Comparative Transcriptome Analysis of Rutabaga (Brassica napus) Cultivars Indicates Activation of Salicylic Acid and Ethylene-Mediated Defenses in Response to Plasmodiophora brassicae
Abstract
:1. Introduction
2. Results and Discussion
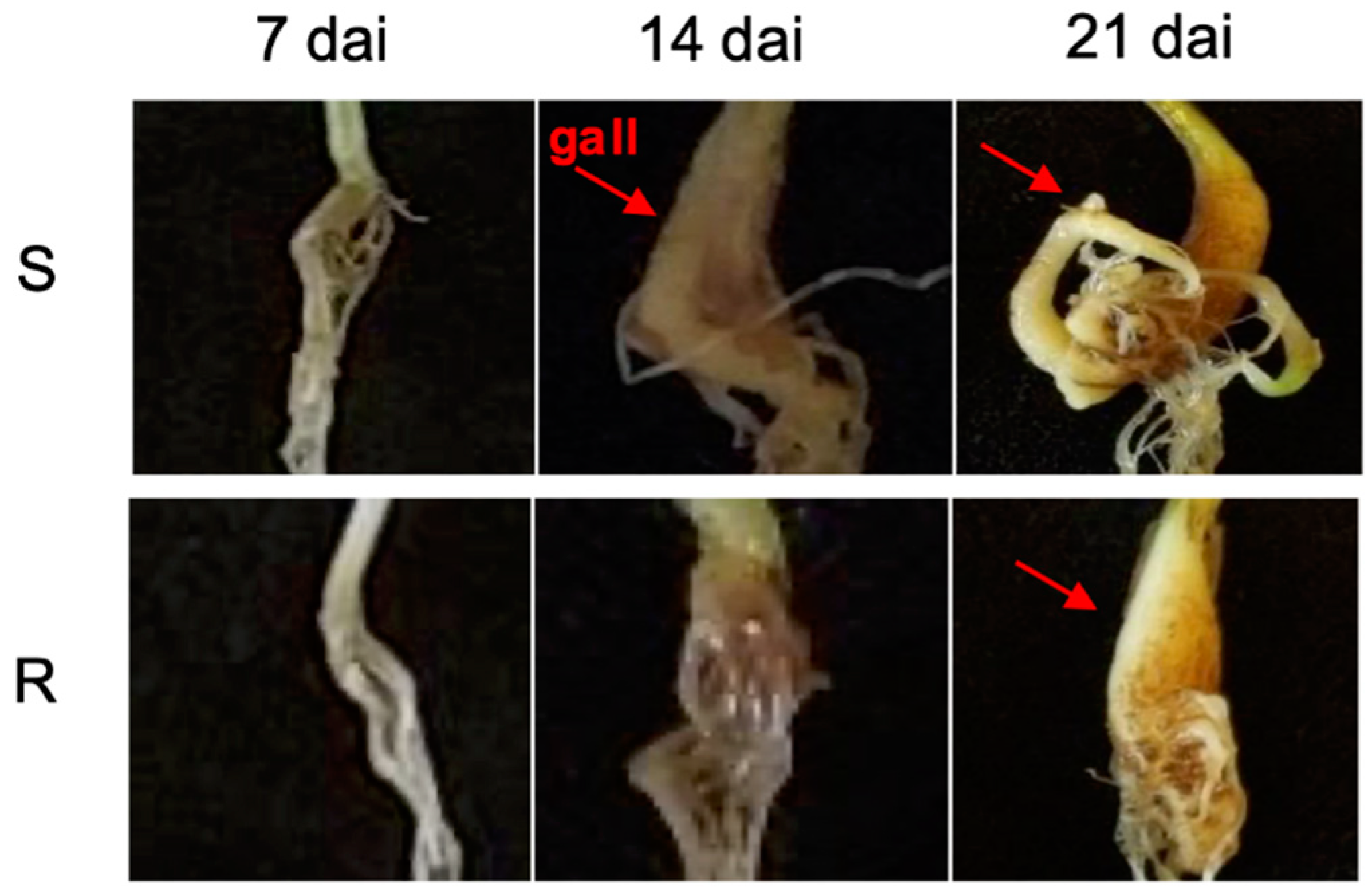
2.1. Disease Assessment
2.2. RNA-Seq Analysis
2.3. Validation of RNA-Seq Data by Quantitative Real-Time PCR (qRT-PCR)
2.4. Genes Related to Biotic Stress Pathways
2.4.1. Overview of Biotic Stress-Related Pathways
2.4.2. Genes Related to SA, ET, and JA Metabolism
2.4.3. Pathogenesis-Related (PR) Genes
2.4.4. Signaling
2.4.5. Transcription Factors
2.4.6. Protein Degradation
2.5. Analysis of Genes with Opposite Regulation in the Resistant vs. Susceptible Hosts
2.6. A Model of the Molecular Response in the Resistant Cultivar ‘Wilhemsburger’ to P. brassicae
3. Materials and Methods
3.1. Pathogen Material
3.2. Plant Material and Inoculation
3.3. RNA Extraction
3.4. RNA-Seq Analysis
3.5. Validation of RNA-Seq Data by qRT-PCR
3.6. Bioinformatic Analyses
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ABA | Abscisic acid |
| ACS2 | 1-amino-cyclopropane-1-carboxylate synthase 2 |
| AOC2 | Allene oxide cyclase 2 |
| AOS | Allene oxide synthase |
| ATL | Arabidopsis Tóxicos en Levadura |
| BAT5 | Bile acid transporter 5 |
| BTL10 | BCA2Â zinc finger ATL 10 |
| bZIP | Basic leucine zipper |
| CAC | Clathrin adaptor complex |
| CaMBP | Calmodulin-binding protein |
| CBP | Calcium binding protein |
| CBP60g | CaMBP 60-like G |
| CCD | Canadian Clubroot Differential |
| CR | Clubroot resistant |
| CS | Clubroot susceptible |
| Ct | Cycle threshold |
| CYP79F1 | Cytochrome p450 79f1 |
| CYP83A1 | Cytochrome P450, family 83, subfamily A, polypeptide 1 |
| CYP94B1 | Cytochrome P450, family 94, subfamily B, polypeptide 1 |
| CYP94C1 | Cytochrome P450, family 94, subfamily C, polypeptide 1 |
| dai | Days after inoculation |
| DAMP | Damage-associated molecular pattern |
| DEG | Differentially expressed gene |
| dH2O | Distilled water |
| DI | Disease index |
| DIR6 | Dirigent protein 6 |
| E2 | E2 ubiquitin-conjugating enzyme |
| E3 | E3 ubiquitin ligase |
| EREBP | APETALA2/ethylene-responsive element binding protein |
| ERF | Ethylene response factor |
| ET | Ethylene |
| ETI | Effector-triggered immunity |
| FAD7 | Fatty acid desaturase 7 |
| GDI1 | Dissociation inhibitor 1 |
| ICS | Isochorismate synthase |
| JA | Jasmonic acid |
| JA-Ile | Jasmonoyl-L-isoleucine |
| log2FC | Log2 fold-change |
| MAPK | Mitogen-activated protein kinase |
| MEKK | MAPK kinase kinase |
| MKK | MAPK kinase |
| MPK6 | Mitogen-activated protein kinase 6 |
| MYB15 | MYB domain protein 15 |
| NLP | Necrosis and ethylene-inducing peptide 1-like protein |
| NPR1 | Nonexpresser of PR genes 1 |
| NPR3 | NPR1-like protein 3 |
