microRNAs as Promising Biomarkers of Platelet Activity in Antiplatelet Therapy Monitoring
Abstract
1. Introduction
2. Do Activated Platelets Release miRNAs?
2.1. Platelet miRNAs
2.2. Origin and Biological Roles of Platelet miRNAs
2.3. Platelets Release Microvesicles Upon Activation
2.4. Platelets Secrete miRNAs
- The secretion of miRNAs into buffer plasma;
- Alterations of the intracellular platelet miRNome;
- Alterations of miRNA levels in cells after (presumable) uptake of platelet-derived miRNAs;
- Alterations of platelet-related miRNA levels linked to diseases known to be associated with increased platelet activation (such as CVDs);
- Alterations of miRNA levels upon antiplatelet therapy.
3. Are Platelet-Related miRNAs Suitable Biomarkers of CVDs?
Which Other Cell Types of the Cardiovascular System Express Platelet-Related miRNAs?
4. Are Platelet-Related miRNA Levels Affected by Antiplatelet Therapy?
4.1. Plasma miRNAs Responsive to Antiplatelet Agents
4.2. Platelet miRNAs Responsive to Antiplatelet Agents
4.3. Plasma miRNAs Responsive to Antiplatelet Treatment in T2DM
5. Which Technical Challenges Need to Be Addressed Regarding the Quantification of Platelet-Related miRNAs in Plasma?
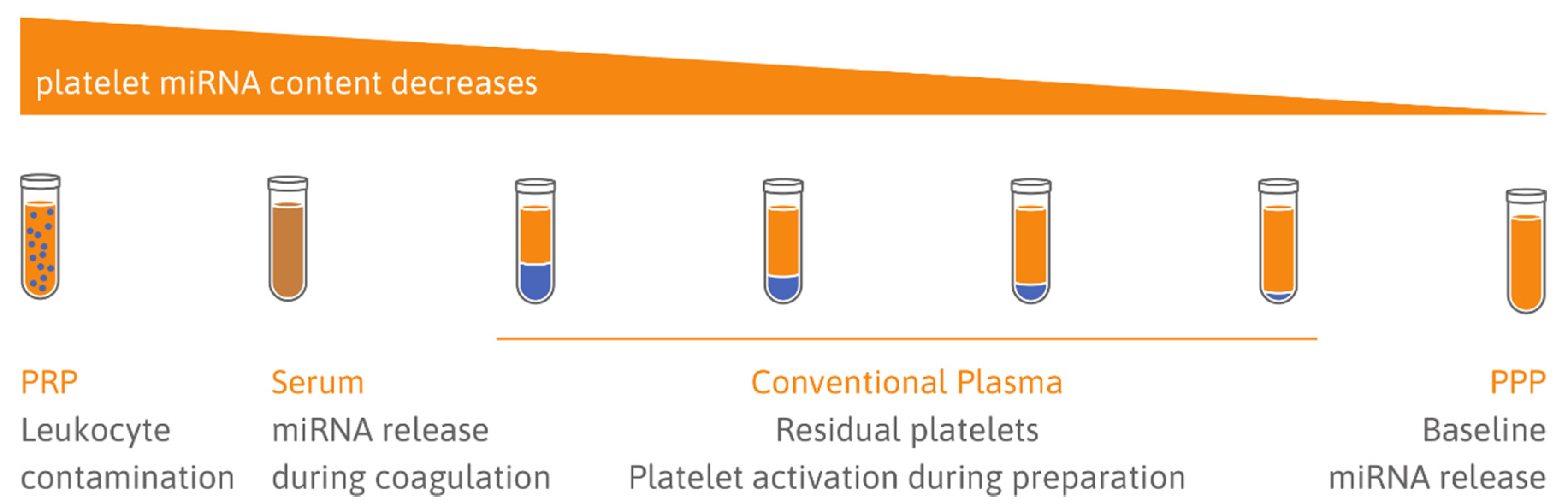
5.1. Choice of Sample Type: Serum vs. Plasma
5.2. Quality of Samples: Cellular Contamination
5.3. Limitations Due to Heparin Use
6. Conclusions
- Do activated platelets release miRNAs?
- Are platelet-related miRNAs suitable biomarkers of CVDs?
- Are platelet-related miRNA levels affected by antiplatelet therapy?
- What technical challenges need to be addressed regarding the quantification of platelet-related miRNAs in plasma?
- Avoid confounding by preanalytical and analytical variation;
- Show a strong correlation between the candidate miRNA and platelet activation;
- Guide treatment decisions to antiplatelet agents.
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AA | arachidonic acid |
| ACM | arrhythmogenic cardiomyopathy |
| ACS | acute coronary syndrome |
| ADP | adenosine diphosphate |
| Ago2 | Argonaute 2 |
| AMI | acute myocardial infarction |
| ASA | acetylsalicylic acid (aspirin) |
| CAD | coronary artery disease |
| CRP-XL | crosslinked collagen-related peptide |
| CTAD | citrate-theophylline-adenosine-dipyridamole |
| CVD | cardiovascular disease |
| DAPT | dual antiplatelet therapy |
| EC | endothelial cell |
| EDTA | ethylenediaminetetraacetic acid |
| EV | extracellular vesicle |
| HSC | hematopoietic stem cell |
| HTPR | high on treatment platelet reactivity |
| HUVEC | human umbilical vein endothelial cell |
| IVD | in vitro diagnostic |
| LD | loading dose |
| LNA | locked nucleic acid |
| LTA | light transmission aggregometry |
| MACE | major adverse cardiac event |
| MD | maintenance dose |
| MEA | multiple electrode aggregometry |
| MI | myocardial infarction |
| miRNA | microRNA |
| mRNA | messenger RNA |
| MRP4 | multidrug resistance protein 4 |
| ncRNA | noncoding RNA |
| PAR2-AP | protease-activated receptor-2 activating peptide |
| PAR4-AP | protease-activated receptor-4 activating peptide |
| PBMC | peripheral blood mononuclear cells |
| PCI | percutaneous coronary intervention |
| PCR | polymerase chain reaction |
| PE | pulmonary embolism |
| PFT | platelet function test |
| PMV | platelet microvesicle |
| POC | point-of-care |
| PPP | platelet-poor plasma |
| pre-miRNA | precursor miRNA |
| PRP | platelet-rich plasma |
| PV | polycythemia vera |
| RT-qPCR | real-time polymerase chain reaction |
| SMC | smooth muscle cell |
| T2DM | type 2 diabetes mellitus |
| TEG | thromboelastography |
| TRAP | thrombin receptor activating peptide |
| UTR | untranslated region |
| VASP | vasodilator-stimulated phosphoprotein phosphorylation |
References
- Gurbel, P.A.; Bliden, K.P.; Hayes, K.M.; Tantry, U. Platelet activation in myocardial ischemic syndromes. Expert Rev. Cardiovasc. Ther. 2004, 2, 535–545. [Google Scholar] [CrossRef] [PubMed]
- Gurbel, P.A.; Jeong, Y.-H.; Navarese, E.P.; Tantry, U.S. Platelet-Mediated Thrombosis: From Bench to Bedside. Circ. Res. 2016, 118, 1380–1391. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Global Health Estimates 2016: Deaths by Cause, Age, Sex, by Country and by Region. 2000–2016; World Health Organization: Geneva, Switzerland, 2018. [Google Scholar]
- Wendelboe, A.M.; Raskob, G.E. Global Burden of Thrombosis: Epidemiologic Aspects. Circ. Res. 2016, 118, 1340–1347. [Google Scholar] [CrossRef] [PubMed]
- Landry, P.; Plante, I.; Ouellet, D.L.; Perron, M.P.; Rousseau, G.; Provost, P. Existence of a microRNA pathway in anucleate platelets. Nat. Struct. Mol. Biol. 2009, 16, 961–966. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1973, 75, 843–854. [Google Scholar] [CrossRef]
- Reinhart, B.J.; Slack, F.J.; Basson, M.; Pasquinelli, A.E.; Bettinger, J.C.; Rougvie, A.E.; Horvitz, H.R.; Ruvkun, G. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 2000, 403, 901–906. [Google Scholar] [CrossRef]
- Baek, D.; Villén, J.; Shin, C.; Camargo, F.D.; Gygi, S.P.; Bartel, D.P. The impact of microRNAs on protein output. Nature 2008, 455, 64–71. [Google Scholar] [CrossRef]
- Hendrickson, D.G.; Hogan, D.J.; McCullough, H.L.; Myers, J.W.; Herschlag, D.; Ferrell, J.E.; Brown, P.O. Concordant Regulation of Translation and mRNA Abundance for Hundreds of Targets of a Human microRNA. PLoS Biol. 2009, 7, e1000238. [Google Scholar] [CrossRef]
