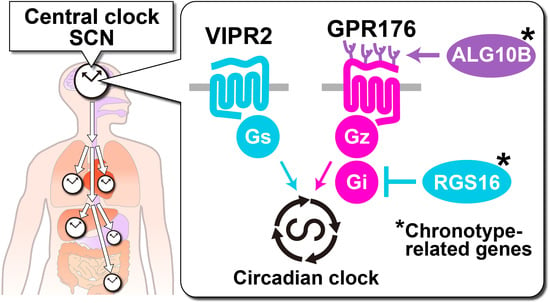
Time-Restricted G-Protein Signaling Pathways via GPR176, Gz, and RGS16 Set the Pace of the Master Circadian Clock in the Suprachiasmatic Nucleus
Abstract
1. Introduction
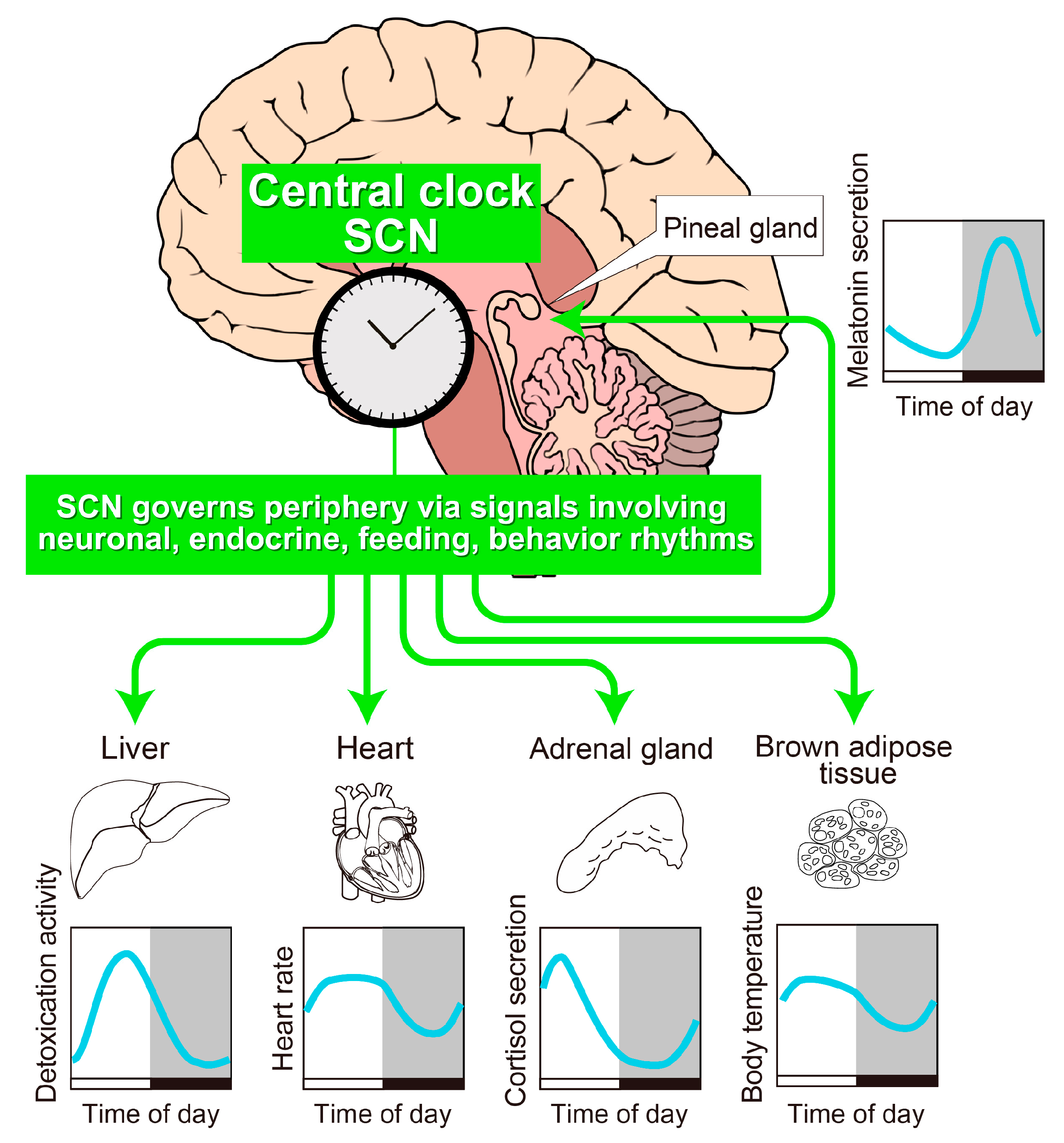
2. The SCN Is the Principal Circadian Pacemaker in Mammals
3. GPR176 Is a Gz-Linked Orphan GPCR that Sets the Pace of the SCN Clock
3.1. GPR176 Colocalizes with VIPR2 and Displays Oscillatory Abundance
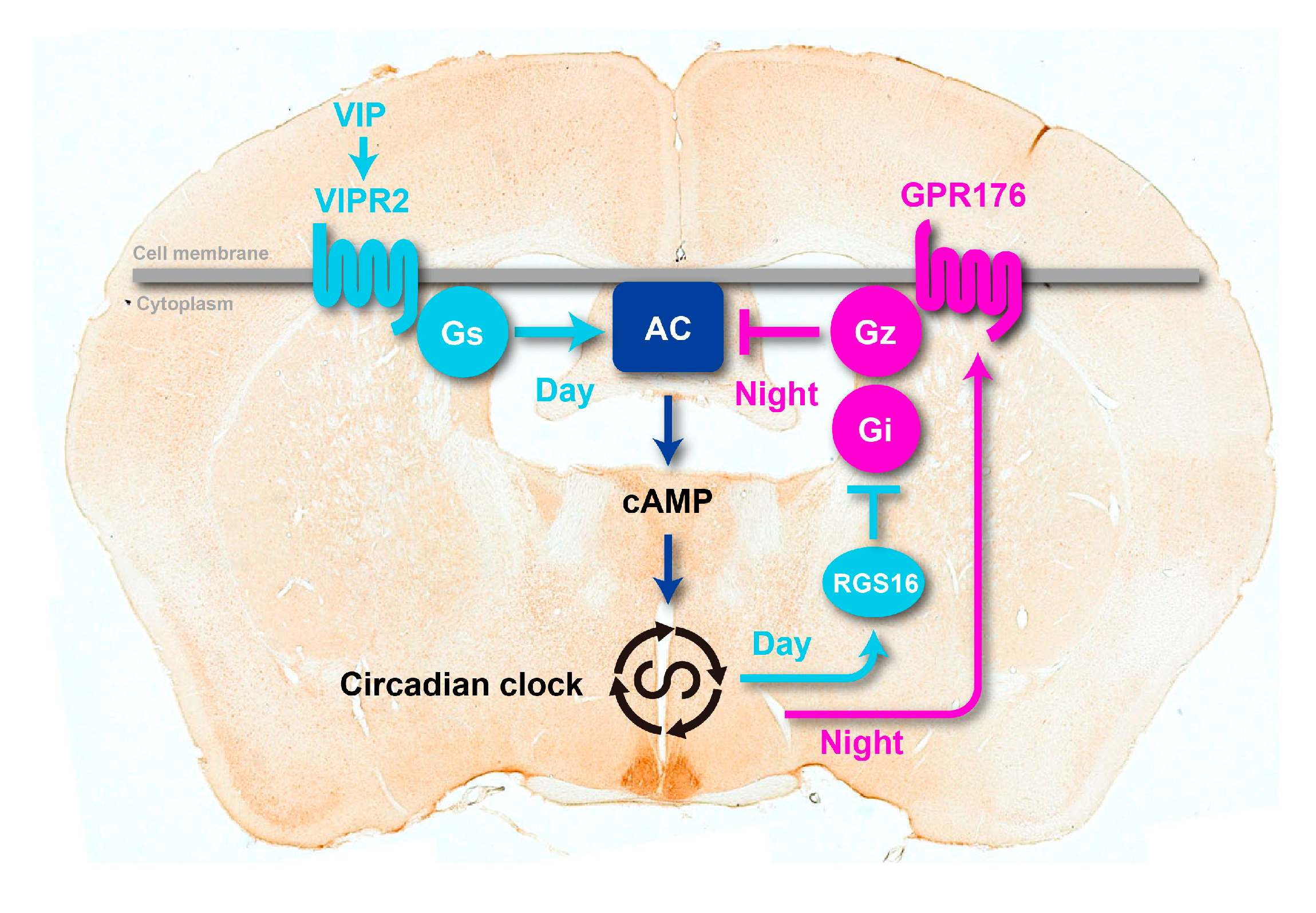
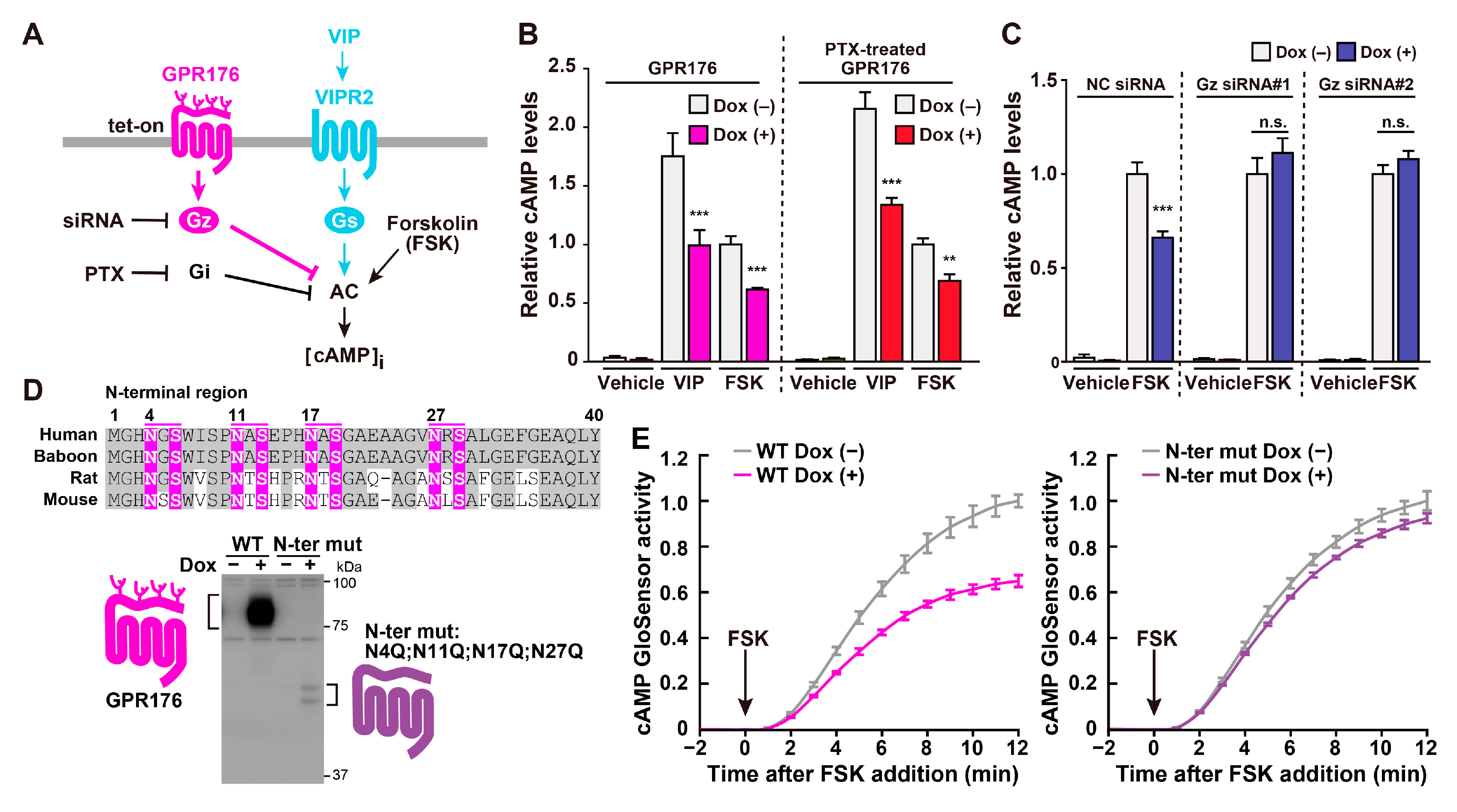
3.2. Agonist-Independent Activity of GPR176 Counteracts VIPR2-cAMP Signaling
3.3. The Unique G-Protein Subclass Gz Is Required for GPR176 Basal Activity
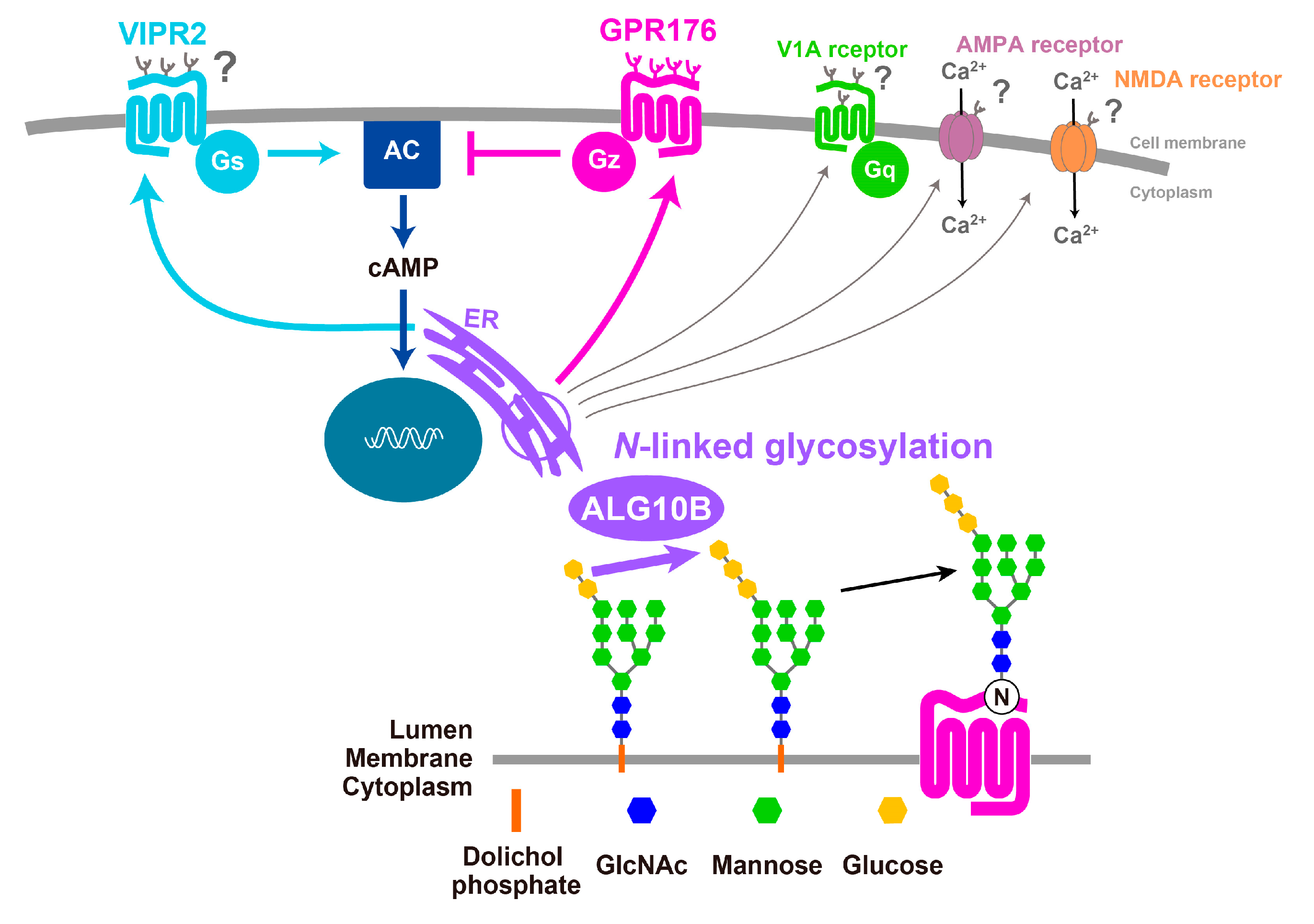
3.4. GPR176 Is an N-Glycosylated GPCR
4. RGS16 Is a Regulator of G-Protein-cAMP Signaling in the SCN
5. Advantage of Focusing on GPCRs in the SCN
6. Deorphanizing GPR176
7. Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Santos, R.; Ursu, O.; Gaulton, A.; Bento, A.P.; Donadi, R.S.; Bologa, C.G.; Karlsson, A.; Al-Lazikani, B.; Hersey, A.; Oprea, T.I.; et al. A comprehensive map of molecular drug targets. Nat. Rev. Drug Discov. 2017, 16, 19–34. [Google Scholar] [CrossRef] [PubMed]
- Hauser, A.S.; Attwood, M.M.; Rask-Andersen, M.; Schioth, H.B.; Gloriam, D.E. Trends in GPCR drug discovery: New agents, targets and indications. Nat. Rev. Drug Discov. 2017, 16, 829–842. [Google Scholar] [CrossRef] [PubMed]
- Wacker, D.; Stevens, R.C.; Roth, B.L. How Ligands Illuminate GPCR Molecular Pharmacology. Cell 2017, 170, 414–427. [Google Scholar] [CrossRef] [PubMed]
- Doi, M.; Takahashi, Y.; Komatsu, R.; Yamazaki, F.; Yamada, H.; Haraguchi, S.; Emoto, N.; Okuno, Y.; Tsujimoto, G.; Kanematsu, A.; et al. Salt-sensitive hypertension in circadian clock-deficient Cry-null mice involves dysregulated adrenal Hsd3b6. Nat. Med. 2010, 16, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Doi, M. Circadian clock-deficient mice as a tool for exploring disease etiology. Biol. Pharm. Bull. 2012, 35, 1385–1391. [Google Scholar] [CrossRef] [PubMed]
- Cederroth, C.R.; Albrecht, U.; Bass, J.; Brown, S.A.; Dyhrfjeld-Johnsen, J.; Gachon, F.; Green, C.B.; Hastings, M.H.; Helfrich-Forster, C.; Hogenesch, J.B.; et al. Medicine in the Fourth Dimension. Cell Metab. 2019, 30, 238–250. [Google Scholar] [CrossRef] [PubMed]