| OPR1 | 12-oxophytodienoate reductase 1 |
| PAMP | Pathogen-associated molecular pattern |
| PCA | Principal component analysis |
| PEPR2 | Pep 1 receptor 2 |
| PLSP2A | Plastidic type I signal peptidase 2A |
| PR | Pathogenesis-related |
| PR1 | Pathogenesis-related gene 1 |
| PRR | Pattern recognition receptor |
| PTI | PAMP-triggered immunity |
| qRT-PCR | Quantitative Real-Time PCR |
| QTL | Quantitative resistance loci |
| R gene | Resistance gene |
| RBOH | Respiratory burst oxidase homolog |
| RIN | RNA integrity number |
| RLP | Receptor-like protein |
| RNA-seq | RNA sequencing |
| ROS | Reactive oxygen species |
| RPKM | Reads per kb of transcript per million mapped reads |
| SA | Salicylic acid |
| SARD1 | SAR deficient 1 |
| SCFE1 | Sclerotinia culture filtrate elicitor 1 |
| SOBIR1 | Protein suppressor of BIR1–1 |
| SRA | Sequence Read Archive |
| TF | Transcription factor |
| TGA | TGACG motif-binding protein |
| TIR-NBS-LRR | Toll-interleukin receptor nucleotide-binding site-leucine-rich repeat |
| TUA5 | Tubulin alpha-5 |
| UBC | Ubiquitin conjugating enzyme |
| VHA-E1 | Vacuolar ATP synthase subunit E1 |
| WAVH1 | WAV3 homolog 1 |
References
- Dixon, G.R. The occurrence and economic impact of Plasmodiophora brassicae and clubroot disease. J. Plant Growth Regul. 2009, 28, 194–202. [Google Scholar] [CrossRef]
- Howard, R.J.; Strelkov, S.E.; Harding, M.W. Clubroot of cruciferous crops - new perspectives on an old disease. Can. J. Plant Pathol. 2010, 32, 43–57. [Google Scholar] [CrossRef]
- Strelkov, S.E.; Hwang, S.-F. Clubroot in the Canadian canola crop: 10 years into the outbreak. Can. J. Plant Pathol. 2014, 36, 27–36. [Google Scholar] [CrossRef]
- Canola Council of Canada—Industry Overview. Available online: https://www.canolacouncil.org/markets-stats/industry-overview/ (accessed on 12 September 2020).
- Strelkov, S.E.; Hwang, S.-F.; Howard, R.J.; Hartman, M.; Turkington, T.K. Progress towards the sustainable management of clubroot (Plasmodiophora brassicae) of canola on the Canadian prairies. Prairie Soils Crop. J. 2011, 4, 114–121. [Google Scholar]
- Hwang, S.-F.; Howard, R.J.; Strelkov, S.E.; Gossen, B.D.; Peng, G. Management of clubroot (Plasmodiophora brassicae) on canola (Brassica napus) in western Canada. Can. J. Plant Pathol. 2014, 36, 49–65. [Google Scholar] [CrossRef]
- Peng, G.; Lahlali, R.; Hwang, S.-F.; Pageau, D.; Hynes, R.K.; McDonald, M.R.; Gossen, B.D.; Strelkov, S.E. Crop rotation, cultivar resistance, and fungicides/biofungicides for managing clubroot (Plasmodiophora brassicae) on canola. Can. J. Plant Pathol. 2014, 36, 99–112. [Google Scholar] [CrossRef]
- Canola Council of Canada—Control Clubroot. Available online: https://www.canolacouncil.org/canola-encyclopedia/diseases/clubroot/control-clubroot/#pathotypes (accessed on 4 July 2019).