- Guo, H.; Ingolia, N.T.; Weissman, J.S.; Bartel, D.P. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 2010, 466, 835–840. [Google Scholar] [CrossRef]
- Eichhorn, S.W.; Guo, H.; McGeary, S.E.; Rodriguez-Mias, R.A.; Shin, C.; Baek, D.; Hsu, S.; Ghoshal, K.; Villén, J.; Bartel, D.P. mRNA Destabilization Is the Dominant Effect of Mammalian MicroRNAs by the Time Substantial Repression Ensues. Mol. Cell 2014, 56, 104–115. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef] [PubMed]
- Nagalla, S.; Shaw, C.; Kong, X.; Kondkar, A.A.; Edelstein, L.C.; Ma, L.; Chen, J.; McKnight, G.S.; Dong, J.; Bray, P.F. Platelet microRNA-mRNA coexpression profiles correlate with platelet reactivity. Blood 2011, 117, 10. [Google Scholar] [CrossRef]
- Born, G.V. Aggregation of Blood Platelets by Adenosine Diphosphate and its Reversal. Nature 1962, 194, 927–929. [Google Scholar] [CrossRef] [PubMed]
- Born, G.V.; Cross, M.J. The aggregation of blood platelets. J. Physiol. 1963, 168, 178–195. [Google Scholar] [CrossRef]
- Bonello, L.; Tantry, U.S.; Marcucci, R.; Blindt, R.; Angiolillo, D.J.; Becker, R.; Bhatt, D.L.; Cattaneo, M.; Collet, J.P.; Cuisset, T.; et al. Consensus and Future Directions on the Definition of High On-Treatment Platelet Reactivity to Adenosine Diphosphate. J. Am. Coll. Cardiol. 2010, 56, 919–933. [Google Scholar] [CrossRef]
- Benjamin, E.J.; Muntner, P.; Alonso, A.; Bittencourt, M.S.; Callaway, C.W.; Carson, A.P.; Chamberlain, A.M.; Chang, A.R.; Cheng, S.; Das, S.R.; et al. Heart Disease and Stroke Statistics—2019 Update: A Report from the American Heart Association. Circulation 2019, 139, e56–e528. [Google Scholar] [CrossRef]
- Stone, G.W.; Witzenbichler, B.; Weisz, G.; Rinaldi, M.J.; Neumann, F.-J.; Metzger, D.C.; Henry, T.D.; Cox, D.A.; Duffy, P.L.; Mazzaferri, E.; et al. Platelet reactivity and clinical outcomes after coronary artery implantation of drug-eluting stents (ADAPT-DES): A prospective multicentre registry study. Lancet 2013, 382, 614–623. [Google Scholar] [CrossRef]
- Aradi, D.; Kirtane, A.; Bonello, L.; Gurbel, P.A.; Tantry, U.S.; Huber, K.; Freynhofer, M.K.; ten Berg, J.; Janssen, P.; Angiolillo, D.J.; et al. Bleeding and stent thrombosis on P2Y12-inhibitors: Collaborative analysis on the role of platelet reactivity for risk stratification after percutaneous coronary intervention. Eur. Heart J. 2015, 36, 1762–1771. [Google Scholar] [CrossRef]
- Sibbing, D.; Aradi, D.; Jacobshagen, C.; Gross, L.; Trenk, D.; Geisler, T.; Orban, M.; Hadamitzky, M.; Merkely, B.; Kiss, R.G.; et al. Guided de-escalation of antiplatelet treatment in patients with acute coronary syndrome undergoing percutaneous coronary intervention (TROPICAL-ACS): A randomised, open-label, multicentre trial. Lancet 2017, 390, 1747–1757. [Google Scholar] [CrossRef]
- Wiviott, S.D.; Braunwald, E.; McCabe, C.H.; Montalescot, G.; Ruzyllo, W.; Gottlieb, S.; Neumann, F.-J.; Ardissino, D.; De Servi, S.; Murphy, S.A.; et al. Prasugrel versus Clopidogrel in Patients with Acute Coronary Syndromes. N. Engl. J. Med. 2007, 357, 2001–2015. [Google Scholar] [CrossRef] [PubMed]
- Antman, E.M.; Wiviott, S.D.; Murphy, S.A.; Voitk, J.; Hasin, Y.; Widimsky, P.; Chandna, H.; Macias, W.; McCabe, C.H.; Braunwald, E. Early and Late Benefits of Prasugrel in Patients with Acute Coronary Syndromes Undergoing Percutaneous Coronary Intervention. J. Am. Coll. Cardiol. 2008, 51, 2028–2033. [Google Scholar] [CrossRef] [PubMed]
- Wallentin, L.; Becker, R.C.; Budaj, A.; Cannon, C.P.; Emanuelsson, H.; Held, C.; Horrow, J.; Husted, S.; James, S.; Katus, H.; et al. Ticagrelor versus Clopidogrel in Patients with Acute Coronary Syndromes. N. Engl. J. Med. 2009, 361, 1045–1057. [Google Scholar] [CrossRef] [PubMed]
- Becker, R.C.; Bassand, J.P.; Budaj, A.; Wojdyla, D.M.; James, S.K.; Cornel, J.H.; French, J.; Held, C.; Horrow, J.; Husted, S.; et al. Bleeding complications with the P2Y12 receptor antagonists clopidogrel and ticagrelor in the PLATelet inhibition and patient Outcomes (PLATO) trial. Eur. Heart J. 2011, 32, 2933–2944. [Google Scholar] [CrossRef] [PubMed]
- Gurbel, P.A.; Bliden, K.P.; Hiatt, B.L.; O’Connor, C.M. Clopidogrel for Coronary Stenting: Response Variability, Drug Resistance, and the Effect of Pretreatment Platelet Reactivity. Circulation 2003, 107, 2908–2913. [Google Scholar] [CrossRef] [PubMed]
- Tantry, U.S.; Bonello, L.; Aradi, D.; Price, M.J.; Jeong, Y.-H.; Angiolillo, D.J.; Stone, G.W.; Curzen, N.; Geisler, T.; ten Berg, J.; et al. Consensus and Update on the Definition of On-Treatment Platelet Reactivity to Adenosine Diphosphate Associated With Ischemia and Bleeding. J. Am. Coll. Cardiol. 2013, 62, 2261–2273. [Google Scholar] [CrossRef] [PubMed]
- Bruchova, H.; Merkerova, M.; Prchal, J.T. Aberrant expression of microRNA in polycythemia vera. Haematologica 2008, 93, 1009–1016. [Google Scholar] [CrossRef][Green Version]
- Merkerova, M.; Belickova, M.; Bruchova, H. Differential expression of microRNAs in hematopoietic cell lineages. Eur. J. Haematol. 2008, 81, 304–310. [Google Scholar] [CrossRef]
- Fink, L.; Hölschermann, H.; Kwapiszewska, G.; Muyal, J.P.; Lengemann, B.; Bohle, R.M.; Santoso, S. Characterization of platelet-specific mRNA by real-time PCR after laser-assisted microdissection. Thromb. Haemost. 2003, 90, 749–756. [Google Scholar] [CrossRef]
- Osman, A.; Fälker, K. Characterization of human platelet microRNA by quantitative PCR coupled with an annotation network for predicted target genes. Platelets 2011, 22, 433–441. [Google Scholar] [CrossRef]
