Δ9-Tetrahydrocannabinol Prevents Mortality from Acute Respiratory Distress Syndrome through the Induction of Apoptosis in Immune Cells, Leading to Cytokine Storm Suppression
Abstract
1. Introduction
2. Results
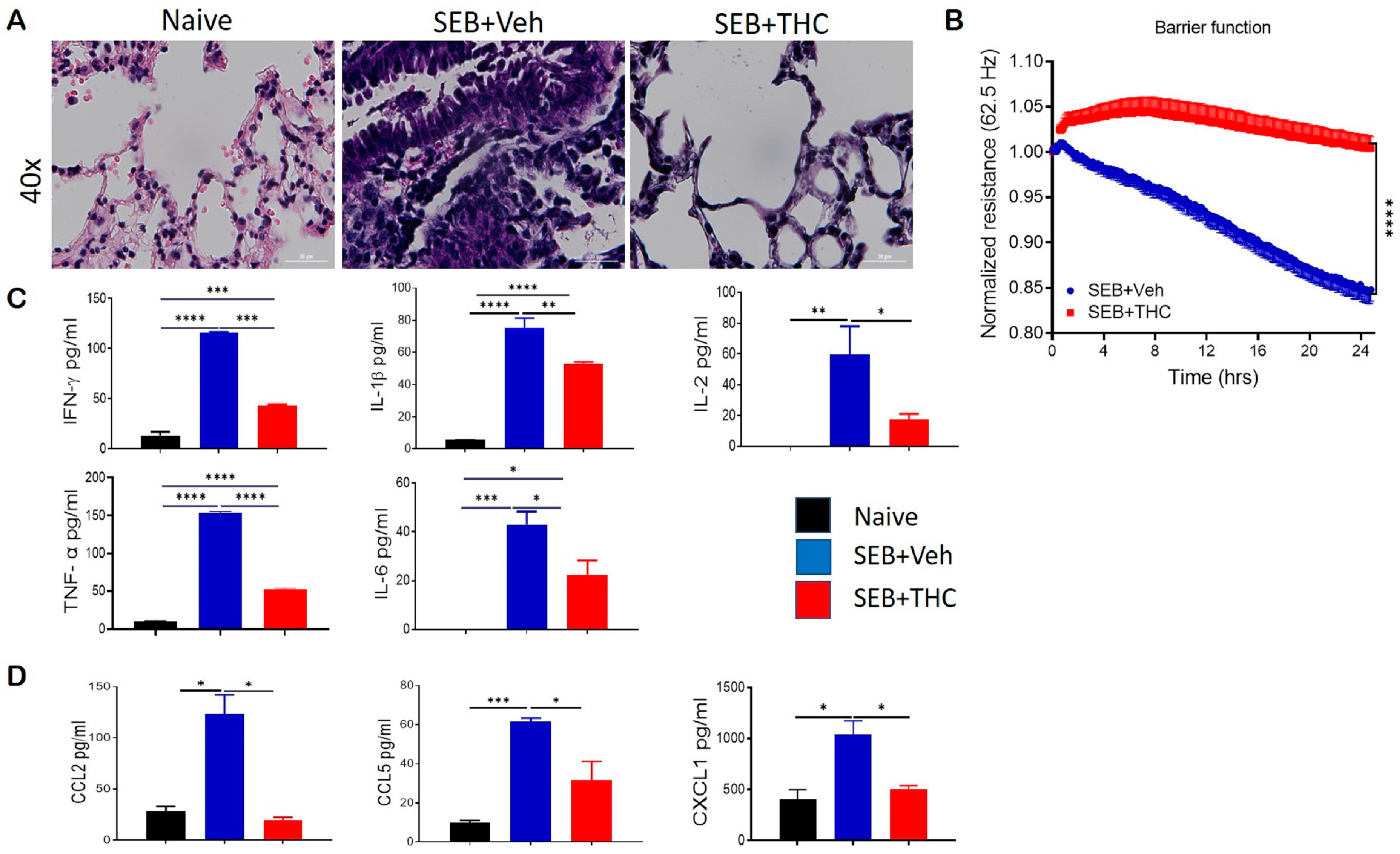
2.1. THC Treatment Prevents SEB-Induced Pulmonary Damage via a Reduction in Infiltrating Immune Cells
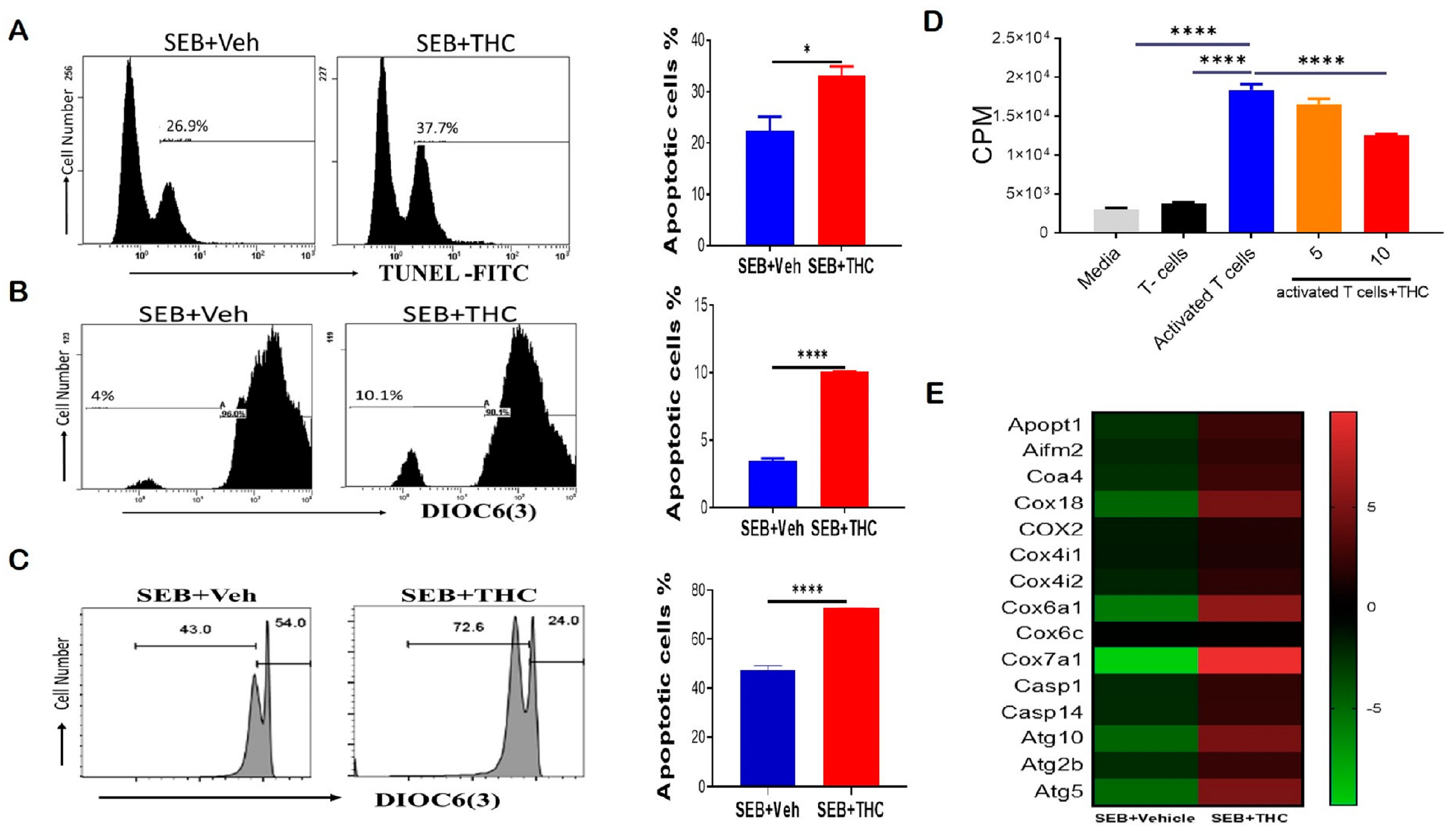
2.2. THC Induces Apoptosis through the Mitochondrial Pathway
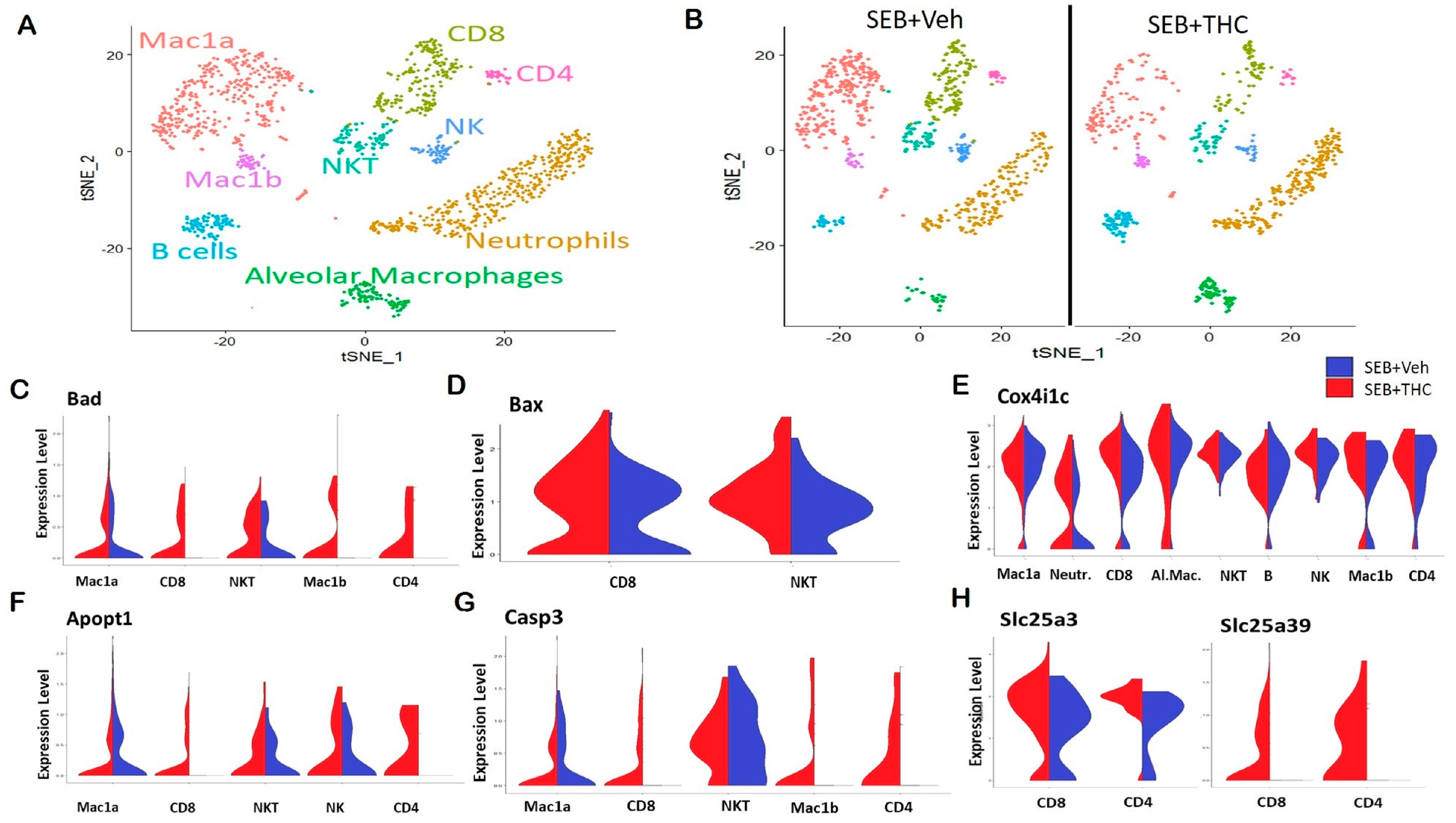
2.3. Pulmonary scRNA-seq Shows a Reduction in Inflammatory Components in SEB+THC-Treated Mice
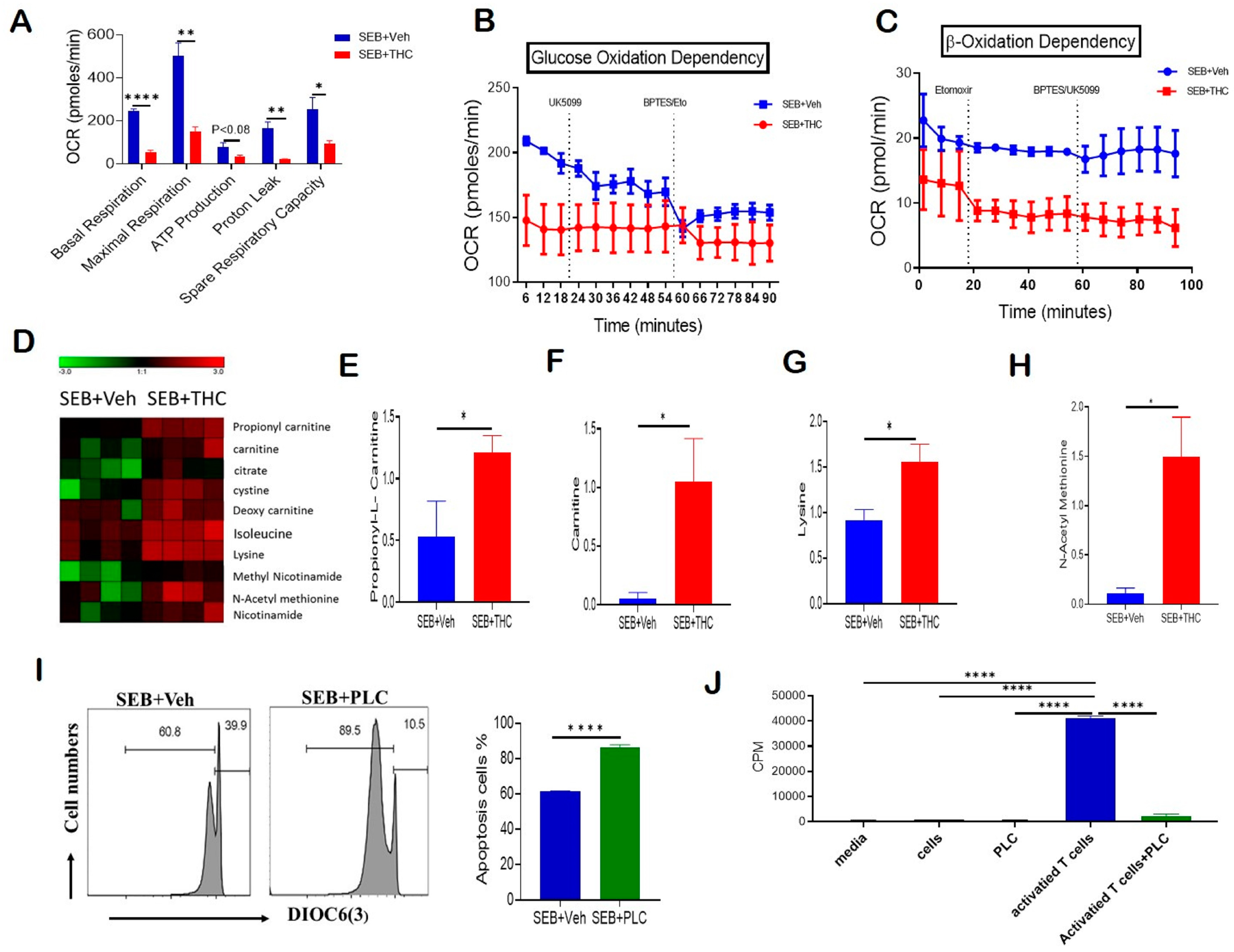
2.4. THC-Mediated Amelioration of ARDS Is Associated with Altered T Cell Metabolism
2.5. Metabolomic Profiling
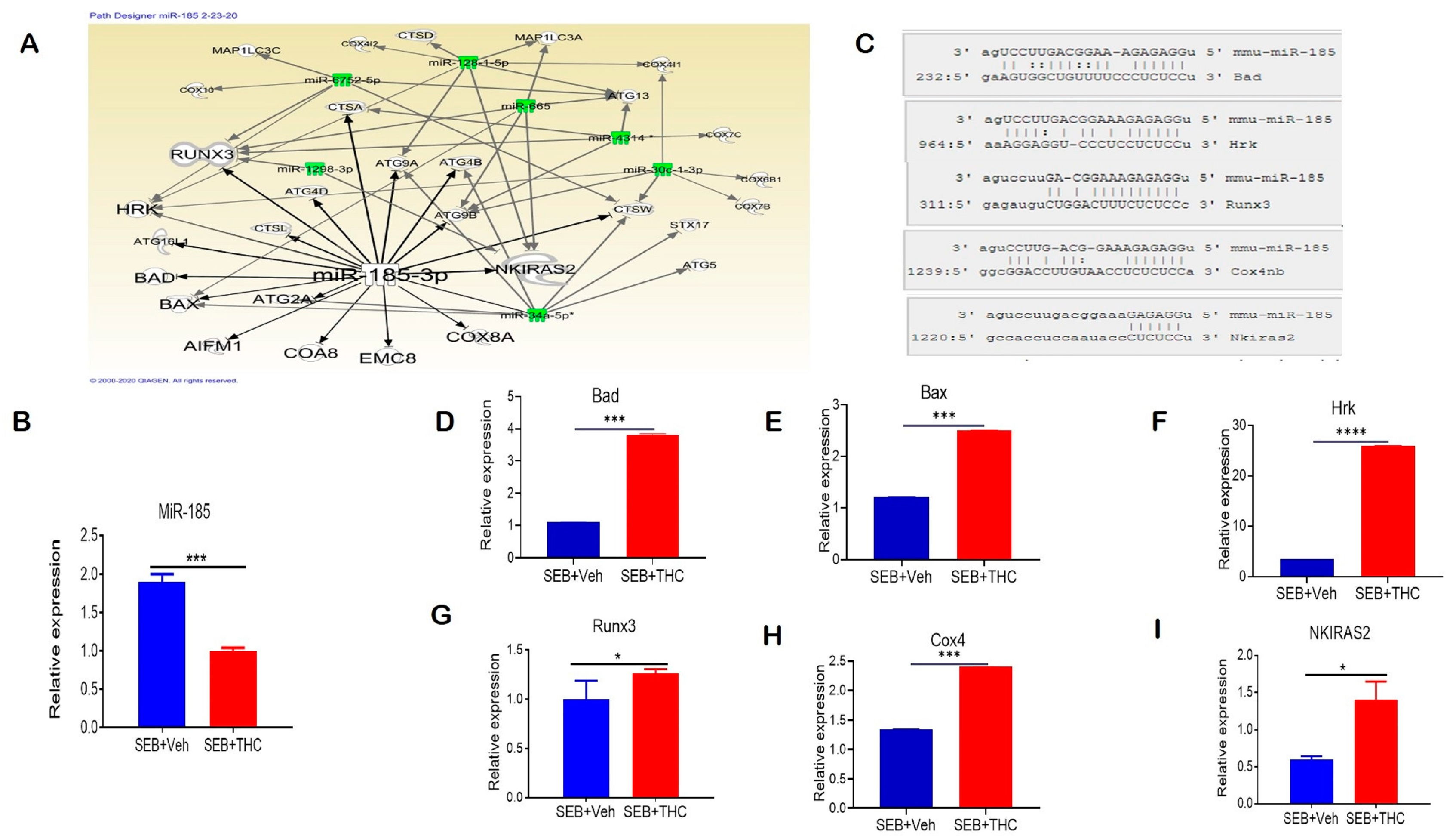
2.6. THC-Mediated Apoptosis of SEB-Activated T Cells Is Epigenetically Regulated
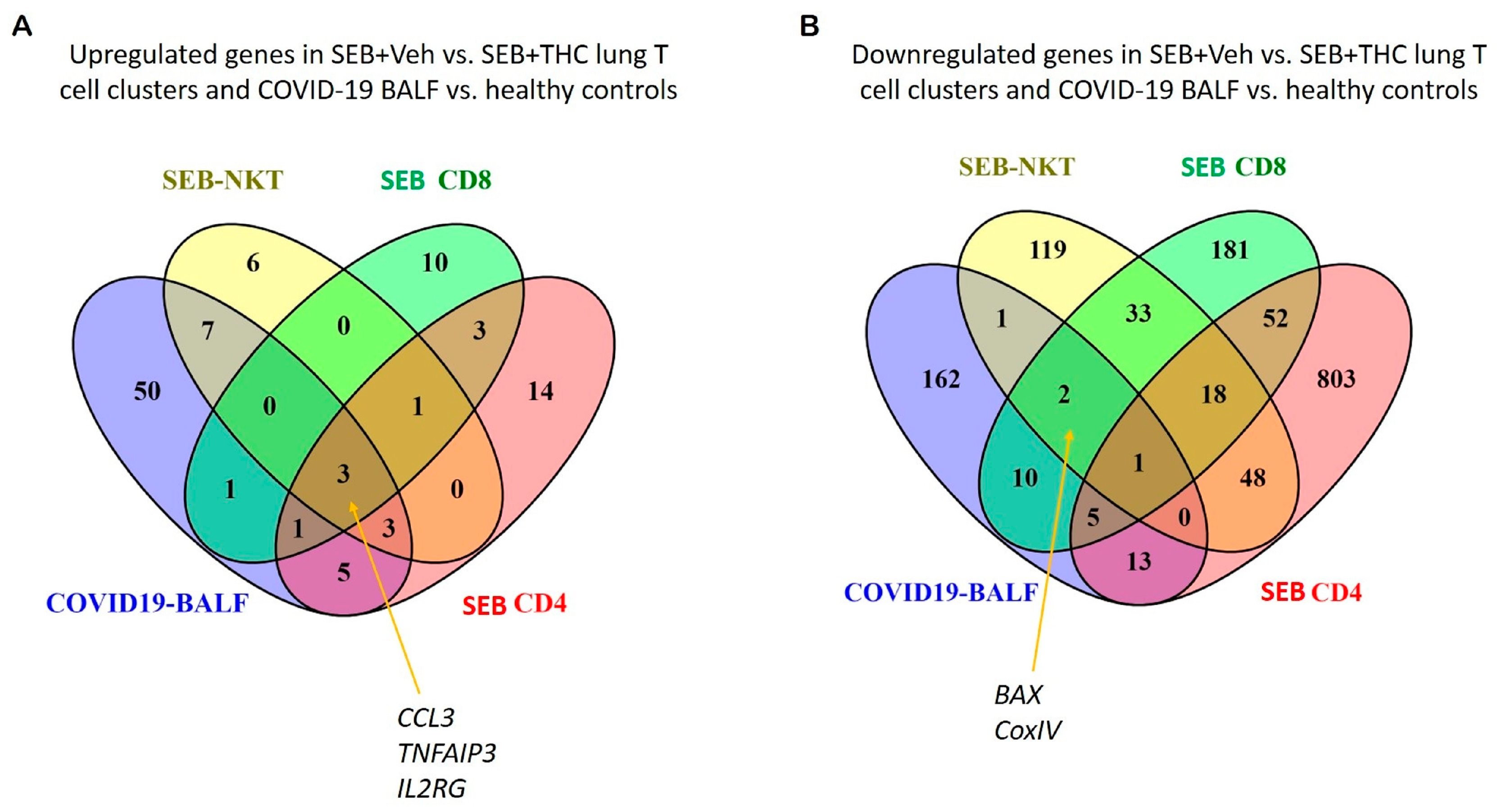
2.7. Meta-Analysis of Genes Altered in SEB-Induced ARDS in Mice vs. COVID-19 BALF Human Datasets
3. Discussion
4. Material and Methods
4.1. Mice Grouping and Housing
4.2. Exposure of Mice with SEB and THC
4.3. Chemicals and Regents
4.4. Histopathology of Lung Tissues
4.5. Analysis of Cytokines×
4.6. Barrier Function Test
4.7. TUNEL Assay
4.8. Mitochondrial Membrane Potential
4.9. Effect of THC-Induced MDSCs on T Cell Proliferation In Vitro
4.10. Purification of CD3+ T Cells
4.11. Metabolic Assays
4.12. Metabolomic Studies
4.13. Separation of Glycolysis, TCA, and Pentose Pathway Metabolites
4.14. Single Cell RNA-Seq and Analysis
4.15. Quantitative Real-Time PCR (qRT-PCR) Validation of Selected miRs and Associated Genes
4.16. Correlation of scRNA-seq Data with Human COVID-19 Datasets
4.17. Data and Analysis
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| 2-DG | 2-Deoxy glucose |
| ALI | acute lung injury |
| ARDS | Acute Respiratory Distress Syndrome |
| Apopt1 or Coa8 | apoptogenic protein 1 or cytochrome c oxidase assembly factor 8 |
| Atg | autophagy |
| BALF | broncho-alveolar lavage fluid |
| Bad | Bcl2-associated agonist of cell death |
| Bax | BCL2 Associated X or Apoptosis Regulator |
| Casp | caspases |
| CDC | Centers for Disease Control |
| Cox | cytochrome c oxidases |
| Aifm2 | apoptosis-inducing factor 2-homologous mitochondrion-associated inducer of death |
| COVID-19 | Corona virus disease – 2019 |
| ECIS | Electric cell-substrate impedance sensing |
| ECAR | extra cellular acidification rate |
| H&E | hematoxylin and eosin |
| Hrk | Activator of apoptosis hara-kiri |
| MDSCs | myeloid-derived suppressor cells |
| MNCs | mononuclear cells |
| miR | miRNA |
| MTA | mouse transcriptome analysis |
| Nkiras2 | NF-κB inhibitor interacting Ras-like 2 |
| OCR | oxygen consumption rate |
| PTSD | post-traumatic stress disorder |
| PLC | propionyl carnitine |
| Rot/AA | Rotenone/antimycin A |
| Runx3 | Runt-related transcription factor 3 |
| scRNA-seq | single-cell RNA sequence |
| SEB | Staphylococcal enterotoxin-B |
| TAC | transcriptome analysis console |
| THC | Δ9-Tetrahydrocannabinol |
| TUNEL | terminal deoxynucleotidyl transferase (TdT)-mediated dUTP nick-end labeling |
References