- Fredua-Agyeman, R.; Hwang, S.-F.; Strelkov, S.E.; Zhou, Q.; Feindel, D. Potential loss of clubroot resistance genes from donor parent Brassica rapa subsp. rapifera (ECD 04) during doubled haploid production. Plant Pathol. 2018, 67, 892–901. [Google Scholar] [CrossRef]
- Strelkov, S.E.; Hwang, S.-F.; Manolii, V.P.; Cao, T.; Feindel, D. Emergence of new virulence phenotypes of Plasmodiophora brassicae on canola (Brassica napus) in Alberta, Canada. Eur. J. Plant Pathol. 2016, 145, 517–529. [Google Scholar] [CrossRef]
- Strelkov, S.E.; Hwang, S.-F.; Manolii, V.P.; Cao, T.; Fredua-Agyeman, R.; Harding, M.W.; Peng, G.; Gossen, B.D.; Mcdonald, M.R.; Feindel, D. Virulence and pathotype classification of Plasmodiophora brassicae populations collected from clubroot resistant canola (Brassica napus) in Canada. Can. J. Plant Pathol. 2018, 40, 284–298. [Google Scholar] [CrossRef]
- Strelkov, S.E.; Hwang, S.-F.; Manolii, V.P.; Turnbull, G.; Fredua-Agyeman, R.; Hollman, K.; Kaus, S. Characterization of clubroot (Plasmodiophora brassicae) from canola (Brassica napus) in the Peace Country of Alberta, Canada. Can. J. Plant Pathol. 2020, 1–7. [Google Scholar] [CrossRef]
- Dodds, P.N.; Rathjen, J.P. Plant immunity: Towards an integrated view of plant-pathogen interactions. Nat. Rev. Genet. 2010, 11, 539–548. [Google Scholar] [CrossRef] [PubMed]
- Zipfel, C. Plant pattern-recognition receptors. Trends Immunol. 2014, 35, 345–351. [Google Scholar] [CrossRef]
- Ueno, H.; Matsumoto, E.; Aruga, D.; Kitagawa, S.; Matsumura, H.; Hayashida, N. Molecular characterization of the CRa gene conferring clubroot resistance in Brassica rapa. Plant Mol. Biol. 2012, 80, 621–629. [Google Scholar] [CrossRef] [Green Version]
- Hatakeyama, K.; Suwabe, K.; Tomita, R.N.; Kato, T.; Nunome, T.; Fukuoka, H.; Matsumoto, S. Identification and characterization of Crr1a, a gene for resistance to clubroot disease (Plasmodiophora brassicae Woronin) in Brassica rapa L. PLoS ONE 2013, 8, e54745. [Google Scholar] [CrossRef] [PubMed]
- Neik, T.X.; Barbetti, M.J.; Batley, J. Current status and challenges in identifying disease resistance genes in Brassica napus. Front. Plant Sci. 2017, 8, 1788. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.; Zhang, X.; Peng, G.; Falk, K.C.; Strelkov, S.E.; Gossen, B.D. Genotyping-by-sequencing reveals three QTL for clubroot resistance to six pathotypes of Plasmodiophora brassicae in Brassica rapa. Sci. Rep. 2017, 7, 4516. [Google Scholar] [CrossRef]
- Yu, F.; Zhang, X.; Huang, Z.; Chu, M.; Song, T.; Falk, K.C.; Deora, A.; Chen, Q.; Zhang, Y.; McGregor, L.; et al. Identification of genome-wide variants and discovery of variants associated with Brassica rapa clubroot resistance gene Rcr1 through bulked segregant RNA sequencing. PLoS ONE 2016, 11, e0153218. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Peng, G.; Liu, X.; Deora, A.; Falk, K.C.; Gossen, B.D.; McDonald, M.R.; Yu, F. Fine mapping of a clubroot resistance gene in Chinese cabbage using SNP markers identified from bulked segregant RNA sequencing. Front. Plant Sci. 2017, 8, 1448. [Google Scholar] [CrossRef] [Green Version]
- Pang, W.; Fu, P.; Li, X.; Zhan, Z.; Yu, S.; Piao, Z. Identification and mapping of the clubroot resistance gene CRd in Chinese cabbage (Brassica rapa ssp. pekinensis). Front. Plant Sci. 2018, 9, 653. [Google Scholar] [CrossRef] [Green Version]
- Chang, A.; Lamara, M.; Wei, Y.; Hu, H.; Parkin, I.A.P.; Gossen, B.D.; Peng, G.; Yu, F. Clubroot resistance gene Rcr6 in Brassica nigra resides in a genomic region homologous to chromosome A08 in B. rapa. BMC Plant Biol. 2019, 19, 224. [Google Scholar] [CrossRef]
- Boyd, L.A.; Ridout, C.; O’Sullivan, D.M.; Leach, J.E.; Leung, H. Plant–pathogen interactions: Disease resistance in modern agriculture. Trends Genet. 2013, 29, 233–240. [Google Scholar] [CrossRef]
- Chen, J.; Pang, W.; Chen, B.; Zhang, C.; Piao, Z. Transcriptome analysis of Brassica rapa near-isogenic lines carrying clubroot-resistant and–susceptible alleles in response to Plasmodiophora brassicae during early infection. Front. Plant Sci. 2016, 6, 1183. [Google Scholar] [CrossRef] [Green Version]
- Luo, Y.; Dong, D.; Su, Y.; Wang, X.; Peng, Y.; Peng, J.; Zhou, C. Transcriptome analysis of Brassica juncea var. tumida Tsen responses to Plasmodiophora brassicae primed by the biocontrol strain Zhihengliuella aestuarii. Funct. Integr. Genom. 2018, 18, 301–314. [Google Scholar] [CrossRef]
- Summanwar, A.; Basu, U.; Rahman, H.; Kav, N. Identification of lncRNAs responsive to infection by Plasmodiophora brassicae in clubroot-susceptible and -resistant Brassica napus lines carrying resistance introgressed from rutabaga. Mol. Plant-Microbe Interact. 2019, 32, 1360–1377. [Google Scholar] [CrossRef]
