Transcriptional and Morpho-Physiological Responses of Marchantia polymorpha upon Phosphate Starvation
Abstract
1. Introduction
2. Results
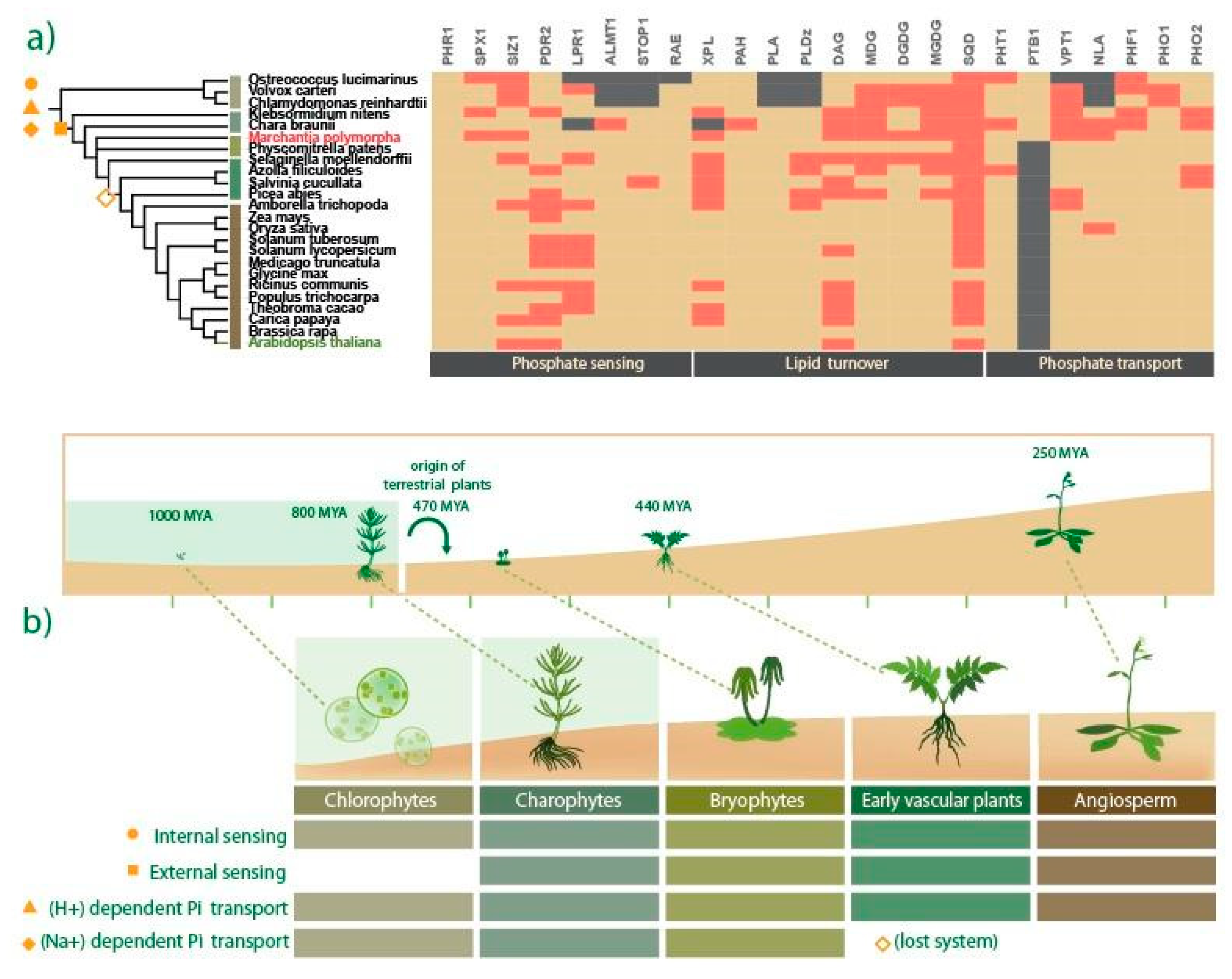
2.1. Evolutionary Landscape of Pi Starvation-Induced Genes across the Plant Kingdom
2.2. M. polymorpha Exhibits Low Genetic Redundancy in Key Pi-Responsive Regulatory Networks
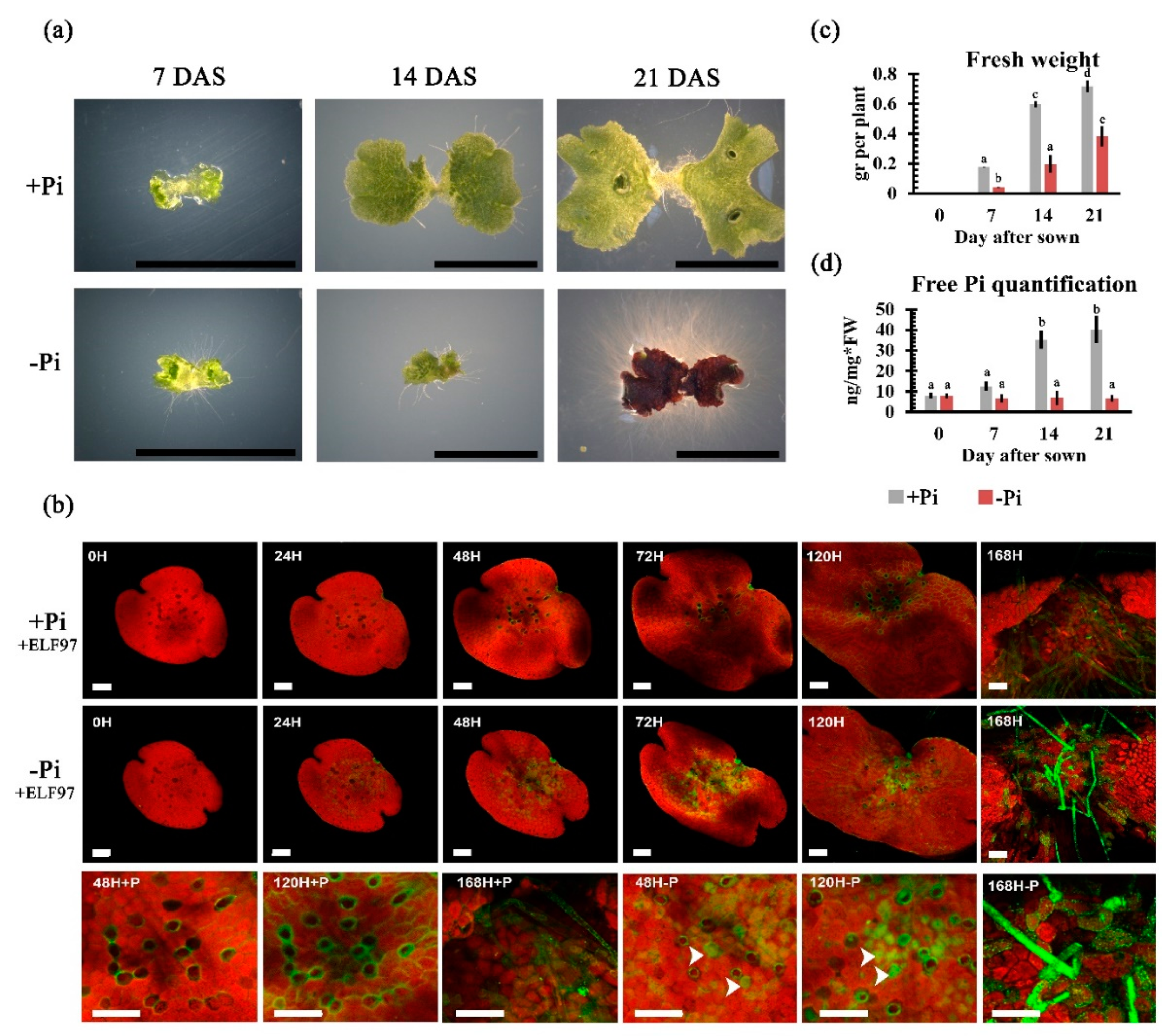
2.3. Phenotypical and Physiological Impacts of Pi Availability on Thallus Development
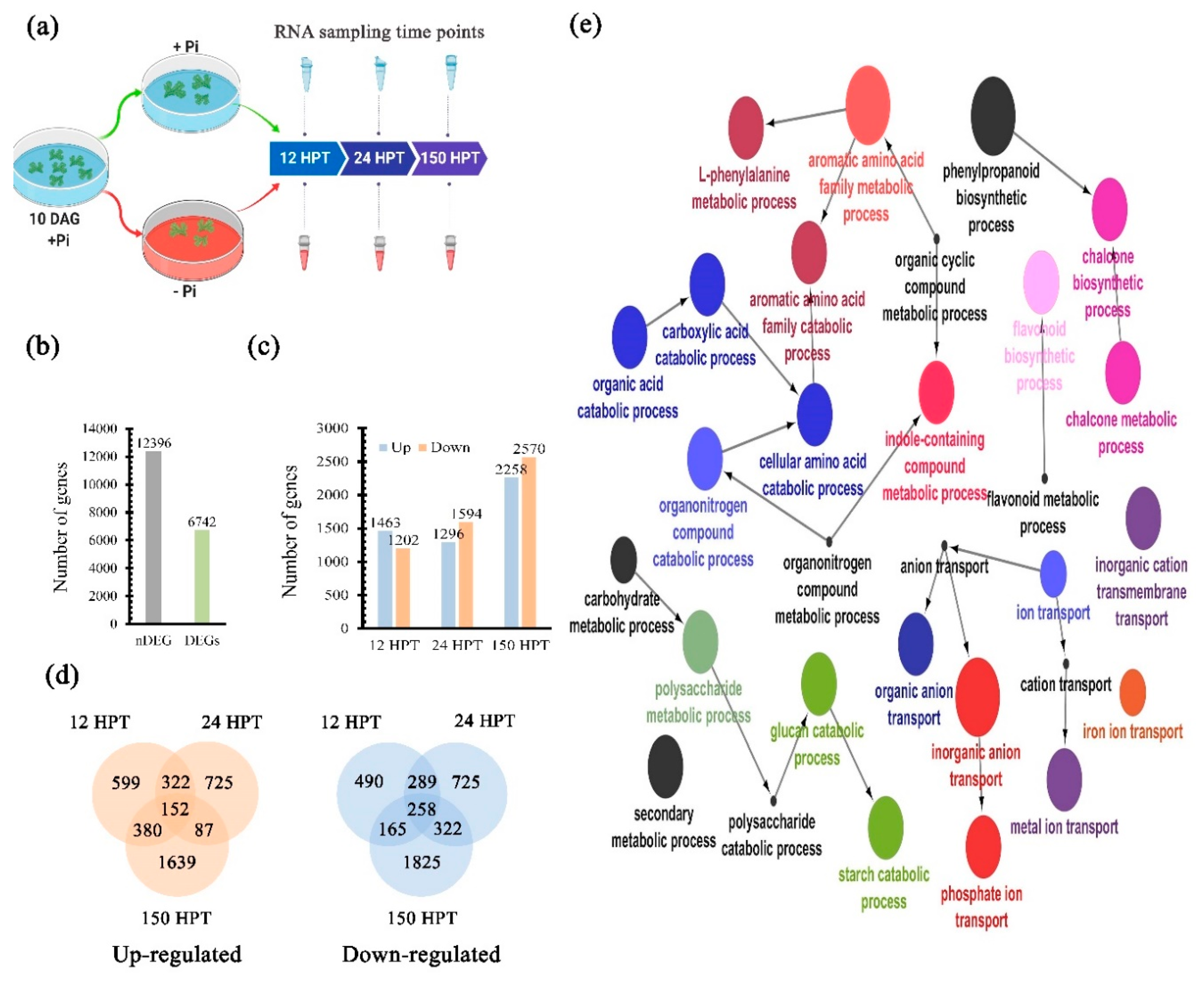
2.4. Transcriptional Dynamics in Response to Pi Availability
2.5. Transcriptional Regulation of the Local and Systemic Pi-Sensing Modules
2.6. Low Pi Promotes the Induction of Genes Related to Organic Acid Synthesis and Exudation
2.7. Pi Starvation Induces the Expression of Genes Involved in Lipid Turnover
2.8. Transcriptional Behavior of Genes Involved in ROS Synthesis in Pi-Starved Thallus
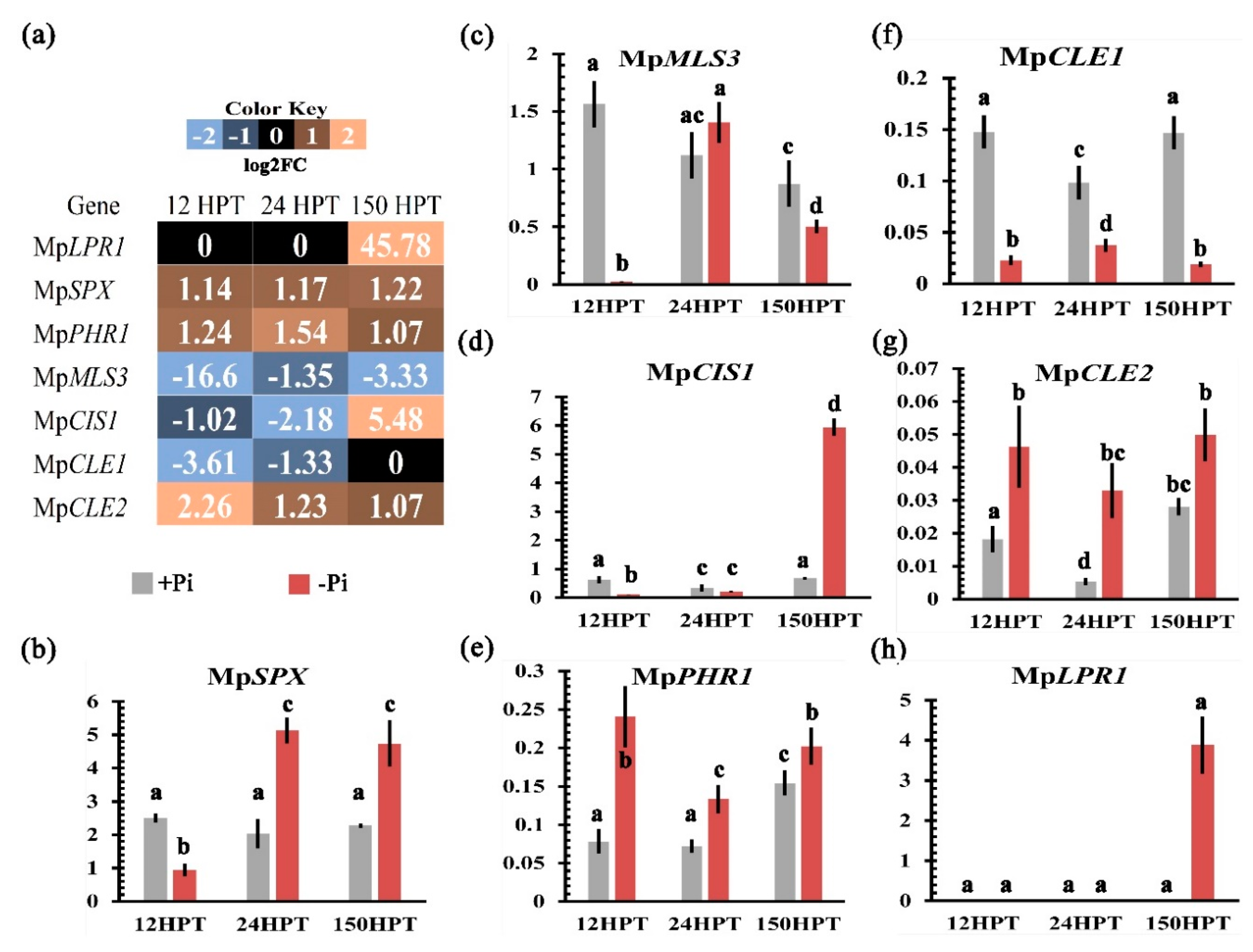
2.9. Validation of Transcriptome by qPCR
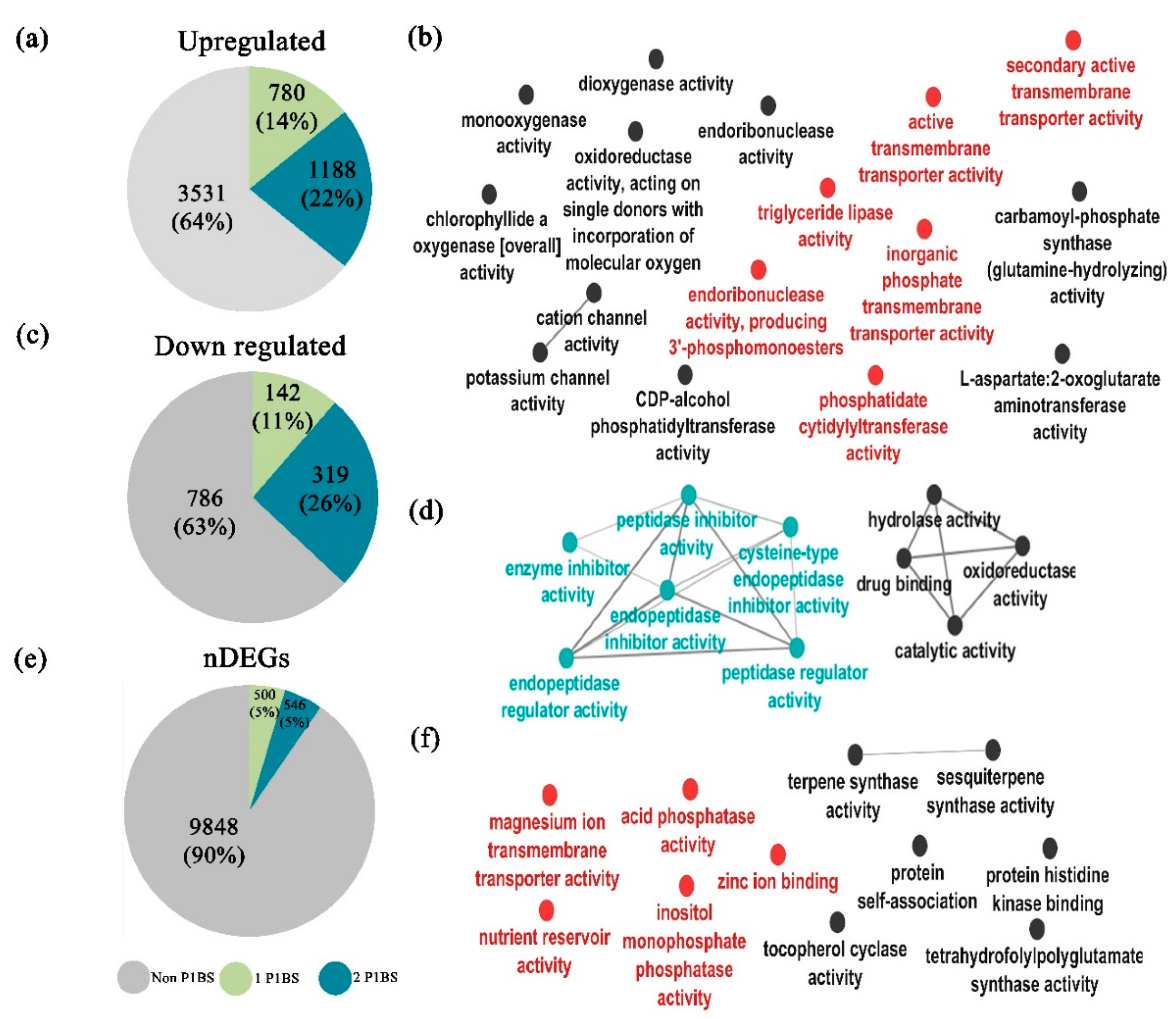
2.10. Determination of PHR1 Binding Sites (P1BS) Enrichment on DEGs
3. Discussion
3.1. Morpho-Physiological Responses upon Pi Starvation
3.2. Evolutionary Perspective of Pi Starvation Responses along Viridiplantae
3.3. Transcriptional Response to Cope with Limited Pi in M. polymorpha
4. Materials and Methods
4.1. Evolutionary Analysis
4.2. Plant Material and Growth Conditions
4.3. In Vivo Phosphatase Activity Assay
4.4. Free Phosphate Quantification
4.5. RNA Extraction, Sequencing, and Quantitative Real-Time qPCR Analysis
4.6. Differential Expressed Genes Determination and GO Enrichment Analysis
4.7. P1BS Search in DEGs
4.8. Analysis of P1BS Enrichment
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| Pi | Phosphate |
| P1BS | PHR1 binding sites |
| Ca | Calcium |
| Al | Aluminium |
| Fe | Iron |
| RSA | Root system architecture |
| OAs | Organic acids |
| TF | Transcription factor |
| IPP | Inositol polyphosphate |
| PM | Plasma membrane |
| +Pi | High phosphate availability |
| -Pi | Low phosphate availability |
| PSR | Phosphate starvation response |
| µM | Micromolar |
| HAS | Hours after sown |
| HPT | Hours post transference |
| DAS | Days after sown |
| FC | Fold change |
| DEGs | Differential expressed genes |
| FDR | False discovery rate |
| GO | Gene ontology |
| qPCR | Quantitative polymerase chain reaction |
| CREs | Cis regulatory elements |
References
- Sanderson, M.J. Molecular data from 27 proteins do not support a Precambrian origin of land plants. Am. J. Bot. 2003, 90, 954–956. [Google Scholar] [CrossRef] [PubMed]
- Delwiche, C.F.; Cooper, E.D. The evolutionary origin of a terrestrial flora. Curr. Biol. 2015, 25, R899–R910. [Google Scholar] [CrossRef] [PubMed]
- Pires, N.D.; Dolan, L. Morphological evolution in land plants: New designs with old genes. Philos. Trans. R. Soc. B 2012, 367, 508–518. [Google Scholar] [CrossRef] [PubMed]
- Ishizaki, K. Evolution of land plants: Insights from molecular studies on basal lineages. Biosci. Biotechnol. Biochem. 2017, 81, 73–80. [Google Scholar] [CrossRef]
