Systematic Analysis of the Maize OSCA Genes Revealing ZmOSCA Family Members Involved in Osmotic Stress and ZmOSCA2.4 Confers Enhanced Drought Tolerance in Transgenic Arabidopsis
Abstract
1. Introduction
2. Results
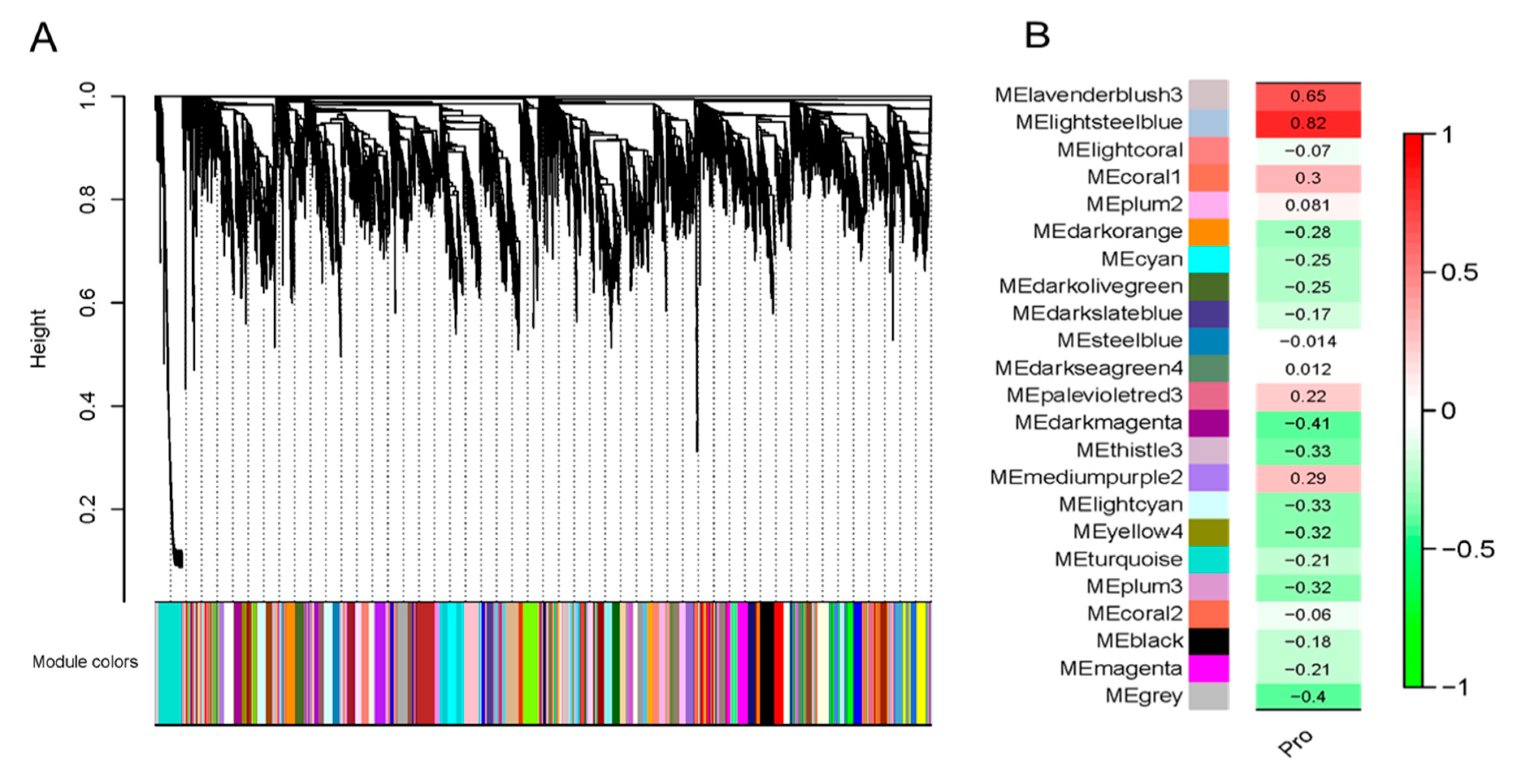
2.1. Transcriptome Co-Expression Network Module Construction and Association Analysis Module and Proline Content
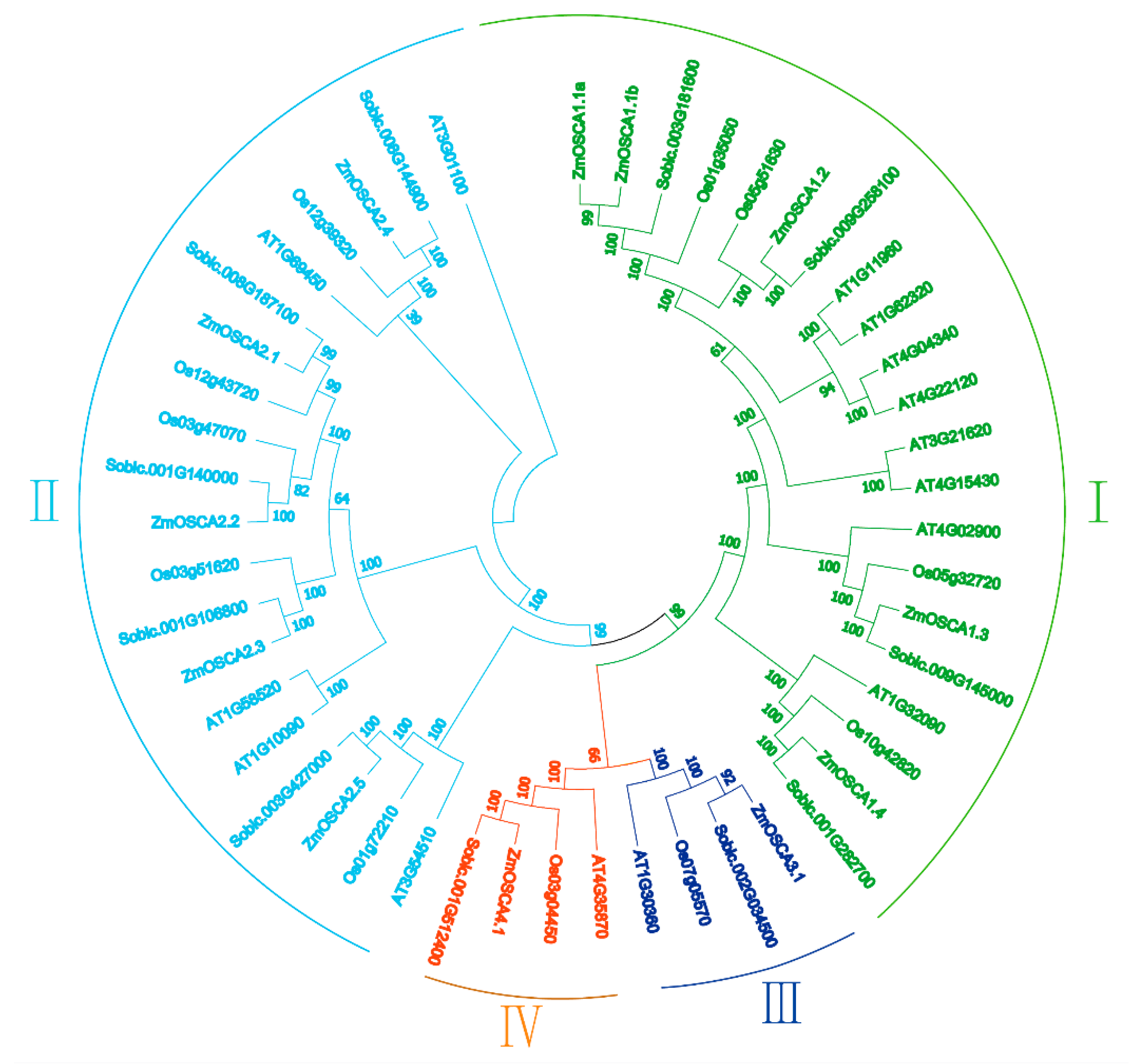
2.2. Genome-Wide Identification and Classification of OSCA Genes in Maize
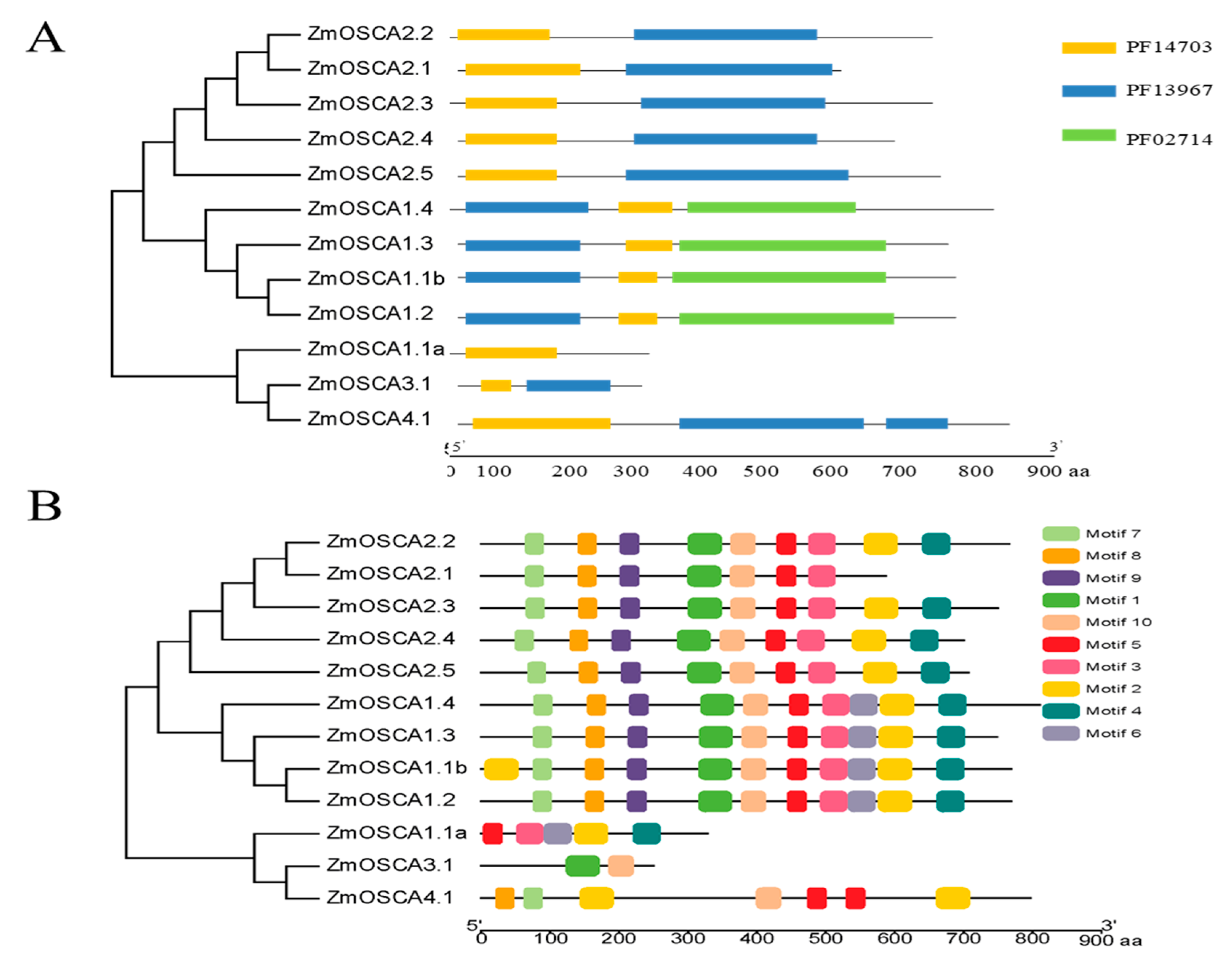
2.3. Additional Conserved Motifs in OSCA Genes in Maize
2.4. Cis-Element Analysis of OSCA Gene Promoter Sequences
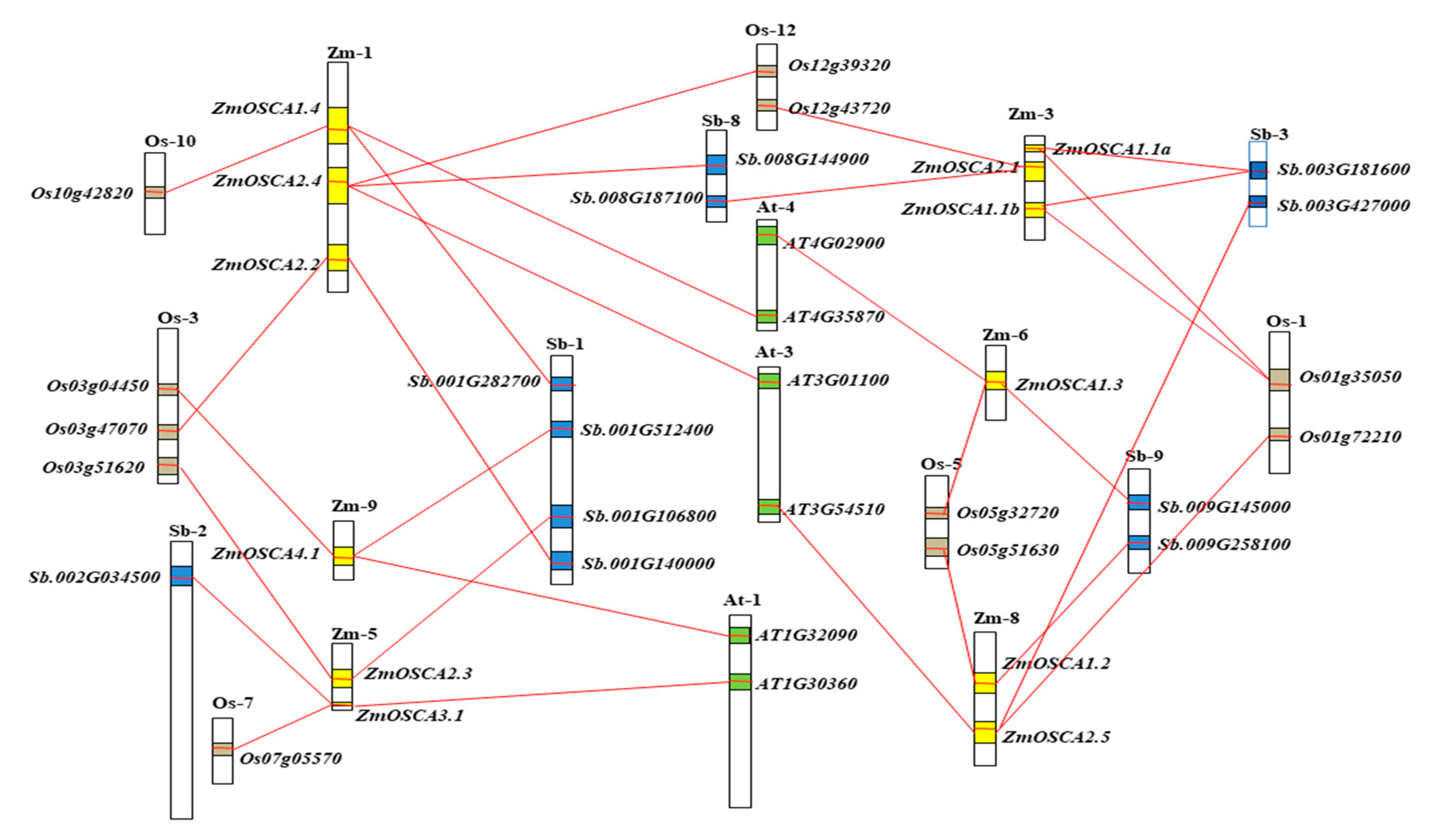
2.5. Collinearity Relationships of OSCA Genes among Maize, Sorghum, Rice, and Arabidopsis
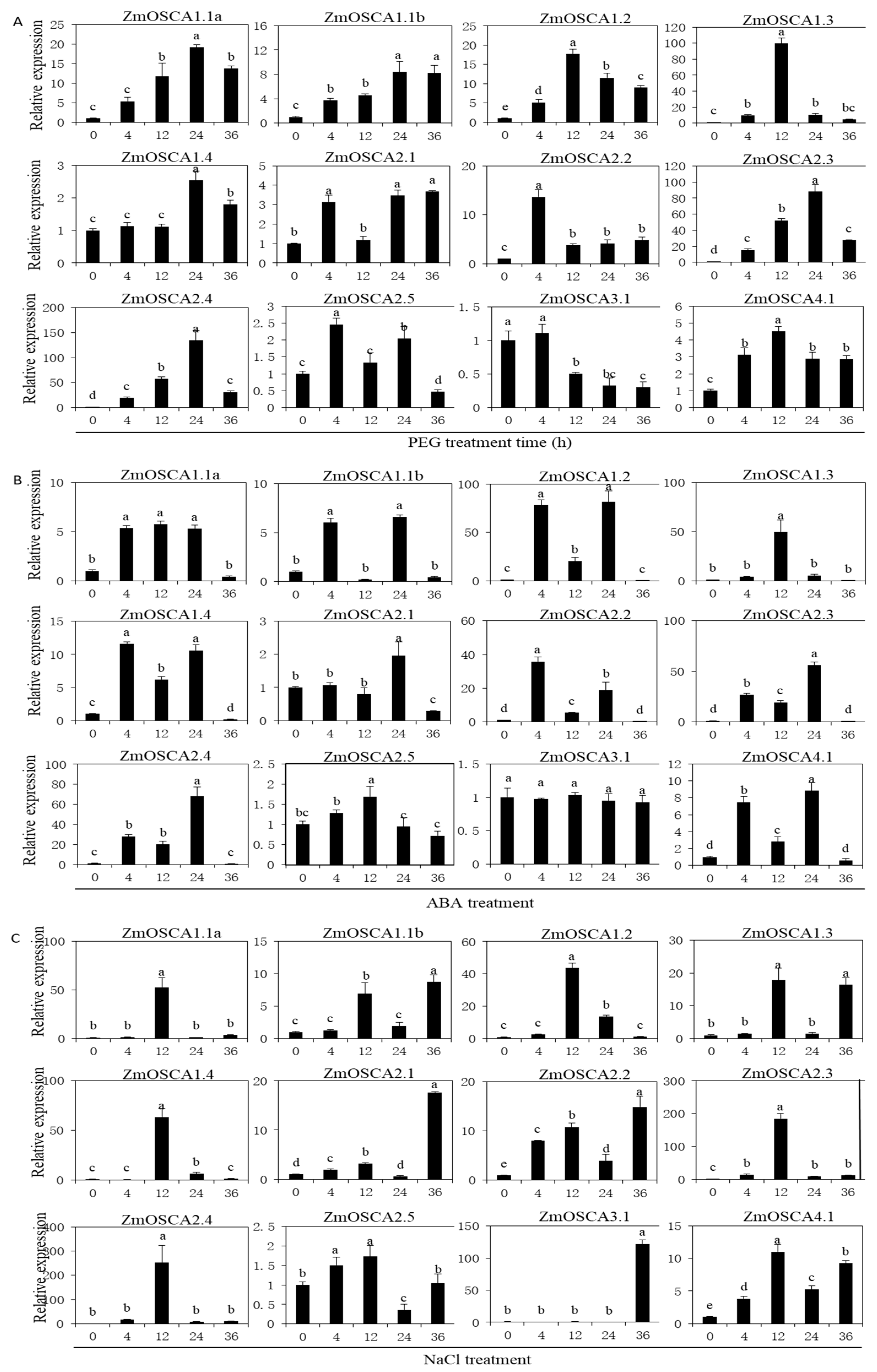
2.6. Expression of ZmOSCA Genes under Abiotic Stress
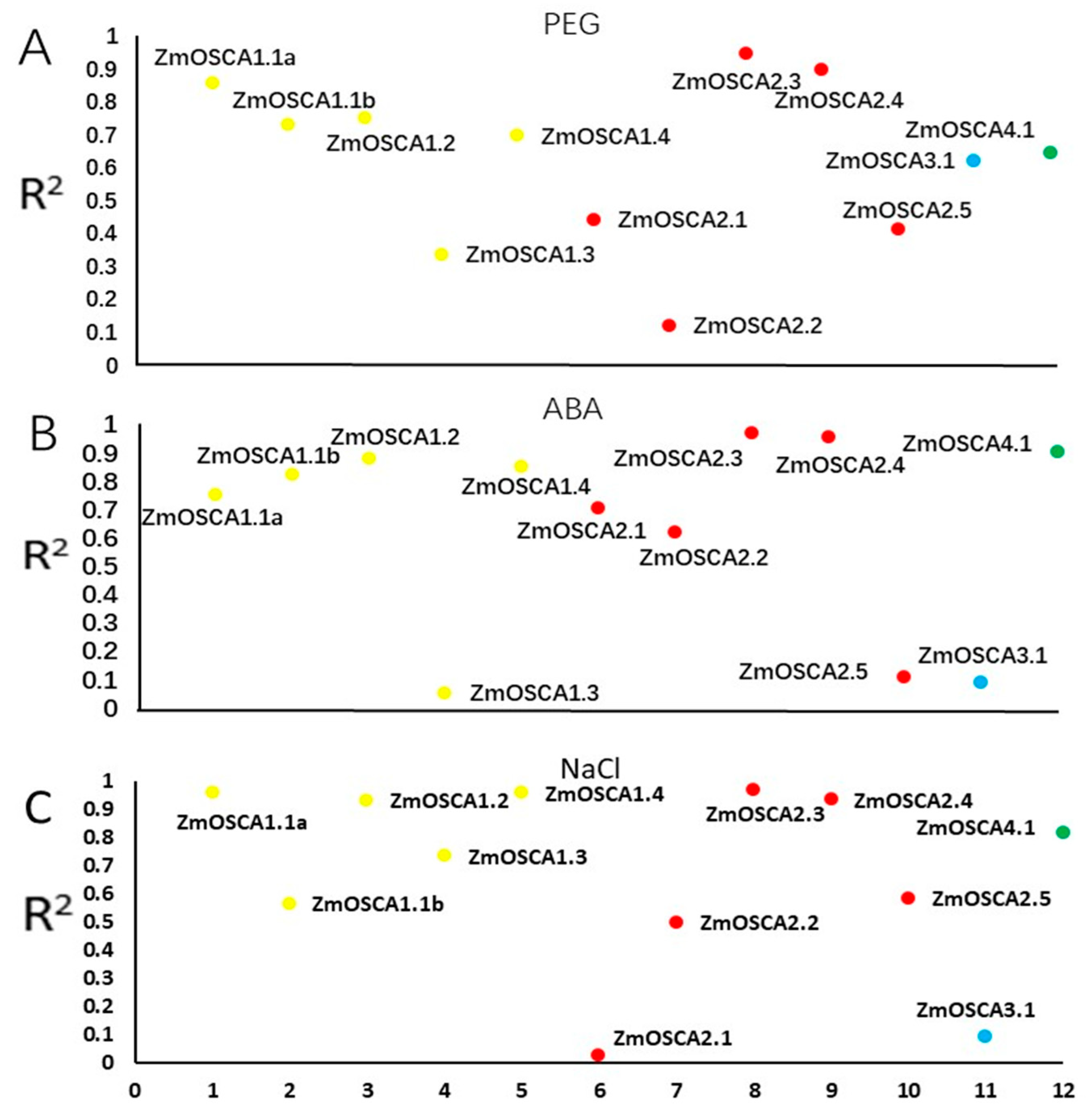
2.7. Analysis of the Relationship between ZmOSCAs Gene and Proline Content
2.8. Predicting the Protein Interaction Network of ZmOSCAs
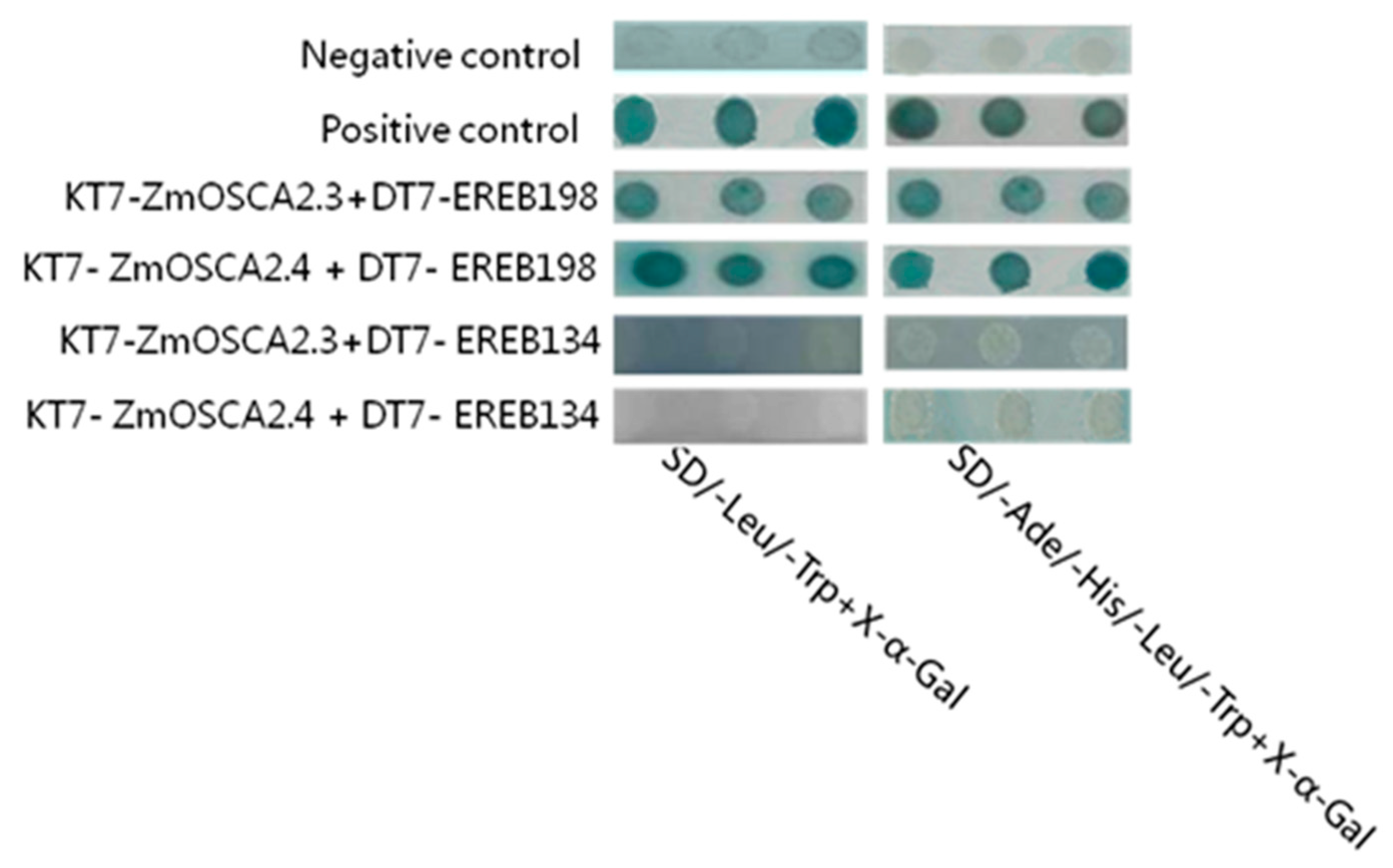
2.9. Yeast Two-Hybrid and Bimolecular Fluorescence Complementation Validate the Interaction Proteins of ZmOSCA2.3 and ZmOSCA2.4
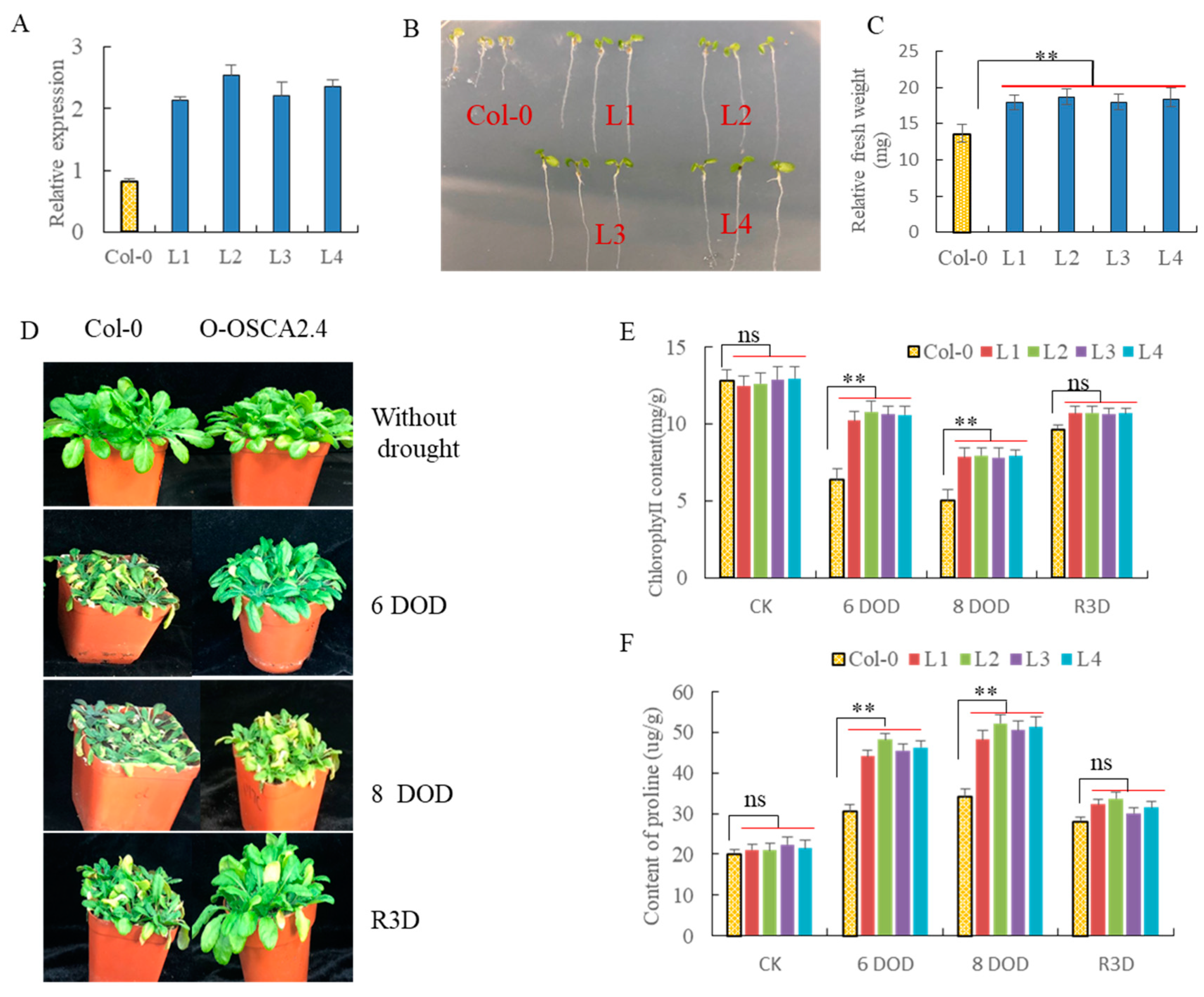
2.10. Over-Expression of ZmOSCA2.4 in Arabidopsis Enhances Plant Tolerance to Drought Stress
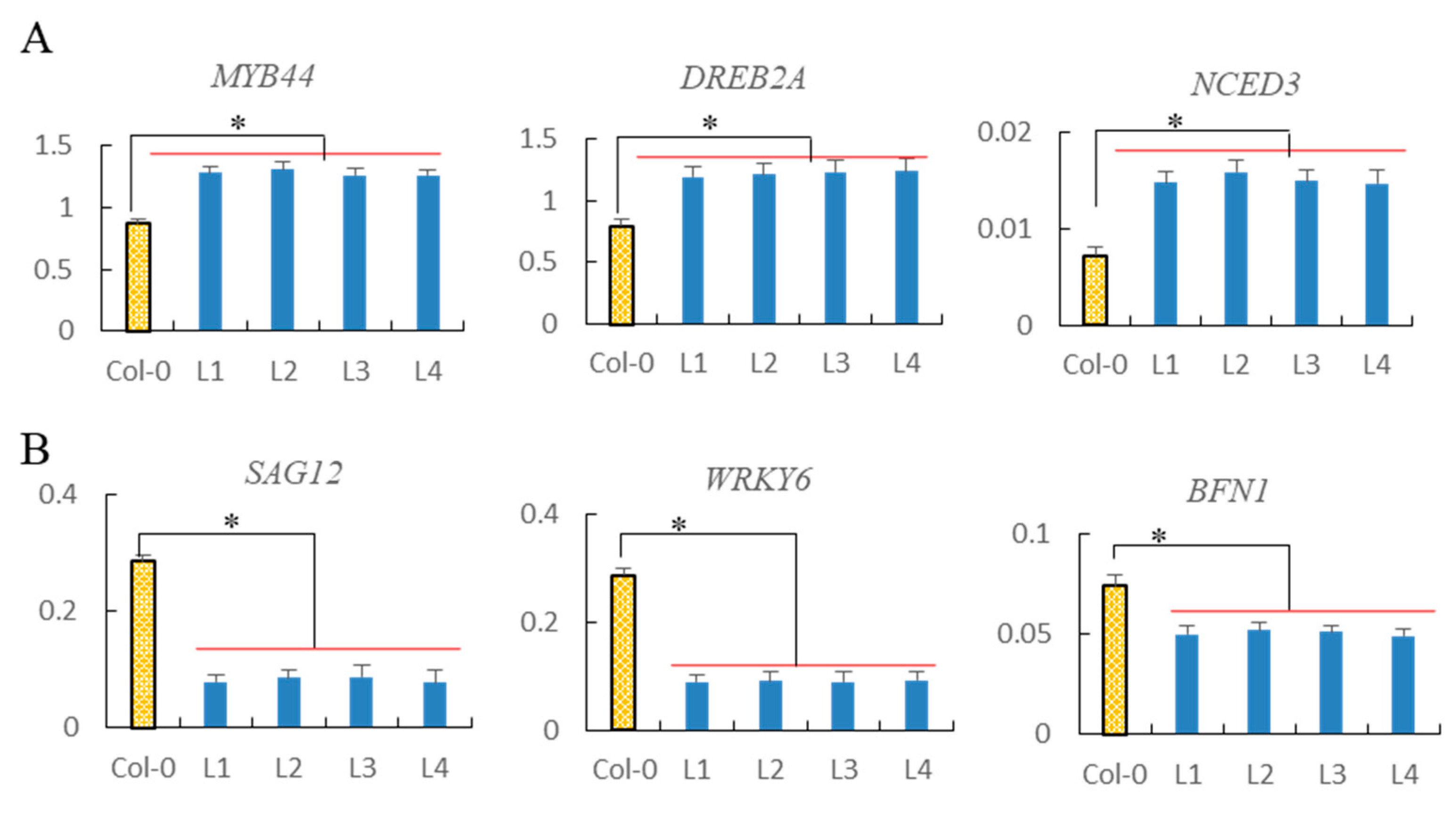
2.11. Analysis of Drought Tolerance-Associated and Senescence-Associated Genes Expression
2.12. Drought Stress and Rewatering Transcriptome and Small RNA Sequencing Predicted the Regulation of ZmOSCA2.4 Gene by miR5054
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Stress Treatment
4.2. Isolation of RNA and Quantitative Real-Time PCR (qPCR) Analysis
4.3. Transcriptome Co-Expression Network Module Construction and Association Analysis Module and Proline Content