- Rahman, H.; Peng, G.; Yu, F.; Falk, K.C.; Kulkarni, M.; Selvaraj, G. Genetics and breeding for clubroot resistance in Canadian spring canola (Brassica napus L.). Can. J. Plant Pathol. 2014, 36, 122–134. [Google Scholar] [CrossRef]
- Fredua-Agyeman, R.; Yu, Z.; Hwang, S.-F.; Strelkov, S.E. Genome-wide mapping of loci associated with resistance to clubroot in Brassica napus ssp. napobrassica (rutabaga) accessions from Nordic countries. Front. Plant Sci. 2020, 11, 742. [Google Scholar] [CrossRef]
- Ayers, C.W.; Lelacheur, K.E. Genetics of resistance in rutabaga to two races of Plasmodiophora brassicae. Can. J. Plant Sci. 1972, 52, 897–900. [Google Scholar] [CrossRef]
- Hasan, M.J.; Rahman, H. Genetics and molecular mapping of resistance to Plasmodiophora brassicae pathotypes 2, 3, 5, 6, and 8 in rutabaga (Brassica napus var. napobrassica). Genome 2016, 59, 805–815. [Google Scholar] [CrossRef] [Green Version]
- Galindo-González, L.; Manolii, V.; Hwang, S.-F.; Strelkov, S.E. Response of Brassica napus to Plasmodiophora brassicae involves salicylic acid-mediated immunity: An RNA-seq-based study. Front. Plant Sci. 2020, 11, 1025. [Google Scholar] [CrossRef]
- Song, T.; Chu, M.; Lahlali, R.; Yu, F.; Peng, G. Shotgun label-free proteomic analysis of clubroot (Plasmodiophora brassicae) resistance conferred by the gene Rcr1 in Brassica rapa. Front. Plant Sci. 2016, 7, 1013. [Google Scholar] [CrossRef] [Green Version]
- Mei, J.; Guo, Z.; Wang, J.; Feng, Y.; Ma, G.; Zhang, C.; Qian, W.; Chen, G. Understanding the resistance mechanism in Brassica napus to Clubroot Caused by Plasmodiophora brassicae. Phytopathology 2019, 109, 810–818. [Google Scholar] [CrossRef]
- Thimm, O.; Bläsing, O.; Gibon, Y.; Nagel, A.; Meyer, S.; Krüger, P.; Selbig, J.; Müller, L.A.; Rhee, S.Y.; Stitt, M. MAPMAN: A user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 2004, 37, 914–939. [Google Scholar] [CrossRef]
- Berens, M.L.; Berry, H.M.; Mine, A.; Argueso, C.T.; Tsuda, K. Evolution of hormone signaling networks in plant defense. Annu. Rev. Phytopathol. 2017, 55, 401–425. [Google Scholar] [CrossRef]
- Wang, J.H.; Gu, K.D.; Han, P.L.; Yu, J.Q.; Wang, C.K.; Zhang, Q.Y.; You, C.X.; Hu, D.G.; Hao, Y.J. Apple ethylene response factor MdERF11 confers resistance to fungal pathogen Botryosphaeria dothidea. Plant Sci. 2020, 291, 110351. [Google Scholar] [CrossRef]
- Fu, P.; Piao, Y.; Zhan, Z.; Zhao, Y.; Pang, W.; Li, X.; Piao, Z. Transcriptome profile of Brassica rapa L. reveals the involvement of jasmonic acid, ethylene, and brassinosteroid signaling pathways in clubroot resistance. Agronomy 2019, 9, 589. [Google Scholar] [CrossRef] [Green Version]
- Guerreiro, A.; Figueiredo, J.; Sousa Silva, M.; Figueiredo, A. Linking jasmonic acid to grapevine resistance against the biotrophic oomycete Plasmopara viticola. Front. Plant Sci. 2016, 7, 565. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jubault, M.; Lariagon, C.; Taconnat, L.; Renou, J.-P.P.; Gravot, A.; Delourme, R.; Manzanares-Dauleux, M.J. Partial resistance to clubroot in Arabidopsis is based on changes in the host primary metabolism and targeted cell division and expansion capacity. Funct. Integr. Genom. 2013, 13, 191–205. [Google Scholar] [CrossRef] [Green Version]
- Lemarié, S.; Robert-Seilaniantz, A.; Lariagon, C.; Lemoine, J.; Marnet, N.; Jubault, M.; Manzanares-Dauleux, M.J.; Gravot, A. Both the jasmonic acid and the salicylic acid pathways contribute to resistance to the biotrophic clubroot agent Plasmodiophora brassicae in Arabidopsis. Plant Cell Physiol. 2015, 56, 2158–2168. [Google Scholar] [CrossRef] [Green Version]
- Jia, H.; Wei, X.; Yang, Y.; Yuan, Y.; Wei, F.; Zhao, Y.; Yang, S.; Yao, Q.; Wang, Z.; Tian, B.; et al. Root RNA-seq analysis reveals a distinct transcriptome landscape between clubroot-susceptible and clubroot-resistant Chinese cabbage lines after Plasmodiophora brassicae infection. Plant Soil 2017, 421, 93–105. [Google Scholar] [CrossRef]
- Garcion, C.; Lohmann, A.; Lamodière, E.; Catinot, J.; Buchala, A.; Doermann, P.; Métraux, J.P. Characterization and biological function of the ISOCHORISMATE SYNTHASE2 gene of Arabidopsis. Plant Physiol. 2008, 147, 1279–1287. [Google Scholar] [CrossRef] [Green Version]
- Qu, L.; Li, S.; Xing, S. Methylation of phytohormones by the SABATH methyltransferases. Chin. Sci. Bull. 2010, 55, 2211–2218. [Google Scholar] [CrossRef]
- Dempsey, D.A.; Vlot, A.C.; Wildermuth, M.C.; Klessig, D.F. Salicylic acid biosynthesis and metabolism. Arab. Book 2011, 9, e0156. [Google Scholar] [CrossRef] [Green Version]
