Adaptive Changes of Glioblastoma Cells Following Exposure to Hypoxic (1% Oxygen) Tumour Microenvironment
Abstract
1. Introduction
2. Results
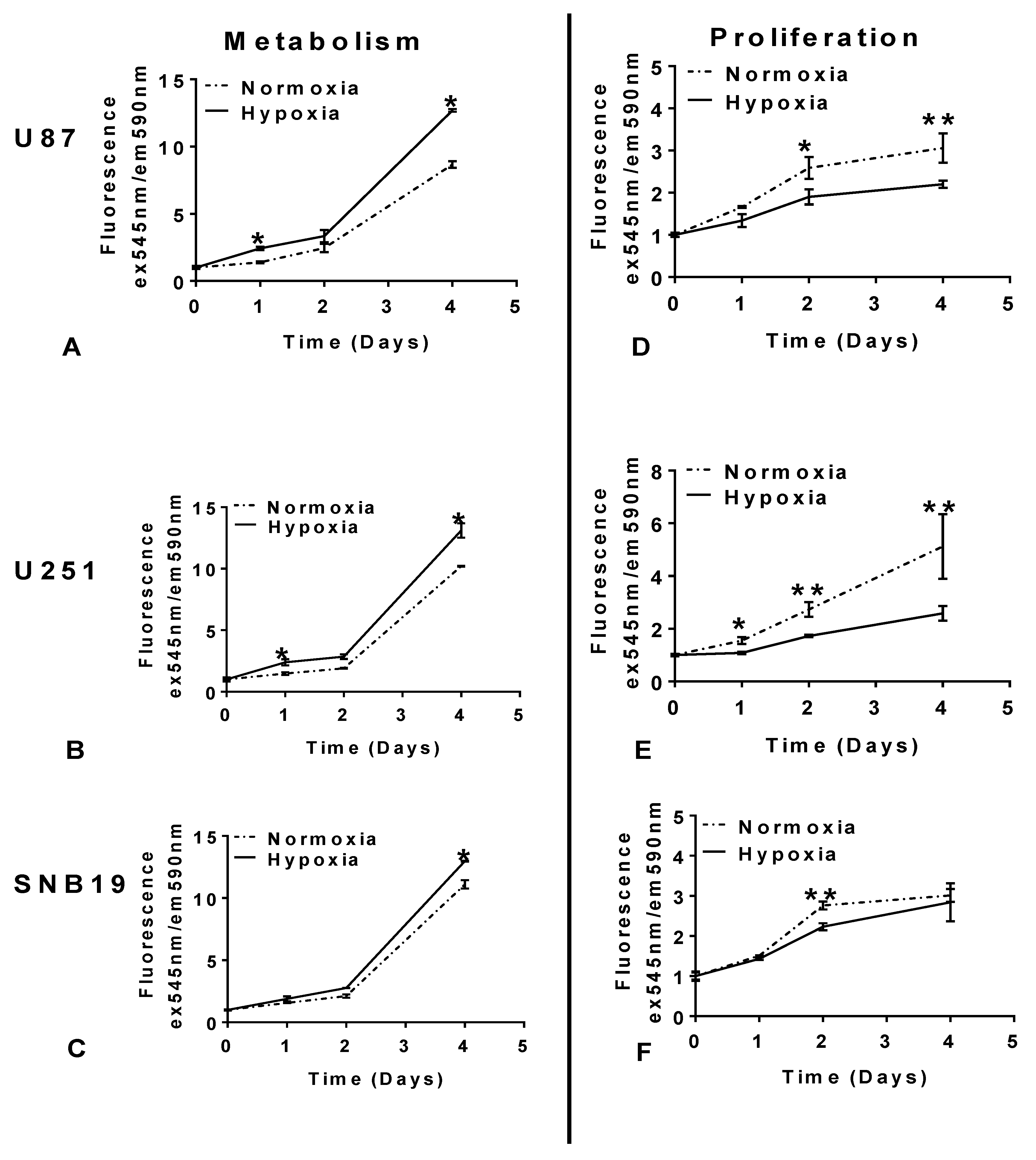
2.1. Glioblastoma Cells Adapt to Hypoxia (1% Oxygen) Tumour Microenvironment by Reducing Proliferation and Enhancing Metabolism
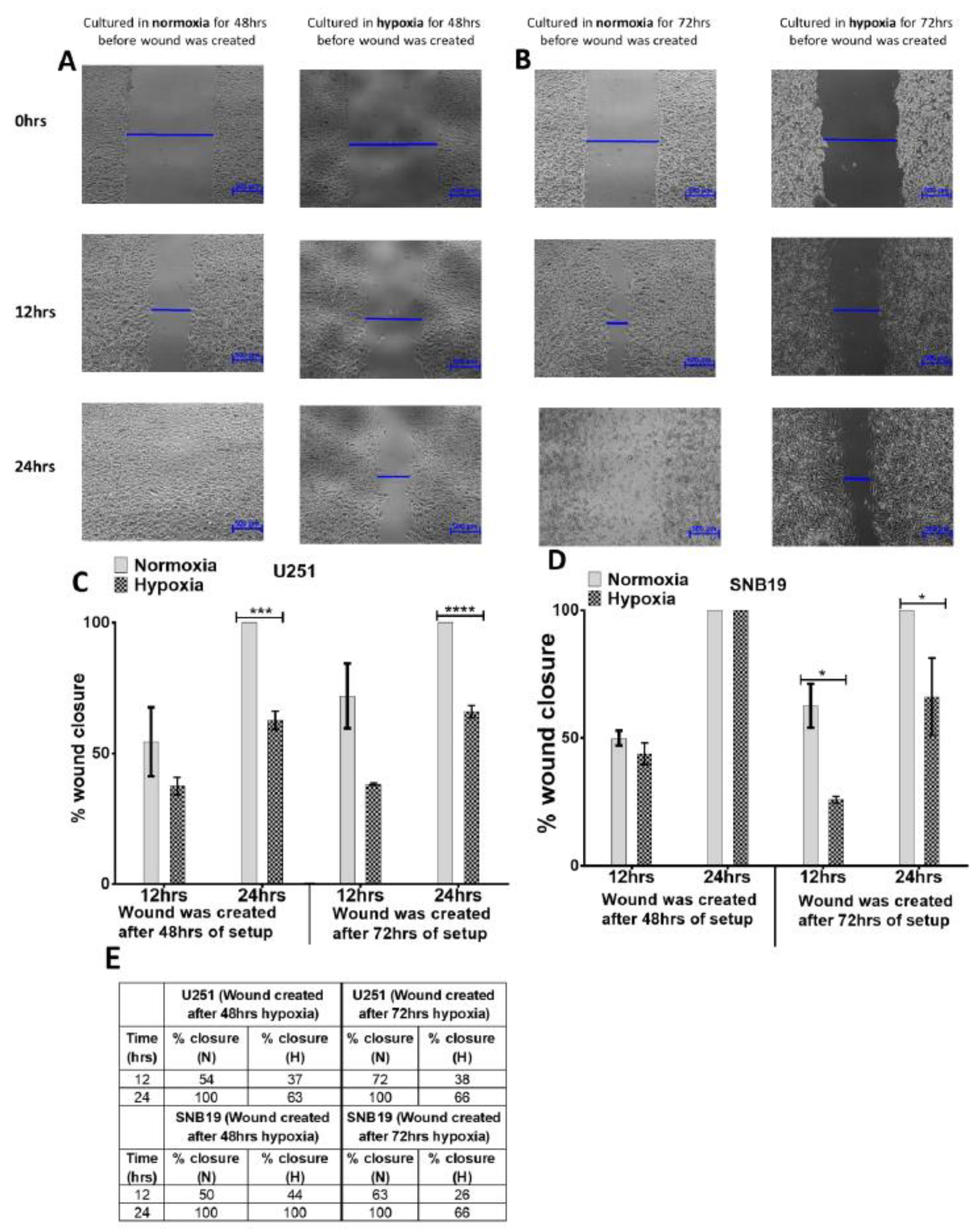
2.2. Glioblastoma Cells Adapt to Hypoxia (1% Oxygen) Tumour Microenvironment by Inhibiting Migration
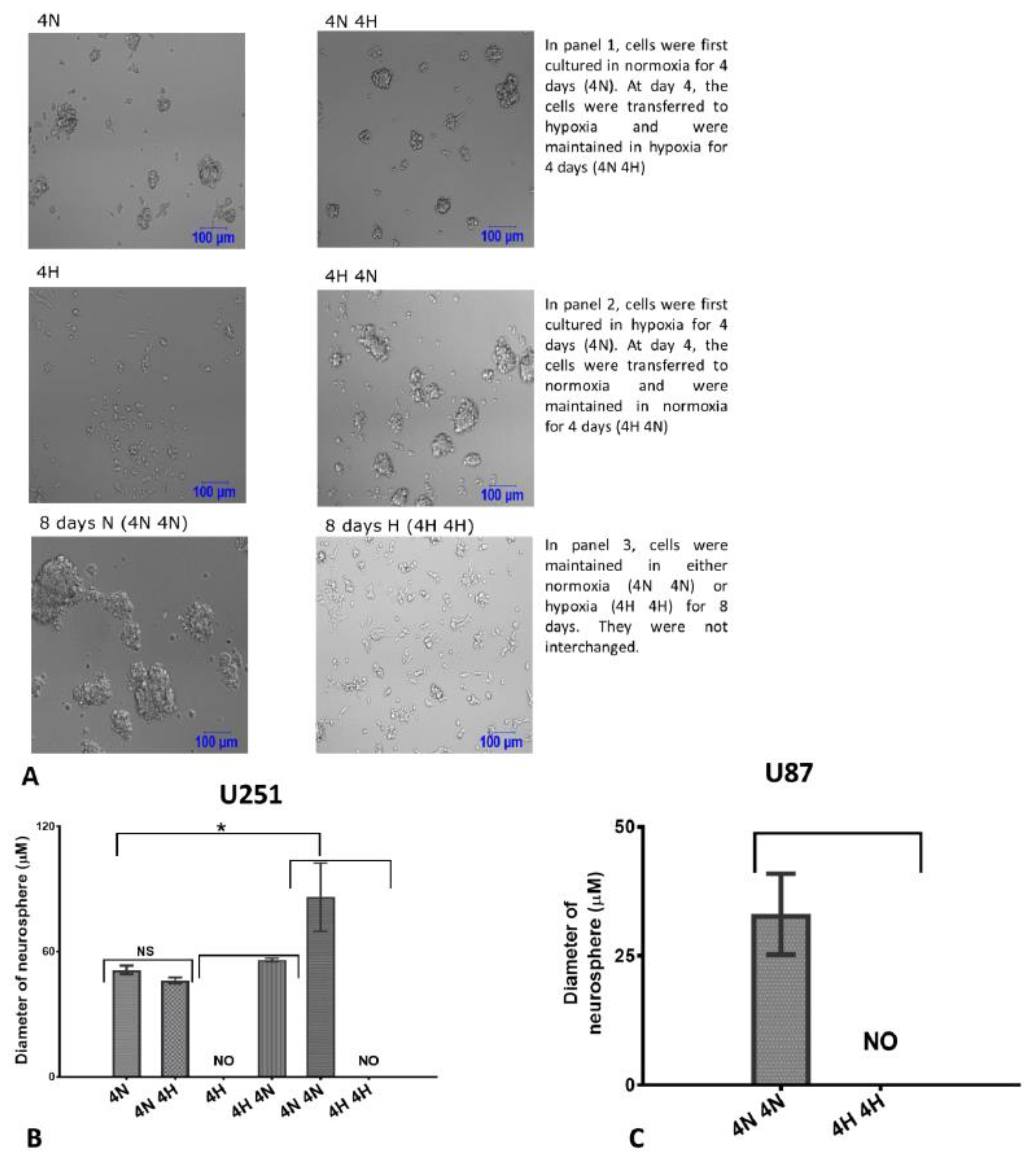
2.3. Glioblastoma Cells Adapt to Hypoxia (1% Oxygen) Tumour Microenvironment by Limiting Neurosphere Growth
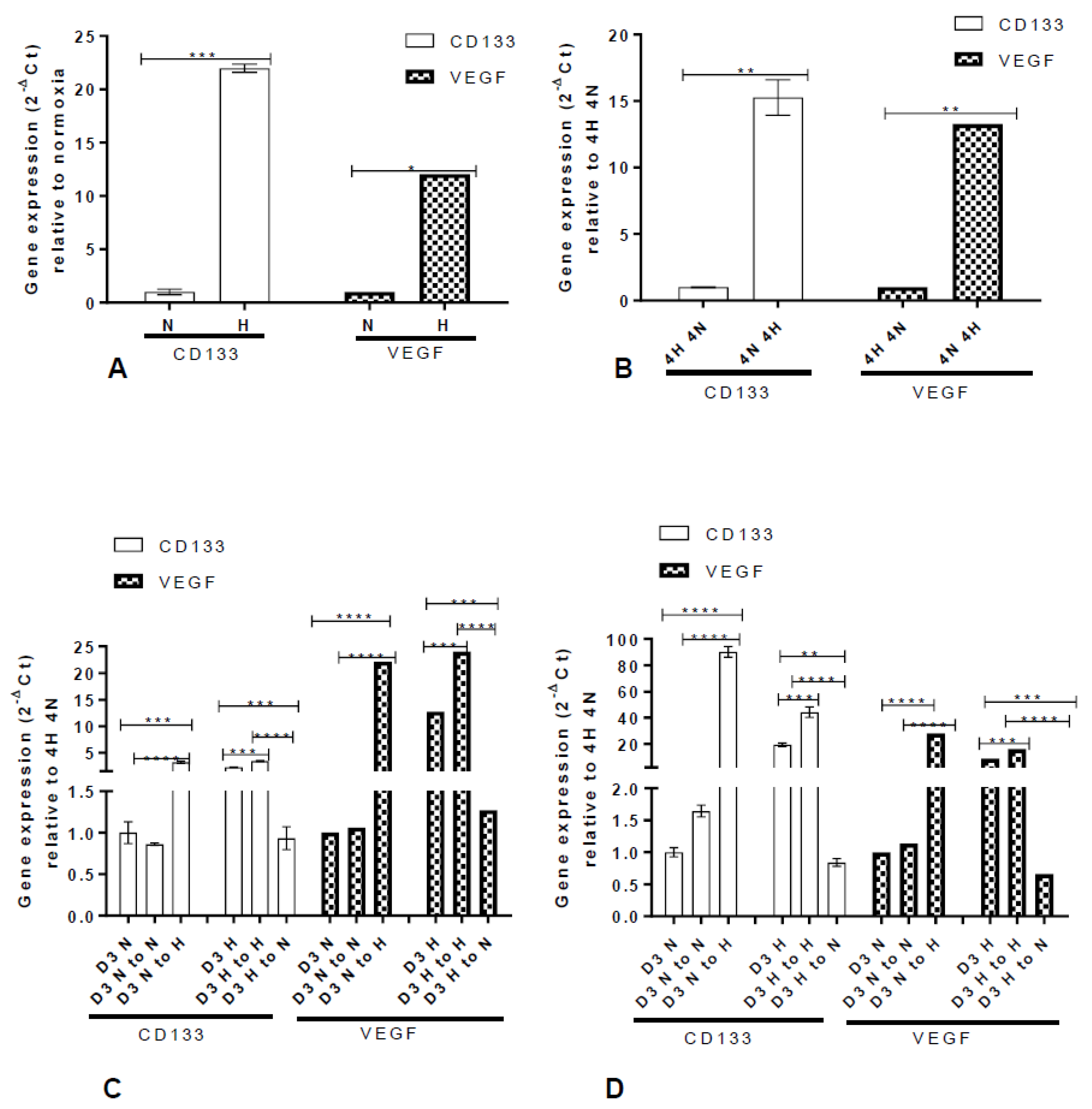
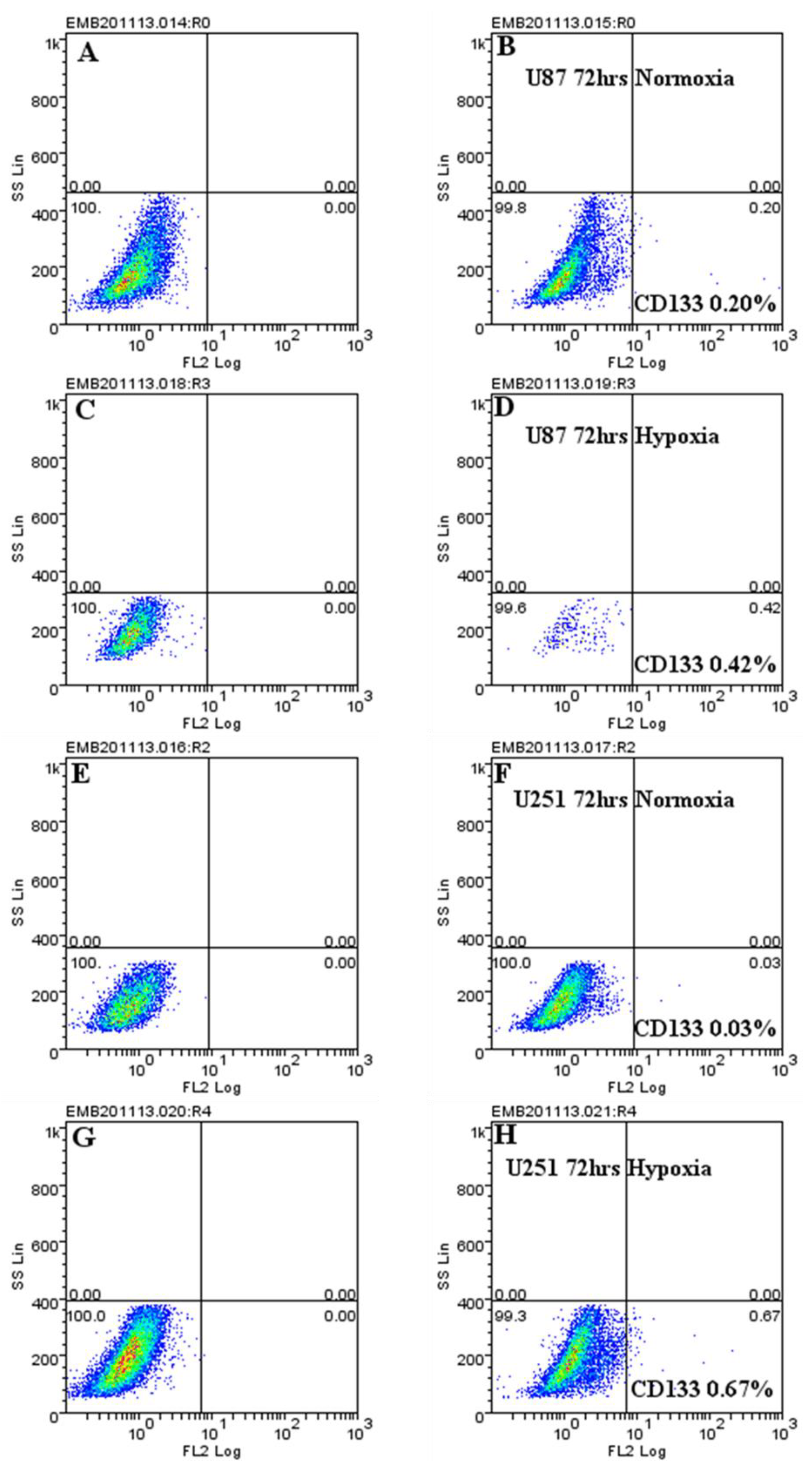
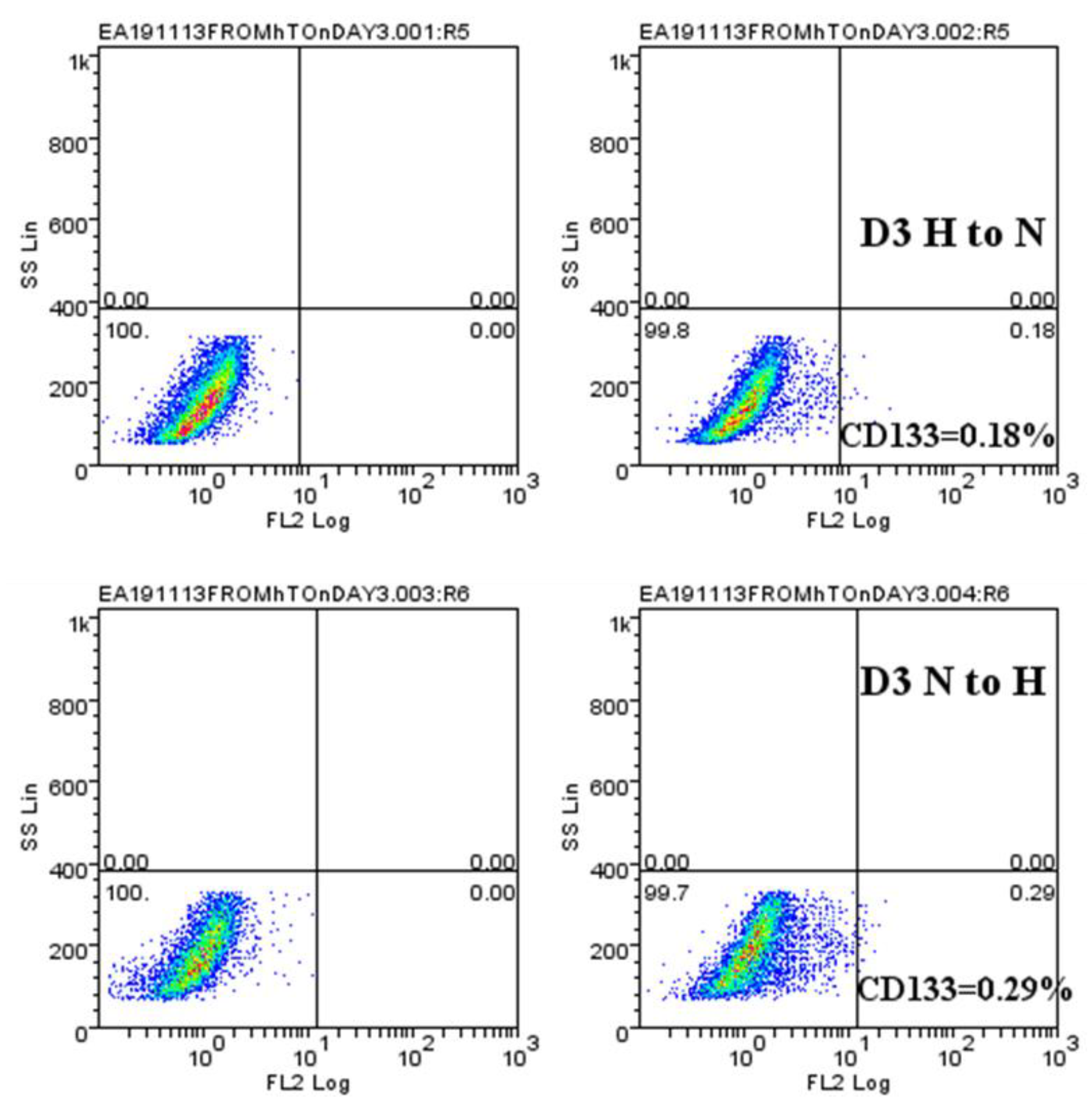
2.4. Hypoxic-Mediated Upregulation of CD133 is Reversible
3. Discussion
4. Material and Methods
4.1. Cell-Lines
4.2. Neurosphere Culture
4.3. Alamar Blue and CyQUANT Assays
4.4. Flow Cytometry
4.5. The Culture of Cells in a Hypoxia Chamber
4.6. Scratch Assay (Wound Healing Assay)
4.7. Quantitative Real-Time PCR
4.8. Statistics
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gilbert, M.R.; Wang, M.; Aldape, K.D.; Stupp, R.; Hegi, M.E.; Jaeckle, K.A.; Armstrong, T.S.; Wefel, J.S.; Won, M.; Blumenthal, D.T. Dose-dense temozolomide for newly diagnosed glioblastoma: A randomized phase III clinical trial. J. Clin. Oncol. 2013, 31, 4085. [Google Scholar] [CrossRef]
- Wang, M.; Dignam, J.J.; Won, M.; Curran, W.; Mehta, M.; Gilbert, M.R. Variation over time and interdependence between disease progression and death among patients with glioblastoma on RTOG 0525. Neuro-Oncology 2015, 17, 999–1006. [Google Scholar] [CrossRef]
- Stupp, R.; Mason, W.P.; Van Den Bent, M.J.; Weller, M.; Fisher, B.; Taphoorn, M.J.; Belanger, K.; Brandes, A.A.; Marosi, C.; Bogdahn, U. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. New Engl. J. Med. 2005, 352, 987–996. [Google Scholar] [CrossRef]
- Stupp, R.; Hegi, M.E.; Mason, W.P.; van den Bent, M.J.; Taphoorn, M.J.; Janzer, R.C.; Ludwin, S.K.; Allgeier, A.; Fisher, B.; Belanger, K. Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. Lancet Oncol. 2009, 10, 459–466. [Google Scholar] [CrossRef]