4.4. Identification of OSCA Protein-Coding Genes in the Maize Genome
4.5. Gene Structure, Additional Conserved Motifs, and Cis-Elements in the Promoter Regions of Abiotic Stress-Responsive Analysis of ZmOSCA Genes
4.6. Interspecies Microsynteny Analysis
4.7. Determination of Chlorophyll and Proline Content
4.8. Predicting the Interaction of ZmOSCA Proteins
4.9. Validation of Interactions between Proteins using Yeast Two-Hybrid and BIFC
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Aceved, O.E.; Hsiao, T.C.; Henderson, D.W. Immediate and subsequent growth responses of maize leaves to changes in water status. Plant Physiol. 1971, 48, 631–636. [Google Scholar] [CrossRef] [PubMed]
- Lang, K.; Lang, P.; Bauer, C.; Duranton, C.; Wieder, T.; Huber, S. Mechanisms of Suicidal Erythrocyte Death. Cell. Physiol. Biochem. 2005, 15, 195–202. [Google Scholar] [CrossRef] [PubMed]
- Dornhorst, A.C. Review of Medical Physiology. Anesthesiology 2001, 52, 959–960. [Google Scholar]
- Neuhofer, W.; Beck, F. Survival in Hostile Environments: Strategies of Renal Medullary Cells. Physiology 2006, 21, 171–180. [Google Scholar] [CrossRef]
- Arakawa, T.; Timasheff, S.N. The stabilization of proteins by osmolytes. Biophys. J. 1985, 47, 411–414. [Google Scholar] [CrossRef]
- Batistie, O.; Kudla, J. Analysis of calcium signaling pathways in plants. Biochim. Biophys. Acta (BBA)-Gen. Subj. 2012, 1820, 1283–1293. [Google Scholar] [CrossRef]
- Steinhorst, L.; Kudla, J. Calcium-A central regulator of pollen germination and tube growth. Biochim. Biophys. Acta (BBA)-Mol. Cell Res. 2013, 1833, 1573–1581. [Google Scholar] [CrossRef]
- Hubbard, K.E.; Siegel, R.S.; Valerio, G.; Brandt, B.; Schroeder, J.I. Abscisic acid and CO2 signaling via calcium sensitivity priming in guard cells, new CDPK mutant phenotypes and a method for improved resolution of stomatal stimulus-response analyses. Ann. Bot. 2012, 109, 5–17. [Google Scholar] [CrossRef]
