Extracellular Vesicles and Carried miRNAs in the Progression of Renal Cell Carcinoma
Abstract
1. Introduction
1.1. Renal Cell Carcinoma
1.2. Renal Cancer Stem Cells
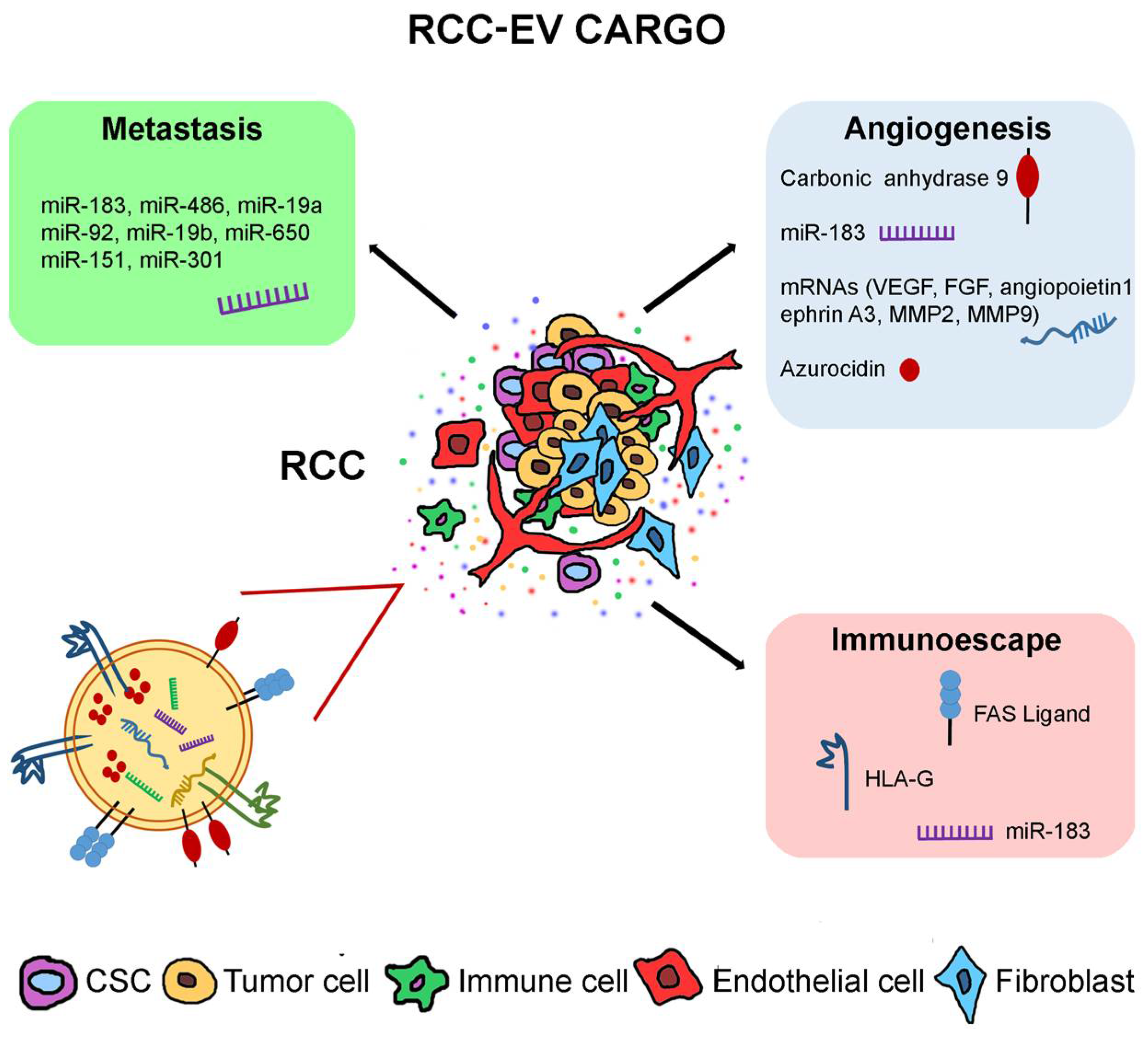
2. Effects of RCC-Derived EVs
2.1. Tumor EVs
2.2. Pro-Angiogenic Ability of RCC-EVs
2.3. Effect of RCC-EVs on Tumor Stroma
2.4. Immunosuppressive Effects of RCC-EVs
2.5. Effect of RCC-EVs on the Premetastatic Niche
2.6. RCC-EV oncomiRNAs
2.6.1. miR-183
2.6.2. miR-486
2.6.3. miR-92, miR-19a and miR-19b
2.6.4. miR-650
2.6.5. miR-301
2.7. Long Non-Coding RNAs in RCC-EVs
3. Circulating miRNAs in RCC
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Petejova, N.; Martinek, A. Renal cell carcinoma: Review of etiology, pathophysiology and risk factors. Biomed. Pap. Med. Fac. Univ. Palacky Olomouc Czech Repub. 2016, 160, 183–194. [Google Scholar] [CrossRef] [PubMed]
- Williamson, S.R.; Taneja, K.; Cheng, L. Renal cell carcinoma staging: Pitfalls, challenges and updates. Histopathology 2019, 74, 18–30. [Google Scholar] [CrossRef]
- Taneja, K.; Williamson, S.R. Updates in Pathologic Staging and Histologic Grading of Renal Cell Carcinoma. Surg. Pathol. Clin. 2018, 11, 797–812. [Google Scholar] [CrossRef] [PubMed]
- Znaor, A.; Lortet-Tieulent, J.; Laversanne, M.; Jemal, A.; Bray, F. International variations and trends in renal cell carcinoma incidence and mortality. Eur. Urol. 2015, 67, 519–530. [Google Scholar] [CrossRef]
- Rini, B.I.; Campbell, S.C.; Escudier, B. Renal cell carcinoma. Lancet. 2009, 373, 1119–1132. [Google Scholar] [CrossRef]
- Patard, J.J.; Leray, E.; Rioux-Leclercq, N.; Cindolo, L.; Ficarra, V.; Zisman, A.; De La Taille, A.; Tostain, J.; Artibani, W.; Abbou, C.C.; et al. Prognostic value of histologic subtypes in renal cell carcinoma: A multicenter experience. J. Clin. Oncol. 2005, 23, 2763–2771. [Google Scholar]
- Schmidt, L.S.; Linehan, W.M. Genetic predisposition to kidney cancer. Semin. Oncol. 2016, 43, 566–574. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.S.; Kim, J.H.; Jang, H.J.; Han, B.; Zang, D.Y. Clinicopathologic Significance of VHL Gene Alteration in Clear-Cell Renal Cell Carcinoma: An Updated Meta-Analysis and Review. Int. J. Mol. Sci. 2018, 19, 2529. [Google Scholar] [CrossRef] [PubMed]
- Maxwell, P.H.; Wiesener, M.S.; Chang, G.W.; Clifford, S.C.; Vaux, E.C.; Cockman, M.E.; Wykoff, C.C.; Pugh, C.W.; Maher, E.R.; Ratcliffe, P.J. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 1999, 399, 271–275. [Google Scholar] [CrossRef]
- Brauch, H.; Weirich, G.; Brieger, J.; Glavac, D.; Rödl, H.; Eichinger, M.; Feurer, M.; Weidt, E.; Puranakanitstha, C.; Neuhaus, C.; et al. VHL alterations in human clear cell renal cell carcinoma: Association with advanced tumor stage and a novel hot spot mutation. Cancer Res. 2000, 60, 1942–1948. [Google Scholar] [PubMed]
- Courtney, K.D.; Choueiri, T.K. Updates on novel therapies for metastatic renal cell carcinoma. Ther. Adv. Med. Oncol. 2010, 2, 209–219. [Google Scholar] [CrossRef] [PubMed]
- Gossage, L.; Murtaza, M.; Slatter, A.F.; Lichtenstein, C.P.; Warren, A.; Haynes, B.; Marass, F.; Roberts, I.; Shanahan, S.J.; Claas, A.; et al. Clinical and pathological impact of VHL, PBRM1, BAP1, SETD2, KDM6A and JARID1c in clear cell renal cell carcinoma. Genes Chromosomes Cancer 2014, 53, 38–51. [Google Scholar] [CrossRef] [PubMed]
- Rausch, S.; Schollenberger, D.; Hennenlotter, J.; Stühler, V.; Kruck, S.; Stenzl, A.; Bedke, J. mTOR and mTOR phosphorylation status in primary and metastatic renal cell carcinoma tissue: Differential expression and clinical relevance. J. Cancer Res. Clin. Oncol. 2019, 145, 153–163. [Google Scholar] [CrossRef] [PubMed]
- Grange, C.; Collino, F.; Tapparo, M.; Camussi, G. Oncogenic micro-RNAs and Renal Cell Carcinoma. Front. Oncol. 2014, 17, 49. [Google Scholar] [CrossRef] [PubMed]
- Mytsyk, Y.; Dosenko, V.; Skrzypczyk, M.A.; Borys, Y.; Diychuk, Y.; Kucher, A.; Kowalskyy, V.; Pasichnyk, S.; Mytsyk, O.; Manyuk, L. Potential clinical applications of microRNAs as biomarkers for renal cell carcinoma. Cent. Eur. J. Urol. 2018, 71, 295–303. [Google Scholar]
