Associations of TIMP-3 Genetic Polymorphisms with EGFR Statuses and Cancer Clinicopathologic Development in Lung Adenocarcinoma Patients
Abstract
:1. Introduction
2. Results
2.1. General Characteristics of LADC Patients Harboring the Wild-Type (WT) or Mutant EGFR
2.2. Associations between TIMP-3 Candidate SNPs (rs9619311, rs9862, and rs11547635) and EGFR Mutations in LADC Patients with or without Cigarette Consumption
2.3. Correlations between Polymorphic Genotypes of TIMP-3 and Clinicopathological Characteristics of LADC Patients of Different Genders with the WT or Mutant EGFR
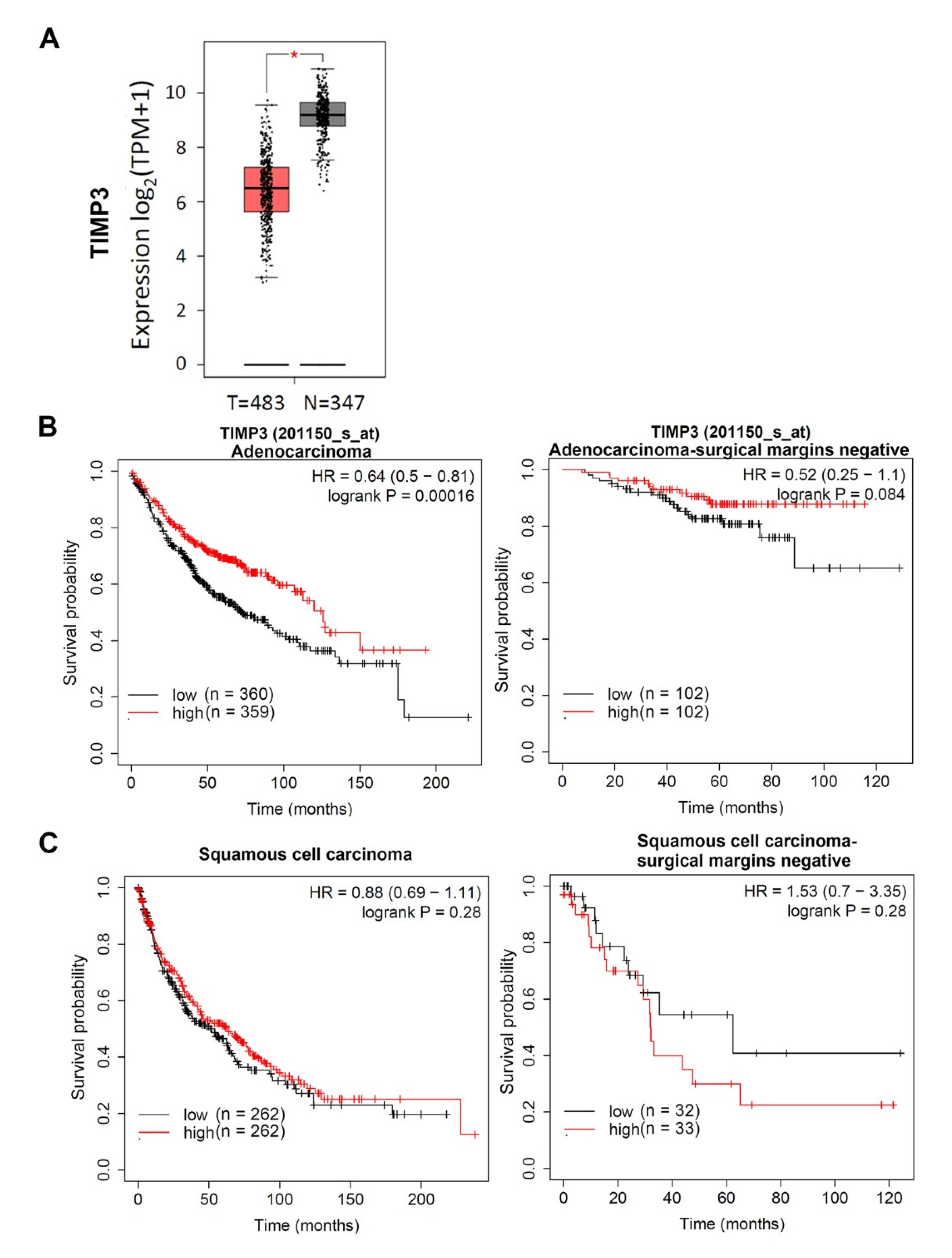
3. Discussion
4. Materials and Methods
4.1. Patient Specimens
4.2. DNA Extraction and EGFR Gene Sequencing from Tumor Tissues
4.3. Genomic TIMP3 SNPs Detected from Blood
4.4. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AORs | Adjusted odds ratios |
| CIs | Confidence intervals |
| EGFR | Epidermal growth factor receptor |
| GTEx | Genotype-Tissue Expression |
| LADC | Lung adenocarcinoma |
| MMP | Matrix metalloproteinase |
| NSCLC | Non-small-cell lung cancer |
| SCC | Squamous cell carcinoma |
| SNPs | Single-nucleotide polymorphisms |
| TCGA | The Cancer Genome Atlas |
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2020. CA Cancer J. Clin. 2020, 70, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Siegelin, M.D.; Borczuk, A.C. Epidermal growth factor receptor mutations in lung adenocarcinoma. Lab. Investig. 2014, 94, 129–137. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shigematsu, H.; Lin, L.; Takahashi, T.; Nomura, M.; Suzuki, M.; Wistuba, I.I.; Fong, K.M.; Lee, H.; Toyooka, S.; Shimizu, N.; et al. Clinical and biological features associated with epidermal growth factor receptor gene mutations in lung cancers. J. Natl. Cancer Inst. 2005, 97, 339–346. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dearden, S.; Stevens, J.; Wu, Y.L.; Blowers, D. Mutation incidence and coincidence in non small-cell lung cancer: Meta-analyses by ethnicity and histology (mutMap). Ann. Oncol. 2013, 24, 2371–2376. [Google Scholar] [CrossRef]
- Riely, G.J.; Pao, W.; Pham, D.; Li, A.R.; Rizvi, N.; Venkatraman, E.S.; Zakowski, M.F.; Kris, M.G.; Ladanyi, M.; Miller, V.A. Clinical course of patients with non-small cell lung cancer and epidermal growth factor receptor exon 19 and exon 21 mutations treated with gefitinib or erlotinib. Clin. Cancer Res. 2006, 12, 839–844. [Google Scholar] [CrossRef] [Green Version]
- Karachaliou, N.; Casas, C.; Queralt, C.M.D.L.; de Aguirre, I.; Melloni, B.; Cardenal, F.; Garcia-Gomez, R.; Massuti, B.; Sanchez, J.M.; Porta, R.; et al. Association of EGFR L858R Mutation in Circulating Free DNA with Survival in the EURTAC Trial. JAMA Oncol. 2015, 1, 149–157. [Google Scholar] [CrossRef] [Green Version]
- Gazdar, A.F. Activating and resistance mutations of EGFR in non-small-cell lung cancer: Role in clinical response to EGFR tyrosine kinase inhibitors. Oncogene 2009, 28, S24–S31. [Google Scholar] [CrossRef] [Green Version]
- Saito, M.; Suzuki, H.; Kono, K.; Takenoshita, S.; Kohno, T. Treatment of lung adenocarcinoma by molecular-targeted therapy and immunotherapy. Surg. Today 2018, 48, 1–8. [Google Scholar] [CrossRef]
