The Potential of CD16 on Plasma-Derived Exosomes as a Liquid Biomarker in Head and Neck Cancer
Abstract
1. Introduction
2. Results
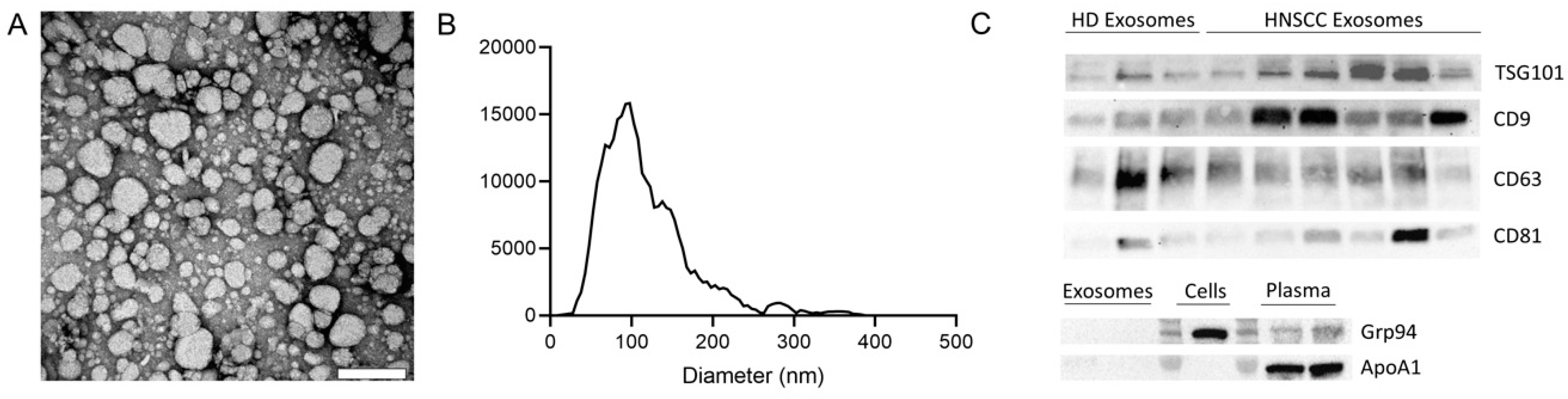
2.1. Characterization of Exosomes
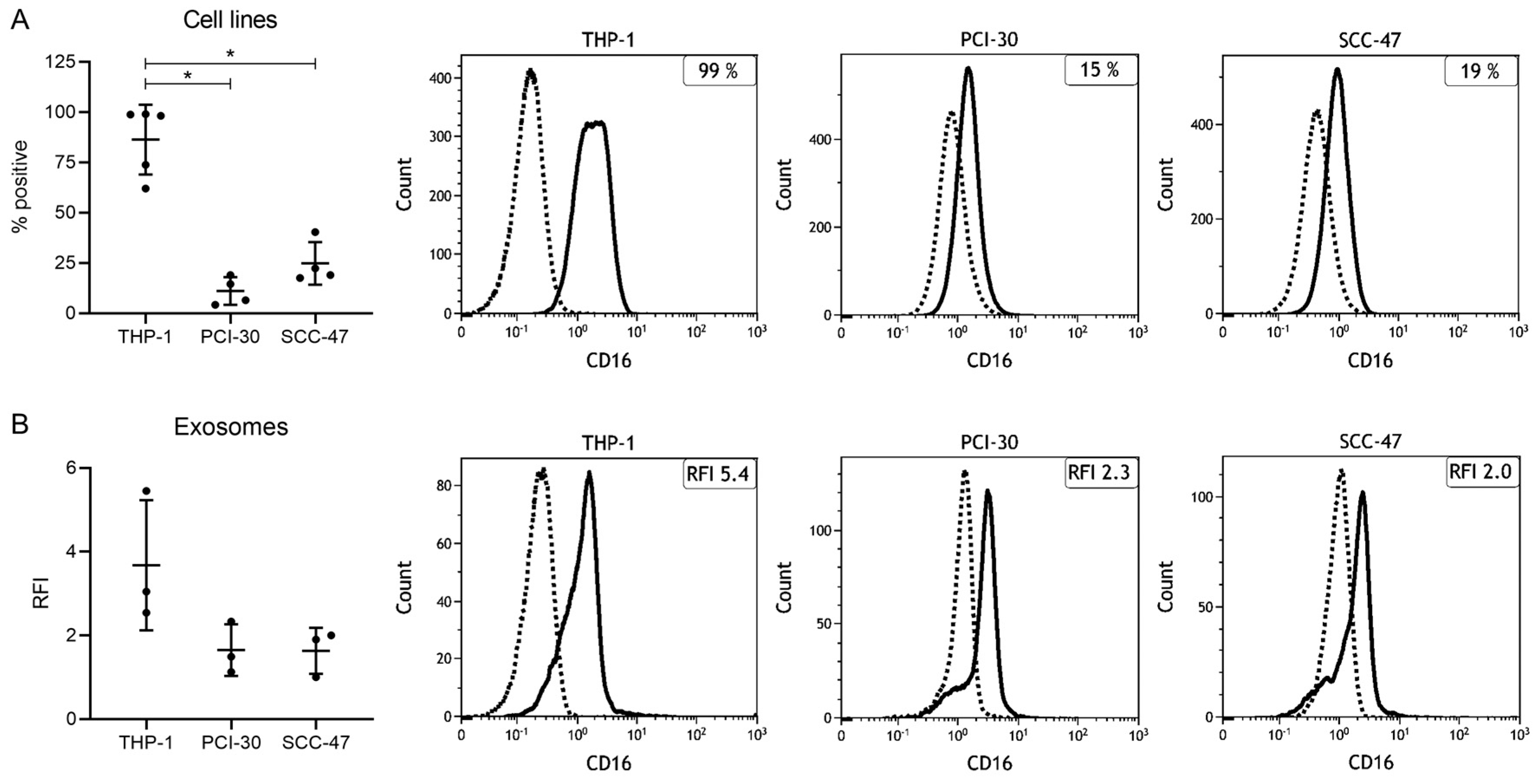
2.2. Expression Levels of CD16 in Cell Lines and Cell Line-Derived Exosomes
2.3. Clinicopathological Characteristics of HNSCC Patients
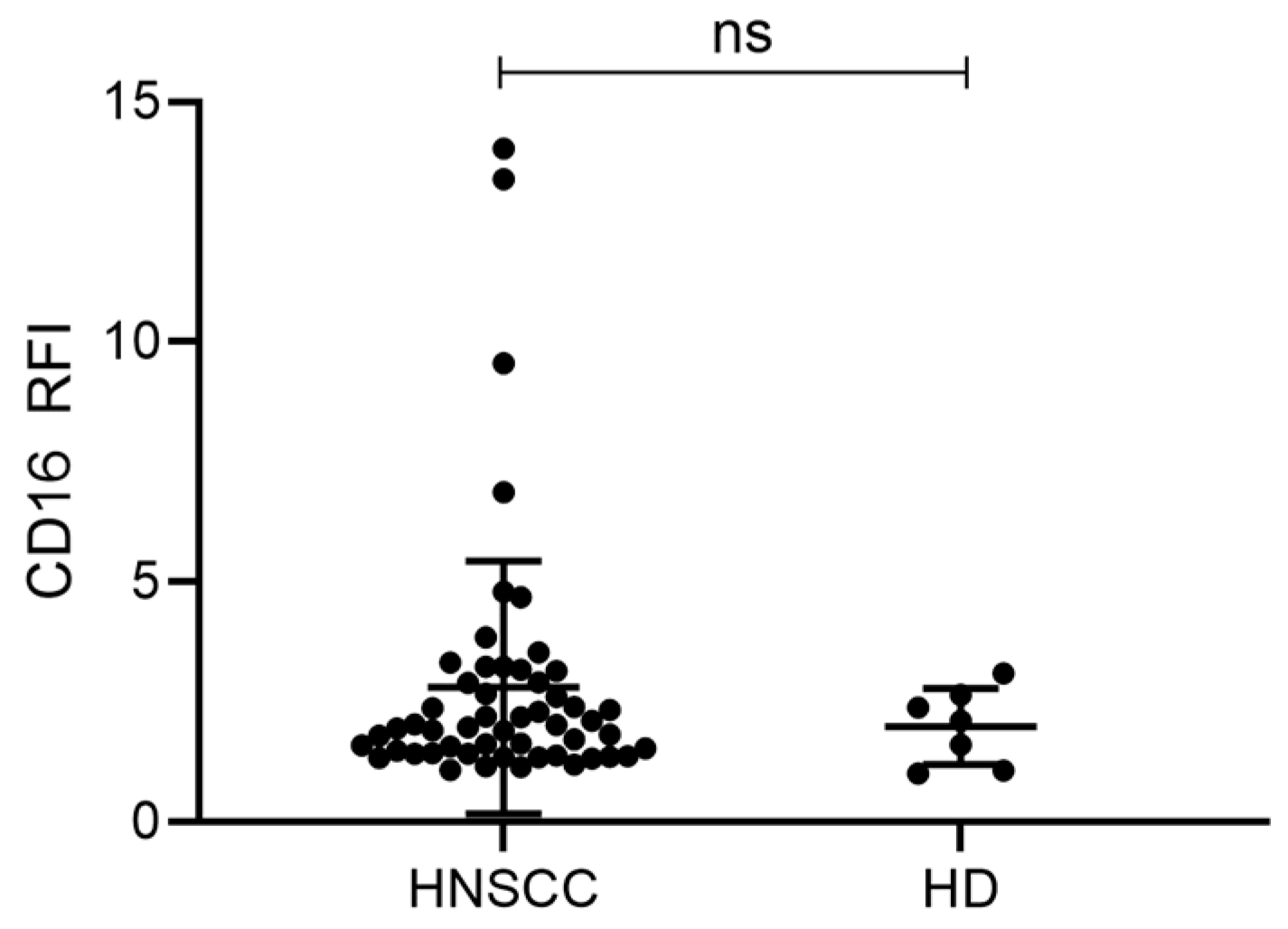
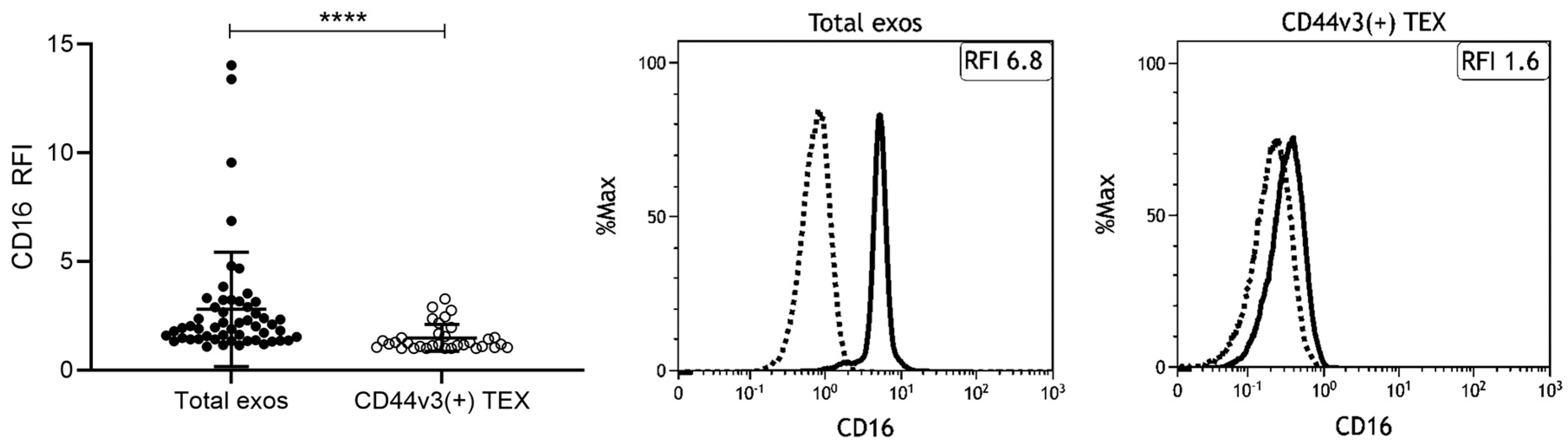
2.4. Exosomes from HNSCC Carry Surface CD16
2.5. Correlation of CD16 Surface Levels on Exosomes with Clinicopathological Parameters
3. Discussion
4. Materials and Methods
4.1. Cell Lines
4.2. Patients
4.3. Exosome Isolation by Mini Size Exclusion Chromatography (Mini-SEC)
4.4. BCA and Exosome Concentration
4.5. Characterization of Exosomes
4.6. Flow Cytometry of Cells
4.7. Immune Capture and On-Bead Flow Cytometry of Exosomes
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AD | Active disease |
| ADCC | Antibody-dependent cell-mediated cytotoxicity |
| ApoA1 | Apolipoprotein A1 |
| EV | Extracellular vesicle |
| FcR | Fc receptor |
| Grp94 | Glucose-regulated protein 94 |
| HD | Healthy donor |
| HNSCC | Head and neck squamous cell carcinoma |
| HPV | Human papilloma virus |
| mAB | Monoclonal antibody |
| MFI | Mean fluorescence intensity |
| NED | No evidence of disease |
| NK | Natural killer |
| NTA | Nanoparticle tracking analysis |
| RFI | Relative fluorescence intensity |
| RT | Room temperature |
| SEC | Size exclusion chromatography |
| TEM | Transmission electron microscopy |
| TEX | Tumor-derived exosomes |
| TME | Tumor microenvironment |
| TSG101 | Tumor susceptibility gene 101 |
| UICC | Union for International Cancer Control |
References
- Chakraborty, P.; Karmakar, T.; Arora, N.; Mukherjee, G. Immune and genomic signatures in oral (head and neck) cancer. Heliyon 2018, 4, e00880. [Google Scholar] [CrossRef]
- Bose, A.; Chakraborty, T.; Chakraborty, K.; Pal, S.; Baral, R. Dysregulation in immune functions is reflected in tumor cell cytotoxicity by peripheral blood mononuclear cells from head and neck squamous cell carcinoma patients. Cancer Immun. 2008, 8, 10. [Google Scholar]
