Genomic Instability Is an Early Event in Aluminium-Induced Tumorigenesis
Abstract
1. Introduction
2. Results
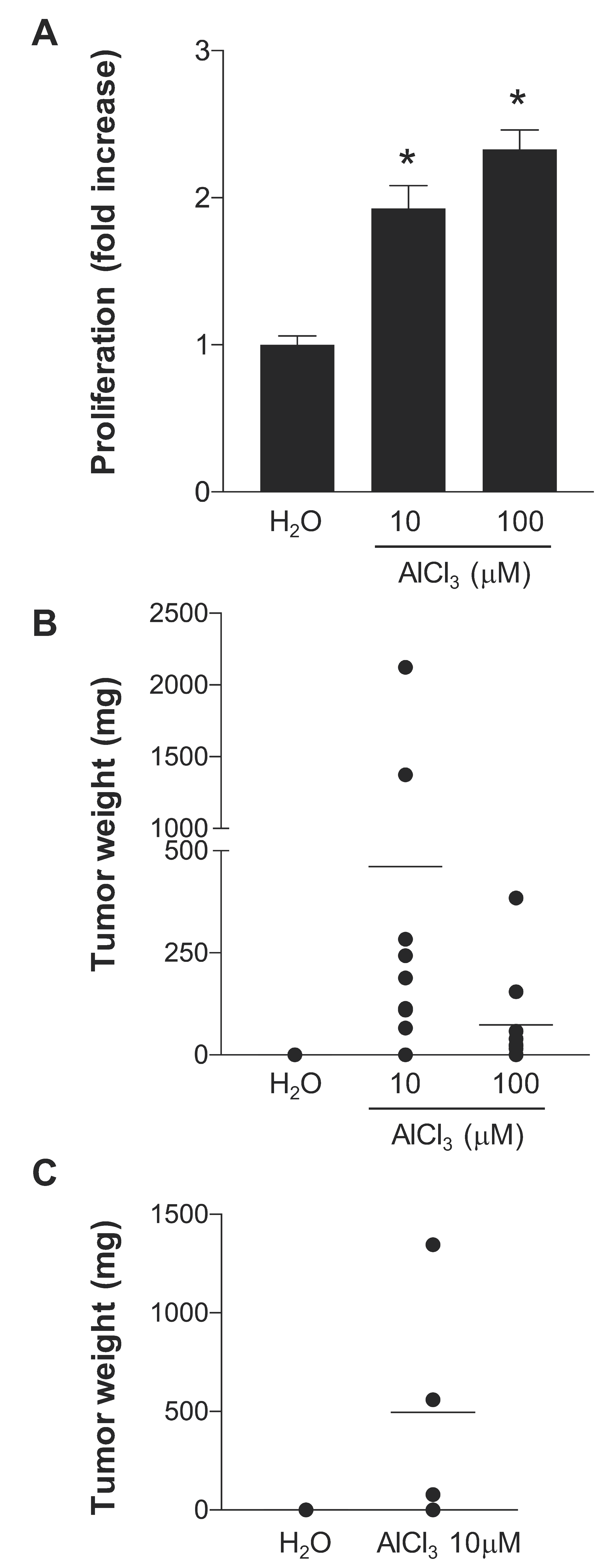
2.1. AlCl3 Transforms HC11 Mouse Mammary Epithelial Cells In Vitro
2.2. Analysis of mRNA Expression Patterns in AlCl3-Treated HC11 or NMuMG Cells
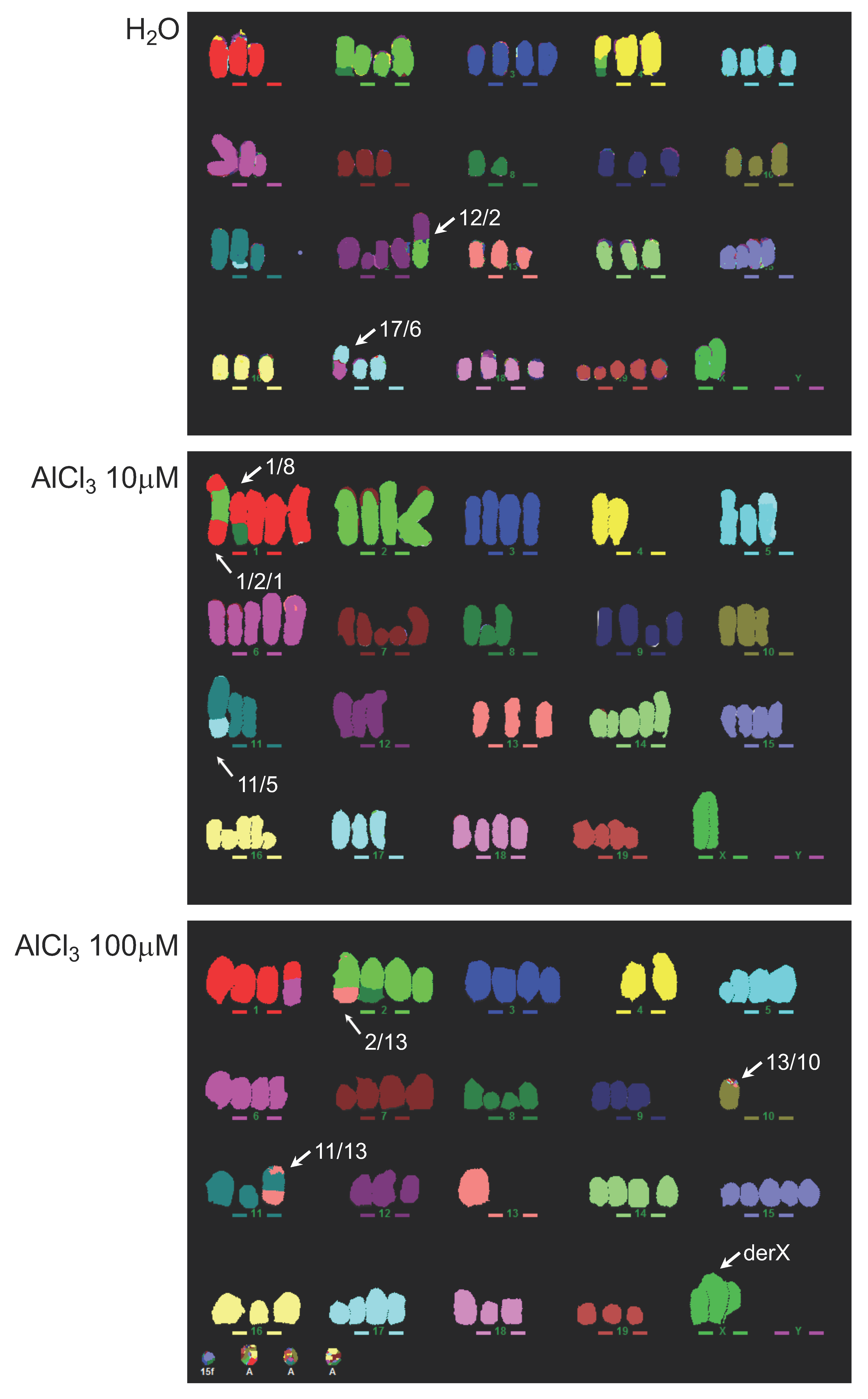
2.3. Analysis of Genomic Instability in AlCl3-Transformed HC11 or NMuMG Cells by Multicolour Fluorescence In Situ Hybridization (MFISH)
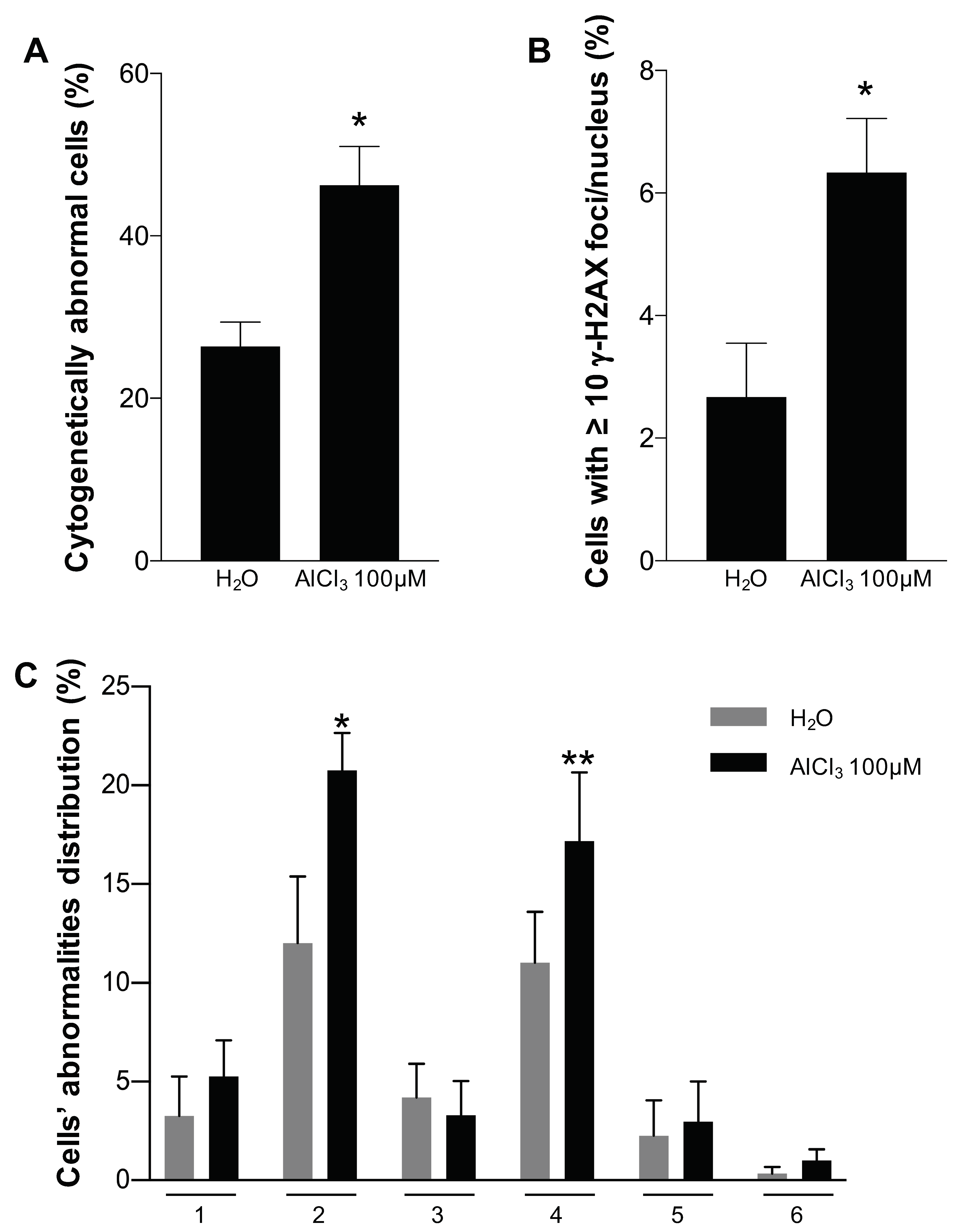
2.4. Analysis of Chromosomal Structural Abnormalities after Short AlCl3 Exposure
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. Growth-In-Low-Attachment (GILA) Assay
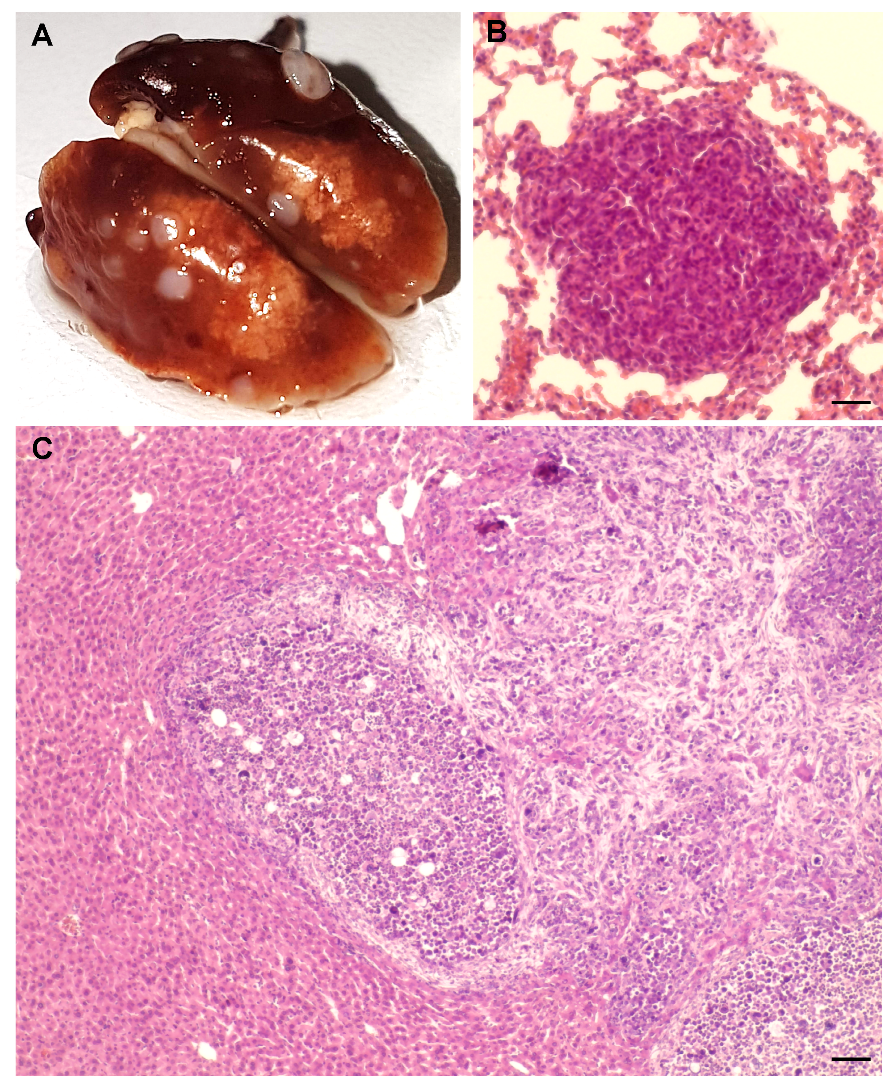
4.3. Syngeneic Grafts
4.4. Ki67 Immunohistochemistry
4.5. RNA Analysis
4.6. Real-Time Quantitative PCR
4.7. MFISH
4.8. Quantification of γ-H2AX Foci and Chromosome Analysis after Short Aluminium Exposure
4.9. Data Availability
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AlCl3 | Aluminium chloride |
| DAPI | 4′,6-Diamidino-2-phenylindole dihydrochloride |
| DSB | DNA double strand breaks |
| γ-H2AX | phosphorylated histone H2AX |
| gDNA | genomic DNA |
| GILA | Growth-in-low-attachment |
| HE | hematoxylin/eosin |
| MFISH | Multicolor Fluorescence in situ hybridization |
| NMuMG | Normal murine mammary gland |
| PBS | Phosphate buffered saline |
| PCC | premature chromosome condensation |
References
- Darbre, P.D. Underarm cosmetics are a cause of breast cancer. Eur. J. Cancer Prev. 2001, 10, 389–393. [Google Scholar] [CrossRef] [PubMed]
