An Overview of ADAM9: Structure, Activation, and Regulation in Human Diseases
Abstract
1. Introduction
2. Structure and Molecular Function of ADAM9
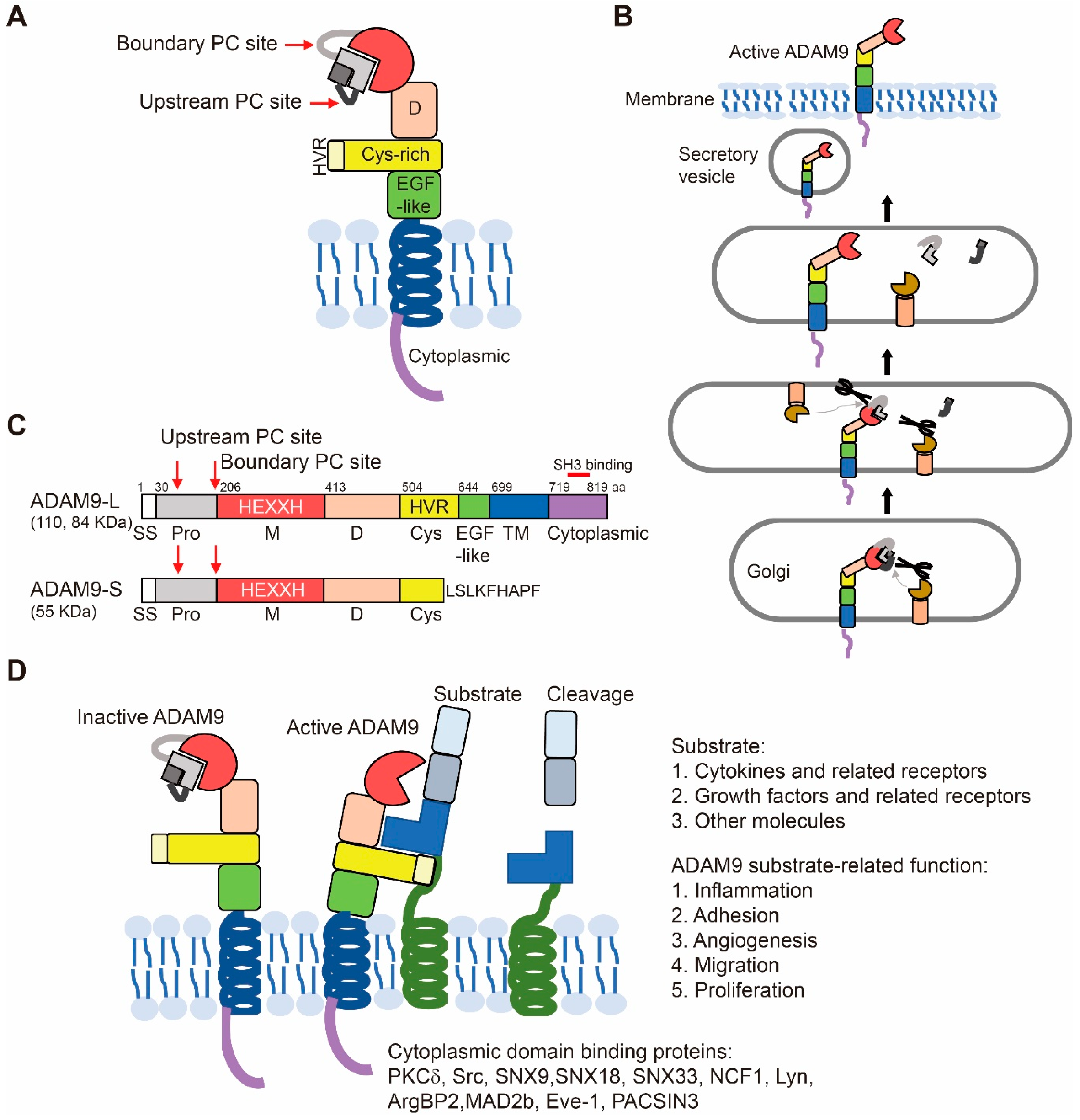
2.1. ADAM9 Structure
2.1.1. Pro-Domain and Its Cleavage for Protease Activation
2.1.2. Metalloproteases, Disintegrin, Cysteine-Rich, EGF-Like, and Transmembrane Domains
2.1.3. Cytoplasmic Domain
2.1.4. Two Isoforms of ADAM9
2.2. Substrate Recognition and Cleavage
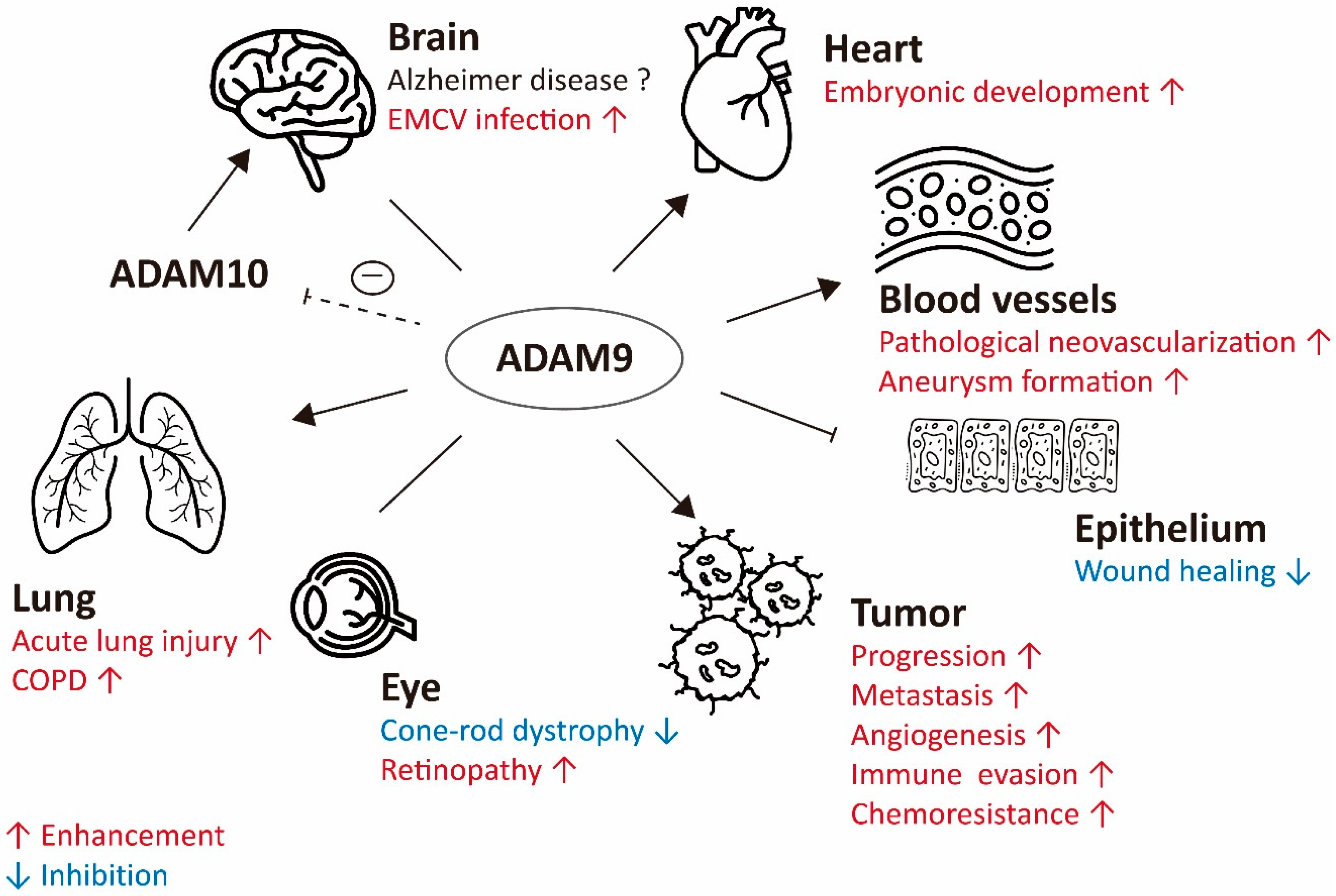
3. Pathological Roles of ADAM9 in Neurodegenerative and Retinal Diseases
3.1. ADAM9-Mediated Neurodegenerative Diseases
3.2. ADAM9-Mediated Retinal Diseases
3.3. ADAM9-Mediated Vascular Disease
4. ADAM9 in Infectious and Inflammatory Diseases
4.1. Infection and Tissue Damage
4.2. Lung Injury and COPD
4.3. Chronic Wounds
5. Role of ADAM9 in Cancers
5.1. Lung Cancer
5.2. Prostate Cancer
5.3. Liver Cancer
5.4. Breast Cancer
5.5. Pancreatic Cancer
5.6. Glioma
5.7. Other Cancers
6. ADAM9 in the Tumor Microenvironment
6.1. Neutrophil
6.2. Monocytes/Macrophages
6.3. Platelet
7. Therapeutic Potential of Manipulating ADAM9 Expression in Cancer Treatment
7.1. MicroRNA and Proteins Targeting ADAM9
7.2. MMP Inhibitors
7.3. Natural Flavonoids
7.4. Sorafenib and Regorafenib
8. Concluding Remarks
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ADAM9 | A disintegrin and a metalloprotease 9 |
| ArgBP2 | Sorbin and SH3 domain-containing protein 2 |
| AIPC | Androgen-independent prostate cancer |
| APP | Amyloid-precursor-protein |
| CDCP1 | CUB domain containing protein 1 |
| COPD | Chronic obstructive pulmonary disease |
| CRD | Cone-rod dystrophy |
| ECM | Extracellular matrix |
| EGF | Epidermal growth factor |
| EMCV | Encephalomyocarditis virus |
| EMT | Epithelial-mesenchymal transition |
| EphB4 | Ephrin type-B receptor 4 |
| GBM | Glioblastoma |
| HB-EGF | Heparin-binding EGF-like growth factor |
| HCC | Hepatocellular carcinoma |
| HVR | Hyper-variable region |
| MDC9 | Metalloprotease/disintegrin/cysteine-rich protein 9 |
| MGCs | Multinucleated giant cells |
| MICA | MHC Class I polypeptide-related sequence A |
| MiRNA | microRNA |
| MMP | Matrix metalloproteinase |
| Naa10p | N-α-Acetyltransferase 10 protein |
| PC | Proprotein convertase |
| PMNs | Polymorphonuclear neutrophils |
| PrPc | Cellular prion protein |
| SH3 | Src-homology 3 |
| SNX | Sorting nexin |
| TNBC | Triple-negative breast cancer |
| TNC | Tenascin-C |
| TNF-α | Tumor necrosis factor-α |
| tPA | Tissue-type plasminogen activator |
| VE-cadherin | Vascular endothelial cadherin |
References
- Hsia, H.E.; Tüshaus, J.; Brummer, T.; Zheng, Y.; Scilabra, S.D.; Lichtenthaler, S.F. Functions of ‘A disintegrin and metalloproteases (ADAMs)’ in the mammalian nervous system. Cell. Mol. Life Sci. 2019, 76, 3055–3081. [Google Scholar] [CrossRef] [PubMed]
- Seals, D.F.; Courtneidge, S.A. The ADAMs family of metalloproteases: Multidomain proteins with multiple functions. Genes Dev. 2003, 17, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Weber, S.; Saftig, P. Ectodomain shedding and ADAMs in development. Development 2012, 139, 3693–3709. [Google Scholar] [CrossRef] [PubMed]
- Giebeler, N.; Zigrino, P. A Disintegrin and Metalloprotease (ADAM): Historical Overview of Their Functions. Toxins 2016, 8, 122. [Google Scholar] [CrossRef]
- Janes, P.W.; Saha, N.; Barton, W.A.; Kolev, M.V.; Wimmer-Kleikamp, S.H.; Nievergall, E.; Blobel, C.P.; Himanen, J.P.; Lackmann, M.; Nikolov, D.B. Adam meets Eph: An ADAM substrate recognition module acts as a molecular switch for ephrin cleavage in trans. Cell 2005, 123, 291–304. [Google Scholar] [CrossRef]
- Matthews, A.L.; Noy, P.J.; Reyat, J.S.; Tomlinson, M.G. Regulation of A disintegrin and metalloproteinase (ADAM) family sheddases ADAM10 and ADAM17: The emerging role of tetraspanins and rhomboids. Platelets 2017, 28, 333–341. [Google Scholar] [CrossRef]
- Edwards, D.R.; Handsley, M.M.; Pennington, C.J. The ADAM metalloproteinases. Mol. Asp. Med. 2008, 29, 258–289. [Google Scholar] [CrossRef]
- Lambrecht, B.N.; Vanderkerken, M.; Hammad, H. The emerging role of ADAM metalloproteinases in immunity. Nat. Rev. Immunol. 2018, 18, 745–758. [Google Scholar] [CrossRef]
- English, W.R.; Siviter, R.J.; Hansen, M.; Murphy, G. ADAM9 is present at endothelial cell-cell junctions and regulates monocyte-endothelial transmigration. Biochem. Biophys. Res. Commun. 2017, 493, 1057–1062. [Google Scholar] [CrossRef]
- Weskamp, G.; Krätzschmar, J.; Reid, M.S.; Blobel, C.P. MDC9, a widely expressed cellular disintegrin containing cytoplasmic SH3 ligand domains. J. Cell Biol. 1996, 132, 717–726. [Google Scholar] [CrossRef]
- Rinchai, D.; Kewcharoenwong, C.; Kessler, B.; Lertmemongkolchai, G.; Chaussabel, D. Increased abundance of ADAM9 transcripts in the blood is associated with tissue damage. F1000Res 2015, 4, 89. [Google Scholar] [CrossRef] [PubMed]