- Lam-ubol, A.; Hopkin, D.; Letuchy, E.M.; Kurago, Z.B. Squamous carcinoma cells influence monocyte phenotype and suppress lipopolysaccharide-induced TNF-alpha in monocytes. Inflammation 2010, 33, 207–223. [Google Scholar] [CrossRef][Green Version]
- Klöß, S.; Chambron, N.; Gardlowski, T.; Arseniev, L.; Koch, J.; Esser, R.; Glienke, W.; Seitz, O.; Köhl, U. Increased sMICA and TGFβ(1) levels in HNSCC patients impair NKG2D-dependent functionality of activated NK cells. Oncoimmunology 2015, 4, e1055993. [Google Scholar] [CrossRef]
- Wulff, S.; Pries, R.; Börngen, K.; Trenkle, T.; Wollenberg, B. Decreased levels of circulating regulatory NK cells in patients with head and neck cancer throughout all tumor stages. Anticancer Res. 2009, 29, 3053–3057. [Google Scholar]
- Millrud, C.R.; Månsson Kvarnhammar, A.; Uddman, R.; Björnsson, S.; Riesbeck, K.; Cardell, L.O. The Activation Pattern of Blood Leukocytes in Head and Neck Squamous Cell Carcinoma Is Correlated to Survival. PLoS ONE 2012, 7, e51120. [Google Scholar] [CrossRef]
- Naeim, F.; Nagesh Rao, P.; Song, S.X.; Phan, R.T. Chapter 2—Principles of Immunophenotyping. In Atlas of Hematopathology, 2nd ed.; Naeim, F., Nagesh Rao, P., Song, S.X., Phan, R.T., Eds.; Academic Press: Cambridge, MA, USA, 2018; pp. 29–56. [Google Scholar]
- Cooper, M.A.; Fehniger, T.A.; Caligiuri, M.A. The biology of human natural killer-cell subsets. Trends Immunol. 2001, 22, 633–640. [Google Scholar] [CrossRef]
- Lowry, L.E.; Zehring, W.A. Potentiation of Natural Killer Cells for Cancer Immunotherapy: A Review of Literature. Front. Immunol. 2017, 8, 1061. [Google Scholar] [CrossRef]
- James, A.; Cohen, A.; Campbell, K. Combination Immune Therapies to Enhance Anti-Tumor Responses by NK Cells. Front. Immunol. 2013, 4, 481. [Google Scholar]
- Whiteside, T.L. The effect of tumor-derived exosomes on immune regulation and cancer immunotherapy. Future Oncol. 2017, 13, 2583–2592. [Google Scholar] [CrossRef]
- Whiteside, T.L. Exosomes carrying immunoinhibitory proteins and their role in cancer. Clin. Exp. Immunol. 2017, 189, 259–267. [Google Scholar] [CrossRef] [PubMed]