- Dipp-Álvarez, M.; Cruz-Ramírez, A. A phylogenetic study of the ANT Family points to a preANT gene as the ancestor of basal and euANT transcription factors in land plants. Front. Plant Sci. 2019, 10, 17. [Google Scholar] [CrossRef] [PubMed]
- López-Bucio, J.; Cruz-Ramırez, A.; Herrera-Estrella, L. The role of nutrient availability in regulating root architecture. Curr. Opin. Plant Biol. 2003, 6, 280–287. [Google Scholar] [CrossRef]
- Puga, M.I.; Rojas-Triana, M.; de Lorenzo, L.; Leyva, A.; Rubio, V.; Paz-Ares, J. Novel signals in the regulation of Pi starvation responses in plants: Facts and promises. Curr. Opin. Plant Biol. 2017, 39, 40–49. [Google Scholar] [CrossRef]
- Gutiérrez-Alanís, D.; Ojeda-Rivera, J.O.; Yong-Villalobos, L.; Cárdenas-Torres, L.; Herrera-Estrella, L. Adaptation to phosphate scarcity: Tips from arabidopsis roots. Trends Plant Sci. 2018, 23, 721–730. [Google Scholar] [CrossRef]
- Raghothama, K.G. Phosphate acquisition. Annu. Rev. Plant Biol. 1999, 50, 665–693. [Google Scholar] [CrossRef]
- Bieleski, R.L. Phosphate pools, phosphate transport, and phosphate availability. Annu. Rev. Plant Physiol. 1973, 24, 225–252. [Google Scholar] [CrossRef]
- Marschner, H. Marschner’s Mineral Nutrition of Higher Plants; Academic Press: Amsterdam, The Netherlands, 2011. [Google Scholar]
- Lynch, J. Root architecture and plant productivity. Plant Physiol. 1995, 109, 7. [Google Scholar] [CrossRef] [PubMed]
- Thibaud, M.C.; Arrighi, J.F.; Bayle, V.; Chiarenza, S.; Creff, A.; Bustos, R.; Nussaume, L. Dissection of local and systemic transcriptional responses to phosphate starvation in Arabidopsis. Plant J. 2010, 64, 775–789. [Google Scholar] [CrossRef] [PubMed]
- Svistoonoff, S.; Creff, A.; Reymond, M.; Sigoillot-Claude, C.; Ricaud, L.; Blanchet, A.; Desnos, T. Root tip contact with low-phosphate media reprograms plant root architecture. Nat. Genet. 2007, 39, 792. [Google Scholar] [CrossRef] [PubMed]
- Balzergue, C.; Dartevelle, T.; Godon, C.; Laugier, E.; Meisrimler, C.; Teulon, J.M.; Müller, J. Low phosphate activates STOP1-ALMT1 to rapidly inhibit root cell elongation. Nat. Commun. 2017, 8, 15300. [Google Scholar] [CrossRef] [PubMed]
- Mora-Macías, J.; Ojeda-Rivera, J.O.; Gutiérrez-Alanís, D.; Yong-Villalobos, L.; Oropeza-Aburto, A.; Raya-González, J.; Herrera-Estrella, L. Malate-dependent Fe accumulation is a critical checkpoint in the root developmental response to low phosphate. Proc. Natl. Acad. Sci. USA 2017, 114, E3563–E3572. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, J.; Guo, J.; Zhou, F.; Singh, S.; Xu, X.; Huang, C.F. F-box protein RAE1 regulates the stability of the aluminum-resistance transcription factor STOP1 in Arabidopsis. Proc. Natl. Acad. Sci. USA 2019, 116, 319–327. [Google Scholar] [CrossRef]
- Kochian, L.V.; Piñeros, M.A.; Liu, J.; Magalhaes, J.V. Plant adaptation to acid soils: The molecular basis for crop aluminum resistance. Annu. Rev. Plant Biol. 2015, 66, 571–598. [Google Scholar] [CrossRef]
- Müller, J.; Toev, T.; Heisters, M.; Teller, J.; Moore, K.L.; Hause, G.; Abel, S. Iron-dependent callose deposition adjusts root meristem maintenance to phosphate availability. Dev. Cell 2015, 33, 216–230. [Google Scholar] [CrossRef]
