Contribution of Immune Cells to Glucocorticoid Receptor Expression in Breast Cancer
Abstract
1. Introduction
2. Results
2.1. Demographic and Clinical Characteristics
2.2. GR-high Breast Cancer has Better Survival
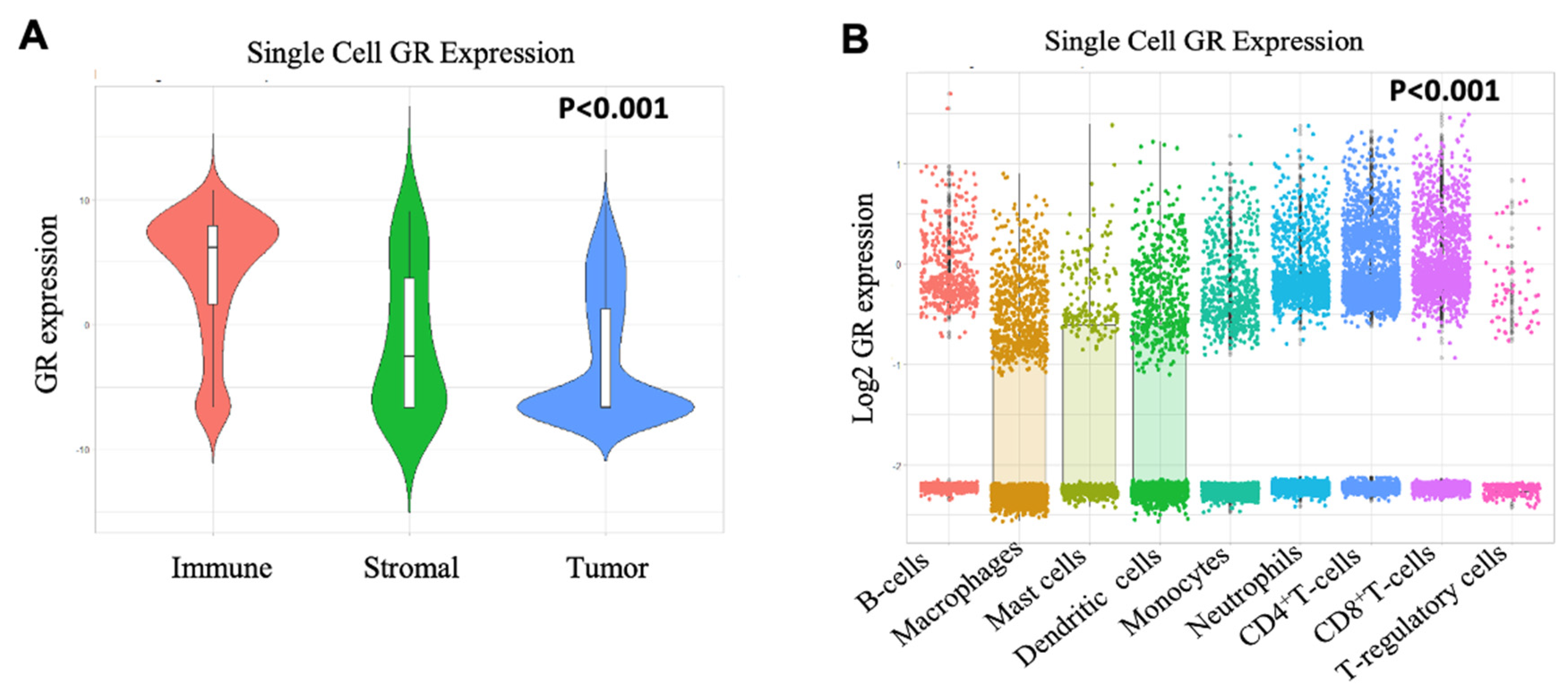
2.3. Immune Cells have High GR Expression than Tumor and Stromal Cells
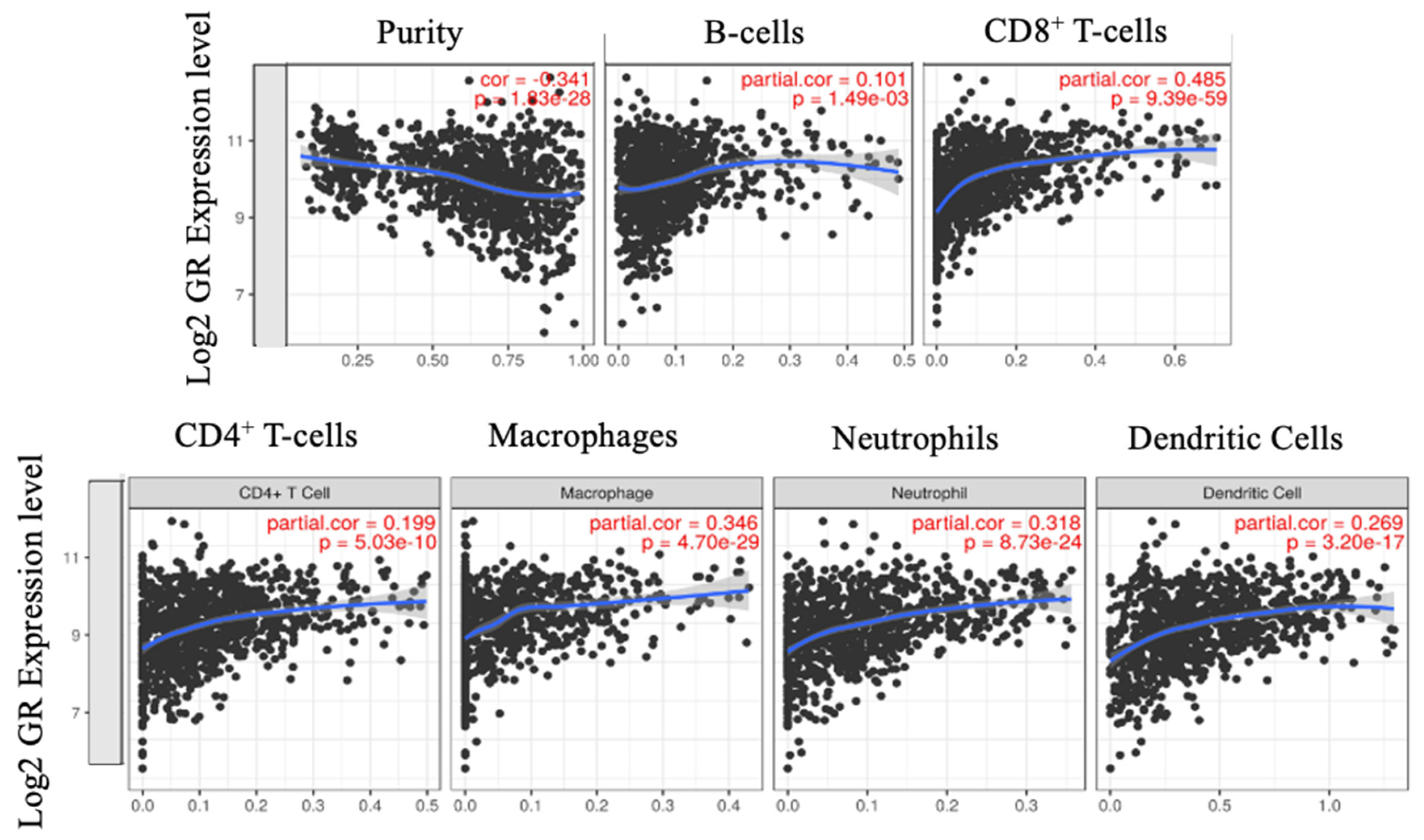
2.4. CD8+ T-cells Significantly Correlate with GR Expression
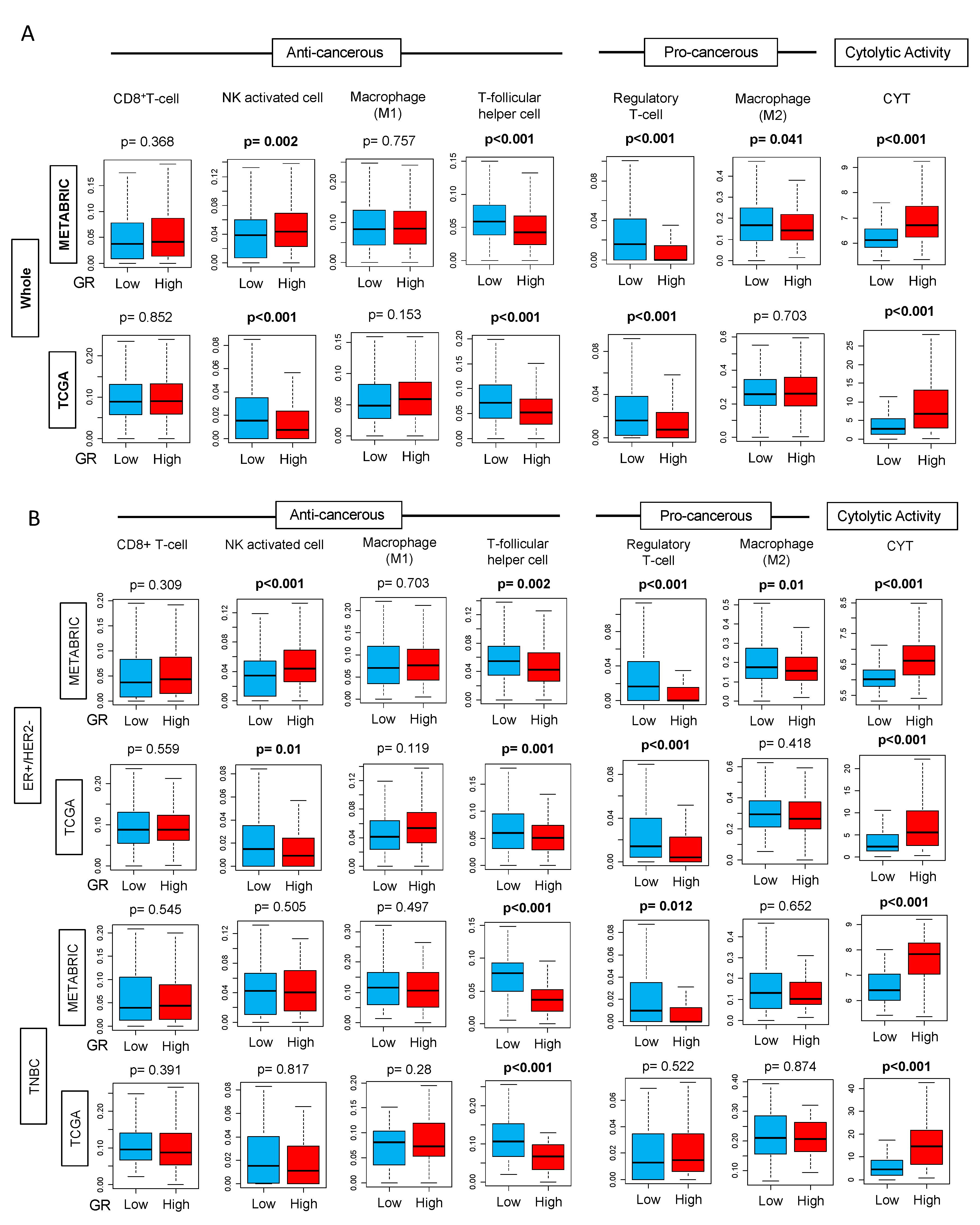
2.5. GR-high Breast Cancer has Higher Cytolytic Activity
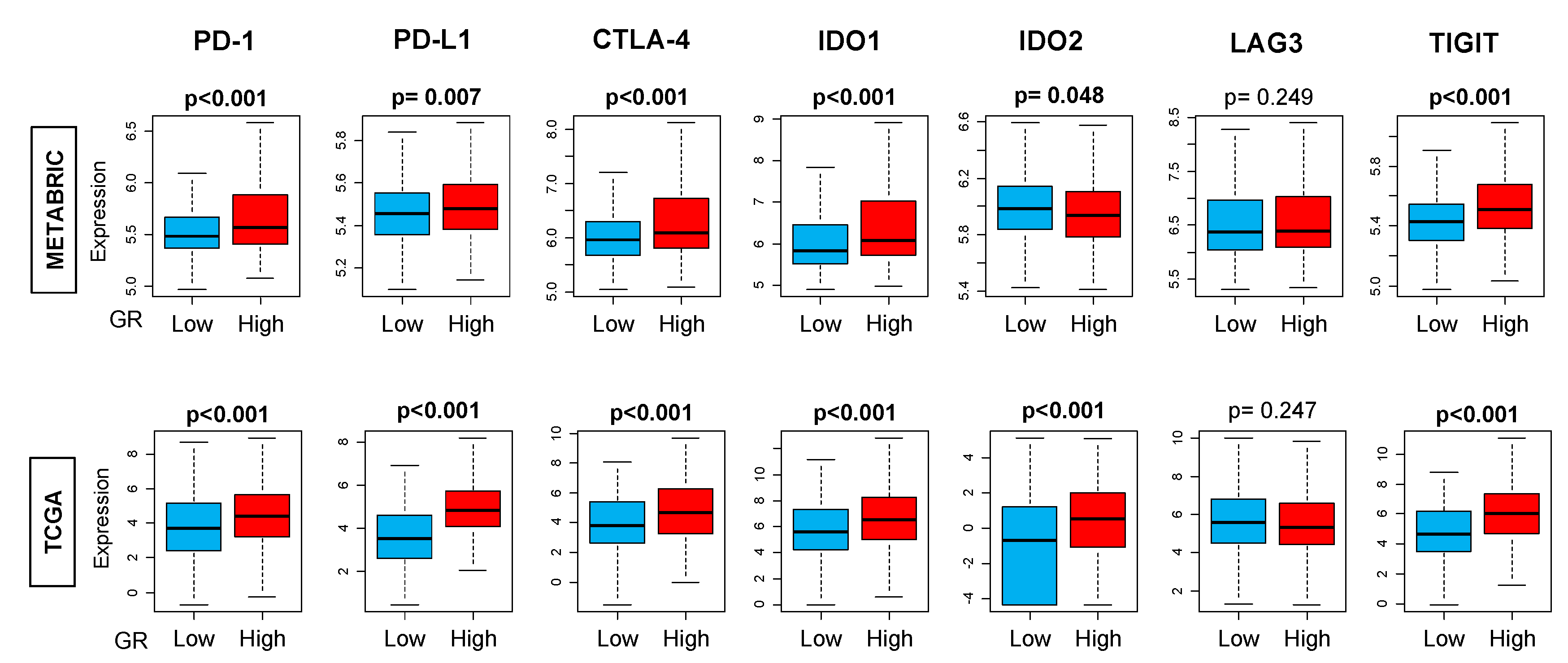
2.6. GR-high Breast Cancer has More T-cell Exhaustion Markers
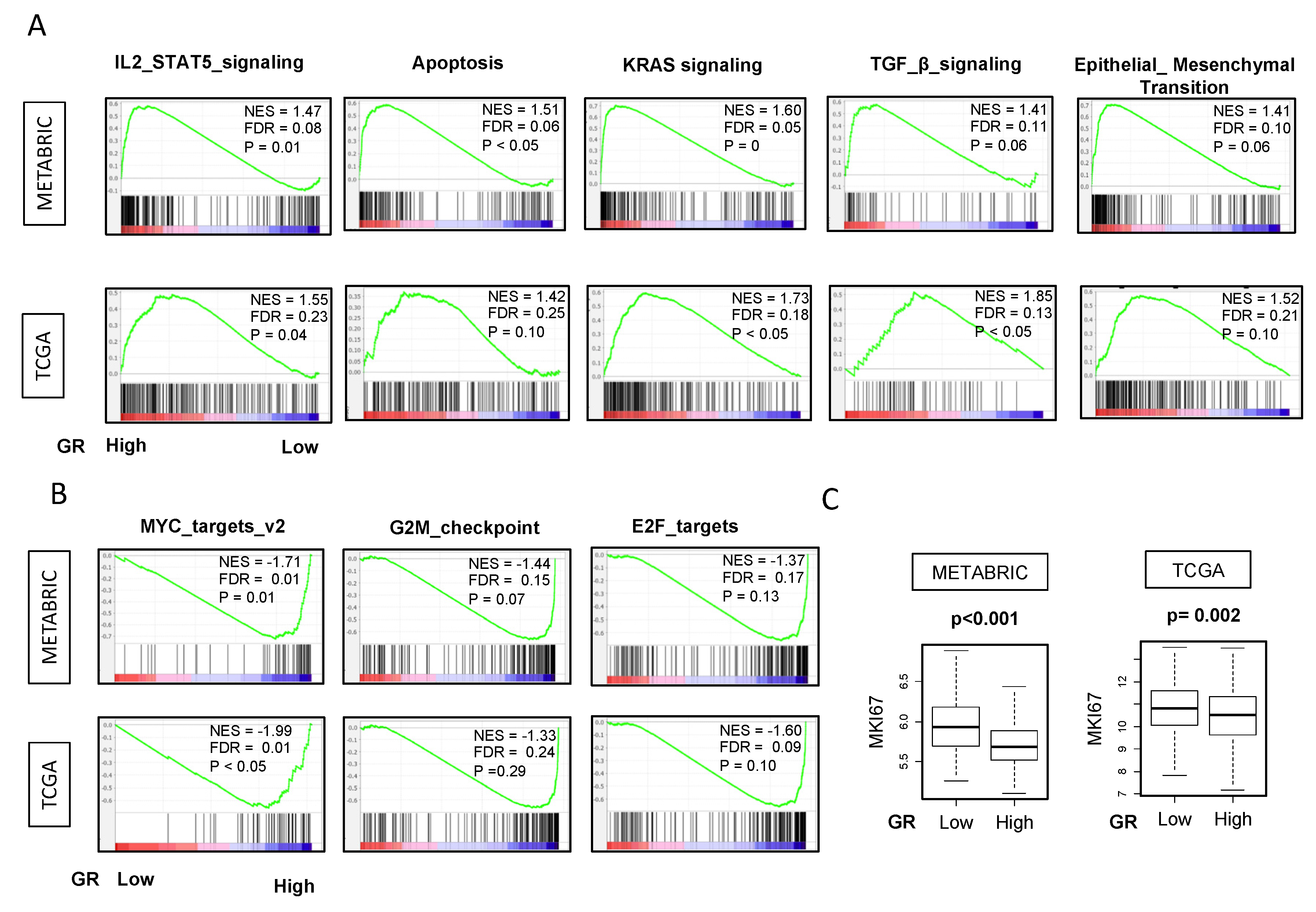
2.7. GR-high Breast Cancer is Enriched in IL2 STAT5, Apoptosis, KRAS, TGF-β, EMT Pathways and GR-low Breast Cancer has Higher Proliferation Markers
3. Discussion
4. Materials and Methods
4.1. Obtaining Data of METABRIC and TCGA
4.2. Immune Analysis Using CIBEROSRT Algorithm
4.3. Gene Set Expression Analysis
4.4. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ACTH | Adrenocorticotrophic hormone |
| DSS | Disease-specific survival |
| CYT | Cytolytic activity |
| DFS | Disease-free survival |
| ER | Estrogen receptor |
| EMT | Epithelial-to-mesenchymal transition |
| GR | Glucocorticoid receptor |
| GSEA | Gene set enrichment analysis |
| METABRIC | Molecular Taxonomy of Breast Cancer International Consortium |
| NK | Natural killer |
| OS | Overall survival |
| RFS | Relapse-free survival |
| TCGA | The Cancer Genome Atlas |
| TGF-β | Transforming growth factor-beta |
| TME | Tumor immune microenvironment |
| TNBC | Triple negative breast cancer |
| T-regs | Regulatory T-cells |
| GEO | Gene Expression Omnibus |
References
- Hwang, J.H.; Kim, W.Y.; Ahmed, M.; Choi, S.; Kim, J.; Han, D.W. The Use of Complementary and Alternative Medicine by Korean Breast Cancer Women: Is It Associated with Severity of Symptoms? Evid Based Complement. Alternat. Med. 2015, 2015, 182475. [Google Scholar] [CrossRef] [PubMed]
- Pinquart, M.; Duberstein, P.R. Depression and cancer mortality: A meta-analysis. Psychol. Med. 2010, 40, 1797–1810. [Google Scholar] [CrossRef] [PubMed]
- Palesh, O.; Butler, L.D.; Koopman, C.; Giese-Davis, J.; Carlson, R.; Spiegel, D. Stress history and breast cancer recurrence. J. Psychosom. Res. 2007, 63, 233–239. [Google Scholar] [CrossRef] [PubMed]
- Sephton, S.E.; Sapolsky, R.M.; Kraemer, H.C.; Spiegel, D. Diurnal cortisol rhythm as a predictor of breast cancer survival. J. Natl. Cancer Inst. 2000, 92, 994–1000. [Google Scholar] [CrossRef] [PubMed]
- Oakley, R.H.; Cidlowski, J.A. Cellular processing of the glucocorticoid receptor gene and protein: New mechanisms for generating tissue-specific actions of glucocorticoids. J. Biol. Chem. 2011, 286, 3177–3184. [Google Scholar] [CrossRef] [PubMed]