- Moch, H.; Montironi, R.; Lopez-Beltran, A.; Cheng, L.; Mischo, A. Oncotargets in different renal cancer subtypes. Curr. Drug Targets 2015, 16, 125–135. [Google Scholar] [CrossRef] [PubMed]
- Ahrens, M.; Scheich, S.; Hartmann, A.; Bergmann, L.; IAG-N Interdisciplinary Working Group Kidney Cancer of the German Cancer Society. Non-Clear Cell Renal Cell Carcinoma—Pathology and Treatment Options. Oncol. Res. Treat. 2019, 42, 128–135. [Google Scholar] [CrossRef]
- Barata, P.C.; Rini, B.I. Treatment of renal cell carcinoma: Current status and future directions. CA Cancer J. Clin. 2017, 67, 507–524. [Google Scholar] [CrossRef]
- Cairns, P. Renal cell carcinoma. Cancer Biomark. 2010, 9, 461–473. [Google Scholar] [CrossRef]
- Graves, A.; Hessamodini, H.; Wong, G.; Lim, W.H. Metastatic renal cell carcinoma: Update on epidemiology, genetics and therapeutic modalities. Immunotargets Ther. 2013, 2, 73–90. [Google Scholar]
- Corrò, C.; Moch, H. Biomarker discovery for renal cancer stem cells. J. Pathol. Clin. Res. 2018, 4, 3–18. [Google Scholar] [CrossRef]
- Bussolati, B.; Bruno, S.; Grange, C.; Ferrando, U.; Camussi, G. Identification of a tumor-initiating stem cell population in human renal carcinomas. FASEB J. 2008, 22, 3696–3705. [Google Scholar] [CrossRef]
- Bakhshinyan, D.; Adile, A.A.; Qazi, M.A.; Singh, M.; Kameda-Smith, M.M.; Yelle, N.; Chokshi, C.; Venugopal, C.; Singh, S.K. Introduction to Cancer Stem Cells: Past, Present and Future. Methods Mol. Biol. 2018, 1692, 1–16. [Google Scholar]
- Melzer, C.; Von Der Ohe, J.; Lehnert, H.; Ungefroren, H.; Hass, R. Cancer stem cell niche models and contribution by mesenchymal stroma/stem cells. Mol. Cancer. 2017, 16, 28. [Google Scholar] [CrossRef]
- Bussolati, B.; Dekel, B.; Azzarone, B.; Camussi, G. Human renal cancer stem cells. Cancer Lett. 2013, 338, 141–146. [Google Scholar] [CrossRef]
- Cheng, B.; Yang, G.; Jiang, R.; Cheng, Y.; Yang, H.; Pei, L.; Qiu, X. Cancer stem cell markers predict a poor prognosis in renal cell carcinoma: A meta-analysis. Oncotarget 2016, 7, 65862–65875. [Google Scholar] [CrossRef]
- Rasti, A.; Abolhasani, M.; Zanjani, L.S.; Asgari, M.; Mehrazma, M.; Madjd, Z. Reduced expression of CXCR4, a novel renal cancer stem cell marker, is associated with high-grade renal cell carcinoma. J. Cancer. Res. Clin. Oncol. 2017, 143, 95–104. [Google Scholar] [CrossRef]
- Addla, S.K.; Brown, M.D.; Hart, C.A.; Ramani, V.A.; Clarke, N.W. Characterization of the Hoechst 33342 side population from normal and malignant human renal epithelial cells. Am. J. Physiol. Ren. Physiol. 2008, 295, F680–F687. [Google Scholar] [CrossRef]
- Zhong, Y.; Guan, K.; Guo, S.; Zhou, C.; Wang, D.; Ma, W.; Zhang, Y.; Li, C.; Zhang, S. Spheres derived from the human SK-RC-42 renal cell carcinoma cell line are enriched in cancer stem cells. Cancer Lett. 2010, 299, 150–160. [Google Scholar] [CrossRef] [PubMed]
- Grange, C.; Tapparo, M.; Collino, F.; Vitillo, L.; Damasco, C.; Deregibus, M.C.; Tetta, C.; Bussolati, B.; Camussi, G. Microvesicles released from human renal cancer stem cells stimulate angiogenesis and formation of lung premetastatic niche. Cancer Res. 2011, 71, 5346–5356. [Google Scholar] [CrossRef]
- Gould, S.J.; Raposo, G. As we wait: Coping with an imperfect nomenclature for extracellular vesicles. J. Extracell. Vesicles 2013, 2, 20389. [Google Scholar] [CrossRef]
- Gai, C.; Pomatto, M.A.C.; Grange, C.; Deregibus, M.C.; Camussi, G. Extracellular vesicles in onco-nephrology. Exp. Mol. Med. 2019, 51, 29. [Google Scholar] [CrossRef]
- Kohlhapp, F.J.; Mitra, A.K.; Lengyel, E.; Peter, M.E. MicroRNAs as mediators and communicators between cancer cells and the tumor microenvironment. Oncogene 2015, 34, 5857–5868. [Google Scholar] [CrossRef] [PubMed]
- Camussi, G.; Deregibus, M.C.; Tetta, C. Tumor-derived microvesicles and the cancer microenvironment. Curr. Mol. Med. 2013, 13, 58–67. [Google Scholar] [CrossRef]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef]