- Wang, K.; Yuan, Y.; Cho, J.-H.; McClarty, S.; Baxter, D.; Galas, D.J. Comparing the MicroRNA Spectrum between Serum and Plasma. PLoS ONE 2012, 7, e41561. [Google Scholar] [CrossRef] [PubMed]
- Bray, P.F.; McKenzie, S.E.; Edelstein, L.C.; Nagalla, S.; Delgrosso, K.; Ertel, A.; Kupper, J.; Jing, Y.; Londin, E.; Loher, P.; et al. The complex transcriptional landscape of the anucleate human platelet. BMC Genom. 2013, 14, 1. [Google Scholar] [CrossRef]
- Pan, Y.; Liang, H.; Liu, H.; Li, D.; Chen, X.; Li, L.; Zhang, C.-Y.; Zen, K. Platelet-secreted microRNA-223 promotes endothelial cell apoptosis induced by advanced glycation end products via targeting the insulin-like growth factor 1 receptor. J. Immunol. 2014, 192, 437–446. [Google Scholar] [CrossRef]
- Simon, L.M.; Edelstein, L.C.; Nagalla, S.; Woodley, A.B.; Chen, E.S.; Kong, X.; Ma, L.; Fortina, P.; Kunapuli, S.; Holinstat, M.; et al. Human platelet microRNA-mRNA networks associated with age and gender revealed by integrated plateletomics. Blood 2014, 123, e37–e45. [Google Scholar] [CrossRef]
- Ambrose, A.R.; Alsahli, M.A.; Kurmani, S.A.; Goodall, A.H. Comparison of the release of microRNAs and extracellular vesicles from platelets in response to different agonists. Platelets 2018, 29, 446–454. [Google Scholar] [CrossRef]
- Plé, H.; Landry, P.; Benham, A.; Coarfa, C.; Gunaratne, P.H.; Provost, P. The Repertoire and Features of Human Platelet microRNAs. PLoS ONE 2012, 7, e50746. [Google Scholar] [CrossRef] [PubMed]
- Teruel-Montoya, R.; Kong, X.; Abraham, S.; Ma, L.; Kunapuli, S.P.; Holinstat, M.; Shaw, C.A.; McKenzie, S.E.; Edelstein, L.C.; Bray, P.F. MicroRNA expression differences in human hematopoietic cell lineages enable regulated transgene expression. PLoS ONE 2014, 9, e102259. [Google Scholar] [CrossRef] [PubMed]
- Clancy, L.; Beaulieu, L.M.; Tanriverdi, K.; Freedman, J.E. The role of RNA uptake in platelet heterogeneity. Thromb. Haemost. 2017, 117, 948–961. [Google Scholar] [CrossRef] [PubMed]
- Edelstein, L.C.; Bray, P.F. MicroRNAs in platelet production and activation. Blood 2011, 117, 9. [Google Scholar] [CrossRef]
- Edelstein, L.C.; McKenzie, S.E.; Shaw, C.; Holinstat, M.A.; Kunapuli, S.P.; Bray, P.F. MicroRNAs in platelet production and activation. J. Thromb. Haemost. 2013, 11, 340–350. [Google Scholar] [CrossRef]
- Rowley, J.W.; Oler, A.J.; Tolley, N.D.; Hunter, B.N.; Low, E.N.; Nix, D.A.; Yost, C.C.; Zimmerman, G.A.; Weyrich, A.S. Genome-wide RNA-seq analysis of human and mouse platelet transcriptomes. Blood 2011, 118, e101–e111. [Google Scholar] [CrossRef] [PubMed]
- Cecchetti, L.; Tolley, N.D.; Michetti, N.; Bury, L.; Weyrich, A.S.; Gresele, P. Megakaryocytes differentially sort mRNAs for matrix metalloproteinases and their inhibitors into platelets: A mechanism for regulating synthetic events. Blood 2011, 118, 1903–1911. [Google Scholar] [CrossRef]
- Zucker-Franklin, D. Endocytosis by human platelets: Metabolic and freeze-fracture studies. J. Cell Biol. 1981, 91, 706–715. [Google Scholar] [CrossRef]
- Risitano, A.; Beaulieu, L.M.; Vitseva, O.; Freedman, J.E. Platelets and platelet-like particles mediate intercellular RNA transfer. Blood 2012, 119, 6288–6295. [Google Scholar] [CrossRef]
- Gidlöf, O.; van der Brug, M.; Öhman, J.; Gilje, P.; Olde, B.; Wahlestedt, C.; Erlinge, D. Platelets activated during myocardial infarction release functional miRNA, which can be taken up by endothelial cells and regulate ICAM1 expression. Blood 2013, 121, 3908–3917. [Google Scholar] [CrossRef] [PubMed]
- Laffont, B.; Corduan, A.; Rousseau, M.; Duchez, A.-C.; Lee, C.H.C.; Boilard, E.; Provost, P. Platelet microparticles reprogram macrophage gene expression and function. Thromb. Haemost. 2016, 115, 311–323. [Google Scholar] [CrossRef] [PubMed]
- Leierseder, S.; Petzold, T.; Zhang, L.; Loyer, X.; Massberg, S.; Engelhardt, S. MiR-223 is dispensable for platelet production and function in mice. Thromb. Haemost. 2013, 110, 1207–1214. [Google Scholar] [CrossRef] [PubMed]
- Elgheznawy, A.; Shi, L.; Hu, J.; Wittig, I.; Laban, H.; Pircher, J.; Mann, A.; Provost, P.; Randriamboavonjy, V.; Fleming, I. Dicer Cleavage by Calpain Determines Platelet microRNA Levels and Function in Diabetes. Circ. Res. 2015, 117, 157–165. [Google Scholar] [CrossRef]
- Kondkar, A.A.; Bray, M.S.; Leal, S.M.; Nagalla, S.; Liu, D.J.; Jin, Y.; Dong, J.F.; Ren, Q.; Whiteheart, S.W.; Shaw, C.; et al. VAMP8/endobrevin is overexpressed in hyperreactive human platelets: Suggested role for platelet microRNA. J. Thromb. Haemost. 2010, 8, 369–378. [Google Scholar] [CrossRef]
- Barwari, T.; Eminaga, S.; Mayr, U.; Lu, R.; Armstrong, P.C.; Chan, M.V.; Sahraei, M.; Fernández-Fuertes, M.; Moreau, T.; Barallobre-Barreiro, J.; et al. Inhibition of profibrotic microRNA-21 affects platelets and their releasate. JCI Insight 2018, 3. [Google Scholar] [CrossRef]
- Kaudewitz, D.; Skroblin, P.; Bender, L.H.; Barwari, T.; Willeit, P.; Pechlaner, R.; Sunderland, N.P.; Willeit, K.; Morton, A.C.; Armstrong, P.C.; et al. Association of MicroRNAs and YRNAs With Platelet Function. Circ. Res. 2016, 118, 420–432. [Google Scholar] [CrossRef]
- Cominetti, M.R.; Martin, A.C.B.M.; Ribeiro, J.U.; Djaafri, I.; Fauvel-Lafève, F.; Crépin, M.; Selistre-de-Araujo, H.S. Inhibition of platelets and tumor cell adhesion by the disintegrin domain of human ADAM9 to collagen I under dynamic flow conditions. Biochimie 2009, 91, 1045–1052. [Google Scholar] [CrossRef]