- Doi, M.; Murai, I.; Kunisue, S.; Setsu, G.; Uchio, N.; Tanaka, R.; Kobayashi, S.; Shimatani, H.; Hayashi, H.; Chao, H.W.; et al. Gpr176 is a Gz-linked orphan G-protein-coupled receptor that sets the pace of circadian behaviour. Nat. Commun. 2016, 7, 10583. [Google Scholar] [CrossRef]
- Wang, T.; Nakagawa, S.; Miyake, T.; Setsu, G.; Kunisue, S.; Goto, K.; Hirasawa, A.; Okamura, H.; Yamaguchi, Y.; Doi, M. Identification and functional characterisation of N-linked glycosylation of the orphan G protein-coupled receptor Gpr176. Sci. Rep. 2020, 10, 4429. [Google Scholar] [CrossRef]
- Martin, A.L.; Steurer, M.A.; Aronstam, R.S. Constitutive Activity among Orphan Class-A G Protein Coupled Receptors. PLoS ONE 2015, 10, e0138463. [Google Scholar] [CrossRef]
- Doi, M.; Ishida, A.; Miyake, A.; Sato, M.; Komatsu, R.; Yamazaki, F.; Kimura, I.; Tsuchiya, S.; Kori, H.; Seo, K.; et al. Circadian regulation of intracellular G-protein signalling mediates intercellular synchrony and rhythmicity in the suprachiasmatic nucleus. Nat. Commun. 2011, 2, 327. [Google Scholar] [CrossRef]
- Hayasaka, N.; Aoki, K.; Kinoshita, S.; Yamaguchi, S.; Wakefield, J.K.; Tsuji-Kawahara, S.; Horikawa, K.; Ikegami, H.; Wakana, S.; Murakami, T.; et al. Attenuated food anticipatory activity and abnormal circadian locomotor rhythms in Rgs16 knockdown mice. PLoS ONE 2011, 6, e17655. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Shmygelska, A.; Tran, D.; Eriksson, N.; Tung, J.Y.; Hinds, D.A. GWAS of 89,283 individuals identifies genetic variants associated with self-reporting of being a morning person. Nat. Commun. 2016, 7, 10448. [Google Scholar] [CrossRef] [PubMed]
- Lane, J.M.; Vlasac, I.; Anderson, S.G.; Kyle, S.D.; Dixon, W.G.; Bechtold, D.A.; Gill, S.; Little, M.A.; Luik, A.; Loudon, A.; et al. Genome-wide association analysis identifies novel loci for chronotype in 100,420 individuals from the UK Biobank. Nat. Commun. 2016, 7, 10889. [Google Scholar] [CrossRef] [PubMed]
- Jones, S.E.; Tyrrell, J.; Wood, A.R.; Beaumont, R.N.; Ruth, K.S.; Tuke, M.A.; Yaghootkar, H.; Hu, Y.; Teder-Laving, M.; Hayward, C.; et al. Genome-Wide Association Analyses in 128,266 Individuals Identifies New Morningness and Sleep Duration Loci. PLoS Genet. 2016, 12, e1006125. [Google Scholar] [CrossRef] [PubMed]
