Effect of 15 BMI-Associated Polymorphisms, Reported for Europeans, across Ethnicities and Degrees of Amerindian Ancestry in Mexican Children
Abstract
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
4.1. Data Analysis
4.2. Association Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| SNP | Single Nucleotide Polymorphism |
| BMI | Body Mass Index |
| AIMs | Ancestry-Informative Markers |
| WHO | World Health Organization |
| AMA | Amerindian Ancestry |
| PCA | Principal Component Analysis |
| GRS | Genetic Risk Score |
References
- Ng, M.; Fleming, T.; Robinson, M.; Thomson, B.; Graetz, N.; Margono, C.; Mullany, E.C.; Biryukov, S.; Abbafati, C.; Abera, S.F.; et al. Global, regional, and national prevalence of overweight and obesity in children and adults during 1980–2013: A systematic analysis for the Global Burden of Disease Study 2013. Lancet 2014, 384, 766–781. [Google Scholar] [CrossRef]
- ENSANUT. Encuesta Nacional de Salud y Nutrición de Medio Camino (National Mid-Way Health and Nutrition Survey) 2016. (ENSANUT MC 2016). Inst. Nac. Salud Públ. 2016, 2016, 151. [Google Scholar] [CrossRef]
- Fernández, J.R.; Pearson, K.E.; Kell, K.P.; Brown, M.M. Genetic admixture and obesity: Recent perspectives and future applications. Hum. Hered. 2013, 75, 98–105. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lu, Y.; Loos, R.J.F. Obesity genomics: Assessing the transferability of susceptibility loci across diverse populations. Genome Med. 2013, 5, 55. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Estrada, A.; Gignoux, C.R.; Fernández-López, J.C.; Zakharia, F.; Sikora, M.; Contreras, A.V.; Acuña-Alonzo, V.; Sandoval, K.; Eng, C.; Romero-Hidalgo, S.; et al. The genetics of Mexico recapitulates Native American substructure and affects biomedical traits. Science 2014, 344, 1280–1285. [Google Scholar] [CrossRef] [PubMed]
- Silva-Zolezzi, I.; Hidalgo-Miranda, A.; Estrada-Gil, J.; Fernandez-Lopez, J.C.; Uribe-Figueroa, L.; Contreras, A.; Balam-Ortiz, E.; del Bosque-Plata, L.; Velazquez-Fernandez, D.; Lara, C.; et al. Analysis of genomic diversity in Mexican Mestizo populations to develop genomic medicine in Mexico. Proc. Natl. Acad. Sci. USA 2009, 106, 8611–8616. [Google Scholar] [CrossRef]
- Villalobos-Comparán, M.; Villamil-Ramírez, H.; Villarreal-Molina, T.; Larrieta-Carrasco, E.; León-Mimila, P.; Romero-Hidalgo, S.; Jacobo-Albavera, L.; Liceaga-Fuentes, A.E.; Campos-Pérez, F.J.; López-Contreras, B.E.; et al. PCSK1 rs6232 is associated with childhood and adult class III obesity in the Mexican population. PLoS ONE 2012, 7, e39037. [Google Scholar] [CrossRef]
