Proteomics and Metabolomics for Cystic Fibrosis Research
Abstract
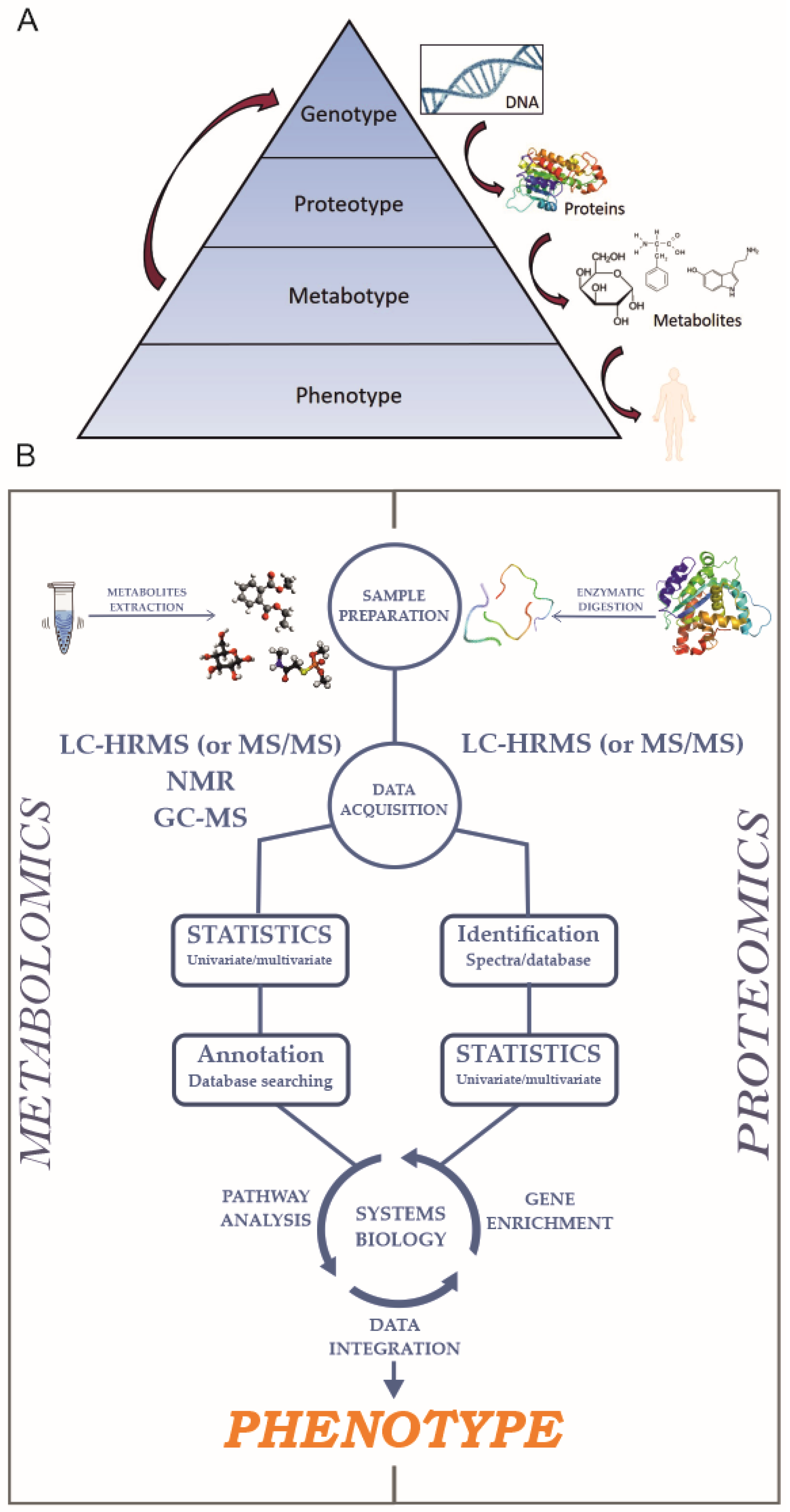
1. Introduction
2. Proteomics
3. Metabolomics
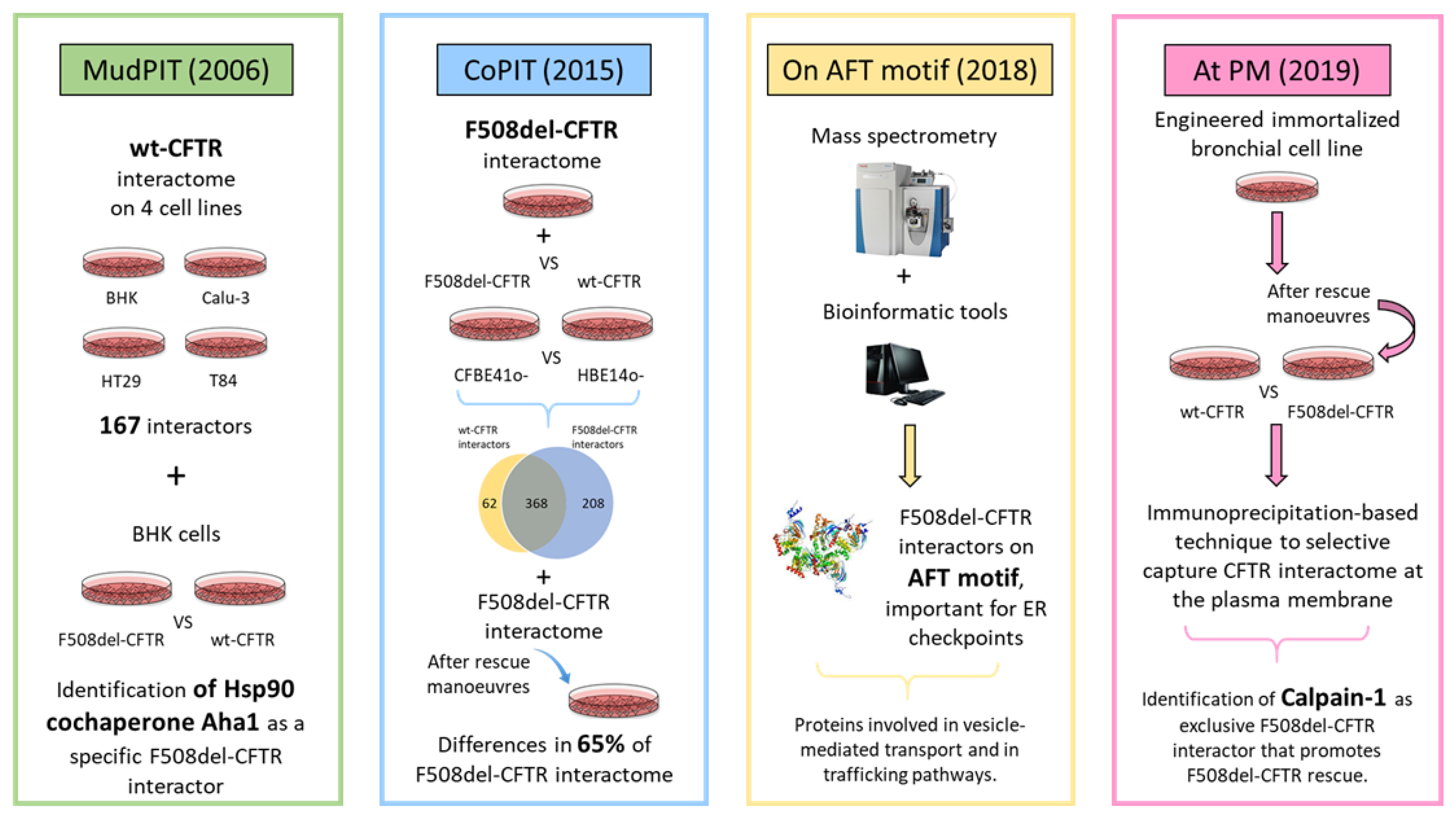
4. Proteomics to Investigate CFTR Network: Interactomics
4.1. MudPIT-Based Interactomics
4.2. CoPIT-Based Interactomics
4.3. Elucidating the AFT-Based CFTR Interactome
4.4. CFTR Interactomics at Plasma Membrane
4.5. G551D-CFTR Interactomics
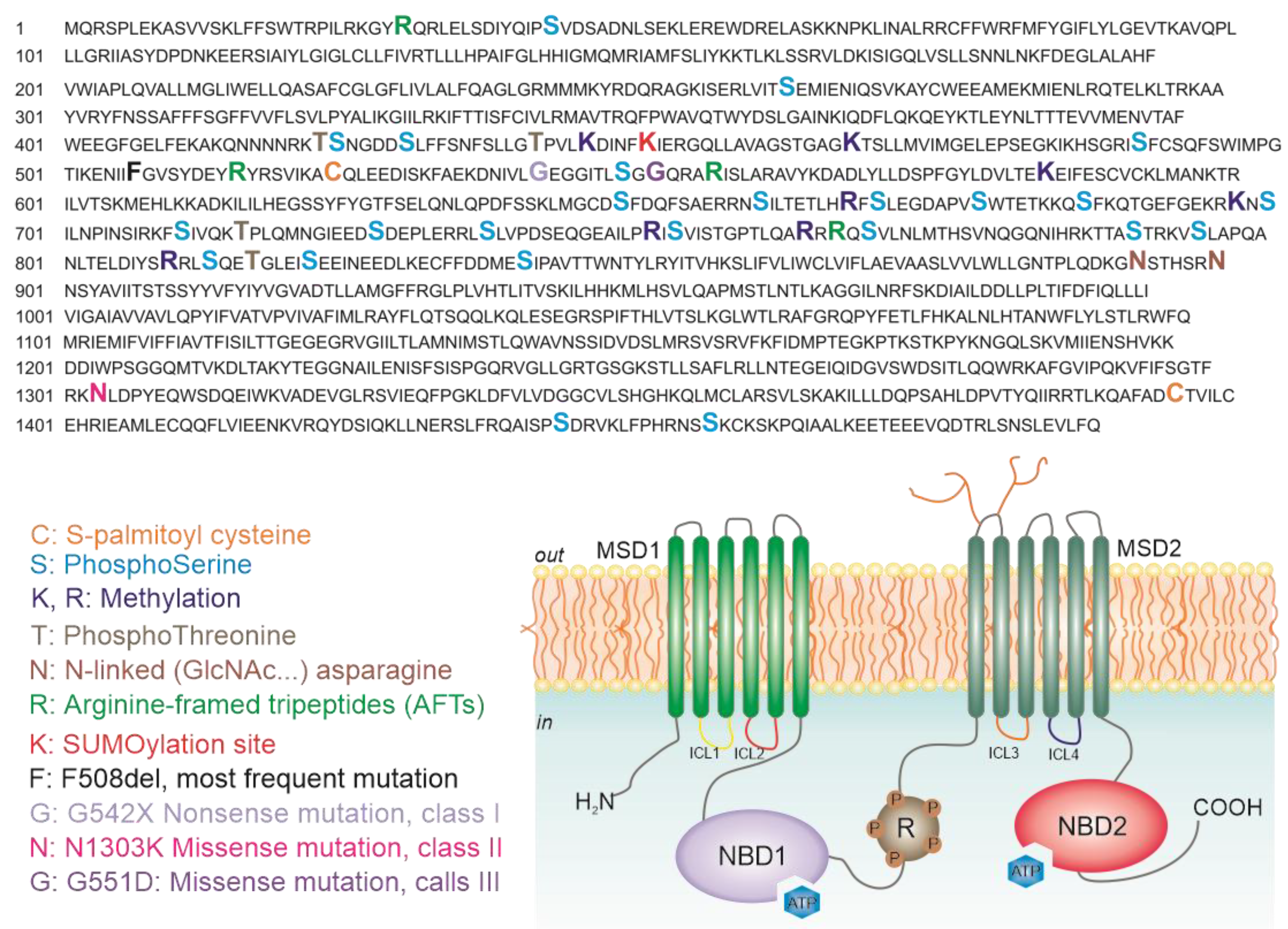
5. Analysis of CFTR Post-Translational Modifications
6. Global Changes in Protein Expression Observed in CF
7. Metabolomics and CFTR Research
- Nucleotide metabolism, with a decrease in purine biosynthesis, essential for the control of ASL (airway surface liquid) volume. The authors claimed that these results are in contrast with previous results from EBC (exhaled breath condensate) purinergic receptors [51].
- Increase in the catabolism of tryptophan, causing accumulation of molecules associated with oxidative stress [52].
- Reduction of glutathione biosynthesis, a key regulator of the oxidative status of the cells.
- Decrease in osmolytes, such as sorbitol and glycerophosphorylcoline, which have a role in the regulation of cell volume.
- Low levels of glucose metabolism, a possible cause of an increase in cell sensitivity to oxidative stress.