- Brew, K.; Nagase, H. The tissue inhibitors of metalloproteinases (TIMPs): An ancient family with structural and functional diversity. Biochim. Biophys. Acta 2010, 1803, 55–71. [Google Scholar] [CrossRef] [Green Version]
- Hsu, C.Y.; Chang, G.C.; Chen, Y.J.; Hsu, Y.C.; Hsiao, Y.J.; Su, K.Y.; Chen, H.Y.; Lin, C.Y.; Chen, J.S.; Chen, Y.J.; et al. FAM198B is Associated with Prolonged Survival and Inhibits Metastasis in Lung Adenocarcinoma via Blockage of ERK-Mediated MMP-1 Expression. Clin. Cancer Res. 2018, 24, 916–926. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.J.; Chang, Y.J.; Kuo, Y.T.; Liang, P.H. Targeting beta-tubulin/CCT-beta complex induces apoptosis and suppresses migration and invasion of highly metastatic lung adenocarcinoma. Carcinogenesis 2020, 41, 699–710. [Google Scholar] [CrossRef]
- Kong, L.; Zhang, P.; Li, W.; Yang, Y.; Tian, Y.; Wang, X.; Chen, S.; Yang, Y.; Huang, T.; Zhao, T.; et al. KDM1A promotes tumor cell invasion by silencing TIMP3 in non-small cell lung cancer cells. Oncotarget 2016, 7, 27959–27974. [Google Scholar] [CrossRef] [Green Version]
- Zhu, Z.; Qian, Q.; Zhao, X.; Ma, L.; Chen, P. N(6)-methyladenosine ALKBH5 promotes non-small cell lung cancer progress by regulating TIMP3 stability. Gene 2020, 731, 144348. [Google Scholar] [CrossRef] [PubMed]
- Licchesi, J.D.; Westra, W.H.; Hooker, C.M.; Herman, J.G. Promoter hypermethylation of hallmark cancer genes in atypical adenomatous hyperplasia of the lung. Clin. Cancer Res. 2008, 14, 2570–2578. [Google Scholar] [CrossRef] [Green Version]
- Xu, P.W.; Xu, H.Y.; Liu, X.N.; Zhang, C.Y.; Tan, C.; Chen, C.M.; Zhang, H.; Jin, Y.T. Aberrant promoter methylation of cell adhesion-related genes associated with clinicopathologic features in non-small cell lung cancer in China. Cancer Biomark. 2013, 13, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.L.; Liu, Y.M.; Sung, T.Y.; Huang, T.C.; Cheng, Y.W.; Liou, J.P.; Pan, S.L. TIMP3 expression associates with prognosis in colorectal cancer and its novel arylsulfonamide inducer, MPT0B390, inhibits tumor growth, metastasis and angiogenesis. Theranostics 2019, 9, 6676–6689. [Google Scholar] [CrossRef] [PubMed]
- Lei, Y.; Liu, Z.; Yang, W. Negative correlation of cytoplasm TIMP3 with miR-222 indicates a good prognosis for NSCLC. Oncol. Targets Ther. 2018, 11, 5551–5557. [Google Scholar] [CrossRef] [Green Version]
- Gu, X.; Fu, M.; Ding, Y.; Ni, H.; Zhang, W.; Zhu, Y.; Tang, X.; Xiong, L.; Li, J.; Qiu, L.; et al. TIMP-3 expression associates with malignant behaviors and predicts favorable survival in HCC. PLoS ONE 2014, 9, e106161. [Google Scholar] [CrossRef]
- Qi, J.H.; Ebrahem, Q.; Moore, N.; Murphy, G.; Claesson-Welsh, L.; Bond, M.; Baker, A.; Anand-Apte, B. A novel function for tissue inhibitor of metalloproteinases-3 (TIMP3): Inhibition of angiogenesis by blockage of VEGF binding to VEGF receptor-2. Nat. Med. 2003, 9, 407–415. [Google Scholar] [CrossRef]
- Hammoud, L.; Burger, D.E.; Lu, X.; Feng, Q. Tissue inhibitor of metalloproteinase-3 inhibits neonatal mouse cardiomyocyte proliferation via EGFR/JNK/SP-1 signaling. Am. J. Physiol. Cell Physiol. 2009, 296, C735–C745. [Google Scholar] [CrossRef] [Green Version]