- Key, T.J.; Verkasalo, P.K.; Banks, E. Epidemiology of breast cancer. Lancet Oncol. 2001, 3, 133–140. [Google Scholar] [CrossRef]
- King, M.C.; Marks, J.H.; Mandell, J.B.; New York Breast Cancer Study Group. Breast and ovarian cancer risks due to inherited mutations in BRCA1 and BRCA2. Science 2003, 302, 643–646. [Google Scholar] [CrossRef] [PubMed]
- McGrath, K.G. An earlier age of breast cancer diagnosis related to more frequent use of antiperspirants/deodorants and underarm shaving. Eur. J. Cancer Prev. 2003, 12, 479–485. [Google Scholar] [CrossRef] [PubMed]
- Linhart, C.; Talasz, H.; Morandi, E.M.; Exley, C.; Lindner, H.H.; Taucher, S.; Egle, D.; Hubalek, M.; Concin, N.; Ulmer, H. Use of Underarm Cosmetic Products in Relation to Risk of Breast Cancer: A Case-Control Study. EBioMedicine 2017, 21, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Darbre, P.D. Aluminium and the human breast. Morphologie 2016, 100, 65–74. [Google Scholar] [CrossRef]
- Sappino, A.P.; Buser, R.; Lesne, L.; Gimelli, S.; Béna, F.; Belin, D.; Mandriota, S.J. Aluminium chloride promotes anchorage-independent growth in human mammary epithelial cells. J. Appl. Toxicol. 2012, 32, 233–243. [Google Scholar] [CrossRef]
- Mandriota, S.J.; Tenan, M.; Ferrari, P.; Sappino, A.P. Aluminium chloride promotes tumorigenesis and metastasis in normal murine mammary gland epithelial cells. Int. J. Cancer 2016, 139, 2781–2790. [Google Scholar] [CrossRef]
- Darbre, P.D.; Bakir, A.; Iskakova, E. Effect of aluminium on migratory and invasive properties of MCF-7 human breast cancer cells in culture. J. Inorg. Biochem. 2013, 128, 245–249. [Google Scholar] [CrossRef]
