TNF-Block Genotypes Influence Susceptibility to HIV-Associated Sensory Neuropathy in Indonesians and South Africans
Abstract
1. Introduction
2. Results
2.1. Measures of the Severity of HIV Disease Predict HIV-SN in Indonesians and Africans
2.2. Two Alleles Associated with HIV-SN in Indonesians but not Africans
2.3. One Haplotype Containing rs9281523*C and rs1800629*A Associated with HIV-SN in Indonesians but Not Africans
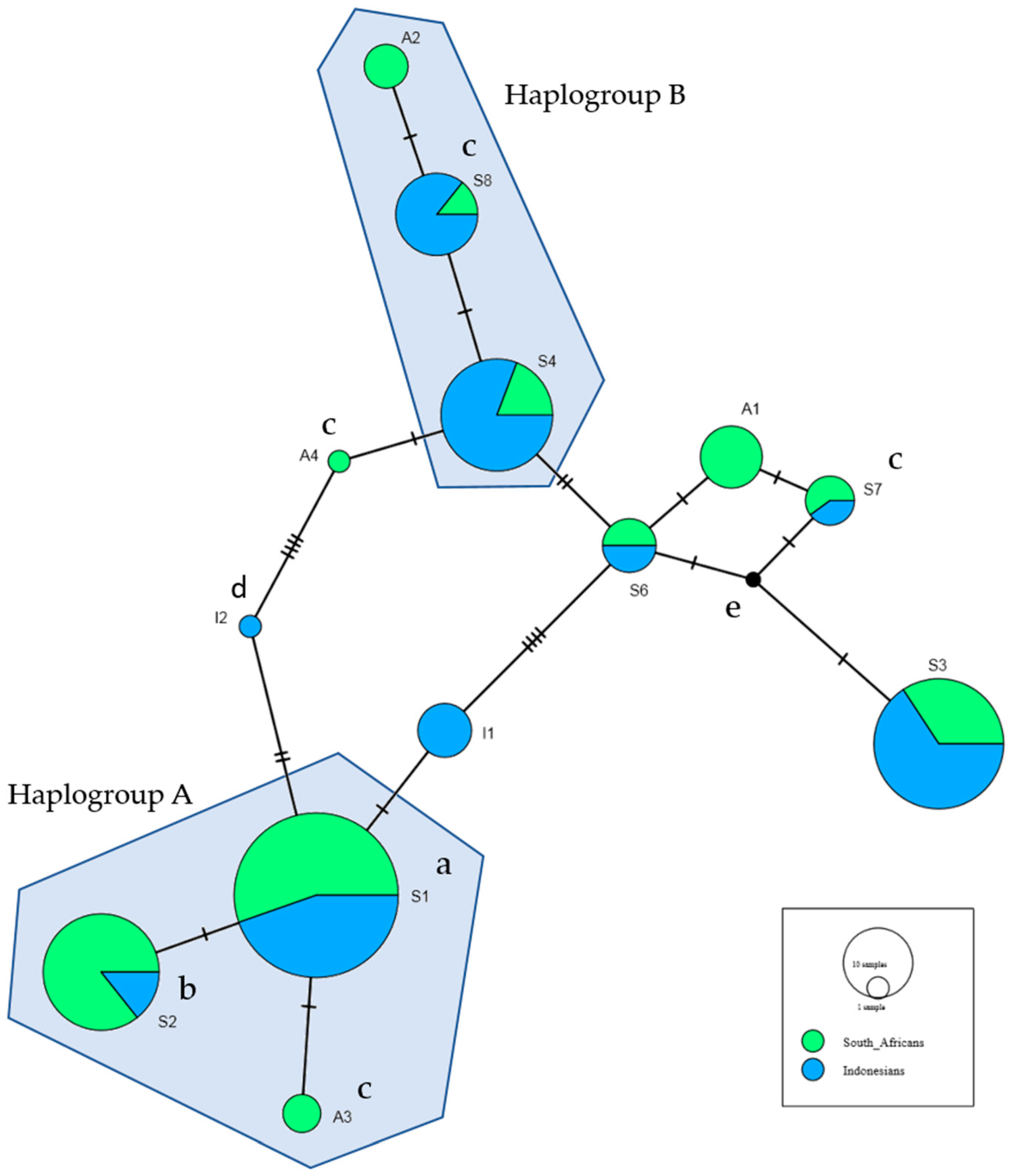
2.4. One Haplogroup Contained the Two Haplotypes Associated with HIV-SN in Africans and Indonesians
3. Discussion
4. Materials and Methods
4.1. Participants and Phenotypes
4.2. Genotyping
4.3. Haplotype Analyses
4.4. Statistical Analyses
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| HIV | Human immunodeficiency virus |
| HIV-SN | HIV-associated sensory neuropathy |
| BPNS | Brief peripheral neuropathy screening tool |
| TNF | Tumor necrosis factor |
| CXCR4 | C-X-C chemokine receptor type 4 |
| CXCL12 | C-X-C chemokine ligand 12 |
| MHC | Major histocompatibility complex |
| LD | Linkage disequilibrium |
| LTB | Lymphotoxin beta |
| LTA | Lymphotoxin alpha |
| NCR3 | Natural Cytotoxicity Triggering Receptor 3 |
| LST1 | Leukocyte-specific transcript 1 |
| NFKBIL1 | Nuclear Factor if Kappa Light Polypeptide Gene Enhancer In B-Cells Inhibitor-Like 1 |
| DDX39B | DExD-Box Helicase 39B |
| MCCD1 | Mitochondrial Coiled-Coil Domain 1 |
| AH | Ancestral Haplotype |
| S1-8 | Shared haplotypes 1-8 |
| A1-4 | African haplotypes 1-4 |
| I1-2 | Indonesian haplotypes 1 and 2 |
| GTEX | Genotype-Tissue Expression |
| eQTL | Expression quantitative trait loci |
| C4A | Complement Component 4A |
| HLA | Human leukocyte antigen |
| MICB | MHC class I polypeptide-related sequence |
References
- Cherry, C.L.; Wadley, A.L.; Kamerman, P.R. Painful HIV-associated sensory neuropathy. Pain Manag. 2012, 2, 543–552. [Google Scholar] [CrossRef]
- Phillips, T.J.C.; Brown, M.; Ramirez, J.D.; Perkins, J.; Woldeamanuel, Y.W.; Williams, A.C.; Orengo, C.; Bennett, D.L.H.; Bodi, I.; Cox, S.; et al. Sensory, psychological, and metabolic dysfunction in HIV-associated peripheral neuropathy: A cross-sectional deep profiling study. Pain 2014, 155, 1846–1860. [Google Scholar] [CrossRef] [PubMed]
- Wadley, A.L.; Mitchell, D.; Kamerman, P.R. Resilience does not explain the dissociation between chronic pain and physical activity in South Africans living with HIV. PeerJ 2016, 4, e2464. [Google Scholar] [CrossRef] [PubMed]
- Pujades-Rodríguez, M.; Dantony, E.; Pinoges, L.; Ecochard, R.; Etard, J.-F.; Carrillo-Casas, E.; Szumilin, E.; AIDS Working Group of Médecins Sans Frontières. Toxicity associated with stavudine dose reduction from 40 to 30 mg in first-line antiretroviral therapy. PLoS ONE 2011, 6, e28112. [Google Scholar]