- Mejía-Benítez, A.; Klünder-Klünder, M.; Yengo, L.; Meyre, D.; Aradillas, C.; Cruz, E.; Pérez-Luque, E.; Malacara, J.M.; Garay, M.E.; Peralta-Romero, J.; et al. Analysis of the contribution of FTO, NPC1, ENPP1, NEGR1, GNPDA2 and MC4R genes to obesity in Mexican children. BMC Med. Genet. 2013, 14, 21. [Google Scholar] [CrossRef]
- Aradillas-García, C.; Cruz, M.; Pérez-Luque, E.; Garay-Sevilla, M.E.; Malacara, J.M. Obesity is associated with the Arg389Gly ADRB1 but not with the Trp64Arg ADRB3 polymorphism in children from San Luis Potosí and León, México. J. Biomed. Res. 2017, 31, 40–46. [Google Scholar] [CrossRef]
- Abadi, A.; Peralta-Romero, J.; Suarez, F.; Gomez-Zamudio, J.; Burguete-Garcia, A.I.; Cruz, M.; Meyre, D. Assessing the effects of 35 European-derived BMI-associated SNPs in Mexican children: Effects of European BMI SNPs in M … Assessing the Effects of 35 European-Derived BMI-Associated SNPs in Mexican Children. Obesity 2016, 24, 1989–1995. [Google Scholar] [CrossRef]
- Liu, H.Y.; Alyass, A.; Abadi, A.; Peralta-romero, J. Fine-mapping of 98 obesity loci in Mexican children. Int. J. Obes. 2019, 43, 23–32. [Google Scholar] [CrossRef] [PubMed]
- Torres, J.M.; de la Fuente, G.M.; Frías, H. Comportamiento alimentario y obesidad infantil en Sonora, México. Revista Latinoamericana de Ciencias Sociales Niñez y Juventud 2010, 8, 1131–1147. [Google Scholar]
- Robles-Ordaz, M.D.; Gallegos-Aguilar, A.C.; Urquidez-Romero, R.; Diaz-Zavala, R.G.; Lavandera-Torres, M.G.; Esparza-Romero, J. Prevalence of prediabetes and modifiable factors in an ethnic group of Mexico: The Comcáac Project. Publ. Health Nutr. 2018, 21, 333–338. [Google Scholar] [CrossRef] [PubMed]
- Castro-juarez, A.A.; Serna-gutiérrez, A.; Lozoya-villegas, J.F.; Toledo-domínguez, I.D.J.; Díaz-zavala, R.G.; Esparza-romero, J. Prevalence of previous diagnosis of hypertension and associated factors in the Yaqui indigenous of Sonora. Revista Mexicana de Cardiología 2018, 29, 90–97. [Google Scholar]
- Composto, C.; Navarro, M.L. Territorios en Disputa. Despojo Capitalista, Luchas en Defensa de los Bienes Comunes Naturales y Alternativas Emancipatorias para América Latina, Bajo Tierra Ediciones; Cambridge University Press: Cambridge, UK, 2014; p. 452. [Google Scholar] [CrossRef]
- Sandoval Arenas, C.O.; Guilherme, M.; Dietz, G. Displacement and revitalization of the Nahuatl language in the High Mountains of Veracruz, Mexico. Arts Humanit. High. Educ. 2017, 16, 66–81. [Google Scholar] [CrossRef]
- Bonilla, C.; Gutiérrez, G.; Parra, E.J.; Kline, C.; Shriver, M.D. Admixture analysis of a rural population of the state of Guerrero, Mexico. Am. J. Phys. Anthropol. 2005, 128, 861–869. [Google Scholar] [CrossRef]
