Epigenetic Modulation of Chromatin States and Gene Expression by G-Quadruplex Structures
Abstract
1. Introduction
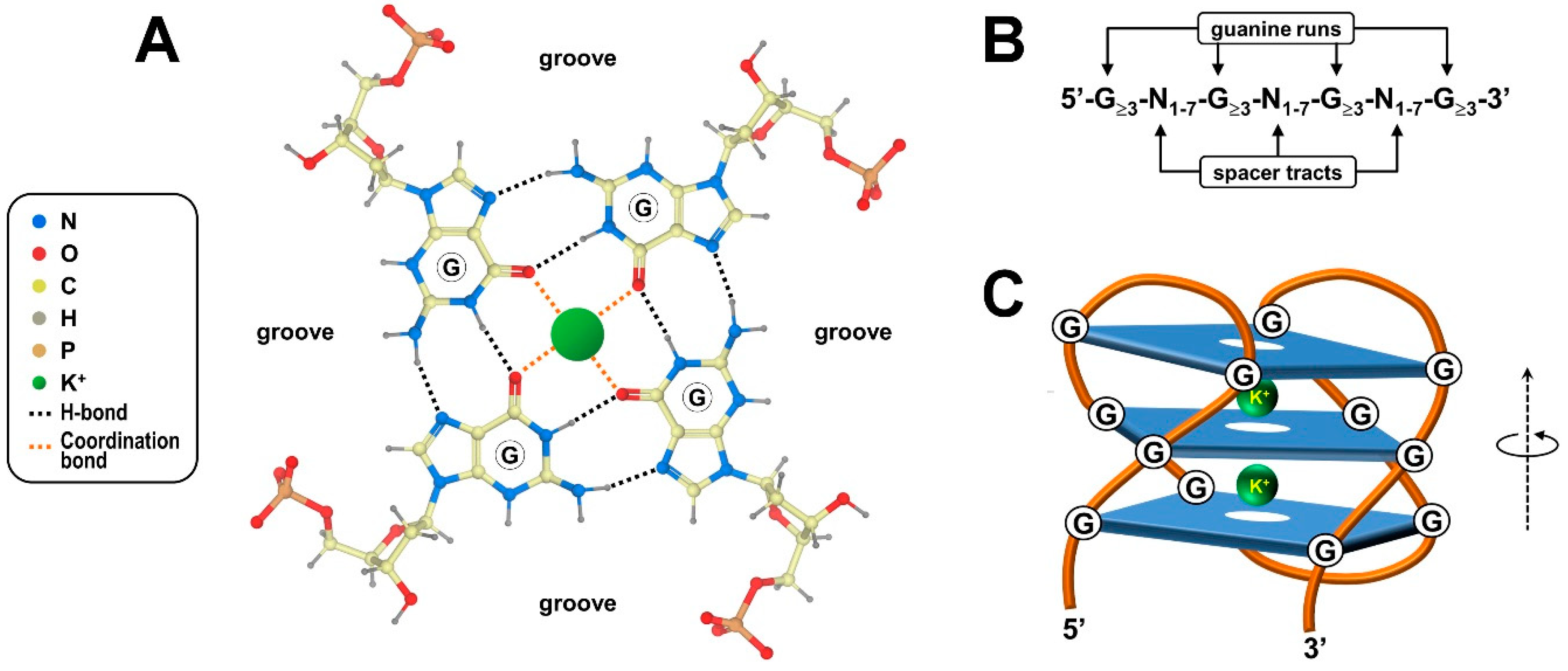
2. Structural Features of G-Quadruplexes and Their Occurrence in Biological Contexts
3. Involvement of G4 Structures in Epigenetic Processes
3.1. Interplay between G4 Structures and Epigenetic Modifications of DNA Bases
3.2. G4 Structures and the Histone Epigenetic Machinery
3.3. Epigenetic Roles of G4 Structures Formed in Coding and Noncoding RNA Transcripts
4. Conclusions and Perspectives
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| 5meC | C5-methyl-deoxycitosine |
| 6meA | N6-methyl-deoxyadenosine |
| 6meG | O6-methyl-deoxyguanine |
| 8oxoG | 8-oxo-7,8-dihydro-guanine |
| APE1 | apurinic-apyrimidinic endonuclease 1 |
| ASXL1 | Additional Sex combs-Like 1 |
| ATRX | ATP- dependent X-linked helicase |
| BRD3 | Bromodomain Containing 3 |
| CTCF | CCCTC-binding Factor |
| DAXX | Death Domain Associated Protein |
| DHX36 | DEAH-Box Helicase 36 |
| DNMT | DNA methyltransferase |
| FACT | Facilitates Chromatin Transcription |
| FANCJ | Fanconi Anemia group J |
| FUS | Fused in Sarcoma protein |
| G4 | G-quadruplex |
| HIF1-α | Hypoxia-inducible factor 1-α |
| HMGB1 | Chromosomal High Mobility Group Box 1 |
| hTERT | human Telomerase Reverse Transcriptase |
| KMT5C | Histone-lysine N-Methyltransferase KMT5C |
| LINE-1 | Long Interspersed Nuclear Element-1 |
| LSD1 | Lysine Specific Demethylase 1 |
| lysC | lysozyme C |
| NuRD | Nucleosome Remodeling and Deacetylase complex |
| OGG1 | 8-oxoguanine DNA glycosylase |
| PADi4 | Peptidylarginine Deiminase 4 |
| PARP1 | Poly ADP-Ribose Polymerase 1 |
| PHF8 | Histone Lysine Demethylase 8 |
| PRC2 | Polycomb Repressive Complex 2 |
| REST | RE1-Silencing Transcription Factor |
| SWI/SNF | SWItch/Sucrose Non-Fermentable |
| TERRA | Telomeric Repeat-containing RNA |
| TLS | Translocated in Liposarcoma protein |
| UTR | Untranslated Region |
| VEGF | Vascular Endothelial Growth Factor |
References
- Levene, P.A.; Jacobs, W.A. Über Guanylsäure. Ber. Dtsch. Chem. Ges. 1909, 42, 2469–2473. [Google Scholar] [CrossRef][Green Version]
- Bang, I. Untersuchungen über die Guanylsäure. Biochem. Z. 1910, 26, 293–311. [Google Scholar]
- Gellert, M.; Lipsett, M.N.; Davies, D.R. Helix formation by guanylic acid. Proc. Natl. Acad. Sci. USA 1962, 48, 2013–2018. [Google Scholar] [CrossRef]
- Arnott, S.; Chandrasekaran, R.; Marttila, C.M. Structures for polyinosinic acid and polyguanylic acid. Biochem. J. 1974, 141, 537–543. [Google Scholar] [CrossRef] [PubMed]
- Zimmerman, S.B.; Cohen, G.H.; Davies, D.R. X-ray fiber diffraction and model-building study of polyguanylic acid and polyinosinic acid. J. Mol. Biol. 1975, 92, 181–192. [Google Scholar] [CrossRef]
- Howard, F.B.; Frazier, J.; Miles, H.T. Stable and metastable forms of poly (G). Biopolymers 1977, 16, 791–809. [Google Scholar] [CrossRef]
- Besnard, E.; Babled, A.; Lapasset, L.; Milhavet, O.; Parrinello, H.; Dantec, C.; Marin, J.M.; Lemaitre, J.M. Unraveling cell type-specific and reprogrammable human replication origin signatures associated with G-quadruplex consensus motifs. Nat. Struct. Mol. Biol. 2012, 19, 837–844. [Google Scholar] [CrossRef]
- Fernando, H.; Sewitz, S.; Darot, J.; Tavare, S.; Huppert, J.L.; Balasubramanian, S. Genome-wide analysis of a G-quadruplex-specific single-chain antibody that regulates gene expression. Nucleic Acids Res. 2009, 37, 6716–6722. [Google Scholar] [CrossRef]
- Paeschke, K.; Simonsson, T.; Postberg, J.; Lipps, H.J. Telomere end-binding proteins control the formation of G-quadruplex DNA structures in vivo. Nat. Struct. Mol. Biol. 2005, 12, 847–854. [Google Scholar] [CrossRef]
- Cavalieri, V.; Di Bernardo, M.; Anello, L.; Spinelli, G. Cis-Regulatory sequences driving the expression of the Hbox12 homeobox-containing gene in the presumptive aboral ectoderm territory of the Paracentrotus lividus sea urchin embryo. Dev. Biol. 2008, 321, 455–469. [Google Scholar] [CrossRef] [PubMed]
- Turturici, G.; La Fiora, V.; Terenzi, A.; Barone, G.; Cavalieri, V. Perturbation of developmental regulatory gene expression by a G-quadruplex DNA inducer in the Sea Urchin embryo. Biochemistry 2018, 57, 4391–4394. [Google Scholar] [CrossRef] [PubMed]
- Ribeyre, C.; Lopes, J.; Boule, J.B.; Piazza, A.; Guedin, A.; Zakian, V.A.; Mergny, J.L.; Nicolas, A. The Yeast Pif1 helicase prevents genomic instability caused by G-Quadruplex-Forming CEB1 sequences in vivo. PLoS Genet. 2009, 5, e1000475. [Google Scholar] [CrossRef] [PubMed]
- Sarkies, P.; Reams, C.; Simpson, L.J.; Sale, J.E. Epigenetic instability due to defective replication of structured DNA. Mol. Cell 2010, 40, 703–713. [Google Scholar] [CrossRef]
- Siddiqui-Jain, A.; Grand, C.L.; Bearss, D.J.; Hurley, L.H. Direct evidence for a G-quadruplex in a promoter region and its targeting with a small molecule to repress c-MYC transcription. Proc. Natl. Acad. Sci. USA 2002, 99, 11593–11598. [Google Scholar] [CrossRef]
- Valton, A.L.; Hassan-Zadeh, V.; Lema, I.; Boggetto, N.; Alberti, P.; Saintome, C.; Riou, J.F.; Prioleau, M.N. G4 motifs affect origin positioning and efficiency in two vertebrate replicators. EMBO J. 2014, 33, 732–746. [Google Scholar] [CrossRef] [PubMed]
- Wieland, M.; Hartig, J.S. RNA quadruplex-based modulation of gene expression. Chem. Biol. 2007, 14, 757–763. [Google Scholar] [CrossRef]
- Sen, D.; Gilbert, W. Formtaion of parallel four-stranded complexes by guanine-rich motifs in DNA and its implications for meiosis. Nature 1988, 334, 364–366. [Google Scholar] [CrossRef]
- Burge, S.; Parkinson, G.N.; Hazel, P.; Todd, A.K.; Neidle, S. Quadruplex DNA: Sequence, topology and structure. Nucleic Acids Res. 2006, 34, 5402–5415. [Google Scholar] [CrossRef]
- Webba da Silva, M. Geometric formalism for DNA quadruplex folding. Chemistry 2007, 13, 9738–9745. [Google Scholar] [CrossRef]
- Huppert, J.L. Structure, location and interactions of G-quadruplexes. FEBS J. 2010, 277, 3452–3458. [Google Scholar] [CrossRef]
- Mergny, J.L.; Sen, D. DNA Quadruple helices in nanotechnology. Chem. Rev. 2019, 119, 6290–6325. [Google Scholar] [CrossRef] [PubMed]
- Neidle, S.; Balasubramanian, S. Quadruplex Nucleic Acids; Royal Society of Chemistry: London, UK, 2006. [Google Scholar]
- Huppert, J.L. Four-stranded nucleic acids: Structure, function and targeting of G-quadruplexes. Chem. Soc. Rev. 2008, 37, 1375–1384. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharyya, D.; Mirihana Arachchilage, G.; Basu, S. metal cations in G-quadruplex folding and stability. Front. Chem. 2016, 4, 38. [Google Scholar] [CrossRef] [PubMed]
- Largy, E.; Mergny, J.L.; Gabelica, V. Role of Alkali Metal Ions in G-Quadruplex Nucleic Acid Structure and Stability. Met. Ions Life Sci. 2016, 16, 203–258. [Google Scholar] [CrossRef]