- Djavaheri, M.; Ma, L.; Klessig, D.F.; Mithöfer, A.; Gropp, G.; Borhan, H. Mimicking the host regulation of salicylic acid: A virulence strategy by the clubroot pathogen Plasmodiophora brassicae. Mol. Plant-Microbe Interact. 2019, 32, 296–305. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chu, M.; Song, T.; Falk, K.C.; Zhang, X.; Liu, X.; Chang, A.; Lahlali, R.; McGregor, L.; Gossen, B.D.; Yu, F. Fine mapping of Rcr1 and analyses of its effect on transcriptome patterns during infection by Plasmodiophora brassicae. BMC Genom. 2014, 15, 1166. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Knaust, A.; Ludwig-Müller, J. The ethylene signaling pathway is needed to restrict root gall growth in Arabidopsis after infection with the obligate biotrophic protist Plasmodiophora brassicae. J. Plant Growth Regul. 2013, 32, 9–21. [Google Scholar] [CrossRef]
- Bethke, G.; Unthan, T.; Uhrig, J.F.; Pöschl, Y.; Gust, A.A.; Scheel, D.; Lee, J. Flg22 regulates the release of an ethylene response factor substrate from MAP kinase 6 in Arabidopsis thaliana via ethylene signaling. Proc. Natl. Acad. Sci. USA 2009, 106, 8067–8072. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Su, T.; Yu, S.; Wang, W.; Li, P.; Zhang, F.; Yu, Y.; Zhang, D.; Zhao, X. ITRAQ analysis of protein profile during the secondary stage of infection of Plasmodiophora brassicae in Chinese cabbage (Brassica rapa subsp. pekinensis). J. Plant Pathol. 2018, 100, 533–542. [Google Scholar] [CrossRef]
- Zhang, W.; Fraiture, M.; Kolb, D.; Löffelhardt, B.; Desaki, Y.; Boutrot, F.F.G.; Tör, M.; Zipfel, C.; Gust, A.A.; Brunner, F. Arabidopsis RECEPTOR-LIKE PROTEIN30 and receptor-like kinase SUPPRESSOR OF BIR1-1/EVERSHED mediate innate immunity to necrotrophic fungi. Plant Cell 2013, 25, 4227–4241. [Google Scholar] [CrossRef] [Green Version]
- Albert, I.; Böhm, H.; Albert, M.; Feiler, C.E.; Imkampe, J.; Wallmeroth, N.; Brancato, C.; Raaymakers, T.M.; Oome, S.; Zhang, H.; et al. An RLP23-SOBIR1-BAK1 complex mediates NLP-triggered immunity. Nat. Plants 2015, 1, 1–9. [Google Scholar] [CrossRef]
- Vlot, A.C.; Dempsey, D.A.; Klessig, D.F. Salicylic acid, a multifaceted hormone to combat disease. Annu. Rev. Phytopathol. 2009, 47, 177–206. [Google Scholar] [CrossRef] [Green Version]
- Ding, Y.; Sun, T.; Ao, K.; Peng, Y.; Zhang, Y.Y.; Li, X.; Zhang, Y.Y. Opposite roles of salicylic acid receptors NPR1 and NPR3/NPR4 in transcriptional regulation of plant immunity. Cell 2018, 173, 1454–1467.e15. [Google Scholar] [CrossRef]
- Paniagua, C.; Bilkova, A.; Jackson, P.; Dabravolski, S.; Riber, W.; Didi, V.; Houser, J.; Gigli-Bisceglia, N.; Wimmerova, M.; Budínská, E.; et al. Dirigent proteins in plants: Modulating cell wall metabolism during abiotic and biotic stress exposure. J. Exp. Bot. 2017, 68, 3287–3301. [Google Scholar] [CrossRef] [Green Version]
- Lahlali, R.; Song, T.; Chu, M.; Yu, F.; Kumar, S.; Karunakaran, C.; Peng, G. Evaluating changes in cell-wall components associated with clubroot resistance using fourier transform infrared spectroscopy and RT-PCR. Int. J. Mol. Sci. 2017, 18, 2058. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ciaghi, S.; Schwelm, A.; Neuhauser, S. Transcriptomic response in symptomless roots of clubroot infected kohlrabi (Brassica oleracea var. gongylodes) mirrors resistant plants. BMC Plant Biol. 2019, 19, 288. [Google Scholar] [CrossRef] [Green Version]
- Dakouri, A.; Zhang, X.; Peng, G.; Falk, K.C.; Gossen, B.D.; Strelkov, S.E.; Yu, F. Analysis of genome-wide variants through bulked segregant RNA sequencing reveals a major gene for resistance to Plasmodiophora brassicae in Brassica oleracea. Sci. Rep. 2018, 8, 17657. [Google Scholar] [CrossRef]
- Liu, Z.; Wu, Y.; Yang, F.; Zhang, Y.; Chen, S.; Xie, Q.; Tian, X.; Zhou, J.M. BIK1 interacts with PEPRs to mediate ethylene-induced immunity. Proc. Natl. Acad. Sci. USA 2013, 110, 6205–6210. [Google Scholar] [CrossRef] [Green Version]
- Zipfel, C. Combined roles of ethylene and endogenous peptides in regulating plant immunity and growth. Proc. Natl. Acad. Sci. USA 2013, 110. [Google Scholar] [CrossRef] [Green Version]
- Liebrand, T.W.H.; Van Den Berg, G.C.M.; Zhang, Z.; Smit, P.; Cordewener, J.H.G.; America, A.H.P.; Sklenar, J.; Jones, A.M.E.; Tameling, W.I.L.; Robatzek, S.; et al. Receptor-like kinase SOBIR1/EVR interacts with receptor-like proteins in plant immunity against fungal infection. Proc. Natl. Acad. Sci. USA 2013, 110, 10010–10015. [Google Scholar] [CrossRef] [Green Version]
