Interference with the Cannabinoid Receptor CB1R Results in Miswiring of GnRH3 and AgRP1 Axons in Zebrafish Embryos
Abstract
1. Introduction
2. Results
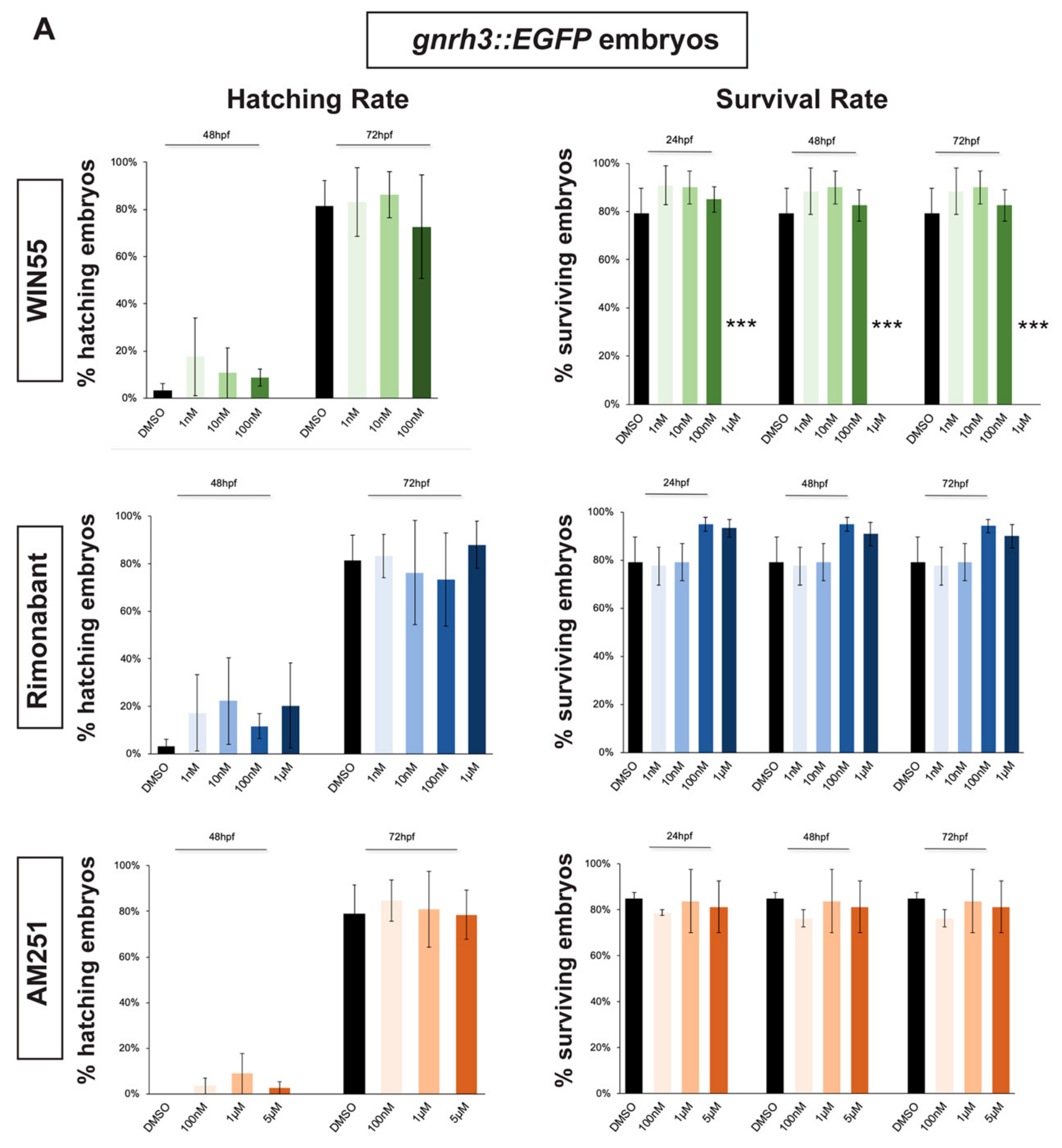
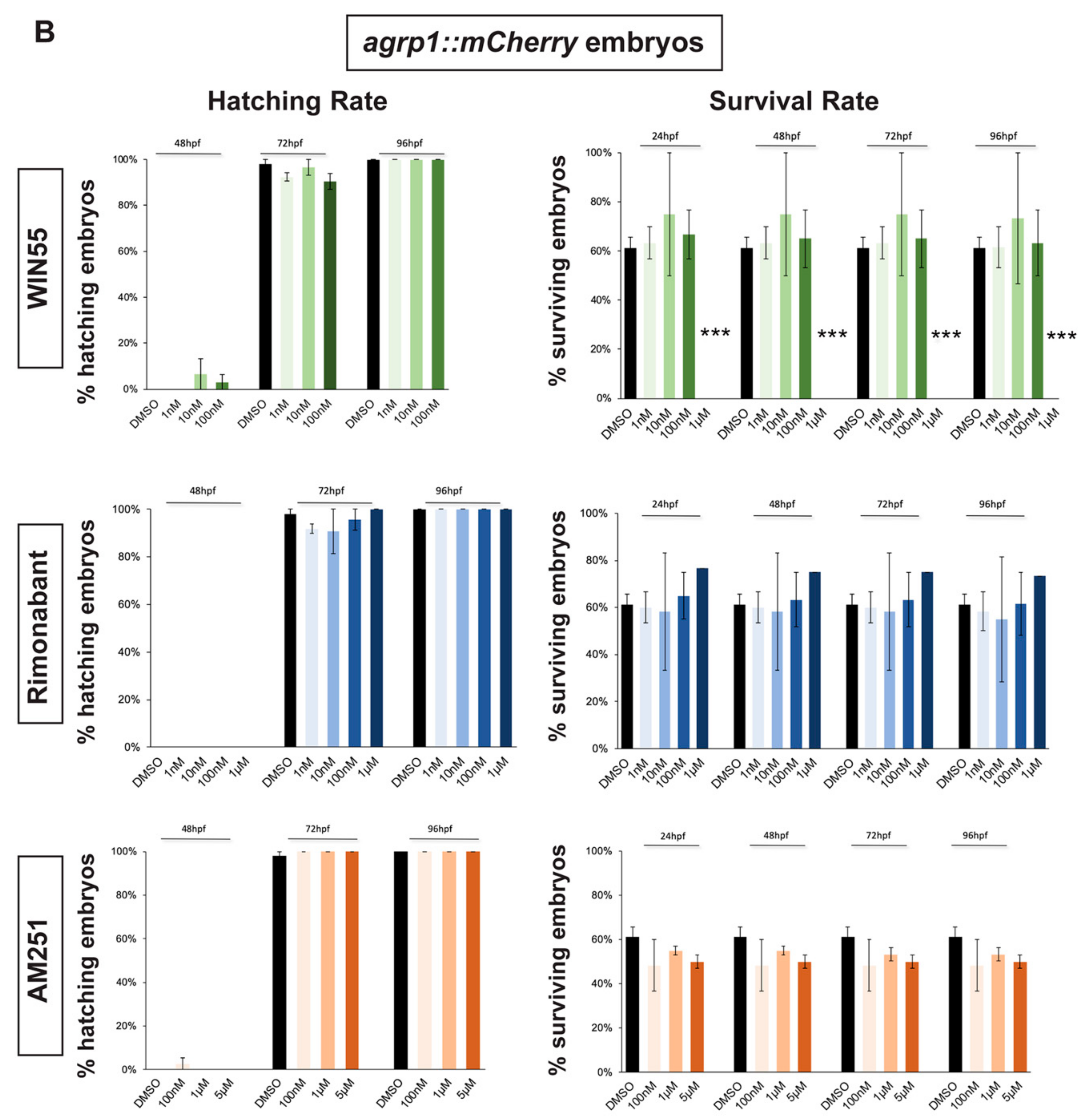
2.1. Survival and Hatching Rates of Zebrafish Embryos Treated with CB1R Ligands
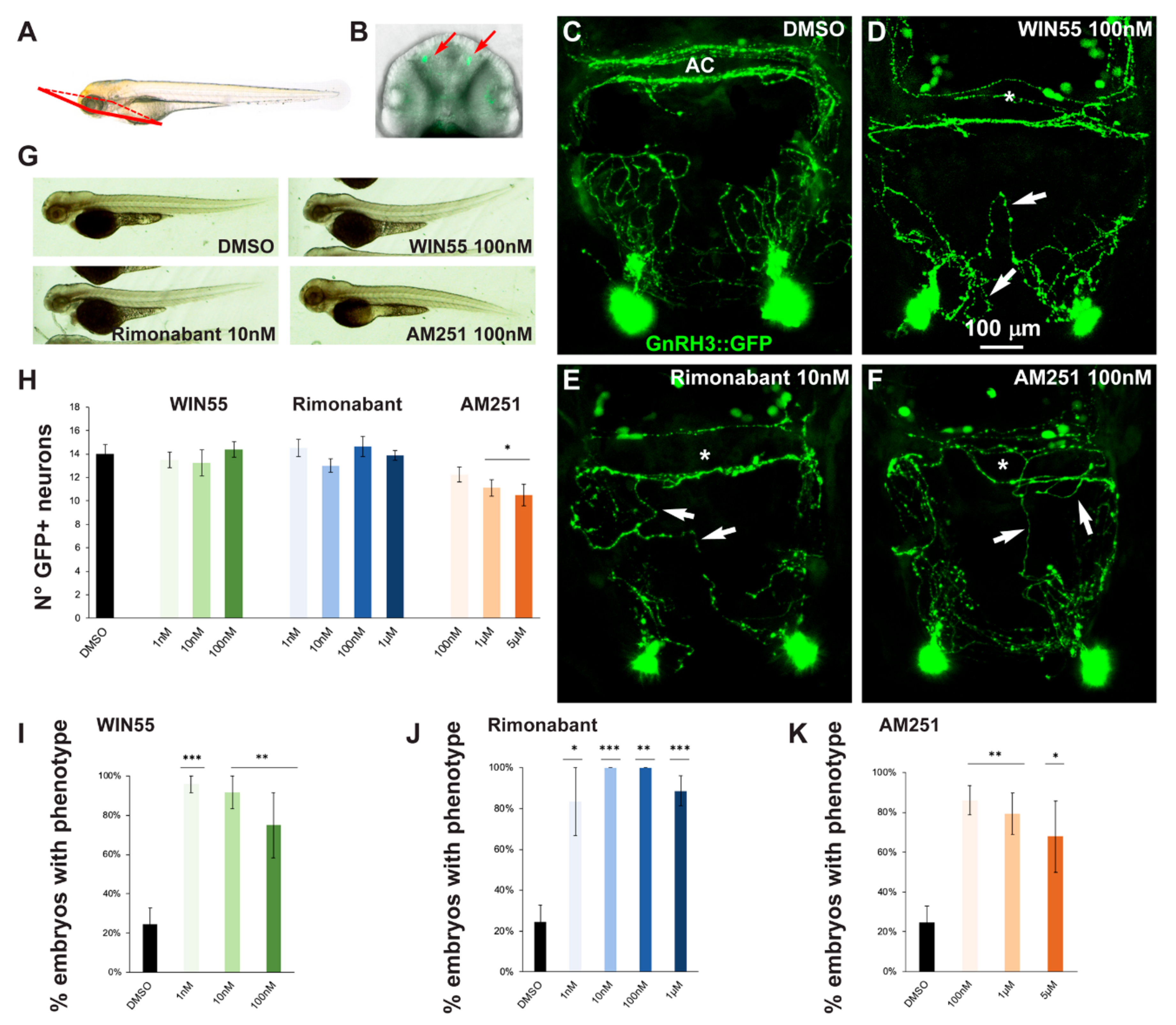
2.2. Effects of CB1R Ligands on Zebrafish GnRH3 Neurons
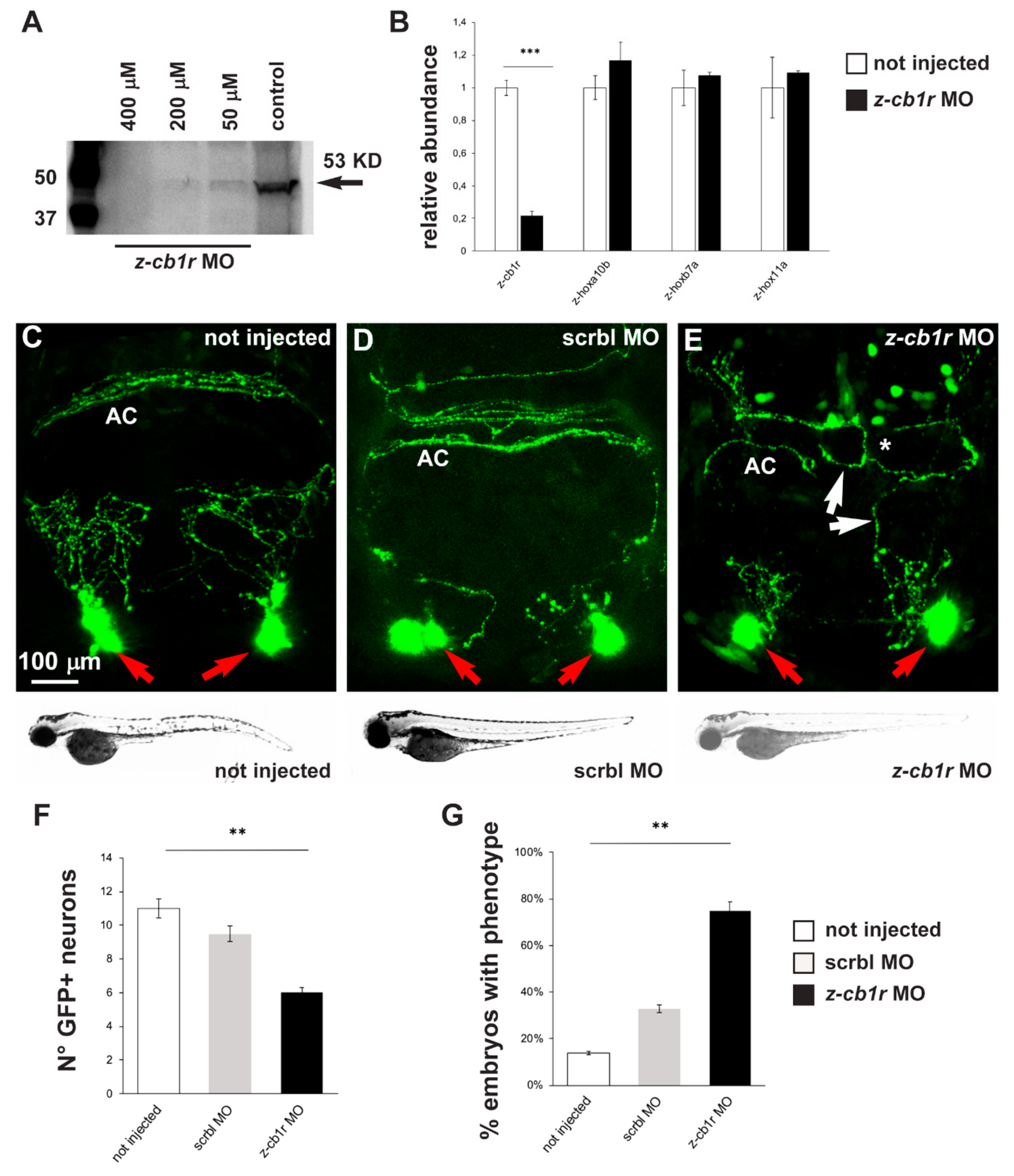
2.3. Effects of CB1R Knockdown on Zebrafish GnRH3 Neurons
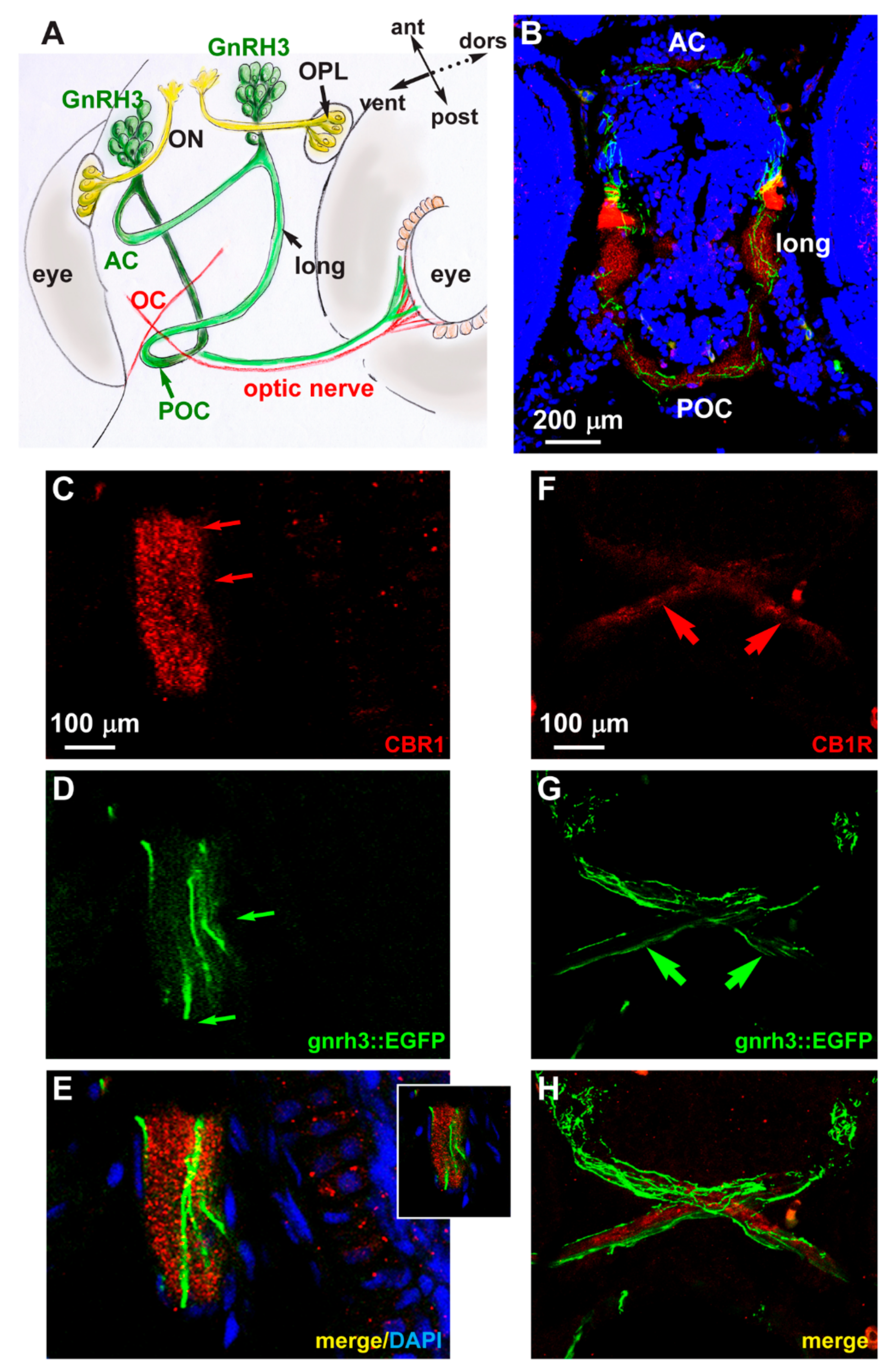
2.4. Expression of CB1R in Developing Zebrafish Brain
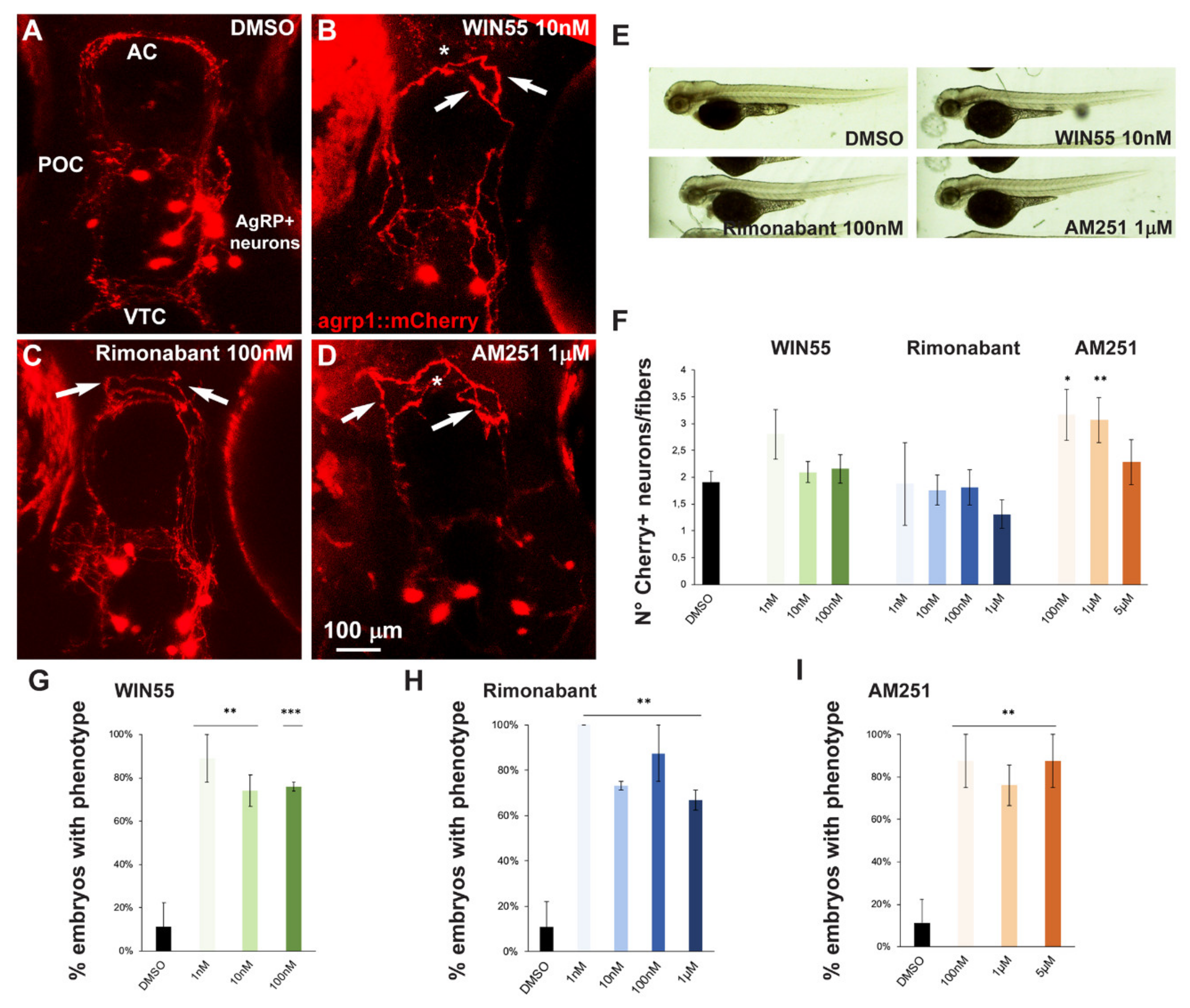
2.5. Effect of Exposure to CB1R Ligands on AgRP1 Neurons
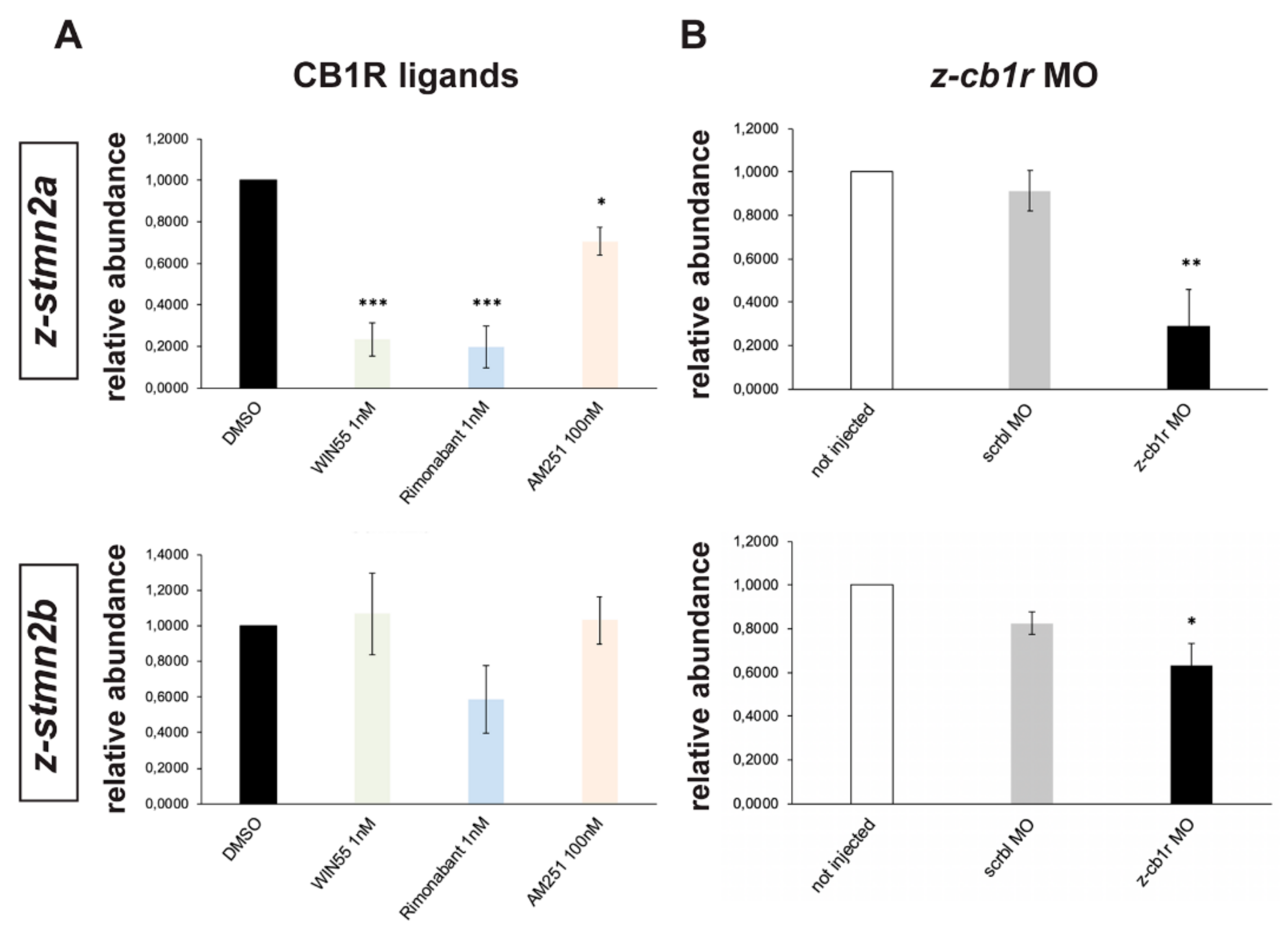
2.6. Search for Genes Functionally Related to CB1R Involved in Axon Fasciculation and Guidance
3. Discussion
4. Materials and Methods
4.1. Zebrafish Strains and Treatments
4.2. Evaluation of Survival Rate and Hatching Rate
4.3. Morpholino Injections
| z-cb1r MO: | 5′ CTAGAGGAAACCTGTCGGAGGAAAT 3′ |
| Mismatch_PPFIA scrambled MO (control): | 5′ TCGTGGCCATCAACTCGAACA 3′. |
4.4. Western Blot
4.5. Immunostaining
4.6. Bioinformatic Searches and Statistical Analyses
4.7. Real-Time qPCR Analysis
| z-β-actin | F | CCCACATAGGAGTCTTTCTG |
| R | TCCCCTTGTTCACAATAACC | |
| z-cb1r | F | CAGAACAGATCATACACCATGA |
| R | TGGTCTTATTCATCATCTACGC | |
| z-hoxb7a | F | GCTGGCGATCTCTGTAAAGC |
| R | TTTTGATGGTAGCCCCTCTG | |
| z-hoxa10b | F | GAGCTAAGGGGGTCCACTG |
| R | CACTTTTGGAATCTCCTGCTTT | |
| z-hox11a | F | CAGCAAACAGTCGACACCAC |
| R | CGGTCGCTCCTTTCCTTC | |
| z-stmn2a | F | TCCATGCTCTCGCTTATTTG |
| R | GGAGGATGTAGAGATGTGCT | |
| z-stmn2b | F | ATGGAGCAGATCAAGGAGAA |
| R | GGAGGATGTAGAGATGTGCT | |
| z-sez6a | F | AGTTTAGCAGTGAACGAACC |
| R | GAAGACCAGTGACTGAAACC | |
| z-sez6b | F | GTCCATTTTCATCCCTGTGG |
| R | GGCTGCTTTGAATCATAGGT | |
| z-negr1 | F | TTTTCGAGTGGTACAAGGGA |
| R | GGGTTGGATGGATTCAGAGG |
4.8. CB1 Ligands Exposure and Confocal Microscope Analysis
4.9. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ∆9-THC | Δ9-tetrahydrocannabinol |
| 2 AG | 2-arachidonoylglycerol |
| AC | anterior commissure |
| AgRP | agouti-related peptide |
| AgRP1 | agouti-related peptide 1 |
| AM251 | (N-(Piperidin-1-yl)-5-(4-iodophenyl)-1-(2,4-dichlorophenyl)-4-methyl-1H-pyrazole-3-carboxamide) |
| CB1R | cannabinoid receptor type 1 |
| CB2R | cannabinoid receptor type 2 |
| CBD | cannabidiol |
| DAGL | diacylglycerol lipase |
| DMSO | dimethyl sulfoxide |
| dpf | days post fertilization |
| eCB | endocannabinoid |
| GnRH | gonadotropin-releasing hormone |
| GTPase | guanosine triphosphatase |
| Hox | homeobox gene |
| hpf | hours post fertilization |
| JNK1 | c-Jun N-terminal kinase 1 |
| MAGL | monoacylglycerol lipase |
| MO | morpholino oligonucleotide |
| Negr1 | neuronal growth regulator 1 |
| POA | pre-optic area |
| POC | post-optic commissure |
