A Toxicity Prediction Tool for Potential Agonist/Antagonist Activities in Molecular Initiating Events Based on Chemical Structures
Abstract
:1. Introduction
2. Results and Discussion
2.1. Distributions of Active and Inactive Compounds
2.2. Models and Performances
2.3. Comparison with the Tox21 Data Challenge 2014
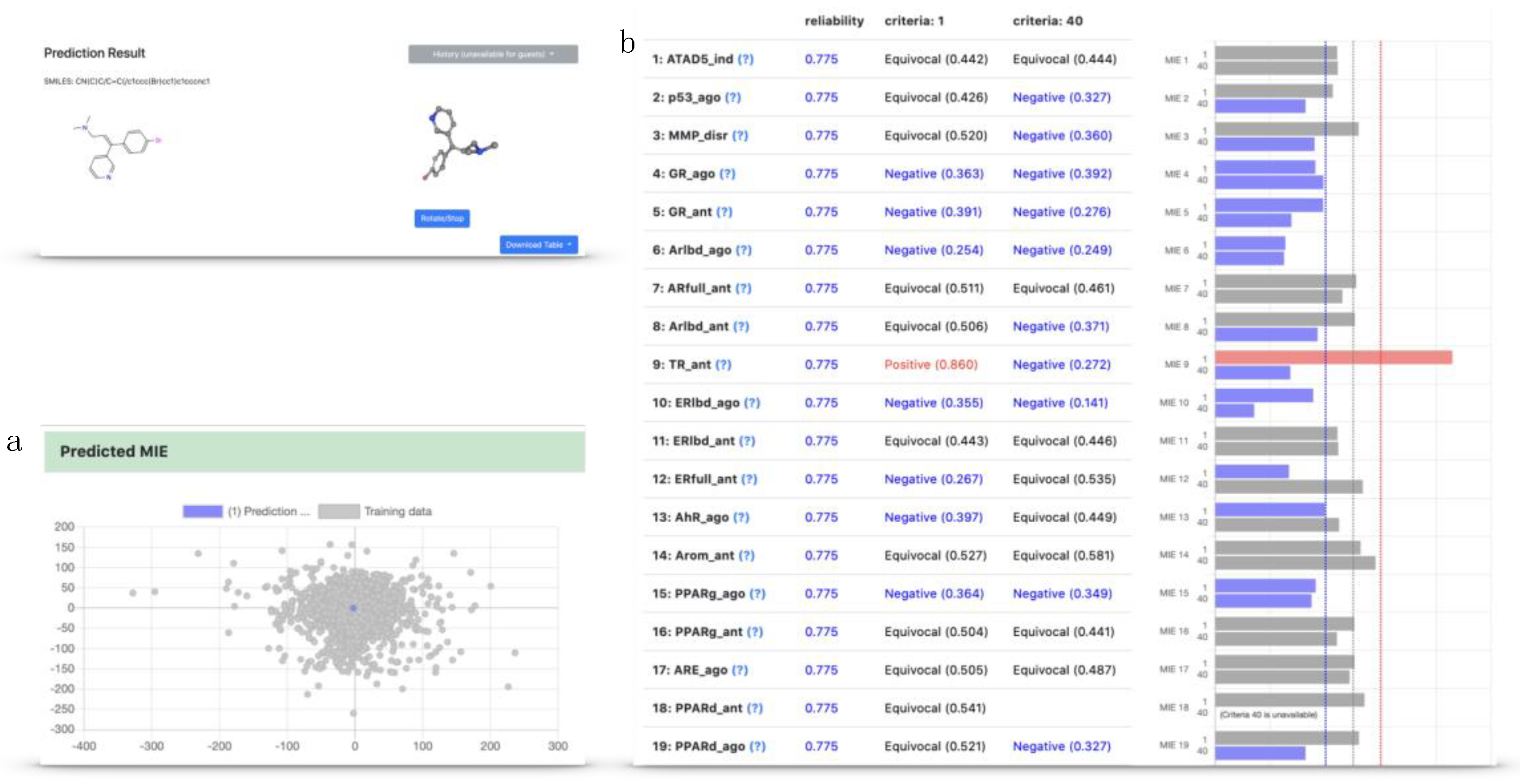
2.4. Implementation of the Models in the Toxicity Predictor
3. Materials and Methods
3.1. Biological Overview of Modeled MIEs
3.2. Data Source
3.3. qHTS Data Analysis
3.4. Conformations and Descriptors
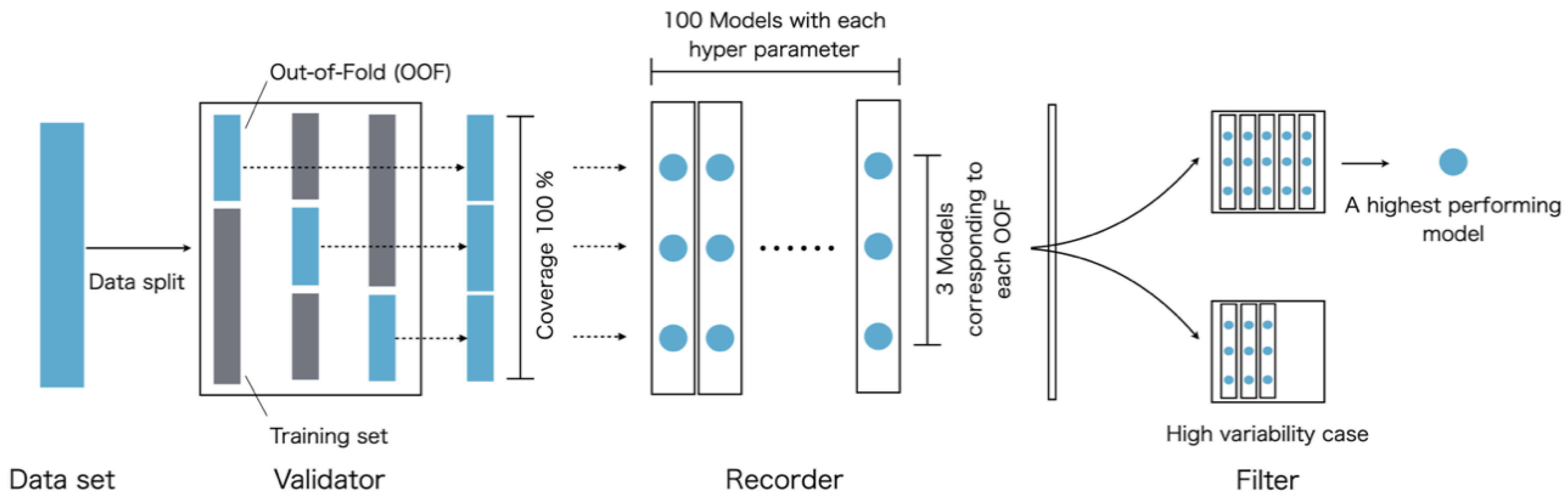
3.5. ML Algorithm and Modeling Scheme
3.6. Evaluation Metrics
- (1)
- SE: accuracy of predicting “positive” (active) when the true outcome is positive.