- Stael, S.; Kmiecik, P.; Willems, P.; Van Der Kelen, K.; Coll, N.S.; Teige, M.; Van Breusegem, F. Plant innate immunity-sunny side up? Trends Plant Sci. 2015, 20, 3–11. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.; Liu, Y.; Fang, Z.; Li, Z.; Yang, L.; Zhuang, M.; Zhang, Y.; Lv, H. Comparative transcriptome analysis between broccoli (Brassica oleracea var. italica) and wild cabbage (Brassica macrocarpa Guss.) in response to Plasmodiophora brassicae during different infection stages. Front. Plant Sci. 2016, 7, 1929. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ning, Y.; Wang, Y.; Fang, Z.; Zhuang, M.; Zhang, Y.; Lv, H.; Liu, Y.; Li, Z.; Yang, L. Comparative transcriptome analysis of cabbage (Brassica oleracea var. capitata) infected by Plasmodiophora brassicae reveals drastic defense response at secondary infection stage. Plant Soil 2019, 443, 167–183. [Google Scholar] [CrossRef]
- Birkenbihl, R.P.; Liu, S.; Somssich, I.E. Transcriptional events defining plant immune responses. Curr. Opin. Plant Biol. 2017, 38, 1–9. [Google Scholar] [CrossRef]
- Hu, Y.; Dong, Q.; Yu, D. Arabidopsis WRKY46 coordinates with WRKY70 and WRKY53 in basal resistance against pathogen Pseudomonas syringae. Plant Sci. 2012, 185, 288–297. [Google Scholar] [CrossRef]
- Asai, T.; Tena, G.; Plotnikova, J.; Willmann, M.R.; Chiu, W.L.; Gomez-Gomez, L.; Boller, T.; Ausubel, F.M.; Sheen, J.; Gomez-Gomez, L.; et al. MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 2002, 415, 977–983. [Google Scholar] [CrossRef]
- Zheng, Z.; Qamar, S.A.; Chen, Z.; Mengiste, T. Arabidopsis WRKY33 transcription factor is required for resistance to necrotrophic fungal pathogens. Plant J. 2006, 48, 592–605. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Meng, X.; Wang, R.; Mao, G.; Han, L.; Liu, Y.; Zhang, S. Dual-level regulation of ACC synthase activity by MPK3/MPK6 cascade and its downstream WRKY transcription factor during ethylene induction in Arabidopsis. PLoS Genet. 2012, 8, e1002767. [Google Scholar] [CrossRef]
- Sheikh, A.H.; Eschen-Lippold, L.; Pecher, P.; Hoehenwarter, W.; Sinha, A.K.; Scheel, D.; Lee, J. Regulation of WRKY46 transcription factor function by mitogen-activated protein kinases in Arabidopsis thaliana. Front. Plant Sci. 2016, 7, 61. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alves, M.; Dadalto, S.; Gonçalves, A.; De Souza, G.; Barros, V.; Fietto, L. Plant bZIP transcription factors responsive to pathogens: A review. Int. J. Mol. Sci. 2013, 14, 7815–7828. [Google Scholar] [CrossRef] [Green Version]
- Mandal, A.; Sharma, N.; Muthamilarasan, M.; Prasad, M. Ubiquitination: A tool for plant adaptation to changing environments. Nucleus 2018, 61, 253–260. [Google Scholar] [CrossRef]
- Zhao, Y.; Bi, K.; Gao, Z.; Chen, T.; Liu, H.; Xie, J.; Cheng, J.; Fu, Y.; Jiang, D. Transcriptome analysis of Arabidopsis thaliana in response to Plasmodiophora brassicae during early infection. Front. Microbiol. 2017, 8, 673. [Google Scholar] [CrossRef]
- Guzmán, P. The prolific ATL family of RING-H2 ubiquitin ligases. Plant Signal. Behav. 2012, 7, 1014–1021. [Google Scholar] [CrossRef] [Green Version]
- Serrano, M.; Guzmán, P. Isolation and gene expression analysis of Arabidopsis thaliana mutants with constitutive expression of ATL2, an early elicitor-response RING-H2 zinc-finger gene. Genetics 2004, 167, 919–929. [Google Scholar] [CrossRef] [Green Version]
- Maekawa, S.; Sato, T.; Asada, Y.; Yasuda, S.; Yoshida, M.; Chiba, Y.; Yamaguchi, J. The Arabidopsis ubiquitin ligases ATL31 and ATL6 control the defense response as well as the carbon/nitrogen response. Plant Mol. Biol. 2012, 79, 217–227. [Google Scholar] [CrossRef]
- Irani, S.; Trost, B.; Waldner, M.; Nayidu, N.; Tu, J.; Kusalik, A.J.; Todd, C.D.; Wei, Y.; Bonham-Smith, P.C. Transcriptome analysis of response to Plasmodiophora brassicae infection in the Arabidopsis shoot and root. BMC Genom. 2018, 19, 23. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bi, K.; He, Z.; Gao, Z.; Zhao, Y.; Fu, Y.; Cheng, J.; Xie, J.; Jiang, D.; Chen, T. Integrated omics study of lipid droplets from Plasmodiophora brassicae. Sci. Rep. 2016, 6, 36965. [Google Scholar] [CrossRef] [Green Version]
- Wasternack, C.; Stenzel, I.; Hause, B.; Hause, G.; Kutter, C.; Maucher, H.; Neumerkel, J.; Feussner, I.; Miersch, O. The wound response in tomato-role of jasmonic acid. J. Plant Physiol. 2006, 163, 297–306. [Google Scholar] [CrossRef]
- Weber, H. Fatty acid-derived signals in plants. Trends Plant Sci. 2002, 7, 217–224. [Google Scholar] [CrossRef]
- Avila, C.A.; Arévalo-Soliz, L.M.; Jia, L.; Navarre, D.A.; Chen, Z.; Howe, G.A.; Meng, Q.W.; Smith, J.E.; Goggin, F.L. Loss of function of FATTY ACID DESATURASE7 in tomato enhances basal aphid resistance in a salicylate-dependent manner. Plant Physiol. 2012, 158, 2028–2041. [Google Scholar] [CrossRef] [Green Version]