- Martinez-Marignac, V.L.; Valladares, A.; Cameron, E.; Chan, A.; Perera, A.; Globus-Goldberg, R.; Wacher, N.; Kumate, J.; McKeigue, P.; O’Donnell, D.; et al. Admixture in Mexico City: Implications for admixture mapping of Type 2 diabetes genetic risk factors. Hum. Genet. 2007, 120, 807–819. [Google Scholar] [CrossRef]
- Tang, H.; Jorgenson, E.; Gadde, M.; Kardia, S.L.; Rao, D.C.; Zhu, X.; Schork, N.J.; Hanis, C.L.; Risch, N. Racial admixture and its impact on BMI and blood pressure in African and Mexican Americans. Hum. Genet. 2006, 119, 624–633. [Google Scholar] [CrossRef]
- Klimentidis, Y.C.; Miller, G.F.; Shriver, M.D. The relationship between European genetic admixture and body composition among Hispanics and Native Americans. Am. J. Hum. Biol. 2009, 21, 377–382. [Google Scholar] [CrossRef]
- Dellasega, C.; Añel-Tiangco, R.M.; Gabbay, R.A. How patients with type 2 diabetes mellitus respond to motivational interviewing. Diabetes Res. Clin. Pract. 2012, 95, 37–41. [Google Scholar] [CrossRef]
- Costa-Urrutia, P.; Álvarez-Fariña, R.; Abud, C.; Franco-Trecu, V.; Esparza-Romero, J.; López-Morales, C.M.; Rodríguez-Arellano, M.E.; Leal, J.V.; Colistro, V.; Granados, J. Effect of multi-component school-based program on body mass index, cardiovascular and diabetes risks in a multi-ethnic study. BMC Pediatr. 2019, 19, 401. [Google Scholar] [CrossRef]
- Bradfield, J.P.; Taal, H.R.; Timpson, N.J.; Scherag, A.; Lecoeur, C.; Warrington, N.M.; Hypponen, E.; Holst, C.; Valcarcel, B.; Thiering, E.; et al. A genome-wide association meta-analysis identifies new childhood obesity loci. Nat. Genet. 2012, 44, 526–531. [Google Scholar] [CrossRef][Green Version]
- Felix, J.F.; Bradfield, J.P.; Monnereau, C.; Van Der Valk, R.J.; Stergiakouli, E.; Chesi, A.; Gaillard, R.; Feenstra, B.; Thiering, E.; Kreiner-Møller, E.; et al. Genome-wide association analysis identifies three new susceptibility loci for childhood body mass index. Hum. Mol. Genet. 2016, 25, 389–403. [Google Scholar] [CrossRef]
- Wang, J.; Mei, H.; Chen, W.; Jiang, Y.; Sun, W.; Li, F.; Fu, Q.; Jiang, F. Study of eight GWAS-identified common variants for association with obesity-related indices in Chinese children at puberty. Int. J. Obes. 2012, 36, 542–547. [Google Scholar] [CrossRef]
- Warrington, N.M.; Howe, L.D.; Paternoster, L.; Kaakinen, M.; Herrala, S.; Huikari, V.; Wu, Y.Y.; Kemp, J.P.; Timpson, N.J.; Pourcain, B.S.; et al. A genome-wide association study of body mass index across early life and childhood. Int. J. Epidemiol. 2015, 44, 700–712. [Google Scholar] [CrossRef]