- Song, S.K.; Hofhuis, H.; Lee, M.M.; Clark, S.E. Key divisions in the early Arabidopsis embryo require POL and PLL1 phosphatases to establish the root stem cell organizer and vascular axis. Dev. Cell 2008, 15, 98–109. [Google Scholar] [CrossRef]
- Gutiérrez-Alanís, D.; Yong-Villalobos, L.; Jiménez-Sandoval, P.; Alatorre-Cobos, F.; Oropeza-Aburto, A.; Mora-Macías, J.; Herrera-Estrella, L. Phosphate starvation-dependent iron mobilization induces CLE14 expression to trigger root meristem differentiation through CLV2/PEPR2 signaling. Dev. Cell 2017, 41, 555–570. [Google Scholar] [CrossRef] [PubMed]
- Rubio, V.; Linhares, F.; Solano, R.; Martín, A.C.; Iglesias, J.; Leyva, A.; Paz-Ares, J. A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev. 2001, 15, 2122–2133. [Google Scholar] [CrossRef]
- Wild, R.; Gerasimaite, R.; Jung, J.Y.; Truffault, V.; Pavlovic, I.; Schmidt, A.; Mayer, A. Control of eukaryotic phosphate homeostasis by inositol polyphosphate sensor domains. Science 2016, 352, 986–990. [Google Scholar] [CrossRef]
- Shimogawara, K.; Wykoff, D.D.; Usuda, H.; Grossman, A.R. Chlamydomonas reinhardtii mutants abnormal in their responses to phosphorus deprivation. Plant Physiol. 1999, 120, 685–694. [Google Scholar] [CrossRef]
- Wykoff, D.D.; Grossman, A.R.; Weeks, D.P.; Usuda, H.; Shimogawara, K. Psr1, a nuclear localized protein that regulates phosphorus metabolism in Chlamydomonas. Proc. Natl. Acad. Sci. USA 1999, 96, 15336–15341. [Google Scholar] [CrossRef]
- Nussaume, L.; Kanno, S.; Javot, H.; Marin, E.; Nakanishi, T.M.; Thibaud, M.C. Phosphate import in plants: Focus on the PHT1 transporters. Front. Plant Sci. 2011, 2, 83. [Google Scholar] [CrossRef]
- Kanno, S.; Arrighi, J.F.; Chiarenza, S.; Bayle, V.; Berthomé, R.; Péret, B.; Thibaud, M.C. A novel role for the root cap in phosphate uptake and homeostasis. Elife 2016, 5, e14577. [Google Scholar] [CrossRef]
- Daram, P.; Brunner, S.; Rausch, C.; Steiner, C.; Amrhein, N.; Bucher, M. Pht2; 1 encodes a low-affinity phosphate transporter from Arabidopsis. Plant Cell 1999, 11, 2153–2166. [Google Scholar] [CrossRef] [PubMed]
- Versaw, W.K.; Harrison, M.J. A chloroplast phosphate transporter, PHT2; 1, influences allocation of phosphate within the plant and phosphate-starvation responses. Plant Cell 2002, 14, 1751–1766. [Google Scholar] [CrossRef]
- Guo, B.; Irigoyen, S.; Fowler, T.B.; Versaw, W.K. Differential expression and phylogenetic analysis suggest specialization of plastid-localized members of the PHT4 phosphate transporter family for photosynthetic and heterotrophic tissues. Plant Signal. Behav. 2008, 3, 784–790. [Google Scholar] [CrossRef] [PubMed]
- Guo, B.; Jin, Y.; Wussler, C.; Blancaflor, E.B.; Motes, C.M.; Versaw, W.K. Functional analysis of the Arabidopsis PHT4 family of intracellular phosphate transporters. New Phytol. 2008, 177, 889–898. [Google Scholar] [CrossRef] [PubMed]
- Takabatake, R.; Hata, S.; Taniguchi, M.; Kouchi, H.; Sugiyama, T.; Izui, K. Isolation and characterization of cDNAs encoding mitochondrial phosphate transporters in soybean, maize, rice, and Arabidopsis. Plant Mol. Biol. 1999, 40, 479–486. [Google Scholar] [CrossRef] [PubMed]