- Keller, S.E.; Weiss, J.M.; Schleifer, S.J.; Miller, N.E.; Stein, M. Stress-induced suppression of immunity in adrenalectomized rats. Science 1983, 221, 1301–1304. [Google Scholar] [CrossRef] [PubMed]
- Homo-Delarche, F.; Fitzpatrick, F.; Christeff, N.; Nunez, E.A.; Bach, J.F.; Dardenne, M. Sex steroids, glucocorticoids, stress and autoimmunity. J. Steroid Biochem. Mol. Biol. 1991, 40, 619–637. [Google Scholar] [CrossRef]
- Pang, D.; Kocherginsky, M.; Krausz, T.; Kim, S.Y.; Conzen, S.D. Dexamethasone decreases xenograft response to Paclitaxel through inhibition of tumor cell apoptosis. Cancer Biol. Ther. 2006, 5, 933–940. [Google Scholar] [CrossRef]
- Skor, M.N.; Wonder, E.L.; Kocherginsky, M.; Goyal, A.; Hall, B.A.; Cai, Y.; Conzen, S.D. Glucocorticoid receptor antagonism as a novel therapy for triple-negative breast cancer. Clin. Cancer Res. 2013, 19, 6163–6172. [Google Scholar] [CrossRef]
- Pan, D.; Kocherginsky, M.; Conzen, S.D. Activation of the glucocorticoid receptor is associated with poor prognosis in estrogen receptor-negative breast cancer. Cancer Res. 2011, 71, 6360–6370. [Google Scholar] [CrossRef]
- Obradovic, M.M.S.; Hamelin, B.; Manevski, N.; Couto, J.P.; Sethi, A.; Coissieux, M.M.; Munst, S.; Okamoto, R.; Kohler, H.; Schmidt, A.; et al. Glucocorticoids promote breast cancer metastasis. Nature 2019, 567, 540–544. [Google Scholar] [CrossRef] [PubMed]
- Nanda, R.; Stringer-Reasor, E.M.; Saha, P.; Kocherginsky, M.; Gibson, J.; Libao, B.; Hoffman, P.C.; Obeid, E.; Merkel, D.E.; Khramtsova, G.; et al. A randomized phase I trial of nanoparticle albumin-bound paclitaxel with or without mifepristone for advanced breast cancer. Springerplus 2016, 5, 947. [Google Scholar] [CrossRef] [PubMed]
- West, D.C.; Kocherginsky, M.; Tonsing-Carter, E.Y.; Dolcen, D.N.; Hosfield, D.J.; Lastra, R.R.; Sinnwell, J.P.; Thompson, K.J.; Bowie, K.R.; Harkless, R.V.; et al. Discovery of a Glucocorticoid Receptor (GR) Activity Signature Using Selective GR Antagonism in ER-Negative Breast Cancer. Clin. Cancer Res. 2018, 24, 3433–3446. [Google Scholar] [CrossRef] [PubMed]
- Loi, S.; Drubay, D.; Adams, S.; Pruneri, G.; Francis, P.A.; Lacroix-Triki, M.; Joensuu, H.; Dieci, M.V.; Badve, S.; Demaria, S.; et al. Tumor-Infiltrating Lymphocytes and Prognosis: A Pooled Individual Patient Analysis of Early-Stage Triple-Negative Breast Cancers. J. Clin. Oncol. 2019, 37, 559–569. [Google Scholar] [CrossRef]
- Glaser, R.; Kiecolt-Glaser, J.K. Stress-induced immune dysfunction: Implications for health. Nat. Rev. Immunol. 2005, 5, 243–251. [Google Scholar] [CrossRef]
- Cain, D.W.; Cidlowski, J.A. Immune regulation by glucocorticoids. Nat. Rev. Immunol. 2017, 17, 233–247. [Google Scholar] [CrossRef]