- Gutterson, N.; Reuber, T.L. Regulation of disease resistance pathways by AP2/ERF transcription factors. Curr. Opin. Plant Biol. 2004, 7, 465–471. [Google Scholar] [CrossRef]
- Li, L.; Long, Y.; Li, H.; Wu, X. Comparative transcriptome analysis reveals key pathways and hub genes in rapeseed during the early stage of Plasmodiophora brassicae infection. Front. Genet. 2020, 10, 1275. [Google Scholar] [CrossRef] [Green Version]
- Chezem, W.R.; Memon, A.; Li, F.S.; Weng, J.K.; Clay, N.K. SG2-type R2R3-MYB transcription factor MYB15 controls defense-induced lignification and basal immunity in Arabidopsis. Plant Cell 2017, 29, 1907–1926. [Google Scholar] [CrossRef] [Green Version]
- Luo, Y.; Bai, R.; Li, J.; Yang, W.; Li, R.; Wang, Q.; Zhao, G.; Duan, D. The transcription factor MYB15 is essential for basal immunity (PTI) in Chinese wild grape. Planta 2019, 1, 1889–1902. [Google Scholar] [CrossRef]
- Hsu, S.-C.; Endow, J.K.; Ruppel, N.J.; Roston, R.L.; Baldwin, A.J.; Inoue, K. Functional diversification of thylakoidal processing peptidases in Arabidopsis thaliana. PLoS ONE 2011, 6, e27258. [Google Scholar] [CrossRef] [PubMed]
- Moon, J.Y.; Kim, S.T.; Choi, G.J.; Kwon, S.-Y.; Cho, H.S.; Kim, H.-S.; Moon, J.S.; Park, J.M. Comparative proteomic analysis of host responses to Plasmodiophora brassicae infection in susceptible and resistant Brassica oleracea. Plant Biotechnol. Rep. 2020, 14, 263–274. [Google Scholar] [CrossRef]
- Fu, F.; Liu, X.; Wang, R.; Zhai, C.; Peng, G.; Yu, F.; Fernando, W.G.D. Fine mapping of Brassica napus blackleg resistance gene Rlm1 through bulked segregant RNA sequencing. Sci. Rep. 2019, 9, 14600. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koo, A.J.K.; Howe, G.A. Catabolism and deactivation of the lipid-derived hormone jasmonoyl-isoleucine. Front. Plant Sci. 2012, 3, 19. [Google Scholar] [CrossRef] [Green Version]
- Hirano, T.; Kimura, S.; Sakamoto, T.; Okamoto, A.; Nakayama, T.; Matsuura, T.; Ikeda, Y.; Takeda, S.; Suzuki, Y.; Ohshima, I.; et al. Reprogramming of the developmental program of Rhus javanica during initial stage of gall induction by Schlechtendalia chinensis. Front. Plant Sci. 2020, 11, 471. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Glawischnig, E.; Jørgensen, K.; Naur, P.; Jørgensen, B.; Olsen, C.E.; Hansen, C.H.; Rasmussen, H.; Pickett, J.A.; Halkier, B.A. CYP79F1 and CYP79F2 have distinct functions in the biosynthesis of aliphatic glucosinolates in Arabidopsis. Plant J. 2003, 33, 923–937. [Google Scholar] [CrossRef] [Green Version]
- Ludwig-Müller, J.; Prinsen, E.; Rolfe, S.A.; Scholes, J.D. Metabolism and plant hormone action during clubroot disease. J. Plant Growth Regul. 2009, 28, 229–244. [Google Scholar] [CrossRef]
- Gigolashvili, T.; Yatusevich, R.; Rollwitz, I.; Humphry, M.; Gershenzon, J.; Flügge, U.I. The plastidic bile acid transporter 5 is required for the biosynthesis of methionine-derived glucosinolates in Arabidopsis thaliana. Plant Cell 2009, 21, 1813–1829. [Google Scholar] [CrossRef] [Green Version]
- Hemm, M.R.; Ruegger, M.O.; Chapple, C. The Arabidopsis ref2 mutant is defective in the gene encoding CYP83A1 and shows both phenylpropanoid and glucosinolate phenotypes. Plant Cell 2003, 15, 179–194. [Google Scholar] [CrossRef] [Green Version]
- Bishop, J.G.; Dean, A.M.; Mitchell-Olds, T. Rapid evolution in plant chitinases: Molecular targets of selection in plant-pathogen coevolution. Proc. Natl. Acad. Sci. USA 2000, 97, 5322–5327. [Google Scholar] [CrossRef] [Green Version]
- Schwelm, A.; Fogelqvist, J.; Knaust, A.; Jülke, S.; Lilja, T.; Bonilla-Rosso, G.; Karlsson, M.; Shevchenko, A.; Dhandapani, V.; Choi, S.R.; et al. The Plasmodiophora brassicae genome reveals insights in its life cycle and ancestry of chitin synthases. Sci. Rep. 2015, 5, 11153. [Google Scholar] [CrossRef] [Green Version]
- Chen, J.; Piao, Y.; Liu, Y.; Li, X.; Piao, Z. Genome-wide identification and expression analysis of chitinase gene family in Brassica rapa reveals its role in clubroot resistance. Plant Sci. 2018, 270, 257–267. [Google Scholar] [CrossRef]
- Peng, Y.; Van Wersch, R.; Zhang, Y. Convergent and divergent signaling in PAMP-triggered immunity and effector-triggered immunity. Mol. Plant-Microbe Interact. 2018, 31, 403–409. [Google Scholar] [CrossRef] [Green Version]
- Turjanski, A.G.; Vaqueánd, J.P.; Gutkind, J.S. MAP kinases and the control of nuclear events. Oncogene 2007, 26, 3240–3253. [Google Scholar] [CrossRef] [Green Version]
- Strelkov, S.E.; Tewari, J.P.; Smith-Degenhardt, E. Characterization of Plasmodiophora brassicae populations from Alberta, Canada. Can. J. Plant Pathol. 2006, 28, 467–474. [Google Scholar] [CrossRef]