- Jia, F.; Wan, X.; Zhu, W.; Sun, D.; Zheng, C.; Liu, P.; Huang, J. Overexpression of mitochondrial phosphate transporter 3 severely hampers plant development through regulating mitochondrial function in Arabidopsis. PLoS ONE 2015, 10, e0129717. [Google Scholar] [CrossRef]
- Liu, J.; Yang, L.; Luan, M.; Wang, Y.; Zhang, C.; Zhang, B.; Luan, S. A vacuolar phosphate transporter essential for phosphate homeostasis in Arabidopsis. Proc. Natl. Acad. Sci. USA 2015, 112, E6571–E6578. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Zhao, H.; Wan, R.; Liu, Y.; Xu, Z.; Tian, W.; Dolan, L. Identification of vacuolar phosphate efflux transporters in land plants. Nat. Plants 2019, 5, 84. [Google Scholar] [CrossRef]
- Essigmann, B.; Güler, S.; Narang, R.A.; Linke, D.; Benning, C. Phosphate availability affects the thylakoid lipid composition and the expression of SQD1, a gene required for sulfolipid biosynthesis in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 1998, 95, 1950–1955. [Google Scholar] [CrossRef]
- Cruz-Ramírez, A.; Oropeza-Aburto, A.; Razo-Hernández, F.; Ramírez-Chávez, E.; Herrera-Estrella, L. Phospholipase DZ2 plays an important role in extraplastidic galactolipid biosynthesis and phosphate recycling in Arabidopsis roots. Proc. Natl. Acad. Sci. USA 2006, 103, 6765–6770. [Google Scholar]
- Andersson, M.X.; Stridh, M.H.; Larsson, K.E.; Liljenberg, C.; Sandelius, A.S. Phosphate-deficient oat replaces a major portion of the plasma membrane phospholipids with the galactolipid digalactosyldiacylglycerol. FEBS Lett. 2003, 537, 128–132. [Google Scholar] [CrossRef]
- Bariola, P.A.; Howard, C.J.; Taylor, C.B.; Verburg, M.T.; Jaglan, V.D.; Green, P.J. The Arabidopsis ribonuclease gene RNS1 is tightly controlled in response to phosphate limitation. Plant J. 1994, 6, 673–685. [Google Scholar] [CrossRef]
- Li, D.; Zhu, H.; Liu, K.; Liu, X.; Leggewie, G.; Udvardi, M.; Wang, D. Purple acid phosphatases of Arabidopsis thaliana comparative analysis and differential regulation by phosphate deprivation. J. Biol. Chem. 2002, 277, 27772–27781. [Google Scholar] [CrossRef]
- Puga, M.I.; Mateos, I.; Charukesi, R.; Wang, Z.; Franco-Zorrilla, J.M.; de Lorenzo, L.; Leyva, A. SPX1 is a phosphate-dependent inhibitor of Phosphate Starvation Response 1 in Arabidopsis. Proc. Natl. Acad. Sci. USA 2014, 111, 14947–14952. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Ruan, W.; Shi, J.; Zhang, L.; Xiang, D.; Yang, C.; Shou, H. Rice SPX1 and SPX2 inhibit phosphate starvation responses through interacting with PHR2 in a phosphate-dependent manner. Proc. Natl. Acad. Sci. USA 2014, 111, 14953–14958. [Google Scholar] [CrossRef] [PubMed]
- Lv, Q.; Zhong, Y.; Wang, Y.; Wang, Z.; Zhang, L.; Shi, J.; Wu, P. SPX4 negatively regulates phosphate signaling and homeostasis through its interaction with PHR2 in rice. Plant Cell 2014, 26, 1586–1597. [Google Scholar] [CrossRef]
- Osorio, M.B.; Ng, S.; Berkowitz, O.; De Clercq, I.; Mao, C.; Shou, H.; Jost, R. SPX4 Acts on PHR1-Dependent and-Independent Regulation of Shoot Phosphorus Status in Arabidopsis. Plant Physiol. 2019, 332–352. [Google Scholar] [CrossRef] [PubMed]
- Shull, C. Phosphorus and the early maturity of plants. Trans. Illinois State Acad. Sci. 1925, 18, 165–171. [Google Scholar]