- (2)
- SP: accuracy of predicting “negative” (inactive) when the true outcome is negative.
- (3)
- ACC: the number of correctly predicted samples divided by the total number of samples.
- (4)
- BAC: average between SE and SP.
- (5)
- MCC: used as a measure to assess the classification accuracy of the models for an unbalanced dataset [71].
- (6)
- AUC: a graph showing the performance of a classification model at all classification thresholds. This curve plots two parameters: (i) SE and (ii) 1–SP [72].
3.7. Applicability Domain
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hansch, C.; Maloney, P.; Fujita, T.; Muir, R.M. Correlation of Biological Activity of Phenoxyacetic Acids with Hammett Substituent Constants and Partition Coefficients. Nature 1962, 194, 178–180. [Google Scholar] [CrossRef]
- Hansch, C.; Fujita, T. p-σ-π Analysis. A Method for the Correlation of Biological Activity and Chemical Structure. J. Am. Chem. Soc. 1964, 86, 1616–1626. [Google Scholar] [CrossRef]
- Gombar, V.K.; Enslein, K.; Blake, B.W. Assessment of developmental toxicity potential of chemicals by quantitative structure-toxicity relationship models. Chemosphere 1995, 31, 2499–2510. [Google Scholar] [CrossRef]
- van de Waterbeemd, H.; Gifford, E. ADMET in-silico modelling: Towards prediction paradise? Nat. Rev. Drug Discov. 2003, 2, 192–204. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S. Computer-aided drug discovery and development. Methods Mol. Biol. 2011, 716, 23–38. [Google Scholar] [CrossRef]
- Macalino, S.J.; Gosu, V.; Hong, S.; Choi, S. Role of computer-aided drug design in modern drug discovery. Arch. Pharmacal Res. 2015, 38, 1686–1701. [Google Scholar] [CrossRef]
- Flecknell, P. Replacement, reduction and refinement. ALTEX 2002, 19, 73–78. [Google Scholar]
- Contrera, J.F.; Matthews, E.J.; Kruhlak, N.L.; Benz, R.D. In silico screening of chemicals for bacterial mutagenicity using electrotopological E-state indices and MDL QSAR software. Regul. Toxicol. Pharmacol. 2005, 43, 313–323. [Google Scholar] [CrossRef]
- Ambure, P.; Halder, A.K.; Díaz, H.G.; Cordeiro, N. QSAR-Co: An Open Source Software for Developing Robust Multitasking or Multitarget Classification-Based QSAR Models. J. Chem. Inf. Model. 2019, 59, 2538–2544. [Google Scholar] [CrossRef]
- Mansouri, K.; Grulke, C.M.; Judson, R.S.; Williams, A.J. OPERA models for predicting physicochemical properties and environmental fate endpoints. J. Cheminformatics 2018, 10, 10. [Google Scholar] [CrossRef] [Green Version]
- Thomas, R.S.; Paules, R.S.; Simeonov, A.; Fitzpatrick, S.C.; Crofton, K.M.; Casey, W.M.; Mendrick, D.L. The US Federal Tox21 Program: A strategic and operational plan for continued leadership. ALTEX 2018, 35, 163–168. [Google Scholar] [CrossRef] [PubMed]
- Xia, M.; Huang, R.; Shi, Q.; Boyd, W.A.; Zhao, J.; Sun, N.; Rice, J.R.; Dunlap, P.E.; Hackstadt, A.J.; Bridge, M.F.; et al. Comprehensive Analyses and Prioritization of Tox21 10K Chemicals Affecting Mitochondrial Function by in-Depth Mechanistic Studies. Environ. Health Perspect. 2018, 126, 077010. [Google Scholar] [CrossRef]
- Ankley, G.T.; Bennett, R.S.; Erickson, R.J.; Hoff, D.J.; Hornung, M.W.; Johnson, R.D.; Mount, D.R.; Nichols, J.W.; Russom, C.L.; Schmieder, P.K.; et al. Adverse outcome pathways: A conceptual framework to support ecotoxicology research and risk assessment. Environ. Toxicol. Chem. 2010, 29, 730–741. [Google Scholar] [CrossRef]
- Allen, T.E.; Goodman, J.M.; Gutsell, S.; Russell, P.J. Defining molecular initiating events in the adverse outcome pathway framework for risk assessment. Chem. Res. Toxicol. 2014, 27, 2100–2112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dix, D.J.; Houck, K.A.; Martin, M.T.; Richard, A.M.; Setzer, R.W.; Kavlock, R.J. The ToxCast Program for Prioritizing Toxicity Testing of Environmental Chemicals. Toxicol. Sci. 2007, 95, 5–12. [Google Scholar] [CrossRef] [PubMed]