- Affandi, J.S.; Price, P.; Imran, D.; Yunihastuti, E.; Djauzi, S.; Cherry, C.L. Can we predict neuropathy risk before stavudine prescription in a resource-limited setting? AIDS Res. Hum. Retrovir. 2008, 24, 1281–1284. [Google Scholar] [CrossRef]
- Cherry, C.L.; Rosenow, A.; Affandi, J.S.; McArthur, J.C.; Wesselingh, S.L.; Price, P. Cytokine genotype suggests a role for inflammation in nucleoside analog-associated sensory neuropathy (NRTI-SN) and predicts an individual’s NRTI-SN risk. AIDS Res. Hum. Retrovir. 2008, 24, 117–123. [Google Scholar] [CrossRef]
- Maritz, J.; Benatar, M.; Dave, J.A.; Harrison, T.B.; Badri, M.; Levitt, N.S.; Heckmann, J.M. HIV neuropathy in South Africans: Frequency, characteristics, and risk factors. Muscle Nerve 2010, 41, 599–606. [Google Scholar] [CrossRef]
- Pettersen, J.A.; Jones, G.; Worthington, C.; Krentz, H.B.; Keppler, O.T.; Hoke, A.; Gill, M.J.; Power, C. Sensory neuropathy in human immunodeficiency virus/acquired immunodeficiency syndrome patients: Protease inhibitor-mediated neurotoxicity. Ann. Neurol. 2006, 59, 816–824. [Google Scholar] [CrossRef]
- Wadley, A.L.; Cherry, C.L.; Price, P.; Kamerman, P.R. HIV neuropathy risk factors and symptom characterization in stavudine-exposed South Africans. J. Pain Symptom Manag. 2011, 41, 700–706. [Google Scholar] [CrossRef]
- Octaviana, F.; Safri, A.Y.; Setiawan, D.D.; Estiasari, R.; Imran, D.; Ranakusuma, T.; Affandi, J.; Cherry, C.L.; Price, P. Detectable plasma HIV RNA is associated with sensory neuropathy in patients with HIV treated without stavudine. J. AIDS 2018, 79, e108–e110. [Google Scholar] [CrossRef]
- Empl, M.; Renaud, S.; Erne, B.; Fuhr, P.; Straube, A.; Schaeren-Wiemers, N.; Steck, A.J. TNF-alpha expression in painful and nonpainful neuropathies. Neurology 2001, 56, 1371–1377. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Zheng, W.; Liu, S.; Zeng, W.; Levitt, R.C.; Candiotti, K.A.; Lubarsky, D.A.; Hao, S. HSV-mediated p55TNFSR reduces neuropathic pain induced by HIV gp120 in rats through CXCR4 activity. Gene Ther. 2014, 21, 328–336. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Rizzardi, G.P.; Barcellini, W.; Tambussi, G.; Lillo, F.; Malnati, M.; Perrin, L.; Lazzarin, A. Plasma levels of soluble CD30, tumour necrosis factor (TNF)-alpha and TNF receptors during primary HIV-1 infection: Correlation with HIV-1 RNA and the clinical outcome. AIDS 1996, 10, F45–F50. [Google Scholar] [CrossRef] [PubMed]