- Maeda, N.; Maruhashi, T.; Sugiura, D.; Shimizu, K.; Okazaki, I.M.; Okazaki, T. Glucocorticoids potentiate the inhibitory capacity of programmed cell death 1 by up-regulating its expression on T cells. J. Biol. Chem. 2019, 294, 19896–19906. [Google Scholar] [CrossRef]
- Hinrichs, C.S.; Palmer, D.C.; Rosenberg, S.A.; Restifo, N.P. Glucocorticoids do not inhibit antitumor activity of activated CD8+ T cells. J. Immunother. 2005, 28, 517–524. [Google Scholar] [CrossRef][Green Version]
- Eberwine, J.; Sul, J.Y.; Bartfai, T.; Kim, J. The promise of single-cell sequencing. Nat. Methods 2014, 11, 25–27. [Google Scholar] [CrossRef]
- Kach, J.; Conzen, S.D.; Szmulewitz, R.Z. Targeting the glucocorticoid receptor in breast and prostate cancers. Sci. Transl. Med. 2015, 7, 305ps19. [Google Scholar] [CrossRef]
- Zhou, Y.; Shao, N.; Aierken, N.; Xie, C.; Ye, R.; Qian, X.; Hu, Z.; Zhang, J.; Lin, Y. Prognostic value of tumor-infiltrating Foxp3+ regulatory T cells in patients with breast cancer: A meta-analysis. J. Cancer 2017, 8, 4098–4105. [Google Scholar] [CrossRef]
- Gu-Trantien, C.; Loi, S.; Garaud, S.; Equeter, C.; Libin, M.; de Wind, A.; Ravoet, M.; Le Buanec, H.; Sibille, C.; Manfouo-Foutsop, G.; et al. CD4(+) follicular helper T cell infiltration predicts breast cancer survival. J. Clin. Invest. 2013, 123, 2873–2892. [Google Scholar] [CrossRef] [PubMed]
- Wan, Y.Y.; Flavell, R.A. The roles for cytokines in the generation and maintenance of regulatory T cells. Immunol. Rev. 2006, 212, 114–130. [Google Scholar] [CrossRef] [PubMed]
- Imamura, T.; Hikita, A.; Inoue, Y. The roles of TGF-β signaling in carcinogenesis and breast cancer metastasis. Breast Cancer 2012, 19, 118–124. [Google Scholar] [CrossRef] [PubMed]
- Taylor, M.A.; Lee, Y.-H.; Schiemann, W.P. Role of TGF-β and the tumor microenvironment during mammary tumorigenesis. Gene Expr. J. Liver Res. 2011, 15, 117–132. [Google Scholar] [CrossRef] [PubMed]
- Felipe Lima, J.; Nofech-Mozes, S.; Bayani, J.; Bartlett, J.M. EMT in Breast Carcinoma-A Review. J. Clin. Med. 2016, 5, 65. [Google Scholar] [CrossRef]
- Hwang, K.T.; Kim, B.H.; Oh, S.; Park, S.Y.; Jung, J.; Kim, J.; Choi, I.S.; Jeon, S.Y.; Kim, W.Y. Prognostic Role of KRAS mRNA Expression in Breast Cancer. J. Breast Cancer 2019, 22, 548–561. [Google Scholar] [CrossRef]
- Denkert, C.; von Minckwitz, G.; Darb-Esfahani, S.; Lederer, B.; Heppner, B.I.; Weber, K.E.; Budczies, J.; Huober, J.; Klauschen, F.; Furlanetto, J.; et al. Tumour-infiltrating lymphocytes and prognosis in different subtypes of breast cancer: A pooled analysis of 3771 patients treated with neoadjuvant therapy. Lancet Oncol. 2018, 19, 40–50. [Google Scholar] [CrossRef]