- Deshmukh, H.S.; Case, L.M.; Wesselkamper, S.C.; Borchers, M.T.; Martin, L.D.; Shertzer, H.G.; Nadel, J.A.; Leikauf, G.D. Metalloproteinases mediate mucin 5AC expression by epidermal growth factor receptor activation. Am. J. Respir. Crit. Care Med. 2005, 171, 305–314. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Middlebrooks, C.D.; Banday, A.R.; Matsuda, K.; Udquim, K.I.; Onabajo, O.O.; Paquin, A.; Figueroa, J.D.; Zhu, B.; Koutros, S.; Kubo, M.; et al. Association of germline variants in the APOBEC3 region with cancer risk and enrichment with APOBEC-signature mutations in tumors. Nat. Genet. 2016, 48, 1330–1338. [Google Scholar] [CrossRef]
- Wei, R.; Cao, L.; Pu, H.; Wang, H.; Zheng, Y.; Niu, X.; Weng, X.; Zhang, H.; Favus, M.; Zhang, L.; et al. TERT Polymorphism rs2736100-C is Associated with EGFR Mutation-Positive Non-Small Cell Lung Cancer. Clin. Cancer Res. 2015, 21, 5173–5180. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shiraishi, K.; Okada, Y.; Takahashi, A.; Kamatani, Y.; Momozawa, Y.; Ashikawa, K.; Kunitoh, H.; Matsumoto, S.; Takano, A.; Shimizu, K.; et al. Association of variations in HLA class II and other loci with susceptibility to EGFR-mutated lung adenocarcinoma. Nat. Commun. 2016, 7, 12451. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.L.; Chien, M.H.; Chou, Y.E.; Chang, J.H.; Liu, T.C.; Tsao, T.C.; Chou, M.C.; Yang, S.F. Association of EGFR mutations and HMGB1 genetic polymorphisms in lung adenocarcinoma patients. J. Cancer 2019, 10, 2907–2914. [Google Scholar] [CrossRef] [Green Version]
- Tsai, H.T.; Hsieh, M.J.; Chiou, H.L.; Lee, H.L.; Hsin, M.C.; Liou, Y.S.; Yang, C.C.; Yang, S.F.; Kuo, W.H. TIMP-3 −1296 T>C and TIMP-4 −55 T>C gene polymorphisms play a role in the susceptibility of hepatocellular carcinoma among women. Tumor Biol. 2014, 35, 8999–9007. [Google Scholar] [CrossRef]
- Bashash, M.; Shah, A.; Hislop, G.; Treml, M.; Bretherick, K.; Janoo-Gilani, R.; Leach, S.; Le, N.; Bajdik, C.; Brooks-Wilson, A. Genetic polymorphisms at TIMP3 are associated with survival of adenocarcinoma of the gastroesophageal junction. PLoS ONE 2013, 8, e59157. [Google Scholar] [CrossRef] [Green Version]
- Tseng, C.H.; Chiang, C.J.; Tseng, J.S.; Yang, T.Y.; Hsu, K.H.; Chen, K.C.; Wang, C.L.; Chen, C.Y.; Yen, S.H.; Tsai, C.M.; et al. EGFR mutation, smoking, and gender in advanced lung adenocarcinoma. Oncotarget 2017, 8, 98384–98393. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.L.; Yuan, J.Q.; Wang, K.F.; Fu, X.H.; Han, X.R.; Threapleton, D.; Yang, Z.Y.; Mao, C.; Tang, J.L. The prevalence of EGFR mutation in patients with non-small cell lung cancer: A systematic review and meta-analysis. Oncotarget 2016, 7, 78985–78993. [Google Scholar] [CrossRef] [Green Version]
- Yoon, H.Y.; Ryu, J.S.; Sim, Y.S.; Kim, D.; Lee, S.Y.; Choi, J.; Park, S.; Ryu, Y.J.; Lee, J.H.; Chang, J.H. Clinical significance of EGFR mutation types in lung adenocarcinoma: A multi-centre Korean study. PLoS ONE 2020, 15, e0228925. [Google Scholar] [CrossRef]
- Shi, Y.; Li, J.; Zhang, S.; Wang, M.; Yang, S.; Li, N.; Wu, G.; Liu, W.; Liao, G.; Cai, K.; et al. Molecular Epidemiology of EGFR Mutations in Asian Patients with Advanced Non-Small-Cell Lung Cancer of Adenocarcinoma Histology—Mainland China Subset Analysis of the PIONEER study. PLoS ONE 2015, 10, e0143515. [Google Scholar] [CrossRef]