- Tran, P.L.; De Cian, A.; Gros, J.; Moriyama, R.; Mergny, J.L. Tetramolecular quadruplex stability and assembly. Top. Curr. Chem. 2013, 330, 243–273. [Google Scholar] [CrossRef]
- Mukundan, V.T.; Phan, A.T. Bulges in G-quadruplexes: Broadening the definition of G-quadruplex-forming sequences. J. Am. Chem. Soc. 2013, 135, 5017–5028. [Google Scholar] [CrossRef]
- Kolesnikova, S.; Curtis, E.A. Structure and Function of Multimeric G-Quadruplexes. Molecules 2019, 24, 3074. [Google Scholar] [CrossRef]
- Zhou, J.; Amrane, S.; Korkut, D.N.; Bourdoncle, A.; He, H.Z.; Ma, D.L.; Mergny, J.L. Combination of i-motif and G-quadruplex structures within the same strand: Formation and application. Angew. Chem. 2013, 52, 7742–7746. [Google Scholar] [CrossRef]
- Zheng, K.W.; Xiao, S.; Liu, J.Q.; Zhang, J.Y.; Hao, Y.H.; Tan, Z. Co-transcriptional formation of DNA:RNA hybrid G-quadruplex and potential function as constitutional cis element for transcription control. Nucleic Acids Res. 2013, 41, 5533–5541. [Google Scholar] [CrossRef]
- Bose, K.; Lech, C.J.; Heddi, B.; Phan, A.T. High-resolution AFM structure of DNA G-wires in aqueous solution. Nat. Commun. 2018, 9, 1959. [Google Scholar] [CrossRef]
- Sondermann, A.; Holste, C.; Moller, R.; Fritzsche, W. Assembly of G-quartet based DNA superstructures (G-wires). AIP Conf. Proc. 2002, 640, 93–98. [Google Scholar]
- Guedin, A.; Gros, J.; Alberti, P.; Mergny, J.L. How long is too long? Effects of loop size on G-quadruplex stability. Nucleic Acids Res. 2010, 38, 7858–7868. [Google Scholar] [CrossRef]
- Lightfoot, H.L.; Hagen, T.; Tatum, N.J.; Hall, J. The diverse structural landscape of quadruplexes. FEBS Lett. 2019, 593, 2083–2102. [Google Scholar] [CrossRef] [PubMed]
- Rachwal, P.A.; Findlow, I.S.; Werner, J.M.; Brown, T.; Fox, K.R. Intramolecular DNA quadruplexes with different arrangements of short and long loops. Nucleic Acids Res. 2007, 35, 4214–4222. [Google Scholar] [CrossRef] [PubMed]
- Simonsson, T. G-quadruplex DNA structures—Variations on a theme. Biol. Chem. 2001, 382, 621–628. [Google Scholar] [CrossRef] [PubMed]
- Wlodarczyk, A.; Grzybowski, P.; Patkowski, A.; Dobek, A. Effect of ions on the polymorphism, effective charge, and stability of human telomeric DNA. Photon correlation spectroscopy and circular dichroism studies. J. Phys. Chem. B 2005, 109, 3594–3605. [Google Scholar] [CrossRef] [PubMed]
- Saccà, B.; Lacroix, L.; Mergny, J.L. The effect of chemical modifications on the thermal stability of different G-quadruplex-forming oligonucleotides. Nucleic Acids Res. 2005, 33, 1182–1192. [Google Scholar] [CrossRef] [PubMed]
- Tran, P.L.; Virgilio, A.; Esposito, V.; Citarella, G.; Mergny, J.L.; Galeone, A. Effects of 8-methylguanine on structure, stability and kinetics of formation of tetramolecular quadruplexes. Biochimie 2011, 93, 399–408. [Google Scholar] [CrossRef]
- Virgilio, A.; Esposito, V.; Citarella, G.; Pepe, A.; Mayol, L.; Galeone, A. The insertion of two 8-methyl-2′-deoxyguanosine residues in tetramolecular quadruplex structures: Trying to orientate the strands. Nucleic Acids Res. 2012, 40, 461–475. [Google Scholar] [CrossRef]
- Yan, Y.Y.; Tan, J.H.; Lu, Y.J.; Yan, S.C.; Wong, K.Y.; Li, D.; Gu, L.Q.; Huang, Z.S. G-Quadruplex conformational change driven by pH variation with potential application as a nanoswitch. Biochim. et Biophys. Acta 2013, 1830, 4935–4942. [Google Scholar] [CrossRef]
- Miyoshi, D.; Nakao, A.; Sugimoto, N. Molecular crowding regulates the structural switch of the DNA G-quadruplex. Biochemistry 2002, 41, 15017–15024. [Google Scholar] [CrossRef] [PubMed]
- Borgognone, M.; Armas, P.; Calcaterra, N.B. Cellular nucleic-acid-binding protein, a transcriptional enhancer of c-Myc, promotes the formation of parallel G-quadruplexes. Biochem. J. 2010, 428, 491–498. [Google Scholar] [CrossRef] [PubMed]
- Rajendran, A.; Endo, M.; Hidaka, K.; Tran, P.L.; Mergny, J.L.; Gorelick, R.J.; Sugiyama, H. HIV-1 nucleocapsid proteins as molecular chaperones for tetramolecular antiparallel G-quadruplex formation. J. Am. Chem. Soc. 2013, 135, 18575–18585. [Google Scholar] [CrossRef] [PubMed]
- Amrane, S.; Kerkour, A.; Bedrat, A.; Vialet, B.; Andreola, M.L.; Mergny, J.L. Topology of a DNA G-quadruplex structure formed in the HIV-1 promoter: A potential target for anti-HIV drug development. J. Am. Chem. Soc. 2014, 136, 5249–5252. [Google Scholar] [CrossRef]
- Bhartiya, D.; Chawla, V.; Ghosh, S.; Shankar, R.; Kumar, N. Genome-wide regulatory dynamics of G-quadruplexes in human malaria parasite Plasmodium falciparum. Genomics 2016, 108, 224–231. [Google Scholar] [CrossRef]
- Biswas, B.; Kandpal, M.; Jauhari, U.K.; Vivekanandan, P. Genome-wide analysis of G-quadruplexes in herpesvirus genomes. BMC Genom. 2016, 17, 949. [Google Scholar] [CrossRef]
- Chambers, V.S.; Marsico, G.; Boutell, J.M.; Di Antonio, M.; Smith, G.P.; Balasubramanian, S. High-throughput sequencing of DNA G-quadruplex structures in the human genome. Nat. Biotechnol. 2015, 33, 877–881. [Google Scholar] [CrossRef]
- Fleming, A.M.; Ding, Y.; Alenko, A.; Burrows, C.J. Zika virus genomic RNA possesses conserved G-quadruplexes characteristic of the flaviviridae family. ACS Infect. Dis. 2016, 2, 674–681. [Google Scholar] [CrossRef]
- Garg, R.; Aggarwal, J.; Thakkar, B. Genome-wide discovery of G-quadruplex forming sequences and their functional relevance in plants. Sci. Rep. 2016, 6, 28211. [Google Scholar] [CrossRef]
- Hershman, S.G.; Chen, Q.; Lee, J.Y.; Kozak, M.L.; Yue, P.; Wang, L.S.; Johnson, F.B. Genomic distribution and functional analyses of potential G-quadruplex-forming sequences in Saccharomyces cerevisiae. Nucleic Acids Res. 2008, 36, 144–156. [Google Scholar] [CrossRef]
- Huppert, J.L.; Balasubramanian, S. Prevalence of quadruplexes in the human genome. Nucleic Acids Res. 2005, 33, 2908–2916. [Google Scholar] [CrossRef]
- Perrone, R.; Nadai, M.; Frasson, I.; Poe, J.A.; Butovskaja, E.; Smithgall, T.E.; Palumbo, M.; Palù, G.; Richter, S.N. A Dynamic G-quadruplex region regulates the HIV-1 long terminal repeat promoter. J. Med. Chem. 2013, 56, 6521–6530. [Google Scholar] [CrossRef] [PubMed]
- Rawal, P.; Kummarasetti, V.B.; Ravindran, J.; Kumar, N.; Halder, K.; Sharma, R.; Mukerji, M.; Das, S.K.; Chowdhury, S. Genome-wide prediction of G4 DNA as regulatory motifs: Role in Escherichia coli global regulation. Genome Res. 2006, 16, 644–655. [Google Scholar] [CrossRef] [PubMed]
- Saad, M.; Guédin, A.; Amor, S.; Bedrat, A.; Tourasse, N.J.; Fayyad-Kazan, H.; Pratviel, G.; Lacroix, L.; Mergny, J.L. Mapping and characterization of G-quadruplexes in the genome of the social amoeba Dictyostelium discoideum. Nucleic Acids Res. 2019, 47, 4363–4374. [Google Scholar] [CrossRef] [PubMed]
- Capra, J.A.; Paeschke, K.; Singh, M.; Zakian, V.A. G-quadruplex DNA sequences are evolutionarily conserved and associated with distinct genomic features in Saccharomyces cerevisiae. PLoS Comput. Biol. 2010, 6, e1000861. [Google Scholar] [CrossRef]