- Jones, S.E.; Lane, J.M.; Wood, A.R.; van Hees, V.T.; Tyrrell, J.; Beaumont, R.N.; Jeffries, A.R.; Dashti, H.S.; Hillsdon, M.; Ruth, K.S.; et al. Genome-wide association analyses of chronotype in 697,828 individuals provides insights into circadian rhythms. Nat. Commun. 2019, 10, 343. [Google Scholar] [CrossRef] [PubMed]
- Herzog, E.D.; Hermanstyne, T.; Smyllie, N.J.; Hastings, M.H. Regulating the Suprachiasmatic Nucleus (SCN) Circadian Clockwork: Interplay between Cell-Autonomous and Circuit-Level Mechanisms. Cold Spring Harb. Perspect Biol. 2017, 9, a027706. [Google Scholar] [CrossRef] [PubMed]
- Lehman, M.N.; Silver, R.; Gladstone, W.R.; Kahn, R.M.; Gibson, M.; Bittman, E.L. Circadian rhythmicity restored by neural transplant. Immunocytochemical characterization of the graft and its integration with the host brain. J. Neurosci. 1987, 7, 1626–1638. [Google Scholar] [CrossRef]
- Ralph, M.R.; Foster, R.G.; Davis, F.C.; Menaker, M. Transplanted suprachiasmatic nucleus determines circadian period. Science 1990, 247, 975–978. [Google Scholar] [CrossRef]
- Sujino, M.; Masumoto, K.H.; Yamaguchi, S.; van der Horst, G.T.; Okamura, H.; Inouye, S.T. Suprachiasmatic nucleus grafts restore circadian behavioral rhythms of genetically arrhythmic mice. Curr. Biol. 2003, 13, 664–668. [Google Scholar] [CrossRef]
- Lee, I.T.; Chang, A.S.; Manandhar, M.; Shan, Y.; Fan, J.; Izumo, M.; Ikeda, Y.; Motoike, T.; Dixon, S.; Seinfeld, J.E.; et al. Neuromedin s-producing neurons act as essential pacemakers in the suprachiasmatic nucleus to couple clock neurons and dictate circadian rhythms. Neuron 2015, 85, 1086–1102. [Google Scholar] [CrossRef]
- Mieda, M.; Ono, D.; Hasegawa, E.; Okamoto, H.; Honma, K.; Honma, S.; Sakurai, T. Cellular clocks in AVP neurons of the SCN are critical for interneuronal coupling regulating circadian behavior rhythm. Neuron 2015, 85, 1103–1116. [Google Scholar] [CrossRef]
- Smyllie, N.J.; Chesham, J.E.; Hamnett, R.; Maywood, E.S.; Hastings, M.H. Temporally chimeric mice reveal flexibility of circadian period-setting in the suprachiasmatic nucleus. Proc. Natl. Acad. Sci. USA 2016, 113, 3657–3662. [Google Scholar] [CrossRef] [PubMed]