- Diamanti-Kandarakis, E.; Bourguignon, J.P.; Giudice, L.C.; Hauser, R.; Prins, G.S.; Soto, A.M.; Zoeller, R.T.; Gore, A.C. Endocrine-disrupting chemicals: An Endocrine Society scientific statement. Endocr. Rev. 2009, 30, 293–342. [Google Scholar] [CrossRef]
- Min, J.; Lee, S.K.; Gu, M.B. Effects of endocrine disrupting chemicals on distinct expression patterns of estrogen receptor, cytochrome P450 aromatase and p53 genes in oryzias latipes liver. J. Biochem. Mol. Toxicol. 2003, 17, 272–277. [Google Scholar] [CrossRef]
- Mansouri, K.; Kleinstreuer, N.; Abdelaziz, A.M.; Alberga, D.; Alves, V.M.; Andersson, P.L.; Andrade, C.H.; Bai, F.; Balabin, I.; Ballabio, D.; et al. CoMPARA: Collaborative Modeling Project for Androgen Receptor Activity. Environ. Health Perspect. 2020, 128, 27002. [Google Scholar] [CrossRef]
- Zhang, J.; Mucs, D.; Norinder, U.; Svensson, F. LightGBM: An Effective and Scalable Algorithm for Prediction of Chemical Toxicity-Application to the Tox21 and Mutagenicity Data Sets. J. Chem. Inf. Model. 2019, 59, 4150–4158. [Google Scholar] [CrossRef]
- Norinder, U.; Boyer, S. Conformal Prediction Classification of a Large Data Set of Environmental Chemicals from ToxCast and Tox21 Estrogen Receptor Assays. Chem. Res. Toxicol. 2016, 29, 1003–1010. [Google Scholar] [CrossRef]
- Banerjee, P.; Siramshetty, V.B.; Drwal, M.N.; Preissner, R. Computational methods for prediction of in vitro effects of new chemical structures. J. Cheminformatics 2016, 8, 51. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, T.; Guestrin, C. XGBoost: A scalable tree boosting system. arXiv 2016, arXiv:1603.02754. [Google Scholar]
- Sheridan, R.P.; Wang, W.M.; Liaw, A.; Ma, J.; Gifford, E.M. Extreme Gradient Boosting as a Method for Quantitative Structure–Activity Relationships. J. Chem. Inf. Model. 2016, 56, 2353–2360. [Google Scholar] [CrossRef] [PubMed]
- Attene-Ramos, M.S.; Miller, N.; Huang, R.; Michael, S.; Itkin, M.; Kavlock, R.J.; Austin, C.P.; Shinn, P.; Simeonov, A.; Tice, R.R.; et al. The Tox21 robotic platform for assessment of environmental chemicals–from vision to reality. Drug Discov. Today 2013, 18, 716–723. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Joonho, G.; Hyunjoong, K. RHSBoost: Improving classification performance in imbalance data. Comput. Stat. Data Anal. 2017, 111. [Google Scholar] [CrossRef]
- Ezzat, A.; Wu, M.; Li, X.; Kwoh, C. Drug-target interaction prediction via class imbalance-aware ensemble learning. BMC Bioinform. 2016, 17, 509. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Judson, R.; Houck, K.; Martin, M.; Richard, A.M.; Knudsen, T.B.; Shah, I.; Little, S.; Wambaugh, J.; Setzer, R.W.; Kothya, P.; et al. Editor’s Highlight: Analysis of the Effects of Cell Stress and Cytotoxicity on In Vitro Assay Activity Across a Diverse Chemical and Assay Space. Toxicol. Sci. 2016, 152, 323–339. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Available online: https://tripod.nih.gov/tox21/challenge/index.jsp (accessed on 31 July 2020).
- Available online: https://tripod.nih.gov/tox21/challenge/leaderboard.jsp (accessed on 31 July 2020).
- Sahigara, F.; Mansouri, K.; Ballabio, D.; Mauri, A.; Consonni, V.; Todeschini, R. Comparison of Different Approaches to Define the Applicability Domain of QSAR Models. Molecules 2012, 17, 4791–4810. [Google Scholar] [CrossRef] [Green Version]
- Dragos, H.; Gilles, M.; Alexandre, V. Predicting the predictability: A unified approach to the applicability domain problem of qsar models. J. Chem. Inf. Model. 2009, 49, 1762–1776. [Google Scholar] [CrossRef]
- Barouki, R.; Aggerbeck, M.; Aggerbeck, L.; Coumoul, X. The aryl hydrocarbon receptor system. Drug Metab. Drug Interact. 2012, 27, 3–8. [Google Scholar] [CrossRef]
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/720719 (accessed on 31 July 2020).