- Wang, J.; Lin, M.; Crenshaw, A.; Hutchinson, A.; Hicks, B.; Yeager, M.; Berndt, S.; Huang, W.Y.; Hayes, R.B.; Chanock, S.J.; et al. High-throughput single nucleotide polymorphism genotyping using nanofluidic Dynamic Arrays. BMC Genom. 2009, 10, 561. [Google Scholar] [CrossRef]
- De Onis, M.; Onyango, A.W.; Borghi, E.; Siyam, A.; Nishida, C.; Siekmann, J. Development of a WHO growth reference for school-aged children and adolescents. Bull. World Health Organ. 2007, 85, 812–819. [Google Scholar] [CrossRef]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.; Bender, D.; Maller, J.; Sklar, P.; De Bakker, P.I.; Daly, M.J.; et al. PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef]
- Patterson, N.; Price, A.L.; Reich, D. Population structure and eigenanalysis. PLoS Genet. 2006, 2, 2074–2093. [Google Scholar] [CrossRef]
- Alexander, D.H.; Novembre, J.; Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 2009, 19, 1655–1664. [Google Scholar] [CrossRef]
- Speliotes, E.K.; Willer, C.J.; Berndt, S.I.; Monda, K.L.; Thorleifsson, G.; Jackson, A.U.; Allen, H.L.; Lindgren, C.M.; Luan, J.A.; Mägi, R.; et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat. Genet. 2010, 42, 937–948. [Google Scholar] [CrossRef]
- Costa-Urrutia, P.; Abud, C.; Franco-Trecu, V.; Colistro, V.; Rodríguez-Arellano, M.E.; Vázquez-Pérez, J.; Granados, J.; Seelaender, M. Genetic Obesity Risk and Attenuation Effect of Physical Fitness in Mexican-Mestizo Population: A Case-Control Study. Ann. Hum. Genet. 2017, 81, 106–116. [Google Scholar] [CrossRef]
- He, M.; Cornelis, M.C.; Franks, P.W.; Zhang, C.; Hu, F.B.; Qi, L. Obesity genotype score and cardiovascular risk in women with type 2 diabetes mellitus. Arterioscler. Thromb. Vasc. Biol. 2010, 30, 327–332. [Google Scholar] [CrossRef]
- Anderson, D.R. Multimodel Inference Understanding AIC and BIC in Model Selection. Sociol. Methods Res. 2004, 33, 261–304. [Google Scholar] [CrossRef]
| Ethnic Group | Sex (Number) | BMI | Prevalence (%) | |
|---|---|---|---|---|
| Mean (SD) | Overweight | Obesity | ||
| NMM | Girls (n = 100) | 19.0 (4.6) | 17.0 | 29.0 |
| Boys (n = 78) | 18.9 (4.1) | 20.5 | 28.2 | |
| Yaquis | Girls (n = 51) | 17.3 (4.5) | 9.8 | 7.8 |
| Boys (n = 72) | 17.3 (4.2) | 20.8 | 9.7 | |
| Seris | Girls (n = 41) | 18.2 (3.3) | 14.6 | 17.1 |
| Boys (n = 27) | 19.2 (5.6) | 14.8 | 25.9 | |
| CMM, indigenous school | Girls (n = 531) | 17.2 (3.0) | 11.9 | 15.4 |
| Boys (n = 480) | 17.2 (3.1) | 12.5 | 15.4 | |
| CMM, regular school | Girls (n = 3761) | 17.6 (3.2) | 14.6 | 20.3 |
| Boys (n = 3773) | 17.7 (3.2) | 16.4 | 19.3 | |
| Gene | Chr | SNP | AA | NMM/Yaquis (n = 301) | Seris (n = 68) | p | CM (n = 8545) | p | |