- Ruivo, C.F.; Adem, B.; Silva, M.; Melo, S.A. The Biology of Cancer Exosomes: Insights and New Perspectives. Cancer Res. 2017, 77, 6480–6488. [Google Scholar] [CrossRef] [PubMed]
- Milane, L.; Singh, A.; Mattheolabakis, G.; Suresh, M.; Amiji, M.M. Exosome mediated communication within the tumor microenvironment. J. Control. Release 2015, 219, 278–294. [Google Scholar] [CrossRef]
- Abels, E.R.; Breakefield, X.O. Introduction to Extracellular Vesicles: Biogenesis, RNA Cargo Selection, Content, Release, and Uptake. Cell. Mol. Neurobiol. 2016, 36, 301–312. [Google Scholar] [CrossRef]
- Ludwig, S.; Floros, T.; Theodoraki, M.-N.; Hong, C.-S.; Jackson, E.K.; Lang, S.; Whiteside, T.L. Suppression of Lymphocyte Functions by Plasma Exosomes Correlates with Disease Activity in Patients with Head and Neck Cancer. Clin. Cancer Res. 2017, 23, 4843–4854. [Google Scholar] [CrossRef] [PubMed]
- Theodoraki, M.-N.; Yerneni, S.S.; Hoffmann, T.K.; Gooding, W.E.; Whiteside, T.L. Clinical Significance of PD-L1+ Exosomes in Plasma of Head and Neck Cancer Patients. Clin. Cancer Res. 2018, 24, 896–905. [Google Scholar] [CrossRef]
- Theodoraki, M.-N.; Yerneni, S.S.; Gooding, W.E.; Ohr, J.; Clump, D.A.; Bauman, J.E.; Ferris, R.L.; Whiteside, T.L. Circulating exosomes measure responses to therapy in head and neck cancer patients treated with cetuximab, ipilimumab, and IMRT. Oncoimmunology 2019, 8, 1593805. [Google Scholar] [CrossRef]
- Theodoraki, M.-N.; Hoffmann, T.K.; Whiteside, T.L. Separation of plasma-derived exosomes into CD3((+)) and CD3((−)) fractions allows for association of immune cell and tumour cell markers with disease activity in HNSCC patients. Clin. Exp. Immunol. 2018, 192, 271–283. [Google Scholar] [CrossRef]
- Theodoraki, M.-N.; Yerneni, S.S.; Brunner, C.; Theodorakis, J.; Hoffmann, T.K.; Whiteside, T.L. Plasma-derived Exosomes Reverse Epithelial-to-Mesenchymal Transition after Photodynamic Therapy of Patients with Head and Neck Cancer. Oncoscience 2018, 5, 75–87. [Google Scholar]
- Theodoraki, M.-N.; Matsumoto, A.; Beccard, I.; Hoffmann, T.K.; Whiteside, T.L. CD44v3 protein-carrying tumor-derived exosomes in HNSCC patients’ plasma as potential noninvasive biomarkers of disease activity. Oncoimmunology 2020, 9, 1747732. [Google Scholar] [CrossRef]