- Krakauer, T. Staphylococcal Superantigens: Pyrogenic Toxins Induce Toxic Shock. Toxins 2019, 11, 178. [Google Scholar] [CrossRef] [PubMed]
- Alghetaa, H.; Mohammed, A.; Sultan, M.; Busbee, P.; Murphy, A.; Chatterjee, S.; Nagarkatti, M.; Nagarkatti, P. Resveratrol protects mice against SEB-induced acute lung injury and mortality by miR-193a modulation that targets TGF-β signalling. J. Cell. Mol. Med. 2018, 22, 2644–2655. [Google Scholar] [CrossRef] [PubMed]
- Mohammed, A.; Alghetaa, H.; Sultan, M.; Singh, N.P.; Nagarkatti, P.; Nagarkatti, M. Administration of Δ9-Tetrahydrocannabinol (THC) Post-Staphylococcal Enterotoxin B Exposure Protects Mice from Acute Respiratory Distress Syndrome and Toxicity. Front. Pharmacol. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Fan, E.; Brodie, D.; Slutsky, A.S. Acute Respiratory Distress Syndrome: Advances in Diagnosis and Treatment. JAMA 2018, 319, 698–710. [Google Scholar] [CrossRef] [PubMed]
- Henry, B.M.; Lippi, G. Poor survival with extracorporeal membrane oxygenation in acute respiratory distress syndrome (ARDS) due to coronavirus disease 2019 (COVID-19): Pooled analysis of early reports. J. Crit. Care 2020, 58, 27–28. [Google Scholar] [CrossRef]
- Channappanavar, R.; Perlman, S. Pathogenic human coronavirus infections: Causes and consequences of cytokine storm and immunopathology. Semin. Immunopathol. 2017, 39, 529–539. [Google Scholar] [CrossRef]
- Watson, S.J. Marijuana and Medicine: Assessing the Science Base: A Summary of the 1999 Institute of Medicine Report. Arch. Gen. Psychiatry 2000, 57, 547–552. [Google Scholar] [CrossRef]
- Rao, R.; Nagarkatti, P.S.; Nagarkatti, M. Delta(9) Tetrahydrocannabinol attenuates Staphylococcal enterotoxin B-induced inflammatory lung injury and prevents mortality in mice by modulation of miR-17-92 cluster and induction of T-regulatory cells. Br. J. Pharmacol. 2015, 172, 1792–1806. [Google Scholar] [CrossRef]
- Elmore, S.A. Apoptosis: A Review of Programmed Cell Death. Toxicol. Pathol. 2007, 35, 495–516. [Google Scholar] [CrossRef]
- McKallip, R.J.; Lombard, C.; Martin, B.R.; Nagarkatti, M.; Nagarkatti, P.S. Delta(9)-tetrahydrocannabinol-induced apoptosis in the thymus and spleen as a mechanism of immunosuppression in vitro and in vivo. J. Pharmacol. Exp. Ther. 2002, 302, 451–465. [Google Scholar] [CrossRef]
- Rieder, S.A.; Chauhan, A.; Singh, U.; Nagarkatti, M.; Nagarkatti, P. Cannabinoid-induced apoptosis in immune cells as a pathway to immunosuppression. Immunobiology 2010, 215, 598–605. [Google Scholar] [CrossRef]
- Zhu, W.; Friedman, H.; Klein, T.W. Delta9-tetrahydrocannabinol induces apoptosis in macrophages and lymphocytes: Involvement of Bcl-2 and caspase-1. J. Pharmacol. Exp. Ther. 1998, 286, 1103–1109. [Google Scholar] [PubMed]
- Lombard, C.; Nagarkatti, M.; Nagarkatti, P. Targeting cannabinoid receptors to treat leukemia: Role of cross-talk between extrinsic and intrinsic pathways in Δ9-tetrahydrocannabinol (THC)-induced apoptosis of Jurkat cells. Leuk. Res. 2005, 29, 915–922. [Google Scholar] [CrossRef] [PubMed]
- Papalexi, E.; Satija, R. Single-cell RNA sequencing to explore immune cell heterogeneity. Nat. Rev. Immunol. 2018, 18, 35–45. [Google Scholar] [CrossRef] [PubMed]