- Hepler, P.K. Calcium: A Central Regulator of Plant Growth and Development. Plant Cell 2005, 17, 2142–2155. [Google Scholar] [CrossRef]
- Boudsocq, M.; Sheen, J. Stress Signaling II: Calcium Sensing and Signaling. Abiotic Stress Adapt. Plant 2009, 75–90. [Google Scholar]
- Trewavas, A.; Knight, M. Mechanical signaling, calcium and plant form. Plant Mol. Biol. 1994, 26, 1329–1341. [Google Scholar] [CrossRef] [PubMed]
- Hou, C.C.; Tian, W.; Kleist, T.; He, K.; Garcia, V.; Bai, F. DUF221 proteins are a family of osmosensitive calcium-permeable cation channels conserved across eukaryotes. Cell Res. 2014, 24, 632–635. [Google Scholar] [CrossRef] [PubMed]
- Yuan, F.; Yang, H.; Xue, Y.; Kong, D.; Ye, R.; Li, C.; Zhang, J.; Theprungsirikul, L.; Shrift, T.; Krichilsky, B.; et al. OSCA1 mediates osmotic-stress-evoked Ca2+ increases vital for osmosensing in Arabidopsis. Nature 2014, 514, 367–371. [Google Scholar] [CrossRef] [PubMed]
- Li, J.W.; Yang, J.K.; Jia, B.W. Evolution and expression analysis of OSCA gene family in soybean. Chin. J. Oil Crop Sci. 2017, 16, 16–26. [Google Scholar]
- Liu, Y.; Li, H.; Shi, Y. A Maize Early Responsive to Dehydration Gene, ZmERD4, Provides Enhanced Drought and Salt Tolerance in Arabidopsis. Plant Mol. Biol. Reporter 2009, 27, 542–548. [Google Scholar] [CrossRef]
- Lv, G.D. Cloning, Molecular Msrkers and Function of the Gene TaOSCA1.4 in Common Wheat; Shandong Agricultural University: Taian, China, 2015. [Google Scholar]
- Ding, S.; Feng, X.; Du, H.; Wang, H. Genome-wide analysis of maize OSCA family members and their involvement in drought stress. Peer J. 2019, 7, e6765. [Google Scholar] [CrossRef]
- Chen, B.; Li, G.F.; Shen, Y.; Huang, X.; Xu, Y.J. Reducing iron accumulation: A potential approach for the prevention and treatment of postmenopausal osteoporosis. Exp. Ther. Med. 2015, 2484, 7–11. [Google Scholar] [CrossRef]