- Bakir, A.; Darbre, P.D. Effect of aluminium on migration of oestrogen unresponsive MDA-MB-231 human breast cancer cells in culture. J. Inorg. Biochem. 2015, 152, 180–185. [Google Scholar] [CrossRef]
- Pereira, S.; Cavalie, I.; Camilleri, V.; Gilbin, R.; Adam-Guillermin, C. Comparative genotoxicity of aluminium and cadmium in embryonic zebrafish cells. Mutat. Res. 2013, 750, 19–26. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.; Sjogren, C.A.; Larsen, P.B.; Schnittger, A. A multi-level response to DNA damage induced by aluminium. Plant J. 2019, 98, 479–491. [Google Scholar] [CrossRef] [PubMed]
- Ball, R.K.; Friis, R.R.; Schoenenberger, C.A.; Doppler, W.; Groner, B. Prolactin regulation of beta-casein gene expression and of a cytosolic 120-kd protein in a cloned mouse mammary epithelial cell line. EMBO J. 1988, 7, 2089–2095. [Google Scholar] [CrossRef] [PubMed]
- Rotem, A.; Janzer, A.; Izar, B.; Ji, Z.; Doench, J.G.; Garraway, L.A.; Struhl, K. Alternative to the soft-agar assay that permits high-throughput drug and genetic screens for cellular transformation. Proc. Natl. Acad. Sci. USA 2015, 112, 5708–5713. [Google Scholar] [CrossRef] [PubMed]
- Saleem, M.; Qadir, M.I.; Perveen, N.; Ahmad, B.; Saleem, U.; Irshad, T.; Ahmad, B. Inhibitors of apoptotic proteins: New targets for anticancer therapy. Chem. Biol. Drug Des. 2013, 82, 243–251. [Google Scholar] [CrossRef] [PubMed]
- Stenman, G. Fusion oncogenes in salivary gland tumors: Molecular and clinical consequences. Head Neck Pathol. 2013, 7 (Suppl. 1), S12–S19. [Google Scholar] [CrossRef]
