Characterization and Immobilization of a Novel SGNH Family Esterase (LaSGNH1) from Lactobacillus acidophilus NCFM
Abstract
1. Introduction
2. Results and Discussion
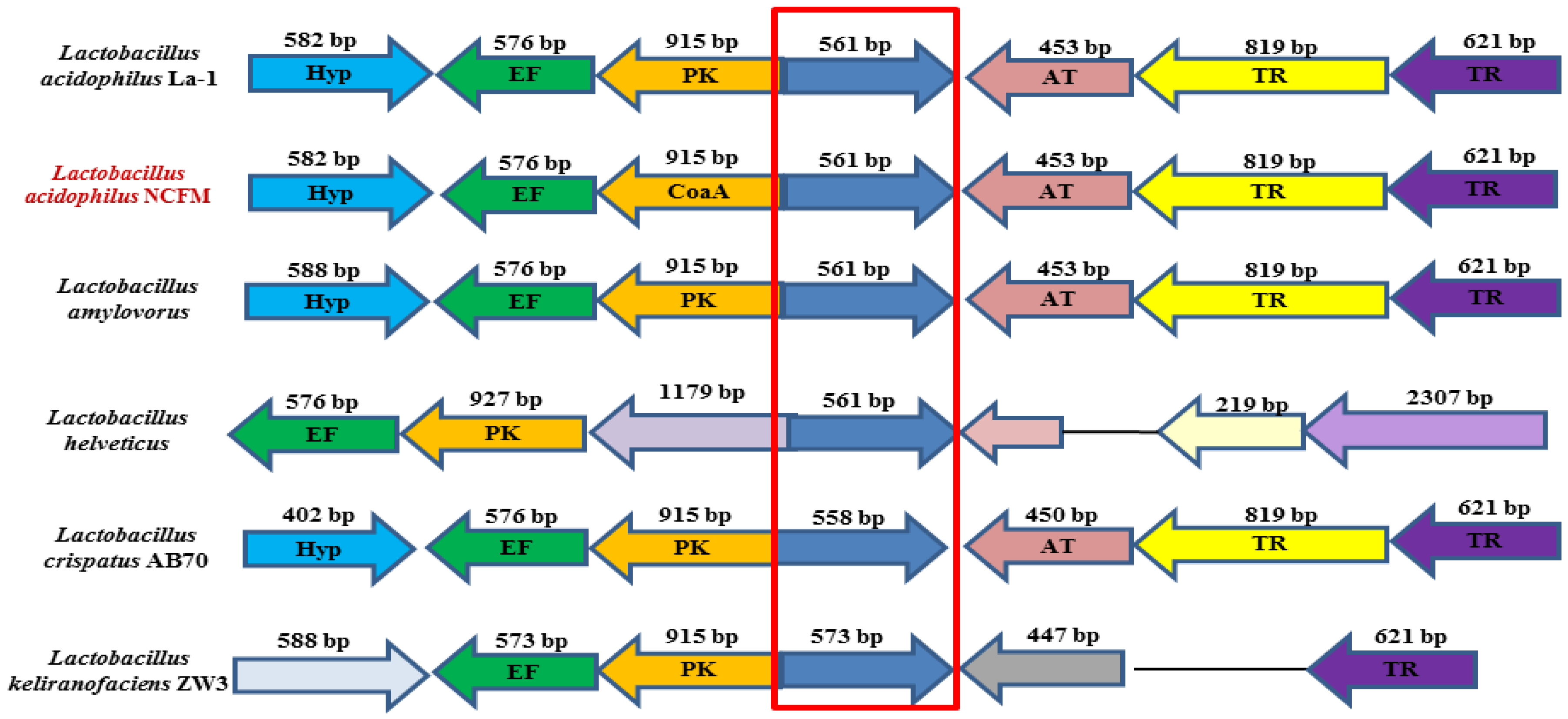
2.1. Bioinformatic Analysis of LaSGNH1
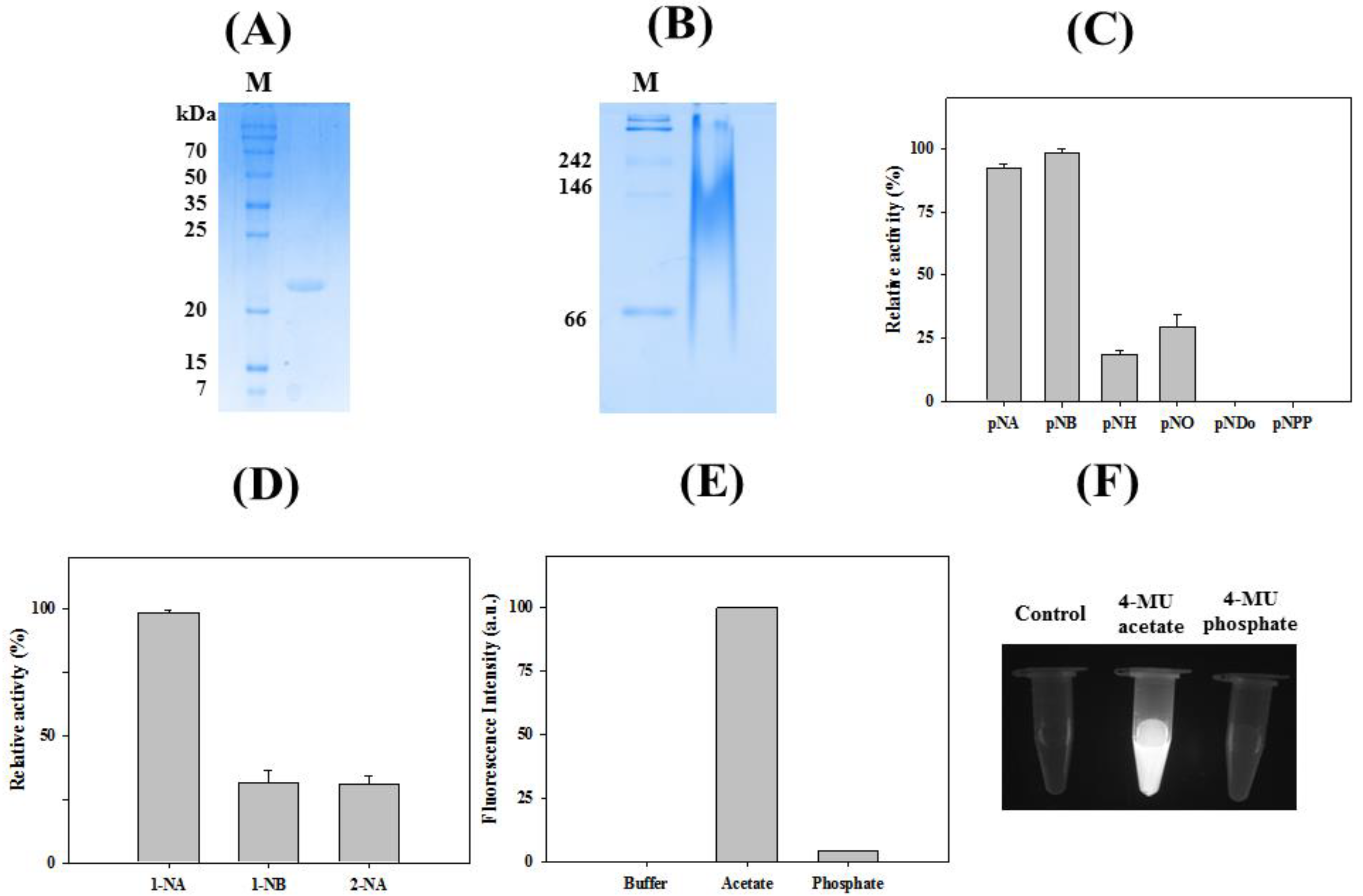
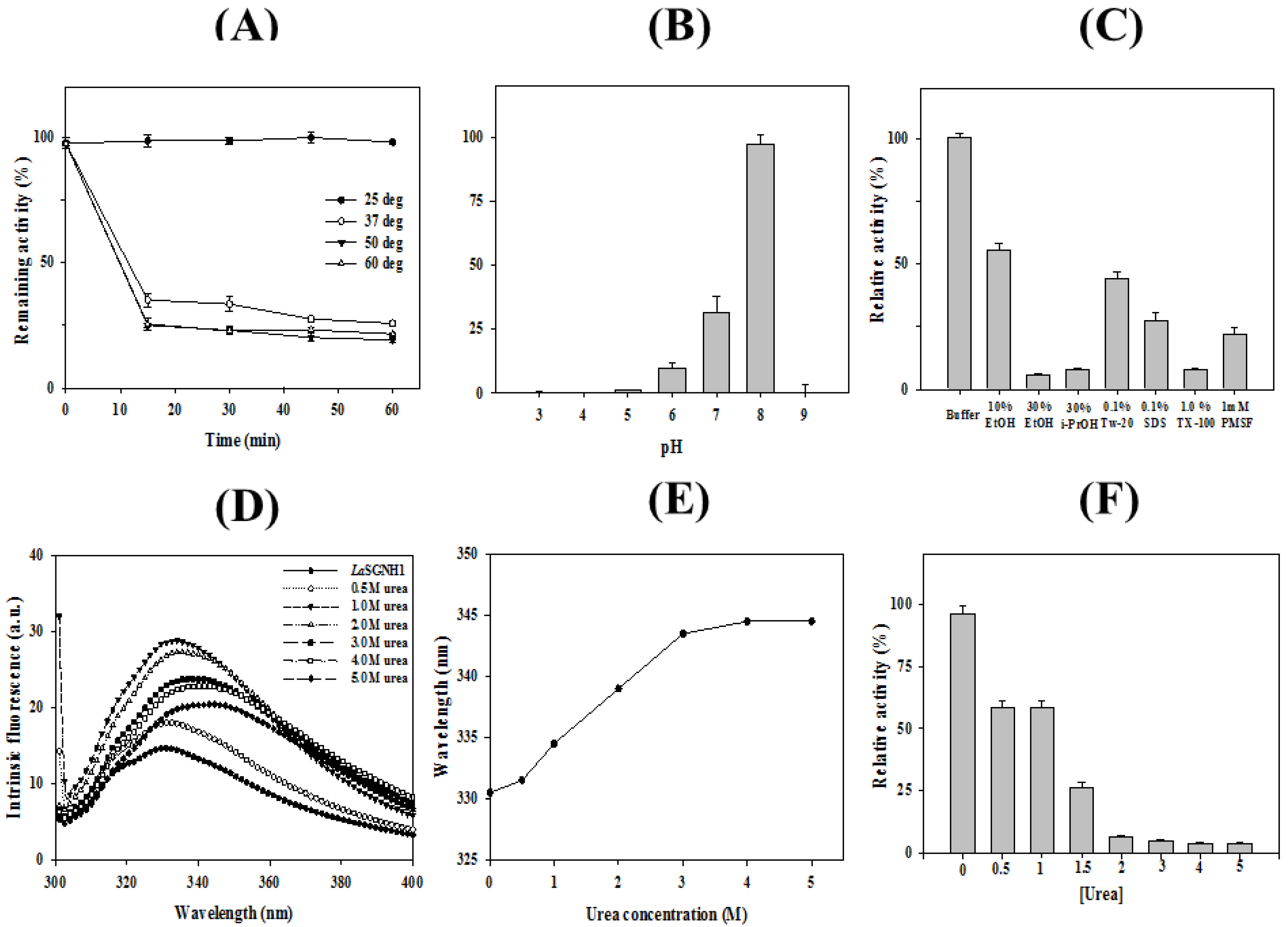
2.2. Characterizations of LaSGNH1
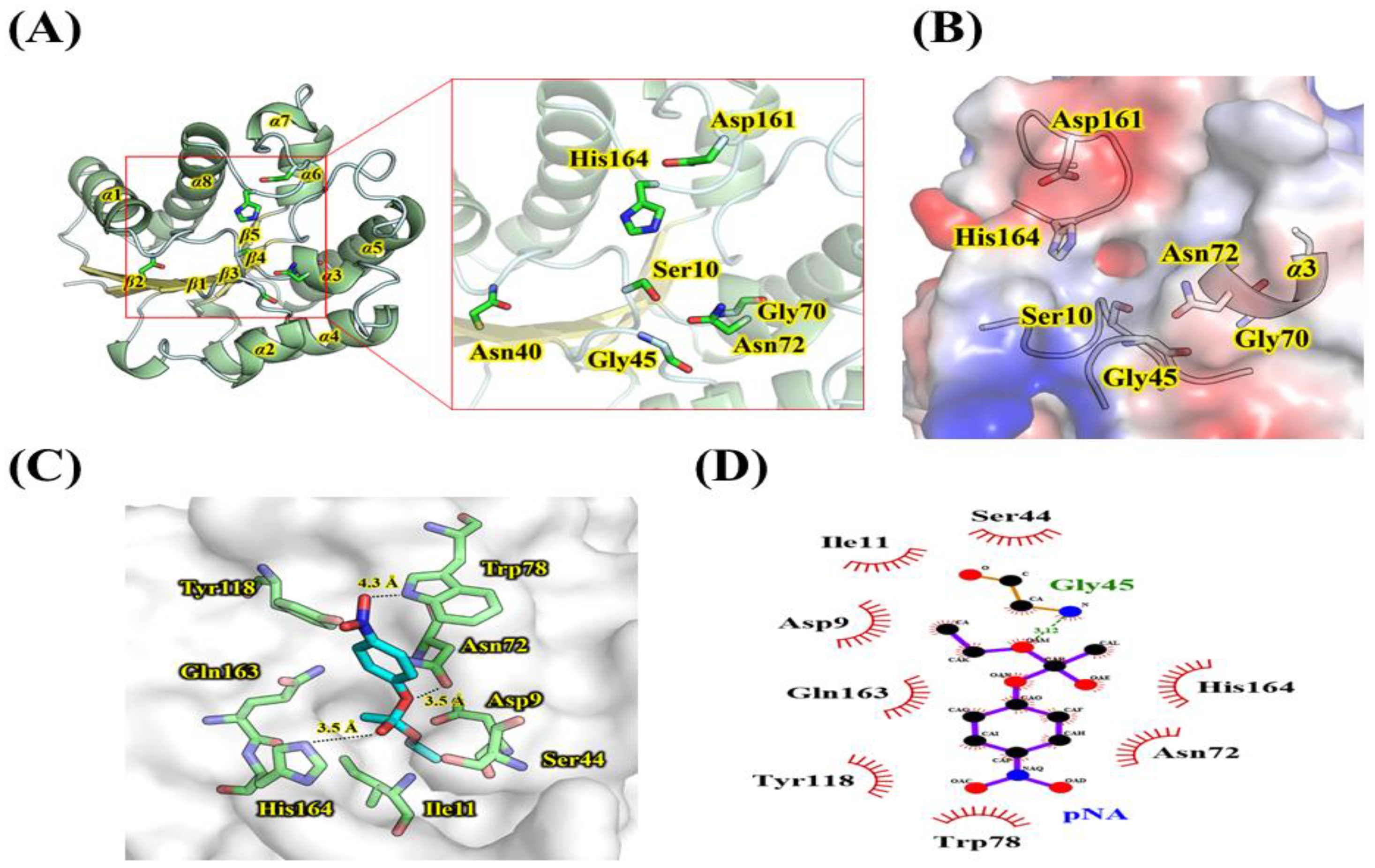
2.3. Homology Modeling and Substrate Analysis of LaSGNH1
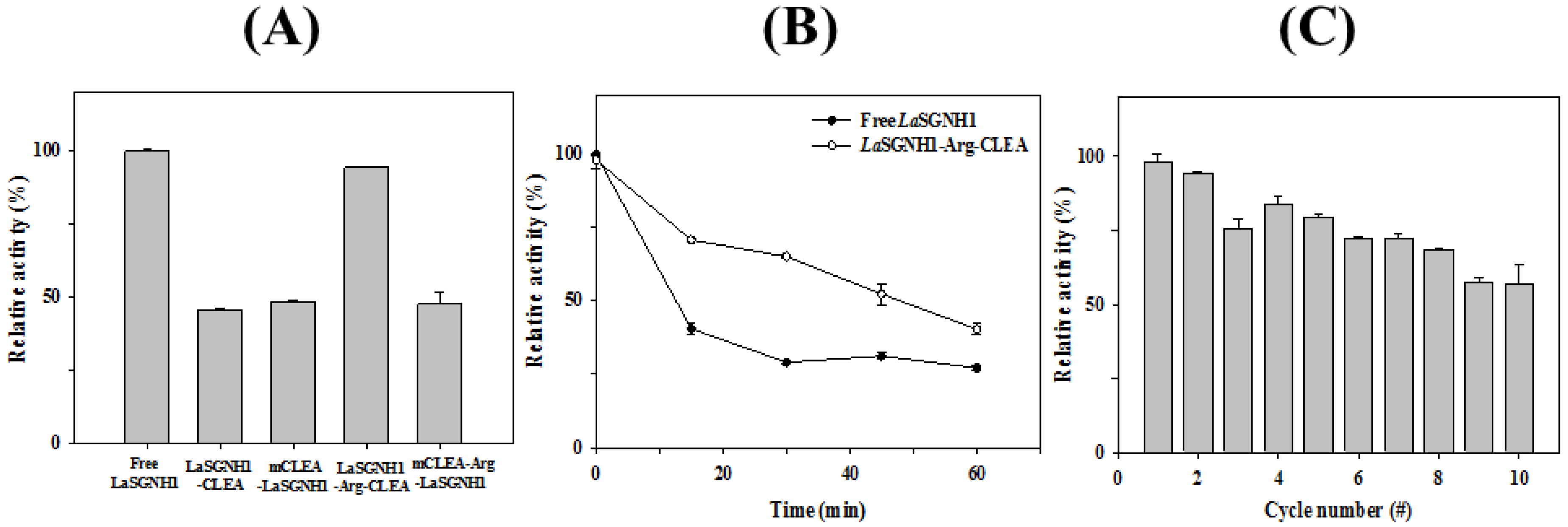
2.4. Immobilization of LaSGNH1
3. Materials and Methods
3.1. Reagents
3.2. Bioinformatic Analysis
3.3. Cloning and Purification
3.4. Biochemical Characterization of LaSGNH1
3.5. Immobilization of LaSGNH1
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Sarmah, N.; Revathi, D.; Sheelu, G.; Yamuna Rani, K.; Sridhar, S.; Mehtab, V.; Sumana, C. Recent advances on sources and industrial applications of lipases. Biotechnol. Prog. 2018, 34, 5–28. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Liu, Y.; Liang, X.; Yang, Y.; Li, Q. Enantio-, regio-, and chemoselective lipase-catalyzed polymer synthesis. Macromol. Biosci. 2018, 18, e1800131. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Black, D.S.; Reilly, P.J. Carboxylic ester hydrolases: Classification and database derived from their primary, secondary, and tertiary structures. Protein Sci. 2016, 25, 1942–1953. [Google Scholar] [CrossRef] [PubMed]