- Garcia, A.; Dunoyer-Geindre, S.; Zapilko, V.; Nolli, S.; Reny, J.-L.; Fontana, P. Functional Validation of microRNA-126-3p as a Platelet Reactivity Regulator Using Human Haematopoietic Stem Cells. Thromb. Haemost. 2019, 119, 254–263. [Google Scholar] [CrossRef] [PubMed]
- Zaldivia, M.T.K.; McFadyen, J.D.; Lim, B.; Wang, X.; Peter, K. Platelet-Derived Microvesicles in Cardiovascular Diseases. Front. Cardiovasc. Med. 2017, 4. [Google Scholar] [CrossRef] [PubMed]
- Wolf, P. The nature and significance of platelet products in human plasma. Br. J. Haematol. 1967, 13, 269–288. [Google Scholar] [CrossRef] [PubMed]
- Warren, B.A.; Vales, O. The release of vesicles from platelets following adhesion to vessel walls in vitro. Br. J. Exp. Pathol. 1972, 53, 206–215. [Google Scholar] [PubMed]
- Sandberg, H.; Bode, A.P.; Dombrose, F.A.; Hoechli, M.; Lentz, B.R. Expression of coagulant activity in human platelets: Release of membranous vesicles providing platelet factor 1 and platelet factor 3. Thromb. Res. 1985, 39, 63–79. [Google Scholar] [CrossRef]
- Sims, P.J.; Wiedmer, T.; Esmon, C.T.; Weiss, H.J.; Shattil, S.J. Assembly of the platelet prothrombinase complex is linked to vesiculation of the platelet plasma membrane. Studies in Scott syndrome: An isolated defect in platelet procoagulant activity. J. Biol. Chem. 1999, 264, 17049–17057. [Google Scholar]
- Wiedmer, T.; Shattil, S.J.; Cunningham, M.; Sims, P.J. Role of calcium and calpain in complement-induced vesiculation of the platelet plasma membrane and in the exposure of the platelet factor Va receptor. Biochemistry 1990, 29, 623–632. [Google Scholar] [CrossRef]
- Kahner, B.N.; Dorsam, R.T.; Kunapuli, S.P. Role of P2Y receptor subtypes in platelet-derived microparticle generation. Front. Biosci. 2008, 13, 433–439. [Google Scholar] [CrossRef]
- Italiano, J.E.; Mairuhu, A.T.; Flaumenhaft, R. Clinical relevance of microparticles from platelets and megakaryocytes. Curr. Opin. Hematol. 2010, 17, 578–584. [Google Scholar] [CrossRef]
- Aatonen, M.T.; Ohman, T.; Nyman, T.A.; Laitinen, S.; Grönholm, M.; Siljander, P.R.-M. Isolation and characterization of platelet-derived extracellular vesicles. J. Extracell. Vesicles 2014, 3. [Google Scholar] [CrossRef]
- Brisson, A.R.; Tan, S.; Linares, R.; Gounou, C.; Arraud, N. Extracellular vesicles from activated platelets: A semiquantitative cryo-electron microscopy and immuno-gold labeling study. Platelets 2017, 28, 263–271. [Google Scholar] [CrossRef]
- Heijnen, H.F.; Schiel, A.E.; Fijnheer, R.; Geuze, H.J.; Sixma, J.J. Activated platelets release two types of membrane vesicles: Microvesicles by surface shedding and exosomes derived from exocytosis of multivesicular bodies and alpha-granules. Blood 1999, 94, 3791–3799. [Google Scholar] [CrossRef] [PubMed]
- Van der Pol, E.; Böing, A.N.; Gool, E.L.; Nieuwland, R. Recent developments in the nomenclature, presence, isolation, detection and clinical impact of extracellular vesicles. J. Thromb. Haemost. 2016, 14, 48–56. [Google Scholar] [CrossRef] [PubMed]
- Cauwenberghs, S.; Feijge, M.A.H.; Harper, A.G.S.; Sage, S.O.; Curvers, J.; Heemskerk, J.W.M. Shedding of procoagulant microparticles from unstimulated platelets by integrin-mediated destabilization of actin cytoskeleton. FEBS Lett. 2006, 580, 5313–5320. [Google Scholar] [CrossRef] [PubMed]
- Flaumenhaft, R. Formation and fate of platelet microparticles. Blood Cells Mol. Dis. 2006, 36, 182–187. [Google Scholar] [CrossRef] [PubMed]
- Flaumenhaft, R.; Dilks, J.R.; Richardson, J.; Alden, E.; Patel-Hett, S.R.; Battinelli, E.; Klement, G.L.; Sola-Visner, M.; Italiano, J.E. Megakaryocyte-derived microparticles: Direct visualization and distinction from platelet-derived microparticles. Blood 2009, 113, 1112–1121. [Google Scholar] [CrossRef]
- Hunter, M.P.; Ismail, N.; Zhang, X.; Aguda, B.D.; Lee, E.J.; Yu, L.; Xiao, T.; Schafer, J.; Lee, M.-L.T.; Schmittgen, T.D.; et al. Detection of microRNA expression in human peripheral blood microvesicles. PLoS ONE 2008, 3, e3694. [Google Scholar] [CrossRef]
- Willeit, P.; Zampetaki, A.; Dudek, K.; Kaudewitz, D.; King, A.; Kirkby, N.S.; Crosby-Nwaobi, R.; Prokopi, M.; Drozdov, I.; Langley, S.R.; et al. Circulating MicroRNAs as Novel Biomarkers for Platelet Activation. Circ. Res. 2013, 112, 595–600. [Google Scholar] [CrossRef]
- Arroyo, J.D.; Chevillet, J.R.; Kroh, E.M.; Ruf, I.K.; Pritchard, C.C.; Gibson, D.F.; Mitchell, P.S.; Bennett, C.F.; Pogosova-Agadjanyan, E.L.; Stirewalt, D.L.; et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. USA 2011, 108, 5003–5008. [Google Scholar] [CrossRef] [PubMed]
- Vickers, K.C.; Palmisano, B.T.; Shoucri, B.M.; Shamburek, R.D.; Remaley, A.T. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat. Cell Biol. 2011, 13, 423–433. [Google Scholar] [CrossRef] [PubMed]
- Wagner, J.; Riwanto, M.; Besler, C.; Knau, A.; Fichtlscherer, S.; Röxe, T.; Zeiher, A.M.; Landmesser, U.; Dimmeler, S. Characterization of levels and cellular transfer of circulating lipoprotein-bound microRNAs. Arterioscler. Thromb. Vasc. Biol. 2013, 33, 1392–1400. [Google Scholar] [CrossRef] [PubMed]
- Diehl, P.; Fricke, A.; Sander, L.; Stamm, J.; Bassler, N.; Htun, N.; Ziemann, M.; Helbing, T.; El-Osta, A.; Jowett, J.B.M.; et al. Microparticles: Major transport vehicles for distinct microRNAs in circulation. Cardiovasc. Res. 2012, 93, 633–644. [Google Scholar] [CrossRef] [PubMed]