- Bucsek, M.J.; Giridharan, T.; MacDonald, C.R.; Hylander, B.L.; Repasky, E.A. An overview of the role of sympathetic regulation of immune responses in infectious disease and autoimmunity. Int. J. Hyperth. 2018, 34, 135–143. [Google Scholar] [CrossRef]
- Cao, G.; Yang, Q.; Zhang, S.; Xu, C.; Roberts, A.I.; Wang, Y.; Shi, Y. Mesenchymal stem cells prevent restraint stress-induced lymphocyte depletion via interleukin-4. Brain Behav. Immun. 2014, 38, 125–132. [Google Scholar] [CrossRef]
- Lu, K.D.; Radom-Aizik, S.; Haddad, F.; Zaldivar, F.; Kraft, M.; Cooper, D.M. Glucocorticoid receptor expression on circulating leukocytes differs between healthy male and female adults. J. Clin. Transl. Sci. 2017, 1, 108–114. [Google Scholar] [CrossRef] [PubMed]
- Gola, H.; Engler, A.; Morath, J.; Adenauer, H.; Elbert, T.; Kolassa, I.T.; Engler, H. Reduced peripheral expression of the glucocorticoid receptor alpha isoform in individuals with posttraumatic stress disorder: A cumulative effect of trauma burden. PLoS ONE 2014, 9, e86333. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Liberman, A.C.; Budzinski, M.L.; Sokn, C.; Gobbini, R.P.; Steininger, A.; Arzt, E. Regulatory and Mechanistic Actions of Glucocorticoids on T and Inflammatory Cells. Front. Endocrinol. (Lausanne) 2018, 9, 235. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Chen, Y.A.; Conejo-Garcia, J.R.; Chung, C.H.; Wang, X. Estimation of immune cell content in tumor using single-cell RNA-seq reference data. BMC Cancer 2019, 19, 715. [Google Scholar] [CrossRef] [PubMed]
- Schelker, M.; Feau, S.; Du, J.; Ranu, N.; Klipp, E.; MacBeath, G.; Schoeberl, B.; Raue, A. Estimation of immune cell content in tumour tissue using single-cell RNA-seq data. Nat. Commun. 2017, 8, 2032. [Google Scholar] [CrossRef]
- Baslan, T.; Hicks, J. Unravelling biology and shifting paradigms in cancer with single-cell sequencing. Nat. Rev. Cancer 2017, 17, 557–569. [Google Scholar] [CrossRef]
- Schmid, P.; Adams, S.; Rugo, H.S.; Schneeweiss, A.; Barrios, C.H.; Iwata, H.; Dieras, V.; Hegg, R.; Im, S.A.; Shaw Wright, G.; et al. Atezolizumab and Nab-Paclitaxel in Advanced Triple-Negative Breast Cancer. N. Engl. J. Med. 2018, 379, 2108–2121. [Google Scholar] [CrossRef]
- Zhang, S.; Chen, L.; Cui, B.; Chuang, H.Y.; Yu, J.; Wang-Rodriguez, J.; Tang, L.; Chen, G.; Basak, G.W.; Kipps, T.J. ROR1 is expressed in human breast cancer and associated with enhanced tumor-cell growth. PLoS ONE 2012, 7, e31127. [Google Scholar] [CrossRef]
- Curtis, C.; Shah, S.P.; Chin, S.F.; Turashvili, G.; Rueda, O.M.; Dunning, M.J.; Speed, D.; Lynch, A.G.; Samarajiwa, S.; Yuan, Y.; et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 2012, 486, 346–352. [Google Scholar] [CrossRef]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 2013, 6, pl1. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, H.; Oshi, M.; Asaoka, M.; Yan, L.; Endo, I.; Takabe, K. Molecular Biological Features of Nottingham Histological Grade 3 Breast Cancers. Ann. Surg. Oncol. 2020. [Google Scholar] [CrossRef]