- Namba, K.; Nishio, M.; Mori, K.; Miyamoto, N.; Tsurudome, M.; Ito, M.; Kawano, M.; Uchida, A.; Ito, Y. Involvement of ADAM9 in multinucleated giant cell formation of blood monocytes. Cell. Immunol. 2001, 213, 104–113. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Polverino, F.; Rojas-Quintero, J.; Zhang, D.; Sánchez, J.; Yambayev, I.; Lindqvist, E.; Virtala, R.; Djukanovic, R.; Davies, D.E.; et al. A Disintegrin and A Metalloproteinase-9 (ADAM9): A Novel Proteinase Culprit with Multifarious Contributions to COPD. Am. J. Respir. Crit. Care Med. 2018, 198, 1500–1518. [Google Scholar] [CrossRef] [PubMed]
- Roychaudhuri, R.; Hergrueter, A.H.; Polverino, F.; Laucho-Contreras, M.E.; Gupta, K.; Borregaard, N.; Owen, C.A. ADAM9 is a novel product of polymorphonuclear neutrophils: Regulation of expression and contributions to extracellular matrix protein degradation during acute lung injury. J. Immunol. 2014, 193, 2469–2482. [Google Scholar] [CrossRef] [PubMed]
- Zigrino, P.; Steiger, J.; Fox, J.W.; Löffek, S.; Schild, A.; Nischt, R.; Mauch, C. Role of ADAM-9 disintegrin-cysteine-rich domains in human keratinocyte migration. J. Biol. Chem. 2007, 282, 30785–30793. [Google Scholar] [CrossRef]
- Abety, A.N.; Fox, J.W.; Schönefuß, A.; Zamek, J.; Landsberg, J.; Krieg, T.; Blobel, C.; Mauch, C.; Zigrino, P. Stromal fibroblast-specific expression of ADAM-9 modulates proliferation and apoptosis in melanoma cells in vitro and in vivo. J. Investig. Dermatol. 2012, 132, 2451–2458. [Google Scholar] [CrossRef]
- Dreymueller, D.; Uhlig, S.; Ludwig, A. ADAM-family metalloproteinases in lung inflammation: Potential therapeutic targets. Am. J. Physiol. Lung Cell. Mol. Physiol. 2015, 308, L325–L343. [Google Scholar] [CrossRef]
- Fu, Q.; Cheng, J.; Zhang, J.; Zhang, Y.; Chen, X.; Luo, S.; Xie, J. miR-20b reduces 5-FU resistance by suppressing the ADAM9/EGFR signaling pathway in colon cancer. Oncol. Rep. 2017, 37, 123–130. [Google Scholar] [CrossRef]
- Mahimkar, R.M.; Visaya, O.; Pollock, A.S.; Lovett, D.H. The disintegrin domain of ADAM9: A ligand for multiple beta1 renal integrins. Biochem. J. 2005, 385, 461–468. [Google Scholar] [CrossRef]
- Shen, G.; Sun, Q.; Yao, Y.; Li, S.; Liu, G.; Yuan, C.; Li, H.; Xu, Y.; Wang, H. Role of ADAM9 and miR-126 in the development of abdominal aortic aneurysm. Atherosclerosis 2020, 297, 47–54. [Google Scholar] [CrossRef]
- Zhang, P.; Shen, M.; Fernandez-Patron, C.; Kassiri, Z. ADAMs family and relatives in cardiovascular physiology and pathology. J. Mol. Cell. Cardiol. 2016, 93, 186–199. [Google Scholar] [CrossRef] [PubMed]
- Fadl, N.N.; Ahmed, H.H.; Booles, H.F.; Sayed, A.H. Serrapeptase and nattokinase intervention for relieving Alzheimer’s disease pathophysiology in rat model. Hum. Exp. Toxicol. 2013, 32, 721–735. [Google Scholar] [CrossRef]
- Cho, C. Testicular and epididymal ADAMs: Expression and function during fertilization. Nat. Rev. Urol. 2012, 9, 550–560. [Google Scholar] [CrossRef] [PubMed]
- Grützmann, R.; Lüttges, J.; Sipos, B.; Ammerpohl, O.; Dobrowolski, F.; Alldinger, I.; Kersting, S.; Ockert, D.; Koch, R.; Kalthoff, H.; et al. ADAM9 expression in pancreatic cancer is associated with tumour type and is a prognostic factor in ductal adenocarcinoma. Br. J. Cancer 2004, 90, 1053–1058. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.Y.; Chen, H.J.; Huang, C.C.; Lai, L.C.; Lu, T.P.; Tseng, G.C.; Kuo, T.T.; Kuok, Q.Y.; Hsu, J.L.; Sung, S.Y.; et al. ADAM9 promotes lung cancer metastases to brain by a plasminogen activator-based pathway. Cancer Res. 2014, 74, 5229–5243. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.Y.; Cho, C.F.; Bai, S.T.; Liu, J.P.; Kuo, T.T.; Wang, L.J.; Lin, Y.S.; Lin, C.C.; Lai, L.C.; Lu, T.P.; et al. ADAM9 promotes lung cancer progression through vascular remodeling by VEGFA, ANGPT2, and PLAT. Sci. Rep. 2017, 7, 15108. [Google Scholar] [CrossRef]
- Schouten, L.R.; Helmerhorst, H.J.; Wagenaar, G.T.; Haltenhof, T.; Lutter, R.; Roelofs, J.J.; van Woensel, J.B.; van Kaam, A.H.; Bos, A.P.; Schultz, M.J.; et al. Age-Dependent Changes in the Pulmonary Renin-Angiotensin System Are Associated With Severity of Lung Injury in a Model of Acute Lung Injury in Rats. Crit. Care Med. 2016, 44, e1226–e1235. [Google Scholar] [CrossRef]
- Mauch, C.; Zamek, J.; Abety, A.N.; Grimberg, G.; Fox, J.W.; Zigrino, P. Accelerated wound repair in ADAM-9 knockout animals. J. Investig. Dermatol. 2010, 130, 2120–2130. [Google Scholar] [CrossRef]
- Mehta, V.; Fields, L.; Evans, I.M.; Yamaji, M.; Pellet-Many, C.; Jones, T.; Mahmoud, M.; Zachary, I. VEGF (Vascular Endothelial Growth Factor) Induces NRP1 (Neuropilin-1) Cleavage via ADAMs (a Disintegrin and Metalloproteinase) 9 and 10 to Generate Novel Carboxy-Terminal NRP1 Fragments That Regulate Angiogenic Signaling. Arter. Thromb. Vasc. Biol. 2018, 38, 1845–1858. [Google Scholar] [CrossRef]