| Rimonabant | SR141716, (N-(Piperidin-1-yl)-5-(4-chlorophenyl)-1-(2,4-dichlorophenyl)4-methyl-1H-pyrazole-3-carboxamide hydrochloride) |
| Sez6 | Seizure protein 6 |
| Stmn2/SCG10 | Stathmin2/Superior Cervical Ganglion-10 Protein |
| VTC | ventral tegmental commissure |
| WIN55 | WIN 55,212-2; (R)-( + )-[2,3-Dihydro-5-methyl-3-(4-morpholinylmethyl)pyrrolo [1,2,3-de]-1,4-benzoxazin-6-yl]-1-naphthalenylmethanone mesylate |
References
- Grotenhermen, F. Pharmacokinetics and Pharmacodynamics of Cannabinoids. Clin. Pharmacokinet. 2003, 42, 327–360. [Google Scholar] [CrossRef] [PubMed]
- Scheyer, A.F.; Melis, M.; Trezza, V.; Manzoni, O.J.J. Consequences of Perinatal Cannabis Exposure. Trends Neurosci. 2019, 42, 871–884. [Google Scholar] [CrossRef] [PubMed]
- Howlett, A.C. The cannabinoid receptors. Prostaglandins Other Lipid Mediat. 2002, 68, 619–631. [Google Scholar] [CrossRef]
- Kano, M.; Ohno-Shosaku, T.; Hashimotodani, Y.; Uchigashima, M.; Watanabe, M. Endocannabinoid-Mediated Control of Synaptic Transmission. Physiol. Rev. 2009, 89, 309–380. [Google Scholar] [CrossRef]
- Pertwee, R.G.; Howlett, A.C.; Abood, M.E.; Alexander, S.P.H.; Di Marzo, V.; Elphick, M.R.; Greasley, P.J.; Hansen, H.S.; Kunos, G.; Mackie, K.; et al. International Union of Basic and Clinical Pharmacology. LXXIX. Cannabinoid Receptors and Their Ligands: Beyond CB1 and CB2. Physiol. Rev. 2010, 62, 588–631. [Google Scholar] [CrossRef]
- Di Marzo, V. Endocannabinoid signaling in the brain: Biosynthetic mechanisms in the limelight. Nat. Neurosci. 2011, 14, 9–15. [Google Scholar] [CrossRef]
- Maccarone, M.; Guzman, M.; Mackie, K.; Doherty, P.; Harkany, T. Programming and reprogramming neural cells by (endo-)cannabinoids: From physiological rules to emerging therapies. Nat. Rev. Neurosci. 2014, 91, 165–171. [Google Scholar]
- Keimpema, E.; Barabas, K.; Morozov, Y.M.; Tortoriello, G.; Torii, M.; Cameron, G.; Yanagawa, Y.; Watanabe, M.; Mackie, K.; Harkany, T. Differential Subcellular Recruitment of Monoacylglycerol Lipase Generates Spatial Specificity of 2-Arachidonoyl Glycerol Signaling during Axonal Pathfinding. J. Neurosci. 2010, 30, 13992–14007. [Google Scholar] [CrossRef]
- Roland, A.B.; Ricobaraza, A.; Carrel, D.; Jordan, B.M.; Rico, F.; Simon, A.; Humbert-Claude, M.; Ferrier, J.; McFadden, M.H.; Scheuring, S.; et al. Cannabinoid-induced actomyosin contractility shapes neuronal morphology and growth. eLife 2014, 3, e03159. [Google Scholar] [CrossRef]
- Njoo, C.; Agarwal, N.; Lutz, B.; Kuner, R. The Cannabinoid Receptor CB1 Interacts with the WAVE1 Complex and Plays a Role in Actin Dynamics and Structural Plasticity in Neurons. PLoS Biol. 2015, 13, e1002286. [Google Scholar] [CrossRef]
- Grenningloh, G.; Soehrman, S.; Bondallaz, P.; Ruchti, E.; Cadas, H. Role of the microtubule destabilizing proteins SCG10 and stathmin in neuronal growth. J. Neurobiol. 2004, 58, 60–69. [Google Scholar] [CrossRef] [PubMed]