- Sagawa, K.; Uwa, N.; Daimon, T.; Sakagami, M.; Tsujimura, T. Expression of CD44 variant isoforms, CD44v3 and CD44v6, are associated with prognosis in nasopharyngeal carcinoma. J. Laryngol. Otol. 2016, 130, 843–849. [Google Scholar] [CrossRef] [PubMed]
- Spiegelberg, D.; Kuku, G.; Selvaraju, R.; Nestor, M. Characterization of CD44 variant expression in head and neck squamous cell carcinomas. Tumour Biol. 2014, 35, 2053–2062. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.J.; Wreesmann, V.B.; Bourguignon, L.Y.W. Association of CD44 V3-containing isoforms with tumor cell growth, migration, matrix metalloproteinase expression, and lymph node metastasis in head and neck cancer. Head Neck 2007, 29, 550–558. [Google Scholar] [CrossRef] [PubMed]
- Yoshioka, Y.; Konishi, Y.; Kosaka, N.; Katsuda, T.; Kato, T.; Ochiya, T. Comparative marker analysis of extracellular vesicles in different human cancer types. J. Extracell. Vesicles 2013, 2, 20424. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, N.; Hong, C.-S.; Ludwig, S.; Azambuja, J.H.; Sharma, P.; Theodoraki, M.-N.; Whiteside, T.L. Isolation and Analysis of Tumor-Derived Exosomes. Curr. Protoc. Immunol. 2019, 127, e91. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, N.; Razzo, B.M.; Yerneni, S.S.; Whiteside, T.L. Optimization of cell culture conditions for exosome isolation using mini-size exclusion chromatography (mini-SEC). Exp. Cell Res. 2019, 378, 149–157. [Google Scholar] [CrossRef]
- Hong, C.-S.; Funk, S.; Muller, L.; Boyiadzis, M.; Whiteside, T.L. Isolation of biologically active and morphologically intact exosomes from plasma of patients with cancer. J. Extracell. Vesicles 2016, 5, 29289. [Google Scholar] [CrossRef]
- Théry, C.; Witwer, K.W.; Aikawa, E.; Alcaraz, M.J.; Anderson, J.D.; Andriantsitohaina, R. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): A position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 2018, 7, 1535750. [Google Scholar] [CrossRef]