- Becker, A.; Thakur, B.K.; Weiss, J.M.; Kim, H.S.; Peinado, H.; Lyden, D. Extracellular Vesicles in Cancer: Cell-to-Cell Mediators of Metastasis. Cancer Cell 2016, 30, 836–848. [Google Scholar] [CrossRef]
- Al-Nedawi, K.; Meehan, B.; Kerbel, R.S.; Allison, A.C.; Rak, J. Endothelial expression of autocrine VEGF upon the uptake of tumor-derived microvesicles containing oncogenic EGFR. Proc. Natl. Acad. Sci. USA 2009, 106, 3794–3799. [Google Scholar] [CrossRef]
- Hood, J.L.; Pan, H.; Lanza, G.M.; Wickline, S.A. Paracrine induction of endothelium by tumor exosomes. Lab. Investig. 2009, 89, 1317–1328. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Z.; Li, Y.; Pan, Y.; Lan, X.; Song, F.; Sun, J.; Zhou, K.; Liu, X.; Ren, X.; Wang, F.; et al. Cancer-derived exosomal miR-25-3p promotes pre-metastatic niche formation by inducing vascular permeability and angiogenesis. Nat. Commun. 2018, 9, 5395. [Google Scholar] [CrossRef]
- Hsu, Y.L.; Hung, J.Y.; Chang, W.A.; Lin, Y.S.; Pan, Y.C.; Tsai, P.H.; Wu, C.Y.; Kuo, P.L. Hypoxic lung cancer-secreted exosomal miR-23a increased angiogenesis and vascular permeability by targeting prolyl hydroxylase and tight junction protein ZO-1. Oncogene 2017, 36, 4929–4942. [Google Scholar] [CrossRef] [PubMed]
- Ho, A.S.; Huang, X.; Cao, H.; Christman-Skieller, C.; Bennewith, K.; Le, Q.T.; Koong, A.C. Circulating miR-210 as a Novel Hypoxia Marker in Pancreatic Cancer. Transl. Oncol. 2010, 3, 109–113. [Google Scholar]
- Jung, K.O.; Youn, H.; Lee, C.H.; Kang, K.W.; Chung, J.K. Visualization of exosome-mediated miR-210 transfer from hypoxic tumor cells. Oncotarget. 2017, 8, 9899–9910. [Google Scholar]
- Horie, K.; Kawakami, K.; Fujita, Y.; Sugaya, M.; Kameyama, K.; Mizutani, K.; Deguchi, T.; Ito, M. Exosomes expressing carbonic anhydrase 9 promote angiogenesis. Biochem. Biophys. Res. Commun. 2017, 492, 356–361. [Google Scholar]
- Svastová, E.; Hulíková, A.; Rafajová, M.; Zat’ovicová, M.; Gibadulinová, A.; Casini, A.; Cecchi, A.; Scozzafava, A.; Supuran, C.T.; Pastorek, J.; Pastoreková, S. Hypoxia activates the capacity of tumor-associated carbonic anhydrase IX to acidify extracellular pH. FEBS Lett. 2004, 577, 439–445. [Google Scholar]
- Jingushi, K.; Uemura, M.; Ohnishi, N.; Nakata, W.; Fujita, K.; Naito, T.; Fujii, R.; Saichi, N.; Nonomura, N.; Tsujikawa, K.; et al. Extracellular vesicles isolated from human renal cell carcinoma tissues disrupt vascular endothelial cell morphology via azurocidin. Int. J. Cancer. 2018, 142, 607–617. [Google Scholar]
- Lindoso, R.S.; Collino, F.; Camussi, G. Extracellular vesicles derived from renal cancer stem cells induce a pro-tumorigenic phenotype in mesenchymal stromal cells. Oncotarget 2015, 6, 7959–7969. [Google Scholar]
- Bruno, S.; Collino, F.; Iavello, A.; Camussi, G. Effects of mesenchymal stromal cell-derived extracellular vesicles on tumor growth. Front. Immunol. 2014, 5, 382. [Google Scholar]
- Egeblad, M.; Werb, Z. New functions for the matrix metalloproteinases in cancer progression. Nat. Rev. Cancer 2002, 2, 161–174. [Google Scholar]
- Park, S.A.; Ryu, C.H.; Kim, S.M.; Lim, J.Y.; Park, S.I.; Jeong, C.H.; Jun, J.A.; Oh, J.H.; Park, S.H.; Oh, W.; et al. CXCR4-transfected human umbilical cord blood-derived mesenchymal stem cells exhibit enhanced migratory capacity toward gliomas. Int. J. Oncol. 2011, 38, 97–103. [Google Scholar]
- Liu, H.; Liu, S.; Li, Y.; Wang, X.; Xue, W.; Ge, G.; Luo, X. The role of SDF-1 CXCR4/CXCR7 axis in the therapeutic effects of hypoxia-preconditioned mesenchymal stem cells for renal ischemia/reperfusion injury. PLoS ONE 2012, 7, e34608. [Google Scholar]
- Peinado, H.; Alečković, M.; Lavotshkin, S.; Matei, I.; Costa-Silva, B.; Moreno-Bueno, G.; Hergueta-Redondo, M.; Williams, C.; García-Santos, G.; Ghajar, C.; et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat. Med. 2012, 18, 883–891. [Google Scholar] [CrossRef]
- Strioga, M.; Schijns, V.; Powell, D.J., Jr.; Pasukoniene, V.; Dobrovolskiene, N.; Michalek, J. Dendritic cells and their role in tumor immunosurveillance. Innate Immun. 2013, 19, 98–111. [Google Scholar] [CrossRef]