- Balkwill, F.R.; Capasso, M.; Hagemann, T. The Tumor Microenvironment at a Glance; The Company of Biologists Ltd.: Cambridge, UK, 2012. [Google Scholar]
- Brown, J.M.; Wilson, W.R. Exploiting tumour hypoxia in cancer treatment. Nat. Rev. Cancer 2004, 4, 437. [Google Scholar] [CrossRef] [PubMed]
- Guerrero-Cázares, H.; Chen, L.; Quiñones-Hinojosa, A. Glioblastoma heterogeneity and more accurate representation in research models. World Neurosurg. 2012, 78, 594–596. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Liu, Y.; Xu, X.; Yin, L.; Zhang, X.; Li, L.; Lu, H. Relationship between glioblastoma heterogeneity and survival time: An MR imaging texture analysis. Am. J. Neuroradiol. 2017, 38, 1695–1701. [Google Scholar] [CrossRef] [PubMed]
- Spence, A.M.; Muzi, M.; Swanson, K.R.; O’Sullivan, F.; Rockhill, J.K.; Rajendran, J.G.; Adamsen, T.C.; Link, J.M.; Swanson, P.E.; Yagle, K.J. Regional hypoxia in glioblastoma multiforme quantified with [18F] fluoromisonidazole positron emission tomography before radiotherapy: Correlation with time to progression and survival. Clin. Cancer Res. 2008, 14, 2623–2630. [Google Scholar] [CrossRef]
- Seidel, S.; Garvalov, B.K.; Wirta, V.; von Stechow, L.; Schänzer, A.; Meletis, K.; Wolter, M.; Sommerlad, D.; Henze, A.-T.; Nister, M. A hypoxic niche regulates glioblastoma stem cells through hypoxia inducible factor 2α. Brain 2010, 133, 983–995. [Google Scholar] [CrossRef] [PubMed]
- Piccirillo, S.; Combi, R.; Cajola, L.; Patrizi, A.; Redaelli, S.; Bentivegna, A.; Baronchelli, S.; Maira, G.; Pollo, B.; Mangiola, A. Distinct pools of cancer stem-like cells coexist within human glioblastomas and display different tumorigenicity and independent genomic evolution. Oncogene 2009, 28, 1807. [Google Scholar] [CrossRef] [PubMed]
- Beig, N.; Patel, J.; Prasanna, P.; Hill, V.; Gupta, A.; Correa, R.; Bera, K.; Singh, S.; Partovi, S.; Varadan, V. Radiogenomic analysis of hypoxia pathway is predictive of overall survival in Glioblastoma. Sci. Rep. 2018, 8, 7. [Google Scholar] [CrossRef]