- Moss, M.L.; Powell, G.; Miller, M.A.; Edwards, L.; Qi, B.; Sang, Q.X.; De Strooper, B.; Tesseur, I.; Lichtenthaler, S.F.; Taverna, M.; et al. ADAM9 inhibition increases membrane activity of ADAM10 and controls α-secretase processing of amyloid precursor protein. J. Biol. Chem. 2011, 286, 40443–40451. [Google Scholar] [CrossRef]
- Asai, M.; Hattori, C.; Szabó, B.; Sasagawa, N.; Maruyama, K.; Tanuma, S.; Ishiura, S. Putative function of ADAM9, ADAM10, and ADAM17 as APP alpha-secretase. Biochem. Biophys. Res. Commun. 2003, 301, 231–235. [Google Scholar] [CrossRef]
- Horiuchi, K.; Zhou, H.M.; Kelly, K.; Manova, K.; Blobel, C.P. Evaluation of the contributions of ADAMs 9, 12, 15, 17, and 19 to heart development and ectodomain shedding of neuregulins beta1 and beta2. Dev. Biol. 2005, 283, 459–471. [Google Scholar] [CrossRef] [PubMed]
- Sammel, M.; Peters, F.; Lokau, J.; Scharfenberg, F.; Werny, L.; Linder, S.; Garbers, C.; Rose-John, S.; Becker-Pauly, C. Differences in Shedding of the Interleukin-11 Receptor by the Proteases ADAM9, ADAM10, ADAM17, Meprin α, Meprin β and MT1-MMP. Int. J. Mol. Sci. 2019, 20, 3677. [Google Scholar] [CrossRef] [PubMed]
- Weskamp, G.; Cai, H.; Brodie, T.A.; Higashyama, S.; Manova, K.; Ludwig, T.; Blobel, C.P. Mice lacking the metalloprotease-disintegrin MDC9 (ADAM9) have no evident major abnormalities during development or adult life. Mol. Cell. Biol. 2002, 22, 1537–1544. [Google Scholar] [CrossRef] [PubMed]
- Goldstein, O.; Mezey, J.G.; Boyko, A.R.; Gao, C.; Wang, W.; Bustamante, C.D.; Anguish, L.J.; Jordan, J.A.; Pearce-Kelling, S.E.; Aguirre, G.D.; et al. An ADAM9 mutation in canine cone-rod dystrophy 3 establishes homology with human cone-rod dystrophy 9. Mol. Vis. 2010, 16, 1549–1569. [Google Scholar] [PubMed]
- Parry, D.A.; Toomes, C.; Bida, L.; Danciger, M.; Towns, K.V.; McKibbin, M.; Jacobson, S.G.; Logan, C.V.; Ali, M.; Bond, J.; et al. Loss of the metalloprotease ADAM9 leads to cone-rod dystrophy in humans and retinal degeneration in mice. Am. J. Hum. Genet. 2009, 84, 683–691. [Google Scholar] [CrossRef]
- Guaiquil, V.; Swendeman, S.; Yoshida, T.; Chavala, S.; Campochiaro, P.A.; Blobel, C.P. ADAM9 is involved in pathological retinal neovascularization. Mol. Cell. Biol. 2009, 29, 2694–2703. [Google Scholar] [CrossRef]
- Moss, M.L.; Bomar, M.; Liu, Q.; Sage, H.; Dempsey, P.; Lenhart, P.M.; Gillispie, P.A.; Stoeck, A.; Wildeboer, D.; Bartsch, J.W. The ADAM10 prodomain is a specific inhibitor of ADAM10 proteolytic activity and inhibits cellular shedding events. J. Biol. Chem. 2007, 282, 35712–35721. [Google Scholar] [CrossRef]
- Gonzales, P.E.; Solomon, A.; Miller, A.B.; Leesnitzer, M.A.; Sagi, I.; Milla, M.E. Inhibition of the tumor necrosis factor-α-converting enzyme by its pro domain. J. Biol. Chem. 2004, 279, 31638–31645. [Google Scholar] [CrossRef]
- Wong, E.; Maretzky, T.; Peleg, Y.; Blobel, C.P.; Sagi, I. The Functional Maturation of A Disintegrin and Metalloproteinase (ADAM) 9, 10, and 17 Requires Processing at a Newly Identified Proprotein Convertase (PC) Cleavage Site. J. Biol. Chem. 2015, 290, 12135–12146. [Google Scholar] [CrossRef]
- Wichert, R.; Scharfenberg, F.; Colmorgen, C.; Koudelka, T.; Schwarz, J.; Wetzel, S.; Potempa, B.; Potempa, J.; Bartsch, J.W.; Sagi, I.; et al. Meprin β induces activities of A disintegrin and metalloproteinases 9, 10, and 17 by specific prodomain cleavage. FASEB J. 2019, 33, 11925–11940. [Google Scholar] [CrossRef] [PubMed]
- Smith, K.M.; Gaultier, A.; Cousin, H.; Alfandari, D.; White, J.M.; DeSimone, D.W. The cysteine-rich domain regulates ADAM protease function in vivo. J. Cell Biol. 2002, 159, 893–902. [Google Scholar] [CrossRef] [PubMed]
- Gaultier, A.; Cousin, H.; Darribère, T.; Alfandari, D. ADAM13 disintegrin and cysteine-rich domains bind to the second heparin-binding domain of fibronectin. J. Biol. Chem. 2002, 277, 23336–23344. [Google Scholar] [CrossRef] [PubMed]
- Kleino, I.; Järviluoma, A.; Hepojoki, J.; Huovila, A.P.; Saksela, K. Preferred SH3 domain partners of ADAM metalloproteases include shared and ADAM-specific SH3 interactions. PLoS ONE 2015, 10, e0121301. [Google Scholar] [CrossRef] [PubMed]
- Soulet, F.; Yarar, D.; Leonard, M.; Schmid, S.L. SNX9 regulates dynamin assembly and is required for efficient clathrin-mediated endocytosis. Mol. Biol. Cell 2005, 16, 2058–2067. [Google Scholar] [CrossRef] [PubMed]
- Mygind, K.J.; Störiko, T.; Freiberg, M.L.; Samsøe-Petersen, J.; Schwarz, J.; Andersen, O.M.; Kveiborg, M. Sorting nexin 9 (SNX9) regulates levels of the transmembrane ADAM9 at the cell surface. J. Biol. Chem. 2018, 293, 8077–8088. [Google Scholar] [CrossRef]