- Catalanotto, C.; Cogoni, C.; Zardo, G. MicroRNA in Control of Gene Expression: An Overview of Nuclear Functions. Int. J. Mol. Sci. 2016, 17, 1712. [Google Scholar] [CrossRef]
- Al-Ghezi, Z.Z.; Miranda, K.; Nagarkatti, M.; Nagarkatti, P. Combination of Cannabinoids, Δ9- Tetrahydrocannabinol and Cannabidiol, Ameliorates Experimental Multiple Sclerosis by Suppressing Neuroinflammation Through Regulation of miRNA-Mediated Signaling Pathways. Front. Immunol. 2019, 10. [Google Scholar] [CrossRef]
- Hegde, V.L.; Tomar, S.; Jackson, A.; Rao, R.; Yang, X.; Singh, U.P.; Singh, N.P.; Nagarkatti, P.S.; Nagarkatti, M. Distinct microRNA expression profile and targeted biological pathways in functional myeloid-derived suppressor cells induced by Delta9-tetrahydrocannabinol in vivo: Regulation of CCAAT/enhancer-binding protein alpha by microRNA-690. J. Biol. Chem. 2013, 288, 36810–36826. [Google Scholar] [CrossRef]
- Huzella, L.M.; Buckley, M.J.; Alves, D.A.; Stiles, B.G.; Krakauer, T. Central roles for IL-2 and MCP-1 following intranasal exposure to SEB: A new mouse model. Res. Vet. Sci. 2009, 86, 241–247. [Google Scholar] [CrossRef]
- Mohammed, A.; Alghetaa, H.; Zhou, J.; Chatterjee, S.; Nagarkatti, P.; Nagarkatti, M. Protective Effects of Delta9-Tetrahydrocannabinol Against Enterotoxin-induced Acute Respiratory Distress Syndrome is Mediated by Modulation of Microbiota. Br. J. Pharmacol. 2020. [Google Scholar] [CrossRef]
- Orlandi, A.; Francesconi, A.; Marcellini, M.; Di Lascio, A.; Spagnoli, L.G. Propionyl-L-carnitine reduces proliferation and potentiates Bax-related apoptosis of aortic intimal smooth muscle cells by modulating nuclear factor-kappaB activity. J. Biol. Chem. 2007, 282, 4932–4942. [Google Scholar] [CrossRef]
- Dushianthan, A.; Grocott, M.; Postle, A.D.; Cusack, R. Acute respiratory distress syndrome and acute lung injury. Postgrad. Med. J. 2011, 87, 612–622. [Google Scholar] [CrossRef] [PubMed]
- Berkowitz, D.M.; Danai, P.A.; Eaton, S.; Moss, M.; Martin, G.S. Alcohol Abuse Enhances Pulmonary Edema in Acute Respiratory Distress Syndrome. Alcohol. Clin. Exp. Res. 2009, 33, 1690–1696. [Google Scholar] [CrossRef]
- Gajic, O.; Dabbagh, O.; Park, P.K.; Adesanya, A.; Chang, S.Y.; Hou, P.; Anderson, H., III; Hoth, J.J.; Mikkelsen, M.E.; Gentile, N.T.; et al. Early identification of patients at risk of acute lung injury: Evaluation of lung injury prediction score in a multicenter cohort study. Am. J. Respir. Crit. Care Med. 2011, 183, 462–470. [Google Scholar] [CrossRef] [PubMed]
- Luhr, O.R.; Antonsen, K.; Karlsson, M.; Aardal, S.; Thorsteinsson, A.; Frostell, C.G.; Bonde, J. Incidence and Mortality after Acute Respiratory Failure and Acute Respiratory Distress Syndrome in Sweden, Denmark, and Iceland. Am. J. Respir. Crit. Care Med. 1999, 159, 1849–1861. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Barnes, K.C. Recent advances in genetic predisposition to clinical acute lung injury. Am. J. Physiol. Cell. Mol. Physiol. 2009, 296, L713–L725. [Google Scholar] [CrossRef] [PubMed]