- Valente, F.P.; Tan, C.R.; Temple, S.E.; Phipps, M.; Witt, C.S.; Kaur, G.; Gut, I.; McGinn, S.; Allcock, R.J.; Chew, C.S.; et al. The evolution and diversity of TNF block haplotypes in European, Asian and Australian Aboriginal populations. Genes Immun. 2009, 10, 607–615. [Google Scholar] [CrossRef][Green Version]
- Chew, C.S.; Cherry, C.L.; Imran, D.; Yunihastuti, E.; Kamarulzaman, A.; Varna, S.; Ismail, R.; Phipps, M.; Aghafar, Z.; Gut, I.; et al. Tumour necrosis factor haplotypes associated with sensory neuropathy in Asian and Caucasian human immunodeficiency virus patients. Tissue Antigens 2011, 77, 126–130. [Google Scholar] [CrossRef]
- Wadley, A.L.; Hendry, L.M.; Kamerman, P.R.; Chew, C.S.N.; Price, P.; Cherry, C.L.; Lombard, Z. Role of TNF block genetic variants in HIV-associated sensory neuropathy in black Southern Africans. Eur. J. Hum. Genet. 2015, 23, 363–368. [Google Scholar] [CrossRef]
- Price, P.; Witt, C.; Allcock, R.; Sayer, D.; Garlepp, M.; Kok, C.C.; French, M.; Mallal, S.; Christiansen, F. The genetic basis for the association of the 8.1 ancestral haplotype (A1, B8, DR3) with multiple immunopathological diseases. Immunol. Rev. 1999, 167, 257–274. [Google Scholar] [CrossRef]
- Bayley, J.P.; de Rooij, H.; van den Elsen, P.J.; Huizinga, T.W.; Verweij, C.L. Functional analysis of linker-scan mutants spanning the −376, −308, −244, and −238 polymorphic sites of the TNF-α promoter. Cytokine 2001, 14, 316–323. [Google Scholar] [CrossRef]
- Witt, C.S.; Price, P.; Kaur, G.; Cheong, K.; Kanga, U.; Sayer, D.; Christiansen, F.; Mehra, N.K. Common HLA-B8-DR3 haplotype in Northern India is different from that found in Europe. Tissue Antigens 2002, 60, 474–480. [Google Scholar] [CrossRef]
- Zhang, J.; Zhao, L.; Wang, B.; Gao, J.; Wang, L.; Li, L.; Cui, B.; Hu, M.; Hong, J.; Gu, W.; et al. HLA-A*33-DR3 and A*33-DR9 haplotypes enhance the risk of type 1 diabetes in Han Chinese. J. Diabetes Investig. 2016, 7, 514–521. [Google Scholar] [CrossRef]
- Gaff, J.; Octaviana, F.; Ariyanto, I.; Cherry, C.; Laws, S.M.; Price, P. Polymorphisms in CAMKK2 associate with susceptibility to sensory neuropathy in HIV patients treated without stavudine. J. Neurovirol. 2019, 25, 814–824. [Google Scholar] [CrossRef] [PubMed]
- Van der Watt, J.J.; Benatar, M.G.; Harrison, T.B.; Carrara, H.; Heckman, J.M. Isoniazid exposure and pyridoxine levels in human immunodeficiency virus associated distal sensory neuropathy. Int. J. Tuberc. Lung Dis. 2015, 19, 1312–1319. [Google Scholar] [CrossRef] [PubMed]
- Childs, E.A.; Lyles, R.H.; Selnes, O.A.; Chen, B.; Miller, E.N.; Cohen, B.A.; Becker, J.T.; Mellors, J.; McArthur, J.C. Plasma viral load and CD4 lymphocytes predict HIV-associated dementia and sensory neuropathy. Neurology 1999, 52, 607–613. [Google Scholar] [CrossRef] [PubMed]