- Konstantina, K.V.; Peter, I.P.; Georgi, I.G. Physiological and anatomical responses of wheat to induced dehydration and rehydration. Cent. Eur. J. Biol. 2013, 8, 499–503. [Google Scholar]
- Ji, L.X.; Wang, J.; Ye, M.; Li, Y.; Guo, B.; Chen, Z.; Li, H.; An, X. Identification and characterization of the Populus AREB/ABF subfamily. J. Integr. Pl. Biol. 2013, 55, 177–186. [Google Scholar] [CrossRef]
- Fujita, Y.; Yoshida, T.; Yamaguchi-Shinozaki, K. Pivotal role of the AREB/ABF-SnRK2 pathway in ABRE-mediated transcription in response to osmotic stress in plants. Physiol. Plant 2013, 147, 15–27. [Google Scholar] [CrossRef]
- Wang, W.; Vinocur, B.; Altman, A. Plant responses to drought, salinity and extreme temperatures: towards genetic engineering for stress tolerance. Planta 2003, 218, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Shavrukov, Y. Salt stress or salt shock: Which genes are we studying? J. Ex Bot. 2013, 64, 119–127. [Google Scholar] [CrossRef] [PubMed]
- Finkelstein, R. Abscisic Acid synthesis and response. Arabidopsis Book. 2013, 11, e0166. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, M.S.A.; Javed, F.; Ashraf, M. Iso-osmotic effect of NaCl and PEG on growth, cations and free proline accumulation in callus tissue of two indicia rice (Oryza sativa L.) genotypes. Plant Growth Regul. 2007, 53, 53–63. [Google Scholar] [CrossRef]
- Li, Y.S.; Yuan, F.; Wen, Z.H. Genome -wide survey and expression analysis of the OSCA gene family in rice. BMC Plant Biol. 2015, 15, 261. [Google Scholar] [CrossRef]
- Jojoa-Cruz, S.; Saotome, K.; Murthy, S.E.; Tsui, C.C.A.; Sansom, M.S.P.; Patapoutian, A.; Ward, A.B. Cryo-EM structure of the mechanically activated ion channel OSCA1.2. eLife Sci. 2018, 7, e41845. [Google Scholar] [CrossRef]