- Anania, M.C.; Sensi, M.; Radaelli, E.; Miranda, C.; Vizioli, M.G.; Pagliardini, S.; Favini, E.; Cleris, L.; Supino, R.; Formelli, F.; et al. TIMP3 regulates migration, invasion and in vivo tumorigenicity of thyroid tumor cells. Oncogene 2011, 30, 3011–3023. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Das, A.M.; Bolkestein, M.; van der Klok, T.; Oude Ophuis, C.M.; Vermeulen, C.E.; Rens, J.A.; Dinjens, W.N.; Atmodimedjo, P.N.; Verhoef, C.; Koljenovic, S.; et al. Tissue inhibitor of metalloproteinase-3 (TIMP3) expression decreases during melanoma progression and inhibits melanoma cell migration. Eur. J. Cancer 2016, 66, 34–46. [Google Scholar] [CrossRef]
- Shen, B.; Jiang, Y.; Chen, Y.R.; Zheng, H.C.; Zeng, W.; Li, Y.Y.; Yin, A.; Nie, Y. Expression and inhibitory role of TIMP-3 in hepatocellular carcinoma. Oncol. Rep. 2016, 36, 494–502. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shinde, A.; Paez, J.S.; Libring, S.; Hopkins, K.; Solorio, L.; Wendt, M.K. Transglutaminase-2 facilitates extracellular vesicle-mediated establishment of the metastatic niche. Oncogenesis 2020, 9, 16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shinde, A.; Hardy, S.D.; Kim, D.; Akhand, S.S.; Jolly, M.K.; Wang, W.H.; Anderson, J.C.; Khodadadi, R.B.; Brown, W.S.; George, J.T.; et al. Spleen Tyrosine Kinase-Mediated Autophagy is Required for Epithelial-Mesenchymal Plasticity and Metastasis in Breast Cancer. Cancer Res. 2019, 79, 1831–1843. [Google Scholar] [CrossRef] [Green Version]
- Hardy, S.D.; Shinde, A.; Wang, W.H.; Wendt, M.K.; Geahlen, R.L. Regulation of epithelial-mesenchymal transition and metastasis by TGF-β, P-bodies, and autophagy. Oncotarget 2017, 8, 103302–103314. [Google Scholar] [CrossRef] [Green Version]
- Thomson, S.; Buck, E.; Petti, F.; Griffin, G.; Brown, E.; Ramnarine, N.; Iwata, K.K.; Gibson, N.; Haley, J.D. Epithelial to mesenchymal transition is a determinant of sensitivity of non-small-cell lung carcinoma cell lines and xenografts to epidermal growth factor receptor inhibition. Cancer Res. 2005, 65, 9455–9462. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Su, C.W.; Chang, Y.C.; Chien, M.H.; Hsieh, Y.H.; Chen, M.K.; Lin, C.W.; Yang, S.F. Loss of TIMP3 by promoter methylation of Sp1 binding site promotes oral cancer metastasis. Cell Death Dis. 2019, 10, 793. [Google Scholar] [CrossRef] [Green Version]
- Libring, S.; Shinde, A.; Chanda, M.K.; Nuru, M.; George, H.; Saleh, A.M.; Abdullah, A.; Kinzer-Ursem, T.L.; Calve, S.; Wendt, M.K.; et al. The Dynamic Relationship of Breast Cancer Cells and Fibroblasts in Fibronectin Accumulation at Primary and Metastatic Tumor Sites. Cancers 2020, 12, 1270. [Google Scholar] [CrossRef]
- Pereira, I.T.; Ramos, E.A.; Costa, E.T.; Camargo, A.A.; Manica, G.C.; Klassen, L.M.; Chequin, A.; Braun-Prado, K.; Pedrosa, F.D.O.; Souza, E.M.; et al. Fibronectin affects transient MMP2 gene expression through DNA demethylation changes in non-invasive breast cancer cell lines. PLoS ONE 2014, 9, e105806. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shinde, A.; Libring, S.; Alpsoy, A.; Abdullah, A.; Schaber, J.A.; Solorio, L.; Wendt, M.K. Autocrine Fibronectin Inhibits Breast Cancer Metastasis. Mol. Cancer Res. 2018, 16, 1579–1589. [Google Scholar] [CrossRef] [Green Version]