- Izumi, Y.; Hirata, M.; Hasuwa, H.; Iwamoto, R.; Umata, T.; Miyado, K.; Tamai, Y.; Kurisaki, T.; Sehara-Fujisawa, A.; Ohno, S.; et al. A metalloprotease-disintegrin, MDC9/meltrin-gamma/ADAM9 and PKCdelta are involved in TPA-induced ectodomain shedding of membrane-anchored heparin-binding EGF-like growth factor. EMBO J. 1998, 17, 7260–7272. [Google Scholar] [CrossRef]
- Fry, J.L.; Toker, A. Secreted and membrane-bound isoforms of protease ADAM9 have opposing effects on breast cancer cell migration. Cancer Res. 2010, 70, 8187–8198. [Google Scholar] [CrossRef] [PubMed]
- Mazzocca, A.; Coppari, R.; De Franco, R.; Cho, J.Y.; Libermann, T.A.; Pinzani, M.; Toker, A. A secreted form of ADAM9 promotes carcinoma invasion through tumor-stromal interactions. Cancer Res. 2005, 65, 4728–4738. [Google Scholar] [CrossRef]
- Seegar, T.C.; Killingsworth, L.B.; Saha, N.; Meyer, P.A.; Patra, D.; Zimmerman, B.; Janes, P.W.; Rubinstein, E.; Nikolov, D.B.; Skiniotis, G.J.C. Structural basis for regulated proteolysis by the α-secretase ADAM10. Cell 2017, 171, 1638–1648.e1637. [Google Scholar] [CrossRef]
- Grötzinger, J.; Lorenzen, I.; Düsterhöft, S. Molecular insights into the multilayered regulation of ADAM17: The role of the extracellular region. Biochim. Biophys. Acta (BBA) Mol. Cell Res. 2017, 1864, 2088–2095. [Google Scholar] [CrossRef] [PubMed]
- Takeda, S.; Igarashi, T.; Mori, H.; Araki, S. Crystal structures of VAP1 reveal ADAMs’ MDC domain architecture and its unique C-shaped scaffold. EMBO J. 2006, 25, 2388–2396. [Google Scholar] [CrossRef]
- Igarashi, T.; Araki, S.; Mori, H.; Takeda, S. Crystal structures of catrocollastatin/VAP2B reveal a dynamic, modular architecture of ADAM/adamalysin/reprolysin family proteins. FEBS Lett. 2007, 581, 2416–2422. [Google Scholar] [CrossRef] [PubMed]
- Maretzky, T.; Swendeman, S.; Mogollon, E.; Weskamp, G.; Sahin, U.; Reiss, K.; Blobel, C.P. Characterization of the catalytic properties of the membrane-anchored metalloproteinase ADAM9 in cell-based assays. Biochem. J. 2017, 474, 1467–1479. [Google Scholar] [CrossRef]
- Mullooly, M.; McGowan, P.M.; Crown, J.; Duffy, M.J. The ADAMs family of proteases as targets for the treatment of cancer. Cancer Biol. Ther. 2016, 17, 870–880. [Google Scholar] [CrossRef]
- Mohan, S.; Thompson, G.R.; Amaar, Y.G.; Hathaway, G.; Tschesche, H.; Baylink, D. ADAM-9 is an insulin-like growth factor binding protein-5 protease produced and secreted by human osteoblasts. Biochemistry 2002, 41, 15394–15403. [Google Scholar] [CrossRef] [PubMed]
- Dyczynska, E.; Sun, D.; Yi, H.; Sehara-Fujisawa, A.; Blobel, C.P.; Zolkiewska, A. Proteolytic processing of delta-like 1 by ADAM proteases. J. Biol. Chem. 2007, 282, 436–444. [Google Scholar] [CrossRef]
- English, W.R.; Corvol, P.; Murphy, G. LPS activates ADAM9 dependent shedding of ACE from endothelial cells. Biochem. Biophys. Res. Commun. 2012, 421, 70–75. [Google Scholar] [CrossRef]
- Caescu, C.I.; Jeschke, G.R.; Turk, B. Active-site determinants of substrate recognition by the metalloproteinases TACE and ADAM10. Biochem. J. 2009, 424, 79–88. [Google Scholar] [CrossRef]
- Ray, B.; Maloney, B.; Sambamurti, K.; Karnati, H.K.; Nelson, P.T.; Greig, N.H.; Lahiri, D.K. Rivastigmine modifies the α-secretase pathway and potentially early Alzheimer’s disease. Transl. Psychiatry 2020, 10, 47. [Google Scholar] [CrossRef]
- Sennvik, K.; Fastbom, J.; Blomberg, M.; Wahlund, L.O.; Winblad, B.; Benedikz, E. Levels of alpha- and beta-secretase cleaved amyloid precursor protein in the cerebrospinal fluid of Alzheimer’s disease patients. Neurosci. Lett. 2000, 278, 169–172. [Google Scholar] [CrossRef]
- Cong, L.; Jia, J. Promoter polymorphisms which regulate ADAM9 transcription are protective against sporadic Alzheimer’s disease. Neurobiol. Aging 2011, 32, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Guaiquil, V.H.; Hewing, N.J.; Chiang, M.F.; Rosenblatt, M.I.; Chan, R.V.; Blobel, C.P. A murine model for retinopathy of prematurity identifies endothelial cell proliferation as a potential mechanism for plus disease. Investig. Ophthalmol. Vis. Sci. 2013, 54, 5294–5302. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Opdenakker, G.; Abu El-Asrar, A. Metalloproteinases mediate diabetes-induced retinal neuropathy and vasculopathy. Cell. Mol. Life Sci. 2019, 76, 3157–3166. [Google Scholar] [CrossRef]
- Folkesson, M.; Li, C.; Frebelius, S.; Swedenborg, J.; Wågsäter, D.; Williams, K.J.; Eriksson, P.; Roy, J.; Liu, M.L. Proteolytically active ADAM10 and ADAM17 carried on membrane microvesicles in human abdominal aortic aneurysms. Thromb. Haemost. 2015, 114, 1165–1174. [Google Scholar] [CrossRef] [PubMed]
- Geng, L.; Wang, W.; Chen, Y.; Cao, J.; Lu, L.; Chen, Q.; He, R.; Shen, W. Elevation of ADAM10, ADAM17, MMP-2 and MMP-9 expression with media degeneration features CaCl2-induced thoracic aortic aneurysm in a rat model. Exp. Mol. Pathol. 2010, 89, 72–81. [Google Scholar] [CrossRef]