- Teng, L.; Chen, Y.; Ding, D.; Dai, H.; Liu, G.; Li, C. Immunosuppressive effect of renal cell carcinoma on phenotype and function of dendritic cells. Int. Urol. Nephrol. 2014, 46, 915–920. [Google Scholar] [CrossRef]
- Grange, C.; Tapparo, M.; Tritta, S.; Deregibus, M.C.; Battaglia, A.; Gontero, P.; Frea, B.; Camussi, G. Role of HLA-G and extracellular vesicles in renal cancer stem cell-induced inhibition of dendritic cell differentiation. BMC Cancer. 2015, 15, 1009. [Google Scholar] [CrossRef]
- Banas, R.; Miller, C.; Guzik, L.; Zeevi, A. Amnion-derived multipotent progenitor cells inhibit blood monocyte differentiation into mature dendritic cells. Cell. Transplant. 2014, 23, 1111–1125. [Google Scholar] [CrossRef]
- Dunker, K.; Schlaf, G.; Bukur, J.; Altermann, W.W.; Handke, D.; Seliger, B. Expression and regulation of non-classical HLA-G in renal cell carcinoma. Tissue Antigens. 2008, 72, 137–148. [Google Scholar] [CrossRef]
- Yang, L.; Wu, X.; Wang, D.; Luo, C.; Chen, L. Renal carcinoma cell-derived exosomes induce human immortalized line of Jurkat T lymphocyte apoptosis in vitro. Urol. Int. 2013, 91, 363–369. [Google Scholar] [CrossRef]
- Liu, Y.; Cao, X. Characteristics and Significance of the Pre-metastatic Niche. Cancer Cell. 2016, 30, 668–681. [Google Scholar] [CrossRef]
- Ingangi, V.; Minopoli, M.; Ragone, C.; Motti, M.L.; Carriero, M.V. Role of Microenvironment on the Fate of Disseminating Cancer Stem Cells. Front. Oncol. 2019, 9, 82. [Google Scholar] [CrossRef]
- Valcz, G.; Buzás, E.I.; Szállási, Z.; Kalmár, A.; Krenács, T.; Tulassay, Z.; Igaz, P.; Molnár, B. Perspective: Bidirectional exosomal transport between cancer stem cells and their fibroblast-rich microenvironment during metastasis formation. NPJ Breast Cancer. 2018, 4, 18. [Google Scholar] [CrossRef]
- Ma, J.; Wang, P.; Liu, Y.; Zhao, L.; Li, Z.; Xue, Y. Krüppel-like factor 4 regulates blood-tumor barrier permeability via ZO-1, occludin and claudin-5. J. Cell. Physiol. 2014, 229, 916–926. [Google Scholar] [CrossRef]
- Sánchez, C.A.; Andahur, E.I.; Valenzuela, R.; Castellón, E.A.; Fullá, J.A.; Ramos, C.G.; Triviño, J.C. Exosomes from bulk and stem cells from human prostate cancer have a differential microRNA content that contributes cooperatively over local and pre-metastatic niche. Oncotarget 2016, 7, 3993–4008. [Google Scholar] [CrossRef]
- Gebeshuber, C.A.; Zatloukal, K.; Martinez, J. miR-29a suppresses tristetraprolin which is a regulator of epithelial polarity and metastasis. EMBO Rep. 2009, 10, 400–405. [Google Scholar] [CrossRef]
- Zhang, X.; Zhu, W.; Zhang, J.; Huo, S.; Zhou, L.; Gu, Z.; Zhang, M. MicroRNA-650 targets ING4 to promote gastric cancer tumorigenicity. Biochem. Biophys. Res. Commun. 2010, 395, 275–280. [Google Scholar] [CrossRef]
- Luedde, T. MicroRNA-151 and its hosting gene FAK (focal adhesion kinase) regulate tumor cell migration and spreading of hepatocellular carcinoma. Hepatology 2010, 52, 1164–1166. [Google Scholar] [CrossRef]
- Di Leva, G.; Garofalo, M.; Croce, C.M. MicroRNAs in cancer. Annu. Rev. Pathol. 2014, 9, 287–314. [Google Scholar] [CrossRef]
- Hirschberger, S.; Hinske, L.C.; Kreth, S. MiRNAs: Dynamic regulators of immune cell functions in inflammation and cancer. Cancer Lett. 2018, 431, 11–21. [Google Scholar] [CrossRef]
- Frixa, T.; Donzelli, S.; Blandino, G. Oncogenic MicroRNAs: Key Players in Malignant Transformation. Cancers 2015, 7, 2466–2485. [Google Scholar] [CrossRef]
- Zhang, B.; Pan, X.; Cobb, G.P.; Anderson, T.A. Micrornas as oncogenes and tumor suppressors. Dev. Biol. 2007, 302, 1–12. [Google Scholar] [CrossRef]
- Petrozza, V.; Carbone, A.; Bellissimo, T.; Porta, N.; Palleschi, G.; Pastore, A.L.; Di Carlo, A.; Della Rocca, C.; Fazi, F. Oncogenic MicroRNAs Characterization in Clear Cell Renal Cell Carcinoma. Int. J. Mol. Sci. 2015, 16, 29219–29225. [Google Scholar] [CrossRef]
- Qiu, M.; Liu, L.; Chen, L.; Tan, G.; Liang, Z.; Wang, K.; Liu, J.; Chen, H. microRNA-183 plays as oncogenes by increasing cell proliferation, migration and invasion via targeting protein phosphatase 2A in renal cancer cells. Biochem. Biophys. Res. Commun. 2014, 452, 163–169. [Google Scholar] [CrossRef]