- Zanconato, F.; Cordenonsi, M.; Piccolo, S. YAP/TAZ at the Roots of Cancer. Cancer Cell 2016, 29, 783–803. [Google Scholar] [CrossRef]
- Merlo, G.R.; Venesio, T.; Taverna, D.; Callahan, R.; Hynes, N.E. Growth suppression of normal mammary epithelial cells by wild-type p53. Ann. N. Y. Acad. Sci. 1993, 698, 108–113. [Google Scholar] [CrossRef]
- Termén, S.; Tan, E.J.; Heldin, C.H.; Moustakas, A. p53 regulates epithelial-mesenchymal transition induced by transforming growth factor β. J. Cell Physiol. 2013, 228, 801–813. [Google Scholar] [CrossRef]
- Nicholson, S.; Exley, C. Aluminum: A potential pro-oxidant in sunscreens/sunblocks? Free Radic. Biol. Med. 2007, 43, 1216–1217. [Google Scholar] [CrossRef]
- Chen, Q.Y.; DesMarais, T.; Costa, M. Metals and Mechanisms of Carcinogenesis. Annu. Rev. Pharmacol. Toxicol. 2019, 59, 537–554. [Google Scholar] [CrossRef] [PubMed]
- Klotz, K.; Weistenhöfer, W.; Neff, F.; Hartwig, A.; van Thriel, C.; Drexler, H. The Health Effects of Aluminum Exposure. Dtsch. Arztebl. Int. 2017, 114, 653–659. [Google Scholar] [CrossRef] [PubMed]
- Brandt, R.; Wong, A.M.; Hynes, N.E. Mammary glands reconstituted with Neu/ErbB2 transformed HC11 cells provide a novel orthotopic tumor model for testing anti-cancer agents. Oncogene 2001, 20, 5459–5465. [Google Scholar] [CrossRef] [PubMed]