- Tararuk, T.; Östman, N.; Li, W.; Björkblom, B.; Padzik, A.; Zdrojewska, J.; Hongisto, V.; Herdegen, T.; Konopka, W.; Courtney, M.J.; et al. JNK1 phosphorylation of SCG10 determines microtubule dynamics and axodendritic length. J. Cell Biol. 2006, 173, 265–277. [Google Scholar] [CrossRef] [PubMed]
- Manna, T.; Grenningloh, G.; Miller, H.P.; Wilson, L. Stathmin Family Protein SCG10 Differentially Regulates the Plus and Minus End Dynamics of Microtubules at Steady State In Vitro: Implications for Its Role in Neurite Outgrowth. Biochemistry 2007, 46, 3543–3552. [Google Scholar] [CrossRef] [PubMed]
- Oudin, M.J.; Hobbs, C.; Doherty, P. DAGL-dependent endocannabinoid signalling: Roles in axonal pathfinding, synaptic plasticity and adult neurogenesis: DAGL-dependent endocannabinoid signalling. Eur. J. Neurosci. 2011, 34, 1634–1646. [Google Scholar] [CrossRef] [PubMed]
- Keimpema, E.; Mackie, K.; Harkany, T. Molecular model of cannabis sensitivity in developing neuronal circuits. Trends Pharmacol. Sci. 2011, 32, 551–561. [Google Scholar] [CrossRef] [PubMed]
- Harkany, T.; Keimpema, E.; Barabás, K.; Mulder, J. Endocannabinoid functions controlling neuronal specification during brain development. Mol. Cell. Endocrinol. 2008, 286, S84–S90. [Google Scholar] [CrossRef]
- Huizink, A.C.; Mulder, E.J.H. Maternal smoking, drinking or cannabis use during pregnancy and neurobehavioral and cognitive functioning in human offspring. Neurosci. Biobehav. Rev. 2006, 30, 24–41. [Google Scholar] [CrossRef]
- Calvigioni, D.; Hurd, Y.L.; Harkany, T.; Keimpema, E. Neuronal substrates and functional consequences of prenatal cannabis exposure. Eur. Child Adolesc. Psychiatry 2014, 23, 931–941. [Google Scholar] [CrossRef]
- Leech, S.L.; Richardson, G.A.; Goldschmidt, L.; Day, N.L. Prenatal Substance Exposure: Effects on Attention and Impulsivity of 6-year-olds. Neurotoxicol. Teratol. 1999, 21, 109–118. [Google Scholar] [CrossRef]
- Leech, S.L.; Larkby, C.A.; Day, R.; Day, N.L. Predictors and Correlates of High Levels of Depression and Anxiety Symptoms Among Children at Age 10. J. Am. Acad. Child Adolesc. Psychiatry 2006, 45, 223–230. [Google Scholar] [CrossRef]
- Hwang, H.M.; Ku, R.Y.; Hashimoto-Torii, K. Prenatal Environment That Affects Neuronal Migration. Front. Cell Dev. Biol. 2019, 7, 138. [Google Scholar] [CrossRef] [PubMed]
- Berghuis, P.; Rajnicek, A.M.; Morozov, Y.M.; Ross, R.A.; Mulder, J.; Urban, G.M.; Monory, K.; Marsicano, G.; Matteoli, M.; Canty, A.; et al. Hardwiring the Brain: Endocannabinoids Shape Neuronal Connectivity. Science 2007, 316, 1212–1216. [Google Scholar] [CrossRef] [PubMed]
- Mulder, J.; Aguado, T.; Keimpema, E.; Barabas, K.; Ballester Rosado, C.J.; Nguyen, L.; Monory, K.; Marsicano, G.; Di Marzo, V.; Hurd, Y.L.; et al. Endocannabinoid signaling controls pyramidal cell specification and long-range axon patterning. Proc. Natl. Acad. Sci. USA 2008, 105, 8760–8765. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.-S.; Zhu, J.; Wager-Miller, J.; Wang, S.; O’Leary, D.; Monory, K.; Lutz, B.; Mackie, K.; Lu, H.-C. Requirement of cannabinoid CB1 receptors in cortical pyramidal neurons for appropriate development of corticothalamic and thalamocortical projections: CB1R in the development of sensory networks. Eur. J. Neurosci. 2010, 32, 693–706. [Google Scholar] [CrossRef] [PubMed]