- Turchinovich, A.; Weiz, L.; Langheinz, A.; Burwinkel, B. Characterization of extracellular circulating microRNA. Nucleic Acids Res. 2011, 39, 7223–7233. [Google Scholar] [CrossRef] [PubMed]
- Laffont, B.; Corduan, A.; Plé, H.; Duchez, A.-C.; Cloutier, N.; Boilard, E.; Provost, P. Activated platelets can deliver mRNA regulatory Ago2•microRNA complexes to endothelial cells via microparticles. Blood 2013, 122, 253–261. [Google Scholar] [CrossRef] [PubMed]
- Coppinger, J.A.; Cagney, G.; Toomey, S.; Kislinger, T.; Belton, O.; McRedmond, J.P.; Cahill, D.J.; Emili, A.; Fitzgerald, D.J.; Maguire, P.B. Characterization of the proteins released from activated platelets leads to localization of novel platelet proteins in human atherosclerotic lesions. Blood 2004, 103, 2096–2104. [Google Scholar] [CrossRef]
- Zampetaki, A.; Willeit, P.; Tilling, L.; Drozdov, I.; Prokopi, M.; Renard, J.-M.; Mayr, A.; Weger, S.; Schett, G.; Shah, A.; et al. Prospective Study on Circulating MicroRNAs and Risk of Myocardial Infarction. J. Am. Coll. Cardiol. 2012, 60, 290–299. [Google Scholar] [CrossRef]
- Shi, R.; Ge, L.; Zhou, X.; Ji, W.-J.; Lu, R.-Y.; Zhang, Y.-Y.; Zeng, S.; Liu, X.; Zhao, J.-H.; Zhang, W.-C.; et al. Decreased platelet miR-223 expression is associated with high on-clopidogrel platelet reactivity. Thromb. Res. 2013, 131, 508–513. [Google Scholar] [CrossRef]
- De Boer, H.C.; van Solingen, C.; Prins, J.; Duijs, J.M.G.J.; Huisman, M.V.; Rabelink, T.J.; van Zonneveld, A.J. Aspirin treatment hampers the use of plasma microRNA-126 as a biomarker for the progression of vascular disease. Eur. Heart J. 2013, 34, 3451–3457. [Google Scholar] [CrossRef]
- Zhang, Y.-Y.; Zhou, X.; Ji, W.-J.; Shi, R.; Lu, R.-Y.; Li, J.-L.; Yang, G.-H.; Luo, T.; Zhang, J.-Q.; Zhao, J.-H.; et al. Decreased circulating microRNA-223 level predicts high on-treatment platelet reactivity in patients with troponin-negative non-ST elevation acute coronary syndrome. J. Thromb. Thrombolysis 2014, 38, 65–72. [Google Scholar] [CrossRef] [PubMed]
- Cimmino, G.; Tarallo, R.; Nassa, G.; Filippo, M.R.D.; Giurato, G.; Ravo, M.; Rizzo, F.; Conte, S.; Pellegrino, G.; Cirillo, P.; et al. Activating stimuli induce platelet microRNA modulation and proteome reorganisation. Thromb. Haemost. 2015, 114, 96–108. [Google Scholar] [CrossRef]
- Chyrchel, B.; Totoń-Żurańska, J.; Kruszelnicka, O.; Chyrchel, M.; Mielecki, W.; Kołton-Wróż, M.; Wołkow, P.; Surdacki, A. Association of plasma miR-223 and platelet reactivity in patients with coronary artery disease on dual antiplatelet therapy: A preliminary report. Platelets 2015, 26, 593–597. [Google Scholar] [CrossRef] [PubMed]
- Carino, A.; De Rosa, S.; Sorrentino, S.; Polimeni, A.; Sabatino, J.; Caiazzo, G.; Torella, D.; Spaccarotella, C.; Mongiardo, A.; Strangio, A.; et al. Modulation of Circulating MicroRNAs Levels during the Switch from Clopidogrel to Ticagrelor. Biomed. Res. Int. 2016, 2016, 3968206. [Google Scholar] [CrossRef] [PubMed]
- Peng, L.; Liu, J.; Qin, L.; Liu, J.; Xi, S.; Lu, C.; Yin, T. Interaction between platelet-derived microRNAs and CYP2C19*2 genotype on clopidogrel antiplatelet responsiveness in patients with ACS. Thromb. Res. 2017, 157, 97–102. [Google Scholar] [CrossRef]
- Jäger, B.; Stojkovic, S.; Haller, P.M.; Piackova, E.; Kahl, B.S.; Andric, T.; Vargas, K.G.; Wojta, J.; Huber, K. Course of platelet miRNAs after cessation of P2Y12 antagonists. Eur. J. Clin. Investig. 2019, 49, e13149. [Google Scholar] [CrossRef]
- Parker, W.A.E.; Schulte, C.; Barwari, T.; Phoenix, F.; Pearson, S.M.; Mayr, M.; Grant, P.J.; Storey, R.F.; Ajjan, R.A. Aspirin, clopidogrel and prasugrel monotherapy in patients with type 2 diabetes mellitus: A double-blind randomised controlled trial of the effects on thrombotic markers and microRNA levels. Cardiovasc. Diabetol. 2020, 19, 3. [Google Scholar] [CrossRef]
- Liu, J.; Qin, L.; Wang, Z.; Peng, L.; Liu, J.; Wang, X.; Du, R.; Zou, Y.; Wu, Y.; Yin, T. Platelet-derived miRNAs as determinants of the antiplatelet response in clopidogrel-treated patients with ACS. Thromb. Res. 2020, 186, 71–74. [Google Scholar] [CrossRef]
- Stratz, C.; Nührenberg, T.G.; Binder, H.; Valina, C.M.; Trenk, D.; Hochholzer, W.; Neumann, F.; Fiebich, B.L. Micro-array profiling exhibits remarkable intra-individual stability of human platelet micro-RNA. Thromb. Haemost. 2012, 107, 634–641. [Google Scholar] [CrossRef]
- Zhou, S.; Jin, J.; Wang, J.; Zhang, Z.; Freedman, J.H.; Zheng, Y.; Cai, L. miRNAS in cardiovascular diseases: Potential biomarkers, therapeutic targets and challenges. Acta Pharm. Sin. 2018, 39, 1073–1084. [Google Scholar] [CrossRef]
- Hulsmans, M.; Holvoet, P. MicroRNA-containing microvesicles regulating inflammation in association with atherosclerotic disease. Cardiovasc. Res. 2013, 100, 7–18. [Google Scholar] [CrossRef] [PubMed]
- Zampetaki, A.; Kiechl, S.; Drozdov, I.; Willeit, P.; Mayr, U.; Prokopi, M.; Mayr, A.; Weger, S.; Oberhollenzer, F.; Bonora, E.; et al. Plasma microRNA profiling reveals loss of endothelial miR-126 and other microRNAs in type 2 diabetes. Circ. Res. 2010, 107, 810–817. [Google Scholar] [CrossRef] [PubMed]
- Olivieri, F.; Spazzafumo, L.; Bonafè, M.; Recchioni, R.; Prattichizzo, F.; Marcheselli, F.; Micolucci, L.; Mensà, E.; Giuliani, A.; Santini, G.; et al. MiR-21-5p and miR-126a-3p levels in plasma and circulating angiogenic cells: Relationship with type 2 diabetes complications. Oncotarget 2015, 6, 35372. [Google Scholar] [CrossRef] [PubMed]