- Tokumaru, Y.; Asaoka, M.; Oshi, M.; Katsuta, E.; Yan, L.; Narayanan, S.; Sugito, N.; Matsuhashi, N.; Futamura, M.; Akao, Y.; et al. High Expression of microRNA-143 is Associated with Favorable Tumor Immune Microenvironment and Better Survival in Estrogen Receptor Positive Breast Cancer. Int. J. Mol. Sci. 2020, 21, 3213. [Google Scholar] [CrossRef] [PubMed]
- Tokumaru, Y.; Katsuta, E.; Oshi, M.; Sporn, J.C.; Yan, L.; Le, L.; Matsuhashi, N.; Futamura, M.; Akao, Y.; Yoshida, K.; et al. High Expression of miR-34a Associated with Less Aggressive Cancer Biology but Not with Survival in Breast Cancer. Int. J. Mol. Sci. 2020, 21, 3045. [Google Scholar] [CrossRef] [PubMed]
- Okano, M.; Oshi, M.; Butash, A.L.; Katsuta, E.; Tachibana, K.; Saito, K.; Okayama, H.; Peng, X.; Yan, L.; Kono, K.; et al. Triple-Negative Breast Cancer with High Levels of Annexin A1 Expression Is Associated with Mast Cell Infiltration, Inflammation, and Angiogenesis. Int. J. Mol. Sci. 2019, 20, 4197. [Google Scholar] [CrossRef] [PubMed]
- Okano, M.; Oshi, M.; Butash, A.L.; Asaoka, M.; Katsuta, E.; Peng, X.; Qi, Q.; Yan, L.; Takabe, K. Estrogen Receptor Positive Breast Cancer with High Expression of Androgen Receptor has Less Cytolytic Activity and Worse Response to Neoadjuvant Chemotherapy but Better Survival. Int. J. Mol. Sci. 2019, 20, 2655. [Google Scholar] [CrossRef]
- Narayanan, S.; Kawaguchi, T.; Peng, X.; Qi, Q.; Liu, S.; Yan, L.; Takabe, K. Tumor Infiltrating Lymphocytes and Macrophages Improve Survival in Microsatellite Unstable Colorectal Cancer. Sci. Rep. 2019, 9, 13455. [Google Scholar] [CrossRef]
- Terakawa, T.; Katsuta, E.; Yan, L.; Turaga, N.; McDonald, K.A.; Fujisawa, M.; Guru, K.A.; Takabe, K. High expression of SLCO2B1 is associated with prostate cancer recurrence after radical prostatectomy. Oncotarget 2018, 9, 14207–14218. [Google Scholar] [CrossRef][Green Version]
- Hirose, Y.; Nagahashi, M.; Katsuta, E.; Yuza, K.; Miura, K.; Sakata, J.; Kobayashi, T.; Ichikawa, H.; Shimada, Y.; Kameyama, H.; et al. Generation of sphingosine-1-phosphate is enhanced in biliary tract cancer patients and is associated with lymphatic metastasis. Sci. Rep. 2018, 8, 10814. [Google Scholar] [CrossRef]
- Young, J.; Kawaguchi, T.; Yan, L.; Qi, Q.; Liu, S.; Takabe, K. Tamoxifen sensitivity-related microRNA-342 is a useful biomarker for breast cancer survival. Oncotarget 2017, 8, 99978–99989. [Google Scholar] [CrossRef]
- Hoki, T.; Katsuta, E.; Yan, L.; Takabe, K.; Ito, F. Low DMT1 Expression Associates With Increased Oxidative Phosphorylation and Early Recurrence in Hepatocellular Carcinoma. J. Surg. Res. 2019, 234, 343–352. [Google Scholar] [CrossRef] [PubMed]