- Marsico, G.; Chambers, V.S.; Sahakyan, A.B.; McCauley, P.; Boutell, J.M.; Antonio, M.D.; Balasubramanian, S. Whole genome experimental maps of DNA G-quadruplexes in multiple species. Nucleic Acids Res. 2019, 47, 3862–3874. [Google Scholar] [CrossRef] [PubMed]
- Puig Lombardi, E.; Holmes, A.; Verga, D.; Teulade-Fichou, M.P.; Nicolas, A.; Londoño-Vallejo, A. Thermodynamically stable and genetically unstable G-quadruplexes are depleted in genomes across species. Nucleic Acids Res. 2019, 47, 6098–6113. [Google Scholar] [CrossRef] [PubMed]
- Verma, A.; Halder, K.; Halder, R.; Yadav, V.K.; Rawal, P.; Thakur, R.K.; Mohd, F.; Sharma, A.; Chowdhury, S. Genome-wide computational and expression analyses reveal G-quadruplex DNA motifs as conserved cis-regulatory elements in human and related species. J. Med. Chem. 2008, 51, 5641–5649. [Google Scholar] [CrossRef]
- Ravichandran, S.; Kim, Y.E.; Bansal, V.; Ghosh, A.; Hur, J.; Subramani, V.K.; Pradhan, S.; Lee, M.K.; Kim, K.K.; Ahn, J.H. Genome-wide analysis of regulatory G-quadruplexes affecting gene expression in human cytomegalovirus. PLoS Pathog. 2018, 14, e1007334. [Google Scholar] [CrossRef]
- Fleming, A.M.; Zhu, J.; Ding, Y.; Visser, J.A.; Zhu, J.; Burrows, C.J. Human DNA repair genes possess potential G-quadruplex sequences in their promoters and 5′-untranslated regions. Biochemistry 2018, 57, 991–1002. [Google Scholar] [CrossRef]
- Hansel-Hertsch, R.; Beraldi, D.; Lensing, S.V.; Marsico, G.; Zyner, K.; Parry, A.; Di Antonio, M.; Pike, J.; Kimura, H.; Narita, M.; et al. G-quadruplex structures mark human regulatory chromatin. Nat. Genet. 2016, 48, 1267–1272. [Google Scholar] [CrossRef] [PubMed]
- Maizels, N.; Gray, L.T. The G4 genome. PLoS Genet. 2013, 9, e1003468. [Google Scholar] [CrossRef] [PubMed]
- Piazza, A.; Adrian, M.; Samazan, F.; Heddi, B.; Hamon, F.; Serero, A.; Lopes, J.; Teulade-Fichou, M.P.; Phan, A.T.; Nicolas, A. Short loop length and high thermal stability determine genomic instability induced by G-quadruplex-forming minisatellites. EMBO J. 2015, 34, 1718–1734. [Google Scholar] [CrossRef]
- Yadav, V.K.; Abraham, J.K.; Mani, P.; Kulshrestha, R.; Chowdhury, S. QuadBase: Genome-wide database of G4 DNAeoccurrence and conservation in human, chimpanzee, mouse and rat promoters and 146 microbes. Nucleic Acids Res. 2008, 36, D381–D385. [Google Scholar] [CrossRef]
- Artusi, S.; Perrone, R.; Lago, S.; Raffa, P.; Di Iorio, E.; Palù, G.; Richter, S.N. Visualization of DNA G-quadruplexes in herpes simplex virus 1-infected cells. Nucleic Acids Res. 2016, 44, 10343–10353. [Google Scholar] [CrossRef]
- Biffi, G.; Tannahill, D.; McCafferty, J.; Balasubramanian, S. Quantitative visualization of DNA G-quadruplex structures in human cells. Nat. Chem. 2013, 5, 182–186. [Google Scholar] [CrossRef]
- Henderson, A.; Wu, Y.; Huang, Y.C.; Chavez, E.A.; Platt, J.; Johnson, F.B.; Brosh, R.M., Jr.; Sen, D.; Lansdorp, P.M. Detection of G-quadruplex DNA in mammalian cells. Nucleic Acids Res. 2014, 42, 860–869. [Google Scholar] [CrossRef]
- Schaffitzel, C.; Berger, I.; Postberg, J.; Hanes, J.; Lipps, H.J.; Plückthun, A. In vitro generated antibodies specific for telomeric guanine-quadruplex DNA react with Stylonychia lemnae macronuclei. Proc. Natl. Acad. Sci. USA 2001, 98, 8572–8577. [Google Scholar] [CrossRef]
- Bao, H.L.; Ishizuka, T.; Sakamoto, T.; Fujimoto, K.; Uechi, T.; Kenmochi, N.; Xu, Y. Characterization of human telomere RNA G-quadruplex structures in vitro and in living cells using 19F NMR spectroscopy. Nucleic Acids Res. 2017, 45, 5501–5511. [Google Scholar] [CrossRef]
- Hänsel, R.; Foldynová-Trantírková, S.; Dötsch, V.; Trantírek, L. Investigation of quadruplex structure under physiological conditions using in-cell NMR. Top. Curr. Chem. 2013, 330, 47–65. [Google Scholar] [CrossRef]
- Chen, M.C.; Murat, P.; Abecassis, K.; Ferré-D’Amaré, A.R.; Balasubramanian, S. Insights into the mechanism of a Gquadruplex-unwinding DEAH-box helicase. Nucleic Acids Res. 2015, 43, 2223–2231. [Google Scholar] [CrossRef] [PubMed]
- Gray, L.T.; Vallur, A.C.; Eddy, J.; Maizels, N. G-quadruplexes are genomewide targets of transcriptional helicases XPB and XPD. Nat. Chem. Biol. 2014, 10, 313–318. [Google Scholar] [CrossRef] [PubMed]
- Saha, T.; Shukla, K.; Thakur, R.S.; Desingu, A.; Nagaraju, G. Mycobacterium tuberculosis UvrD1 and UvrD2 helicases unwind G-quadruplex DNA. FEBS J. 2019, 286, 2062–2086. [Google Scholar] [CrossRef]
- Tippana, R.; Chen, M.C.; Demeshkina, N.A.; Ferré-D’Amaré, A.R.; Myong, S. RNA G-quadruplex is resolved by repetitive and ATP-dependent mechanism of DHX36. Nat. Commun. 2019, 10, 1855. [Google Scholar] [CrossRef] [PubMed]
- Voter, A.F.; Qiu, Y.; Tippana, R.; Myong, S.; Keck, J.L. A guanine-flipping and sequestration mechanism for G-quadruplex unwinding by RecQ helicases. Nat. Commun. 2018, 9, 4201. [Google Scholar] [CrossRef]
- Wu, G.; Xing, Z.; Tran, E.J.; Yang, D. DDX5 helicase resolves G-quadruplex and is involved in MYC gene transcriptional activation. Proc. Natl. Acad. Sci. USA 2019, 116, 20453–20461. [Google Scholar] [CrossRef] [PubMed]
- Zhou, R.; Zhang, J.; Bochman, M.L.; Zakian, V.A.; Ha, T. Periodic DNA patrolling underlies diverse functions of Pif1 on R-loops and G-rich DNA. eLife 2014, 3, e02190. [Google Scholar] [CrossRef]
- Cavalieri, V.; Spinelli, G. Environmental epigenetics in zebrafish. Epigenet. Chromatin 2017, 10, 46. [Google Scholar] [CrossRef]
- Cavalieri, V.; Spinelli, G. Histone-mediated transgenerational epigenetics. In Translational Epigenetics, 2nd ed.; Academic Press: Cambridge, MA, USA, 2007; pp. 157–183. [Google Scholar] [CrossRef]
- Halder, R.; Riou, J.F.; Teulade-Fichou, M.P.; Frickey, T.; Hartig, J.S. Bisquinolinium compounds induce quadruplex-specific transcriptome changes in HeLa S3 cell lines. BMC Res. Notes 2012, 5, 138. [Google Scholar] [CrossRef]
- Mikami-Terao, Y.; Akiyama, M.; Yuza, Y.; Yanagisawa, T.; Yamada, O.; Yamada, H. Antitumor activity of G-quadruplex-interactive agent TMPyP4 in K562 leukemic cells. Cancer Lett. 2008, 261, 226–234. [Google Scholar] [CrossRef]
- Bhattacharjee, S.; Chakraborty, S.; Sengupta, P.K.; Bhowmik, S. Exploring the interactions of the dietary plant flavonoids fisetin and naringenin with G-quadruplex and duplex DNA, showing contrasting binding behavior: Spectroscopic and molecular modeling approaches. J. Phys. Chem. B 2016, 120, 8942–8952. [Google Scholar] [CrossRef] [PubMed]
- François, M.; Leifert, W.; Tellam, R.; Fenech, M. G-quadruplexes: A possible epigenetic target for nutrition. Mutat. Res. Rev. Mutat. Res. 2015, 764, 101–107. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Zhang, M.; Zhang, J.L.; Li, H.Q.; Zhang, X.C.; Sun, Q.; Qiu, C.M. Interactions of daidzin with intramolecular G-quadruplex. FEBS Lett. 2006, 580, 4905–4910. [Google Scholar] [CrossRef] [PubMed]