8. Metabolomics for Biomarker Discovery: Support and Aid CF Diagnosis
- Ceramides, known to be involved in proinflammatory and proapoptotic signaling, were thought to be increased in CF patients [77]. Based on these studies, amitriptyline and fenretinide, two drugs modulating ceramide metabolism, are undergoing clinical trials. Nevertheless, recent studies showed that the plasma levels of ceramides in CF murine models are decreased [78]; these unclear results underline the need of further investigations of the role of sphingolipids in CF. Indeed, sphingosines levels are lower in CF mouse epithelial cells compared to no-CF epithelia [79] and gangliosides are reduced in Calu-3 bronchial epithelial cells (found to be decreased) [80].
9. Omics to Study the CF Microbiome
10. Conclusions and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| LC–HRMS | Liquid chromatography coupled to high resolution mass spectrometry |
| MS/MS | Tandem mass analysis |
| NMR | Nuclear magnetic resonance |
| LD | Linear dichroism |
References
- Ratjen, F.; Döring, G. Cystic fibrosis. Lancet 2003, 361, 681–689. [Google Scholar] [CrossRef]
- Hwang, T.C.; Yeh, J.T.; Zhang, J.; Yu, Y.C.; Yeh, H.I.; Destefano, S. Structural mechanisms of CFTR function and dysfunction. J. Gen. Physiol. 2018, 150, 539–570. [Google Scholar] [CrossRef] [PubMed]
- Scanlin, F.T.; Glick, M.C. Terminal glycosylation in cystic fibrosis. Biochim. Biophys. Acta 1999, 1455, 241–253. [Google Scholar] [CrossRef]
- Vergani, P.; Lockless, S.W.; Nairn, A.C.; Gadsby, D.C. CFTR channel opening by ATP-driven tight dimerization of its nucleotide-binding domains. Nature 2005, 433, 876–880. [Google Scholar] [CrossRef]
- Derichs, N. Targeting a genetic defect: Cystic fibrosis transmembrane conductance regulator modulators in cystic fibrosis. Eur. Respir. Rev. 2013, 22, 58–65. [Google Scholar] [CrossRef]
- Amaral, M.D. Novel personalized therapies for cystic fibrosis: Treating the basic defect in all patients. J. Intern Med. 2015, 277, 155–166. [Google Scholar] [CrossRef]
- Lukacs, G.L.; Chang, X.B.; Bear, C.; Kartner, N.; Mohamed, A.; Riordan, J.R.; Grinstein, S. The delta F508 mutation decreases the stability of cystic fibrosis transmembrane conductance regulator in the plasma membrane. Determination of functional half-lives on transfected cells. J. Biol. Chem. 1993, 268, 21592–21598. [Google Scholar]
- Sanders, D.B.; Fink, A.K. Background and Epidemiology. Pediatr. Clin. N. Am. 2016, 63, 567–584. [Google Scholar] [CrossRef]
- Andersen, D.H. Cystic fibrosis of the pancreas and its relation to celiac disease. Am. J. Dis. Child 1938, 56, 344–399. [Google Scholar] [CrossRef]
- Riordan, J.R.; Rommens, J.M.; Kerem, B.S.; Alon, N.; Rozmahel, R.; Grzelczak, Z.; Zielenski, J.; Lok, S.; Plavsic, N.; Chou, J.L. Identification of the Cystic Fibrosis Gene cloning and characterization of complementary DNA. Science 1989, 245, 1066–1073. [Google Scholar] [CrossRef]
- Lander, E.S.; Linton, L.M.; Birren, B.; Nusbaum, C.; Zody, M.C.; Baldwin, J.; Devon, K.; Dewar, K.; Doyle, M.; FitzHugh, W.; et al. Initial sequencing and analysis of the human genome. Nature 2001, 409, 860–921. [Google Scholar] [PubMed]
- Goldstein, J.L.; Brown, M.S. Regulation of the mevalonate pathway. Nature 1990, 343, 425–430. [Google Scholar] [CrossRef] [PubMed]
- Boztepe, T.; Gulec, S. Investigation of the influence of high glucose on molecular and genetic responses: An in vitro study using a human intestine model. Genes Nutr. 2018, 13, 11. [Google Scholar] [CrossRef] [PubMed]
- Monte, A.A.; Brocker, C.; Nebert, D.W.; Gonzalez, F.J.; Thompson, D.C.; Vasiliou, V. Improved drug therapy: Triangulating phenomics with genomics and metabolomics. Hum. Genom. 2014, 8, 16. [Google Scholar] [CrossRef]
- Zhang, Y.; Fonslow, B.R.; Shan, B.; Baek, M.C.; Yates, J.R., III. Protein analysis by shotgun/bottom-up proteomics. Chem. Rev. 2013, 113, 2343–2394. [Google Scholar] [CrossRef]
- Bantscheff, M.; Lemeer, S.; Savitski, M.M.; Kuster, B. Quantitative mass spectrometry in proteomics: Critical review update from 2007 to the present. Anal. Bioanal. Chem. 2012, 404, 939–965. [Google Scholar] [CrossRef]
- Jabeen, A.; Mohamedali, A.; Ranganathan, S. Looking for Missing Proteins. Ref. Modul. Life Sci. 2019. [Google Scholar] [CrossRef]
- Griffiths, W.J.; Wang, Y. Mass spectrometry: From proteomics to metabolomics and lipidomics. Chem. Soc. Rev. 2009, 38, 1882–1896. [Google Scholar] [CrossRef]
- Beale, D.J.; Pinu, F.R.; Kouremenos, K.A.; Poojary, M.M.; Narayana, V.K.; Boughton, B.A.; Kanojia, K.; Dayalan, S.; Jones, O.A.; Dias, D.A. Review of recent developments in GC-MS approaches to metabolomics-based research. Metabolomics 2018, 14, 152. [Google Scholar] [CrossRef]
- Fiehn, O. Metabolomics by Gas Chromatography-Mass Spectrometry: Combined Targeted and Untargeted Profiling. Curr. Protoc. Mol. Biol. 2016, 114, 30–34. [Google Scholar] [CrossRef]