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/743053 (accessed on 31 July 2020).
- Fuentes, N.; Silveyra, P. Estrogen receptor signaling mechanisms. Adv. Protein Chem. Struct. Biol. 2019, 116, 135–170. [Google Scholar] [CrossRef] [PubMed]
- Ranhotra, H.S. Estrogen-related receptor alpha and cancer: Axis of evil. J. Recept Signal. Transduct Res. 2015, 35, 505–508. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.R.; Jeung, E.B.; Cho, M.H.; Kim, T.H.; Leung, P.C.; Choi, K.C. Molecular mechanism(s) of endocrine-disrupting chemicals and their potent oestrogenicity in diverse cells and tissues that express oestrogen receptors. J. Cell Mol. Med. 2013, 17, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/743139 (accessed on 31 July 2020).
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/1347030 (accessed on 31 July 2020).
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/1224895 (accessed on 31 July 2020).
- Brucker-Davis, F. Effects of environmental synthetic chemicals on thyroid function. Thyroid 1998, 8, 827–856. [Google Scholar] [CrossRef] [PubMed]
- Howdeshell, K.L. A model of the development of the brain as a construct of the thyroid system. Environ. Health Perspect. 2002, 110 (Suppl. 3), 337–348. [Google Scholar] [CrossRef] [Green Version]
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/743140 (accessed on 31 July 2020).
- Li, G.; Guo, G.L. Farnesoid X receptor, the bile acid sensing nuclear receptor, in liver regeneration. Acta Pharm. Sin. B 2015, 5, 93–98. [Google Scholar] [CrossRef] [Green Version]
- Huang, X.F.; Zhao, W.Y.; Huang, W.D. FXR and liver carcinogenesis. Acta Pharm. Sin. 2015, 36, 37–43. [Google Scholar] [CrossRef] [Green Version]
- Hakkola, J.; Bernasconi, C.; Coecke, S.; Richert, L.; Andersson, T.B.; Pelkonen, O. Cytochrome P450 Induction and Xeno-Sensing Receptors Pregnane X Receptor, Constitutive Androstane Receptor, Aryl Hydrocarbon Receptor and Peroxisome Proliferator-Activated Receptor α at the Crossroads of Toxicokinetics and Toxicodynamics. Basic Clin. Pharmacol. Toxicol. 2018, 123, 42–50. [Google Scholar] [CrossRef] [Green Version]
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/1347033 (accessed on 31 July 2020).
- Ghyselinck, N.B.; Duester, G. Retinoic acid signaling pathways. Dev. Camb. Engl. 2019, 146, dev167502. [Google Scholar] [CrossRef] [Green Version]
- Toporova, L.; Balaguer, P. Nuclear receptors are the major targets of endocrine disrupting chemicals. Mol. Cell. Endocrinol. 2020, 502, 15. [Google Scholar] [CrossRef]
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/1159531 (accessed on 31 July 2020).
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/743241 (accessed on 31 July 2020).
- Buendia, I.; Michalska, P.; Navarro, E.; Gameiro, I.; Egea, J.; León, R. Nrf2-ARE pathway: An emerging target against oxidative stress and neuroinflammation in neurodegenerative diseases. Pharmacology 2016, 157, 84–104. [Google Scholar] [CrossRef]
- Reuter, S.; Gupta, S.C.; Chaturvedi, M.M.; Aggarwal, B.B. Oxidative stress, inflammation, and cancer: How are they linked? Free Radic. Biol. Med. 2010, 49, 1603–1616. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hayden, M.S.; Ghosh, S. NF-κB in immunobiology. Cell Res. 2011, 21, 223–244. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/1224894 (accessed on 31 July 2020).
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/720552 (accessed on 31 July 2020).
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/1347034 (accessed on 31 July 2020).
- Sanaei, M.; Kavoosi, F. Histone Deacetylases and Histone Deacetylase Inhibitors: Molecular Mechanisms of Action in Various Cancers. Adv. Biomed. Res. 2019, 8, 63. [Google Scholar] [CrossRef]
- Siddiquiab, M.S.; François, M.; Fenech, M.F.; Leiferta, W.R. Persistent γH2AX: A promising molecular marker of DNA damage and aging. Mutat. Res. Rev. Mutat. Res. 2015, 766, 1–19. [Google Scholar] [CrossRef]
- Akerfelt, M.; Morimoto, R.I.; Sistonen, L. Heat shock factors: Integrators of cell stress, development and lifespan. Nat. Rev. Mol. Cell Biol. 2010, 11, 545–555. [Google Scholar] [CrossRef]
- Jin, G.; Sivaraman, A.; Lee, K. Development of taladegib as a sonic hedgehog signaling pathway inhibitor. Arch. Pharm. Res. 2017, 40, 1390–1393. [Google Scholar] [CrossRef] [PubMed]
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/1347032 (accessed on 31 July 2020).
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/720637 (accessed on 31 July 2020).
- Oakes, S.A.; Papa, F.R. The role of endoplasmic reticulum stress in human pathology. Annu. Rev. Pathol. 2015, 10, 173–194. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Available online: https://pubchem.ncbi.nlm.nih.gov/bioassay/720516 (accessed on 31 July 2020).
- Available online: https://www.rdkit.org/docs/index.html (accessed on 31 July 2020).