- Jansen, F.; Yang, X.; Proebsting, S.; Hoelscher, M.; Przybilla, D.; Baumann, K.; Schmitz, T.; Dolf, A.; Endl, E.; Franklin, B.S.; et al. MicroRNA Expression in Circulating Microvesicles Predicts Cardiovascular Events in Patients With Coronary Artery Disease. JAHA 2014, 3, e001249. [Google Scholar] [CrossRef]
- Sygitowicz, G.; Tomaniak, M.; Błaszczyk, O.; Kołtowski, Ł.; Filipiak, K.J.; Sitkiewicz, D. Circulating microribonucleic acids miR-1, miR-21 and miR-208a in patients with symptomatic heart failure: Preliminary results. Arch. Cardiovasc. Dis. 2015, 108, 634–642. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, Y.-J.; Liu, T.; Zhang, H.; Yang, S.-J. Plasma microRNA-21 is a potential diagnostic biomarker of acute myocardial infarction. Eur. Rev. Med. Pharm. Sci. 2016, 20, 323–329. [Google Scholar]
- Liu, X.; Dong, Y.; Chen, S.; Zhang, G.; Zhang, M.; Gong, Y.; Li, X. Circulating MicroRNA-146a and MicroRNA-21 Predict Left Ventricular Remodeling after ST-Elevation Myocardial Infarction. Cardiology 2015, 132, 233–241. [Google Scholar] [CrossRef]
- Sommariva, E.; D’Alessandra, Y.; Farina, F.M.; Casella, M.; Cattaneo, F.; Catto, V.; Chiesa, M.; Stadiotti, I.; Brambilla, S.; Dello Russo, A.; et al. MiR-320a as a Potential Novel Circulating Biomarker of Arrhythmogenic CardioMyopathy. Sci. Rep. 2017, 7, 1–10. [Google Scholar] [CrossRef]
- Zhang, H.; Liu, S.; Dong, T.; Yang, J.; Xie, Y.; Wu, Y.; Kang, K.; Hu, S.; Gou, D.; Wei, Y. Profiling of differentially expressed microRNAs in arrhythmogenic right ventricular cardiomyopathy. Sci. Rep. 2016, 6, 1–11. [Google Scholar] [CrossRef]
- Scherrer, N.; Fays, F.; Mueller, B.; Luft, A.; Fluri, F.; Christ-Crain, M.; Devaux, Y.; Katan, M. MicroRNA 150-5p Improves Risk Classification for Mortality within 90 Days after Acute Ischemic Stroke. J. Stroke 2017, 19, 323–332. [Google Scholar] [CrossRef]
- Zhou, X.; Wen, W.; Shan, X.; Qian, J.; Li, H.; Jiang, T.; Wang, W.; Cheng, W.; Wang, F.; Qi, L.; et al. MiR-28-3p as a potential plasma marker in diagnosis of pulmonary embolism. Thromb. Res. 2016, 138, 91–95. [Google Scholar] [CrossRef] [PubMed]
- Schulte, C.; Molz, S.; Appelbaum, S.; Karakas, M.; Ojeda, F.; Lau, D.M.; Hartmann, T.; Lackner, K.J.; Westermann, D.; Schnabel, R.B.; et al. miRNA-197 and miRNA-223 Predict Cardiovascular Death in a Cohort of Patients with Symptomatic Coronary Artery Disease. PLoS ONE 2015, 10, e0145930. [Google Scholar] [CrossRef]
- Ji, R.; Cheng, Y.; Yue, J.; Yang, J.; Liu, X.; Chen, H.; Dean, D.B.; Zhang, C. MicroRNA expression signature and antisense-mediated depletion reveal an essential role of MicroRNA in vascular neointimal lesion formation. Circ. Res. 2007, 100, 1579–1588. [Google Scholar] [CrossRef] [PubMed]
- Kuosmanen, S.M.; Kansanen, E.; Sihvola, V.; Levonen, A.-L. MicroRNA Profiling Reveals Distinct Profiles for Tissue-Derived and Cultured Endothelial Cells. Sci. Rep. 2017, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Juzenas, S.; Venkatesh, G.; Hübenthal, M.; Hoeppner, M.P.; Du, Z.G.; Paulsen, M.; Rosenstiel, P.; Senger, P.; Hofmann-Apitius, M.; Keller, A.; et al. A comprehensive, cell specific microRNA catalogue of human peripheral blood. Nucleic Acids Res. 2017, 45, 9290–9301. [Google Scholar] [CrossRef]
- Thum, T.; Galuppo, P.; Wolf, C.; Fiedler, J.; Kneitz, S.; van Laake, L.W.; Doevendans, P.A.; Mummery, C.L.; Borlak, J.; Haverich, A.; et al. MicroRNAs in the Human Heart: A Clue to Fetal Gene Reprogramming in Heart Failure. Circulation 2007, 116, 258–267. [Google Scholar] [CrossRef]
- Humphreys, D.T.; Hynes, C.J.; Patel, H.R.; Wei, G.H.; Cannon, L.; Fatkin, D.; Suter, C.M.; Clancy, J.L.; Preiss, T. Complexity of Murine Cardiomyocyte miRNA Biogenesis, Sequence Variant Expression and Function. PLoS ONE 2012, 7, e30933. [Google Scholar] [CrossRef]
- Matkovich, S.J.; Van Booven, D.J.; Eschenbacher, W.H.; Dorn, G.W. RISC RNA sequencing for context-specific identification of in vivo miR targets. Circ. Res. 2011, 108, 18–26. [Google Scholar] [CrossRef]
- O’Connell, R.M.; Chaudhuri, A.A.; Rao, D.S.; Gibson, W.S.J.; Balazs, A.B.; Baltimore, D. MicroRNAs enriched in hematopoietic stem cells differentially regulate long-term hematopoietic output. Proc. Natl. Acad. Sci. USA 2010, 107, 14235–14240. [Google Scholar] [CrossRef] [PubMed]
- Wisman, P.P.; Roest, M.; Asselbergs, F.W.; de Groot, P.G.; Moll, F.L.; van der Graaf, Y.; de Borst, G.J. Platelet-reactivity tests identify patients at risk of secondary cardiovascular events: A systematic review and meta-analysis. J. Thromb. Haemost. 2014, 12, 736–747. [Google Scholar] [CrossRef] [PubMed]
- López Farré, A.J.; Tamargo, J.; Mateos-Cáceres, P.J.; Azcona, L.; Macaya, C. Old and New Molecular Mechanisms Associated with Platelet Resistance to Antithrombotics. Pharm. Res. 2010, 27, 2365–2373. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Wang, H.; Chen, Y.-F.; Lin, W.-W.; Wang, C.-L.; Lin, C.-H. High maintenance dose of clopidogrel in patients with high on-treatment platelet reactivity after a percutaneous coronary intervention: A meta-analysis. Coron. Artery Dis. 2015, 26, 386–395. [Google Scholar] [CrossRef] [PubMed]
- Fiolaki, A.; Katsanos, A.H.; Kyritsis, A.P.; Papadaki, S.; Kosmidou, M.; Moschonas, I.C.; Tselepis, A.D.; Giannopoulos, S. High on treatment platelet reactivity to aspirin and clopidogrel in ischemic stroke: A systematic review and meta-analysis. J. Neurol. Sci. 2017, 376, 112–116. [Google Scholar] [CrossRef] [PubMed]