- Sporn, J.C.; Katsuta, E.; Yan, L.; Takabe, K. Expression of MicroRNA-9 is Associated With Overall Survival in Breast Cancer Patients. J. Surg. Res. 2019, 233, 426–435. [Google Scholar] [CrossRef] [PubMed]
- Katsuta, E.; Qi, Q.; Peng, X.; Hochwald, S.N.; Yan, L.; Takabe, K. Pancreatic adenocarcinomas with mature blood vessels have better overall survival. Sci. Rep. 2019, 9, 1310. [Google Scholar] [CrossRef] [PubMed]
- Kawaguchi, T.; Yan, L.; Qi, Q.; Peng, X.; Gabriel, E.M.; Young, J.; Liu, S.; Takabe, K. Overexpression of suppressive microRNAs, miR-30a and miR-200c are associated with improved survival of breast cancer patients. Sci. Rep. 2017, 7, 15945. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.Y.; Kawaguchi, T.; Yan, L.; Young, J.; Qi, Q.; Takabe, K. Clinical Relevance of microRNA Expressions in Breast Cancer Validated Using the Cancer Genome Atlas (TCGA). Ann. Surg. Oncol. 2017, 24, 2943–2949. [Google Scholar] [CrossRef]
- Kawaguchi, T.; Yan, L.; Qi, Q.; Peng, X.; Edge, S.B.; Young, J.; Yao, S.; Liu, S.; Otsuji, E.; Takabe, K. Novel MicroRNA-Based Risk Score Identified by Integrated Analyses to Predict Metastasis and Poor Prognosis in Breast Cancer. Ann. Surg. Oncol. 2018, 25, 4037–4046. [Google Scholar] [CrossRef]
- Ramanathan, R.; Olex, A.L.; Dozmorov, M.; Bear, H.D.; Fernandez, L.J.; Takabe, K. Angiopoietin pathway gene expression associated with poor breast cancer survival. Breast Cancer Res. Treat. 2017, 162, 191–198. [Google Scholar] [CrossRef]
- Thorsson, V.; Gibbs, D.L.; Brown, S.D.; Wolf, D.; Bortone, D.S.; Ou Yang, T.H.; Porta-Pardo, E.; Gao, G.F.; Plaisier, C.L.; Eddy, J.A.; et al. The Immune Landscape of Cancer. Immunity 2018, 48, 812–830.e14. [Google Scholar] [CrossRef]
- Newman, A.M.; Liu, C.L.; Green, M.R.; Gentles, A.J.; Feng, W.; Xu, Y.; Hoang, C.D.; Diehn, M.; Alizadeh, A.A. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 2015, 12, 453–457. [Google Scholar] [CrossRef]
- McDonald, K.A.; Kawaguchi, T.; Qi, Q.; Peng, X.; Asaoka, M.; Young, J.; Opyrchal, M.; Yan, L.; Patnaik, S.; Otsuji, E.; et al. Tumor Heterogeneity Correlates with Less Immune Response and Worse Survival in Breast Cancer Patients. Ann. Surg. Oncol. 2019, 26, 2191–2199. [Google Scholar] [CrossRef]
- Narayanan, S.; Kawaguchi, T.; Yan, L.; Peng, X.; Qi, Q.; Takabe, K. Cytolytic Activity Score to Assess Anticancer Immunity in Colorectal Cancer. Ann. Surg. Oncol. 2018, 25, 2323–2331. [Google Scholar] [CrossRef] [PubMed]
- Takeshita, T.; Yan, L.; Asaoka, M.; Rashid, O.; Takabe, K. Late recurrence of breast cancer is associated with pro-cancerous immune microenvironment in the primary tumor. Sci. Rep. 2019, 9, 16942. [Google Scholar] [CrossRef] [PubMed]