- Ivanovic, Z. Hypoxia or in situ normoxia: The stem cell paradigm. J. Cell. Physiol. 2009, 219, 271–275. [Google Scholar] [CrossRef]
- Evans, S.M.; Judy, K.D.; Dunphy, I.; Jenkins, W.T.; Hwang, W.-T.; Nelson, P.T.; Lustig, R.A.; Jenkins, K.; Magarelli, D.P.; Hahn, S.M. Hypoxia is important in the biology and aggression of human glial brain tumors. Clin. Cancer Res. 2004, 10, 8177–8184. [Google Scholar] [CrossRef]
- Dings, J.; Meixensberger, J.; Jäger, A.; Roosen, K. Clinical experience with 118 brain tissue oxygen partial pressure catheter probes. Neurosurgery 1998, 43, 1082–1094. [Google Scholar] [CrossRef]
- Li, J.Z.; Gao, W.; Chan, J.Y.-W.; Ho, W.-K.; Wong, T.-S. Hypoxia in head and neck squamous cell carcinoma. Isrn Otolaryngol. 2012, 2012, 708974. [Google Scholar] [CrossRef]
- Kawai, N.; Lin, W.; Cao, W.-D.; Ogawa, D.; Miyake, K.; Haba, R.; Maeda, Y.; Yamamoto, Y.; Nishiyama, Y.; Tamiya, T. Correlation between 18F-fluoromisonidazole PET and expression of HIF-1α and VEGF in newly diagnosed and recurrent malignant gliomas. Eur. J. Nucl. Med. Mol. Imaging 2014, 41, 1870–1878. [Google Scholar] [CrossRef]
- Hashimoto, O.; Shimizu, K.; Semba, S.; Chiba, S.; Ku, Y.; Yokozaki, H.; Hori, Y. Hypoxia induces tumor aggressiveness and the expansion of CD133-positive cells in a hypoxia-inducible factor-1α-dependent manner in pancreatic cancer cells. Pathobiology 2011, 78, 181–192. [Google Scholar] [CrossRef]
- Iida, H.; Suzuki, M.; Goitsuka, R.; Ueno, H. Hypoxia induces CD133 expression in human lung cancer cells by up-regulation of OCT3/4 and SOX2. Int. J. Oncol. 2012, 40, 71–79. [Google Scholar]
- Ahmed, E.M.; Bandopadhyay, G.; Coyle, B.; Grabowska, A. A HIF-independent, CD133-mediated mechanism of cisplatin resistance in glioblastoma cells. Cell. Oncol. 2018, 41, 319–328. [Google Scholar] [CrossRef]