- Emwas, A.H.; Roy, R.; McKay, R.T.; Tenori, L.; Saccenti, E.; Gowda, G.A.; Raftery, D.; Alahmari, F.; Jaremko, L.; Jaremko, M.; et al. NMR Spectroscopy for Metabolomics Research. Metabolites 2019, 9, 123. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.R.; Kwon, H.N.; Nam, H.; Kim, J.J.; Park, S.; Kim, Y.H. Urine-NMR metabolomics for screening of advanced colorectal adenoma and early stage colorectal cancer. Sci. Rep. 2019, 9, 4786. [Google Scholar] [CrossRef] [PubMed]
- Doerr, A. Tracking the protein–metabolite interactome. Nat. Methods 2018, 15, 160. [Google Scholar] [CrossRef]
- Breitling, R. What is systems biology? Front. Physiol. 2010, 1, 9. [Google Scholar] [CrossRef]
- Kiemer, L.; Cesareni, G. Comparative interactomics: Comparing apples and pears? Trends Biotechnol. 2007, 25, 448–454. [Google Scholar] [CrossRef]
- Wang, X.; Venable, J.; LaPointe, P.; Hutt, D.M.; Koulov, A.V.; Coppinger, J.; Gurkan, C.; Kellner, W.; Matteson, J.; Plutner, H.; et al. Hsp90 cochaperone Aha1 downregulation rescues misfolding of CFTR in cystic fibrosis. Cell 2006, 127, 803–815. [Google Scholar] [CrossRef]
- Pankow, S.; Bamberger, C.; Calzolari, D.; Martínez-Bartolomé, S.; Lavallée-Adam, M.; Balch, W.E.; Yates, J.R., 3rd. Delta F508 CFTR interactome remodelling promotes rescue of cystic fibrosis. Nature 2015, 528, 510–516. [Google Scholar] [CrossRef]
- Hutt, D.M.; Herman, D.; Rodrigues, A.P.C.; Noel, S.; Pilewski, J.M.; Matteson, J.; Hoch, B.; Kellner, W.; Kelly, J.W.; Schmidt, A. Reduced histone deacetylase 7 activity restores function to misfolded CFTR in cystic fibrosis. Nat. Chem. Biol. 2010, 6, 25–33. [Google Scholar] [CrossRef]
- Sondo, E.; Tomati, V.; Caci, E.; Esposito, A.I.; Pfeffer, U.; Pedemonte, N.; Galietta, L.J. Rescue of the mutant CFTR chloride channel by pharmacological correctors and low temperature analyzed by gene expression profiling. Am. J. Physiol. Cell Physiol. 2011, 301, C872–C885. [Google Scholar] [CrossRef]
- Santos, J.D.; Canato, S.; Carvalho, A.S.; Botelho, H.M.; Aloria, K.; Amaral, M.D.; Matthiesen, R.; Falcao, A.O.; Farinha, C.M. Folding Status Is Determinant over Traffic-Competence in Defining CFTR Interactors in the Endoplasmic Reticulum. Cells 2019, 8, 353. [Google Scholar] [CrossRef]
- Roxo-Rosa, M.; Xu, Z.; Schmidt, A.; Neto, M.; Cai, Z.; Soares, C.M.; Sheppard, D.N.; Amaral, M.D. Revertant mutants G550E and 4RK rescue cystic fibrosis mutants in the first nucleotide-binding domain of CFTR by different mechanisms. Proc. Natl. Acad. Sci. USA 2006, 103, 17891–17896. [Google Scholar] [CrossRef] [PubMed]
- Farinha, C.M.; King-Underwood, J.; Sousa, M.; Correia, A.R.; Henriques, B.J.; Roxo-Rosa, M.; Da Paula, A.C.; Williams, J.; Hirst, S.; Gomes, C.M.; et al. Revertants, low temperature, and correctors reveal the mechanism of F508del-CFTR rescue by VX-809 and suggest multiple agents for full correction. Chem. Biol. 2013, 20, 943–955. [Google Scholar] [CrossRef] [PubMed]
- Denic, V.; Dotsch, V.; Sinning, I. Endoplasmic reticulum targeting and insertion of tail-anchored membrane proteins by the GET pathway. Cold Spring Harb. Perspect Biol. 2013, 5, a013334. [Google Scholar] [CrossRef] [PubMed]
- Tomati, V.; Pesce, E.; Caci, E.; Sondo, E.; Scudieri, P.; Marini, M.; Amato, F.; Castaldo, G.; Ravazzolo, R.; Galietta, L.; et al. High-throughput screening identifies FAU protein as a regulator of mutant cystic fibrosis transmembrane conductance regulator channel. J. Biol. Chem. 2018, 293, 1203–1217. [Google Scholar] [CrossRef] [PubMed]
- Matos, A.M.; Pinto, F.R.; Barros, P.; Amaral, M.D.; Pepperkok, R.; Matos, P. Inhibition of calpain 1 restores plasma membrane stability to pharmacologically rescued Phe508del-CFTR variant. J. Biol. Chem. 2019, 294, 13396–13410. [Google Scholar] [CrossRef]
- Botelho, H.M.; Uliyakina, I.; Awatade, N.T.; Proença, M.C.; Tischer, C.; Sirianant, L.; Kunzelmann, K.; Pepperkok, R.; Amaral, M.D. Protein traffic disorders: An effective high-throughput fluorescence microscopy pipeline for drug discovery. Sci. Rep. 2015, 5, 9038. [Google Scholar] [CrossRef]
- Averna, M.; Stifanese, R.; De Tullio, R.; Minicucci, L.; Cresta, F.; Palena, S.; Salamino, F.; Pontremoli, S.; Melloni, E. Evidence for alteration of calpain/calpastatin system in PBMC of cystic fibrosis patients. Biochim. Biophys. Acta 2011, 1812, 1649–1657. [Google Scholar] [CrossRef]
- Teng, L.; Kerbiriou, M.; Taiya, M.; Le Hir, S.; Mignen, O.; Benz, N.; Trouvé, P.; Férec, C. Proteomic Identification of Calumenin as a G551D-CFTR Associated Protein. PLoS ONE 2012, 7, e40173. [Google Scholar] [CrossRef]
- Pankow, S.; Bamberger, C.; Yates, J.R., III. A posttranslational modification code for CFTR maturation is altered in cystic fibrosis. Sci. Signal. 2019, 12, eaan7984. [Google Scholar] [CrossRef]
- Gadsby, D.C.; Nairn, A.C. Control of CFTR channel gating by phosphorylation and nucleotide hydrolysis. Physiol. Rev. 1999, 79 (Suppl. 1), S77–S107. [Google Scholar] [CrossRef]