- Voth, P.D.; Hamner, K.C. Responses of Marchantia polymorpha to nutrient supply and photoperiod. Bot. Gaz. 1940, 102, 169–205. [Google Scholar] [CrossRef]
- Voth, P.D. Gemmae-cup production in Marchantia polymorpha and its response to calcium deficiency and supply of other nutrients. Bot. Gaz. 1941, 103, 310–325. [Google Scholar] [CrossRef]
- Bonnot, C.; Proust, H.; Pinson, B.; Colbalchini, F.P.; Lesly-Veillard, A.; Breuninger, H.; Dolan, L. Functional PTB phosphate transporters are present in streptophyte algae and early diverging land plants. New Phytol. 2017, 214, 1158–1171. [Google Scholar] [CrossRef] [PubMed]
- Shimamura, M. Marchantia polymorpha: Taxonomy, phylogeny and morphology of a model system. Plant Cell Physiol. 2016, 57, 230–256. [Google Scholar] [CrossRef] [PubMed]
- Bowman, J.L.; Kohchi, T.; Yamato, K.T.; Jenkins, J.; Shu, S.; Ishizaki, K.; Adam, C. Insights into land plant evolution garnered from the Marchantia polymorpha genome. Cell 2017, 171, 287–304. [Google Scholar] [CrossRef]
- Péret, B.; Clément, M.; Nussaume, L.; Desnos, T. Root developmental adaptation to phosphate starvation: Better safe than sorry. Trends Plant Sci. 2011, 16, 442–450. [Google Scholar] [CrossRef]
- Albert, N.W.; Thrimawithana, A.H.; McGhie, T.K.; Clayton, W.A.; Deroles, S.C.; Schwinn, K.E.; Davies, K.M. Genetic analysis of the liverwort Marchantia polymorpha reveals that R2R3 MYB activation of flavonoid production in response to abiotic stress is an ancient character in land plants. New Phytol. 2018, 218, 554–566. [Google Scholar] [CrossRef]
- Berland, H.; Albert, N.W.; Stavland, A.; Jordheim, M.; McGhie, T.K.; Zhou, Y.; Davies, K.M. Auronidins are a previously unreported class of flavonoid pigments that challenges when anthocyanin biosynthesis evolved in plants. Proc. Natl. Acad. Sci. USA 2019, 116, 20232–20239. [Google Scholar] [CrossRef]
- Del Pozo, J.C.; Allona, I.; Rubio, V.; Leyva, A.; De La Peña, A.; Aragoncillo, C.; Paz-Ares, J. A type 5 acid phosphatase gene from Arabidopsis thaliana is induced by phosphate starvation and by some other types of phosphate mobilising/oxidative stress conditions. Plant J. 1999, 19, 579–589. [Google Scholar] [CrossRef]
- Huang, Z.; Terpetschnig, E.; You, W.; Haugland, R.P. 2-(2′-Phosphoryloxyphenyl)-4 (3H)-quinazolinone derivatives as fluorogenic precipitating substrates of phosphatases. Anal. Biochem. 1992, 207, 32–39. [Google Scholar] [CrossRef]
- Misson, J.; Raghothama, K.G.; Jain, A.; Jouhet, J.; Block, M.A.; Bligny, R.; Doumas, P. A genome-wide transcriptional analysis using Arabidopsis thaliana Affymetrix gene chips determined plant responses to phosphate deprivation. Proc. Natl. Acad. Sci. USA 2005, 102, 11934–11939. [Google Scholar] [CrossRef]
- Secco, D.; Jabnoune, M.; Walker, H.; Shou, H.; Wu, P.; Poirier, Y.; Whelan, J. Spatio-temporal transcript profiling of rice roots and shoots in response to phosphate starvation and recovery. Plant Cell 2013, 25, 4285–4304. [Google Scholar] [CrossRef]
- Calderón-Vázquez, C.; Sawers, R.J.; Herrera-Estrella, L. Phosphate deprivation in maize: Genetics and genomics. Plant Physiol. 2011, 156, 1067–1077. [Google Scholar] [CrossRef]