- Kuginuki, Y.; Yoshikawa, H.; Hirai, M. Variation in virulence of Plasmodiophora brassicae in Japan tested with clubroot-resistant cultivars of Chinese cabbage (Brassica rapa L. ssp. pekinensis). Eur. J. Plant Pathol. 1999, 105, 327–332. [Google Scholar] [CrossRef]
- Horiuchi, S.; Hori, M. A simple greenhouse technique for obtaining high levels of clubroot incidence. Bull. Chugoku Natl. Agric. Exp. Stn. Ser. E 1980, 17, 33–35. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [Green Version]
- Ewels, P.; Magnusson, M.; Lundin, S.; Käller, M. MultiQC: Summarize analysis results for multiple tools and samples in a single report. Bioinformatics 2016, 32, 3047–3048. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chalhoub, B.; Denoeud, F.; Liu, S.; Parkin, I.A.P.; Tang, H.; Wang, X.; Chiquet, J.; Belcram, H.; Tong, C.; Samans, B.; et al. Early allopolyploid evolution in the post-neolithic Brassica napus oilseed genome. Science 2014, 345, 950–953. [Google Scholar] [CrossRef] [Green Version]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Yang, H.; Liu, J.; Huang, S.; Guo, T.; Deng, L.; Hua, W. Selection and evaluation of novel reference genes for quantitative reverse transcription PCR (qRT-PCR) based on genome and transcriptome data in Brassica napus L. Gene 2014, 538, 113–122. [Google Scholar] [CrossRef] [PubMed]
- Chandna, R.; Augustine, R.; Bisht, N.C. Evaluation of candidate reference genes for gene expression normalization in Brassica juncea using real time quantitative RT-PCR. PLoS ONE 2012, 7, e36918. [Google Scholar] [CrossRef]
- Han, P.P.; Qin, L.; Li, Y.S.; Liao, X.S.; Xu, Z.X.; Hh, X.J.; Xie, L.H.; Yu, C.B.; Wu, Y.F.; Xing, L. Identification of suitable reference genes in leaves and roots of rapeseed (Brassica napus L.) under different nutrient deficiencies. J. Integr. Agric. 2017, 16, 809–819. [Google Scholar] [CrossRef]
- Pfaffl, M.W.; Tichopad, A.; Prgomet, C.; Neuvians, T.P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper—Excel-based tool using pair-wise correlations. Biotechnol. Lett. 2004, 26, 509–515. [Google Scholar] [CrossRef]
- Lamesch, P.; Berardini, T.Z.; Li, D.; Swarbreck, D.; Wilks, C.; Sasidharan, R.; Muller, R.; Dreher, K.; Alexander, D.L.; Garcia-Hernandez, M.; et al. The Arabidopsis Information Resource (TAIR): Improved gene annotation and new tools. Nucleic Acids Res. 2012, 40, D1202–D1210. [Google Scholar] [CrossRef]
- Bardou, P.; Mariette, J.; Escudié, F.; Djemiel, C.; Klopp, C. Jvenn: An interactive Venn diagram viewer. BMC Bioinformatics 2014, 15, 293. [Google Scholar] [CrossRef] [Green Version]
- Howe, E.; Holton, K.; Nair, S.; Schlauch, D.; Sinha, R.; Quackenbush, J. MeV: MultiExperiment viewer. In Biomedical Informatics for Cancer Research; Springer: Boston, MA, USA, 2010; pp. 267–277. [Google Scholar]
- Pathan, M.; Keerthikumar, S.; Ang, C.S.; Gangoda, L.; Quek, C.Y.J.; Williamson, N.A.; Mouradov, D.; Sieber, O.M.; Simpson, R.J.; Salim, A.; et al. FunRich: An open access standalone functional enrichment and interaction network analysis tool. Proteomics 2015, 15, 2597–2601. [Google Scholar] [CrossRef] [PubMed]







Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhou, Q.; Galindo-González, L.; Manolii, V.; Hwang, S.-F.; Strelkov, S.E. Comparative Transcriptome Analysis of Rutabaga (Brassica napus) Cultivars Indicates Activation of Salicylic Acid and Ethylene-Mediated Defenses in Response to Plasmodiophora brassicae. Int. J. Mol. Sci. 2020, 21, 8381. https://doi.org/10.3390/ijms21218381
Zhou Q, Galindo-González L, Manolii V, Hwang S-F, Strelkov SE. Comparative Transcriptome Analysis of Rutabaga (Brassica napus) Cultivars Indicates Activation of Salicylic Acid and Ethylene-Mediated Defenses in Response to Plasmodiophora brassicae. International Journal of Molecular Sciences. 2020; 21(21):8381. https://doi.org/10.3390/ijms21218381
Chicago/Turabian StyleZhou, Qinqin, Leonardo Galindo-González, Victor Manolii, Sheau-Fang Hwang, and Stephen E. Strelkov. 2020. "Comparative Transcriptome Analysis of Rutabaga (Brassica napus) Cultivars Indicates Activation of Salicylic Acid and Ethylene-Mediated Defenses in Response to Plasmodiophora brassicae" International Journal of Molecular Sciences 21, no. 21: 8381. https://doi.org/10.3390/ijms21218381
APA StyleZhou, Q., Galindo-González, L., Manolii, V., Hwang, S.-F., & Strelkov, S. E. (2020). Comparative Transcriptome Analysis of Rutabaga (Brassica napus) Cultivars Indicates Activation of Salicylic Acid and Ethylene-Mediated Defenses in Response to Plasmodiophora brassicae. International Journal of Molecular Sciences, 21(21), 8381. https://doi.org/10.3390/ijms21218381