- Winter, M.C.; Welsh, M.J. Stimulation of CFTR activity by its phosphorylated R domain. Nature 1997, 389, 294–296. [Google Scholar] [CrossRef] [PubMed]
- Treharne, K.J.; Crawford, R.M.; Xu, Z.; Chen, J.H.; Best, O.G.; Schulte, E.A.; Gruenert, D.C.; Wilson, S.M.; Sheppard, D.N.; Kunzelmann, K.; et al. Protein kinase CK2, cystic fibrosis transmembrane conductance regulator, and the deltaF508 mutation: F508 deletion disrupts a kinase-binding site. J. Biol. Chem. 2008, 283, 25103. [Google Scholar] [PubMed]
- O’Connell, J.D.; Paulo, J.A.; O’Brien, J.J.; Gygi, S.P. Proteome-Wide Evaluation of Two Common Protein Quantification Methods. J. Proteome Res. 2018, 17, 1934–1942. [Google Scholar] [CrossRef] [PubMed]
- Frischer, T.; Myung, J.K.; Maurer, G.; Eichler, I.; Szepfalusi, Z.; Lubec, G. Possible dysregulation of chaperon and metabolic proteins in cystic fibrosis bronchial tissue. Proteomics 2006, 6, 3381–3388. [Google Scholar] [CrossRef]
- Puglia, M.; Landi, C.; Gagliardi, A.; Breslin, L.; Armini, A.; Brunetti, J.; Pini, A.; Bianchi, L.; Bini, L. The proteome speciation of an immortalized cystic fibrosis cell line: New perspectives on the pathophysiology of the disease. J. Proteom. 2018, 170, 28–42. [Google Scholar] [CrossRef]
- Rauniyar, N.; Gupta, V.; Balch, W.E.; Yates, J.R., 3rd. Quantitative proteomic profiling reveals differentially regulated proteins in cystic fibrosis cells. J. Proteome Res. 2014, 13, 4668–4675. [Google Scholar] [CrossRef]
- Braccia, C.; Tomati, V.; Caci, E.; Pedemonte, N.; Armirotti, A. SWATH label-free proteomics for cystic fibrosis research. J. Cyst. Fibros. 2019, 18, 501–506. [Google Scholar] [CrossRef]
- Chin, S.; Ramjeesingh, M.; Hung, M.; Ereño-Oreba, J.; Cui, H.; Laselva, O.; Julien, J.P.; Bear, C.E. Cholesterol Interaction Directly Enhances Intrinsic Activity of the Cystic Fibrosis Transmembrane Conductance Regulator (CFTR). Cells 2019, 8, 804. [Google Scholar] [CrossRef]
- Malik, F.A.; Meissner, A.; Semenkov, I.; Molinski, S.; Pasyk, S.; Ahmadi, S.; Bui, H.H.; Bear, C.E.; Lidington, D.; Bolz, S.S. Sphingosine-1-Phosphate Is a Novel Regulator of Cystic Fibrosis Transmembrane Conductance Regulator (CFTR) Activity. PLoS ONE 2015, 10, e0130313. [Google Scholar] [CrossRef]
- Wetmore, D.R.; Joseloff, E.; Pilewski, J.; Lee, D.P.; Lawton, K.A.; Mitchell, M.W.; Milburn, M.V.; Ryals, J.A.; Guo, L. Metabolomic profiling reveals biochemical pathways and biomarkers associated with pathogenesis in cystic fibrosis cells. J. Biol. Chem. 2010, 285, 30516–30522. [Google Scholar] [CrossRef]
- Esther, C.R., Jr.; Alexis, N.E.; Clas, M.L.; Lazarowski, E.R.; Donaldson, S.H.; Ribeiro, C.M.; Moore, C.G.; Davis, S.D.; Boucher, R.C. Extracellular purines are biomarkers of neutrophilic airway inflammation. Eur. Respir. J. 2008, 31, 949–956. [Google Scholar] [CrossRef] [PubMed]
- Pawlak, K.; Kowalewska, A.; Mysliwiec, M.; Pawlak, D. Kynurenine and its metabolites--kynurenic acid and anthranilic acid are associated with soluble endothelial adhesion molecules and oxidative status in patients with chronic kidney disease. Am. J. Med. Sci. 2009, 338, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Joseloff, E.; Sha, W.; Bell, S.C.; Wetmore, D.R.; Lawton, K.A.; Milburn, M.V.; Ryals, J.A.; Guo, L.; Muhlebach, M.S. Serum metabolomics indicate altered cellular energy metabolism in children with cystic fibrosis. Pediatr. Pulmonol. 2014, 49, 463–472. [Google Scholar] [CrossRef] [PubMed]
- Al-Qahtani, W.; Abdel Jabar, M.; Masood, A.; Jacob, M.; Nizami, I.; Dasouki, M.; Abdel Rahman, A.M. Dried Blood Spot-Based Metabolomic Profiling in Adults with Cystic Fibrosis. J. Proteome Res. 2020, 19, 2346–2357. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, B.C. Inhaled dry powder mannitol: A solution for cystic fibrosis. Am. J. Respir. Crit. Care Med. 2012, 185, 596–598. [Google Scholar] [CrossRef]
- Kopp, B.T.; McCulloch, S.; Shrestha, C.L.; Zhang, S.; Sarzynski, L.; Woodley, F.W.; Hayes, D., Jr. Metabolomic responses to lumacaftor/ivacaftor in cystic fibrosis. Pediatr. Pulmonol. 2018, 53, 583–591. [Google Scholar] [CrossRef]
- Grassme, H.; Henry, B.; Ziobro, R.; Becker, K.A.; Riethmüller, J.; Gardner, A.; Seitz, A.P.; Steinmann, J.; Lang, S.; Ward, C.; et al. Beta1-Integrin Accumulates in Cystic Fibrosis Luminal Airway Epithelial Membranes and Decreases Sphingosine, Promoting Bacterial Infections. Cell Host Microbe 2017, 21, 707–718.e8. [Google Scholar] [CrossRef]
- Tomati, V.; Sondo, E.; Armirotti, A.; Caci, E.; Pesce, E.; Marini, M.; Gianotti, A.; Jeon, Y.J.; Cilli, M.; Pistorio, A.; et al. Genetic Inhibition Of The Ubiquitin Ligase Rnf5 Attenuates Phenotypes Associated To F508del Cystic Fibrosis Mutation. Sci. Rep. 2015, 5, 12138. [Google Scholar] [CrossRef]
- Castellani, C.; Massie, J.; Sontag, M.; Southern, K.W. Newborn screening for cystic fibrosis. Lancet Respir. Med. 2016, 4, 653–661. [Google Scholar] [CrossRef]