- Matetzky, S.; Shenkman, B.; Guetta, V.; Shechter, M.; Beinart, R.; Goldenberg, I.; Novikov, I.; Pres, H.; Savion, N.; Varon, D.; et al. Clopidogrel Resistance Is Associated With Increased Risk of Recurrent Atherothrombotic Events in Patients With Acute Myocardial Infarction. Circulation 2004, 109, 3171–3175. [Google Scholar] [CrossRef]
- La Rosa, G.; Biasucci, L.M.; Mandolini, C.; Massimi, I.; Copponi, G.; Pulcinelli, F.M.; Crea, F. Platelet miRNA-26b down-regulates multidrug resistance protein 4 in patients on chronic aspirin treatment. J. Cardiovasc. Med. (Hagerstown) 2018, 19, 611–613. [Google Scholar] [CrossRef] [PubMed]
- Weber, M.J. New human and mouse microRNA genes found by homology search: New human and mouse microRNA gene. FEBS J. 2004, 272, 59–73. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, B.M.; Heimberg, A.M.; Moy, V.N.; Sperling, E.A.; Holstein, T.W.; Heber, S.; Peterson, K.J. The deep evolution of metazoan microRNAs. Evol. Dev. 2009, 11, 50–68. [Google Scholar] [CrossRef] [PubMed]
- Braza-Boïls, A.; Barwari, T.; Gutmann, C.; Thomas, M.R.; Judge, H.M.; Joshi, A.; Pechlaner, R.; Shankar-Hari, M.; Ajjan, R.A.; Sabroe, I.; et al. Circulating MicroRNA Levels Indicate Platelet and Leukocyte Activation in Endotoxemia Despite Platelet P2Y12 Inhibition. Int. J. Mol. Sci. 2020, 21, 2897. [Google Scholar] [CrossRef]
- Mussbacher, M.; Schrottmaier, W.C.; Salzmann, M.; Brostjan, C.; Schmid, J.A.; Starlinger, P.; Assinger, A. Optimized plasma preparation is essential to monitor platelet-stored molecules in humans. PLoS ONE 2017, 12, e0188921. [Google Scholar] [CrossRef]
- Glinge, C.; Clauss, S.; Boddum, K.; Jabbari, R.; Jabbari, J.; Risgaard, B.; Tomsits, P.; Hildebrand, B.; Kääb, S.; Wakili, R.; et al. Stability of Circulating Blood-Based MicroRNAs—Pre-Analytic Methodological Considerations. PLoS ONE 2017, 12, e0167969. [Google Scholar] [CrossRef]
- Faraldi, M.; Sansoni, V.; Perego, S.; Gomarasca, M.; Kortas, J.; Ziemann, E.; Banfi, G.; Lombardi, G. Study of the preanalytical variables affecting the measurement of clinically relevant free-circulating microRNAs: Focus on sample matrix, platelet depletion, and storage conditions. Biochem. Med. (Zagreb) 2020, 30, 010703. [Google Scholar] [CrossRef] [PubMed]
- Köberle, V.; Pleli, T.; Schmithals, C.; Augusto Alonso, E.; Haupenthal, J.; Bönig, H.; Peveling-Oberhag, J.; Biondi, R.M.; Zeuzem, S.; Kronenberger, B.; et al. Differential stability of cell-free circulating microRNAs: Implications for their utilization as biomarkers. PLoS ONE 2013, 8, e75184. [Google Scholar] [CrossRef] [PubMed]
- Sunderland, N.; Skroblin, P.; Barwari, T.; Huntley, R.P.; Lu, R.; Joshi, A.; Lovering, R.C.; Mayr, M. MicroRNA Biomarkers and Platelet Reactivity: The Clot Thickens. Circ. Res. 2017, 120, 418–435. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, C.C.; Kroh, E.; Wood, B.; Arroyo, J.D.; Dougherty, K.J.; Miyaji, M.M.; Tait, J.F.; Tewari, M. Blood cell origin of circulating microRNAs: A cautionary note for cancer biomarker studies. Cancer Prev. Res. (Phila) 2012, 5, 492–497. [Google Scholar] [CrossRef] [PubMed]
- Kirschner, M.B.; Kao, S.C.; Edelman, J.J.; Armstrong, N.J.; Vallely, M.P.; van Zandwijk, N.; Reid, G. Haemolysis during Sample Preparation Alters microRNA Content of Plasma. PLoS ONE 2011, 6, e24145. [Google Scholar] [CrossRef] [PubMed]
- McDonald, J.S.; Milosevic, D.; Reddi, H.V.; Grebe, S.K.; Algeciras-Schimnich, A. Analysis of circulating microRNA: Preanalytical and analytical challenges. Clin. Chem. 2011, 57, 833–840. [Google Scholar] [CrossRef]
- Binderup, H.G.; Houlind, K.; Madsen, J.S.; Brasen, C.L. Pre-storage centrifugation conditions have significant impact on measured microRNA levels in biobanked EDTA plasma samples. Biochem. Biophys. Rep. 2016, 7, 195–200. [Google Scholar] [CrossRef]
- Mitchell, A.J.; Gray, W.D.; Hayek, S.S.; Ko, Y.-A.; Thomas, S.; Rooney, K.; Awad, M.; Roback, J.D.; Quyyumi, A.; Searles, C.D. Platelets confound the measurement of extracellular miRNA in archived plasma. Sci. Rep. 2016, 6, 32651. [Google Scholar] [CrossRef]
- Basso, D.; Padoan, A.; Laufer, T.; Aneloni, V.; Moz, S.; Schroers, H.; Pelloso, M.; Saiz, A.; Krapp, M.; Fogar, P.; et al. Relevance of pre-analytical blood management on the emerging cardiovascular protein biomarkers TWEAK and HMGB1 and on miRNA serum and plasma profiling. Clin. Biochem. 2017, 50, 186–193. [Google Scholar] [CrossRef]
- Binderup, H.G.; Madsen, J.S.; Heegaard, N.H.H.; Houlind, K.; Andersen, R.F.; Brasen, C.L. Quantification of microRNA levels in plasma—Impact of preanalytical and analytical conditions. PLoS ONE 2018, 13, e0201069. [Google Scholar] [CrossRef]
- McSHINE, R.L.; Das, P.C.; Sibinga, C.T.S.; Brozovic, B. Differences between the effects of EDTA and citrate anticoagulants on platelet count and mean platelet volume. Clin. Lab. Haematol. 1990, 12, 277–285. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.H.; Yi, H.S.; Kim, Y.; Kroh, E.M.; Chien, J.W.; Eaton, K.D.; Goodman, M.T.; Tait, J.F.; Tewari, M.; Pritchard, C.C. Plasma Processing Conditions Substantially Influence Circulating microRNA Biomarker Levels. PLoS ONE 2013, 8, e64795. [Google Scholar] [CrossRef] [PubMed]
- Kaudewitz, D.; Lee, R.; Willeit, P.; McGregor, R.; Markus, H.S.; Kiechl, S.; Zampetaki, A.; Storey, R.F.; Channon, K.M.; Mayr, M. Impact of intravenous heparin on quantification of circulating microRNAs in patients with coronary artery disease. Thromb. Haemost. 2013, 110, 609–615. [Google Scholar] [CrossRef]