- Grimes, D.R.; Macauley, R.J.; Currell, F.; Scott, J.; Basanta, D. Hypoxia increases the tempo of evolution in the peri-necrotic niche in glioblastoma. bioRxiv 2018, 293712. [Google Scholar] [CrossRef]
- Sforna, L.; Cenciarini, M.; Belia, S.; D’Adamo, M.C.; Pessia, M.; Franciolini, F.; Catacuzzeno, L. The role of ion channels in the hypoxia-induced aggressiveness of glioblastoma. Front. Cell. Neurosci. 2014, 8, 467. [Google Scholar] [CrossRef]
- Joseph, J.V.; Conroy, S.; Pavlov, K.; Sontakke, P.; Tomar, T.; Eggens-Meijer, E.; Balasubramaniyan, V.; Wagemakers, M.; den Dunnen, W.F.; Kruyt, F.A. Hypoxia enhances migration and invasion in glioblastoma by promoting a mesenchymal shift mediated by the HIF1α-ZEB1 axis. Cancer Lett. 2015, 359, 107–116. [Google Scholar] [CrossRef] [PubMed]
- Clarke, R.H.; Moosa, S.; Anzivino, M.; Wang, Y.; Floyd, D.H.; Purow, B.W.; Lee, K.S. Sustained radiosensitization of hypoxic glioma cells after oxygen pretreatment in an animal model of glioblastoma and in vitro models of tumor hypoxia. PLoS ONE 2014, 9, e111199. [Google Scholar] [CrossRef]
- Musah-Eroje, A.; Watson, S. A novel 3D in vitro model of glioblastoma reveals resistance to temozolomide which was potentiated by hypoxia. J. Neuro-Oncol. 2019, 142, 231–240. [Google Scholar] [CrossRef] [PubMed]
- Terry, S.; Faouzi Zaarour, R.; Hassan Venkatesh, G.; Francis, A.; El-Sayed, W.; Buart, S.; Bravo, P.; Thiery, J.; Chouaib, S. Role of Hypoxic Stress in Regulating Tumor Immunogenicity, Resistance and Plasticity. Int. J. Mol. Sci. 2018, 19, 3044. [Google Scholar] [CrossRef] [PubMed]
- Monteiro, A.; Hill, R.; Pilkington, G.; Madureira, P. The role of hypoxia in glioblastoma invasion. Cells 2017, 6, 45. [Google Scholar] [CrossRef]
- Luoto, K.R.; Kumareswaran, R.; Bristow, R.G. Tumor hypoxia as a driving force in genetic instability. Genome Integr. 2013, 4, 5. [Google Scholar] [CrossRef]
- Wang, X.; Prager, B.C.; Wu, Q.; Kim, L.J.; Gimple, R.C.; Shi, Y.; Yang, K.; Morton, A.R.; Zhou, W.; Zhu, Z. Reciprocal signaling between glioblastoma stem cells and differentiated tumor cells promotes malignant progression. Cell Stem Cell 2018, 22, 514–528.e5. [Google Scholar] [CrossRef]