- Rooney, M.S.; Shukla, S.A.; Wu, C.J.; Getz, G.; Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 2015, 160, 48–61. [Google Scholar] [CrossRef]
- Takahashi, H.; Kawaguchi, T.; Yan, L.; Peng, X.; Qi, Q.; Morris, L.G.T.; Chan, T.A.; Tsung, A.; Otsuji, E.; Takabe, K. Immune Cytolytic Activity for Comprehensive Understanding of Immune Landscape in Hepatocellular Carcinoma. Cancers (Basel) 2020, 12, 1221. [Google Scholar] [CrossRef]
- Asaoka, M.; Patnaik, S.K.; Zhang, F.; Ishikawa, T.; Takabe, K. Lymphovascular invasion in breast cancer is associated with gene expression signatures of cell proliferation but not lymphangiogenesis or immune response. Breast Cancer Res. Treat. 2020, 181, 309–322. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, H.; Asaoka, M.; Yan, L.; Rashid, O.M.; Oshi, M.; Ishikawa, T.; Nagahashi, M.; Takabe, K. Biologically Aggressive Phenotype and Anti-cancer Immunity Counterbalance in Breast Cancer with High Mutation Rate. Sci. Rep. 2020, 10, 1852. [Google Scholar] [CrossRef] [PubMed]
- Asaoka, M.; Ishikawa, T.; Takabe, K.; Patnaik, S.K. APOBEC3-Mediated RNA Editing in Breast Cancer is Associated with Heightened Immune Activity and Improved Survival. Int. J. Mol. Sci. 2019, 20, 5621. [Google Scholar] [CrossRef]
- Li, T.; Fan, J.; Wang, B.; Traugh, N.; Chen, Q.; Liu, J.S.; Li, B.; Liu, X.S. TIMER: A Web Server for Comprehensive Analysis of Tumor-Infiltrating Immune Cells. Cancer Res. 2017, 77, e108–e110. [Google Scholar] [CrossRef]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef]
- Oshi, M.; Katsuta, E.; Yan, L.; Ebos, J.M.L.; Rashid, O.M.; Matsuyama, R.; Endo, I.; Takabe, K. A Novel 4-Gene Score to Predict Survival, Distant Metastasis and Response to Neoadjuvant Therapy in Breast Cancer. Cancers (Basel) 2020, 12, 1148. [Google Scholar] [CrossRef]
- Oshi, M.; Takahashi, H.; Tokumaru, Y.; Yan, L.; Rashid, O.M.; Matsuyama, R.; Endo, I.; Takabe, K. G2M Cell Cycle Pathway Score as a Prognostic Biomarker of Metastasis in Estrogen Receptor (ER)-Positive Breast Cancer. Int. J. Mol. Sci. 2020, 21, 2921. [Google Scholar] [CrossRef] [PubMed]
- Tokumaru, Y.; Oshi, M.; Katsuta, E.; Yan, L.; Satyananda, V.; Matsuhashi, N.; Futamura, M.; Akao, Y.; Yoshida, K.; Takabe, K. KRAS signaling enriched triple negative breast cancer is associated with favorable tumor immune microenvironment and better survival. Am. J. Cancer Res. 2020, 10, 897–907. [Google Scholar] [PubMed]
- Katsuta, E.; Yan, L.; Takeshita, T.; McDonald, K.A.; Dasgupta, S.; Opyrchal, M.; Takabe, K. High MYC mRNA Expression Is More Clinically Relevant than MYC DNA Amplification in Triple-Negative Breast Cancer. Int. J. Mol. Sci. 2019, 21, 217. [Google Scholar] [CrossRef] [PubMed]