- Davydow, D.S.; Desai, S.V.; Needham, D.M.; Bienvenu, O.J. Psychiatric Morbidity in Survivors of the Acute Respiratory Distress Syndrome: A Systematic Review. Psychosom. Med. 2008, 70, 512–519. [Google Scholar] [CrossRef]
- Herridge, M.; Cheung, A.M.; Tansey, C.M.; Matte-Martyn, A.; Diaz-Granados, N.; Al-Saidi, F.; Cooper, A.B.; Guest, C.B.; Mazer, C.D.; Mehta, S.; et al. One-Year Outcomes in Survivors of the Acute Respiratory Distress Syndrome. N. Engl. J. Med. 2003, 348, 683–693. [Google Scholar] [CrossRef]
- Hopkins, R.O.; Weaver, L.K.; Collingridge, D.; Parkinson, R.B.; Chan, K.J.; Orme, J.F., Jr. Two-year cognitive, emotional, and quality-of-life outcomes in acute respiratory distress syndrome. Am. J. Respir. Crit. Care Med. 2005, 171, 340–347. [Google Scholar] [CrossRef]
- Rubenfeld, G.D.; Caldwell, E.; Peabody, E.; Weaver, J.; Martin, D.P.; Neff, M.; Stern, E.J.; Hudson, L.D. Incidence and Outcomes of Acute Lung Injury. N. Engl. J. Med. 2005, 353, 1685–1693. [Google Scholar] [CrossRef]
- Rodríguez-Morales, A.J.; Cardona-Ospina, J.A.; Gutiérrez-Ocampo, E.; Villamizar-Peña, R.; Holguin-Rivera, Y.; Escalera-Antezana, J.P.; Alvarado-Arnez, L.E.; Bonilla-Aldana, D.K.; Franco-Paredes, C.; Henao-Martinez, A.F.; et al. Clinical, laboratory and imaging features of COVID-19: A systematic review and meta-analysis. Travel Med. Infect. Dis. 2020, 34, 101623. [Google Scholar] [CrossRef]
- Zhang, J.; Zhou, L.; Yang, Y.; Peng, W.; Wang, W.; Chen, X. Therapeutic and triage strategies for 2019 novel coronavirus disease in fever clinics. Lancet Respir. Med. 2020, 8, e11–e12. [Google Scholar] [CrossRef]
- Hanada, S.; Pirzadeh, M.; Carver, K.Y.; Deng, J.C. Respiratory Viral Infection-Induced Microbiome Alterations and Secondary Bacterial Pneumonia. Front. Immunol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Madsen, L.J.M. Toxins as Weapons of Mass Destruction: A Comparison and Contrast With Biological-Warfare and Chemical-Warfare Agents. Clin. Lab. Med. 2001, 21, 593–606. [Google Scholar] [CrossRef]
- Saeed, A.I.; Rieder, S.A.; Price, R.L.; Barker, J.; Nagarkatti, P.; Nagarkatti, M. Acute lung injury induced by Staphylococcal enterotoxin B: Disruption of terminal vessels as a mechanism of induction of vascular leak. Microsc. Microanal. 2012, 18, 445–452. [Google Scholar] [CrossRef][Green Version]
- Hall, W.; Degenhardt, L. Prevalence and correlates of cannabis use in developed and developing countries. Curr. Opin. Psychiatry 2007, 20, 393–397. [Google Scholar] [CrossRef]
- Nagarkatti, P.; Pandey, R.; Rieder, S.A.; Hegde, V.L.; Nagarkatti, M. Cannabinoids as novel anti-inflammatory drugs. Future Med. Chem. 2009, 1, 1333–1349. [Google Scholar] [CrossRef]
- Salazar-Roa, M.; Carracedo, A.; Salanueva, I.J.; Hernandez-Tiedra, S.; Lorente, M.; Egia, A.; Vázquez, P.; Blázquez, C.; Torres, S.; Garcia, S.; et al. Cannabinoid action induces autophagy-mediated cell death through stimulation of ER stress in human glioma cells. J. Clin. Investig. 2009, 119, 1359–1372. [Google Scholar] [CrossRef]
- Johnson, E.R.; Matthay, M.A. Acute Lung Injury: Epidemiology, Pathogenesis, and Treatment. J. Aerosol Med. Pulm. Drug Deliv. 2010, 23, 243–252. [Google Scholar] [CrossRef]