- Bar, E.E.; Lin, A.; Mahairaki, V.; Matsui, W.; Eberhart, C.G. Hypoxia increases the expression of stem-cell markers and promotes clonogenicity in glioblastoma neurospheres. Am. J. Pathol. 2010, 177, 1491–1502. [Google Scholar] [CrossRef] [PubMed]
- Soeda, A.; Park, M.; Lee, D.; Mintz, A.; Androutsellis-Theotokis, A.; McKay, R.; Engh, J.; Iwama, T.; Kunisada, T.; Kassam, A. Hypoxia promotes expansion of the CD133-positive glioma stem cells through activation of HIF-1α. Oncogene 2009, 28, 3949. [Google Scholar] [CrossRef] [PubMed]
- Claes, A.; Idema, A.J.; Wesseling, P. Diffuse glioma growth: A guerilla war. Acta Neuropathol. 2007, 114, 443–458. [Google Scholar] [CrossRef] [PubMed]
- Page, B.; Page, M.; Noel, C. A new fluorometric assay for cytotoxicity measurements in-vitro. Int. J. Oncol. 1993, 3, 473–476. [Google Scholar] [CrossRef] [PubMed]
- Lakowicz, J.R.; Szmacinski, H.; Nowaczyk, K.; Johnson, M.L. Fluorescence lifetime imaging of free and protein-bound NADH. Proc. Natl. Acad. Sci. USA 1992, 89, 1271–1275. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Wang, S.-Y.; Nottke, A.C.; Rocheleau, J.V.; Piston, D.W.; Goodman, R.H. Redox sensor CtBP mediates hypoxia-induced tumor cell migration. Proc. Natl. Acad. Sci. USA 2006, 103, 9029–9033. [Google Scholar] [CrossRef]
- Garofalo, O.; Cox, D.; Bachelard, H. Brain levels of NADH and NAD+ under hypoxic and hypoglycaemic conditions in vitro. J. Neurochem. 1988, 51, 172–176. [Google Scholar] [CrossRef] [PubMed]
- Santilli, G.; Lamorte, G.; Carlessi, L.; Ferrari, D.; Nodari, L.R.; Binda, E.; Delia, D.; Vescovi, A.L.; De Filippis, L. Mild hypoxia enhances proliferation and multipotency of human neural stem cells. PLoS ONE 2010, 5, e8575. [Google Scholar] [CrossRef] [PubMed]