- Mishra, A.; Greaves, R.; Massie, J. The relevance of sweat testing for the diagnosis of cystic fibrosis in the genomic era. Clin. Biochem. Rev. 2005, 26, 135–153. [Google Scholar]
- Knowles, M.; Gatzy, J.; Boucher, R. Increased bioelectric potential difference across respiratory epithelia in cystic fibrosis. N. Engl. J. Med. 1981, 305, 1489–1495. [Google Scholar] [CrossRef] [PubMed]
- Castellani, C.; Linnane, B.; Pranke, I.; Cresta, F.; Sermet-Gaudelus, I.; Peckham, D. Cystic Fibrosis Diagnosis in Newborns, Children, and Adults. Semin. Respir. Crit. Care Med. 2019, 40, 701–714. [Google Scholar] [CrossRef] [PubMed]
- DiBattista, A.; McIntosh, N.; Lamoureux, M.; Al-Dirbashi, O.Y.; Chakraborty, P.; Britz-McKibbin, P. Metabolic Signatures of Cystic Fibrosis Identified in Dried Blood Spots For Newborn Screening Without Carrier Identification. J. Proteome Res. 2019, 18, 841–854. [Google Scholar] [CrossRef]
- Zhang, Q.; Ames, J.M.; Smith, R.D.; Baynes, J.W.; Metz, T.O. A perspective on the Maillard reaction and the analysis of protein glycation by mass spectrometry: Probing the pathogenesis of chronic disease. J. Proteome Res. 2009, 8, 754–769. [Google Scholar] [CrossRef]
- Hudson, V.M. Rethinking cystic fibrosis pathology: The critical role of abnormal reduced glutathione (GSH) transport caused by CFTR mutation. Free Radic. Biol. Med. 2001, 30, 1440–1461. [Google Scholar] [CrossRef]
- Kogan, I.; Ramjeesingh, M.; Li, C.; Kidd, J.F.; Wang, Y.; Leslie, E.M.; Cole, S.P.; Bear, C.E. CFTR directly mediates nucleotide-regulated glutathione flux. EMBO J. 2003, 22, 1981–1989. [Google Scholar] [CrossRef]
- Macedo, A.N.; Mathiaparanam, S.; Brick, L.; Keenan, K.; Gonska, T.; Pedder, L.; Hill, S.; Britz-McKibbin, P. The Sweat Metabolome of Screen-Positive Cystic Fibrosis Infants: Revealing Mechanisms beyond Impaired Chloride Transport. ACS Cent. Sci. 2017, 3, 904–913. [Google Scholar] [CrossRef]
- Hioki, T.; Fukami, T.; Nakajima, M.; Yokoi, T. Human paraoxonase 1 is the enzyme responsible for pilocarpine hydrolysis. Drug Metab Dispos. 2011, 39, 1345–1352. [Google Scholar] [CrossRef]
- Xie, C.; Jin, R.; Zhao, Y.; Lin, L.; Li, L.; Chen, J.; Zhang, Y. Paraoxonase 2 gene polymorphisms and prenatal phthalates’ exposure in Chinese newborns. Environ. Res. 2015, 140, 354–359. [Google Scholar] [CrossRef]
- Zhou, Z.; Alvarez, D.; Milla, C.; Zare, R.N. Proof of concept for identifying cystic fibrosis from perspiration samples. Proc. Natl. Acad. Sci. USA 2019, 116, 24408–24412. [Google Scholar] [CrossRef]
- Sharom, F.J. Complex Interplay between the P-Glycoprotein Multidrug Efflux Pump and the Membrane: Its Role in Modulating Protein Function. Front. Oncol. 2014, 4, 41. [Google Scholar] [CrossRef] [PubMed]
- Cottrill, K.A.; Farinha, C.M.; McCarty, N.A. The bidirectional relationship between CFTR and lipids. Commun. Biol. 2020, 3, 179. [Google Scholar] [CrossRef] [PubMed]
- Al-Turkmani, M.R.; Andersson, C.; Alturkmani, R.; Katrangi, W.; Cluette-Brown, J.E.; Freedman, S.D.; Laposata, M. A mechanism accounting for the low cellular level of linoleic acid in cystic fibrosis and its reversal by DHA. J. Lipid Res. 2008, 49, 1946–1954. [Google Scholar] [CrossRef] [PubMed]
- Freedman, S.D.; Blanco, P.G.; Zaman, M.M.; Shea, J.C.; Ollero, M.; Hopper, I.K.; Weed, D.A.; Gelrud, A.; Regan, M.M.; Laposata, M.; et al. Association of cystic fibrosis with abnormalities in fatty acid metabolism. N. Engl. J. Med. 2004, 350, 560–569. [Google Scholar] [CrossRef]
- Ernst, W.L.; Shome, K.; Wu, C.C.; Gong, X.; Frizzell, R.A.; Aridor, M. VAMP-associated Proteins (VAP) as Receptors That Couple Cystic Fibrosis Transmembrane Conductance Regulator (CFTR) Proteostasis with Lipid Homeostasis. J. Biol. Chem. 2016, 291, 5206–5220. [Google Scholar] [CrossRef]
- Slomiany, A.; Murty, V.L.; Aono, M.; Snyder, C.E.; Herp, A.; Slomiany, B.L. Lipid composition of tracheobronchial secretions from normal individuals and patients with cystic fibrosis. Biochim. Biophys. Acta 1982, 710, 106–111. [Google Scholar] [CrossRef]
- Teichgraber, V.; Ulrich, M.; Endlich, N.; Riethmüller, J.; Wilker, B.; De Oliveira-Munding, C.C.; van Heeckeren, A.M.; Barr, M.L.; von Kürthy, G.; Schmid, K.W.; et al. Ceramide accumulation mediates inflammation, cell death and infection susceptibility in cystic fibrosis. Nat. Med. 2008, 14, 382–391. [Google Scholar] [CrossRef]
- Garic, D.; De Sanctis, J.B.; Wojewodka, G.; Houle, D.; Cupri, S.; Abu-Arish, A.; Hanrahan, J.W.; Hajduch, M.; Matouk, E.; Radzioch, D. Fenretinide differentially modulates the levels of long- and very long-chain ceramides by downregulating Cers5 enzyme: Evidence from bench to bedside. J. Mol. Med. 2017, 95, 1053–1064. [Google Scholar] [CrossRef]
- Boujaoude, L.C.; Bradshaw-Wilder, C.; Mao, C.; Cohn, J.; Ogretmen, B.; Hannun, Y.A.; Obeid, L.M. Cystic fibrosis transmembrane regulator regulates uptake of sphingoid base phosphates and lysophosphatidic acid: Modulation of cellular activity of sphingosine 1-phosphate. J. Biol. Chem. 2001, 276, 35258–35264. [Google Scholar] [CrossRef]