- Argaw, A.; Duff, G.; Zabouri, N.; Cecyre, B.; Chaine, N.; Cherif, H.; Tea, N.; Lutz, B.; Ptito, M.; Bouchard, J.-F. Concerted Action of CB1 Cannabinoid Receptor and Deleted in Colorectal Cancer in Axon Guidance. J. Neurosci. 2011, 31, 1489–1499. [Google Scholar] [CrossRef]
- Duff, G.; Argaw, A.; Cecyre, B.; Cherif, H.; Tea, N.; Zabouri, N.; Casanova, C.; Ptito, M.; Bouchard, J.-F. Cannabinoid Receptor CB2 Modulates Axon Guidance. PLoS ONE 2013, 8, e70849. [Google Scholar] [CrossRef]
- Abbas Farishta, R.; Robert, C.; Turcot, O.; Thomas, S.; Vanni, M.P.; Bouchard, J.-F.; Casanova, C. Impact of CB1 Receptor Deletion on Visual Responses and Organization of Primary Visual Cortex in Adult Mice. Investig. Opthalmol. Vis. Sci. 2015, 56, 7697. [Google Scholar] [CrossRef][Green Version]
- Diaz-Alonso, J.; Aguado, T.; Wu, C.-S.; Palazuelos, J.; Hofmann, C.; Garcez, P.; Guillemot, F.; Lu, H.-C.; Lutz, B.; Guzman, M.; et al. The CB1 Cannabinoid Receptor Drives Corticospinal Motor Neuron Differentiation through the Ctip2/Satb2 Transcriptional Regulation Axis. J. Neurosci. 2012, 32, 16651–16665. [Google Scholar] [CrossRef]
- Tortoriello, G.; Morris, C.V.; Alpar, A.; Fuzik, J.; Shirran, S.L.; Calvigioni, D.; Keimpema, E.; Botting, C.H.; Reinecke, K.; Herdegen, T.; et al. Miswiring the brain: 9-tetrahydrocannabinol disrupts cortical development by inducing an SCG10/stathmin-2 degradation pathway. EMBO J. 2014, 33, 668–685. [Google Scholar] [CrossRef]
- Watson, S.; Chambers, D.; Hobbs, C.; Doherty, P.; Graham, A. The endocannabinoid receptor, CB1, is required for normal axonal growth and fasciculation. Mol. Cell. Neurosci. 2008, 38, 89–97. [Google Scholar] [CrossRef]
- Sufian, M.S.; Amin, M.R.; Kanyo, R.; Allison, W.T.; Ali, D.W. CB1 and CB2 receptors play differential roles in early zebrafish locomotor development. J. Exp. Biol. 2019, 222, jeb206680. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, K.T.; Amin, M.R.; Shah, P.; Ali, D.W. Motor neuron development in zebrafish is altered by brief (5-hr) exposures to THC (∆9-tetrahydrocannabinol) or CBD (cannabidiol) during gastrulation. Sci. Rep. 2018, 8, 10581. [Google Scholar] [CrossRef] [PubMed]
- Martella, A.; Sepe, R.M.; Silvestri, C.; Zang, J.; Fasano, G.; Carnevali, O.; De Girolamo, P.; Neuhauss, S.C.F.; Sordino, P.; Di Marzo, V. Important role of endocannabinoid signaling in the development of functional vision and locomotion in zebrafish. FASEB J. 2016, 30, 4275–4288. [Google Scholar] [CrossRef] [PubMed]