- Yu, J.; Liu, R.; Huang, J.; Wang, L.; Wang, W. Inhibition of Phosphatidylinositol 3-kinease suppresses formation and progression of experimental abdominal aortic aneurysms. Sci. Rep. 2017, 7, 15208. [Google Scholar] [CrossRef]
- Baggen, J.; Thibaut, H.J.; Hurdiss, D.L.; Wahedi, M.; Marceau, C.D.; van Vliet, A.L.W.; Carette, J.E.; van Kuppeveld, F.J.M. Identification of the Cell-Surface Protease ADAM9 as an Entry Factor for Encephalomyocarditis Virus. mBio 2019, 10, e01780-19. [Google Scholar] [CrossRef]
- Bazzone, L.E.; King, M.; MacKay, C.R.; Kyawe, P.P.; Meraner, P.; Lindstrom, D.; Rojas-Quintero, J.; Owen, C.A.; Wang, J.P.; Brass, A.L.; et al. A Disintegrin and Metalloproteinase 9 Domain (ADAM9) Is a Major Susceptibility Factor in the Early Stages of Encephalomyocarditis Virus Infection. mBio 2019, 10, e02734-18. [Google Scholar] [CrossRef]
- Liu, R.; Wang, F.; Guo, Y.; Yang, J.; Chen, S.; Gao, X.; Wang, X. MicroRNA-425 promotes the development of lung adenocarcinoma via targeting A disintegrin and metalloproteinases 9 (ADAM9). Onco Targets Ther. 2018, 11, 4065–4073. [Google Scholar] [CrossRef]
- Chang, J.H.; Lai, S.L.; Chen, W.S.; Hung, W.Y.; Chow, J.M.; Hsiao, M.; Lee, W.J.; Chien, M.H. Quercetin suppresses the metastatic ability of lung cancer through inhibiting Snail-dependent Akt activation and Snail-independent ADAM9 expression pathways. Biochim. Biophys. Acta Mol. Cell Res. 2017, 1864, 1746–1758. [Google Scholar] [CrossRef] [PubMed]
- Kossmann, C.M.; Annereau, M.; Thomas-Schoemann, A.; Nicco-Overney, C.; Chéreau, C.; Batteux, F.; Alexandre, J.; Lemare, F. ADAM9 expression promotes an aggressive lung adenocarcinoma phenotype. Tumour Biol. 2017, 39, 1010428317716077. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Qi, J.; Chen, N.; Fu, W.; Zhou, B.; He, A. High expression of a disintegrin and metalloproteinase-9 predicts a shortened survival time in completely resected stage I non-small cell lung cancer. Oncol. Lett. 2013, 5, 1461–1466. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Chen, N.; Qi, J.; Zhou, B.; Qiu, X. HDGF and ADAM9 are novel molecular staging biomarkers, prognostic biomarkers and predictive biomarkers for adjuvant chemotherapy in surgically resected stage I non-small cell lung cancer. J. Cancer Res. Clin. Oncol. 2014, 140, 1441–1449. [Google Scholar] [CrossRef]
- Fritzsche, F.R.; Jung, M.; Tölle, A.; Wild, P.; Hartmann, A.; Wassermann, K.; Rabien, A.; Lein, M.; Dietel, M.; Pilarsky, C.; et al. ADAM9 expression is a significant and independent prognostic marker of PSA relapse in prostate cancer. Eur. Urol. 2008, 54, 1097–1106. [Google Scholar] [CrossRef]
- Hua, Y.; Liang, C.; Miao, C.; Wang, S.; Su, S.; Shao, P.; Liu, B.; Bao, M.; Zhu, J.; Xu, A.; et al. MicroRNA-126 inhibits proliferation and metastasis in prostate cancer via regulation of ADAM9. Oncol. Lett. 2018, 15, 9051–9060. [Google Scholar] [CrossRef]
- Lin, Y.-W.; Wen, Y.-C.; Chu, C.-Y.; Tung, M.-C.; Yang, Y.-C.; Hua, K.-T.; Pan, K.-F.; Hsiao, M.; Lee, W.-J.; Chien, M.-H. Stabilization of ADAM9 by N-α-acetyltransferase 10 protein contributes to promoting progression of androgen-independent prostate cancer. Cell Death Dis. 2020, 11, 591. [Google Scholar] [CrossRef]
- Mygind, K.J.; Schwarz, J.; Sahgal, P.; Ivaska, J.; Kveiborg, M. Loss of ADAM9 expression impairs β1 integrin endocytosis, focal adhesion formation and cancer cell migration. J. Cell Sci. 2018, 131, jcs205393. [Google Scholar] [CrossRef]
- Oh, S.; Park, Y.; Lee, H.J.; Lee, J.; Lee, S.H.; Baek, Y.S.; Chun, S.K.; Lee, S.M.; Kim, M.; Chon, Y.E.; et al. A Disintegrin and Metalloproteinase 9 (ADAM9) in Advanced Hepatocellular Carcinoma and Their Role as a Biomarker During Hepatocellular Carcinoma Immunotherapy. Cancers 2020, 12, 745. [Google Scholar] [CrossRef]
- Kohga, K.; Takehara, T.; Tatsumi, T.; Ishida, H.; Miyagi, T.; Hosui, A.; Hayashi, N. Sorafenib inhibits the shedding of major histocompatibility complex class I-related chain A on hepatocellular carcinoma cells by down-regulating a disintegrin and metalloproteinase 9. Hepatology 2010, 51, 1264–1273. [Google Scholar] [CrossRef]
- Dong, Y.; Wu, Z.; He, M.; Chen, Y.; Chen, Y.; Shen, X.; Zhao, X.; Zhang, L.; Yuan, B.; Zeng, Z. ADAM9 mediates the interleukin-6-induced Epithelial-Mesenchymal transition and metastasis through ROS production in hepatoma cells. Cancer Lett. 2018, 421, 1–14. [Google Scholar] [CrossRef] [PubMed]
- O’Shea, C.; McKie, N.; Buggy, Y.; Duggan, C.; Hill, A.D.; McDermott, E.; O’Higgins, N.; Duffy, M.J. Expression of ADAM-9 mRNA and protein in human breast cancer. Int. J. Cancer 2003, 105, 754–761. [Google Scholar] [CrossRef]