- Miyake, T.; Doi, M. Reconstitution of organismal liver clock function requires light. Trends Endocrinol. Metab. 2019, 30, 569–571. [Google Scholar] [CrossRef] [PubMed]
- Koronowski, K.B.; Kinouchi, K.; Welz, P.S.; Smith, J.G.; Zinna, V.M.; Shi, J.; Samad, M.; Chen, S.; Magnan, C.N.; Kinchen, J.M.; et al. Defining the Independence of the Liver Circadian Clock. Cell 2019, 177, 1448–1462.e14. [Google Scholar] [CrossRef] [PubMed]
- Welz, P.S.; Zinna, V.M.; Symeonidi, A.; Koronowski, K.B.; Kinouchi, K.; Smith, J.G.; Guillen, I.M.; Castellanos, A.; Crainiciuc, G.; Prats, N.; et al. BMAL1-driven tissue clocks respond independently to light to maintain homeostasis. Cell 2019, 177, 1436–1447.e12. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, J.S. Transcriptional architecture of the mammalian circadian clock. Nat. Rev. Genet. 2017, 18, 164–179. [Google Scholar] [CrossRef]
- O’Neill, J.S.; Reddy, A.B. Circadian clocks in human red blood cells. Nature 2011, 469, 498–503. [Google Scholar] [CrossRef]
- Edgar, R.S.; Green, E.W.; Zhao, Y.; van Ooijen, G.; Olmedo, M.; Qin, X.; Xu, Y.; Pan, M.; Valekunja, U.K.; Feeney, K.A.; et al. Peroxiredoxins are conserved markers of circadian rhythms. Nature 2012, 485, 459–464. [Google Scholar] [CrossRef]
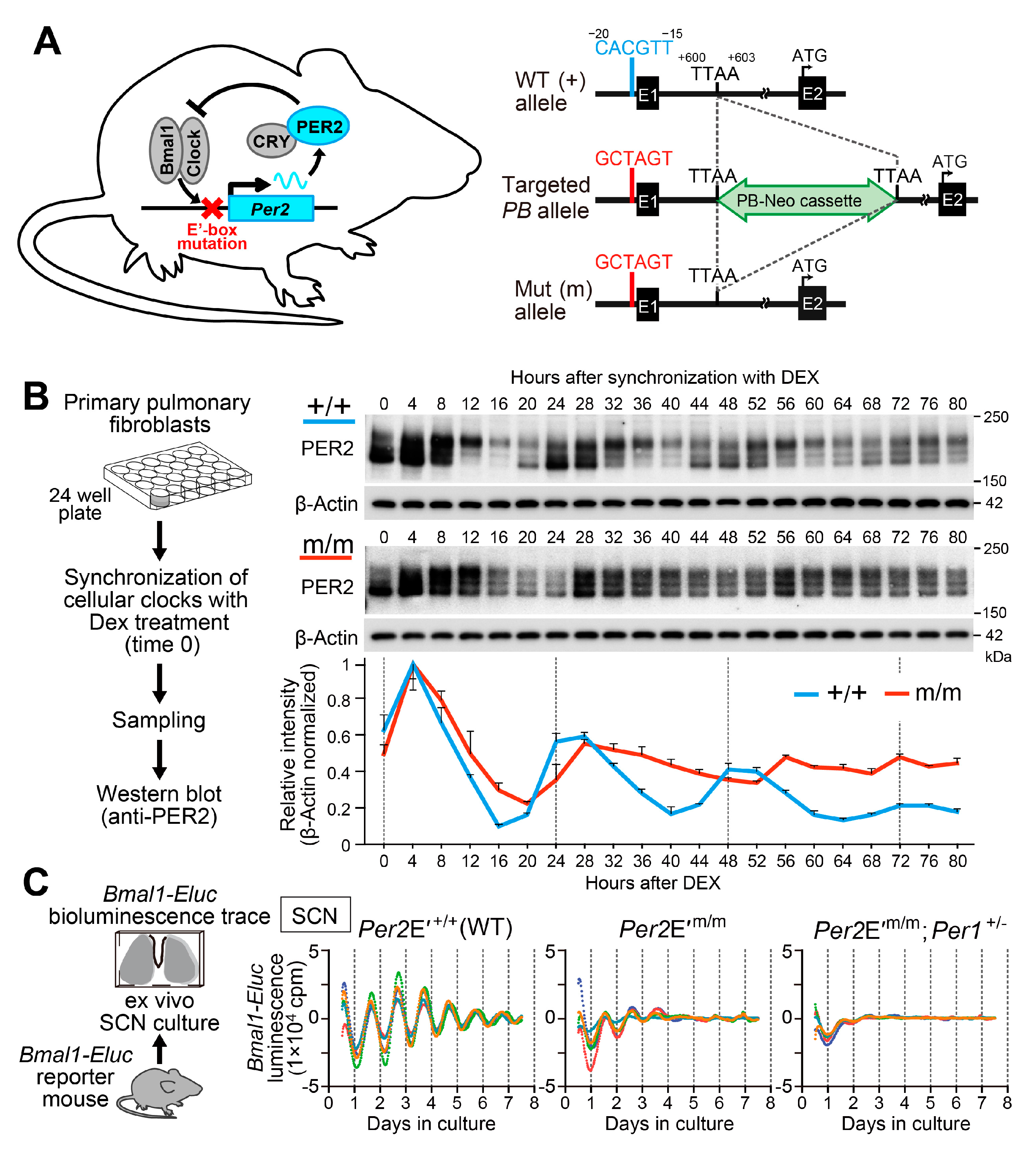
- Doi, M.; Shimatani, H.; Atobe, Y.; Murai, I.; Hayashi, H.; Takahashi, Y.; Fustin, J.M.; Yamaguchi, Y.; Kiyonari, H.; Koike, N.; et al. Non-coding cis-element of Period2 is essential for maintaining organismal circadian behaviour and body temperature rhythmicity. Nat. Commun. 2019, 10, 2563. [Google Scholar] [CrossRef]
- Mermet, J.; Yeung, J.; Hurni, C.; Mauvoisin, D.; Gustafson, K.; Jouffe, C.; Nicolas, D.; Emmenegger, Y.; Gobet, C.; Franken, P.; et al. Clock-dependent chromatin topology modulates circadian transcription and behavior. Genes Dev. 2018, 32, 347–358. [Google Scholar] [CrossRef]
- Hastings, M.H.; Maywood, E.S.; Brancaccio, M. Generation of circadian rhythms in the suprachiasmatic nucleus. Nat. Rev. Neurosci. 2018, 19, 453–469. [Google Scholar] [CrossRef] [PubMed]
- Maywood, E.S.; Chesham, J.E.; O’Brien, J.A.; Hastings, M.H. A diversity of paracrine signals sustains molecular circadian cycling in suprachiasmatic nucleus circuits. Proc. Natl. Acad. Sci. USA 2011, 108, 14306–14311. [Google Scholar] [CrossRef] [PubMed]