- Frezza, C.; Zheng, L.; Tennant, D.A.; Papkovsky, D.B.; Hedley, B.A.; Kalna, G.; Watson, D.G.; Gottlieb, E. Metabolic profiling of hypoxic cells revealed a catabolic signature required for cell survival. PLoS ONE 2011, 6, e24411. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, T.G.; Diogo, M.M.; Fernandes-Platzgummer, A.; da Silva, C.L.; Cabral, J.M. Effect of hypoxia on proliferation and neural commitment of embryonic stem cells at different stages of pluripotency. In Proceedings of the 1st Portuguese Biomedical Engineering Meeting, Lisbon, Portugal, 1–4 March 2011; pp. 1–4. [Google Scholar]
- Richards, R.; Jenkinson, M.D.; Haylock, B.J.; See, V. Cell cycle progression in glioblastoma cells is unaffected by pathophysiological levels of hypoxia. PeerJ 2016, 4, e1755. [Google Scholar] [CrossRef]
- Holzwarth, C.; Vaegler, M.; Gieseke, F.; Pfister, S.M.; Handgretinger, R.; Kerst, G.; Müller, I. Low physiologic oxygen tensions reduce proliferation and differentiation of human multipotent mesenchymal stromal cells. BMC Cell Biol. 2010, 11, 11. [Google Scholar] [CrossRef]
- Li, P.; Zhou, C.; Xu, L.; Xiao, H. Hypoxia enhances stemness of cancer stem cells in glioblastoma: An in vitro study. Int. J. Med. Sci. 2013, 10, 399. [Google Scholar] [CrossRef]
- Jones, L.J.; Gray, M.; Yue, S.T.; Haugland, R.P.; Singer, V.L. Sensitive determination of cell number using the CyQUANT® cell proliferation assay. J. Immunol. Methods 2001, 254, 85–98. [Google Scholar] [CrossRef]
- Vogler, M.; Vogel, S.; Krull, S.; Farhat, K.; Leisering, P.; Lutz, S.; Wuertz, C.M.; Katschinski, D.M.; Zieseniss, A. Hypoxia modulates fibroblastic architecture, adhesion and migration: A role for HIF-1α in cofilin regulation and cytoplasmic actin distribution. PLoS ONE 2013, 8, e69128. [Google Scholar] [CrossRef]
- Zagzag, D.; Lukyanov, Y.; Lan, L.; Ali, M.A.; Esencay, M.; Mendez, O.; Yee, H.; Voura, E.B.; Newcomb, E.W. Hypoxia-inducible factor 1 and VEGF upregulate CXCR4 in glioblastoma: Implications for angiogenesis and glioma cell invasion. Lab. Investig. 2006, 86, 1221. [Google Scholar] [CrossRef]
- Shen, W.; Zhu, J.; Yu, Z.; Xue, Q. Effect of hypoxia on migration, invasion and adhesion to endothelium of human pulmonary adenocarcinoma A549 cells. Chin. Ger. J. Clin. Oncol. 2008, 7, 509. [Google Scholar] [CrossRef]