- Ishikawa, S.; Takenaka, K.; Yanagihara, K.; Miyahara, R.; Kawano, Y.; Otake, Y.; Hasegawa, S.; Wada, H.; Tanaka, F. Matrix metalloproteinase-2 status in stromal fibroblasts, not in tumor cells, is a significant prognostic factor in non-small-cell lung cancer. Clin. Cancer Res. 2004, 10, 6579–6585. [Google Scholar] [CrossRef] [Green Version]
- Ma, D.H.; Chen, J.I.; Zhang, F.; Hwang, D.G.; Chen, J.K. Inhibition of fibroblast-induced angiogenic phenotype of cultured endothelial cells by the overexpression of tissue inhibitor of metalloproteinase (TIMP)-3. J. Biomed. Sci. 2003, 10, 526–534. [Google Scholar] [CrossRef] [PubMed]
- Ahonen, M.; Baker, A.H.; Kahari, V.M. Adenovirus-mediated gene delivery of tissue inhibitor of metalloproteinases-3 inhibits invasion and induces apoptosis in melanoma cells. Cancer Res. 1998, 58, 2310–2315. [Google Scholar]
- Wilmanski, T.; Zhou, X.; Zheng, W.; Shinde, A.; Donkin, S.S.; Wendt, M.; Burgess, J.R.; Teegarden, D. Inhibition of pyruvate carboxylase by 1α,25-dihydroxyvitamin D promotes oxidative stress in early breast cancer progression. Cancer Lett. 2017, 411, 171–181. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nguyen, H.; Ivanova, V.S.; Kavandi, L.; Rodriguez, G.C.; Maxwell, G.L.; Syed, V. Progesterone and 1,25-dihydroxyvitamin D3; inhibit endometrial cancer cell growth by upregulating semaphorin 3B and semaphorin 3F. Mol. Cancer Res. 2011, 9, 1479–1492. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kornfeld, J.W.; Meder, S.; Wohlberg, M.; Friedrich, R.E.; Rau, T.; Riethdorf, L.; Löning, T.; Pantel, K.; Riethdorf, S. Overexpression of TACE and TIMP3 mRNA in head and neck cancer: Association with tumour development and progression. Br. J. Cancer 2011, 104, 138–145. [Google Scholar] [CrossRef] [Green Version]
- Bachman, K.E.; Herman, J.G.; Corn, P.G.; Merlo, A.; Costello, J.F.; Cavenee, W.K.; Baylin, S.B.; Graff, J.R. Methylation-associated silencing of the tissue inhibitor of metalloproteinase-3 gene suggest a suppressor role in kidney, brain, and other human cancers. Cancer Res. 1999, 59, 798–802. [Google Scholar]
- Czarnecka, K.H.; Szmyd, B.; Barańska, M.; Kaszkowiak, M.; Kordiak, J.; Antczak, A.; Pastuszak-Lewandoska, D.; Brzeziańska-Lasota, E. A Strong Decrease in TIMP3 Expression Mediated by the Presence of miR-17 and 20a Enables Extracellular Matrix Remodeling in the NSCLC Lesion Surroundings. Front. Oncol. 2019, 9, 1372. [Google Scholar] [CrossRef] [Green Version]
- Hu, J.; Ni, S.; Cao, Y.; Zhang, T.; Wu, T.; Yin, X.; Lang, Y.; Lu, H. The Angiogenic Effect of microRNA-21 Targeting TIMP3 through the Regulation of MMP2 and MMP9. PLoS ONE 2016, 11, e0149537. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, B.; Hsu, S.H.; Majumder, S.; Kutay, H.; Huang, W.; Jacob, S.T.; Ghoshal, K. TGFbeta-mediated upregulation of hepatic miR-181b promotes hepatocarcinogenesis by targeting TIMP3. Oncogene 2010, 29, 1787–1797. [Google Scholar] [CrossRef] [Green Version]
- Su, C.W.; Huang, Y.W.; Chen, M.K.; Su, S.C.; Yang, S.F.; Lin, C.W. Polymorphisms and Plasma Levels of Tissue Inhibitor of Metalloproteinase-3: Impact on Genetic Susceptibility and Clinical Outcome of Oral Cancer. Medicine 2015, 94, e2092. [Google Scholar] [CrossRef] [PubMed]
- Yongxin, S.; Wenjun, D.; Qiang, W.; Yunqing, S.; Liming, Z.; Chunsheng, W. Heavy smoking before coronary surgical procedures affects the native matrix metalloproteinase-2 and matrix metalloproteinase-9 gene expression in saphenous vein conduits. Ann. Thorac. Surg. 2013, 95, 55–61. [Google Scholar] [CrossRef]