- Maity, K.; Heumann, J.; McGrath, AP.; Kopcho, N.J.; Hsu, P.K.; Lee, C.W.; Mapes, J.H.; Garza, D.; Krishnan, S.; Morgan, G.P.; et al. Cryo-EM structure of OSCA1.2 from Oryza sativa: mechanical basis of hyperosmolality-gating in plants. Proc. Natl. Acad. Sci. USA 2019, 116, 14309–14318. [Google Scholar] [CrossRef]
- Kiyosue, T.; Yamaguchishinozaki, K.; Shinozaki, K. Cloning of cDNAs for genes that are early-responsive to dehydration stress (ERDs) in Arabidopsis thaliana L.: identification of three ERDs as HSP cognate genes. Plant Mol. Biol. 1994, 25, 791–798. [Google Scholar] [CrossRef]
- Chatzigeorgiou, M.; Bang, S.; Sun, W.H. Tmc-1 encodes a sodium-sensitive channel required for salt chemosensation in C. elegans. Nature 2013, 494, 95–99. [Google Scholar] [CrossRef]
- Yoshida, T.; Mogami, J.; Yamaguchi-Shinozaki, K. ABA-dependent and ABA-independent signaling in response to osmotic stress in plants. Curr. Opin. Plant Biol. 2014, 21, 133–139. [Google Scholar] [CrossRef]
- Fujita, Y.; Fujita, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. ABA-mediated transcriptional regulation in response to osmotic stress in plants. J. Plant Res. 2011, 124, 509–525. [Google Scholar] [CrossRef] [PubMed]
- Cui, Z.H.; Wang, Y.F.; Fan, J.J. Research Advances of plant osmotic regulation. J. Maize Sci. 2007, 15, 140–143. [Google Scholar]
- Qu, F.F. Physiological and biochemical mechanisms related to salt tolerance in plants. J. Henan Sci. Technol. 2015, 141–143. [Google Scholar]
- Andrew, C.A.; Arnaud, C.; Daniele, P. VirusMINT: A viral protein interaction database. Nucleic Acids Res. 2009, 37, D669–D673. [Google Scholar]
- Jofuku, K.D.; Den Boer, B.; Van Montagu, M.; Okamuro, J.K. Control of Arabidopsis flower and seed development by the homeotic gene APETALA2. Plant Cell 1994, 6, 1211–1225. [Google Scholar] [PubMed]