- Zhu, J.; Wang, K.; Li, T.; Chen, J.; Xie, D.; Chang, X.; Yao, J.; Wu, J.; Zhou, Q.; Jia, Y. Hypoxia-induced TET1 facilitates trophoblast cell migration and invasion through HIF1α signaling pathway. Sci. Rep. 2017, 7, 8077. [Google Scholar] [CrossRef]
- Guo, M.; Cai, C.; Zhao, G.; Qiu, X.; Zhao, H.; Ma, Q.; Tian, L.; Li, X.; Hu, Y.; Liao, B. Hypoxia promotes migration and induces CXCR4 expression via HIF-1α activation in human osteosarcoma. PLoS ONE 2014, 9, e90518. [Google Scholar] [CrossRef]
- Gilkes, D.M.; Semenza, G.L.; Wirtz, D. Hypoxia and the extracellular matrix: Drivers of tumour metastasis. Nat. Rev. Cancer 2014, 14, 430. [Google Scholar] [CrossRef]
- An, Q.; Han, C.; Zhou, Y.; Li, F.; Li, D.; Zhang, X.; Yu, Z.; Duan, Z.; Kan, Q. In vitro effects of mitomycin C on the proliferation of the non-small-cell lung cancer line A549. Int. J. Clin. Exp. Med. 2015, 8, 20516. [Google Scholar]
- Ali, I.; Braun, D.P. Resveratrol enhances mitomycin C-mediated suppression of human colorectal cancer cell proliferation by up-regulation of p21WAF1/CIP1. Anticancer Res. 2014, 34, 5439–5446. [Google Scholar]
- Olivotto, M.; Sbarba, P.D. Environmental restrictions within tumor ecosystems select for a convergent, hypoxia-resistant phenotype of cancer stem cells. Cell Cycle 2008, 7, 176–187. [Google Scholar] [CrossRef]
- Mohyeldin, A.; Garzón-Muvdi, T.; Quiñones-Hinojosa, A. Oxygen in stem cell biology: A critical component of the stem cell niche. Cell Stem Cell 2010, 7, 150–161. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.K.; Hawkins, C.; Clarke, I.D.; Squire, J.A.; Bayani, J.; Hide, T.; Henkelman, R.M.; Cusimano, M.D.; Dirks, P.B. Identification of human brain tumour initiating cells. Nature 2004, 432, 396. [Google Scholar] [CrossRef]
- Zhou, Y.; Zhou, Y.; Shingu, T.; Feng, L.; Chen, Z.; Ogasawara, M.; Keating, M.J.; Kondo, S.; Huang, P. Metabolic alterations in highly tumorigenic glioblastoma cells preference for hypoxia and high dependency on glycolysis. J. Biol. Chem. 2011, 286, 32843–32853. [Google Scholar] [CrossRef]