- Itokazu, Y.; Pagano, R.E.; Schroeder, A.S.; O’Grady, S.M.; Limper, A.H.; Marks, D.L. Reduced GM1 ganglioside in CFTR-deficient human airway cells results in decreased beta1-integrin signaling and delayed wound repair. Am. J. Physiol. Cell Physiol. 2014, 306, C819–C830. [Google Scholar] [CrossRef]
- Rogers, G.B.; KBruce, D.; Hoffman, L.R. How can the cystic fibrosis respiratory microbiome influence our clinical decision-making? Curr. Opin. Pulm. Med. 2017, 23, 536–543. [Google Scholar] [CrossRef] [PubMed]
- Vernocchi, P.; Del Chierico, F.; Russo, A.; Majo, F.; Rossitto, M.; Valerio, M.; Casadei, L.; La Storia, A.; De Filippis, F.; Rizzo, C.; et al. Gut microbiota signatures in cystic fibrosis: Loss of host CFTR function drives the microbiota enterophenotype. PLoS ONE 2018, 13, e0208171. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Leong, L.; Keating, R.L.; Kanno, T.; Abell, G.; Mobegi, F.M.; Choo, J.M.; Wesselingh, S.L.; Mason, A.J.; Burr, L.D.; et al. Opportunistic bacteria confer the ability to ferment prebiotic starch in the adult cystic fibrosis gut. Gut Microbes 2019, 10, 367–381. [Google Scholar] [CrossRef] [PubMed]
- Kaakoush, N.O.; Pickford, R.; Jaffe, A.; Ooi, C.Y. Is there a role for stool metabolomics in cystic fibrosis? Pediatr. Int. 2016, 58, 808–811. [Google Scholar] [CrossRef]
- Blanchard, A.C.; Waters, V.J. Microbiology of Cystic Fibrosis Airway Disease. Semin. Respir. Crit. Care Med. 2019, 40, 727–736. [Google Scholar] [CrossRef]
- Dong, K.; Moon, K.; Chen, V.; Raymond, N.G.; Foster, L.J.; Tebbutt, S.J.; Quon, B.S. Identification of novel blood biomarkers of treatment response in cystic fibrosis pulmonary exacerbations by label-free quantitative proteomics. Sci. Rep. 2019, 9, 1–9. [Google Scholar] [CrossRef]
- Roberts, J.M.; Darlene, L.Y.D.; Hollander, Z.; Ng, R.T.; Tebbutt, S.J.; Wilcox, P.G.; Sin, D.D.; Quon, B.S. Multiple reaction monitoring mass spectrometry to identify novel plasma protein biomarkers of treatment response in cystic fibrosis pulmonary exacerbations. J. Cyst. Fibros. 2018, 17, 333–340. [Google Scholar] [CrossRef]
- Shoki, A.H.; Mayer-Hamblett, N.; Wilcox, P.G.; Sin, D.D.; Quon, B.S. Systematic review of blood biomarkers in cystic fibrosis pulmonary exacerbations. Chest 2013, 144, 1659–1670. [Google Scholar] [CrossRef]
- Singh, A.; He, M.; Chen, V.; Hollander, Z.; Tebbutt, S.J.; Ng, R.T.; McManus, B.M.; Sin, D.D.; Ratjen, F.; Quon, B.S. Blood biomarkers to predict short-term pulmonary exacerbation risk in children and adolescents with CF: A pilot study. J. Cyst. Fibros. 2020, 19, 49–51. [Google Scholar] [CrossRef]
- Quon, B.S.; Dai, D.L.; Hollander, Z.; Ng, R.T.; Tebbutt, S.J.; Man, S.F.; Wilcox, P.G.; Sin, D.D. Discovery of novel plasma protein biomarkers to predict imminent cystic fibrosis pulmonary exacerbations using multiple reaction monitoring mass spectrometry. Thorax 2016, 71, 216–222. [Google Scholar] [CrossRef]
- Zang, X.; Monge, M.E.; Gaul, D.A.; McCarty, N.A.; Stecenko, A.; Fernández, F.M. Early Detection of Cystic Fibrosis Acute Pulmonary Exacerbations by Exhaled Breath Condensate Metabolomics. J. Proteome Res. 2020, 19, 144–152. [Google Scholar] [CrossRef] [PubMed]
- Zang, X.; Monge, M.E.; McCarty, N.A.; Stecenko, A.A.; Fernández, F.M. Feasibility of Early Detection of Cystic Fibrosis Acute Pulmonary Exacerbations by Exhaled Breath Condensate Metabolomics: A Pilot Study. J. Proteome Res. 2017, 16, 550–558. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.; Spaněl, P.; Gilchrist, F.J.; Lenney, W. Hydrogen cyanide, a volatile biomarker of Pseudomonas aeruginosa infection. J. Breath Res. 2013, 7, 044001. [Google Scholar] [CrossRef] [PubMed]
- Gilchrist, F.J.; Bright-Thomas, R.J.; Jones, A.M.; Smith, D.; Spaněl, P.; Webb, A.K.; Lenney, W. Hydrogen cyanide concentrations in the breath of adult cystic fibrosis patients with and without Pseudomonas aeruginosa infection. J. Breath Res. 2013, 7, 026010. [Google Scholar] [CrossRef]
- Dummer, J.; Storer, M.; Sturney, S.; Scott-Thomas, A.; Chambers, S.; Swanney, M.; Epton, M. Quantification of hydrogen cyanide (HCN) in breath using selected ion flow tube mass spectrometry--HCN is not a biomarker of Pseudomonas in chronic suppurative lung disease. J. Breath Res. 2013, 7, 017105. [Google Scholar] [CrossRef]
- Shestivska, V.; Nemec, A.; Dřevínek, P.; Sovová, K.; Dryahina, K.; Spaněl, P. Quantification of methyl thiocyanate in the headspace of Pseudomonas aeruginosa cultures and in the breath of cystic fibrosis patients by selected ion flow tube mass spectrometry. Rapid Commun. Mass Spectrom. 2011, 25, 2459–2467. [Google Scholar] [CrossRef]
- Spanel, P.; Sovová, K.; Dryahina, K.; Doušová, T.; Dřevínek, P.; Smith, D. Do linear logistic model analyses of volatile biomarkers in exhaled breath of cystic fibrosis patients reliably indicate Pseudomonas aeruginosa infection? J. Breath Res. 2016, 10, 036013. [Google Scholar] [CrossRef]