- Grommes, J.; Alard, J.-E.; Drechsler, M.; Wantha, S.; Mörgelin, M.; Kuebler, W.M.; Jacobs, M.; Von Hundelshausen, P.; Markart, P.; Wygrecka, M.; et al. Disruption of Platelet-derived Chemokine Heteromers Prevents Neutrophil Extravasation in Acute Lung Injury. Am. J. Respir. Crit. Care Med. 2012, 185, 628–636. [Google Scholar] [CrossRef]
- Lai, C.; Wang, K.; Zhao, Z.; Zhang, L.; Gu, H.; Yang, P.; Wang, X. C-C Motif Chemokine Ligand 2 (CCL2) Mediates Acute Lung Injury Induced by Lethal Influenza H7N9 Virus. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Gorczyca, W.; Bruno, S.; Darzynkiewicz, R.; Gong, J.; Darzynkiewicz, Z. DNA strand breaks occurring during apoptosis—Their early insitu detection by the terminal deoxynucleotidyl transferase and nick translation assays and prevention by serine protease inhibitors. Int. J. Oncol. 1992, 1. [Google Scholar] [CrossRef] [PubMed]
- Downer, E.; Boland, B.; Fogarty, M.; Campbell, V. Delta 9-tetrahydrocannabinol induces the apoptotic pathway in cultured cortical neurones via activation of the CB1 receptor. Neuroreport 2001, 12, 3973–3978. [Google Scholar] [CrossRef]
- Downer, E.J.; Fogarty, M.P.; Campbell, V.A. Tetrahydrocannabinol-induced neurotoxicity depends on CB1receptor-mediated c-Jun N-terminal kinase activation in cultured cortical neurons. Br. J. Pharmacol. 2003, 140, 547–557. [Google Scholar] [CrossRef] [PubMed]
- Do, Y.; McKallip, R.J.; Nagarkatti, M.; Nagarkatti, P. Activation through cannabinoid receptors 1 and 2 on dendritic cells triggers NF-kappaB-dependent apoptosis: Novel role for endogenous and exogenous cannabinoids in immunoregulation. J. Immunol. 2004, 173, 2373–2382. [Google Scholar] [CrossRef]
- Nachshon-Kedmi, M.; Yannai, S.; Fares, F.A. Induction of apoptosis in human prostate cancer cell line, PC3, by 3,3′-diindolylmethane through the mitochondrial pathway. Br. J. Cancer 2004, 91, 1358–1363. [Google Scholar] [CrossRef]
- Chandra, D.; Liu, J.W.; Tang, D.G. Early mitochondrial activation and cytochrome cup-regulation during apoptosis. J. Biol. Chem. 2002, 277, 50842–50854. [Google Scholar] [CrossRef]
- Mick, D.U.; Fox, T.D.; Rehling, P. Inventory control: Cytochrome c oxidase assembly regulates mitochondrial translation. Nat. Rev. Mol. Cell Biol. 2010, 12, 14–20. [Google Scholar] [CrossRef]
- Nyongo, A.; Huntrakoon, M. Microcystic Adenoma of the Pancreas with Myoepithelial Cells: A Hitherto Undescribed Morphologic Feature. Am. J. Clin. Pathol. 1985, 84, 114–120. [Google Scholar] [CrossRef][Green Version]
- Cui, B.; Lin, H.; Yu, J.; Yu, J.; Hu, Z.-W. Autophagy and the Immune Response. Adv. Exp. Med. Biol. 2019, 1206, 595–634. [Google Scholar] [CrossRef]
- Cosentino, K.; Garcia-Saez, A.J. Bax and Bak Pores: Are We Closing the Circle? Trends Cell Biol. 2017, 27, 266–275. [Google Scholar] [CrossRef] [PubMed]
- Jiang, X.; Wang, X. Cytochrome c promotes caspase-9 activation by inducing nucleotide binding to Apaf. J. Biol. Chem. 2000, 275, 31199–31203. [Google Scholar] [CrossRef]