- Whiteside, T.L. The emerging role of plasma exosomes in diagnosis, prognosis and therapies of patients with cancer. Contemp. Oncol. (Pozn.) 2018, 22, 38–40. [Google Scholar] [CrossRef]
- Whiteside, T.L. Proteomic Analysis of Plasma-Derived Exosomes in Defining Their Role as Biomarkers of Disease Progression, Response to Therapy and Outcome. Proteomes 2019, 7, 27. [Google Scholar] [CrossRef]
- Yeap, W.H.; Wong, K.L.; Shimasaki, N.; Teo, E.C.Y.; Quek, J.K.S.; Yong, H.X.; Diong, C.P.; Bertoletti, A.; Linn, Y.C.; Wong, S.C. CD16 is indispensable for antibody-dependent cellular cytotoxicity by human monocytes. Sci. Rep. 2016, 6, 34310. [Google Scholar] [CrossRef] [PubMed]
- Kapellos, T.S.; Bonaguro, L.; Gemünd, I.; Reusch, N.; Saglam, A.; Hinkley, E.R.; Schultze, J.L. Human Monocyte Subsets and Phenotypes in Major Chronic Inflammatory Diseases. Front. Immunol. 2019, 10, 2035. [Google Scholar] [CrossRef]
- Wong, K.L.; Yeap, W.H.; Tai, J.J.Y.; Ong, S.M.; Dang, T.M.; Wong, S.C. The three human monocyte subsets: Implications for health and disease. Immunol. Res. 2012, 53, 41–57. [Google Scholar] [CrossRef] [PubMed]
- Clémenceau, B.; Vivien, R.; Debeaupuis, E.; Esbelin, J.; Biron, C.; Levy, Y.; Vié, H. FcγRIIIa (CD16) Induction on Human T Lymphocytes and CD16pos T-Lymphocyte Amplification. J. Immunother. 2011, 34, 542–549. [Google Scholar] [CrossRef] [PubMed]
- Björkström, N.K.; Gonzalez, V.D.; Malmberg, K.-J.; Falconer, K.; Alaeus, A.; Nowak, G.; Jorns, C.; Ericzon, B.-G.; Weiland, O.; Sandberg, J.K.; et al. Elevated Numbers of FcγRIIIA+ (CD16+) Effector CD8 T Cells with NK Cell-Like Function in Chronic Hepatitis C Virus Infection. J. Immunol. 2008, 181, 4219–4228. [Google Scholar] [CrossRef]
- Barbarin, A.; Cayssials, E.; Jacomet, F.; Nunez, N.G.; Basbous, S.; Lefèvre, L.; Abdallah, M.; Piccirilli, N.; Morin, B.; Lavoue, V.; et al. Phenotype of NK-Like CD8(+) T Cells with Innate Features in Humans and Their Relevance in Cancer Diseases. Front. Immunol. 2017, 8, 316. [Google Scholar] [CrossRef] [PubMed]
- Lo Nigro, C.; Macagno, M.; Sangiolo, D.; Bertolaccini, L.; Aglietta, M.; Merlano, M.C. NK-mediated antibody-dependent cell-mediated cytotoxicity in solid tumors: Biological evidence and clinical perspectives. Ann. Transl. Med. 2019, 7, 105. [Google Scholar] [CrossRef]