- Jochen, H.; Hesseler, M.; Stiba, K.; Padhi, S.K.; Kazlauskas, R.J.; Bronscheuer, U.T. Protein engineering of α/β-hydrolase fold enzymes. Chembiochem 2011, 12, 1508–1517. [Google Scholar] [CrossRef] [PubMed]
- Khan, F.I.; Lan, D.; Durrani, R.; Huan, W.; Zhao, Z.; Wang, Y. The lid domain in lipases: Structural and functional determinant of enzymatic properties. Front. Bioeng. Biotechnol. 2017, 5, 16. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Meng, X.; Xu, X.; Liu, W.; Li, S. The molecular basis for lipase stereoselectivity. Appl. Microbiol. Biotechnol. 2018, 102, 3487–3495. [Google Scholar] [CrossRef]
- De Miranda, A.S.; Miranda, L.S.; de Souza, R.O. Lipases: Valuable catalysts for dynamic kinetic resolutions. Biotechnol. Adv. 2015, 33, 372–393. [Google Scholar] [CrossRef]
- Akoh, C.C.; Lee, G.C.; Liaw, Y.C.; Huang, T.H.; Shaw, J.F. GDSL family of serine esterases/lipases. Prog. Lipid Res. 2004, 43, 534–552. [Google Scholar] [CrossRef]
- Leščić Ašler, I.; Ivić, N.; Kovačić, F.; Schell, S.; Knorr, J.; Krauss, U.; Wilhelm, S.; Kojić-Prodić, B.; Jaeger, K.E. Probing enzyme promiscuity of SGNH hydrolases. Chembiochem 2010, 11, 2158–2167. [Google Scholar] [CrossRef]
- Wilhelm, S.; Rosenau, F.; Kolmar, H.; Jaeger, K.E. Autotransporters with GDSL passenger domains: Molecular physiology and biotechnological applications. Chembiochem 2011, 12, 1476–1485. [Google Scholar] [CrossRef]