- Krizek, B. AINTEGUMENTA and AINTEGUMENTA-LIKE6 act redundantly to regulate Arabidopsis floral growth and patterning. Plant Physiol. 2009, 150, 1916–1929. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.S.; Chen, M.; Li, L.C.; Ma, Y.Z. Functions and application of the AP2/ERF transcription factor family in crop improvement. J. Integr. Plant Biol. 2011, 7, 64–79. [Google Scholar]
- Gutterson, N.; Reuber, T.L. Regulation of disease resistance pathways by AP2/ERF transcription factors. Curr. Opin. Plant Biol. 2004, 7, 465–471. [Google Scholar] [CrossRef]
- Jung, C.; Seo, J.S.; Han, S.W.; Koo, Y.J.; Kim, C.H.; Song, S.I.; Nahm, B.H.; Choi, Y.D.; Cheong, J.J. Overexpression of AtMYB44 Enhances Stomatal Closure to Confer Abiotic Stress Tolerance in Transgenic Arabidopsis. Plant Physiol. 2008, 146, 623–635. [Google Scholar] [CrossRef]
- Sakuma, Y.; Maruyama, K.; Osakabe, Y.; Qin, F.; Seki, M.; Shinozaki, K.; YamaguchiShinozaki, K. Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 2006, 18, 1292–1309. [Google Scholar] [CrossRef]
- Weaver, L.M.; Gan, S.; Quirino, B.; Amasino, R.M. A comparison of the expression patterns of several senescence-associated genes in response to stress and hormone treatment. Plant Mol. Biol. 1998, 37, 455–469. [Google Scholar] [CrossRef] [PubMed]
- Robatzek, S.; Somssich, I.E. Targets of AtWRKY6 regulation during plant senescence and pathogen defense. Genes Dev. 2002, 16, 1139–1149. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Amador, M.A.; Abler, M.L.; De Rocher, E.J.; Thompson, D.M.; van Hoof, A.; LeBrasseur, N.D.; Lers, A.; Green, P.J. Identification of BFN1, a Bifunctional Nuclease Induced during Leaf and Stem Senescence in Arabidopsis. Plant Physiol. 2000, 122, 169–180. [Google Scholar]
- Trapnell, C.; Williams, BA.; Pertea, G. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511. [Google Scholar] [CrossRef]
- Pei, G.; Chen, L.; Zhang, W. WGCNA application to proteomic and metabolomic data analysis. Methods Enzymol. 2017, 585, 135–158. [Google Scholar]