- Zhang, Q.H.; Sun, H.M.; Zheng, R.Z.; Li, Y.C.; Zhang, Q.; Cheng, P.; Tang, Z.H.; Huang, F. Meta-analysis of microRNA-183 family expression in human cancer studies comparing cancer tissues with noncancerous tissues. Gene 2013, 527, 26–32. [Google Scholar] [CrossRef]
- He, Y.; Liu, J.; Wang, Y.; Zhu, X.; Fan, Z.; Li, C.; Yin, H.; Liu, Y. Role of miR-486-5p in regulating renal cell carcinoma cell proliferation and apoptosis via TGF-β-activated kinase 1. J. Cell. Biochem. 2019, 120, 2954–2963. [Google Scholar] [CrossRef]
- Goto, K.; Oue, N.; Shinmei, S.; Sentani, K.; Sakamoto, N.; Naito, Y.; Hayashi, T.; Teishima, J.; Matsubara, A.; Yasui, W. Expression of miR-486 is a potential prognostic factor after nephrectomy in advanced renal cell carcinoma. Mol. Clin. Oncol. 2013, 1, 235–240. [Google Scholar] [CrossRef][Green Version]
- Ma, Q.; Peng, Z.; Wang, L.; Li, Y.; Wang, K.; Zheng, J.; Liang, Z.; Liu, T. miR-19a correlates with poor prognosis of clear cell renal cell carcinoma patients via promoting cell proliferation and suppressing PTEN/SMAD4 expression. Int. J. Oncol. 2016, 49, 2589–2599. [Google Scholar] [CrossRef]
- Xiao, W.; Gao, Z.; Duan, Y.; Yuan, W.; Ke, Y. Downregulation of miR-19a exhibits inhibitory effects on metastatic renal cell carcinoma by targeting PIK3CA and inactivating Notch signaling in vitro. Oncol. Rep. 2015, 34, 739–746. [Google Scholar] [CrossRef]
- Niu, S.; Ma, X.; Zhang, Y.; Liu, Y.N.; Chen, X.; Gong, H.; Yao, Y.; Liu, K.; Zhang, X. MicroRNA-19a and microRNA-19b promote the malignancy of clear cell renal cell carcinoma through targeting the tumor suppressor RhoB. PLoS ONE 2018, 13, e0192790. [Google Scholar] [CrossRef]
- Valera, V.A.; Walter, B.A.; Linehan, W.M.; Merino, M.J. Regulatory Effects of microRNA-92 (miR-92) on VHL Gene Expression and the Hypoxic Activation of miR-210 in Clear Cell Renal Cell Carcinoma. Cancer 2011, 2, 515–526. [Google Scholar] [CrossRef]
- Liu, G.L.; Yang, H.J.; Liu, B.; Liu, T. Effects of MicroRNA-19b on the Proliferation, Apoptosis and Migration of Wilms’ Tumor Cells Via the PTEN/PI3K/AKT Signaling Pathway. J. Cell. Biochem. 2017, 118, 3424–3434. [Google Scholar] [CrossRef]
- Zhang, X.L.; Xu, G.; Zhou, Y.; Yan, J.J. MicroRNA-183 promotes the proliferation and metastasis of renal cell carcinoma through targeting Dickkopf-related protein 3. Oncol. Lett. 2018, 15, 6003–6008. [Google Scholar] [CrossRef]
- Li, H.; Pan, X.; Gui, Y.; Quan, J.; Li, Z.; Zhao, L.; Guan, X.; Xu, J.; Xu, W.; Lai, Y. Upregulation of miR-183-5p predicts worse survival in patients with renal cell cancer after surgery. Cancer Biomark. 2019, 24, 153–158. [Google Scholar] [CrossRef]
- Yang, Y.; Ji, C.; Guo, S.; Su, X.; Zhao, X.; Zhang, S.; Liu, G.; Qiu, X.; Zhang, Q.; Guo, H.; et al. The miR-486-5p plays a causative role in prostate cancer through negative regulation of multiple tumor suppressor pathways. Oncotarget 2017, 8, 72835–72846. [Google Scholar] [CrossRef]
- Chow, T.F.; Mankaruos, M.; Scorilas, A.; Youssef, Y.; Girgis, A.; Mossad, S.; Metias, S.; Rofael, Y.; Honey, R.J.; Stewart, R.; et al. The miR-17-92 cluster is over expressed in and has an oncogenic effect on renal cell carcinoma. J. Urol. 2010, 183, 743–751. [Google Scholar] [CrossRef]
- Fang, L.L.; Wang, X.H.; Sun, B.F.; Zhang, X.D.; Zhu, X.H.; Yu, Z.J.; Luo, H. Expression, regulation and mechanism of action of the miR-17-92 cluster in tumor cells. Int. J. Mol. Med. 2017, 40, 1624–1630. [Google Scholar] [CrossRef]
- Zuo, Z.H.; Yu, Y.P.; Ding, Y.; Liu, S.; Martin, A.; Tseng, G.; Luo, J.H. Oncogenic Activity of miR-650 in Prostate Cancer Is Mediated by Suppression of CSR1 Expression. Am. J. Pathol. 2015, 185, 1991–1999. [Google Scholar] [CrossRef]
- You, Q.; Li, H.; Liu, Y.; Xu, Y.; Miao, S.; Yao, G.; Xue, Y.; Geng, J.; Jin, X.; Meng, H. MicroRNA-650 targets inhibitor of growth 4 to promote colorectal cancer progression via mitogen activated protein kinase signaling. Oncol. Lett. 2018, 16, 2326–2334. [Google Scholar] [CrossRef]