- Oria, V.O.; Lopatta, P.; Schilling, O. The pleiotropic roles of ADAM9 in the biology of solid tumors. Cell. Mol. Life Sci. 2018, 75, 2291–2301. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.J.; Zou, J.X.; Wang, H.; Duan, Z.J.; Wang, H.B.; Chen, P.; Liu, P.Q.; Xu, J.Z.; Chen, H.W. Histone methyltransferase NSD2 mediates the survival and invasion of triple-negative breast cancer cells via stimulating ADAM9-EGFR-AKT signaling. Acta Pharmacol. Sin. 2019, 40, 1067–1075. [Google Scholar] [CrossRef]
- van Kampen, J.G.M.; van Hooij, O.; Jansen, C.F.; Smit, F.P.; van Noort, P.I.; Schultz, I.; Schaapveld, R.Q.J.; Schalken, J.A.; Verhaegh, G.W. miRNA-520f Reverses Epithelial-to-Mesenchymal Transition by Targeting ADAM9 and TGFBR2. Cancer Res. 2017, 77, 2008–2017. [Google Scholar] [CrossRef] [PubMed]
- Oria, V.O.; Lopatta, P.; Schmitz, T.; Preca, B.T.; Nyström, A.; Conrad, C.; Bartsch, J.W.; Kulemann, B.; Hoeppner, J.; Maurer, J.; et al. ADAM9 contributes to vascular invasion in pancreatic ductal adenocarcinoma. Mol. Oncol. 2019, 13, 456–479. [Google Scholar] [CrossRef]
- Yuan, P.; He, X.H.; Rong, Y.F.; Cao, J.; Li, Y.; Hu, Y.P.; Liu, Y.; Li, D.; Lou, W.; Liu, M.F. KRAS/NF-κB/YY1/miR-489 Signaling Axis Controls Pancreatic Cancer Metastasis. Cancer Res. 2017, 77, 100–111. [Google Scholar] [CrossRef]
- Xing, C.; Ye, H.; Wang, W.; Sun, M.; Zhang, J.; Zhao, Z.; Jiang, G. Circular RNA ADAM9 facilitates the malignant behaviours of pancreatic cancer by sponging miR-217 and upregulating PRSS3 expression. Artif. Cells Nanomed. Biotechnol. 2019, 47, 3920–3928. [Google Scholar] [CrossRef]
- Fan, X.; Wang, Y.; Zhang, C.; Liu, L.; Yang, S.; Wang, Y.; Liu, X.; Qian, Z.; Fang, S.; Qiao, H.; et al. ADAM9 Expression Is Associate with Glioma Tumor Grade and Histological Type, and Acts as a Prognostic Factor in Lower-Grade Gliomas. Int. J. Mol. Sci. 2016, 17, 1276. [Google Scholar] [CrossRef]
- Sarkar, S.; Zemp, F.J.; Senger, D.; Robbins, S.M.; Yong, V.W. ADAM-9 is a novel mediator of tenascin-C-stimulated invasiveness of brain tumor-initiating cells. Neuro Oncol. 2015, 17, 1095–1105. [Google Scholar] [CrossRef]
- Shintani, Y.; Higashiyama, S.; Ohta, M.; Hirabayashi, H.; Yamamoto, S.; Yoshimasu, T.; Matsuda, H.; Matsuura, N. Overexpression of ADAM9 in Non-Small Cell Lung Cancer Correlates with Brain Metastasis. Cancer Res. 2004, 64, 4190–4196. [Google Scholar] [CrossRef] [PubMed]
- Chiu, K.L.; Kuo, T.T.; Kuok, Q.Y.; Lin, Y.S.; Hua, C.H.; Lin, C.Y.; Su, P.Y.; Lai, L.C.; Sher, Y.P. ADAM9 enhances CDCP1 protein expression by suppressing miR-218 for lung tumor metastasis. Sci. Rep. 2015, 5, 16426. [Google Scholar] [CrossRef]
- Chiu, K.L.; Lin, Y.S.; Kuo, T.T.; Lo, C.C.; Huang, Y.K.; Chang, H.F.; Chuang, E.Y.; Lin, C.C.; Cheng, W.C.; Liu, Y.N.; et al. ADAM9 enhances CDCP1 by inhibiting miR-1 through EGFR signaling activation in lung cancer metastasis. Oncotarget 2017, 8, 47365–47378. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.F.; Wang, S.; Xue, W.H.; Cheng, J.L. microRNA-590 suppresses the tumorigenesis and invasiveness of non-small cell lung cancer cells by targeting ADAM9. Mol. Cell. Biochem. 2016, 423, 29–37. [Google Scholar] [CrossRef] [PubMed]
- Wan, J.; Hao, L.; Zheng, X.; Li, Z. Circular RNA circ_0020123 promotes non-small cell lung cancer progression by acting as a ceRNA for miR-488-3p to regulate ADAM9 expression. Biochem. Biophys. Res. Commun. 2019, 515, 303–309. [Google Scholar] [CrossRef]
- Zhang, J.; Pan, Y.F.; Ding, Z.W.; Yang, G.Z.; Tan, Y.X.; Yang, C.; Jiang, T.Y.; Liu, L.J.; Zhang, B.; Han, T.; et al. RMP promotes venous metastases of hepatocellular carcinoma through promoting IL-6 transcription. Oncogene 2015, 34, 1575–1583. [Google Scholar] [CrossRef]
- Xiang, L.Y.; Ou, H.H.; Liu, X.C.; Chen, Z.J.; Li, X.H.; Huang, Y.; Yang, D.H. Loss of tumor suppressor miR-126 contributes to the development of hepatitis B virus-related hepatocellular carcinoma metastasis through the upregulation of ADAM9. Tumour Biol. 2017, 39, 1010428317709128. [Google Scholar] [CrossRef] [PubMed]
- Wan, D.; Shen, S.; Fu, S.; Preston, B.; Brandon, C.; He, S.; Shen, C.; Wu, J.; Wang, S.; Xie, W.; et al. miR-203 suppresses the proliferation and metastasis of hepatocellular carcinoma by targeting oncogene ADAM9 and oncogenic long non-coding RNA HULC. Anticancer Agents Med. Chem. 2016, 16, 414–423. [Google Scholar] [CrossRef] [PubMed]
- Hu, D.; Shen, D.; Zhang, M.; Jiang, N.; Sun, F.; Yuan, S.; Wan, K. MiR-488 suppresses cell proliferation and invasion by targeting ADAM9 and lncRNA HULC in hepatocellular carcinoma. Am. J. Cancer Res. 2017, 7, 2070–2080. [Google Scholar]
- Micocci, K.C.; Moritz, M.N.; Lino, R.L.; Fernandes, L.R.; Lima, A.G.; Figueiredo, C.C.; Morandi, V.; Selistre-de-Araujo, H.S. ADAM9 silencing inhibits breast tumor cells transmigration through blood and lymphatic endothelial cells. Biochimie 2016, 128, 174–182. [Google Scholar] [CrossRef]