- Tasker, J.G.; Chen, C.; Fisher, M.O.; Fu, X.; Rainville, J.R.; Weiss, G.L. Endocannabinoid Regulation of Neuroendocrine Systems. In International Review of Neurobiology; Elsevier: Amsterdam, The Netherlands, 2015; Volume 125, pp. 163–201. ISBN 978-0-12-801278-9. [Google Scholar]
- Hillard, C.J. Endocannabinoids and the Endocrine System in Health and Disease. In Endocannabinoids; Pertwee, R.G., Ed.; Springer International Publishing: Cham, Switzerland, 2015; Volume 231, pp. 317–339. ISBN 978-3-319-20824-4. [Google Scholar]
- Bovolin, P.; Cottone, E.; Pomatto, V.; Fasano, S.; Pierantoni, R.; Cobellis, G.; Meccariello, R. Endocannabinoids are Involved in Male Vertebrate Reproduction: Regulatory Mechanisms at Central and Gonadal Level. Front. Endocrinol. 2014, 5, 54. [Google Scholar] [CrossRef] [PubMed]
- Glanowska, K.M.; Moenter, S.M. Endocannabinoids and prostaglandins both contribute to GnRH neuron-GABAergic afferent local feedback circuits. J. Neurophysiol. 2011, 106, 3073–3081. [Google Scholar] [CrossRef] [PubMed]
- Farkas, I.; Kalló, I.; Deli, L.; Vida, B.; Hrabovszky, E.; Fekete, C.; Moenter, S.M.; Watanabe, M.; Liposits, Z. Retrograde Endocannabinoid Signaling Reduces GABAergic Synaptic Transmission to Gonadotropin-Releasing Hormone Neurons. Endocrinology 2010, 151, 5818–5829. [Google Scholar] [CrossRef]
- Wu, Q.; Clark, M.S.; Palmiter, R.D. Deciphering a neuronal circuit that mediates appetite. Nature 2012, 483, 594–597. [Google Scholar] [CrossRef]
- Horn, H.; Böhme, B.; Dietrich, L.; Koch, M. Endocannabinoids in Body Weight Control. Pharmaceuticals 2018, 11, 55. [Google Scholar] [CrossRef]
- Stefanucci, A.; Macedonio, G.; Dvorácskó, S.; Tömböly, C.; Mollica, A. Novel Fubinaca/Rimonabant hybrids as endocannabinoid system modulators. Amino Acids 2018, 50, 1595–1605. [Google Scholar] [CrossRef]
- Dimmit, M.P.; Stefanucci, A.; Pieretti, S.; Minosi, P.; Dvorácskó, S.; Tömböly, C.; Zengin, G.; Mollica, A. Mollica Discovery of Orexant and Anorexant Agents with Indazole Scaffold Endowed with Peripheral Antiedema Activity. Biomolecules 2019, 9, 492. [Google Scholar] [CrossRef]
- Morozov, Y.M.; Koch, M.; Rakic, P.; Horvath, T.L. Cannabinoid type 1 receptor-containing axons innervate NPY/AgRP neurons in the mouse arcuate nucleus. Mol. Metab. 2017, 6, 374–381. [Google Scholar] [CrossRef] [PubMed]
- Manfredi-Lozano, M.; Roa, J.; Tena-Sempere, M. Connecting metabolism and gonadal function: Novel central neuropeptide pathways involved in the metabolic control of puberty and fertility. Front. Neuroendocrinol. 2018, 48, 37–49. [Google Scholar] [CrossRef] [PubMed]
- Abraham, E.; Palevitch, O.; Gothilf, Y.; Zohar, Y. The zebrafish as a model system for forebrain GnRH neuronal development. Gen. Comp. Endocrinol. 2009, 164, 151–160. [Google Scholar] [CrossRef] [PubMed]
- Abraham, E.; Palevitch, O.; Gothilf, Y.; Zohar, Y. Targeted Gonadotropin-Releasing Hormone-3 Neuron Ablation in Zebrafish: Effects on Neurogenesis, Neuronal Migration, and Reproduction. Endocrinology 2010, 151, 332–340. [Google Scholar] [CrossRef]