- Zeng, Z.L.; Li, F.J.; Gao, F.; Sun, D.S.; Yao, L. Upregulation of miR-650 is correlated with downregulation of ING4 and progression of hepatocellular carcinoma. J. Surg. Oncol. 2013, 107, 105–110. [Google Scholar] [CrossRef]
- Ningning, S.; Libo, S.; Chuanbin, W.; Haijiang, S.; Qing, Z. MiR-650 regulates the proliferation, migration and invasion of human oral cancer by targeting growth factor independent 1 (Gfi1). Biochimie 2019, 156, 69–78. [Google Scholar] [CrossRef]
- Shi, W.; Gerster, K.; Alajez, N.M.; Tsang, J.; Waldron, L.; Pintilie, M.; Hui, A.B.; Sykes, J.; P’ng, C.; Miller, N.; et al. MicroRNA-301 mediates proliferation and invasion in human breast cancer. Cancer. Res. 2011, 71, 2926–2937. [Google Scholar] [CrossRef]
- Hui, A.B.; Shi, W.; Boutros, P.C.; Miller, N.; Pintilie, M.; Fyles, T.; McCready, D.; Wong, D.; Gerster, K.; Waldron, L.; et al. Robust global micro-RNA profiling with formalin-fixed paraffin-embedded breast cancer tissues. Lab. Investig. 2009, 89, 597–606. [Google Scholar] [CrossRef]
- Miko, E.; Czimmerer, Z.; Csánky, E.; Boros, G.; Buslig, J.; Dezso, B.; Scholtz, B. Differentially expressed microRNAs in small cell lung cancer. Exp. Lung. Res. 2009, 35, 646–664. [Google Scholar] [CrossRef]
- Silva, J.; García, V.; Zaballos, Á.; Provencio, M.; Lombardía, L.; Almonacid, L.; García, J.M.; Domínguez, G.; Peña, C.; Diaz, R.; et al. Vesicle-related microRNAs in plasma of nonsmall cell lung cancer patients and correlation with survival. Eur. Respir. J. 2011, 37, 617–623. [Google Scholar] [CrossRef] [PubMed]
- Turchinovich, A.; Drapkina, O.; Tonevitsky, A. Transcriptome of Extracellular Vesicles: State-of-the-Art. Front. Immunol. 2019, 10, 202. [Google Scholar] [CrossRef]
- Cech, T.R.; Steitz, J.A. The noncoding RNA revolution-trashing old rules to forge new ones. Cell 2014, 157, 77–94. [Google Scholar] [CrossRef]
- Qi, P.; Zhou, X.Y.; Du, X. Circulating long non-coding RNAs in cancer: Current status and future perspectives. Mol. Cancer 2016, 15, 39. [Google Scholar] [CrossRef]
- Qu, L.; Ding, J.; Chen, C.; Wu, Z.J.; Liu, B.; Gao, Y.; Chen, W.; Liu, F.; Sun, W.; Li, X.F.; et al. Exosome-Transmitted lncARSR Promotes Sunitinib Resistance in Renal Cancer by Acting as a Competing Endogenous RNA. Cancer Cell. 2016, 29, 653–668. [Google Scholar] [CrossRef]
- Wulfken, L.M.; Moritz, R.; Ohlmann, C.; Holdenrieder, S.; Jung, V.; Becker, F.; Herrmann, E.; Walgenbach-Brünagel, G.; von Ruecker, A.; Müller, S.C.; et al. MicroRNAs in renal cell carcinoma: Diagnostic implications of serum miR-1233 levels. PLoS ONE 2011, 6, e25787. [Google Scholar] [CrossRef]
- Redova, M.; Poprach, A.; Nekvindova, J.; Iliev, R.; Radova, L.; Lakomy, R.; Svoboda, M.; Vyzula, R.; Slaby, O. Circulating miR-378 and miR-451 in serum are potential biomarkers for renal cell carcinoma. J. Transl. Med. 2012, 10, 55. [Google Scholar] [CrossRef]
- Zhao, A.; Li, G.; Péoc’h, M.; Genin, C.; Gigante, M. Serum miR-210 as a novel biomarker for molecular diagnosis of clear cell renal cell carcinoma. Exp. Mol. Pathol. 2013, 94, 115–120. [Google Scholar] [CrossRef]
- Iwamoto, H.; Kanda, Y.; Sejima, T.; Osaki, M.; Okada, F.; Takenaka, A. Serum miR-210 as a potential biomarker of early clear cell renal cell carcinoma. Int. J. Oncol. 2014, 44, 53–58. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, A.L.; Ferreira, M.; Silva, J.; Gomes, M.; Dias, F.; Santos, J.I.; Maurício, J.; Lobo, F.; Medeiros, R. Higher circulating expression levels of miR-221 associated with poor overall survival in renal cell carcinoma patients. Tumour. Biol. 2014, 35, 4057–4066. [Google Scholar] [CrossRef]
- Wang, C.; Hu, J.; Lu, M.; Gu, H.; Zhou, X.; Chen, X.; Zen, K.; Zhang, C.Y.; Zhang, T.; Ge, J.; et al. A panel of five serum miRNAs as a potential diagnostic tool for early-stage renal cell carcinoma. Sci. Rep. 2015, 5, 7610. [Google Scholar] [CrossRef]
- Heinemann, F.G.; Tolkach, Y.; Deng, M.; Schmidt, D.; Perner, S.; Kristiansen, G.; Müller, S.C.; Ellinger, J. Serum miR-122-5p and miR-206 expression: Non-invasive prognostic biomarkers for renal cell carcinoma. Clin. Epigenet. 2018, 10, 11. [Google Scholar] [CrossRef]