- Jewett, A.; Teruel, A.; Romero, M.; Head, C.; Cacalano, N. Rapid and potent induction of cell death and loss of NK cell cytotoxicity against oral tumors by F(ab′)2 fragment of anti-CD16 antibody. Cancer Immunol. Immunother. 2008, 57, 1053–1066. [Google Scholar] [CrossRef]
- Watanabe, M.; Kono, K.; Kawaguchi, Y.; Mizukami, Y.; Mimura, K.; Maruyama, T.; Izawa, S.; Fujii, H. NK cell dysfunction with down-regulated CD16 and up-regulated CD56 molecules in patients with esophageal squamous cell carcinoma. Dis. Esophagus 2010, 23, 675–681. [Google Scholar] [CrossRef]
- Türkseven, M.R.; Oygür, T. Evaluation of natural killer cell defense in oral squamous cell carcinoma. Oral Oncol. 2010, 46, e34–e37. [Google Scholar] [CrossRef]
- Dasgupta, S.; Bhattacharya-Chatterjee, M.; O’Malley, B.W.J.; Chatterjee, S.K. Inhibition of NK cell activity through TGF-beta 1 by down-regulation of NKG2D in a murine model of head and neck cancer. J. Immunol. 2005, 175, 5541–5550. [Google Scholar] [CrossRef]
- Bauernhofer, T.; Kuss, I.; Henderson, B.; Baum, A.S.; Whiteside, T.L. Preferential apoptosis of CD56dim natural killer cell subset in patients with cancer. Eur. J. Immunol. 2003, 33, 119–124. [Google Scholar] [CrossRef]
- Trivedi, S.; Srivastava, R.M.; Concha-Benavente, F.; Ferrone, S.; Garcia-Bates, T.M.; Li, J.; Ferris, R.L. Anti-EGFR Targeted Monoclonal Antibody Isotype Influences Antitumor Cellular Immunity in Head and Neck Cancer Patients. Clin. Cancer Res. 2016, 22, 5229–5237. [Google Scholar] [CrossRef] [PubMed]
- Ferris, R.L.; Lenz, H.-J.; Trotta, A.M.; García-Foncillas, J.; Schulten, J.; Audhuy, F.; Merlano, M.; Milano, G. Rationale for combination of therapeutic antibodies targeting tumor cells and immune checkpoint receptors: Harnessing innate and adaptive immunity through IgG1 isotype immune effector stimulation. Cancer Treat. Rev. 2018, 63, 48–60. [Google Scholar] [CrossRef] [PubMed]
- Veluchamy, J.P.; Spanholtz, J.; Tordoir, M.; Thijssen, V.L.; Heideman, D.A.M.; Verheul, H.M.W.; De Gruijl, T.D.; Van der Vliet, H.J. Combination of NK Cells and Cetuximab to Enhance Anti-Tumor Responses in RAS Mutant Metastatic Colorectal Cancer. PLoS ONE 2016, 11, e0157830. [Google Scholar] [CrossRef]