- Zeng, H.; Wang, G.; Zhang, Y.; Hu, X.; Pi, E.; Zhu, Y.; Du, L. Genome-wide identification of phosphate-deficiency-responsive genes in soybean roots by high-throughput sequencing. Plant Soil 2016, 398, 207–227. [Google Scholar] [CrossRef]
- Bindea, G.; Mlecnik, B.; Hackl, H.; Charoentong, P.; Tosolini, M.; Kirilovsky, A.; Galon, J. ClueGO: A Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 2009, 25, 1091–1093. [Google Scholar] [CrossRef]
- Hirakawa, Y.; Uchida, N.; Yamaguchi, Y.L.; Tabata, R.; Ishida, S.; Ishizaki, K.; Bowman, J.L. Control of proliferation in the haploid meristem by CLE peptide signaling in Marchantia polymorpha. PLoS Genet. 2019, 15, e1007997. [Google Scholar] [CrossRef]
- Hirakawa, Y.; Fujimoto, T.; Ishida, S.; Uchida, N.; Sawa, S.; Kiyosue, T.; Bowman, J.L. Induction of Multichotomous Branching by CLAVATA Peptide in Marchantia polymorpha. Curr. Biol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Magalhaes, J.V.; Shaff, J.; Kochian, L.V. Aluminum-activated citrate and malate transporters from the MATE and ALMT families function independently to confer Arabidopsis aluminum tolerance. Plant J. 2019, 57, 389–399. [Google Scholar] [CrossRef]
- Mittler, R.; Vanderauwera, S.; Suzuki, N.; Miller, G.A.D.; Tognetti, V.B.; Vandepoele, K.; Van Breusegem, F. ROS signaling: The new wave? Trends Plant Sci. 2011, 16, 300–309. [Google Scholar] [CrossRef]
- Foreman, J.; Demidchik, V.; Bothwell, J.H.; Mylona, P.; Miedema, H.; Torres, M.A.; Davies, J.M. Reactive oxygen species produced by NADPH oxidase regulate plant cell growth. Nature 2003, 422, 442–446. [Google Scholar] [CrossRef]
- Carol, R.J.; Takeda, S.; Linstead, P.; Durrant, M.C.; Kakesova, H.; Derbyshire, P.; Dolan, L. A RhoGDP dissociation inhibitor spatially regulates growth in root hair cells. Nature 2005, 438, 1013–1016. [Google Scholar] [CrossRef]
- Saint-Marcoux, D.; Proust, H.; Dolan, L.; Langdale, J.A. Identification of reference genes for real-time quantitative PCR experiments in the liverwort Marchantia polymorpha. PLoS ONE 2015, 10, e0118678. [Google Scholar] [CrossRef]
- Bustos, R.; Castrillo, G.; Linhares, F.; Puga, M.I.; Rubio, V.; Pérez-Pérez, J.; Paz-Ares, J. A central regulatory system largely controls transcriptional activation and repression responses to phosphate starvation in Arabidopsis. PLoS Genet. 2010, 6, e1001102. [Google Scholar] [CrossRef]
- McLeay, R.C.; Bailey, T.L. Motif Enrichment Analysis: A unified framework and an evaluation on ChIP data. BMC Bioinform. 2010, 11, 165. [Google Scholar] [CrossRef]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37 (Suppl. 2), W202–W208. [Google Scholar] [CrossRef] [PubMed]
- Castrillo, G.; Teixeira, P.J.P.L.; Paredes, S.H.; Law, T.F.; de Lorenzo, L.; Feltcher, M.E.; Paz-Ares, J. Root microbiota drive direct integration of phosphate stress and immunity. Nature 2017, 543, 513–518. [Google Scholar] [CrossRef]
- Briat, J.F.; Rouached, H.; Tissot, N.; Gaymard, F.; Dubos, C. Integration of P, S, Fe, and Zn nutrition signals in Arabidopsis thaliana: Potential involvement of PHOSPHATE STARVATION RESPONSE 1 (PHR1). Front. Plant Sci. 2015, 6, 290. [Google Scholar] [CrossRef]
- Li, W.; Lan, P. Genome-wide analysis of overlapping genes regulated by iron deficiency and phosphate starvation reveals new interactions in Arabidopsis roots. BMC Res. Notes 2015, 8, 555. [Google Scholar] [CrossRef]