|---|---|---|---|---|---|---|---|---|---|
| β (SE) | p | β (SE) | β (SE) | ||||||
| SEC16B | 1 | rs543874 | G | 0.21 (0.14) | 0.15 | −0.29 (0.25) | 0.24 | 0.10 (0.02) | 3.4 × 10−8 |
| OLFM4 | 13 | rs12429545 | G | −0.20 (0.11) | 0.08 | 0.02 (0.25) | 0.94 | −0.07 (0.01) | 1.2 × 10−6 |
| FTO | 16 | rs9939609 | A | 0.03 (0.13) | 0.79 | 0.34 (0.36) | 0.35 | 0.11 (0.02) | 2.4 × 10−6 |
| MC4R | 18 | rs6567160 | C | −0.03 (0.20) | 0.87 | −0.24 (0.51) | 0.64 | 0.13 (0.03) | 4.5 × 10−5 |
| GNPDA2 | 4 | rs13130484 | T | −0.08 (0.11) | 0.46 | 0.12 (0.18) | 0.53 | 0.06 (0.01) | 3.4 × 10−4 |
| OLFM4 | 13 | rs9568856 | G | 0.14 (0.10) | 0.19 | 0.05 (0.23) | 0.82 | −0.05 (0.01) | 1.0 × 10−3 |
| FAIM2 | 5 | rs7132908 | A | −0.24 (0.14) | 0.08 | 0.12 (0.37) | 0.75 | 0.06 (0.02) | 3.0 × 10−3 |
| FAM120AOS | 12 | rs944990 | A | −0.08 (0.12) | 0.48 | 0.32 (0.25) | 0.21 | 0.05 (0.01) | 0.01 |
| LMX1B | 9 | rs3829849 | A | 0.05 (0.13) | 0.69 | 0.02 (0.34) | 0.99 | 0.06 (0.02) | 0.02 |
| HOXB5 | 9 | rs9299 | A | 0.02 (0.10) | 0.86 | −0.36 (0.21) | 0.10 | 0.03 (0.01) | 0.03 |
| ADAM23 | 17 | rs13387838 | G | 0.01 (0.35) | 0.99 | M | −0.23 (0.01) | 0.04 | |
| ELP3 | 4 | rs13253111 | G | −0.13 (0.10) | 0.20 | −0.07 (0.20) | 0.73 | −0.02 (0.01) | 0.12 |
| RAB27B | 2 | rs8092503 | G | 0.05 (0.11) | 0.66 | 0.13 (0.28) | 0.66 | 0.02 (0.02) | 0.17 |
| GPR61 | 4 | rs7550711 | T | −0.31 (0.65) | 0.64 | M | 0.15 (0.012) | 0.20 | |
| TNNI3K | 8 | rs12041852 | A | 0.07 (0.11) | 0.53 | 0.67 (0.42) | 0.12 | −0.01 (0.01) | 0.54 |
| NMM/Yaquis (n = 301) | Seris (n = 68) | CM (n = 8545) | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Gene | Chr | SNP | AA | OR (CI) | p | OR (CI) | p | OR (CI) | p |
| SEC16B | 1 | rs543874 | G | 1.79 (1.04, 3.08) | 0.04 | 0.31 (0.08, 1.24) | 0.09 | 1.26 (1.13, 1.32) | 1.0 × 10−5 |
| OLFM4 | 13 | rs12429545 | G | 0.84 (0.53, 1.34) | 0.47 | 0.44 (0.13, 1.65) | 0.21 | 0.85 (0.78, 0.93) | 2.2 × 10−4 |
| FTO | 16 | rs9939609 | A | 0.93 (0.55, 1.57) | 0.78 | 2.17 (0.43, 10.87) | 0.34 | 1.26 (1.12 1.42) | 2.2 × 10−4 |
| MC4R | 18 | rs6567160 | C | 1.73 (0.79, 3.81) | 0.17 | 1.00 (0.08, 11.99) | 1.00 | 1.25 (1.06, 1.48) | 8.0 × 10−3 |
| GNPDA2 | 4 | rs13130484 | T | 0.86 (0.54, 1.36) | 0.52 | 1.43 (0.63, 3.25) | 0.40 | 1.12 (1.01, 1.21) | 0.03 |
| OLFM4 | 13 | rs9568856 | G | 1.04 (0.67, 1.59) | 0.87 | 0.78 (0.28, 2.11) | 0.62 | 0.90 (0.82, 0.98) | 0.01 |
| FAIM2 | 12 | rs7132908 | A | 0.91 (0.52, 1.57) | 0.73 | 1.41 (0.32, 6.08) | 0.65 | 1.048 (0.93, 1.18) | 0.46 |
| FAM120AOS | 9 | rs944990 | A | 0.96 (0.57, 1.59) | 0.78 | 0.42 (0.12, 1.46) | 0.17 | 1.076 (0.97, 1.19) | 0.17 |