- Sánchez-Alcázar, J.A.; Ault, J.G.; Khodjakov, A.; Schneider, E. Increased mitochondrial cytochrome c levels and mitochondrial hyperpolarization precede camptothecin-induced apoptosis in Jurkat cells. Cell Death Differ. 2000, 7, 1090–1100. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, O.; Fukuo, K.; Sun, X.; Nishitani, M.; Yotsui, T.; Higuchi, M.; Suzuki, T.; Rakugi, H.; Smithies, O.; Maeda, N.; et al. Apop-1, a Novel Protein Inducing Cyclophilin D-dependent but Bax/Bak-related Channel-independent Apoptosis. J. Biol. Chem. 2006, 281, 23899–23907. [Google Scholar] [CrossRef] [PubMed]
- Michalek, R.D.; Gerriets, V.A.; Jacobs, S.R.; MacIntyre, A.; Maciver, N.; Mason, E.F.; Sullivan, S.A.; Nichols, A.G.; Rathmell, J.C. Cutting edge: Distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J. Immunol. 2011, 186, 3299–3303. [Google Scholar] [CrossRef] [PubMed]
- Dominguez-Amorocho, O.; Takiishi, T.; da Cunha, F.F.; Camara, N.O.S. Immunometabolism: A target for the comprehension of immune response toward transplantation. World J. Transplant. 2019, 9, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.Y.; Liu, Y.-Y.; Liu, G.-H.; Lu, H.-B.; Mao, C.-Y. L-Carnitine and heart disease. Life Sci. 2018, 194, 88–97. [Google Scholar] [CrossRef] [PubMed]
- Rebouche, C.J. Kinetics, Pharmacokinetics, and Regulation of l-Carnitine and Acetyl-l-carnitine Metabolism. Ann. N. Y. Acad. Sci. 2004, 1033, 30–41. [Google Scholar] [CrossRef]
- Martínez, Y.; Li, X.; Liu, G.; Bin, P.; Yan, W.; Más, D.; Valdivié, M.; Hu, C.-A.A.; Ren, W.; Yin, J. The role of methionine on metabolism, oxidative stress, and diseases. Amino Acids 2017, 49, 2091–2098. [Google Scholar] [CrossRef]
- Li, G.; Wang, Y.; Liu, Y.; Su, Z.; Liu, C.; Ren, S.; Deng, T.; Huang, D.; Tian, Y.; Qiu, Y. miR-185-3p regulates nasopharyngeal carcinoma radioresistance by targeting WNT2Bin vitro. Cancer Sci. 2014, 105, 1560–1568. [Google Scholar] [CrossRef]
- Zhao, Z.-T.; Zhou, W.; Liu, L.-Y.; Lan, T.; Zhan, Q.; Song, Y.-M. Molecular mechanism and effect of microRNA185 on proliferation, migration and invasion of esophageal squamous cell carcinoma. Zhonghua Yi Xue Za Zhi 2013, 93, 1426–1431. [Google Scholar] [PubMed]
- Li, Q.; Wang, J.-X.; He, Y.-Q.; Feng, C.; Zhang, X.-J.; Sheng, J.-Q.; Li, P.-F. MicroRNA-185 regulates chemotherapeutic sensitivity in gastric cancer by targeting apoptosis repressor with caspase recruitment domain. Cell Death Dis. 2014, 5, e1197. [Google Scholar] [CrossRef] [PubMed]
- Qadir, X.V.; Han, C.; Lu, D.; Zhang, J.; Wu, T. miR-185 inhibits hepatocellular carcinoma growth by targeting the DNMT1/PTEN/Akt pathway. Am. J. Pathol. 2014, 184, 2355–2364. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Lee, H.; Cao, Y.; Cruz, C.S.D.; Jin, Y. miR-185 mediates lung epithelial cell death after oxidative stress. Am. J. Physiol. Cell. Mol. Physiol. 2016, 310, L700–L710. [Google Scholar] [CrossRef]
- Khodadadi, H.; Salles Évila, L.; Jarrahi, A.; Chibane, F.; Costigliola, V.; Yu, J.C.; Vaibhav, K.; Hess, D.C.; Dhandapani, K.M.; Baban, B. Cannabidiol Modulates Cytokine Storm in Acute Respiratory Distress Syndrome Induced by Simulated Viral Infection Using Synthetic RNA. Cannabis Cannabinoid Res. 2020. [Google Scholar] [CrossRef]