- D’Angelo, S.P.; Pietanza, M.C.; Johnson, M.L.; Riely, G.J.; Miller, V.A.; Sima, C.S.; Zakowski, M.F.; Rusch, V.W.; Ladanyi, M.; Kris, M.G. Incidence of EGFR exon 19 deletions and L858R in tumor specimens from men and cigarette smokers with lung adenocarcinomas. J. Clin. Oncol. 2011, 29, 2066–2070. [Google Scholar]
- Kim, I.A.; Lee, J.S.; Kim, H.J.; Kim, W.S.; Lee, K.Y. Cumulative smoking dose affects the clinical outcomes of EGFR-mutated lung adenocarcinoma patients treated with EGFR-TKIs: A retrospective study. BMC Cancer 2018, 18, 768. [Google Scholar] [CrossRef]
- Lee, Y.S.; Kim, J.K.; Park, T.H.; Kim, Y.R.; Myeong, H.S.; Kwon, K.; Ro, Y.T.; Noh, Y.H.; Kim, S.Y. Systematic identification of novel biomarker signatures associated with acquired erlotinib resistance in cancer cells. Mol. Cell. Toxicol. 2016, 12, 139–148. [Google Scholar] [CrossRef]
- Su, K.Y.; Chen, H.Y.; Li, K.C.; Kuo, M.L.; Yang, J.C.; Chan, W.K.; Ho, B.C.; Chang, G.C.; Shih, J.Y.; Yu, S.L.; et al. Pretreatment epidermal growth factor receptor (EGFR) T790M mutation predicts shorter EGFR tyrosine kinase inhibitor response duration in patients with non-small-cell lung cancer. J. Clin. Oncol. 2012, 30, 433–440. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hsu, K.H.; Ho, C.C.; Hsia, T.C.; Tseng, J.S.; Su, K.Y.; Wu, M.F.; Chiu, K.L.; Yang, T.Y.; Chen, K.C.; Ooi, H.; et al. Identification of five driver gene mutations in patients with treatment-naïve lung adenocarcinoma in Taiwan. PLoS ONE 2015, 10, e0120852. [Google Scholar] [CrossRef] [Green Version]
| Subject Characteristic | Wild-Type (N = 109) | Mutation Type (N = 168) | p Value |
|---|---|---|---|
| Age, | |||
| mean ± SD (years) | 65.45 ± 13.34 | 65.69 ± 13.58 | 0.885 |
| Gender, n (%) | |||
| Male | 65 (59.6%) | 60 (35.7%) | <0.001 |
| Female | 44 (40.4%) | 108 (64.3%) | |
| Cigarette smoking, n (%) | |||
| Non-smoker | 49 (45.0%) | 130 (77.4%) | <0.001 |
| Ever-smoker | 60 (55.0%) | 38 (22.6%) | |
| Stage, n (%) | |||
| I+II | 25 (22.9%) | 47 (28.0%) | 0.350 |
| III+IV | 84 (77.1%) | 121 (72.0%) | |
| Tumor T status, n (%) | |||
| T1+T2 | 59 (54.1%) | 107 (63.7%) | 0.113 |
| T3+T4 | 50 (45.9%) | 61 (36.3%) | |
| Lymph node status, n (%) | |||
| Negative | 28 (25.7%) | 53 (31.5%) | 0.295 |
| Positive | 81 (74.3%) | 115 (68.5%) | |
| Distant metastasis, n (%) | |||
| Negative | 53 (48.6%) | 80 (47.6%) | 0.870 |
| Positive | 56 (51.4%) | 88 (52.4%) | |
| Cell differentiation, n (%) | |||
| Well/Moderately | 87 (79.8%) | 158 (94.0%) | <0.001 |
| Poorly | 22 (20.2%) | 10 (6.0%) |
| Genotype SNP | Wild-Type (N = 109) | Mutation Type (N = 168) | AOR (95% CI) | p-Value |
|---|---|---|---|---|
| rs9619311 | ||||
| TT | 89 (81.7%) | 133 (79.2%) | 1.00 | |
| TC | 19 (17.4%) | 33 (19.6%) | 1.228 (0.621–2.428) | 0.555 |
| CC | 1 (0.9%) | 2 (1.2%) | 1.229 (0.103–14.708) | 0.871 |
| TC+CC | 20 (18.3%) | 35 (20.8%) | 1.228 (0.631–2.390) | 0.545 |
| rs9862 | ||||
| CC | 43 (39.4%) | 49 (29.2%) | 1.00 | |
| CT | 47 (43.1%) | 69 (41.1%) | 1.423 (0.778–2.602) | 0.252 |
| TT | 19 (17.4%) | 50 (29.7%) | 2.530 (1.230–5.205) | 0.012 * |
| CT+TT | 66 (60.6%) | 119 (70.8%) | 1.748 (1.002–3.048) | 0.049 * |
| rs11547635 | ||||