- Kato, H.; Ishizaki, K.; Kouno, M.; Shirakawa, M.; Bowman, J.L.; Nishihama, R.; Kohchi, T. Auxin-mediated transcriptional system with a minimal set of components is critical for morphogenesis through the life cycle in Marchantia polymorpha. PLoS Genet. 2015, 11, e1005084. [Google Scholar] [CrossRef]
- Flores-Sandoval, E.; Eklund, D.M.; Bowman, J.L. A simple auxin transcriptional response system regulates multiple morphogenetic processes in the liverwort Marchantia polymorpha. PLoS Genet. 2015, 11, e1005207. [Google Scholar] [CrossRef] [PubMed]
- López-Bucio, J.; Hernández-Abreu, E.; Sánchez-Calderón, L.; Nieto-Jacobo, M.F.; Simpson, J.; Herrera-Estrella, L. Phosphate availability alters architecture and causes changes in hormone sensitivity in the Arabidopsis root system. Plant Physiol. 2002, 129, 244–256. [Google Scholar] [CrossRef]
- Huang, K.L.; Ma, G.J.; Zhang, M.L.; Xiong, H.; Wu, H.; Zhao, C.Z.; Li, X.B. The ARF7 and ARF19 transcription factors positively regulate PHOSPHATE STARVATION RESPONSE1 in Arabidopsis roots. Plant Physiol. 2018, 178, 413–427. [Google Scholar] [CrossRef]
- Clayton, W.A.; Albert, N.W.; Thrimawithana, A.H.; McGhie, T.K.; Deroles, S.C.; Schwinn, K.E.; Davies, K.M. UVR8-mediated induction of flavonoid biosynthesis for UVB tolerance is conserved between the liverwort Marchantia polymorpha and flowering plants. Plant J. 2018, 96, 503–517. [Google Scholar] [CrossRef]
- Kubo, H.; Nozawa, S.; Hiwatashi, T.; Kondou, Y.; Nakabayashi, R.; Mori, T.; Ishizaki, K. Biosynthesis of riccionidins and marchantins is regulated by R2R3-MYB transcription factors in Marchantia polymorpha. J. Plant Res. 2018, 131, 849–864. [Google Scholar] [CrossRef]
- Moseley, J.; Grossman, A.R. Phosphate metabolism and responses to phosphorus deficiency. In The Chlamydomonas Sourcebook; Harris, E.H., Ed.; Academic Press: Amsterdam, The Netherlands, 2009; pp. 189–215. [Google Scholar]
- González-Muñoz, E.; Avendaño-Vázquez, A.O.; Montes, R.A.C.; de Folter, S.; Andrés-Hernández, L.; Abreu-Goodger, C.; Sawers, R.J. The maize (Zea mays ssp. mays var. B73) genome encodes 33 members of the purple acid phosphatase family. Front. Plant Sci. 2015, 6, 341. [Google Scholar]
- Mehra, P.; Pandey, B.K.; Giri, J. Improvement in phosphate acquisition and utilization by a secretory purple acid phosphatase (OsPAP21b) in rice. Plant Biotechnol. J. 2017, 15, 1054–1067. [Google Scholar] [CrossRef]
- Reinhard, C.T.; Planavsky, N.J.; Gill, B.C.; Ozaki, K.; Robbins, L.J.; Lyons, T.W.; Konhauser, K.O. Evolution of the global phosphorus cycle. Nature 2017, 541, 386–389. [Google Scholar] [CrossRef] [PubMed]
- Alcaraz, L.D.; Olmedo, G.; Bonilla, G.; Cerritos, R.; Hernández, G.; Cruz, A.; López, V. The genome of Bacillus coahuilensis reveals adaptations essential for survival in the relic of an ancient marine environment. Proc. Natl. Acad. Sci. USA 2008, 105, 5803–5808. [Google Scholar] [CrossRef]
- Gargouri, M.; Park, J.J.; Holguin, F.O.; Kim, M.J.; Wang, H.; Deshpande, R.R.; Gang, D.R. Identification of regulatory network hubs that control lipid metabolism in Chlamydomonas reinhardtii. J. Exp. Bot. 2015, 66, 4551–4566. [Google Scholar] [CrossRef]