- Elliott, D.M.; Singh, N.; Nagarkatti, M.; Nagarkatti, P. Cannabidiol Attenuates Experimental Autoimmune Encephalomyelitis Model of Multiple Sclerosis through Induction of Myeloid-Derived Suppressor Cells. Front. Immunol. 2018, 9. [Google Scholar] [CrossRef]
- Hegde, V.L.; Nagarkatti, P.; Nagarkatti, M. Role of Myeloid-Derived Suppressor Cells in Amelioration of Experimental Autoimmune Hepatitis Following Activation of TRPV1 Receptors by Cannabidiol. PLoS ONE 2011, 6, e18281. [Google Scholar] [CrossRef]
- Kilkenny, C.; Browne, W.; Cuthill, I.C.; Emerson, M.; Altman, D.G.; Group NCRRGW. Animal research: Reporting in vivo experiments: The ARRIVE guidelines. Br. J. Pharmacol. 2010, 160, 1577–1579. [Google Scholar] [CrossRef]
- McGrath, J.; Drummond, G.; McLachlan, E.M.; Kilkenny, C.; Wainwright, C.L. Guidelines for reporting experiments involving animals: The ARRIVE guidelines. Br. J. Pharmacol. 2010, 160, 1573–1576. [Google Scholar] [CrossRef]
- Rieder, S.A.; Nagarkatti, P.; Nagarkatti, M. Multiple anti-inflammatory pathways triggered by resveratrol lead to amelioration of staphylococcal enterotoxin B-induced lung injury. Br. J. Pharmacol. 2012, 167, 1244–1258. [Google Scholar] [CrossRef]
- Sido, J.M.; Nagarkatti, P.S.; Nagarkatti, M. Delta(9)-Tetrahydrocannabinol attenuates allogeneic host-versus-graft response and delays skin graft rejection through activation of cannabinoid receptor 1 and induction of myeloid-derived suppressor cells. J. Leukoc. Biol. 2015, 98, 435–447. [Google Scholar] [CrossRef] [PubMed]
- Vantaku, V.; Putluri, V.; Bader, D.A.; Maity, S.; Ma, J.; Arnold, J.M.; Rajapakshe, K.; Donepudi, S.R.; Von Rundstedt, F.-C.; Devarakonda, V.; et al. Epigenetic loss of AOX1 expression via EZH2 leads to metabolic deregulations and promotes bladder cancer progression. Oncogene 2019. [Google Scholar] [CrossRef] [PubMed]
- Vantaku, V.; Dong, J.; Ambati, C.R.; Perera, D.; Donepudi, S.R.; Amara, C.S.; Putluri, V.; Ravi, S.S.; Robertson, M.J.; Piyarathna, D.W.B.; et al. Multi-omics Integration Analysis Robustly Predicts High-Grade Patient Survival and Identifies CPT1B Effect on Fatty Acid Metabolism in Bladder Cancer. Clin. Cancer Res. 2019, 25, 3689–3701. [Google Scholar] [CrossRef] [PubMed]
- Amara, C.S.; Ambati, C.R.; Vantaku, V.; Piyarathna, D.W.B.; Donepudi, S.R.; Ravi, S.S.; Arnold, J.M.; Putluri, V.; Chatta, G.; Guru, K.A.; et al. Serum Metabolic Profiling Identified a Distinct Metabolic Signature in Bladder Cancer Smokers: A Key Metabolic Enzyme Associated with Patient Survival. Cancer Epidemiol. Biomark. Prev. 2019, 28, 770–781. [Google Scholar] [CrossRef] [PubMed]
- Kornberg, M.D.; Bhargava, P.; Kim, P.M.; Putluri, V.; Snowman, A.M.; Putluri, N.; Calabresi, P.A.; Snyder, S.H. Dimethyl fumarate targets GAPDH and aerobic glycolysis to modulate immunity. Science 2018, 360, 449–453. [Google Scholar] [CrossRef] [PubMed]
- Wangler, M.F.; Chao, Y.-H.; Bayat, V.; Giagtzoglou, N.; Shinde, A.B.; Putluri, N.; Coarfa, C.; Donti, T.; Graham, B.H.; Faust, J.E.; et al. Peroxisomal biogenesis is genetically and biochemically linked to carbohydrate metabolism in Drosophila and mouse. PLoS Genet. 2017, 13, e1006825. [Google Scholar] [CrossRef]