| LMX1B | 9 | rs3829849 | A | 0.92 (0.53, 1.58) | 0.51 | 0.73 (0.15, 3.68) | 0.71 | 1.18 (1.02, 1.37) | 0.03 |
| ADAM23 | 2 | rs13387838 | A | 1.52 (0.44, 5.27) | 0.78 | M | 1.45 (0.81, 2.58) | 0.21 | |
| HOXB5 | 17 | rs9299 | G | 0.94 (0.60, 1.46) | 0.77 | 0.54 (0.20, 1.43) | 0.21 | 0.94 (0.86, 1.02) | 0.14 |
| ELP3 | 8 | rs13253111 | G | 0.61 (0.39, 0.94) | 0.33 | 0.86 (0.36, 2.07) | 0.74 | 0.94 (0.86, 1.03) | 0.20 |
| RAB27B | 18 | rs8092503 | G | 1.24 (0.81, 1.90) | 1.00 | 0.42 (0.08, 2.16) | 0.30 | 1.04 (0.96, 1.14) | 0.33 |
| GPR61 | 1 | rs7550711 | T | VLF | M | 1.03 (0.52, 2.04) | 0.93 | ||
| TNNI3K | 1 | rs12041852 | A | 1.18 (0.74, 1.88) | 0.48 | 4.35 (0.77, 24.43) | 0.09 | 0.97 (0.88, 1.07) | 0.54 |
| Variables | β | SE | p |
|---|---|---|---|
| Intercept | −0.01 | 0.01 | 0.26 |
| GRS | 0.11 | 0.01 | 0.1 × 10−16 |
| AMA | −0.05 | 0.01 | 6.8 × 10−7 |
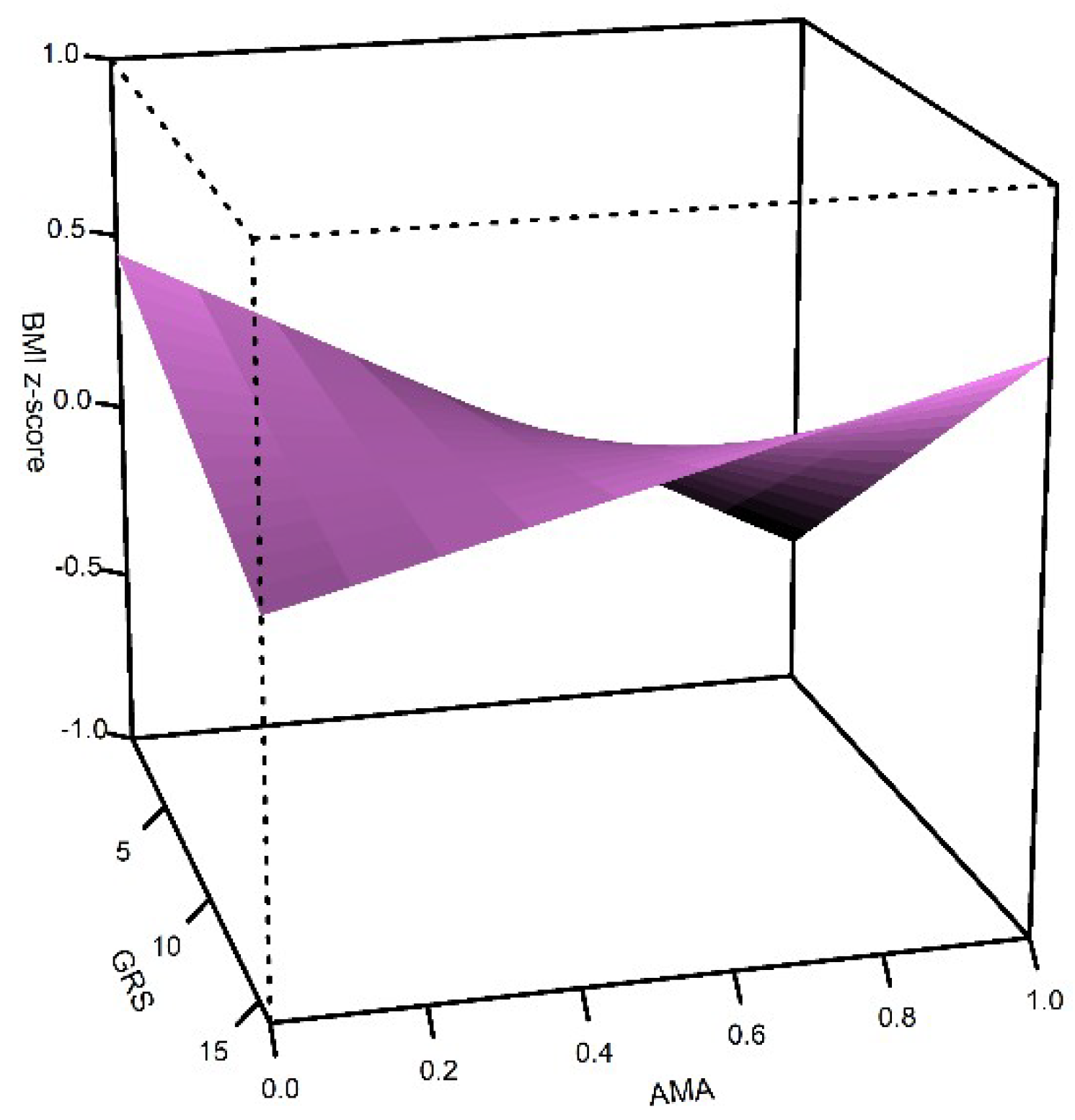
| GRS*AMA | 0.03 | 0.01 | 6.0 × 10−3 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Costa-Urrutia, P.; Abud, C.; Franco-Trecu, V.; Colistro, V.; Rodríguez-Arellano, M.E.; Alvarez-Fariña, R.; Acuña Alonso, V.; Bertoni, B.; Granados, J. Effect of 15 BMI-Associated Polymorphisms, Reported for Europeans, across Ethnicities and Degrees of Amerindian Ancestry in Mexican Children. Int. J. Mol. Sci. 2020, 21, 374. https://doi.org/10.3390/ijms21020374
Costa-Urrutia P, Abud C, Franco-Trecu V, Colistro V, Rodríguez-Arellano ME, Alvarez-Fariña R, Acuña Alonso V, Bertoni B, Granados J. Effect of 15 BMI-Associated Polymorphisms, Reported for Europeans, across Ethnicities and Degrees of Amerindian Ancestry in Mexican Children. International Journal of Molecular Sciences. 2020; 21(2):374. https://doi.org/10.3390/ijms21020374
Chicago/Turabian StyleCosta-Urrutia, Paula, Carolina Abud, Valentina Franco-Trecu, Valentina Colistro, Martha Eunice Rodríguez-Arellano, Rafael Alvarez-Fariña, Víctor Acuña Alonso, Bernardo Bertoni, and Julio Granados. 2020. "Effect of 15 BMI-Associated Polymorphisms, Reported for Europeans, across Ethnicities and Degrees of Amerindian Ancestry in Mexican Children" International Journal of Molecular Sciences 21, no. 2: 374. https://doi.org/10.3390/ijms21020374
APA StyleCosta-Urrutia, P., Abud, C., Franco-Trecu, V., Colistro, V., Rodríguez-Arellano, M. E., Alvarez-Fariña, R., Acuña Alonso, V., Bertoni, B., & Granados, J. (2020). Effect of 15 BMI-Associated Polymorphisms, Reported for Europeans, across Ethnicities and Degrees of Amerindian Ancestry in Mexican Children. International Journal of Molecular Sciences, 21(2), 374. https://doi.org/10.3390/ijms21020374