- Ciravolo, V.; Huber, V.; Ghedini, G.C.; Venturelli, E.; Bianchi, F.; Campiglio, M.; Morelli, D.; Villa, A.; Mina, P.D.; Menard, S.; et al. Potential role of HER2-overexpressing exosomes in countering trastuzumab-based therapy. J. Cell. Physiol. 2012, 227, 658–667. [Google Scholar] [CrossRef]
- Battke, C.; Ruiss, R.; Welsch, U.; Wimberger, P.; Lang, S.; Jochum, S.; Zeidler, R. Tumour exosomes inhibit binding of tumour-reactive antibodies to tumour cells and reduce ADCC. Cancer Immunol. Immunother. 2011, 60, 639–648. [Google Scholar] [CrossRef]
- Xie, F.; Xu, M.; Lu, J.; Mao, L.; Wang, S. The role of exosomal PD-L1 in tumor progression and immunotherapy. Mol. Cancer 2019, 18, 146. [Google Scholar] [CrossRef]
- Yang, Y.; Li, C.-W.; Chan, L.-C.; Wei, Y.; Hsu, J.-M.; Xia, W.; Cha, J.-H.; Hou, J.; Hsu, J.L.; Sun, L.; et al. Exosomal PD-L1 harbors active defense function to suppress T cell killing of breast cancer cells and promote tumor growth. Cell Res. 2018, 28, 862–864. [Google Scholar] [CrossRef]
- Bergmann, C.; Strauss, L.; Wang, Y.; Szczepanski, M.J.; Lang, S.; Johnson, J.T.; Whiteside, T.L. T regulatory type 1 cells in squamous cell carcinoma of the head and neck: Mechanisms of suppression and expansion in advanced disease. Clin. Cancer Res. 2008, 14, 3706–3715. [Google Scholar] [CrossRef] [PubMed]
- Lathers, D.M.R.; Young, M.R.I. Increased aberrance of cytokine expression in plasma of patients with more advanced squamous cell carcinoma of the head and neck. Cytokine 2004, 25, 220–228. [Google Scholar] [CrossRef] [PubMed]
- Sparano, A.; Lathers, D.M.R.; Achille, N.; Petruzzelli, G.J.; Young, M.R.I. Modulation of Th1 and Th2 Cytokine Profiles and Their Association with Advanced Head and Neck Squamous Cell Carcinoma. Otolaryngol. Neck Surg. 2004, 131, 573–576. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.J.; Grandis, J.R.; Carey, T.E.; Gollin, S.M.; Whiteside, T.L.; Koch, W.M.; Ferris, R.L.; Lai, S.Y. Head and neck squamous cell carcinoma cell lines: Established models and rationale for selection. Head Neck 2007, 29, 163–188. [Google Scholar] [CrossRef] [PubMed]