- Takeshita, T.; Asaoka, M.; Katsuta, E.; Photiadis, S.J.; Narayanan, S.; Yan, L.; Takabe, K. High expression of polo-like kinase 1 is associated with TP53 inactivation, DNA repair deficiency, and worse prognosis in ER positive Her2 negative breast cancer. Am. J. Transl. Res. 2019, 11, 6507–6521. [Google Scholar]


| Demographic and Clinical Characteristics | METABRIC | TCGA | ||||
|---|---|---|---|---|---|---|
| Clinical Variables (percent per GR status) | GR-LOW n = 348 | GR-HIGH n = 348 | p-value | GR-LOW n = 256 | GR-HIGH n = 256 | p-value |
| Age at diagnosis | 0.063 | 0.376 | ||||
| Median | 63 | 61 | 59 | 60 | ||
| IQR | 53–71 | 52–69 | 50–69 | 50–66 | ||
| Stage at diagnosis | 0.017 | 0.019 | ||||
| I | 104 (29.9%) | 140 (40.2%) | 36 (14.0%) | 56 (21.9%) | ||
| II | 215 (61.8%) | 182 (52.3%) | 165 (64.5%) | 135 (52.7%) | ||
| III | 29 (8.3%) | 26 (7.5%) | 55 (21.5%) | 65 (25.4%) | ||
| Clinical Subtypes | ||||||
| ER Positive | 257 (48.1%) | 277 (51.9%) | 0.088 | 183 (47.2%) | 205 (52.8%) | 0.020 |
| PR Positive | 188 (51.8%) | 175 (48.2%) | 0.363 | 152 (45%) | 186 (55%) | 0.001 |
| HER2 Positive | 59 (71.1%) | 24 (28.9%) | <0.001 | 54 (65.1%) | 29 (34.9%) | <0.001 |
| Triple Negative | 55 (49.1%) | 57 (50.9%) | 0.918 | 28 (65.1%) | 15 (34.9%) | 0.033 |
| PAM50 Subtypes | <0.001 | <0.001 | ||||
| Luminal A | 117 (33.6%) | 137 (39.4%) | 65 (25.4%) | 149 (58.2%) | ||
| Luminal B | 102 (29.3%) | 64 (18.4%) | 45 (17.6%) | 34 (13.3%) | ||
| HER2 | 59 (17%) | 24 (6.9%) | 21 (8.2%) | 7 (2.7%) | ||
| Basal | 57 (16.4%) | 47 (13.5%) | 30 (11.7%) | 27 (10.5%) | ||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gandhi, S.; Elkhanany, A.; Oshi, M.; Dai, T.; Opyrchal, M.; Mohammadpour, H.; Repasky, E.A.; Takabe, K. Contribution of Immune Cells to Glucocorticoid Receptor Expression in Breast Cancer. Int. J. Mol. Sci. 2020, 21, 4635. https://doi.org/10.3390/ijms21134635
Gandhi S, Elkhanany A, Oshi M, Dai T, Opyrchal M, Mohammadpour H, Repasky EA, Takabe K. Contribution of Immune Cells to Glucocorticoid Receptor Expression in Breast Cancer. International Journal of Molecular Sciences. 2020; 21(13):4635. https://doi.org/10.3390/ijms21134635
Chicago/Turabian StyleGandhi, Shipra, Ahmed Elkhanany, Masanori Oshi, Tao Dai, Mateusz Opyrchal, Hemn Mohammadpour, Elizabeth A. Repasky, and Kazuaki Takabe. 2020. "Contribution of Immune Cells to Glucocorticoid Receptor Expression in Breast Cancer" International Journal of Molecular Sciences 21, no. 13: 4635. https://doi.org/10.3390/ijms21134635
APA StyleGandhi, S., Elkhanany, A., Oshi, M., Dai, T., Opyrchal, M., Mohammadpour, H., Repasky, E. A., & Takabe, K. (2020). Contribution of Immune Cells to Glucocorticoid Receptor Expression in Breast Cancer. International Journal of Molecular Sciences, 21(13), 4635. https://doi.org/10.3390/ijms21134635