- Riniker, S.; Landrum, G.A. Better Informed Distance Geometry: Using What We Know To Improve Conformation Generation. J. Chem. Inf. Model. 2015, 55, 2562–2574. [Google Scholar] [CrossRef]
- Tosco, P.; Stiefl, N.; Landrum, G. Bringing the MMFF force field to the RDKit: Implementation and validation. J. Cheminformatics 2014, 6, 37. [Google Scholar] [CrossRef]
- Moriwaki, H.; Tian, Y.S.; Kawashita, N.; Takagi, T. Mordred: A molecular descriptor calculator. J. Cheminformatics 2018, 10, 4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Available online: https://mordred-descriptor.github.io/documentation/master/index.html (accessed on 31 July 2020).
- Matthews, B.W. Comparison of the predicted and observed secondary structure of T4 phage lysozyme. Biochim. Et Biophys. Acta 1975, 405, 442–451. [Google Scholar] [CrossRef]
- Fawcett, T. An Introduction to ROC Analysis. Pattern Recognit. Lett. 2006, 27, 861–874. [Google Scholar] [CrossRef]
- Fluss, R.; Faraggi, D.; Reiser, B. Estimation of the Youden Index and its Associated Cutoff Point. Biom. J. 2005, 47, 458–472. [Google Scholar] [CrossRef] [PubMed] [Green Version]






| No. | AID | Abbreviation | Criteria 40 | Criteria 1 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AUC | SE | SP | ACC | BAC | MCC | AUC | SE | SP | ACC | BAC | MCC | |||
| 1 | 720516 | ATAD5_ind | 0.840 | 0.750 | 0.843 | 0.840 | 0.796 | 0.272 | 0.845 | 0.744 | 0.847 | 0.839 | 0.795 | 0.395 |
| 2 | 720552 | p53_ago | 0.899 | 0.824 | 0.830 | 0.830 | 0.827 | 0.356 | 0.845 | 0.804 | 0.793 | 0.794 | 0.799 | 0.458 |
| 3 | 720637 | MMP_disr | 0.919 | 0.845 | 0.846 | 0.846 | 0.845 | 0.501 | 0.795 | 0.698 | 0.788 | 0.758 | 0.743 | 0.475 |
| 4 | 720719 | GR_ago | 0.783 | 0.600 | 0.931 | 0.923 | 0.766 | 0.300 | 0.841 | 0.754 | 0.807 | 0.800 | 0.780 | 0.416 |
| 5 | 720725 | GR_ant | 0.808 | 0.577 | 0.905 | 0.888 | 0.741 | 0.328 | 0.827 | 0.801 | 0.721 | 0.743 | 0.761 | 0.471 |
| 6 | 743053 | Arlbd_ago | 0.878 | 0.765 | 0.947 | 0.941 | 0.856 | 0.481 | 0.766 | 0.582 | 0.843 | 0.806 | 0.712 | 0.357 |
| 7 | 743054 | ARfull_ant | 0.774 | 0.750 | 0.681 | 0.683 | 0.716 | 0.169 | 0.833 | 0.841 | 0.700 | 0.734 | 0.770 | 0.468 |
| 8 | 743063 | Arlbd_ant | 0.844 | 0.786 | 0.791 | 0.790 | 0.788 | 0.338 | 0.833 | 0.805 | 0.724 | 0.745 | 0.765 | 0.469 |
| 9 | 743067 | TR_ant | 0.783 | 0.511 | 0.924 | 0.906 | 0.718 | 0.306 | 0.829 | 0.740 | 0.825 | 0.796 | 0.782 | 0.555 |
| 10 | 743077 | ERlbd_ago | 0.782 | 0.536 | 0.961 | 0.938 | 0.748 | 0.457 | 0.735 | 0.600 | 0.843 | 0.812 | 0.722 | 0.362 |
| 11 | 743078 | ERlbd_ant | 0.810 | 0.815 | 0.684 | 0.691 | 0.750 | 0.237 | 0.805 | 0.696 | 0.789 | 0.767 | 0.743 | 0.444 |
| 12 | 743091 | ERfull_ant | 0.826 | 0.872 | 0.699 | 0.705 | 0.785 | 0.235 | 0.862 | 0.730 | 0.870 | 0.842 | 0.800 | 0.555 |
| 13 | 743122 | AhR_ago | 0.888 | 0.713 | 0.907 | 0.887 | 0.810 | 0.513 | 0.749 | 0.728 | 0.695 | 0.702 | 0.711 | 0.359 |
| 14 | 743139 | Arom_ant | 0.801 | 0.892 | 0.598 | 0.608 | 0.745 | 0.186 | 0.807 | 0.825 | 0.661 | 0.704 | 0.743 | 0.429 |
| 15 | 743140 | PPARg_ago | 0.813 | 0.750 | 0.823 | 0.821 | 0.786 | 0.238 | 0.832 | 0.735 | 0.819 | 0.805 | 0.777 | 0.457 |
| 16 | 743199 | PPARg_ant | 0.829 | 0.786 | 0.798 | 0.798 | 0.792 | 0.290 | 0.810 | 0.824 | 0.645 | 0.682 | 0.734 | 0.383 |
| 17 | 743219 | ARE_ago | 0.785 | 0.794 | 0.652 | 0.672 | 0.723 | 0.317 | 0.795 | 0.770 | 0.715 | 0.733 | 0.742 | 0.461 |
| 18 | 743226 | PPARd_ant | 0.681 | 0.600 | 0.885 | 0.884 | 0.743 | 0.111 | 0.811 | 0.764 | 0.749 | 0.751 | 0.756 | 0.374 |