- Aton, S.J.; Colwell, C.S.; Harmar, A.J.; Waschek, J.; Herzog, E.D. Vasoactive intestinal polypeptide mediates circadian rhythmicity and synchrony in mammalian clock neurons. Nat. Neurosci. 2005, 8, 476–483. [Google Scholar] [CrossRef]
- Goda, T.; Doi, M.; Umezaki, Y.; Murai, I.; Shimatani, H.; Chu, M.L.; Nguyen, V.H.; Okamura, H.; Hamada, F.N. Calcitonin receptors are ancient modulators for rhythms of preferential temperature in insects and body temperature in mammals. Genes Dev. 2018, 32, 140–155. [Google Scholar] [CrossRef]
- Goto, K.; Doi, M.; Wang, T.; Kunisue, S.; Murai, I.; Okamura, H. G-protein-coupled receptor signaling through Gpr176, Gz, and RGS16 tunes time in the center of the circadian clock [Review]. Endocr. J. 2017, 64, 571–579. [Google Scholar] [CrossRef] [PubMed]
- Goda, T.; Hamada, F.N. Drosophila Temperature Preference Rhythms: An Innovative Model to Understand Body Temperature Rhythms. Int. J. Mol. Sci. 2019, 20, 1988. [Google Scholar] [CrossRef] [PubMed]
- An, S.; Tsai, C.; Ronecker, J.; Bayly, A.; Herzog, E.D. Spatiotemporal distribution of vasoactive intestinal polypeptide receptor 2 in mouse suprachiasmatic nucleus. J. Comp. Neurol. 2012, 520, 2730–2741. [Google Scholar] [CrossRef]
- O’Neill, J.S.; Maywood, E.S.; Chesham, J.E.; Takahashi, J.S.; Hastings, M.H. cAMP-dependent signaling as a core component of the mammalian circadian pacemaker. Science 2008, 320, 949–953. [Google Scholar] [CrossRef]
- Ho, M.K.; Wong, Y.H. G(z) signaling: Emerging divergence from G(i) signaling. Oncogene 2001, 20, 1615–1625. [Google Scholar] [CrossRef] [PubMed]
- Graziano, M.P.; Freissmuth, M.; Gilman, A.G. Expression of Gs alpha in Escherichia coli. Purification and properties of two forms of the protein. J. Biol. Chem. 1989, 264, 409–418. [Google Scholar]
- Linder, M.E.; Ewald, D.A.; Miller, R.J.; Gilman, A.G. Purification and characterization of Go alpha and three types of Gi alpha after expression in Escherichia coli. J. Biol. Chem. 1990, 265, 8243–8251. [Google Scholar] [PubMed]
- Casey, P.J.; Fong, H.K.; Simon, M.I.; Gilman, A.G. Gz, a guanine nucleotide-binding protein with unique biochemical properties. J. Biol. Chem. 1990, 265, 2383–2390. [Google Scholar] [PubMed]