| CC | 52 (47.7%) | 85 (50.6%) | 1.00 | |
| CT | 44 (40.4%) | 68 (40.5%) | 0.891 (0.514–1.544) | 0.680 |
| TT | 13 (11.9%) | 15 (8.9%) | 0.611 (0.253–1.474) | 0.273 |
| CT+TT | 57 (52.3%) | 83 (49.4%) | 0.826 (0.492–1.387) | 0.469 |
| Genotype SNP | Non-Smoking (N = 179) | Smoking (N = 98) | ||||
|---|---|---|---|---|---|---|
| Wild-Type (N = 49) | Mutation Type (N = 130) | p-Value | Wild-Type (N = 60) | Mutation Type (N = 38) | p-Value | |
| rs9619311 | ||||||
| TT | 39 (79.6%) | 106 (81.5%) | 50 (83.3%) | 27 (71.1%) | ||
| TC | 9 (18.4%) | 23 (17.7%) | 0.675 | 10 (16.7%) | 10 (26.3%) | 0.268 |
| CC | 1 (2.0%) | 1 (0.8%) | 0.423 | 0 (0.0%) | 1 (2.6%) | – |
| TC+CC | 10 (20.4%) | 24 (18.5%) | 0.556 | 10 (16.7%) | 11 (28.9%) | 0.190 |
| rs9862 | ||||||
| CC | 18 (36.7%) | 42 (32.3%) | 25 (41.7%) | 7 (18.4%) | ||
| CT | 22 (44.9%) | 50 (38.5%) | 0.853 | 25 (41.7%) | 19 (50.0%) | 0.169 |
| TT | 9 (18.4%) | 38 (29.2%) | 0.116 | 10 (16.6%) | 12 (31.6%) | 0.045 *,a |
| CT+TT | 31 (63.3%) | 88 (67.7%) | 0.386 | 35 (58.3%) | 31 (81.6%) | 0.070 |
| rs11547635 | ||||||
| CC | 25 (51.0%) | 62 (47.7%) | 27 (45.0%) | 23 (60.5%) | ||
| CT | 18 (36.7%) | 55 (42.3%) | 0.736 | 26 (43.3%) | 13 (34.2%) | 0.235 |
| TT | 6 (12.3%) | 13 (10.0%) | 0.661 | 7 (11.7%) | 2 (5.3%) | 0.317 |
| CT+TT | 24 (49.0%) | 68 (52.3%) | 0.903 | 33 (55.0%) | 15 (39.5%) | 0.168 |
| SNP Genotypes | Wild-Type (N = 109) | L858R | Exon 19 In-Frame Deletion | ||
|---|---|---|---|---|---|
| (N = 78) | AOR (95% CI) | (N = 81) | AOR (95% CI) | ||
| rs9619311 | |||||
| TT | 89 (81.7%) | 64 (82.1%) | 1.00 | 64 (79.0%) | 1.00 |
| TC | 19 (17.4%) | 13 (16.7%) | 0.701 (0.277–1.772) | 16 (19.8%) | 1.147 (0.514–2.557) |
| CC | 1 (0.9%) | 1 (1.2%) | 0.560 (0.033–9.449) | 1 (1.2%) | 1.340 (0.081–22.202) |
| TC+CC | 20 (18.3%) | 14 (17.9%) | 0.689 (0.280–1.695) | 17 (21.0%) | 1.157 (0.529–2.534) |
| rs9862 | |||||
| CC | 43 (39.4%) | 23 (29.5%) | 1.00 | 23 (28.4%) | 1.00 |
| CT | 47 (43.1%) | 30 (38.5%) | 1.343 (0.614–2.938) | 37 (45.7%) | 1.560 (0.760–3.204) |
| TT | 19 (17.4%) | 25 (32.0%) | 2.975 (1.182–7.488) a | 21 (25.9%) | 2.295 (0.973–5.412) |
| CT+TT | 66 (60.6%) | 55 (70.5%) | 1.787 (0.873–3.661) | 58 (71.6%) | 1.772 (0.908–3.458) |
| rs11547635 | |||||
| CC | 52 (47.7%) | 40 (51.3%) | 1.00 | 39 (48.1%) | 1.00 |
| CT | 44 (40.4%) | 32 (41.0%) | 1.005 (0.493–2.046) | 33 (40.7%) | 0.932 (0.486–1.787) |
| TT | 13 (11.9%) | 6 (7.7%) | 0.536 (0.166–1.731) | 9 (11.2%) | 0.885 (0.325–2.410) |
| CT+TT | 57 (52.3%) | 38 (48.7%) | 0.883 (0.453–1.722) | 42 (51.9%) | 0.922 (0.499–1.702) |
| Variable | All (N = 277) | Males (N = 125) | Females (N = 152) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| CC (N = 92) | CT+TT (N = 185) | p-Value | CC (N = 39) | CT+TT (N = 86) | p-Value | CC (N = 53) | CT+TT (N = 99) | p-Value | |
| Stage | |||||||||
| I+II | 26 (28.3%) | 46 (24.9%) | 0.544 | 13 (33.3%) | 14 (16.3%) | 0.032 *,a | 13 (24.5%) | 32 (32.3%) | 0.316 |
| III+IV | 66 (71.7%) | 107 (75.1%) | 26 (66.7%) | 72 (83.7%) | 40 (75.5%) | 67 (67.7%) | |||
| Tumor T status | |||||||||
| T1+T2 | 58 (63.0%) | 108 (58.4%) | 0.456 | 24 (61.5%) | 47 (54.7%) | 0.471 | 34 (64.2%) | 61 (61.6%) | 0.758 |
| T3+T4 | 34 (37.0%) | 77 (41.6%) | 15 (38.5%) | 39 (45.3%) | 19 (35.8%) | 38 (38.4%) | |||
| Lymph node status | |||||||||