- Hauser, S.; Wulfken, L.M.; Holdenrieder, S.; Moritz, R.; Ohlmann, C.H.; Jung, V.; Becker, F.; Herrmann, E.; Walgenbach-Brünagel, G.; von Ruecker, A.; et al. Analysis of serum microRNAs (miR-26a-2*.; miR-191.; miR-337-3p and miR-378) as potential biomarkers in renal cell carcinoma. Cancer. Epidemiol. 2012, 36, 391–394. [Google Scholar] [CrossRef]
- Fedorko, M.; Stanik, M.; Iliev, R.; Redova-Lojova, M.; Machackova, T.; Svoboda, M.; Pacik, D.; Dolezel, J.; Slaby, O. Combination of MiR-378 and MiR-210 Serum Levels Enables Sensitive Detection of Renal Cell Carcinoma. Int. J. Mol. Sci. 2015, 16, 23382–23389. [Google Scholar] [CrossRef]
- Chanudet, E.; Wozniak, M.B.; Bouaoun, L.; Byrnes, G.; Mukeriya, A.; Zaridze, D.; Brennan, P.; Muller, D.C.; Scelo, G. Large-scale genome-wide screening of circulating microRNAs in clear cell renal cell carcinoma reveals specific signatures in late-stage disease. Int. J. Cancer. 2017, 141, 1730–1740. [Google Scholar] [CrossRef]
- Zhang, W.; Ni, M.; Su, Y.; Wang, H.; Zhu, S.; Zhao, A.; Li, G. MicroRNAs in Serum Exosomes as Potential Biomarkers in Clear-cell Renal Cell Carcinoma. Eur. Urol. Focus 2018, 4, 412–419. [Google Scholar] [CrossRef]
- Chen, Y.; Wang, X.; Zhu, X.; Shao, S. Detection Performance of Circulating MicroRNA-210 for Renal Cell Carcinoma: A Meta-Analysis. Clin. Lab. 2018, 64, 569–576. [Google Scholar] [CrossRef]
- Lukamowicz-Rajska, M.; Mittmann, C.; Prummer, M.; Zhong, Q.; Bedke, J.; Hennenlotter, J.; Stenzl, A.; Mischo, A.; Bihr, S.; Schmidinger, M.; et al. MiR-99b-5p expression and response to tyrosine kinase inhibitor treatment in clear cell renal cell carcinoma patients. Oncotarget. 2016, 7, 78433–78447. [Google Scholar] [CrossRef]
- Tusong, H.; Maolakuerban, N.; Guan, J.; Rexiati, M.; Wang, W.G.; Azhati, B.; Nuerrula, Y.; Wang, Y.J. Functional analysis of serum microRNAs miR-21 and miR-106a in renal cell carcinoma. Cancer Biomark. 2017, 18, 79–85. [Google Scholar] [CrossRef]
| miRNAs | Targets in RCC | Renal Samples | Reference |
|---|---|---|---|
| miR-183 | Dickkopf-related protein-3 (DKK-3) Protein phosphatase 2A | Renal CSC EVs Renal tumor cells Renal cancer tissues Serum of RCC patients | Grange et al. [30] Qiu et al. [71] Zhang et al. [72] |
| miR-486 | TGF-β-activated kinase 1 | Renal CSC EVs Renal tumor cells Renal cancer tissues | Grange et al. [30] He et al. [73] Goto et al. [74] |
| miR-19a | PTEN SMAD4 RhoB PIK3CA | Renal CSC EVs Renal cancer tissues | Grange et al. [30] Ma et al. [75] Xiao et al. [76] Niu et al. [77] |
| miR-92 | VHL | Renal CSC EVs Renal cancer tissues | Valera et al. [78] Grange et al. [30] |
| miR-19b | RhoB PTEN/PI3K/AKT Signalling Pathway | Renal CSC EVs | Grange et al. [30] Niu et al. [77] Liu et al. [79] |
| miR-650 | Not determined | Renal CSC EVs | Grange et al. [30] |
| miR-151 | Not determined | Renal CSC EVs | Grange et al. [30] |
| miR-301 | Not determined | Renal CSC EVs | Grange et al. [30] |
| miRNA | Screening Technique | miRNA Source | Screening Groups | Validation Groups | Study Results | References |
|---|---|---|---|---|---|---|
| miR-378 | Real time PCR | Serum | 25 healthy controls 25 RCC | 123 healthy controls 117 RCC | miR-378 levels were higher in RCC screening group versus healthy controls, but results were not validated. | Hauser et al. [104] |
| Real time PCR | Serum | 15 healthy controls 12 RCC | 35 healthy controls 90 RCC | Validation of the increase of miR-378 in patients’ sera, in combination with the downregulation of the antitumor miR-451. | Redova et al. [98] | |
| Real time PCR | Serum | 100 healthy controls 195 RCC | Not performed | Diagnostic and prognostic value of miR-378 increase, in combination with miR-210 increase. | Fedorko et al. [105] | |
| miR-221 | Real time PCR | Plasma | 34 healthy controls 44 RCC | Not performed | miR-221 levels positively correlate with progression towards a metastatic disease and inversely correlate with survival rate. | Teixeira et al. [101] |
