characterization, expression and chromosomal location of OsGAPDH, a submergence responsive gene in rice (Oryza sativa L.). Theor. Appl. Genet. 2002, 105, 34–42. [Google Scholar] [CrossRef]
- Chinnusamy, V.; Zhu, J.; Zhu, J.-K. Salt stress signaling and mechanisms of plant salt tolerance. Genet. Eng. 2006, 27, 141–177. [Google Scholar]
- Kohli, A.; Sreenivasulu, N.; Lakshmanan, P.; Kumar, P.P. The phytohormone crosstalk paradigm takes center stage in understanding how plants respond to abiotic stresses. Plant Cell Rep. 2013, 32, 945–957. [Google Scholar] [CrossRef]
- Dinneny, J.R. A gateway with a guard: How the endodermis regulates growth through hormone signaling. Plant Sci. 2014, 214, 14–19. [Google Scholar] [CrossRef] [PubMed]
- Kariola, T.; Brader, G.; Helenius, E.; Li, J.; Heino, P.; Palva, E.T. Early Responsive To Dehydration 15, a Negative Regulator of Abscisic Acid Responses in Arabidopsis1. Plant Physiol. 2006, 142, 1559–1573. [Google Scholar] [CrossRef]
- Kazan, K. Diverse roles of jasmonates and ethylene in abiotic stress tolerance. Trends Plant Sci. 2015, 20, 219–229. [Google Scholar] [CrossRef] [PubMed]
- Mahajan, S.; Tuteja, N. Cold, salinity and drought stresses: An overview. Arch. Biochem. Biophys. 2005, 444, 139–158. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Ji, S.; Fang, X.; Wang, Q.; Li, Z.; Yao, F.; Hou, L.; Dai, S. Protein Kinase LTRPK1 Influences Cold Adaptation and Microtubule Stability in Rice. J. Plant Growth Regul. 2013, 32, 483–490. [Google Scholar] [CrossRef]
- Nakashima, K.; Takasaki, H.; Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. NAC transcription factors in plant abiotic stress responses. Biochim. Biophys. Acta (BBA) 2012, 1819, 97–103. [Google Scholar] [CrossRef]
- Wang, Y.X. Characterization of a novel Medicago sativa NAC transcription factor gene involved in response to drought stress. Mol. Biol. Rep. 2013, 40, 6451–6458. [Google Scholar] [CrossRef]
- Niu, Y.; Figueroa, P.; Browse, J. Characterization of JAZ-interacting bHLH transcription factors that regulate jasmonate responses in Arabidopsis. J. Exp. Bot. 2011, 62, 2143–2154. [Google Scholar] [CrossRef]
- Jin, P.; Huang, L.; Wang, D.; Wu, H.; Zhu, L.; Fu, B. Expression profiling of rice AP2/EREBP genes responsive to abiotic stresses. Sci. Agric. Sin. 2009, 42, 3765–3773. [Google Scholar]
- Zhou, Q.Y.; Tian, A.G.; Zou, H.F.; Xie, Z.M.; Lei, G.; Huang, J.; Wang, C.M.; Wang, H.W.; Zhang, J.S.; Chen, S.Y. Soybean WRKY-type transcription factor genes, GmWRKY13, GmWRKY21, and GmWRKY54, confer differential tolerance to abiotic stresses in transgenic Arabidopsis plants. Plant Biotechnol. J. 2010, 6, 486–503. [Google Scholar] [CrossRef]
- Marker, A.F.H. The use of acetone and methanol in the estimation of chlorophyll in the presence of phaeophytin. Freshw. Boil. 1972, 2, 361–385. [Google Scholar] [CrossRef]
- Gambino, G.; Perrone, I.; Gribaudo, I. A Rapid and effective method for RNA extraction from different tissues of grapevine and other woody plants. Phytochem. Anal. 2008, 19, 520–525. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]




| Sample ID | WT-0h | WT-3h | OE-0h | OE-3h |
|---|---|---|---|---|
| Total Reads | 7,639,214(100.00%) | 7,499,090(100.00%) | 7,165,453(100.00%) | 7,318,039(100.00%) |
| Total BasePairs | 366,468,011(100.00%) | 357,615,524(100.00%) | 346,511,389(100.00%) | 358,182,356(100.00%) |
| Total Mapped Reads | 4,547,338(60.80%) | 5,339,751(73.16%) | 4,739,829(67.03%) | 5,396,632(73.83%) |