- Schulte, C.; Barwari, T.; Joshi, A.; Theofilatos, K.; Zampetaki, A.; Barallobre-Barreiro, J.; Singh, B.; Sörensen, N.A.; Neumann, J.T.; Zeller, T.; et al. Comparative Analysis of Circulating Noncoding RNAs Versus Protein Biomarkers in the Detection of Myocardial Injury. Circ. Res. 2019, 125, 328–340. [Google Scholar] [CrossRef]
- Morrow David, A.; de Lemos James, A. Benchmarks for the Assessment of Novel Cardiovascular Biomarkers. Circulation 2007, 115, 949–952. [Google Scholar] [CrossRef] [PubMed]
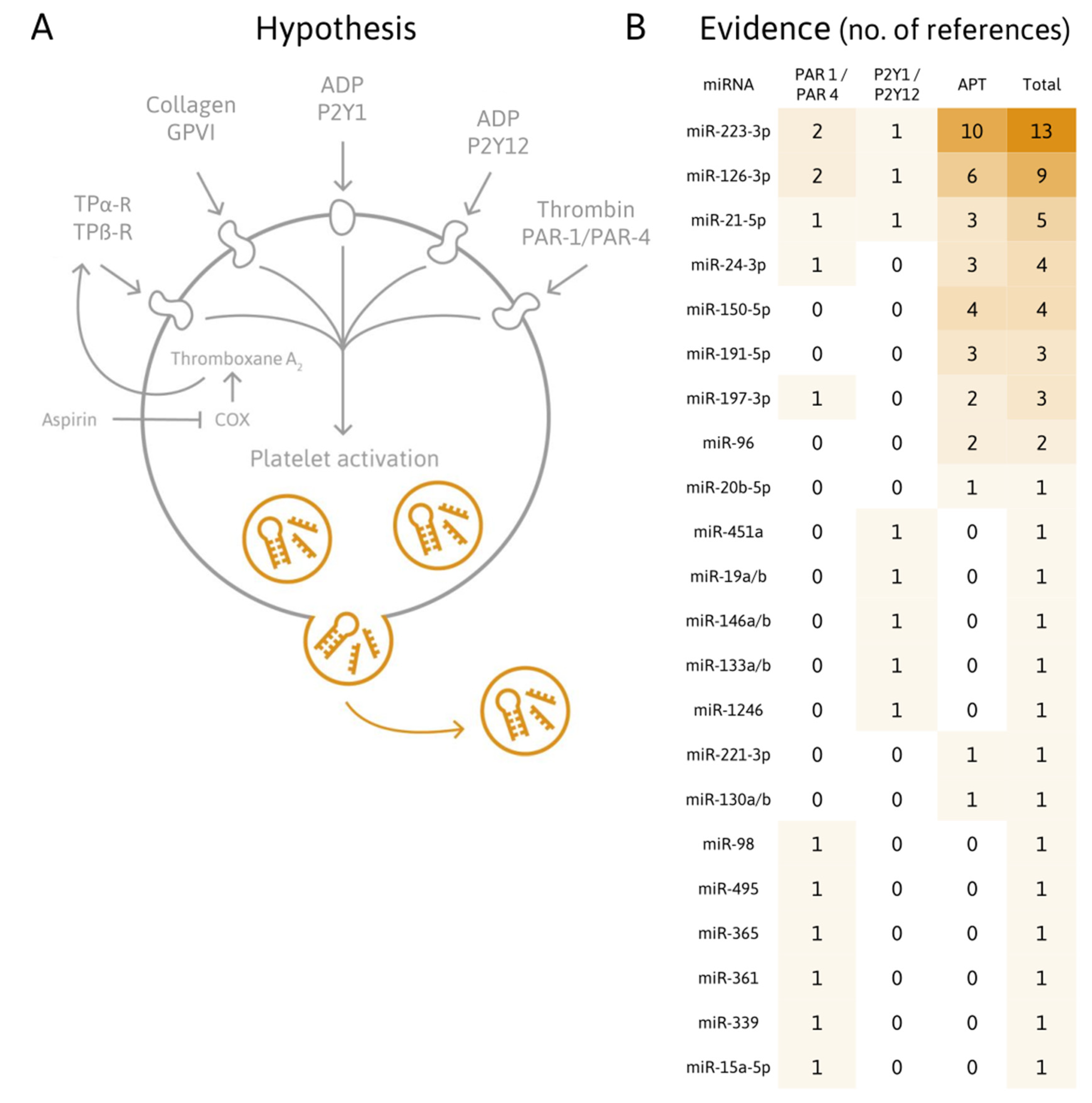
| Evidence | Reference | Year | miRNAs of Interest | Agonist/Condition/Treatment | PFT | Origin of Samples | Method |
|---|---|---|---|---|---|---|---|
| B | [14] | 2011 | 74 miRNAs differentially expressed | Hyperreactive vs. hyporeactive platelets | Maximal aggregation response to ADP and epinephrine | 19 healthy subjects | S |
| B | [31] | 2011 | miR-15a miR-98 miR-339 miR-361 miR-365 miR-495 | Thrombin | P-selectin | 4 stimulated, 6 resting platelet samples from healthy subjects | S |
| A + B + D | [75] | 2012 | miR-1246 miR-451 miR-223 miR-146 miR-133 miR-126 miR-21 miR-19 | ADP and patients with stable CAD vs. patients with ACS | - | Extracted platelets, 5 patients with stable CAD, 5 patients with ACS | T |
| A + B + D | [79] | 2012 | miR-223 miR-197 miR-126 miR-24 miR-21 | Thrombin and healthy subjects: limb ischemia-reperfusion | - | 820 subjects from general population (Bruneck cohort), 11 healthy subjects, extracted platelets and PMVs | S + T |
| A + B D + E | [71] | 2013 | miR-223 miR-197 miR-191 miR-150 miR-126 miR-24 miR-21 miR-20b | Healthy subjects: dose-escalation of ASA combined with prasugrel Patients: ASA at baseline, addition of dipyridamole or clopidogrel | Verify Now, LTA, formation of thromboxane B2 | Platelets, PMVs, serum, PRP, PPP from 3 healthy subjects, serum and PPP from 19 T2DM patients, 9 healthy subjects, 33 patients with symptomatic carotid atherosclerosis | S + T |
| B + E | [80] | 2013 | miR-223 miR-96 | Clopidogrel + ASA | VASP assay, LTA | 33 non-diabetic CVD patients | T |
| A + C | [77] | 2013 | miR-223 | Thrombin and co-incubation of HUVECs with released PMVs | P-selectin | Extracted platelets, HUVEC | T |
| A + D + E | [81] | 2013 | miR-126 | PRP stimulated with AA in presence/absence of aspirin and patients: one period (6 weeks) placebo, one period ASA | P-selectin | 4 healthy subjects, 40 T2DM patients without CVD | T |
| E | [82] | 2014 | miR-223 | DAPT with clopidogrel | VASP assay | 62 ACS patients | T |
| B | [83] | 2015 | ~50 miRNAs differentially expressed | ADP, collagen, TRAP | LTA | 15 healthy subjects | S |
| D | [49] | 2015 | miR-223 | T2DM | Clot retraction & platelet adhesion and spreading assay | 22 patients with T2DM, 22 healthy subjects | T |
| E | [84] | 2015 | miR-223 | DAPT with clopidogrel or prasugrel or ticagrelor | MEA | 21 patients with ACS | T |
| D + E | [52] | 2016 | miR-223 miR-191 miR-126 miR-24 | Healthy subjects: DAPT with prasugrel and patients: ASA only or DAPT with clopidogrel, prasugrel, or ticagrelor | VASP assay, LTA, Verify Now | 669 subjects from general population (Bruneck cohort), 125 ACS patients, additional ACS patients (n = 8/group) and healthy subjects (n = 6) to assess impact of antiplatelet treatment | S |
| C | [47] | 2016 | miR-126 | Thrombin stimulation of platelets, co-incubation of PMVs with primary human macrophages | - | Healthy subjects | S + T |
| E | [85] | 2016 | miR-223 miR-150 miR-126 miR-96 | Switch from DAPT with clopidogrel to ticagrelor “No load”: ASA + ticagrelor without LD“Load”: ticagrelor LD, then ticagrelor MD + ASA | MEA | 16 ACS patients (8 “no load” group, 8 “load” group) | T |
| E | [86] | 2017 | miR-223 miR-221 miR-21 | DAPT with clopidogrel | LTA | 272 subjects included; 21 “high responders”, 18 “low responders” | T |
| A | [36] | 2018 | 46 miRNAs consistently secreted | CRP-XL, PAR1-AP, PAR4-AP, ADP | - | 4 healthy subjects | S |
| E | [87] | 2019 | miR-223 miR-150 miR-126 miR-21 | Cessation of DAPT with clopidogrel or prasugrel or ticagrelor | MEA | 62 CAD patients | T |
| E | [88] | 2020 | miR-223 miR-197 miR-191 miR-24 | ASA + one period (28 days) clopidogrel, one period prasugrel | LTA, P-selectin | 56 T2DM patients | T |
| E | [89] | 2020 | miR-223 miR-150 miR-130 miR-126 | Patients: ASA + clopidogrel | TEG | 214 healthy subjects, 430 ACS patients | T |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Krammer, T.L.; Mayr, M.; Hackl, M. microRNAs as Promising Biomarkers of Platelet Activity in Antiplatelet Therapy Monitoring. Int. J. Mol. Sci. 2020, 21, 3477. https://doi.org/10.3390/ijms21103477
Krammer TL, Mayr M, Hackl M. microRNAs as Promising Biomarkers of Platelet Activity in Antiplatelet Therapy Monitoring. International Journal of Molecular Sciences. 2020; 21(10):3477. https://doi.org/10.3390/ijms21103477
Chicago/Turabian StyleKrammer, Teresa L., Manuel Mayr, and Matthias Hackl. 2020. "microRNAs as Promising Biomarkers of Platelet Activity in Antiplatelet Therapy Monitoring" International Journal of Molecular Sciences 21, no. 10: 3477. https://doi.org/10.3390/ijms21103477
APA StyleKrammer, T. L., Mayr, M., & Hackl, M. (2020). microRNAs as Promising Biomarkers of Platelet Activity in Antiplatelet Therapy Monitoring. International Journal of Molecular Sciences, 21(10), 3477. https://doi.org/10.3390/ijms21103477