- Robroeks, C.M.; van Berkel, J.J.; Dallinga, J.W.; Jöbsis, Q.; Zimmermann, L.J.; Hendriks, H.J.; Wouters, M.F.; van der Grinten, C.P.; van de Kant, K.D.; van Schooten, F.J.; et al. Metabolomics of volatile organic compounds in cystic fibrosis patients and controls. Pediatr. Res. 2010, 68, 75–80. [Google Scholar] [CrossRef]
- Glasser, N.R.; Hunter, R.C.; Liou, T.G.; Newman, D.K.; Mountain West CF Consortium Investigators. Refinement of metabolite detection in cystic fibrosis sputum reveals heme correlates with lung function decline. PLoS ONE 2019, 14, e0226578. [Google Scholar] [CrossRef]
- Quinn, R.A.; Lim, Y.W.; Mak, T.D.; Whiteson, K.; Furlan, M.; Conrad, D.; Rohwer, F.; Dorrestein, P. Metabolomics of pulmonary exacerbations reveals the personalized nature of cystic fibrosis disease. PeerJ 2016, 4, e2174. [Google Scholar] [CrossRef]
- Berkefeld, H.; Sailer, C.A.; Bildl, W.; Rohde, V.; Thumfart, J.O.; Eble, S.; Klugbauer, N.; Reisinger, E.; Bischofberger, J.; Oliver, D.; et al. BKCa-Cav channel complexes mediate rapid and localized Ca2+-activated K+ signaling. Science 2006, 314, 615–620. [Google Scholar] [CrossRef] [PubMed]
- Schwenk, J.; Harmel, N.; Zolles, G.; Bildl, W.; Kulik, A.; Heimrich, B.; Chisaka, O.; Jonas, P.; Schulte, U.; Fakler, B.; et al. Functional proteomics identify cornichon proteins as auxiliary subunits of AMPA receptors. Science 2009, 323, 1313–1319. [Google Scholar] [CrossRef] [PubMed]
- Santos, S.D.; Manadas, B.; Duarte, C.B.; Carvalho, A.L. Proteomic analysis of an interactome for long-form AMPA receptor subunits. J. Proteome Res. 2010, 9, 1670–1682. [Google Scholar] [CrossRef]
- Zeitler, S.; Ye, L.; Andreyeva, A.; Schumacher, F.; Monti, J.; Nürnberg, B.; Nowak, G.; Kleuser, B.; Reichel, M.; Fejtová, A.; et al. Acid sphingomyelinase - a regulator of canonical transient receptor potential channel 6 (TRPC6) activity. J. Neurochem. 2019, 150, 678–690. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, V.; Sidore, M.; Bechara, C.; Duneau, J.P.; Sturgis, J.N. The lipid environment of Escherichia coli Aquaporin Z. Biochim. Biophys. Acta Biomembr. 2019, 1861, 431–440. [Google Scholar] [CrossRef] [PubMed]
- Sinz, A. Cross-Linking/Mass Spectrometry for Studying Protein Structures and Protein-Protein Interactions: Where Are We Now and Where Should We Go from Here? Angew Chem. Int. Ed. Engl. 2018, 57, 6390–6396. [Google Scholar] [CrossRef]
- Chavez, J.D.; Mohr, J.P.; Mathay, M.; Zhong, X.; Keller, A.; Bruce, J.E. Systems structural biology measurements by in vivo cross-linking with mass spectrometry. Nat. Protoc. 2019, 14, 2318–2343. [Google Scholar] [CrossRef]
- Iacobucci, C.; Götze, M.; Ihling, C.H.; Piotrowski, C.; Arlt, C.; Schäfer, M.; Hage, C.; Schmidt, R.; Sinz, A. A cross-linking/mass spectrometry workflow based on MS-cleavable cross-linkers and the MeroX software for studying protein structures and protein-protein interactions. Nat. Protoc. 2018, 13, 2864–2889. [Google Scholar] [CrossRef]
- Bobadilla, J.L.; Macek, M., Jr.; Fine, J.P.; Farrell, P.M. Cystic fibrosis: A worldwide analysis of CFTR mutations--correlation with incidence data and application to screening. Hum. Mutat. 2002, 19, 575–606. [Google Scholar] [CrossRef]
- Vizcaino, J.A.; Csordas, A.; del-Toro, N.; Dianes, J.A.; Griss, J.; Lavidas, I.; Mayer, G.; Perez-Riverol, Y.; Reisinger, F.; Ternent, T.; et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 2016, 44, 11033. [Google Scholar] [CrossRef]
- Kale, N.S.; Haug, K.; Conesa, P.; Jayseelan, K.; Moreno, P.; Rocca-Serra, P.; Nainala, V.C.; Spicer, R.A.; Williams, M.; Li, X.; et al. MetaboLights: An. Open-Access Database Repository for Metabolomics Data. Curr. Protoc. Bioinform. 2016, 53, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Kelchtermans, P.; Bittremieux, W.; De Grave, K.; Degroeve, S.; Ramon, J.; Laukens, K.; Valkenborg, D.; Barsnes, H.; Martens, L. Machine learning applications in proteomics research: How the past can boost the future. Proteomics 2014, 14, 353–366. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liessi, N.; Pedemonte, N.; Armirotti, A.; Braccia, C. Proteomics and Metabolomics for Cystic Fibrosis Research. Int. J. Mol. Sci. 2020, 21, 5439. https://doi.org/10.3390/ijms21155439
Liessi N, Pedemonte N, Armirotti A, Braccia C. Proteomics and Metabolomics for Cystic Fibrosis Research. International Journal of Molecular Sciences. 2020; 21(15):5439. https://doi.org/10.3390/ijms21155439
Chicago/Turabian StyleLiessi, Nara, Nicoletta Pedemonte, Andrea Armirotti, and Clarissa Braccia. 2020. "Proteomics and Metabolomics for Cystic Fibrosis Research" International Journal of Molecular Sciences 21, no. 15: 5439. https://doi.org/10.3390/ijms21155439
APA StyleLiessi, N., Pedemonte, N., Armirotti, A., & Braccia, C. (2020). Proteomics and Metabolomics for Cystic Fibrosis Research. International Journal of Molecular Sciences, 21(15), 5439. https://doi.org/10.3390/ijms21155439