- Mikutis, G.; Karaköse, H.; Jaiswal, R.; LeGresley, A.; Islam, T.; Fernandez-Lahore, M.; Kuhnert, N. Phenolic promiscuity in the cell nucleus—Epigallocatechingallate (EGCG) and theaflavin-3,30-digallate from green and black tea bind to model cell nuclear structures including histone proteins, double stranded DNA and telomeric quadruplex DNA. Food Funct. 2013, 4, 328–337. [Google Scholar] [CrossRef] [PubMed]
- Pattanayak, R.; Basak, P.; Sen, S.; Bhattacharyya, M. Interaction of KRAS G-quadruplex with natural polyphenols: A spectroscopic analysis with molecular modeling. Int. J. Biol. Macromol. 2016, 89, 228–237. [Google Scholar] [CrossRef]
- Platella, C.; Guida, S.; Bonmassar, L.; Aquino, A.; Bonmassar, E.; Ravagnan, G.; Montesarchio, D.; Roviello, G.N.; Musumeci, D.; Fuggetta, M.P. Antitumour activity of resveratrol on human melanoma cells: A possible mechanism related to its interaction with malignant cell telomerase. Biochim. et Biophys. Acta Gen. Subj. 2017, 1861, 2843–2851. [Google Scholar] [CrossRef]
- Benabou, S.; Mazzini, S.; Avino, A.; Eritja, R.; Gargallo, R.A. pH-dependent bolt involving cytosine bases located in the lateral loops of antiparallel G-quadruplex structures within the SMARCA4 gene promotor. Sci. Rep. 2019, 9, 15807. [Google Scholar] [CrossRef]
- Hardin, C.C.; Corregan, M.; Brown, B.A., II.; Frederick, L.N. Cytosine-cytosine+ base pairing stabilizes DNA quadruplexes and cytosine methylation greatly enhances the effect. Biochemistry 1993, 32, 5870–5880. [Google Scholar] [CrossRef]
- Kuo, M.H.; Wang, Z.F.; Tseng, T.Y.; Li, M.H.; Hsu, S.T.; Lin, J.J.; Chang, T.C. Conformational transition of a hairpin structure to G-quadruplex within the WNT1 gene promoter. J. Am. Chem. Soc. 2015, 137, 210–218. [Google Scholar] [CrossRef]
- Stevens, A.J.; Stuffrein-Roberts, S.; Cree, S.L.; Gibb, A.; Miller, A.L.; Doudney, K.; Aitchison, A.; Eccles, M.R.; Joyce, P.R.; Filichev, V.V.; et al. G-quadruplex structures and CpG methylation cause drop-out of the maternal allele in polymerase chain reaction amplification of the imprinted MEST gene promoter. PLoS ONE 2014, 9, e113955. [Google Scholar] [CrossRef]
- Halder, R.; Halder, K.; Sharma, P.; Garg, G.; Sengupta, S.; Chowdhury, S. Guanine quadruplex DNA structure restricts methylation of CpG dinucleotides genome-wide. Mol. Biosyst. 2010, 6, 2439–2447. [Google Scholar] [CrossRef] [PubMed]
- Jara-Espejo, M.; Line, S.R. DNA G-quadruplex stability, position and chromatin accessibility are associated with CpG island methylation. FEBS J. 2020, 287, 483–495. [Google Scholar] [CrossRef] [PubMed]
- Mao, S.Q.; Ghanbarian, A.T.; Spiegel, J.; Martínez Cuesta, S.; Beraldi, D.; Di Antonio, M.; Marsico, G.; Hänsel-Hertsch, R.; Tannahill, D.; Balasubramanian, S. DNA G-quadruplex structures mold the DNA methylome. Nat. Struct. Mol. Biol. 2018, 25, 951–957. [Google Scholar] [CrossRef] [PubMed]
- Cree, S.L.; Fredericks, R.; Miller, A.; Pearce, F.G.; Filichev, V.; Fee, C.; Kennedy, M.A. DNA G-quadruplexes show strong interaction with DNA methyltransferases in vitro. FEBS Lett. 2016, 590, 2870–2883. [Google Scholar] [CrossRef]
- Balasubramanian, S.; Hurley, L.H.; Neidle, S. Targeting G-quadruplexes in gene promoters: A novel anticancer strategy? Nat. Rev. Drug Discov. 2011, 10, 261–275. [Google Scholar] [CrossRef]
- Renaud, S.; Loukinov, D.; Abdullaev, Z.; Guilleret, I.; Bosman, F.T.; Lobanenkov, V.; Benhattar, J. Dual role of DNA methylation inside and outside of CTCF-binding regions in the transcriptional regulation of the telomerase hTERT gene. Nucleic Acids Res. 2007, 35, 1245–1256. [Google Scholar] [CrossRef] [PubMed]
- Li, P.T.; Wang, Z.F.; Chu, I.T.; Kuan, Y.M.; Li, M.H.; Huang, M.C.; Chiang, P.C.; Chang, T.C.; Chen, C.T. Expression of the human telomerase reverse transcriptase gene is modulated by quadruplex formation in its first exon due to DNA methylation. J. Biol. Chem. 2017, 292, 20859–20870. [Google Scholar] [CrossRef] [PubMed]
- Renaud, S.; Loukinov, D.; Bosman, F.T.; Lobanenkov, V.; Benhattar, J. CTCF binds the proximal exonic region of hTERT and inhibits its transcription. Nucleic Acids Res. 2005, 33, 6850–6860. [Google Scholar] [CrossRef]
- Tahiliani, M.; Koh, K.P.; Shen, Y.; Pastor, W.A.; Bandukwala, H.; Brudno, Y.; Agarwal, S.; Iyer, L.M.; Liu, D.R.; Aravind, L.; et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 2009, 324, 930–935. [Google Scholar] [CrossRef]
- Wu, X.; Zhang, Y. TET-mediated active DNA demethylation: Mechanism, function and beyond. Nat. Rev. Genet. 2017, 18, 517–534. [Google Scholar] [CrossRef]
- Pastor, W.A.; Pape, U.J.; Huang, Y.; Henderson, H.R.; Lister, R.; Ko, M.; McLoughlin, E.M.; Brudno, Y.; Mahapatra, S.; Kapranov, P.; et al. Genome-wide mapping of 5-hydroxymethylcytosine in embryonic stem cells. Nature 2011, 473, 394–397. [Google Scholar] [CrossRef] [PubMed]
- Song, C.X.; Szulwach, K.E.; Fu, Y.; Dai, Q.; Yi, C.; Li, X.; Li, Y.; Chen, C.H.; Zhang, W.; Jian, X.; et al. Selective chemical labeling reveals the genome-wide distribution of 5-hydroxymethylcytosine. Nat. Biotechnol. 2011, 29, 68–72. [Google Scholar] [CrossRef] [PubMed]
- Valinluck, V.; Sowers, L.C. Endogenous cytosine damage products alter the site selectivity of human DNA maintenance methyltransferase DNMT1. Cancer Res. 2007, 67, 946–950. [Google Scholar] [CrossRef] [PubMed]
- Bhavsar-Jog, Y.P.; Van Dornshuld, E.; Brooks, T.A.; Tschumper, G.S.; Wadkins, R.M. Epigenetic modification, dehydration, and molecular crowding effects on the thermodynamics of i-motif structure formation from C-rich DNA. Biochemistry 2014, 53, 1586–1594. [Google Scholar] [CrossRef]
- Morgan, R.K.; Molnar, M.M.; Batra, H.; Summerford, B.; Wadkins, R.M.; Brooks, T.A. Effects of 5-Hydroxymethylcytosine epigenetic modification on the stability and molecular recognition of VEGF i-motif and G-quadruplex structures. J. Nucleic Acids 2018, 2018, 9281286. [Google Scholar] [CrossRef]
- Spruijt, C.G.; Gnerlich, F.; Smits, A.H.; Pfaffeneder, T.; Jansen, P.W.; Bauer, C.; Münzel, M.; Wagner, M.; Müller, M.; Khan, F.; et al. Dynamic readers for 5-(hydroxy) methylcytosine and its oxidized derivatives. Cell 2013, 152, 1146–1159. [Google Scholar] [CrossRef] [PubMed]
- Sun, D.; Guo, K.; Shin, Y.J. Evidence of the formation of G-quadruplex structures in the promoter region of the human Vascular Endothelial Growth Factor Gene. Nucleic Acids Res. 2011, 39, 1256–1265. [Google Scholar] [CrossRef]
- Tsukakoshi, K.; Saito, S.; Yoshida, W.; Goto, S.; Ikebukuro, K. CpG methylation changes G-quadruplex structures derived from gene promoters and interaction with VEGF and SP1. Molecules 2018, 23, 944. [Google Scholar] [CrossRef]
- Yu, Y.; Cui, Y.; Niedernhofer, L.J.; Wang, Y. Occurrence, biological consequences, and human health relevance of oxidative stress-induced DNA damage. Chem. Res. Toxicol. 2016, 29, 2008–2039. [Google Scholar] [CrossRef]