- Soni, S.; Sathe, S.S.; Odaneth, A.A.; Lali, A.M.; Chandrayan, S.K. SGNH hydrolase-type esterase domain containing Cbes-AcXE2: A novel and thermostable acetyl xylan esterase from Caldicellulosiruptor bescii. Extremophiles 2017, 21, 687–697. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Ryu, B.H.; Kim, J.; Yoo, W.; An, D.R.; Kim, B.Y.; Kwon, S.; Lee, S.; Wang, Y.; Kim, K.K.; et al. Characterization of a novel SGNH-type esterase from Lactobacillus plantarum. Int. J. Biol. Macromol. 2017, 96, 560–568. [Google Scholar] [CrossRef] [PubMed]
- Shahinyan, G.; Margaryan, A.; Panosyan, H.; Trchounian, A. Identification and sequence analyses of novel lipase encoding novel thermophillic bacilli isolated from Armenian geothermal springs. BMC Microbiol. 2017, 17, 103. [Google Scholar] [CrossRef] [PubMed]
- Yu, N.; Yang, J.C.; Yin, G.T.; Li, R.S.; Zou, W.T.; He, C. Identification and characterization of a novel esterase from Thauera sp. Biotechnol. Appl. Biochem. 2018, 65, 748–755. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.M.; Yu, T.T.; Han, N.Y.; Yu, J.L.; Li, J.J.; Yang, Y.J.; Tang, X.H.; Xu, B.; Zhou, J.P.; Tang, H.Z.; et al. Identification and characterization of a new 7-Aminocephalosporanic acid deacetylase from Thermophilic Bacterium Alicyclobacillus tengchongensis. J. Bacteriol. 2015, 198, 311–320. [Google Scholar] [CrossRef]
- Ding, J.; Zhu, H.; Ye, Y.; Li, J.; Han, N.; Wu, Q.; Huang, Z.; Meng, Z. A thermostable and alkaline GDSL-motif esterase from Bacillus sp. K91: Crystallization and X-ray crystallographic analysis. Acta Crystallogr. Sect. F Struct. Biol. Commun. 2018, 74, 117–121. [Google Scholar] [CrossRef]