- Rosenberg, T.; Aaberg-Jessen, C.; Petterson, S.A.; Kristensen, B.W. Heterogenic expression of stem cell markers in patient-derived glioblastoma spheroid cultures exposed to long-term hypoxia. CNS Oncol. 2018, 7, CNS15. [Google Scholar] [CrossRef]
- Chung, D.; Wong, A.; Hayashi, K.; Yellowley, C. Effect of hypoxia on generation of neurospheres from adipose tissue-derived canine mesenchymal stromal cells. Vet. J. 2014, 199, 123–130. [Google Scholar] [CrossRef]
- Yoshida, Y.; Takahashi, K.; Okita, K.; Ichisaka, T.; Yamanaka, S. Hypoxia enhances the generation of induced pluripotent stem cells. Cell Stem Cell 2009, 5, 237–241. [Google Scholar] [CrossRef]
- Amberger-Murphy, V. Hypoxia helps glioma to fight therapy. Curr. Cancer Drug Targets 2009, 9, 381–390. [Google Scholar] [CrossRef] [PubMed]
- Griguer, C.E.; Oliva, C.R.; Gobin, E.; Marcorelles, P.; Benos, D.J.; Lancaster, J.R., Jr.; Gillespie, G.Y. CD133 is a marker of bioenergetic stress in human glioma. PLoS ONE 2008, 3, e3655. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Musah-Eroje, A.; Watson, S. Adaptive Changes of Glioblastoma Cells Following Exposure to Hypoxic (1% Oxygen) Tumour Microenvironment. Int. J. Mol. Sci. 2019, 20, 2091. https://doi.org/10.3390/ijms20092091
Musah-Eroje A, Watson S. Adaptive Changes of Glioblastoma Cells Following Exposure to Hypoxic (1% Oxygen) Tumour Microenvironment. International Journal of Molecular Sciences. 2019; 20(9):2091. https://doi.org/10.3390/ijms20092091
Chicago/Turabian StyleMusah-Eroje, Ahmed, and Sue Watson. 2019. "Adaptive Changes of Glioblastoma Cells Following Exposure to Hypoxic (1% Oxygen) Tumour Microenvironment" International Journal of Molecular Sciences 20, no. 9: 2091. https://doi.org/10.3390/ijms20092091
APA StyleMusah-Eroje, A., & Watson, S. (2019). Adaptive Changes of Glioblastoma Cells Following Exposure to Hypoxic (1% Oxygen) Tumour Microenvironment. International Journal of Molecular Sciences, 20(9), 2091. https://doi.org/10.3390/ijms20092091