- Vancura, P.; Abdelhadi, S.; Csicsely, E.; Baba, K.; Tosini, G.; Iuvone, P.M.; Spessert, R. Gnaz couples the circadian and dopaminergic system to G protein-mediated signaling in mouse photoreceptors. PLoS ONE 2017, 12, e0187411. [Google Scholar] [CrossRef]
- Barr, A.J.; Brass, L.F.; Manning, D.R. Reconstitution of receptors and GTP-binding regulatory proteins (G proteins) in Sf9 cells. A direct evaluation of selectivity in receptor.G protein coupling. J. Biol. Chem. 1997, 272, 2223–2229. [Google Scholar] [CrossRef] [PubMed]
- Serres, F.; Li, Q.; Garcia, F.; Raap, D.K.; Battaglia, G.; Muma, N.A.; Van de Kar, L.D. Evidence that G(z)-proteins couple to hypothalamic 5-HT(1A) receptors in vivo. J. Neurosci. 2000, 20, 3095–3103. [Google Scholar] [CrossRef] [PubMed]
- Yung, L.Y.; Tsim, S.T.; Wong, Y.H. Stimulation of cAMP accumulation by the cloned Xenopus melatonin receptor through Gi and Gz proteins. FEBS Lett. 1995, 372, 99–102. [Google Scholar] [CrossRef]
- Chan, A.S.; Lai, F.P.; Lo, R.K.; Voyno-Yasenetskaya, T.A.; Stanbridge, E.J.; Wong, Y.H. Melatonin mt1 and MT2 receptors stimulate c-Jun N-terminal kinase via pertussis toxin-sensitive and -insensitive G proteins. Cell Signal. 2002, 14, 249–257. [Google Scholar] [CrossRef]
- Tansky, M.F.; Pothoulakis, C.; Leeman, S.E. Functional consequences of alteration of N-linked glycosylation sites on the neurokinin 1 receptor. Proc. Natl. Acad. Sci. USA 2007, 104, 10691–10696. [Google Scholar] [CrossRef]
- Kozielewicz, P.; Alomar, H.; Yusof, S.; Grafton, G.; Cooper, A.J.; Curnow, S.J.; Ironside, J.W.; Pall, H.; Barnes, N.M. N-glycosylation and expression in human tissues of the orphan GPR61 receptor. FEBS Open Bio 2017, 7, 1982–1993. [Google Scholar] [CrossRef]
- Lutz, E.M.; Sheward, W.J.; West, K.M.; Morrow, J.A.; Fink, G.; Harmar, A.J. The VIP2 receptor: Molecular characterisation of a cDNA encoding a novel receptor for vasoactive intestinal peptide. FEBS Lett. 1993, 334, 3–8. [Google Scholar] [CrossRef]