- Privé, F.; Kaderbhai, N.N.; Girdwood, S.; Worgan, H.J.; Pinloche, E.; Scollan, N.D.; Huws, S.A.; Newbold, C.J. Identification and characterization of three novel lipases belonging to families II and V from Anaerovibrio lipolyticus 5ST. PLoS ONE 2013, 8, e69076. [Google Scholar] [CrossRef]
- Kim, S.; Bae, S.Y.; Kim, S.J.; Ngo, T.D.; Kim, K.K.; Kim, T.D. Characterization, amyloid formation, and immobilization of a novel SGNH hydrolase from Listeria innocua 11262. Int. J. Biol. Macromol. 2012, 50, 103–111. [Google Scholar] [CrossRef]
- Bae, S.Y.; Ryu, B.H.; Jang, E.; Kim, S.; Kim, T.D. Characterization and immobilization of a novel SGNH hydrolase (Est24) from Sinorhizobium meliloti. Appl. Microbiol. Biotechnol. 2013, 97, 1637–1647. [Google Scholar] [CrossRef]
- Brown, L.; Pingitore, E.V.; Mozzi, F.; Saavedra, L.; Villegas, J.M.; Hebert, E.M. Lactic acid bacteria as cell factories for the generation of bioactive peptides. Protein Pept. Lett. 2017, 24, 146–155. [Google Scholar] [CrossRef]
- Hatti-Kaul, R.; Chen, L.; Dishisha, T.; Enshasy, H.E. Lactic acid bacteria: From starter cultures to producers of chemicals. FEMS Microbiol. Lett. 2018, 365, fny213. [Google Scholar] [CrossRef] [PubMed]
- Stefanovic, E.; Fitzgerald, G.; McAuliffe, O. Advances in the genomics and metabolomics of dairy lactobacilli: A review. Food Microbiol. 2017, 61, 33–49. [Google Scholar] [CrossRef] [PubMed]
- Heeney, D.D.; Gareau, M.G.; Marco, M.L. Intestinal Lactobacillus in health and disease, a driver or just along for the ride? Curr. Opin. Biotechnol. 2018, 49, 140–147. [Google Scholar] [CrossRef] [PubMed]