- Shainer, I.; Buchshtab, A.; Hawkins, T.A.; Wilson, S.W.; Cone, R.D.; Gothilf, Y. Novel hypophysiotropic AgRP2 neurons and pineal cells revealed by BAC transgenesis in zebrafish. Sci. Rep. 2017, 7, 44777. [Google Scholar] [CrossRef]
- Antonelli, T.; Tanganelli, S.; Tomasini, M.C.; Finetti, S.; Trabace, L.; Steardo, L.; Sabino, V.; Carratu, M.R.; Cuomo, V.; Ferraro, L. Long-term effects on cortical glutamate release induced by prenatal exposure to the cannabinoid receptor agonist (r)-(+)-[2,3-dihydro-5-methyl-3-(4-morpholinyl-methyl) pyrrolo [1,2,3-de]-1,4-benzoxazin-6-yl]-1-naphthalenylmethanone: An in vivo microdialysis study in the awake rat. Neuroscience 2004, 124, 367–375. [Google Scholar]
- Berghuis, P.; Dobszay, M.B.; Wang, X.; Spano, S.; Ledda, F.; Sousa, K.M.; Schulte, G.; Ernfors, P.; Mackie, K.; Paratcha, G.; et al. Endocannabinoids regulate interneuron migration and morphogenesis by transactivating the TrkB receptor. Proc. Natl. Acad. Sci. USA 2005, 102, 19115–19120. [Google Scholar] [CrossRef]
- Alpár, A.; Tortoriello, G.; Calvigioni, D.; Niphakis, M.J.; Milenkovic, I.; Bakker, J.; Cameron, G.A.; Hanics, J.; Morris, C.V.; Fuzik, J.; et al. Endocannabinoids modulate cortical development by configuring Slit2/Robo1 signalling. Nat. Commun. 2014, 5, 4421. [Google Scholar] [CrossRef]
- Garaffo, G.; Conte, D.; Provero, P.; Tomaiuolo, D.; Luo, Z.; Pinciroli, P.; Peano, C.; D’Atri, I.; Gitton, Y.; Etzion, T.; et al. The Dlx5 and Foxg1 transcription factors, linked via miRNA-9 and-200, are required for the development of the olfactory and GnRH system. Mol. Cell. Neurosci. 2015, 68, 103–119. [Google Scholar] [CrossRef]
- Barresi, M.J.F. Hedgehog regulated Slit expression determines commissure and glial cell position in the zebrafish forebrain. Development 2005, 132, 3643–3656. [Google Scholar] [CrossRef]
- Shainer, I.; Michel, M.; Marquart, G.D.; Bhandiwad, A.A.; Zmora, N.; Ben-Moshe Livne, Z.; Zohar, Y.; Hazak, A.; Mazon, Y.; Förster, D.; et al. Agouti-Related Protein 2 Is a New Player in the Teleost Stress Response System. Curr. Biol. 2019, 29, e7. [Google Scholar] [CrossRef] [PubMed]
- Benowitz, L.I.; Routtenberg, A. GAP-43: An intrinsic determinant of neuronal development and plasticity. Trends Neurosci. 1997, 20, 84–91. [Google Scholar] [CrossRef]
- Bromberg, K.D.; Iyengar, R.; He, J.C. Regulation of neurite outgrowth by G(i/o) signaling pathways. Front. Biosci. 2008, 13, 4544–4557. [Google Scholar] [CrossRef] [PubMed]
- Wiencken-Barger, A.E.; Mavity-Hudson, J.; Bartsch, U.; Schachner, M.; Casagrande, V.A. The role of L1 in axon pathfinding and fasciculation. Cereb. Cortex 2004, 14, 121–131. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Perron, U.; Provero, P.; Molineris, I. In silico prediction of lncRNA function using tissue specific and evolutionary conserved expression. BMC Bioinform. 2017, 18, 144. [Google Scholar] [CrossRef] [PubMed]