- Tao, Y.; Han, T.; Zhang, T.; Sun, C. Sulfatase-2 promotes the growth and metastasis of colorectal cancer by activating Akt and Erk1/2 pathways. Biomed. Pharmacother. 2017, 89, 1370–1377. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Nakamura, I.; Dhanasekaran, R.; Iguchi, E.; Tolosa, E.J.; Romecin, P.A.; Vera, R.E.; Almada, L.L.; Miamen, A.G.; Chaiteerakij, R.; et al. Transcriptional Induction of Periostin by a Sulfatase 2-TGFβ1-SMAD Signaling Axis Mediates Tumor Angiogenesis in Hepatocellular Carcinoma. Cancer Res. 2017, 77, 632–645. [Google Scholar] [CrossRef] [PubMed]
- Jung, C.H.; Ho, J.N.; Park, J.K.; Kim, E.M.; Hwang, S.G.; Um, H.D. Involvement of SULF2 in y-irradiation-induced invasion and resistance of cancer cells by inducing IL-6 expression. Oncotarget 2016, 7, 16090–16103. [Google Scholar] [CrossRef]
- Tavera-Mendoza, L.E.; Brown, M. A less invasive method for orthotopic injection of breast cancer cells into the mouse mammary gland. Lab. Anim. 2017, 51, 85–88. [Google Scholar] [CrossRef]
- Becker, J.R.; Cuella-Martin, R.; Barazas, M.; Liu, R.; Oliveira, C.; Oliver, A.W.; Bilham, K.; Holt, A.B.; Blackford, A.N.; Heierhorst, J.; et al. The ASCIZ-DYNLL1 axis promotes 53BP1-dependent non-homologous end joining and PARP inhibitor sensitivity. Nat. Commun. 2018, 9, 5406. [Google Scholar] [CrossRef]
- Silva Nunes, J.P.; Martins Dias, A.A. ImageJ macros for the user-friendly analysis of soft-agar and wound-healing assays. Biotechniques 2017, 32, 175–179. [Google Scholar] [CrossRef]

| Cell Line | a | b | c | d | e | f |
|---|---|---|---|---|---|---|
| HC11 | 0 | - (parental) | 77 | 17 | 3 | 18 |
| 71 | H2O | 69 | 27 | 6 | 24 | |
| 71 | AlCl3 10 µM | 74 | 34 | 14 | 25 | |
| 71 | AlCl3 100 µM | NA | 41 | 18 | 24 | |
| NMuMG | 0 | - (parental) | 39 | 1 | 1 | 24 |