- Vantaku, V.; Donepudi, S.R.; Piyarathna, D.W.B.; Amara, C.S.; Ambati, C.R.; Tang, W.; Putluri, V.; Chandrashekar, D.S.; Varambally, S.; Terris, M.; et al. Large-scale profiling of serum metabolites in African American and European American patients with bladder cancer reveals metabolic pathways associated with patient survival. Cancer 2019, 125, 921–932. [Google Scholar] [CrossRef]
- Butler, A.; Hoffman, P.; Smibert, P.; Papalexi, E.; Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol. 2018, 36, 411–420. [Google Scholar] [CrossRef]
- Stuart, T.; Butler, A.; Hoffman, P.; Hafemeister, C.; Papalexi, E.; Mauck, W.M., III; Hao, Y.; Stoeckius, M.; Smibert, P.; Satija, R. Comprehensive Integration of Single-Cell Data. Cell 2019, 177, 1888–1902. [Google Scholar] [CrossRef]
- Curtis, M.J.; Alexander, S.P.; Cirino, G.; Docherty, J.R.; George, C.H.; Giembycz, M.A.; Hoyer, D.; A Insel, P.; Izzo, A.A.; Ji, Y.; et al. Experimental design and analysis and their reporting II: Updated and simplified guidance for authors and peer reviewers. Br. J. Pharmacol. 2018, 175, 987–993. [Google Scholar] [CrossRef]
| Gene Name | Forward Primer (5′→3′) | Reverse Primer (5′→3′) |
|---|---|---|
| Runx3 | CAGGTTCAACGACCTTCGATT | GTGGTAGGTAGCCACTTGGG |
| Nkiras2 | TCTGTGGGCAAAACTTCAATCC | CGATGGAGCCTACATAGATGTCC |
| Bad | AAGTCCGATCCCGGAATCC | GCTCACTCGGCTCAAACTCT |
| Bax | TGAAGACAGGGGCCTTTTTG | AATTCGCCGGAGACACTCG |
| Cox4 | CTCTTCGTGGACTGCATCCC | GGTAATAGCCAGCGATCACATAG |
| Hrk | CAAAGCCTGGGAGGTCTGAG | TCTCACAAAGGCTTCGGTCC |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mohammed, A.; F.K. Alghetaa, H.; Miranda, K.; Wilson, K.; P. Singh, N.; Cai, G.; Putluri, N.; Nagarkatti, P.; Nagarkatti, M. Δ9-Tetrahydrocannabinol Prevents Mortality from Acute Respiratory Distress Syndrome through the Induction of Apoptosis in Immune Cells, Leading to Cytokine Storm Suppression. Int. J. Mol. Sci. 2020, 21, 6244. https://doi.org/10.3390/ijms21176244
Mohammed A, F.K. Alghetaa H, Miranda K, Wilson K, P. Singh N, Cai G, Putluri N, Nagarkatti P, Nagarkatti M. Δ9-Tetrahydrocannabinol Prevents Mortality from Acute Respiratory Distress Syndrome through the Induction of Apoptosis in Immune Cells, Leading to Cytokine Storm Suppression. International Journal of Molecular Sciences. 2020; 21(17):6244. https://doi.org/10.3390/ijms21176244
Chicago/Turabian StyleMohammed, Amira, Hasan F.K. Alghetaa, Kathryn Miranda, Kiesha Wilson, Narendra P. Singh, Guoshuai Cai, Nagireddy Putluri, Prakash Nagarkatti, and Mitzi Nagarkatti. 2020. "Δ9-Tetrahydrocannabinol Prevents Mortality from Acute Respiratory Distress Syndrome through the Induction of Apoptosis in Immune Cells, Leading to Cytokine Storm Suppression" International Journal of Molecular Sciences 21, no. 17: 6244. https://doi.org/10.3390/ijms21176244
APA StyleMohammed, A., F.K. Alghetaa, H., Miranda, K., Wilson, K., P. Singh, N., Cai, G., Putluri, N., Nagarkatti, P., & Nagarkatti, M. (2020). Δ9-Tetrahydrocannabinol Prevents Mortality from Acute Respiratory Distress Syndrome through the Induction of Apoptosis in Immune Cells, Leading to Cytokine Storm Suppression. International Journal of Molecular Sciences, 21(17), 6244. https://doi.org/10.3390/ijms21176244