| 19 | 743227 | PPARd_ago | 0.812 | 0.615 | 0.954 | 0.949 | 0.785 | 0.296 | 0.796 | 0.705 | 0.790 | 0.780 | 0.747 | 0.356 |
| 20 | 743228 | HSR_act | 0.788 | 0.576 | 0.922 | 0.910 | 0.749 | 0.315 | 0.790 | 0.667 | 0.808 | 0.789 | 0.737 | 0.370 |
| 21 | 743239 | FXR_ago | 0.775 | 0.727 | 0.836 | 0.835 | 0.782 | 0.163 | 0.817 | 0.689 | 0.834 | 0.825 | 0.762 | 0.325 |
| 22 | 743240 | FXR_ant | 0.757 | 0.933 | 0.565 | 0.577 | 0.749 | 0.178 | 0.843 | 0.788 | 0.799 | 0.798 | 0.794 | 0.481 |
| 23 | 743241 | VDR_ago | N.D | N.D | N.D | N.D | N.D | N.D | 0.826 | 0.769 | 0.727 | 0.730 | 0.748 | 0.297 |
| 24 | 743242 | VDR_ant | 0.716 | 1.000 | 0.399 | 0.403 | 0.699 | 0.066 | 0.701 | 0.630 | 0.689 | 0.678 | 0.660 | 0.258 |
| 25 | 1159518 | NFkB_ago | 0.780 | 0.667 | 0.846 | 0.846 | 0.756 | 0.081 | 0.871 | 0.692 | 0.912 | 0.900 | 0.802 | 0.427 |
| 26 | 1159519 | ERsr_ago | 0.638 | 0.857 | 0.441 | 0.445 | 0.649 | 0.052 | 0.801 | 0.655 | 0.833 | 0.816 | 0.744 | 0.349 |
| 27 | 1159523 | ROR_ant | 0.828 | 0.789 | 0.764 | 0.766 | 0.777 | 0.323 | 0.695 | 0.523 | 0.819 | 0.703 | 0.671 | 0.359 |
| 28 | 1159528 | AP1_ago | 0.777 | 0.553 | 0.877 | 0.851 | 0.715 | 0.319 | 0.799 | 0.765 | 0.722 | 0.729 | 0.743 | 0.372 |
| 29 | 1159531 | RXR_ago | 0.532 | 0.235 | 0.964 | 0.951 | 0.600 | 0.135 | 0.725 | 0.527 | 0.841 | 0.756 | 0.684 | 0.374 |
| 30 | 1159555 | RAR_ant | 0.831 | 0.800 | 0.742 | 0.746 | 0.771 | 0.308 | 0.683 | 0.740 | 0.511 | 0.601 | 0.626 | 0.249 |
| 31 | 1224892 | CAR_ago | 0.889 | 0.826 | 0.808 | 0.810 | 0.817 | 0.455 | 0.847 | 0.684 | 0.889 | 0.845 | 0.787 | 0.556 |
| 32 | 1224893 | CAR_ant | 0.809 | 0.652 | 0.880 | 0.874 | 0.766 | 0.239 | 0.793 | 0.700 | 0.768 | 0.746 | 0.734 | 0.448 |
| 33 | 1224894 | HIF1_ago | 0.556 | 0.250 | 1.000 | 0.997 | 0.625 | 0.499 | 0.854 | 0.769 | 0.829 | 0.824 | 0.799 | 0.395 |
| 34 | 1224895 | TSHR_ago | 0.872 | 0.750 | 0.880 | 0.874 | 0.815 | 0.355 | 0.838 | 0.692 | 0.831 | 0.816 | 0.762 | 0.389 |
| 35 | 1224896 | H2AX_ago | 0.834 | 0.696 | 0.892 | 0.880 | 0.794 | 0.394 | 0.779 | 0.605 | 0.842 | 0.814 | 0.724 | 0.354 |
| 36 | 1259247 | Arfulls_ant | 0.856 | 0.857 | 0.733 | 0.747 | 0.795 | 0.401 | 0.824 | 0.788 | 0.767 | 0.774 | 0.778 | 0.534 |
| 37 | 1259248 | Erfulls_ant | 0.835 | 0.850 | 0.702 | 0.711 | 0.776 | 0.283 | 0.793 | 0.668 | 0.798 | 0.770 | 0.733 | 0.416 |
| 38 | 1259387 | ARant_ago | 0.852 | 0.727 | 0.946 | 0.939 | 0.837 | 0.460 | 0.712 | 0.494 | 0.872 | 0.841 | 0.683 | 0.275 |
| 39 | 1259388 | HDAC_ant | 0.897 | 0.783 | 0.888 | 0.883 | 0.835 | 0.407 | 0.868 | 0.768 | 0.879 | 0.871 | 0.824 | 0.447 |
| 40 | 1259390 | Shh_ago | 0.571 | 1.000 | 0.219 | 0.223 | 0.609 | 0.042 | 0.724 | 0.609 | 0.913 | 0.905 | 0.761 | 0.266 |
| 41 | 1259391 | ERaant_ago | 0.934 | 0.850 | 0.959 | 0.956 | 0.904 | 0.493 | 0.782 | 0.551 | 0.898 | 0.880 | 0.725 | 0.299 |
| 42 | 1259392 | Shh_ant | 0.829 | 0.809 | 0.718 | 0.731 | 0.764 | 0.379 | 0.758 | 0.642 | 0.745 | 0.705 | 0.693 | 0.383 |
| 43 | 1259393 | TSHR_agoant | 0.834 | 0.750 | 0.875 | 0.874 | 0.812 | 0.120 | 0.669 | 0.727 | 0.681 | 0.682 | 0.704 | 0.093 |
| 44 | 1259394 | ERb_ago | 0.980 | 0.923 | 0.973 | 0.972 | 0.948 | 0.531 | 0.729 | 0.444 | 0.937 | 0.900 | 0.691 | 0.348 |
| 45 | 1259395 | TSHR_ant | 0.865 | 0.933 | 0.715 | 0.721 | 0.824 | 0.244 | 0.850 | 0.800 | 0.807 | 0.807 | 0.804 | 0.381 |
| 46 | 1259396 | Erb_ant | 0.825 | 0.677 | 0.863 | 0.851 | 0.770 | 0.352 | 0.798 | 0.743 | 0.763 | 0.758 | 0.753 | 0.462 |
| 47 | 1259401 | ERRPGC_ant | 0.843 | 0.698 | 0.843 | 0.837 | 0.770 | 0.290 | 0.751 | 0.595 | 0.793 | 0.723 | 0.694 | 0.390 |