- Saito, I.; Takayama, M.; Sugiyama, H.; Nakatani, K.; Tsuchida, A.; Yamamoto, M. Photoinduced DNA cleavage via electron transfer: Demonstration that guanine residues located 5′ to guanine are the most electron-donating sites. J. Am. Chem. Soc. 1995, 117, 6406–6407. [Google Scholar] [CrossRef]
- Steenken, S.; Jovanovic, S.V. How easily oxidizable is DNA? One-electron reduction potentials of adenosine and guanosine radicals in aqueous solution. J. Am. Chem. Soc. 1997, 119, 617–618. [Google Scholar] [CrossRef]
- Pfaffeneder, T.; Spada, F.; Wagner, M.; Brandmayr, C.; Laube, S.K.; Eisen, D.; Truss, M.; Steinbacher, J.; Hackner, B.; Kotljarova, O.; et al. Tet oxidizes thymine to 5-hydroxymethyluracil in mouse embryonic stem cell DNA. Nat. Chem. Biol. 2014, 10, 574–581. [Google Scholar] [CrossRef] [PubMed]
- Bielskutė, S.; Plavec, J.; Podbevšek, P. Impact of Oxidative Lesions on the Human Telomeric G-Quadruplex. J. Am. Chem. Soc. 2019, 141, 2594–2603. [Google Scholar] [CrossRef]
- Lech, C.J.; Cheow Lim, J.K.; Wen Lim, J.M.; Amrane, S.; Heddi, B.; Phan, A.T. Effects of site-specific guanine C8-modifications on an intramolecular DNA G-quadruplex. Biophys. J. 2011, 101, 1987–1998. [Google Scholar] [CrossRef]
- An, N.; Fleming, A.M.; Burrows, C.J. Human Telomere G-Quadruplexes with five repeats accommodate 8-Oxo-7,8-dihydroguanine by looping out the DNA damage. ACS Chem. Biol. 2016, 11, 500–507. [Google Scholar] [CrossRef]
- Fleming, A.M.; Zhou, J.; Wallace, S.S.; Burrows, C.J. A role for the fifth G-track in G-quadruplex forming oncogene promoter sequences during oxidative stress: Do these “spare tires” have an evolved function? ACS Cent. Sci. 2015, 1, 226–233. [Google Scholar] [CrossRef] [PubMed]
- Hao, W.; Qi, T.; Pan, L.; Wang, R.; Zhu, B.; Aguilera-Aguirre, L.; Radak, Z.; Hazra, T.K.; Vlahopoulos, S.A.; Bacsi, A.; et al. Effects of the stimuli-dependent enrichment of 8-oxoguanine DNA glycosylase1 on chromatinized DNA. Redox Biol. 2018, 18, 43–53. [Google Scholar] [CrossRef] [PubMed]
- Pan, L.; Zhu, B.; Hao, W.; Zeng, X.; Vlahopoulos, S.A.; Hazra, T.K.; Hegde, M.L.; Radak, Z.; Bacsi, A.; Brasier, A.R.; et al. Oxidized guanine base lesions function in 8-Oxoguanine DNA Glycosylase-1-mediated epigenetic regulation of nuclear factor κb-driven gene expression. J. Biol. Chem. 2016, 291, 25553–25566. [Google Scholar] [CrossRef]
- Antoniali, G.; Lirussi, L.; D’Ambrosio, C.; Dal Piaz, F.; Vascotto, C.; Casarano, E.; Marasco, D.; Scaloni, A.; Fogolari, F.; Tell, G. SIRT1 gene expression upon genotoxic damage is regulated by APE1 through nCaRE-promoter elements. Mol. Biol. Cell 2014, 25, 532–547. [Google Scholar] [CrossRef]
- Cogoi, S.; Xodo, L.E. G-quadruplex formation within the promoter of the KRAS proto-oncogene and its effect on transcription. Nucleic Acids Res. 2006, 34, 2536–2549. [Google Scholar] [CrossRef]
- Cogoi, S.; Ferino, A.; Miglietta, G.; Pedersen, E.B.; Xodo, L.E. The regulatory G4 motif of the Kirsten ras (KRAS) gene is sensitive to guanine oxidation: Implications on transcription. Nucleic Acids Res. 2018, 46, 661–676. [Google Scholar] [CrossRef] [PubMed]
- Fleming, A.M.; Ding, Y.; Burrows, C.J. Oxidative DNA damage is epigenetic by regulating gene transcription via base excision repair. Proc. Natl. Acad. Sci. USA 2017, 114, 2604–2609. [Google Scholar] [CrossRef] [PubMed]
- Fleming, A.M.; Zhu, J.; Howpay Manage, S.A.; Burrows, C.J. Human NEIL3 Gene expression regulated by epigenetic-like oxidative DNA modification. J. Am. Chem. Soc. 2019, 141, 11036–11049. [Google Scholar] [CrossRef] [PubMed]
- Fleming, A.M.; Burrows, C.J. Interplay of guanine oxidation and G-quadruplex folding in gene promoters. J. Am. Chem. Soc. 2020, 142, 1115–1136. [Google Scholar] [CrossRef]
- Pegg, A.E.; Dolan, M.E.; Moschel, R.C. Structure, function, and inhibition of O6-alkylguanine-DNA alkyltransferase. Prog. Nucleic Acid Res. Mol. Biol. 1995, 51, 167–223. [Google Scholar] [CrossRef]
- Mekmaysy, C.S.; Petraccone, L.; Garbett, N.C.; Ragazzon, P.A.; Gray, R.; Trent, J.O.; Chaires, J.B. Effect of O6-methylguanine on the stability of G-quadruplex DNA. J. Am. Chem. Soc. 2008, 130, 6710–6711. [Google Scholar] [CrossRef]
- Engel, J.D.; von Hippel, P.H. Effects of methylation on the stability of nucleic acid conformations: Studies at the monomer level. Biochemistry 1974, 13, 4143–4158. [Google Scholar] [CrossRef]
- Engel, J.D.; von Hippel, P.H. Effects of methylation on the stability of nucleic acid conformations. Studies at the polymer level. J. Biol. Chem. 1978, 253, 927–934. [Google Scholar]
- Laddachote, S.; Nagata, M.; Yoshida, W. Destabilisation of the c-kit1 G-quadruplex structure by N6-methyladenosine modification. Biochem. Biophys. Res. Commun. 2020, 524, 472–476. [Google Scholar] [CrossRef]
- Phan, A.T.; Kuryavyi, V.; Burge, S.; Neidle, S.; Patel, D.J. Structure of an unprecedented G-quadruplex scaffold in the human c-kit promoter. J. Am. Chem. Soc. 2007, 129, 4386–4392. [Google Scholar] [CrossRef]
- Koziol, M.J.; Bradshaw, C.R.; Allen, G.E.; Costa, A.S.H.; Frezza, C.; Gurdon, J.B. Identification of methylated deoxyadenosines in vertebrates reveals diversity in DNA modifications. Nat. Struct. Mol. Biol. 2016, 23, 24–30. [Google Scholar] [CrossRef]
- Cavalieri, V.; Melfi, R.; Spinelli, G. Promoter activity of the sea urchin (Paracentrotus lividus) nucleosomal H3 and H2A and linker H1 α-histone genes is modulated by enhancer and chromatin insulator. Nucleic Acids Res. 2009, 37, 7407–7415. [Google Scholar] [CrossRef]
- Cavalieri, V.; Spinelli, G. Ectopic hbox12 expression evoked by histone deacetylase inhibition disrupts axial specification of the sea urchin embryo. PLoS ONE 2015, 10, e0143860. [Google Scholar] [CrossRef]
- Cavalieri, V. Histones, their variants and post-translational modifications in zebrafish development. Front. Cell Dev. Biol. 2020, 8, 456. [Google Scholar] [CrossRef]
- Di Caro, D.; Melfi, R.; Alessandro, C.; Serio, G.; Di Caro, V.; Cavalieri, V.; Palla, F.; Spinelli, G. Down-regulation of early sea urchin histone H2A gene relies on cis regulative sequences located in the 5′ and 3′ regions and including the enhancer blocker sns. J. Mol. Biol. 2004, 342, 1367–1377. [Google Scholar] [CrossRef]
- Di Caro, V.; Cavalieri, V.; Melfi, R.; Spinelli, G. Regulated sea urchin alpha-H2A histone gene. J. Mol. Biol. 2007, 365, 1285–1297. [Google Scholar] [CrossRef]
- Alabert, C.; Jasencakova, Z.; Groth, A. Chromatin replication and histone dynamics. Adv. Exp. Med. Biol. 2017, 1042, 311–333. [Google Scholar] [CrossRef]
- Hammond, C.M.; Strømme, C.B.; Huang, H.; Patel, D.J.; Groth, A. Histone chaperone networks shaping chromatin function. Nat. Rev. Mol. Cell Biol. 2017, 18, 141–158. [Google Scholar] [CrossRef]
- Brosh, R.M., Jr.; Majumdar, A.; Desai, S.; Hickson, I.D.; Bohr, V.A.; Seidman, M.M. Unwinding of a DNA triple helix by the Werner and Bloom syndrome helicases. J. Biol. Chem. 2001, 276, 3024–3030. [Google Scholar] [CrossRef]