- Garg, R.; Singh, V.K.; Rajkumar, M.S. Global transcriptome and coexpression network analyses reveal cultivar-specific molecular signatures associated with seed development and seed size/weight determination in chickpea. Plant J. 2017, 91, 1088–1107. [Google Scholar] [CrossRef]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J.; et al. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, D222–D230. [Google Scholar] [CrossRef]
- Marchlerbauer, A.; Lu, S.; Anderson, J.B.; Chitsaz, F.; Derbyshire, M.K.; DeWeese-Scott, C.; Fong, J.H.; Geer, L.Y.; Geer, R.C.; Gonzales, N.R. CDD: A conserved domain database for the functional annotation of proteins. Nucleic Acids Res. 2011, 39, D225–D229. [Google Scholar] [CrossRef]
- Bates, L.S.; Waldren, R.P.; Teare, I.D. Rapid determination of free proline for water-stress studies. Plant Soil 1973, 39, 205–207. [Google Scholar] [CrossRef]
- Xiao, D.L.; Deng, H.Y.; Xie, L.Y. Construction of a yeast two-hybrid cDNA library of Laodelphax striatellus. Plant Prot. 2011, 37, 19–23. [Google Scholar]


| Gene Name | Gene Identifier | Chromosome | Protein Length (aa) | ORF (bp) | Number of Exons | Transmembrane Domain | Isoelectric Point | Molecular Weight (KDa) | Class |
|---|---|---|---|---|---|---|---|---|---|
| ZmOSCA1.1a | GRMZM2G064189 | 3 | 327 | 984 | 4 | 4 | 8.54 | 36.95 | I |
| ZmOSCA1.1b | GRMZM2G021194 | 3 | 768 | 2307 | 11 | 9 | 9.05 | 87.51 | I |
| ZmOSCA1.2 | GRMZM2G456000 | 8 | 768 | 2307 | 11 | 9 | 9.15 | 87.93 | I |
| ZmOSCA1.3 | GRMZM2G181206 | 6 | 748 | 2247 | 11 | 10 | 9.15 | 85.65 | I |
| ZmOSCA1.4 | GRMZM2G128641 | 1 | 810 | 2433 | 11 | 9 | 8.66 | 93.88 | I |
| ZmOSCA2.1 | GRMZM2G163059 | 3 | 586 | 1761 | 8 | 7 | 9.35 | 66.76 | II |
| ZmOSCA2.2 | GRMZM2G409093 | 1 | 765 | 2298 | 10 | 9 | 9.3 | 87.38 | II |
| ZmOSCA2.3 | GRMZM2G164470 | 5 | 749 | 2250 | 10 | 10 | 8.8 | 86.15 | II |