- Kato, H.; Mutte, S.K.; Suzuki, H.; Crespo, I.; Das, S.; Radoeva, T.; Lindhoud, S. Design principles of a minimal auxin response system. Nat. Plants 2020, 6, 473–482. [Google Scholar] [CrossRef]
- Höwing, T.; Huesmann, C.; Hoefle, C.; Nagel, M.K.; Isono, E.; Huckelhoven, R.; Gietl, C. Endoplasmic reticulum KDEL-tailed cysteine endopeptidase 1 of Arabidopsis (AtCEP1) is involved in pathogen defense. Front. Plant Sci. 2014, 5, 58. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Capella-Gutiérrez, S.; Silla-Martínez, J.M.; Gabaldón, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef]
- Posada, D.; Crandall, K.A. Modeltest: Testing the model of DNA substitution. Bioinformatics 1998, 14, 817–818. [Google Scholar] [CrossRef]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In Proceedings of the 2010 Gateway Computing Environments Workshop (GCE), New Orleans, LA, USA, 14 November 2010; pp. 1–8. [Google Scholar]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Heinz, S.; Benner, C.; Spann, N.; Bertolino, E.; Lin, Y.C.; Laslo, P.; Cheng, J.X.; Murre, C.; Singh, H.; Glass, C.K. Simple Combinations of Lineage-Determining Transcription Factors Prime cis-Regulatory Elements Required for Macrophage and B Cell Identities. Mol. Cell 2010. [Google Scholar] [CrossRef]
- Wagih, O. ggseqlogo: A versatile R package for drawing sequence logos. Bioinformatics 2017. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rico-Reséndiz, F.; Cervantes-Pérez, S.A.; Espinal-Centeno, A.; Dipp-Álvarez, M.; Oropeza-Aburto, A.; Hurtado-Bautista, E.; Cruz-Hernández, A.; Bowman, J.L.; Ishizaki, K.; Arteaga-Vázquez, M.A.; et al. Transcriptional and Morpho-Physiological Responses of Marchantia polymorpha upon Phosphate Starvation. Int. J. Mol. Sci. 2020, 21, 8354. https://doi.org/10.3390/ijms21218354
Rico-Reséndiz F, Cervantes-Pérez SA, Espinal-Centeno A, Dipp-Álvarez M, Oropeza-Aburto A, Hurtado-Bautista E, Cruz-Hernández A, Bowman JL, Ishizaki K, Arteaga-Vázquez MA, et al. Transcriptional and Morpho-Physiological Responses of Marchantia polymorpha upon Phosphate Starvation. International Journal of Molecular Sciences. 2020; 21(21):8354. https://doi.org/10.3390/ijms21218354
Chicago/Turabian StyleRico-Reséndiz, Félix, Sergio Alan Cervantes-Pérez, Annie Espinal-Centeno, Melissa Dipp-Álvarez, Araceli Oropeza-Aburto, Enrique Hurtado-Bautista, Andrés Cruz-Hernández, John L. Bowman, Kimitsune Ishizaki, Mario A. Arteaga-Vázquez, and et al. 2020. "Transcriptional and Morpho-Physiological Responses of Marchantia polymorpha upon Phosphate Starvation" International Journal of Molecular Sciences 21, no. 21: 8354. https://doi.org/10.3390/ijms21218354
APA StyleRico-Reséndiz, F., Cervantes-Pérez, S. A., Espinal-Centeno, A., Dipp-Álvarez, M., Oropeza-Aburto, A., Hurtado-Bautista, E., Cruz-Hernández, A., Bowman, J. L., Ishizaki, K., Arteaga-Vázquez, M. A., Herrera-Estrella, L., & Cruz-Ramírez, A. (2020). Transcriptional and Morpho-Physiological Responses of Marchantia polymorpha upon Phosphate Starvation. International Journal of Molecular Sciences, 21(21), 8354. https://doi.org/10.3390/ijms21218354