| Perfect Match | 4,063,215(54.33%) | 4,756,366(65.17%) | 4,241,363(59.98%) | 4,835,471(66.15%) |
| <=2bp Mismatch | 484,123(6.47%) | 583,385(7.99%) | 498,466(7.05%) | 561,161(7.68%) |
| Unique Match | 4,291,451(57.38%) | 5,119,890(70.15%) | 4,490,200(63.50%) | 5,162,943(70.63%) |
| Multi-position Match | 255,887(3.42%) | 219,861(3.01%) | 249,629(3.53%) | 233,689(3.20%) |
| Total Unmapped Reads | 2,931,601(39.20%) | 1,958,525(26.84%) | 2,331,832(32.97%) | 1,913,212(26.17%) |
| Clean Reads | 7,478,939 | 7,298,276 | 7,071,661 | 7,309,844 |
| Hormone | Gene | Log2 (WT 0 h/Trx 0 h) | Log2 (WT 3 h/Trx 3 h) | Gene Annotation |
|---|---|---|---|---|
| ABA | AT1G52040.1 | −1.466825674 | −1.650566816 | myrosinase-binding protein 1 |
| BR | AT4G34650.1 | 0.06962026 | −1.262992532 | squalene synthase 2 |
| ETH | AT5G47220.1 | −0.468665009 | 1.963721496 | ethylene-responsive transcription factor 2 |
| AT2G20880.1 | 0.519653186 | −1.327806826 | ethylene-responsive transcription factor ERF053 | |
| AT1G69260.1 | −0.387237409 | −1.327806826 | ABI five binding protein | |
| JA | AT3G55970.1 | −0.457626737 | −2.592176442 | jasmonate-regulated protein JRG21 |
| AT3G50760.1 | 0.502974445 | −1.129387909 | Probable galacturonosyltransferase-like 2 | |
| AT5G24780.1 | −2.57480625 | −2.023514997 | acid phosphatase VSP1 | |
| AT5G24770.1 | −0.608451639 | −1.443297712 | acid phosphatase VSP2 | |
| AT2G39030.1 | −3.23523431 | −2.243926852 | L-ornithine N5-acetyltransferase NATA1 | |
| 6-BA | AT1G21310.1 | −0.4968619 | 1.346863463 | response to cytokinin stimulus |
| SA | AT3G20470.1 | 0.527147723 | 1.015449778 | glycine-rich protein 5 |
| Primer Name | Forward Primer(5′→3′) | Reverse Primer(5′→3′) |
|---|---|---|
| α-Tubulin | 5′-GCACTGGCCTCCAAGGAT-3′ | 5′-TGGGTCGCTCAATGTCAAGG-3′ |
| ThTrx5 | 5′-AATCTATGATTGCTCCTTCC-3′ | 5′-GCCAGCTCATTAACAACC-3′ |
| AT4G25100.2 | 5′-GATGCTTTGGAGCCGCATATG-3′ | 5′-GAAGAACTCGTGGTTCCACG-3′ |
| AT4G34400.1 | 5′-CAGCTCAGAGTTCATGGTGATC-3′ | 5′-GTGAGCTCCATTATAGGCAAAGG-3′ |
| AT1G44970.1 | 5′-CGTGAGTGGATTCCCTAAACAATC-3′ | 5′-CACTCCAAACCCAGAGATTGG-3′ |
| AT4G37990.1 | 5′-CGAAAGACAATTCCGGAGTTCTC-3′ | 5′-CAGTCACCACGCCCACGATC-3′ |
| AT4G36190.1 | 5′-GGTTACTGCAACCTCGTAGGATTTC-3′ | 5′-GGTGATTCCATTGCAAGGTCC-3′ |
| AT4G37410 | 5′-CCTCATCAAACCGCCGGTTC-3′ | 5′-AGCAATGTATTTGGAGGTTAGAAAACG-3′ |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luan, J.; Dong, J.; Song, X.; Jiang, J.; Li, H. Overexpression of Tamarix hispida ThTrx5 Confers Salt Tolerance to Arabidopsis by Activating Stress Response Signals. Int. J. Mol. Sci. 2020, 21, 1165. https://doi.org/10.3390/ijms21031165
Luan J, Dong J, Song X, Jiang J, Li H. Overexpression of Tamarix hispida ThTrx5 Confers Salt Tolerance to Arabidopsis by Activating Stress Response Signals. International Journal of Molecular Sciences. 2020; 21(3):1165. https://doi.org/10.3390/ijms21031165
Chicago/Turabian StyleLuan, Jiayu, Jingxiang Dong, Xin Song, Jing Jiang, and Huiyu Li. 2020. "Overexpression of Tamarix hispida ThTrx5 Confers Salt Tolerance to Arabidopsis by Activating Stress Response Signals" International Journal of Molecular Sciences 21, no. 3: 1165. https://doi.org/10.3390/ijms21031165
APA StyleLuan, J., Dong, J., Song, X., Jiang, J., & Li, H. (2020). Overexpression of Tamarix hispida ThTrx5 Confers Salt Tolerance to Arabidopsis by Activating Stress Response Signals. International Journal of Molecular Sciences, 21(3), 1165. https://doi.org/10.3390/ijms21031165