- Hawtin, S.R.; Davies, A.R.; Matthews, G.; Wheatley, M. Identification of the glycosylation sites utilized on the V1a vasopressin receptor and assessment of their role in receptor signalling and expression. Biochem. J. 2001, 357, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, Y.; Suzuki, T.; Mizoro, Y.; Kori, H.; Okada, K.; Chen, Y.; Fustin, J.M.; Yamazaki, F.; Mizuguchi, N.; Zhang, J.; et al. Mice genetically deficient in vasopressin V1a and V1b receptors are resistant to jet lag. Science 2013, 342, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Lichnerova, K.; Kaniakova, M.; Park, S.P.; Skrenkova, K.; Wang, Y.X.; Petralia, R.S.; Suh, Y.H.; Horak, M. Two N-glycosylation Sites in the GluN1 Subunit Are Essential for Releasing N-methyl-d-aspartate (NMDA) Receptors from the Endoplasmic Reticulum. J. Biol. Chem. 2015, 290, 18379–18390. [Google Scholar] [CrossRef] [PubMed]
- Kandel, M.B.; Yamamoto, S.; Midorikawa, R.; Morise, J.; Wakazono, Y.; Oka, S.; Takamiya, K. N-glycosylation of the AMPA-type glutamate receptor regulates cell surface expression and tetramer formation affecting channel function. J. Neurochem. 2018, 147, 730–747. [Google Scholar] [CrossRef] [PubMed]
- Bansal, G.; Druey, K.M.; Xie, Z. R4 RGS proteins: Regulation of G-protein signaling and beyond. Pharmacol. Ther. 2007, 116, 473–495. [Google Scholar] [CrossRef] [PubMed]
- Mure, L.S.; Le, H.D.; Benegiamo, G.; Chang, M.W.; Rios, L.; Jillani, N.; Ngotho, M.; Kariuki, T.; Dkhissi-Benyahya, O.; Cooper, H.M.; et al. Diurnal transcriptome atlas of a primate across major neural and peripheral tissues. Science 2018, 359, eaao0318. [Google Scholar] [CrossRef]
- Yamazaki, S.; Numano, R.; Abe, M.; Hida, A.; Takahashi, R.; Ueda, M.; Block, G.D.; Sakaki, Y.; Menaker, M.; Tei, H. Resetting central and peripheral circadian oscillators in transgenic rats. Science 2000, 288, 682–685. [Google Scholar] [CrossRef]
- Foster, S.R.; Hauser, A.S.; Vedel, L.; Strachan, R.T.; Huang, X.P.; Gavin, A.C.; Shah, S.D.; Nayak, A.P.; Haugaard-Kedstrom, L.M.; Penn, R.B.; et al. Discovery of human signaling systems: Pairing peptides to G Protein-Coupled receptors. Cell 2019, 179, 895–908.e21. [Google Scholar] [CrossRef]
- Kroeze, W.K.; Sassano, M.F.; Huang, X.P.; Lansu, K.; McCorvy, J.D.; Giguere, P.M.; Sciaky, N.; Roth, B.L. PRESTO-Tango as an open-source resource for interrogation of the druggable human GPCRome. Nat. Struct. Mol. Biol. 2015, 22, 362–369. [Google Scholar] [CrossRef]