- Zhu, N.; Zhang, D.; Xie, H.; Zhou, Z.; Chen, H.; Hu, T.; Bai, Y.; Shen, Y.; Yuan, W.; Jing, Q.; et al. Endothelial-specific intron-derived miR-126 is down-regulated in human breast cancer and targets both VEGFA and PIK3R2. Mol. Cell. Biochem. 2011, 351, 157–164. [Google Scholar] [CrossRef]
- Wang, C.Z.; Yuan, P.; Li, Y. MiR-126 regulated breast cancer cell invasion by targeting ADAM9. Int. J. Clin. Exp. Pathol. 2015, 8, 6547–6553. [Google Scholar]
- Qin, C.; Zhao, Y.; Gong, C.; Yang, Z. MicroRNA-154/ADAM9 axis inhibits the proliferation, migration and invasion of breast cancer cells. Oncol. Lett. 2017, 14, 6969–6975. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Zhang, Y.; Ding, W.; Lin, Y.; Huang, Z.; Luo, Q. MiR-33a suppresses breast cancer cell proliferation and metastasis by targeting ADAM9 and ROS1. Protein Cell 2015, 6, 881–889. [Google Scholar] [CrossRef]
- Kristensen, L.S.; Andersen, M.S.; Stagsted, L.V.W.; Ebbesen, K.K.; Hansen, T.B.; Kjems, J. The biogenesis, biology and characterization of circular RNAs. Nat. Rev. Genet. 2019, 20, 675–691. [Google Scholar] [CrossRef]
- Wu, D.M.; Wen, X.; Han, X.R.; Wang, S.; Wang, Y.J.; Shen, M.; Fan, S.H.; Zhang, Z.F.; Shan, Q.; Li, M.Q.; et al. Bone Marrow Mesenchymal Stem Cell-Derived Exosomal MicroRNA-126-3p Inhibits Pancreatic Cancer Development by Targeting ADAM9. Mol. Ther. Nucleic Acids 2019, 16, 229–245. [Google Scholar] [CrossRef] [PubMed]
- Hamada, S.; Satoh, K.; Fujibuchi, W.; Hirota, M.; Kanno, A.; Unno, J.; Masamune, A.; Kikuta, K.; Kume, K.; Shimosegawa, T. MiR-126 acts as a tumor suppressor in pancreatic cancer cells via the regulation of ADAM9. Mol. Cancer Res. 2012, 10, 3–10. [Google Scholar] [CrossRef]
- Ji, T.; Zhang, X.; Li, W. MicroRNA-543 inhibits proliferation, invasion and induces apoptosis of glioblastoma cells by directly targeting ADAM9. Mol. Med. Rep. 2017, 16, 6419–6427. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Wang, S.; Yuan, A.; Yuan, X.; Liu, B. MicroRNA-140 represses glioma growth and metastasis by directly targeting ADAM9. Oncol. Rep. 2016, 36, 2329–2338. [Google Scholar] [CrossRef]
- Josson, S.; Anderson, C.S.; Sung, S.Y.; Johnstone, P.A.; Kubo, H.; Hsieh, C.L.; Arnold, R.; Gururajan, M.; Yates, C.; Chung, L.W. Inhibition of ADAM9 expression induces epithelial phenotypic alterations and sensitizes human prostate cancer cells to radiation and chemotherapy. Prostate 2011, 71, 232–240. [Google Scholar] [CrossRef]
- Ueno, M.; Shiomi, T.; Mochizuki, S.; Chijiiwa, M.; Shimoda, M.; Kanai, Y.; Kataoka, F.; Hirasawa, A.; Susumu, N.; Aoki, D.; et al. ADAM9 is over-expressed in human ovarian clear cell carcinomas and suppresses cisplatin-induced cell death. Cancer Sci. 2018, 109, 471–482. [Google Scholar] [CrossRef]
- Amendola, R.S.; Martin, A.C.; Selistre-de-Araújo, H.S.; Paula-Neto, H.A.; Saldanha-Gama, R.; Barja-Fidalgo, C. ADAM9 disintegrin domain activates human neutrophils through an autocrine circuit involving integrins and CXCR2. J. Leukoc. Biol. 2015, 97, 951–962. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Liu, Q.; Zhang, X.; Liu, X.; Zhou, B.; Chen, J.; Huang, D.; Li, J.; Li, H.; Chen, F.J.N. DNA of neutrophil extracellular traps promotes cancer metastasis via CCDC25. Nature 2020, 583, 133–138. [Google Scholar] [CrossRef] [PubMed]
- Puissegur, M.-P.; Lay, G.; Gilleron, M.; Botella, L.; Nigou, J.; Marrakchi, H.; Mari, B.; Duteyrat, J.-L.; Guerardel, Y.; Kremer, L. Mycobacterial lipomannan induces granuloma macrophage fusion via a TLR2-dependent, ADAM9-and β1 integrin-mediated pathway. J. Immunol. 2007, 178, 3161–3169. [Google Scholar] [CrossRef] [PubMed]
- Brooks, P.J.; Glogauer, M.; McCulloch, C.A. An overview of the derivation and function of multinucleated giant cells and their role in pathologic processes. Am. J. Pathol. 2019, 189, 1145–1158. [Google Scholar] [CrossRef]
- Nieswandt, B.; Hafner, M.; Echtenacher, B.; Männel, D.N. Lysis of tumor cells by natural killer cells in mice is impeded by platelets. Cancer Res. 1999, 59, 1295–1300. [Google Scholar]
- Palumbo, J.S.; Talmage, K.E.; Massari, J.V.; La Jeunesse, C.M.; Flick, M.J.; Kombrinck, K.W.; Jirousková, M.; Degen, J.L. Platelets and fibrin(ogen) increase metastatic potential by impeding natural killer cell-mediated elimination of tumor cells. Blood 2005, 105, 178–185. [Google Scholar] [CrossRef]
- Labelle, M.; Begum, S.; Hynes, R.O. Direct signaling between platelets and cancer cells induces an epithelial-mesenchymal-like transition and promotes metastasis. Cancer Cell 2011, 20, 576–590. [Google Scholar] [CrossRef]
- Mammadova-Bach, E.; Zigrino, P.; Brucker, C.; Bourdon, C.; Freund, M.; De Arcangelis, A.; Abrams, S.I.; Orend, G.; Gachet, C.; Mangin, P.H. Platelet integrin α6β1 controls lung metastasis through direct binding to cancer cell-derived ADAM9. JCI Insight 2016, 1, e88245. [Google Scholar] [CrossRef]