| ZmOSCA2.4 | GRMZM2G039186 | 1 | 699 | 2100 | 10 | 9 | 8.74 | 79.04 | II |
| ZmOSCA2.5 | GRMZM2G402708 | 8 | 706 | 2121 | 10 | 11 | 9.05 | 79.64 | II |
| ZmOSCA3.1 | GRMZM2G162253 | 5 | 249 | 750 | 2 | 2 | 8.88 | 28.68 | III |
| ZmOSCA4.1 | GRMZM2G059891 | 9 | 796 | 2391 | 1 | 9 | 7.67 | 89.4 | IV |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cao, L.; Zhang, P.; Lu, X.; Wang, G.; Wang, Z.; Zhang, Q.; Zhang, X.; Wei, X.; Mei, F.; Wei, L.; et al. Systematic Analysis of the Maize OSCA Genes Revealing ZmOSCA Family Members Involved in Osmotic Stress and ZmOSCA2.4 Confers Enhanced Drought Tolerance in Transgenic Arabidopsis. Int. J. Mol. Sci. 2020, 21, 351. https://doi.org/10.3390/ijms21010351
Cao L, Zhang P, Lu X, Wang G, Wang Z, Zhang Q, Zhang X, Wei X, Mei F, Wei L, et al. Systematic Analysis of the Maize OSCA Genes Revealing ZmOSCA Family Members Involved in Osmotic Stress and ZmOSCA2.4 Confers Enhanced Drought Tolerance in Transgenic Arabidopsis. International Journal of Molecular Sciences. 2020; 21(1):351. https://doi.org/10.3390/ijms21010351
Chicago/Turabian StyleCao, Liru, Pengyu Zhang, Xiaomin Lu, Guorui Wang, Zhenhua Wang, Qianjin Zhang, Xin Zhang, Xin Wei, Fujian Mei, Li Wei, and et al. 2020. "Systematic Analysis of the Maize OSCA Genes Revealing ZmOSCA Family Members Involved in Osmotic Stress and ZmOSCA2.4 Confers Enhanced Drought Tolerance in Transgenic Arabidopsis" International Journal of Molecular Sciences 21, no. 1: 351. https://doi.org/10.3390/ijms21010351
APA StyleCao, L., Zhang, P., Lu, X., Wang, G., Wang, Z., Zhang, Q., Zhang, X., Wei, X., Mei, F., Wei, L., & Wang, T. (2020). Systematic Analysis of the Maize OSCA Genes Revealing ZmOSCA Family Members Involved in Osmotic Stress and ZmOSCA2.4 Confers Enhanced Drought Tolerance in Transgenic Arabidopsis. International Journal of Molecular Sciences, 21(1), 351. https://doi.org/10.3390/ijms21010351