- Sale, J.E.; Lehmann, A.R.; Woodgate, R. Y-family DNA polymerases and their role in tolerance of cellular DNA damage. Nat. Rev. Mol. Cell Biol. 2012, 13, 141–152. [Google Scholar] [CrossRef]
- Sanders, C.M. Human Pif1 helicase is a G-quadruplex DNA-binding protein with G-quadruplex DNA-unwinding activity. Biochem. J. 2010, 430, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Shin-ya, K.; Brosh, R.M., Jr. FANCJ helicase defective in Fanconia anemia and breast cancer unwinds G-quadruplex DNA to defend genomic stability. Mol. Cell Biol. 2008, 28, 4116–4128. [Google Scholar] [CrossRef] [PubMed]
- Jasencakova, Z.; Scharf, A.N.; Ask, K.; Corpet, A.; Imhof, A.; Almouzni, G.; Groth, A. Replication stress interferes with histone recycling and predeposition marking of new histones. Mol. Cell 2010, 37, 736–743. [Google Scholar] [CrossRef] [PubMed]
- Sarkies, P.; Murat, P.; Phillips, L.G.; Patel, K.J.; Balasubramanian, S.; Sale, J.E. FANCJ coordinates two pathways that maintain epigenetic stability at G-quadruplex DNA. Nucleic Acids Res. 2012, 40, 1485–1498. [Google Scholar] [CrossRef] [PubMed]
- Schiavone, D.; Guilbaud, G.; Murat, P.; Papadopoulou, C.; Sarkies, P.; Prioleau, M.N.; Balasubramanian, S.; Sale, J.E. Determinants of G quadruplex-induced epigenetic instability in REV1-deficient cells. EMBO J. 2014, 33, 2507–2520. [Google Scholar] [CrossRef]
- Schwab, R.A.; Nieminuszczy, J.; Shin-ya, K.; Niedzwiedz, W. FANCJ couples replication past natural fork barriers with maintenance of chromatin structure. J. Cell Biol. 2013, 201, 33–48. [Google Scholar] [CrossRef]
- Takahama, K.; Takada, A.; Tada, S.; Shimizu, M.; Sayama, K.; Kurokawa, R.; Oyoshi, T. Regulation of telomere length by G-quadruplex telomere DNA- and TERRA-binding protein TLS/FUS. Chem. Biol. 2013, 20, 341–350. [Google Scholar] [CrossRef]
- Hou, Y.; Li, F.; Zhang, R.; Li, S.; Liu, H.; Qin, Z.S.; Sun, X. Integrative characterization of G-Quadruplexes in the three-dimensional chromatin structure. Epigenetics 2019, 14, 894–911. [Google Scholar] [CrossRef]
- Hirschi, A.; Martin, W.J.; Luka, Z.; Loukachevitch, L.V.; Reiter, N.J. G-quadruplex RNA binding and recognition by the lysine-specific Histone Demethylase-1 Enzyme. RNA 2016, 22, 1250–1260. [Google Scholar] [CrossRef]
- Hussain, T.; Saha, D.; Purohit, G.; Kar, A.; Kishore Mukherjee, A.; Sharma, S.; Sengupta, S.; Dhapola, P.; Maji, B.; Vedagopuram, S.; et al. Transcription regulation of CDKN1A (p21/CIP1/WAF1) by TRF2 is epigenetically controlled through the REST repressor complex. Sci. Rep. 2017, 7, 11541. [Google Scholar] [CrossRef]
- Saha, D.; Singh, A.; Hussain, T.; Srivastava, V.; Sengupta, S.; Kar, A.; Dhapola, P.; Dhople, V.; Ummanni, R.; Chowdhury, S. Epigenetic suppression of human telomerase (hTERT) is mediated by the metastasis suppressor NME2 in a G-quadruplex-dependent fashion. J. Biol. Chem. 2017, 292, 15205–15215. [Google Scholar] [CrossRef] [PubMed]
- Thakur, R.K.; Kumar, P.; Halder, K.; Verma, A.; Kar, A.; Parent, J.L.; Basundra, R.; Kumar, A.; Chowdhury, S. Metastases suppressor NM23-H2 interaction with G-quadruplex DNA within c-MYC promoter nuclease hypersensitive element induces c-MYC expression. Nucleic Acids Res. 2009, 37, 172–183. [Google Scholar] [CrossRef] [PubMed]
- Vlasenok, M.; Levchenko, O.; Basmanov, D.; Klinov, D.; Varizhuk, A.; Pozmogova, G. Data set on G4 DNA interactions with human proteins. Data Brief 2018, 18, 348–359. [Google Scholar] [CrossRef] [PubMed]
- Cuthbert, G.L.; Daujat, S.; Snowden, A.W.; Erdjument-Bromage, H.; Hagiwara, T.; Yamada, M.; Schneider, R.; Gregory, P.D.; Tempst, P.; Bannister, A.J.; et al. Histone deimination antagonizes arginine methylation. Cell 2004, 118, 545–553. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Wahab, O.; Adli, M.; LaFave, L.M.; Gao, J.; Hricik, T.; Shih, A.H.; Pandey, S.; Patel, J.P.; Chung, Y.R.; Koche, R.; et al. ASXL1 mutations promote myeloid transformation through loss of PRC2-mediated gene repression. Cancer Cell 2012, 22, 180–193. [Google Scholar] [CrossRef]
- Fisher, C.L.; Berger, J.; Randazzo, F.; Brock, H.W. A human homolog of Additional sex combs, ADDITIONAL SEX COMBS-LIKE 1, maps to chromosome 20q11. Gene 2003, 306, 115–126. [Google Scholar] [CrossRef]
- Beltran, M.; Tavares, M.; Justin, N.; Khandelwal, G.; Ambrose, J.; Foster, B.M.; Worlock, K.B.; Tvardovskiy, A.; Kunzelmann, S.; Herrero, J.; et al. G-tract RNA removes Polycomb repressive complex 2 from genes. Nat. Struct. Mol. Biol. 2019, 26, 899–909. [Google Scholar] [CrossRef]
- Wang, X.; Paucek, R.D.; Gooding, A.R.; Brown, Z.Z.; Ge, E.J.; Muir, T.W.; Cech, T.R. Molecular analysis of PRC2 recruitment to DNA in chromatin and its inhibition by RNA. Nat. Struct. Mol. Biol. 2017, 24, 1028–1038. [Google Scholar] [CrossRef]
- Nguyen, D.T.; Voon, H.P.J.; Xella, B.; Scott, C.; Clynes, D.; Babbs, C.; Ayyub, H.; Kerry, J.; Sharpe, J.A.; Sloane-Stanley, J.A.; et al. The chromatin remodelling factor ATRX suppresses R-loops in transcribed telomeric repeats. EMBO Rep. 2017, 18, 914–928. [Google Scholar] [CrossRef]
- Udugama, M.; Chang, F.T.; Chan, F.L.; Tang, M.C.; Pickett, H.A.; McGhie, J.D.; Mayne, L.; Collas, P.; Mann, J.R.; Wong, L.H. Histone variant H3.3 provides the heterochromatic H3 lysine 9 tri-methylation mark at telomeres. Nucleic Acids Res. 2015, 43, 10227–10237. [Google Scholar] [CrossRef]
- Angelov, D.; Bondarenko, V.A.; Almagro, S.; Menoni, H.; Mongélard, F.; Hans, F.; Mietton, F.; Studitsky, V.M.; Hamiche, A.; Dimitrov, S.; et al. Nucleolin is a histone chaperone with FACT-like activity and assists remodeling of nucleosomes. EMBO J. 2006, 25, 1669–1679. [Google Scholar] [CrossRef] [PubMed]
- Dempsey, L.A.; Sun, H.; Hanakahi, L.A.; Maizels, N. G4 DNA binding by LR1 and its subunits, nucleolin and hnRNP D, a role for G-G pairing in immunoglobulin switch recombination. J. Biol. Chem. 1999, 274, 1066–1071. [Google Scholar] [CrossRef] [PubMed]
- González, V.; Guo, K.; Hurley, L.; Sun, D. Identification and characterization of nucleolin as a c-myc G-quadruplex-binding protein. J. Biol. Chem. 2009, 284, 23622–23635. [Google Scholar] [CrossRef] [PubMed]
- Bustin, M. Revised nomenclature for high mobility group (HMG) chromosomal proteins. Trends Biochem. Sci. 2001, 26, 152–153. [Google Scholar] [CrossRef]
- Malarkey, C.S.; Churchill, M.E. The high mobility group box: The ultimate utility player of a cell. Trends Biochem. Sci. 2012, 37, 553–562. [Google Scholar] [CrossRef] [PubMed]
- Postnikov, Y.; Bustin, M. Regulation of chromatin structure and function by HMGN proteins. Biochim. et Biophys. Acta 2010, 1799, 62–68. [Google Scholar] [CrossRef]
- Amato, J.; Madanayake, T.W.; Iaccarino, N.; Novellino, E.; Randazzo, A.; Hurley, L.H.; Pagano, B. HMGB1 binds to the KRAS promoter G-quadruplex: A new player in oncogene transcriptional regulation? Chem. Commun. 2018, 54, 9442–9445. [Google Scholar] [CrossRef] [PubMed]