- Ala, U.; Piro, R.M.; Grassi, E.; Damasco, C.; Silengo, L.; Oti, M.; Provero, P.; Di Cunto, F. Prediction of Human Disease Genes by Human-Mouse Conserved Coexpression Analysis. PLoS Comput. Biol. 2008, 4, e1000043. [Google Scholar] [CrossRef] [PubMed]
- Shimogori, T.; Lee, D.A.; Miranda-Angulo, A.; Yang, Y.; Wang, H.; Jiang, L.; Yoshida, A.C.; Kataoka, A.; Mashiko, H.; Avetisyan, M.; et al. A genomic atlas of mouse hypothalamic development. Nat. Neurosci. 2010, 13, 767–775. [Google Scholar] [CrossRef]
- Giampietro, C.; Luzzati, F.; Gambarotta, G.; Giacobini, P.; Boda, E.; Fasolo, A.; Perroteau, I. Stathmin Expression Modulates Migratory Properties of GN-11 Neurons in Vitro. Endocrinology 2005, 146, 1825–1834. [Google Scholar] [CrossRef][Green Version]
- Szczurkowska, J.; Pischedda, F.; Manago, F.; Haas, C.; Papaleo, F.; Schäfer, M.; Piccoli, G.; Cancedda, L. Negr1 is required for transition of migrating pyramidal neurons from layer V to layer II/III of the mouse cerebral cortex. Int. J. Dev. Neurosci. 2015, 47, 16. [Google Scholar] [CrossRef]
- Kim, M.H.; Gunnersen, J.M.; Tan, S.-S. Localized expression of the seizure-related gene SEZ-6 in developing and adult forebrains. Mech. Dev. 2002, 118, 171–174. [Google Scholar] [CrossRef]
- Osaki, G.; Mitsui, S.; Yuri, K. The distribution of the seizure-related gene 6 (Sez-6) protein during postnatal development of the mouse forebrain suggests multiple functions for this protein: An analysis using a new antibody. Brain Res. 2011, 1386, 58–69. [Google Scholar] [CrossRef] [PubMed]
- Leyva-Díaz, E.; López-Bendito, G. In and out from the cortex: Development of major forebrain connections. Neuroscience 2013, 254, 26–44. [Google Scholar] [CrossRef] [PubMed]
- Stoeckli, E.T. Understanding axon guidance: Are we nearly there yet? Development 2018, 145, dev151415. [Google Scholar] [CrossRef] [PubMed]
- Vitalis, T.; Lainé, J.; Simon, A.; Roland, A.; Leterrier, C.; Lenkei, Z. The type 1 cannabinoid receptor is highly expressed in embryonic cortical projection neurons and negatively regulates neurite growth in Vitro. Eur. J. Neurosci. 2008, 28, 1705–1718. [Google Scholar] [CrossRef] [PubMed]
- Keimpema, E.; Tortoriello, G.; Alpar, A.; Capsoni, S.; Arisi, I.; Calvigioni, D.; Hu, S.S.-J.; Cattaneo, A.; Doherty, P.; Mackie, K.; et al. Nerve growth factor scales endocannabinoid signaling by regulating monoacylglycerol lipase turnover in developing cholinergic neurons. Proc. Natl. Acad. Sci. USA 2013, 110, 1935–1940. [Google Scholar] [CrossRef] [PubMed]
- De Salas-Quiroga, A.; Díaz-Alonso, J.; García-Rincón, D.; Remmers, F.; Vega, D.; Gómez-Cañas, M.; Lutz, B.; Guzmán, M.; Galve-Roperh, I. Prenatal exposure to cannabinoids evokes long-lasting functional alterations by targeting CB1 receptors on developing cortical neurons. Proc. Natl. Acad. Sci. USA 2015, 112, 13693–13698. [Google Scholar] [CrossRef]
- Hedrich, J.; Angamo, E.A.; Conrad, A.; Lutz, B.; Luhmann, H.J. Cell type specific impact of cannabinoid receptor signaling in somatosensory barrel map formation in mice. J. Comp. Neurol. 2020, 528, 7–17. [Google Scholar] [CrossRef]