| Characteristics | Patients (n = 53) | |
|---|---|---|
| n | % | |
| Age (years) | ||
| ≤60 | 24 | 45 |
| >60 | 29 | 55 |
| (range: 36–84) | ||
| Gender | ||
| Male | 43 | 81 |
| Female | 10 | 19 |
| Disease status | ||
| AD | 42 | 79 |
| NED | 11 | 21 |
| Primary tumor site | ||
| Oral cavity | 23 | 43 |
| Pharynx | 15 | 28 |
| Larynx | 15 | 28 |
| Tumor stage | ||
| T1 | 13 | 25 |
| T2 | 14 | 26 |
| T3 | 8 | 15 |
| T4 | 18 | 34 |
| Nodal status | ||
| N0 | 17 | 32 |
| N1 | 16 | 30 |
| N2 | 14 | 26 |
| N3 | 6 | 11 |
| Distant metastasis | ||
| M0 | 53 | 100 |
| UICC stage | ||
| I | 16 | 30 |
| II | 7 | 13 |
| III | 10 | 19 |
| IV | 20 | 38 |
| HPV status | ||
| Positive | 10 | 19 |
| Negative | 17 | 32 |
| Undefined | 26 | 49 |
| Alcohol consumption | ||
| Yes | 37 | 70 |
| No | 14 | 26 |
| Unknown | 2 | 4 |
| Tobacco consumption | ||
| Yes | 46 | 87 |
| No | 7 | 13 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hofmann, L.; Ludwig, S.; Schuler, P.J.; Hoffmann, T.K.; Brunner, C.; Theodoraki, M.-N. The Potential of CD16 on Plasma-Derived Exosomes as a Liquid Biomarker in Head and Neck Cancer. Int. J. Mol. Sci. 2020, 21, 3739. https://doi.org/10.3390/ijms21113739
Hofmann L, Ludwig S, Schuler PJ, Hoffmann TK, Brunner C, Theodoraki M-N. The Potential of CD16 on Plasma-Derived Exosomes as a Liquid Biomarker in Head and Neck Cancer. International Journal of Molecular Sciences. 2020; 21(11):3739. https://doi.org/10.3390/ijms21113739
Chicago/Turabian StyleHofmann, Linda, Sonja Ludwig, Patrick J. Schuler, Thomas K. Hoffmann, Cornelia Brunner, and Marie-Nicole Theodoraki. 2020. "The Potential of CD16 on Plasma-Derived Exosomes as a Liquid Biomarker in Head and Neck Cancer" International Journal of Molecular Sciences 21, no. 11: 3739. https://doi.org/10.3390/ijms21113739
APA StyleHofmann, L., Ludwig, S., Schuler, P. J., Hoffmann, T. K., Brunner, C., & Theodoraki, M.-N. (2020). The Potential of CD16 on Plasma-Derived Exosomes as a Liquid Biomarker in Head and Neck Cancer. International Journal of Molecular Sciences, 21(11), 3739. https://doi.org/10.3390/ijms21113739