| Negative | 29 (31.5%) | 52 (28.1%) | 0.556 | 10 (25.6%) | 19 (22.1%) | 0.663 | 19 (35.8%) | 33 (33.3%) | 0.755 |
| Positive | 63 (68.5%) | 133 (71.9%) | 29 (74.4%) | 67 (77.9%) | 34 (64.2%) | 66 (66.7%) | |||
| Distant metastasis | |||||||||
| Negative | 42 (45.7%) | 91 (49.2%) | 0.579 | 19 (48.7%) | 38 (44.2%) | 0.637 | 23 (43.4%) | 53 (53.5%) | 0.233 |
| Positive | 50 (54.3%) | 94 (50.8%) | 20 (51.3%) | 48 (55.8%) | 30 (56.6%) | 46 (46.5%) | |||
| Cell differentiation | |||||||||
| Well/Moderately | 84 (91.3%) | 161 (87.0%) | 0.294 | 33 (84.6%) | 69 (80.2%) | 0.558 | 51 (96.2%) | 92 (92.9%) | 0.412 |
| Poorly | 8 (8.7%) | 24 (13.0%) | 6 (15.4%) | 17 (19.8%) | 2 (3.8%) | 7 (7.1%) | |||
| Variable | Wild-type (N = 65) | Mutation type (N = 60) | ||||
|---|---|---|---|---|---|---|
| CC (N = 24) | CT+TT (N = 41) | p-Value | CC (N = 15) | CT+TT (N = 45) | p-Value | |
| Stage | ||||||
| I+II | 6 (25.0%) | 6 (14.6%) | 0.299 | 7 (46.7%) | 8 (17.8%) | 0.025 *,a |
| III+IV | 18 (75.0%) | 35 (85.4%) | 8 (53.3%) | 37 (82.2%) | ||
| Tumor T status | ||||||
| T1+T2 | 15 (62.5%) | 21 (51.2%) | 0.377 | 9 (60.0%) | 15 (57.8%) | 0.880 |
| T3+T4 | 9 (37.5%) | 20 (48.8%) | 6 (40.0%) | 19 (42.2%) | ||
| Lymph node status | ||||||
| Negative | 3 (12.5%) | 10 (24.4%) | 0.247 | 7 (46.7%) | 9 (20.0%) | 0.043 *,b |
| Positive | 21 (87.5%) | 31 (75.6%) | 8 (53.3%) | 36 (80.0%) | ||
| Distant metastasis | ||||||
| Negative | 11 (45.8%) | 20 (48.8%) | 0.818 | 8 (53.3%) | 18 (40.0%) | 0.367 |
| Positive | 13 (54.2%) | 21 (51.2%) | 7 (46.7%) | 27 (60.0%) | ||
| Cell differentiation | ||||||
| Well/Moderately | 19 (79.2%) | 30 (73.2%) | 0.588 | 14 (93.3%) | 39 (86.7%) | 0.486 |
| Poorly | 5 (20.8%) | 11 (26.8%) | 1 (6.7%) | 6 (13.3%) | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chang, J.-H.; Lai, T.-C.; Yang, P.-J.; Shih, P.-C.; Yang, Y.-C.; Lee, K.-L.; Liu, T.-C.; Tsao, T.C.-Y.; Yang, S.-F.; Chien, M.-H. Associations of TIMP-3 Genetic Polymorphisms with EGFR Statuses and Cancer Clinicopathologic Development in Lung Adenocarcinoma Patients. Int. J. Mol. Sci. 2020, 21, 8023. https://doi.org/10.3390/ijms21218023
Chang J-H, Lai T-C, Yang P-J, Shih P-C, Yang Y-C, Lee K-L, Liu T-C, Tsao TC-Y, Yang S-F, Chien M-H. Associations of TIMP-3 Genetic Polymorphisms with EGFR Statuses and Cancer Clinicopathologic Development in Lung Adenocarcinoma Patients. International Journal of Molecular Sciences. 2020; 21(21):8023. https://doi.org/10.3390/ijms21218023
Chicago/Turabian StyleChang, Jer-Hwa, Tsung-Ching Lai, Po-Jen Yang, Pei-Chun Shih, Yi-Chieh Yang, Kai-Ling Lee, Tu-Chen Liu, Thomas Chang-Yao Tsao, Shun-Fa Yang, and Ming-Hsien Chien. 2020. "Associations of TIMP-3 Genetic Polymorphisms with EGFR Statuses and Cancer Clinicopathologic Development in Lung Adenocarcinoma Patients" International Journal of Molecular Sciences 21, no. 21: 8023. https://doi.org/10.3390/ijms21218023
APA StyleChang, J.-H., Lai, T.-C., Yang, P.-J., Shih, P.-C., Yang, Y.-C., Lee, K.-L., Liu, T.-C., Tsao, T. C.-Y., Yang, S.-F., & Chien, M.-H. (2020). Associations of TIMP-3 Genetic Polymorphisms with EGFR Statuses and Cancer Clinicopathologic Development in Lung Adenocarcinoma Patients. International Journal of Molecular Sciences, 21(21), 8023. https://doi.org/10.3390/ijms21218023