| miR-210 | Real time PCR | Tissue samples and serum | 32 RCC tissues samples (tumor and adjacent healthy tissue); sera from 42 healthy controls, 68 RCC before and 10 RCC patients after nephrectomy | Not performed | Increased levels of miR-210 both in tissue samples (compared to adjacent healthy tissue) and in RCC sera. | Zhao et al. [99] |
| Real time PCR | Tissue samples and serum | 23 healthy controls 34 RCC | Not performed | Increased levels of miR-210 in RCC sera compared to controls. No correlation with tumor stage. | Iwamoto et al. [100] | |
| Real time PCR | Serum | 100 healthy controls 195 RCC | Not performed | Increased levels of miR-210 in RCC sera compared to controls. Prognostic value in combination with miR-378. | Fedorko et al. [105] | |
| Real time PCR | Serum EV | 80 healthy controls, 82 RCC before nephrectomy 10 RCC after nephrectomy. | Not performed | Higher content in miR-210 and miR-1233 in EV isolated from RCC patients’ sera. | Zhang et al. [107] | |
| Meta analysis | Serum | Not performed | 7 studies: 415 healthy controls 570 RCC | Validation of the potential diagnostic value of miR-210. | Chen et al. [108] | |
| miR-193a | TaqMan Low Density Array | Serum | 25 healthy controls 25 RCC | 107 healthy controls 107 RCC | Increased in RCC sera. | Wang et al. [102] |
| miR-362 | TaqMan Low Density Array | Serum | 25 healthy controls 25 RCC | 107 healthy controls 107 RCC | Increased in RCC sera. | Wang et al. [102] |
| miR-572 | TaqMan Low Density Array | Serum | 25 healthy controls 25 RCC | 107 healthy controls 107 RCC | Increased in RCC sera. | Wang et al. [102] |
| miR-1233 | TaqMan Low Density Array | Tissue samples and serum | 6 healthy controls and 6 RCC | 93 healthy controls and 84 RCC | miR-1233 was identified and validated as increased in RCC. | Wulfken et al. [97] |
| Real Time PCR | Serum EPCAM+ EV | 80 healthy controls, 82 RCC before nephrectomy 10 RCC after nephrectomy. | Not performed | Higher content in miR-210 and miR-1233 in EV isolated from RCC patients’ sera. | Zhang et al. [107] | |
| miR-99b | Sequencing | Tissue samples | 40 RCC (tumor and adjacent healthy tissue) | 65 RCC | miR-99b levels positively correlated with tumor progression after TKI therapy | Lukamowicz et al. [109] |
| miR-21 | Real Time PCR | Serum | 30 healthy controls 30 RCC before and after nephrectomy | Not performed | Increased levels in RCC sera. | Tusong et al. [110] |
| miR-106a | Real Time PCR | Serum | 30 healthy controls 30 RCC before and after nephrectomy | Not performed | Increased levels in RCC sera. | Tusong et al. [110] |
| miR-122-5p | small RNA sequencing | Serum | 8 benign renal tumors 18 RCC | 28 healthy controls 47 benign renal tumors 68 RCC | Correlation of miR-122 levels with metastatic progression and survival | Heinemann et al. [103] |
| miR-206 | small RNA sequencing | Serum | 8 benign renal tumors 18 RCC | 28 healthy controls 47 benign renal tumors 68 RCC | Correlation of miR-122 levels with metastatic progression and survival | Heinemann et al. [103] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Grange, C.; Brossa, A.; Bussolati, B. Extracellular Vesicles and Carried miRNAs in the Progression of Renal Cell Carcinoma. Int. J. Mol. Sci. 2019, 20, 1832. https://doi.org/10.3390/ijms20081832
Grange C, Brossa A, Bussolati B. Extracellular Vesicles and Carried miRNAs in the Progression of Renal Cell Carcinoma. International Journal of Molecular Sciences. 2019; 20(8):1832. https://doi.org/10.3390/ijms20081832
Chicago/Turabian StyleGrange, Cristina, Alessia Brossa, and Benedetta Bussolati. 2019. "Extracellular Vesicles and Carried miRNAs in the Progression of Renal Cell Carcinoma" International Journal of Molecular Sciences 20, no. 8: 1832. https://doi.org/10.3390/ijms20081832
APA StyleGrange, C., Brossa, A., & Bussolati, B. (2019). Extracellular Vesicles and Carried miRNAs in the Progression of Renal Cell Carcinoma. International Journal of Molecular Sciences, 20(8), 1832. https://doi.org/10.3390/ijms20081832