- Cameron, P.U.; Mallal, S.A.; French, M.A.; Dawkins, R.L. Major histocompatibility complex genes influence the outcome of HIV infection. Ancestral haplotypes with C4 null alleles explain diverse HLA associations. Hum. Immunol. 1990, 29, 282–295. [Google Scholar] [CrossRef]
- Keet, I.P.; Klein, M.R.; Just, J.J.; Kaslow, R.A. The role of host genetics in the natural history of HIV-1 infection: The needles in the haystack. AIDS 1996, 10, S59–S67. [Google Scholar] [CrossRef]
- Candore, G.; Lio, D.; Colonna Romano, G.; Caruso, C. Pathogenesis of autoimmune diseases associated with 8.1 ancestral haplotype: Effect of multiple gene interactions. Autoimmun. Rev. 2002, 1, 29–35. [Google Scholar] [CrossRef]
- Tan, J.H.; Temple, S.E.; Kee, C.; Waterer, G.W.; Tan, C.R.; Gut, I.; Price, P. Characterisation of TNF block haplotypes affecting the production of TNF and LTA. Tissue Antigens 2011, 77, 100–106. [Google Scholar] [CrossRef]
- Lam, T.H.; Shen, M.; Tay, M.Z.; Ren, E.C. Unique allelic eQTL clusters in human MHC haplotypes. G3 Genes/Genomes/Genetics 2017, 7, 2595–2604. [Google Scholar] [CrossRef]
- Octaviana, F.; Safri, A.Y.; Setiawan, D.D.; Estiasari, R.; Imran, D.; Ranakusuma, T.; Price, P. Neuropathic pain in HIV patients receiving art without stavudine in an Indonesia referral hospital. J. Neurol. Sci. 2019, 397, 146–149. [Google Scholar] [CrossRef]
- Cherry, C.L.; Wesselingh, S.L.; Lal, L.; McArthur, J.C. Evaluation of a clinical screening tool for HIV-associated sensory neuropathies. Neurology 2005, 65, 1778–1781. [Google Scholar] [CrossRef]
- Woldeamanuel, Y.W.; Kamerman, P.R.; Veliotes, D.G.; Phillips, T.J.; Asboe, D.; Boffito, M.; Rice, A.S. Development, Validation, and Field-Testing of an Instrument for Clinical Assessment of HIV-Associated Neuropathy and Neuropathic Pain in Resource-Restricted and Large Population Study Settings. PLoS ONE 2016, 11, e0164994. [Google Scholar] [CrossRef] [PubMed]
- Pillay, P.; Wadley, A.L.; Cherry, C.L.; Karstaedt, A.S.; Kamerman, P.R. Clinical diagnosis of sensory neuropathy in HIV patients treated with tenofovir: A 6-month follow-up study. J. Peripher. Nerv. Syst. 2019, 24, 304–313. [Google Scholar] [CrossRef] [PubMed]
- Warpe, B.M. Peripheral neuropathy due to tuberculosis. In Tuberculosis of the Central Nervous System: Pathogenesis, Imaging, and Management; Turgut, M., Akhaddar, A., Turgut, A.T., Garg, R.K., Eds.; Springer International Publishing: Cham, Switzerland, 2017; pp. 339–350. [Google Scholar]