- Lili, Z.; Junyan, W.; Hongfei, Z.; Baoqing, Z.; Bolin, Z. Detoxification of cancerogenic compounds by lactic acid bacteria strains. Crit. Rev. Food Sci. Nutr. 2018, 58, 2727–2742. [Google Scholar] [CrossRef]
- Dong, X.; Yi, H.; Han, C.T.; Nou, I.S.; Hur, Y. GDSL esterase/lipase genes in Brassica rapa L.: Genome-wide identification and expression analysis. Mol. Genet. Genomics 2016, 291, 531–542. [Google Scholar] [CrossRef]
- Mozhaev, V.V.; Martinek, K. Structure-stability relationships in proteins: New approaches to stabilizing enzymes. Enzym. Microb. Technol. 1984, 6, 50–59. [Google Scholar] [CrossRef]
- Kang, L.J.; Meng, Z.T.; Hu, C.; Zhang, Y.; Guo, H.L.; Li, Q.; Li, M. Screening, purification, and characterization of a novel organic solvent-tolerant esterase, Lip2, from Monascus purpureus strain M7. Extremophiles 2017, 21, 345–355. [Google Scholar] [CrossRef]
- Shakiba, M.H.; Ali, M.S.; Rahman, R.N.; Salleh, A.B.; Leow, T.C. Cloning, expression and characterization of a novel cold-adapted GDSL family esterase from Photobacterium sp. strain J15. Extremophiles 2016, 20, 44–55. [Google Scholar] [CrossRef]
- Hwang, H.; Kim, S.; Yoon, S.; Ryu, Y.; Lee, S.Y.; Kim, T.D. Characterization of a novel oligomeric SGNH-arylesterase from Sinorhizobium meliloti 1021. Int. J. Biol. Macromol. 2010, 46, 145–152. [Google Scholar] [CrossRef]
- Fritsch, C.; Jänsch, A.; Ehrmann, M.A.; Toelstede, S.; Vogel, R.F. Characterization of cinnamoyl esterases from different Lactobacilli and Bifidobacteria. Curr. Microbiol. 2017, 74, 247–256. [Google Scholar] [CrossRef]