| 48 | 1259402 | ERRPGC_ago | 0.840 | 0.650 | 0.937 | 0.925 | 0.794 | 0.415 | 0.805 | 0.734 | 0.777 | 0.768 | 0.756 | 0.444 |
| 49 | 1259403 | ERR_ant | 0.812 | 0.653 | 0.856 | 0.835 | 0.755 | 0.392 | 0.819 | 0.696 | 0.826 | 0.786 | 0.761 | 0.510 |
| 50 | 1259404 | ERR_ago | 0.884 | 0.880 | 0.814 | 0.816 | 0.847 | 0.274 | 0.803 | 0.680 | 0.820 | 0.777 | 0.750 | 0.491 |
| 51 | 1347030 | TRHR_ago | 0.748 | 0.833 | 0.637 | 0.638 | 0.735 | 0.077 | 0.751 | 0.593 | 0.853 | 0.846 | 0.723 | 0.201 |
| 52 | 1347031 | PR_ant | 0.892 | 0.880 | 0.794 | 0.804 | 0.837 | 0.473 | 0.831 | 0.757 | 0.821 | 0.802 | 0.789 | 0.550 |
| 53 | 1347032 | TGFb_ant | 0.809 | 0.750 | 0.765 | 0.764 | 0.757 | 0.273 | 0.860 | 0.780 | 0.824 | 0.817 | 0.802 | 0.493 |
| 54 | 1347033 | PXR_ago | 0.851 | 0.759 | 0.817 | 0.805 | 0.788 | 0.517 | 0.838 | 0.745 | 0.817 | 0.790 | 0.781 | 0.556 |
| 55 | 1347034 | CaspH_ind | 0.870 | 0.791 | 0.852 | 0.849 | 0.821 | 0.348 | 0.858 | 0.773 | 0.856 | 0.848 | 0.814 | 0.452 |
| 56 | 1347035 | TGFb_ago | 0.968 | 1.000 | 0.938 | 0.938 | 0.969 | 0.174 | 0.900 | 0.818 | 0.937 | 0.936 | 0.878 | 0.311 |
| 57 | 1347036 | PR_ago | 0.943 | 0.833 | 0.989 | 0.986 | 0.911 | 0.701 | 0.799 | 0.537 | 0.986 | 0.967 | 0.761 | 0.564 |
| 58 | 1347037 | CaspC_ind | 0.884 | 0.850 | 0.785 | 0.786 | 0.817 | 0.216 | 0.863 | 0.771 | 0.882 | 0.878 | 0.827 | 0.351 |
| 59 | 1347038 | TRHR_ant | 0.822 | 0.700 | 0.841 | 0.840 | 0.771 | 0.148 | 0.828 | 0.870 | 0.701 | 0.709 | 0.785 | 0.260 |
| Metrics | Criteria 40 | Criteria 1 |
|---|---|---|
| AUC | 0.817 ± 0.088 | 0.802 ± 0.051 |
| SE | 0.750 ± 0.151 | 0.705 ± 0.094 |
| SP | 0.809 ± 0.149 | 0.801 ± 0.082 |
| ACC | 0.807 ± 0.144 | 0.788 ± 0.069 |
| BAC | 0.780 ± 0.069 | 0.753 ± 0.045 |
| MCC | 0.307 ± 0.141 | 0.402 ± 0.096 |
| No. | AID | Molecular Initiating Events | Activity Type | Abbreviation |
|---|---|---|---|---|
| 1 | 720516 | ATAD5 | genotoxic inducer | ATAD5_ind |
| 2 | 720552 | p53 | agonist | p53_ago |
| 3 | 720637 | mitochondrial membrane potential | disruptor | MMP_disr |
| 4 | 720719 | glucocorticoid receptor | agonist | GR_ago |
| 5 | 720725 | glucocorticoid receptor | antagonist | GR_ant |
| 6 | 743053 | androgen receptor lbd | agonist | Arlbd_ago |
| 7 | 743054 | androgen receptor full | antagonist | ARfull_ant |
| 8 | 743063 | androgen receptor lbd | antagonist | Arlbd_ant |
| 9 | 743067 | thyroid receptor | antagonist | TR_ant |
| 10 | 743077 | estrogen receptor alpha lbd | agonist | ERlbd_ago |
| 11 | 743078 | estrogen receptor alpha lbd | antagonist | ERlbd_ant |
| 12 | 743091 | estrogen receptor alpha full | antagonist | ERfull_ant |
| 13 | 743122 | aryl hydrocarbon receptor | agonist | AhR_ago |
| 14 | 743139 | aromatase | antagonist | Arom_ant |
| 15 | 743140 | peroxisome proliferator-activated receptor gamma | agonist | PPARg_ago |
| 16 | 743199 | peroxisome proliferator-activated receptor gamma | antagonist | PPARg_ant |
| 17 | 743219 | antioxidant response element | agonist | ARE_ago |
| 18 | 743226 | peroxisome proliferator-activated receptor delta | antagonist | PPARd_ant |
| 19 | 743227 | peroxisome proliferator-activated receptor delta | agonist | PPARd_ago |
| 20 | 743228 | heat shock response | activator | HSR_act |
| 21 | 743239 | farnesoid-X-receptor | agonist | FXR_ago |
| 22 | 743240 | farnesoid-X-receptor | antagonist | FXR_ant |
| 23 | 743241 | vitamin D receptor | agonist | VDR_ago |
| 24 | 743242 | vitamin D receptor | antagonist | VDR_ant |
| 25 | 1159518 | NFkB | agonist | NFkB_ago |