- Scheet, P.; Stephens, M. A fast and flexible statistical model for large-scale population genotype data: Applications to inferring missing genotypes and haplotypic phase. Am. J. Hum. Genet. 2006, 78, 629–644. [Google Scholar] [CrossRef] [PubMed]
- Bandelt, H.J.; Forster, P.; Rohl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 1999, 16, 37–48. [Google Scholar] [CrossRef] [PubMed]
| Variable | Africans | Indonesians | ||||
|---|---|---|---|---|---|---|
| HIV-SN | HIV-SN | |||||
| +ve (n = 29) | −ve (n = 46) | pa,b | +ve (n = 34) | −ve (n = 167) | pa,b | |
| Age (years) | 40 (24–60) | 37 (19–58) | 0.11 | 36 (21–59) | 35 (19–60) | 0.68 |
| Height (cm) | 168 (147–179) | 163 (135–186) n = 45 | 0.03 | 167 (151–185) | 167 (142–180) n = 166 | 0.71 |
| Weight (kg) | 66 (45–112) | 55 (35–110) n = 44 | 0.03 | 59 (39–88) | 58.5 (37–104) n = 166 | 1.00 |
| Current CD4 T-cells/µL | 221 (22–685) | 300 (8–832) | 0.06 | 326 (44–729) | 458 (84–1166) | 0.003 |
| Nadir CD4 T-cells/µL | 107 (4–575) | 223 (8–771) | 0.002 | 54 (3–428) | 121 (1–599) n = 165 | 0.06 |
| HIV RNA >500 copies/mL | 21/29 (72%) | 25/46 (54%) | 0.12 | 6/29 (17%) | 7/163 (4.1%) | 0.005 |
| History of Tuberculosis | 6/28 (29%) | 3/45 (7%) | 0.08 c | 18/35 (53%) | 66/168 (39%) | 0.09 |
| Female Gender | 15/29 (52%) | 30/46 (65%) | 0.25 | 9/35 (26%) | 49/167 (29%) | 0.98 |
| SNP rsID | Africans (n = 75) | Indonesians (n = 202) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Minor Allele | MAF a | HIV-SN | pe | Minor Allele | MAF a | HIV-SN | pd | |||
| +ve b | −ve | +ve b | −ve c | |||||||
| rs2075582 (MCCD1) | C | 0.13 | 7/28 d (25%) | 12/46 (27%) | 0.87 | C | 0.34 | 19/34 e (56%) | 94/166 (57%) | 0.97 |
| rs9281523 (DDX39B) | - | 0.20 | 9/29 (31%) | 17/46 (37%) | 0.60 | - | 0.03 | 5/29 (17%) | 8/166 (5%) | 0.03 |
| rs11796 (DDX39B) | A | 0.41 | 16/29 (55%) | 31/46 (67%) | 0.27 | T | 0.35 | 21/35 (60%) | 95/167 (57%) | 0.73 |
| rs2523506 (DDX39B) | T | 0.12 | 6/28 (21%) | 9/46 (20%) | 0.85 | T | 0.34 | 13/34 (38%) | 70/165 (42%) | 0.65 |
| rs2523504 (intergenic) | T | 0.17 | 8/29 (28%) | 15/46 (33%) | 0.60 | T | 0.33 | 19/35 (54%) | 92/165 (56%) | 0.87 |
| rs2071594 (intergenic) | G | 0.41 | 16/29 (55%) | 31/46 (67%) | 0.29 | C | 0.37 | 21/35 (60%) | 98/165 (59%) | 0.95 |
| rs2071593 (intergenic) | A | 0.07 | 2/29 f (7%) | 8/46 (17%) | 0.30 | A | 0.12 | 7/34 (21%) | 38/167 (23%) | 1.00 |
| rs2071592 (NFKBIL1) | T | 0.42 | 16/29 (55%) | 30/46 (67%) | 0.32 | A | 0.30 | 19/34 (56%) | 83/166 (50%) | 0.53 |
| rs4947324 (intergenic) | T | 0.16 | 6/29 (21%) | 16/46 (35%) | 0.19 | T | 0.03 | 3/35 (9%) | 8/166 (5%) | 0.41 |
| rs909253 (LTA) | A | 0.41 | 16/29 (55%) | 31/46 (67%) | 0.29 | G | 0.36 | 21/35 (60%) | 99/167 (59%) | 0.94 |