- Cogoi, S.; Paramasivam, M.; Membrino, A.; Yokoyama, K.K.; Xodo, L.E. The KRAS promoter responds to Myc-associated zinc finger and poly (ADP-ribose) polymerase 1 proteins, which recognize a critical quadruplex-forming GA-element. J. Biol. Chem. 2010, 285, 22003–22016. [Google Scholar] [CrossRef]
- Muthurajan, U.M.; Hepler, M.R.; Hieb, A.R.; Clark, N.J.; Kramer, M.; Yao, T.; Luger, K. Automodification switches PARP-1 function from chromatin architectural protein to histone chaperone. Proc. Natl. Acad. Sci. USA 2014, 111, 12752–12757. [Google Scholar] [CrossRef]
- Gushchanskaya, E.S.; Artemov, A.V.; Ulyanov, S.V.; Logacheva, M.D.; Penin, A.A.; Kotova, E.S.; Akopov, S.B.; Nikolaev, L.G.; Iarovaia, O.V.; Sverdlov, E.D.; et al. The clustering of CpG islands may constitute an important determinant of the 3D organization of interphase chromosomes. Epigenetics 2014, 9, 951–963. [Google Scholar] [CrossRef][Green Version]
- Halder, K.; Halder, R.; Chowdhury, S. Genome-wide analysis predicts DNA structural motifs as nucleosome exclusion signals. Mol. Biosyst. 2009, 5, 1703–1712. [Google Scholar] [CrossRef] [PubMed]
- Kouzine, F.; Wojtowicz, D.; Baranello, L.; Yamane, A.; Nelson, S.; Resch, W.; Kieffer-Kwon, K.R.; Benham, C.J.; Casellas, R.; Przytycka, T.M.; et al. Permanganate/S1 Nuclease footprinting reveals non-b dna structures with regulatory potential across a mammalian genome. Cell Syst. 2017, 4, 344–356. [Google Scholar] [CrossRef]
- Wong, H.M.; Huppert, J.L. Stable G-quadruplexes are found outside nucleosome-bound regions. Mol. Biosyst. 2009, 5, 1713–1719. [Google Scholar] [CrossRef]
- Foulk, M.S.; Urban, J.M.; Casella, C.; Gerbi, S.A. Characterizing and controlling intrinsic biases of lambda exonuclease in nascent strand sequencing reveals phasing between nucleosomes and G-quadruplex motifs around a subset of human replication origins. Genome Res. 2015, 25, 725–735. [Google Scholar] [CrossRef] [PubMed]
- Baiamonte, E.; Spinelli, G.; Maggio, A.; Acuto, S.; Cavalieri, V. the sea urchin sns5 chromatin insulator shapes the chromatin architecture of a lentivirus vector integrated in the mammalian genome. Nucleic Acid Ther. 2016, 26, 318–326. [Google Scholar] [CrossRef] [PubMed]
- Cavalieri, V.; Melfi, R.; Spinelli, G. The Compass-like locus, exclusive to the Ambulacrarians, encodes a chromatin insulator binding protein in the Sea Urchin embryo. PLoS Genet. 2013, 9, e1003847. [Google Scholar] [CrossRef] [PubMed]
- Dixon, J.R.; Selvaraj, S.; Yue, F.; Kim, A.; Li, Y.; Shen, Y.; Hu, M.; Liu, J.S.; Ren, B. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 2012, 485, 376–380. [Google Scholar] [CrossRef]
- Lee, B.K.; Iyer, V.R. Genome-wide studies of CCCTC-binding factor (CTCF) and cohesin provide insight into chromatin structure and regulation. J. Biol. Chem. 2012, 287, 30906–30913. [Google Scholar] [CrossRef]
- Ong, C.T.; Corces, V.G. CTCF: An architectural protein bridging genome topology and function. Nat. Rev. Genet. 2014, 15, 234–246. [Google Scholar] [CrossRef]
- Hegyi, H. Enhancer-promoter interaction facilitated by transiently forming G-quadruplexes. Sci. Rep. 2015, 5, 9165. [Google Scholar] [CrossRef]
- Joachimi, A.; Benz, A.; Hartig, J.S. A comparison of DNA and RNA quadruplex structures and stabilities. Bioorg. Med. Chem. 2009, 17, 6811–6815. [Google Scholar] [CrossRef]
- Liu, H.; Kugimiya, A.; Sakurai, T.; Katahira, M.; Uesugi, S. A comparison of the properties and the solution structure for RNA and DNA quadruplexes which contain two GGAGG sequences joined with a tetranucleotide linker. Nucleosides Nucleotides Nucleic Acids 2002, 21, 785–801. [Google Scholar] [CrossRef] [PubMed]
- Olsen, C.M.; Marky, L.A. Energetic and hydration contributions of the removal of methyl groups from thymine to form uracil in G-quadruplexes. J. Phys. Chem. B 2009, 113, 9–11. [Google Scholar] [CrossRef] [PubMed]
- Zaccaria, F.; Fonseca Guerra, C. RNA versus DNA G-Quadruplex: The origin of increased stability. Chemistry 2018, 24, 16315–16322. [Google Scholar] [CrossRef] [PubMed]
- Tang, C.F.; Shafer, R.H. Engineering the quadruplex fold: Nucleoside conformation determines both folding topology and molecularity in Guanine quadruplexes. J. Am. Chem. Soc. 2006, 128, 5966–5973. [Google Scholar] [CrossRef]
- Zhang, J.Y.; Xia, Y.; Hao, Y.H.; Tan, Z. DNA:RNA hybrid G-quadruplex formation upstream of transcription start site. Sci. Rep. 2020, 10, 7429. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, T.; Roy, S.; Kumar, S.; Chakraborty, T.K.; Maiti, S. In the sense of transcription regulation by G-quadruplexes: Asymmetric effects in sense and antisense strands. Biochemistry 2014, 53, 3711–3718. [Google Scholar] [CrossRef]
- Xiao, S.; Zhang, J.Y.; Zheng, K.W.; Hao, Y.H.; Tan, Z. Bioinformatic analysis reveals an evolutional selection for DNA: RNA hybrid G-quadruplex structures as putative transcription regulatory elements in warm-blooded animals. Nucleic Acids Res. 2013, 41, 10379–10390. [Google Scholar] [CrossRef]
- Arora, A.; Dutkiewicz, M.; Scaria, V.; Hariharan, M.; Maiti, S.; Kurreck, J. Inhibition of translation in living eukaryotic cells by an RNA G-quadruplex motif. RNA 2008, 14, 1290–1296. [Google Scholar] [CrossRef]
- Gomez, D.; Guedin, A.; Mergny, J.L.; Salles, B.; Riou, J.F.; Teulade-Fichou, M.P.; Calsou, P.A. G-quadruplex structure within the 50-UTR of TRF2 mRNA represses translation in human cells. Nucleic Acids Res. 2010, 38, 7187–7198. [Google Scholar] [CrossRef]
- Kumari, S.; Bugaut, A.; Huppert, J.L.; Balasubramanian, S. An RNA G-quadruplex in the 5′-UTR of the NRAS proto-oncogene modulates translation. Nat. Chem. Biol. 2007, 3, 218–221. [Google Scholar] [CrossRef]
- Morris, M.J.; Basu, S. An unusually stable G-quadruplex within the 5′-UTR of the MT3 matrix metalloproteinase mRNA represses translation in eukaryotic cells. Biochemistry 2009, 48, 5313–5319. [Google Scholar] [CrossRef] [PubMed]
- Shahid, R.; Bugaut, A.; Balasubramanian, S. The BCL-2 5′ untranslated region contains an RNA G-quadruplex-forming motif that modulates protein expression. Biochemistry 2010, 49, 8300–8306. [Google Scholar] [CrossRef] [PubMed]
- Bugaut, A. Balasubramanian, S. 5′-UTR RNA G-quadruplexes: Translation regulation and targeting. Nucleic Acids Res. 2012, 40, 4727–4741. [Google Scholar] [CrossRef] [PubMed]
- Agarwala, P.; Pandey, S.; Mapa, K.; Maiti, S. The G-quadruplex augments translation in the 5’ untranslated region of transforming growth factor β2. Biochemistry 2013, 52, 1528–1538. [Google Scholar] [CrossRef] [PubMed]
- Agarwala, P.; Pandey, S.; Maiti, S. Role of G-quadruplex located at 5’ end of mRNAs. Biochim. et Biophys. Acta 2014, 1840, 3503–3510. [Google Scholar] [CrossRef]
- Agarwala, P.; Pandey, S.; Ekka, M.K.; Chakraborty, D.; Maiti, S. Combinatorial role of two G-quadruplexes in 5’ UTR of transforming growth factor β2 (TGFβ2). Biochim. et Biophys. Acta Gen. Subj. 2019, 1863, 129416. [Google Scholar] [CrossRef]