- Wang, Y.; Ryu, B.H.; Yoo, W.; Lee, C.W.; Kim, K.K.; Lee, J.H.; Kim, T.D. Identification, characterization, immobilization, and mutational analysis of a novel acetylesterase with industrial potential (LaAcE) from Lactobacillus acidophilus. Biochim. Biophys. Acta Gen. Subj. 2018, 1862, 197–210. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Geng, X.; Egashira, Y.; Sanada, H. Purification and characterization of a feruloyl esterase from the intestinal bacterium Lactobacillus acidophilus. Appl. Environ. Microbiol. 2004, 70, 2367–2372. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lee, C.W.; Kwon, S.; Park, S.H.; Kim, B.Y.; Yoo, W.; Ryu, B.H.; Kim, H.W.; Shin, S.C.; Kim, S.; Park, H.; et al. Crystal structure and functional characterization of an esterase (EaEST) from Exiguobacterium antarcticum. PLoS ONE 2017, 12, e0169540. [Google Scholar] [CrossRef] [PubMed]
- Oh, C.; Ryu, B.H.; Yoo, W.; Nguyen, D.D.; Kim, T.; Ha, S.C.; Kim, T.D.; Kim, K.K. Identification and Crystallization of Penicillin-Binding Protein/β-Lactamase Homolog (Rp46) from Ruegeria Pomeroyi. Crystals 2017, 7, 6. [Google Scholar]
- Kim, K.; Ryu, B.H.; Kim, S.S.; An, D.R.; Ngo, T.D.; Pandian, R.; Kim, K.K.; Kim, T.D. Structural and biochemical characterization of a carbohydrate acetylesterase from Sinorhizobium melioti 1021. FEBS Lett. 2015, 589, 117–122. [Google Scholar] [CrossRef]