| rs1041981 (LTA) | C | 0.39 | 15/29 (52%) | 30/46 (67%) | 0.20 | A | 0.36 | 21/35 (60%) | 99/167 (59%) | 0.94 |
| rs1799964 (intergenic) | C | 0.15 | 6/29 (21%) | 12/46 (27%) | 0.56 | C | 0.27 | 16/35 (46%) | 76/165 (46%) | 0.97 |
| rs1800629 (intergenic) | A | 0.23 | 10/29 (34%) | 19/46 (41%) | 0.56 | A | 0.04 | 6/35 (17%) | 11/167 (7%) | 0.04 |
| Variable | Odds Ratio | p Value | 95% Confidence Interval |
|---|---|---|---|
| SNP Model:n = 193 a, p = 0.0011, Pseudo R2 = 0.09 | |||
| Current CD4 T-cells/µL | 1.00 | 0.02 | 1.00–1.00 |
| >500 copies HIV RNA/mL | 1.86 | 0.12 | 0.85–4.07 |
| rs9281523*C | 2.49 | 0.15 | 0.71–8.65 |
| Haplotype Model: No haplotypes independently associated with HIV-SN after correction for current CD4 T-cells/µL and >500 copies of HIV RNA/mL | |||
| Variable | Odds Ratio | p Value | 95% Confidence Interval |
|---|---|---|---|
| SNP Model:n = 71 a, p = 0.0003, Pseudo R2 = 0.23 | |||
| Weight (kg) | 1.04 | 0.04 | 1.00–1.08 |
| History of Tuberculosis | 5.66 | 0.04 | 1.09–29.36 |
| Nadir CD4 T-cells/µL | 0.99 | 0.02 | 0.99–1.00 |
| rs4947324*T | 0.25 | 0.05 | 0.06–1.01 |
| Haplotype Model:n = 71 a, p = 0.0003, Pseudo R2 = 0.22 | |||
| Weight (kg) | 1.04 | 0.02 | 1.01–1.08 |
| History of Tuberculosis | 5.22 | 0.04 | 1.09–24.86 |
| Nadir CD4 T-cells/µL | 0.99 | 0.02 | 0.99–1.00 |
| S1 (Shared Haplotype 1) | 3.21 | 0.07 | 0.93–11.12 |
| Haplotype a | rs2075582 | rs9281523 | rs11796 | rs2523506 | rs2523504 | rs2071594 | rs2071593 | rs2071592 | rs4947324 | rs909253 | rs1041981 | rs1799964 | rs1800629 | HIV-SN b | pc | |||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| +ve (n = 35) | −ve (n = 167) | |||||||||||||||||
| S1 | T | - | T | G | C | C | G | A | C | G | A | T | G | 15 | 43% | 72 | 43% | 0.69 |
| S3 | T | - | A | T | C | G | G | T | C | A | C | C | G | 14 | 40% | 70 | 42% | 0.85 |
| S4 | C | - | A | G | T | G | G | T | C | A | C | T | G | 12 | 34% | 60 | 36% | 0.87 |
| S8 | C | - | A | G | T | G | A | T | C | A | C | T | G | 7 | 20% | 36 | 22% | 0.97 |
| I1 | T | - | T | G | C | C | G | T | C | G | A | T | G | 3 | 9% | 21 | 13% | 0.77 |
| S6 | T | - | A | G | C | G | G | T | C | A | C | T | G | 2 | 6% | 10 | 6% | 0.99 |
| S2 d | T | C | T | G | C | C | G | A | C | G | A | T | A | 5 | 14% | 8 | 5% | 0.02 |
| S7 | T | - | A | G | C | G | G | T | T | A | C | C | G | 2 | 6% | 8 | 5% | 0.66 |
| I2 | C | - | A | G | C | C | G | A | C | G | A | T | G | 0 | 0% | 5 | 3% | 0.99 |
| S5 | T | - | T | G | C | C | G | A | C | G | A | T | A | 1 | 3% | 3 | 2% | 0.51 |
| Haplotype a | rs2075582 | rs9281523 | rs11796 | rs2523506 | rs2523504 | rs2071594 | rs2071593 | rs2071592 | rs4947324 | rs909253 | rs1041981 | rs1799964 | rs1800629 | HIV-SN b | pc | |||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| +ve (n = 29) | −ve (n = 46) | |||||||||||||||||