- Bonnal, S.; Schaeffer, C.; Créancier, L.; Clamens, S.; Moine, H.; Prats, A.C.; Vagner, S. A single internal ribosome entry site containing a G-quartet RNA structure drives fibroblast growth factor 2 gene expression at four alternative translation initiation codons. J. Biol. Chem. 2003, 278, 39330–39336. [Google Scholar] [CrossRef]
- Bhattacharyya, D.; Diamond, P.; Basu, S. An independently folding RNA G-quadruplex domain directly recruits the 40S ribosomal subunit. Biochemistry 2015, 54, 1879–1885. [Google Scholar] [CrossRef]
- Morris, M.J.; Negishi, Y.; Pazsint, C.; Schonhoft, J.D.; Basu, S. An RNA G-quadruplex is essential for cap-independent translation initiation in human VEGF IRES. J. Am. Chem. Soc. 2010, 132, 17831–17839. [Google Scholar] [CrossRef]
- Reddy, K.; Zamiri, B.; Stanley, S.Y.; Macgregor, R.B., Jr.; Pearson, C.E. The disease-associated r(GGGGCC)n repeat from the C9orf72 gene forms tract length-dependent uni- and multimolecular RNA G-quadruplex structures. J. Biol. Chem. 2013, 288, 9860–9866. [Google Scholar] [CrossRef] [PubMed]
- Beaudoin, J.D.; Perreault, J.P. Exploring mRNA 3’-UTR G-quadruplexes: Evidence of roles in both alternative polyadenylation and mRNA shortening. Nucleic Acids Res. 2013, 41, 5898–5911. [Google Scholar] [CrossRef] [PubMed]
- Didiot, M.C.; Tian, Z.; Schaeffer, C.; Subramanian, M.; Mandel, J.L.; Moine, H. The G-quartet containing FMRP binding site in FMR1mRNAis a potent exonic splicing enhancer. Nucleic Acids Res. 2008, 36, 4902–4912. [Google Scholar] [CrossRef] [PubMed]
- Gomez, D.; Lemarteleur, T.; Lacroix, L.; Mailliet, P.; Mergny, J.L.; Riou, J.F. Telomerase downregulation induced by the G-quadruplex ligand 12459 in A549 cells is mediated by hTERT RNA alternative splicing. Nucleic Acids Res. 2004, 32, 371–379. [Google Scholar] [CrossRef]
- Huang, H.; Zhang, J.; Harvey, S.E.; Hu, X.; Cheng, C. RNA G-quadruplex secondary structure promotes alternative splicing via the RNA-binding protein hnRNPF. Genes Dev. 2017, 31, 2296–2309. [Google Scholar] [CrossRef] [PubMed]
- Endoh, T.; Sugimoto, N. Unusual -1 ribosomal frameshift caused by stable RNA G-quadruplex in open reading frame. Anal. Chem. 2013, 85, 11435–11439. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.H.; Teulade-Fichou, M.P.; Olsthoorn, R.C. Stimulation of ribosomal frameshifting by RNA G-quadruplex structures. Nucleic Acids Res. 2014, 42, 1887–1892. [Google Scholar] [CrossRef]
- Imperatore, J.A.; McAninch, D.S.; Valdez-Sinon, A.N.; Bassell, G.J.; Mihailescu, M.R. FUS recognizes G-quadruplex structures within neuronal mRNAs. Front. Mol. Biosci. 2020, 7, 6. [Google Scholar] [CrossRef]
- Ishiguro, A.; Kimura, N.; Watanabe, Y.; Watanabe, S.; Ishihama, A. TDP-43 binds and transports G-quadruplex-containing mRNAs into neurites for local translation. Genes Cells 2016, 21, 466–481. [Google Scholar] [CrossRef]
- Rihan, K.; Antoine, E.; Maurin, T.; Bardoni, B.; Bordonné, R.; Soret, J.; Rage, F. A new cis-acting motif is required for the axonal SMN-dependent Anxa2 mRNA localization. RNA 2017, 23, 899–909. [Google Scholar] [CrossRef][Green Version]
- Subramanian, M.; Rage, F.; Tabet, R.; Flatter, E.; Mandel, J.L.; Moine, H. G-quadruplex RNA structure as a signal for neurite mRNA targeting. EMBO Rep. 2011, 12, 697–704. [Google Scholar] [CrossRef]
- Zhang, Y.; Gaetano, C.M.; Williams, K.R.; Bassell, G.J.; Mihailescu, M.R. FMRP interacts with G-quadruplex structures in the 3′-UTR of its dendritic target Shank1 mRNA. RNA Biol. 2014, 11, 1364–1374. [Google Scholar] [CrossRef]
- Jayaraj, G.G.; Pandey, S.; Scaria, V.; Maiti, S. Potential G-quadruplexes in the human long non-coding transcriptome. RNA Biol. 2012, 9, 81–86. [Google Scholar] [CrossRef]
- Yang, S.Y.; Lejault, P.; Chevrier, S.; Boidot, R.; Robertson, A.G.; Wong, J.M.Y.; Monchaud, D. Transcriptome-wide identification of transient RNA G-quadruplexes in human cells. Nat. Commun. 2018, 9, 4730. [Google Scholar] [CrossRef] [PubMed]
- Mirihana Arachchilage, G.; Dassanayake, A.C.; Basu, S. A potassium ion-dependent RNA structural switch regulates human pre-miRNA 92b maturation. Chem. Biol. 2015, 22, 262–272. [Google Scholar] [CrossRef]
- Pandey, S.; Agarwala, P.; Jayaraj, G.G.; Gargallo, R.; Maiti, S. The RNA stem-Loop to G-quadruplex equilibrium controls mature microRNA production inside the cell. Biochemistry 2015, 54, 7067–7078. [Google Scholar] [CrossRef]
- Rouleau, S.G.; Garant, J.M.; Bolduc, F.; Bisaillon, M.; Perreault, J.P. G-Quadruplexes influence pri-microRNA processing. RNA Biol. 2018, 15, 198–206. [Google Scholar] [CrossRef]
- Liu, G.; Du, W.; Xu, H.; Sun, Q.; Tang, D.; Zou, S.; Zhang, Y.; Ma, M.; Zhang, G.; Du, X.; et al. RNA G-quadruplex regulates microRNA-26a biogenesis and function. J. Hepatol. 2020. [Google Scholar] [CrossRef]
- Chan, K.L.; Peng, B.; Umar, M.I.; Chan, C.Y.; Sahakyan, A.B.; Le, M.T.N.; Kwok, C.K. Structural analysis reveals the formation and role of RNA G-quadruplex structures in human mature microRNAs. Chem. Commun. 2018, 54, 10878–10881. [Google Scholar] [CrossRef]
- Sahakyan, A.B.; Murat, P.; Mayer, C.; Balasubramanian, S. G-quadruplex structures within the 3′-UTR of LINE-1 elements stimulate retrotransposition. Nat. Struct. Mol. Biol. 2017, 24, 243–247. [Google Scholar] [CrossRef]
- Balaratnam, S.; Hettiarachchilage, M.; West, N.; Piontkivska, H.; Basu, S. A secondary structure within a human piRNA modulates its functionality. Biochimie 2019, 157, 72–80. [Google Scholar] [CrossRef] [PubMed]
- Hirashima, K.; Seimiya, H. Telomeric repeat-containing RNA/G-quadruplex-forming sequences cause genome-wide alteration of gene expression in human cancer cells in vivo. Nucleic Acids Res. 2015, 43, 2022–2032. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Komiyama, M. Structure, function and targeting of human telomere RNA. Methods 2012, 57, 100–105. [Google Scholar] [CrossRef] [PubMed]



© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Reina, C.; Cavalieri, V. Epigenetic Modulation of Chromatin States and Gene Expression by G-Quadruplex Structures. Int. J. Mol. Sci. 2020, 21, 4172. https://doi.org/10.3390/ijms21114172
Reina C, Cavalieri V. Epigenetic Modulation of Chromatin States and Gene Expression by G-Quadruplex Structures. International Journal of Molecular Sciences. 2020; 21(11):4172. https://doi.org/10.3390/ijms21114172
Chicago/Turabian StyleReina, Chiara, and Vincenzo Cavalieri. 2020. "Epigenetic Modulation of Chromatin States and Gene Expression by G-Quadruplex Structures" International Journal of Molecular Sciences 21, no. 11: 4172. https://doi.org/10.3390/ijms21114172
APA StyleReina, C., & Cavalieri, V. (2020). Epigenetic Modulation of Chromatin States and Gene Expression by G-Quadruplex Structures. International Journal of Molecular Sciences, 21(11), 4172. https://doi.org/10.3390/ijms21114172