- Prasad, S.; Roy, I. Converting Enzymes into Tools of Industrial Importance. Recent Pat. Biotechnol. 2018, 12, 33–56. [Google Scholar] [CrossRef]
- Hoarau, M.; Badieyan, S.; Marsh, E.N.G. Immobilized enzymes: understanding enzyme—Surface interactions at the molecular level. Org. Biomol. Chem. 2017, 15, 9539–9551. [Google Scholar] [CrossRef]
- Mukherjee, J.; Majumder, A.B.; Gupta, M.N. Adding an appropriate amino acid during crosslinking results in more stable crosslinked enzyme aggregates. Anal. Biochem. 2016, 507, 27–32. [Google Scholar] [CrossRef]
- Cui, J.; Cui, L.; Jia, S.; Su, Z.; Zhang, S. Hybrid cross-linked lipase aggregates with magnetic nanoparticles: A robust and recyclable biocatalysis for the epoxidation of oleic acid. J. Agric. Food Chem. 2016, 64, 7179–7187. [Google Scholar] [CrossRef]
- Sievers, F.; Higgins, D.G. Clustal Omega for making accurate alignments of many protein sequences. Protein Sci. 2018, 27, 135–145. [Google Scholar] [CrossRef]
- Gouet, P.; Robert, X.; Courcelle, E. ESPript/ENDscript: Extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic Acids Res. 2003, 31, 3320–3323. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Seeliger, D.; de Groot, B.L. Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J. Comput. Aided Mol. Des. 2010, 24, 417–422. [Google Scholar] [CrossRef] [PubMed]
- Oh, C.; Ryu, B.H.; Yoo, W.; Nguyen, D.D.; Kim, T.; Ha, S.; Kim, T.D.; Kim, K.K. Identification and crystallographic analysis of a new carbohydrate acetylesterase (SmAcE1) from Sinorhizobium meliloti. Crystals 2018, 8, 12. [Google Scholar] [CrossRef]