| S1 d | T | - | T | G | C | C | G | A | C | G | A | T | G | 19 | 66% | 23 | 50% | 0.19 |
| S2 | T | C | T | G | C | C | G | A | C | G | A | T | A | 9 | 31% | 16 | 35% | 0.34 |
| S3 | T | - | A | T | C | G | G | T | C | A | C | C | G | 5 | 17% | 8 | 17% | 0.99 |
| A1 | T | - | A | G | C | G | G | T | T | A | C | T | G | 3 | 10% | 8 | 17% | 0.51 |
| S4 | C | - | A | G | T | G | G | T | C | A | C | T | G | 3 | 10% | 5 | 11% | 0.99 |
| S5 | T | - | T | G | C | C | G | A | C | G | A | T | A | 1 | 3% | 4 | 9% | 0.64 |
| A2 | C | - | A | G | T | G | A | T | T | A | C | T | G | 1 | 3% | 4 | 9% | 0.64 |
| S6 | T | - | A | G | C | G | G | T | C | A | C | T | G | 3 | 10% | 2 | 4% | 0.37 |
| A3 | T | - | T | G | T | C | G | A | C | G | A | T | G | 0 | 0% | 4 | 9% | 0.15 |
| S7 | T | - | A | G | C | G | G | T | T | A | C | C | G | 0 | 0% | 4 | 9% | 0.15 |
| S8 | C | - | A | G | T | G | A | T | C | A | C | T | G | 0 | 0% | 2 | 4% | 0.52 |
| A4 | C | - | A | G | T | G | G | T | C | A | A | T | G | 0 | 0% | 2 | 4% | 0.52 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gaff, J.; Octaviana, F.; Pillay, P.; Ngassa Mbenda, H.G.; Ariyanto, I.A.; Gan, J.A.; Cherry, C.L.; Kamerman, P.; Laws, S.M.; Price, P. TNF-Block Genotypes Influence Susceptibility to HIV-Associated Sensory Neuropathy in Indonesians and South Africans. Int. J. Mol. Sci. 2020, 21, 380. https://doi.org/10.3390/ijms21020380
Gaff J, Octaviana F, Pillay P, Ngassa Mbenda HG, Ariyanto IA, Gan JA, Cherry CL, Kamerman P, Laws SM, Price P. TNF-Block Genotypes Influence Susceptibility to HIV-Associated Sensory Neuropathy in Indonesians and South Africans. International Journal of Molecular Sciences. 2020; 21(2):380. https://doi.org/10.3390/ijms21020380
Chicago/Turabian StyleGaff, Jessica, Fitri Octaviana, Prinisha Pillay, Huguette Gaelle Ngassa Mbenda, Ibnu A. Ariyanto, June Anne Gan, Catherine L. Cherry, Peter Kamerman, Simon M. Laws, and Patricia Price. 2020. "TNF-Block Genotypes Influence Susceptibility to HIV-Associated Sensory Neuropathy in Indonesians and South Africans" International Journal of Molecular Sciences 21, no. 2: 380. https://doi.org/10.3390/ijms21020380
APA StyleGaff, J., Octaviana, F., Pillay, P., Ngassa Mbenda, H. G., Ariyanto, I. A., Gan, J. A., Cherry, C. L., Kamerman, P., Laws, S. M., & Price, P. (2020). TNF-Block Genotypes Influence Susceptibility to HIV-Associated Sensory Neuropathy in Indonesians and South Africans. International Journal of Molecular Sciences, 21(2), 380. https://doi.org/10.3390/ijms21020380