| 38 | H2O S. I | 67 | 8 | 5 | 23 | |
| 38 | AlCl3 100 µM S. I | 72 | 14 | 9 | 23 | |
| 40 | H2O S. III | 64 | 23 | 14 | 25 | |
| 40 | AlCl3 100 µM S. III | 65 | 37 | 26 | 23 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mandriota, S.J.; Tenan, M.; Nicolle, A.; Jankowska, J.D.; Ferrari, P.; Tille, J.-C.; Durin, M.-A.; Green, C.M.; Tabruyn, S.; Moralli, D.; et al. Genomic Instability Is an Early Event in Aluminium-Induced Tumorigenesis. Int. J. Mol. Sci. 2020, 21, 9332. https://doi.org/10.3390/ijms21239332
Mandriota SJ, Tenan M, Nicolle A, Jankowska JD, Ferrari P, Tille J-C, Durin M-A, Green CM, Tabruyn S, Moralli D, et al. Genomic Instability Is an Early Event in Aluminium-Induced Tumorigenesis. International Journal of Molecular Sciences. 2020; 21(23):9332. https://doi.org/10.3390/ijms21239332
Chicago/Turabian StyleMandriota, Stefano J., Mirna Tenan, Adeline Nicolle, Julia D. Jankowska, Paolo Ferrari, Jean-Christophe Tille, Mary-Anne Durin, Catherine M. Green, Sebastien Tabruyn, Daniela Moralli, and et al. 2020. "Genomic Instability Is an Early Event in Aluminium-Induced Tumorigenesis" International Journal of Molecular Sciences 21, no. 23: 9332. https://doi.org/10.3390/ijms21239332
APA StyleMandriota, S. J., Tenan, M., Nicolle, A., Jankowska, J. D., Ferrari, P., Tille, J.-C., Durin, M.-A., Green, C. M., Tabruyn, S., Moralli, D., & Sappino, A.-P. (2020). Genomic Instability Is an Early Event in Aluminium-Induced Tumorigenesis. International Journal of Molecular Sciences, 21(23), 9332. https://doi.org/10.3390/ijms21239332