- Park, S.H.; Yoo, W.; Lee, C.W.; Jeong, C.S.; Shin, S.C.; Kim, H.W.; Park, H.; Kim, K.K.; Kim, T.D.; Lee, J.H. Crystal structure and functional characterization of a cold-active acetyl xylan esterase (PbAcE) from psychrophilic soil microbe Paenibacillus sp. PLoS ONE 2018, 13, e0206260. [Google Scholar] [CrossRef] [PubMed]


© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Le, L.T.H.L.; Yoo, W.; Jeon, S.; Kim, K.K.; Kim, T.D. Characterization and Immobilization of a Novel SGNH Family Esterase (LaSGNH1) from Lactobacillus acidophilus NCFM. Int. J. Mol. Sci. 2020, 21, 91. https://doi.org/10.3390/ijms21010091
Le LTHL, Yoo W, Jeon S, Kim KK, Kim TD. Characterization and Immobilization of a Novel SGNH Family Esterase (LaSGNH1) from Lactobacillus acidophilus NCFM. International Journal of Molecular Sciences. 2020; 21(1):91. https://doi.org/10.3390/ijms21010091
Chicago/Turabian StyleLe, Ly Thi Huong Luu, Wanki Yoo, Sangeun Jeon, Kyeong Kyu Kim, and T. Doohun Kim. 2020. "Characterization and Immobilization of a Novel SGNH Family Esterase (LaSGNH1) from Lactobacillus acidophilus NCFM" International Journal of Molecular Sciences 21, no. 1: 91. https://doi.org/10.3390/ijms21010091
APA StyleLe, L. T. H. L., Yoo, W., Jeon, S., Kim, K. K., & Kim, T. D. (2020). Characterization and Immobilization of a Novel SGNH Family Esterase (LaSGNH1) from Lactobacillus acidophilus NCFM. International Journal of Molecular Sciences, 21(1), 91. https://doi.org/10.3390/ijms21010091