- Colosimo, D.A.; Kohn, J.A.; Luo, P.M.; Piscotta, F.J.; Han, S.M.; Pickard, A.J.; Rao, A.; Cross, J.R.; Cohen, L.J.; Brady, S.F. Mapping interactions of microbial metabolites with human G-Protein-Coupled receptors. Cell Host Microbe 2019, 26, 273–282.e7. [Google Scholar] [CrossRef]
- Cohen, L.J.; Esterhazy, D.; Kim, S.H.; Lemetre, C.; Aguilar, R.R.; Gordon, E.A.; Pickard, A.J.; Cross, J.R.; Emiliano, A.B.; Han, S.M.; et al. Commensal bacteria make GPCR ligands that mimic human signalling molecules. Nature 2017, 549, 48–53. [Google Scholar] [CrossRef] [PubMed]
| Subtype | kcat (min–1) | Source | Ref |
|---|---|---|---|
| Gαs | 4.5 | bovine, recombinant, purified from E. coli | [40] |
| Gαi1 | 2.4 | rat, recombinant, purified from E. coli | [41] |
| Gαi2 | 2.7 | rat, recombinant, purified from E. coli | [41] |
| Gαi3 | 1.8 | rat, recombinant, purified from E. coli | [41] |
| Gαo | 2.2 | rat, recombinant, purified from E. coli | [41] |
| Gαz | 0.05 | human, recombinant, purified from E. coli | [42] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nakagawa, S.; Nguyen Pham, K.T.; Shao, X.; Doi, M. Time-Restricted G-Protein Signaling Pathways via GPR176, Gz, and RGS16 Set the Pace of the Master Circadian Clock in the Suprachiasmatic Nucleus. Int. J. Mol. Sci. 2020, 21, 5055. https://doi.org/10.3390/ijms21145055
Nakagawa S, Nguyen Pham KT, Shao X, Doi M. Time-Restricted G-Protein Signaling Pathways via GPR176, Gz, and RGS16 Set the Pace of the Master Circadian Clock in the Suprachiasmatic Nucleus. International Journal of Molecular Sciences. 2020; 21(14):5055. https://doi.org/10.3390/ijms21145055
Chicago/Turabian StyleNakagawa, Shumpei, Khanh Tien Nguyen Pham, Xinyan Shao, and Masao Doi. 2020. "Time-Restricted G-Protein Signaling Pathways via GPR176, Gz, and RGS16 Set the Pace of the Master Circadian Clock in the Suprachiasmatic Nucleus" International Journal of Molecular Sciences 21, no. 14: 5055. https://doi.org/10.3390/ijms21145055
APA StyleNakagawa, S., Nguyen Pham, K. T., Shao, X., & Doi, M. (2020). Time-Restricted G-Protein Signaling Pathways via GPR176, Gz, and RGS16 Set the Pace of the Master Circadian Clock in the Suprachiasmatic Nucleus. International Journal of Molecular Sciences, 21(14), 5055. https://doi.org/10.3390/ijms21145055