| 26 | 1159519 | endoplasmic reticulum stress response | agonist | ERsr_ago |
| 27 | 1159523 | retinoid-related orphan receptor gamma | antagonist | ROR_ant |
| 28 | 1159528 | activator protein-1 | agonist | AP1_ago |
| 29 | 1159531 | retinoid X receptor-alpha | agonist | RXR_ago |
| 30 | 1159555 | retinoic acid receptor | antagonist | RAR_ant |
| 31 | 1224892 | constitutive androstane receptor | agonist | CAR_ago |
| 32 | 1224893 | constitutive androstane receptor | antagonist | CAR_ant |
| 33 | 1224894 | hypoxia | agonist | HIF1_ago |
| 34 | 1224895 | thyroid stimulating hormone receptor | agonist | TSHR_ago |
| 35 | 1224896 | histone variant H2AX | agonist | H2AX_ago |
| 36 | 1259247 | androgen receptor with stimulator | antagonist | Arfulls_ant |
| 37 | 1259248 | estrogen receptor alpha with stimulator | antagonist | Erfulls_ant |
| 38 | 1259387 | androgen receptor with antagonist | agonist | ARant_ago |
| 39 | 1259388 | histone deacetylase | antagonist | HDAC_ant |
| 40 | 1259390 | sonic hedgehog signaling | agonist | Shh_ago |
| 41 | 1259391 | estrogen receptor alpha with antagonist | agonist | ERaant_ago |
| 42 | 1259392 | sonic hedgehog signaling | antagonist | Shh_ant |
| 43 | 1259393 | thyroid stimulating hormone receptor | agonist antagonist | TSHR_agoant |
| 44 | 1259394 | estrogen receptor beta | agonist | ERb_ago |
| 45 | 1259395 | thyroid stimulating hormone receptor | antagonist | TSHR_ant |
| 46 | 1259396 | estrogen receptor beta | antagonist | Erb_ant |
| 47 | 1259401 | estrogen related receptor with PGC | antagonist | ERRPGC_ant |
| 48 | 1259402 | estrogen related receptor with PGC | agonist | ERRPGC_ago |
| 49 | 1259403 | estrogen related receptor | antagonist | ERR_ant |
| 50 | 1259404 | estrogen related receptor | agonist | ERR_ago |
| 51 | 1347030 | thyrotropin releasing hormone receptor | agonist | TRHR_ago |
| 52 | 1347031 | progesterone receptor | antagonist | PR_ant |
| 53 | 1347032 | transforming growth factor beta | antagonist | TGFb_ant |
| 54 | 1347033 | human pregnane X receptor | agonist | PXR_ago |
| 55 | 1347034 | caspase-3/7 in HepG2 | inducer | CaspH_ind |
| 56 | 1347035 | transforming growth factor beta | agonist | TGFb_ago |
| 57 | 1347036 | progesterone receptor | agonist | PR_ago |
| 58 | 1347037 | caspase-3/7 in CHO-K1 | inducer | CaspC_ind |
| 59 | 1347038 | thyrotropin releasing hormone receptor | antagonist | TRHR_ant |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kurosaki, K.; Wu, R.; Uesawa, Y. A Toxicity Prediction Tool for Potential Agonist/Antagonist Activities in Molecular Initiating Events Based on Chemical Structures. Int. J. Mol. Sci. 2020, 21, 7853. https://doi.org/10.3390/ijms21217853
Kurosaki K, Wu R, Uesawa Y. A Toxicity Prediction Tool for Potential Agonist/Antagonist Activities in Molecular Initiating Events Based on Chemical Structures. International Journal of Molecular Sciences. 2020; 21(21):7853. https://doi.org/10.3390/ijms21217853
Chicago/Turabian StyleKurosaki, Kota, Raymond Wu, and Yoshihiro Uesawa. 2020. "A Toxicity Prediction Tool for Potential Agonist/Antagonist Activities in Molecular Initiating Events Based on Chemical Structures" International Journal of Molecular Sciences 21, no. 21: 7853. https://doi.org/10.3390/ijms21217853
APA StyleKurosaki, K., Wu, R., & Uesawa, Y. (2020). A Toxicity Prediction Tool for Potential Agonist/Antagonist Activities in Molecular Initiating Events Based on Chemical Structures. International Journal of Molecular Sciences, 21(21), 7853. https://doi.org/10.3390/ijms21217853