- Migliarini, B.; Carnevali, O. A novel role for the endocannabinoid system during zebrafish development. Mol. Cell. Endocrinol. 2009, 299, 172–177. [Google Scholar] [CrossRef]
- Abraham, E.; Palevitch, O.; Ijiri, S.; Du, S.J.; Gothilf, Y.; Zohar, Y. Early Development of Forebrain Gonadotrophin-Releasing Hormone (GnRH) Neurones and the Role of GnRH as an Autocrine Migration Factor. J. Neuroendocrinol. 2008, 20, 394–405. [Google Scholar] [CrossRef]
- Díaz-Alonso, J.; de Salas-Quiroga, A.; Paraíso-Luna, J.; García-Rincón, D.; Garcez, P.P.; Parsons, M.; Andradas, C.; Sánchez, C.; Guillemot, F.; Guzmán, M.; et al. Loss of Cannabinoid CB1 Receptors Induces Cortical Migration Malformations and Increases Seizure Susceptibility. Cereb. Cortex 2017, 27, 5303–5317. [Google Scholar]
- Williams, E.-J.; Walsh, F.S.; Doherty, P. The FGF receptor uses the endocannabinoid signaling system to couple to an axonal growth response. J. Cell Biol. 2003, 160, 481–486. [Google Scholar] [CrossRef] [PubMed]
- Gomez, T.M.; Letourneau, P.C. Actin dynamics in growth cone motility and navigation. J. Neurochem. 2014, 129, 221–234. [Google Scholar] [CrossRef] [PubMed]
- Brents, L.K. Marijuana, the Endocannabinoid System and the Female Reproductive System. Yale J. Biol. Med. 2016, 89, 175–191. [Google Scholar] [PubMed]
- Koch, M. Cannabinoid Receptor Signaling in Central Regulation of Feeding Behavior: A Mini-Review. Front. Neurosci. 2017, 11, 293. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zuccarini, G.; D’Atri, I.; Cottone, E.; Mackie, K.; Shainer, I.; Gothilf, Y.; Provero, P.; Bovolin, P.; Merlo, G.R. Interference with the Cannabinoid Receptor CB1R Results in Miswiring of GnRH3 and AgRP1 Axons in Zebrafish Embryos. Int. J. Mol. Sci. 2020, 21, 168. https://doi.org/10.3390/ijms21010168
Zuccarini G, D’Atri I, Cottone E, Mackie K, Shainer I, Gothilf Y, Provero P, Bovolin P, Merlo GR. Interference with the Cannabinoid Receptor CB1R Results in Miswiring of GnRH3 and AgRP1 Axons in Zebrafish Embryos. International Journal of Molecular Sciences. 2020; 21(1):168. https://doi.org/10.3390/ijms21010168
Chicago/Turabian StyleZuccarini, Giulia, Ilaria D’Atri, Erika Cottone, Ken Mackie, Inbal Shainer, Yoav Gothilf, Paolo Provero, Patrizia Bovolin, and Giorgio Roberto Merlo. 2020. "Interference with the Cannabinoid Receptor CB1R Results in Miswiring of GnRH3 and AgRP1 Axons in Zebrafish Embryos" International Journal of Molecular Sciences 21, no. 1: 168. https://doi.org/10.3390/ijms21010168
APA StyleZuccarini, G., D’Atri, I., Cottone, E., Mackie, K., Shainer, I., Gothilf, Y., Provero, P., Bovolin, P., & Merlo, G. R. (2020). Interference with the Cannabinoid Receptor CB1R Results in Miswiring of GnRH3 and AgRP1 Axons in Zebrafish Embryos. International Journal of Molecular Sciences, 21(1), 168. https://doi.org/10.3390/ijms21010168