- He, B.; Zhao, Z.; Cai, Q.; Zhang, Y.; Zhang, P.; Shi, S.; Xie, H.; Peng, X.; Yin, W.; Tao, Y.; et al. miRNA-based biomarkers, therapies, and resistance in Cancer. Int. J. Biol. Sci. 2020, 16, 2628–2647. [Google Scholar] [CrossRef]
- Kim, J.M.; Jeung, H.C.; Rha, S.Y.; Yu, E.J.; Kim, T.S.; Shin, Y.K.; Zhang, X.; Park, K.H.; Park, S.W.; Chung, H.C.; et al. The effect of disintegrin-metalloproteinase ADAM9 in gastric cancer progression. Mol. Cancer Ther. 2014, 13, 3074–3085. [Google Scholar] [CrossRef] [PubMed]
- Overall, C.M.; Kleifeld, O. Tumour microenvironment-opinion: Validating matrix metalloproteinases as drug targets and anti-targets for cancer therapy. Nat. Rev. Cancer 2006, 6, 227–239. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, M.H.; Tsai, J.P.; Yang, S.F.; Chiou, H.L.; Lin, C.L.; Hsieh, Y.H.; Chang, H.R. Fisetin Suppresses the Proliferation and Metastasis of Renal Cell Carcinoma through Upregulation of MEK/ERK-Targeting CTSS and ADAM9. Cells 2019, 8, 948. [Google Scholar] [CrossRef] [PubMed]
- Lei, D.; Zhang, F.; Yao, D.; Xiong, N.; Jiang, X.; Zhao, H. Galangin increases ERK1/2 phosphorylation to decrease ADAM9 expression and prevents invasion in A172 glioma cells. Mol. Med. Rep. 2018, 17, 667–673. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.F.; Yang, S.F.; Chiou, H.L.; Hsu, W.H.; Hsu, J.C.; Liu, C.J.; Hsieh, Y.H. Licochalcone A inhibits the invasive potential of human glioma cells by targeting the MEK/ERK and ADAM9 signaling pathways. Food Funct. 2018, 9, 6196–6204. [Google Scholar] [CrossRef] [PubMed]
- Arai, J.; Goto, K.; Stephanou, A.; Tanoue, Y.; Ito, S.; Muroyama, R.; Matsubara, Y.; Nakagawa, R.; Morimoto, S.; Kaise, Y.; et al. Predominance of regorafenib over sorafenib: Restoration of membrane-bound MICA in hepatocellular carcinoma cells. J. Gastroenterol. Hepatol. 2018, 33, 1075–1081. [Google Scholar] [CrossRef]
- Arai, J.; Goto, K.; Otoyama, Y.; Nakajima, Y.; Sugiura, I.; Kajiwara, A.; Tojo, M.; Ichikawa, Y.; Uozumi, S.; Shimozuma, Y.; et al. Leukotriene receptor antagonists enhance HCC treatment efficacy by inhibiting ADAMs and suppressing MICA shedding. Cancer Immunol. Immunother. 2020. [Google Scholar] [CrossRef]
| Type | Role of ADAM9 in Cancer | Reference | |
|---|---|---|---|
| Lung Cancer | Clinical Significance | Overexpressed in cancer | [25,26,70,71] |
| Negative correlation with OS | [25,70,71,72,73,74] | ||
| Mechanism | ADAM9-tPA-CDCP1-Metastasis | [25] | |
| ADAM9-ANGPT2-Metastasis | [26] | ||
| ADAM9-IL8/VEGFA-Angiogenesis | [26,72] | ||
| Prostate Cancer | Clinical Relevance | Overexpressed in cancer | [75,76] |
| Negative correlation with RFS | [75] | ||
| Mechanism | Naa10p-ADAM9-Tumorigenesis/Metastasis | [77] | |
| ADAM9-Integrin Degradation | [78] | ||
| Liver Cancer | Clinical Relevance | Negative correlation with immunotherapy response | [79,80] |
| Mechanism | ADAM9-MICA cleavage-immune evasion | [80] | |
| IL-6-ADAM9-JNK-Metastasis | [81] | ||
| Breast Cancer | Clinical Relevance | Overexpressed in cancer | [82] |
| Positive correlation with progression | [83] | ||
| Mechanism | NSD2-ADAM9-Tumorigenesis | [84] | |
| Pancreatic Cancer | Clinical Relevance | Overexpressed in cancer | [24,85] |
| Positive correlation with progression | [86] | ||
| Negative correlation with OS | [24,85] | ||
| Mechanism | KRAS-ADAM9-Tumorigenesis | [87] | |
| ADAM9-MEK-ERK-Tumorigenesis | [24] | ||
| Circ-ADAM9-ERK-Tumorigenesis | [88] | ||
| Brain Cancer | Clinical Relevance | Overexpressed in cancer | [89,90] |
| Negative correlation with OS/PFS | [89,90] | ||
| Mechanism | TNC-ADAM9-Metastasis | [90] | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chou, C.-W.; Huang, Y.-K.; Kuo, T.-T.; Liu, J.-P.; Sher, Y.-P. An Overview of ADAM9: Structure, Activation, and Regulation in Human Diseases. Int. J. Mol. Sci. 2020, 21, 7790. https://doi.org/10.3390/ijms21207790
Chou C-W, Huang Y-K, Kuo T-T, Liu J-P, Sher Y-P. An Overview of ADAM9: Structure, Activation, and Regulation in Human Diseases. International Journal of Molecular Sciences. 2020; 21(20):7790. https://doi.org/10.3390/ijms21207790
Chicago/Turabian StyleChou, Cheng-Wei, Yu-Kai Huang, Ting-Ting Kuo, Jing-Pei Liu, and Yuh-Pyng Sher. 2020. "An Overview of ADAM9: Structure, Activation, and Regulation in Human Diseases" International Journal of Molecular Sciences 21, no. 20: 7790. https://doi.org/10.3390/ijms21207790
APA StyleChou, C.-W., Huang, Y.-K., Kuo, T.-T., Liu, J.-P., & Sher, Y.-P. (2020). An Overview of ADAM9: Structure, Activation, and Regulation in Human Diseases. International Journal of Molecular Sciences, 21(20), 7790. https://doi.org/10.3390/ijms21207790





