A New Subclade of Leptosphaeria biglobosa Identified from Brassica rapa
Abstract
:1. Introduction
2. Results
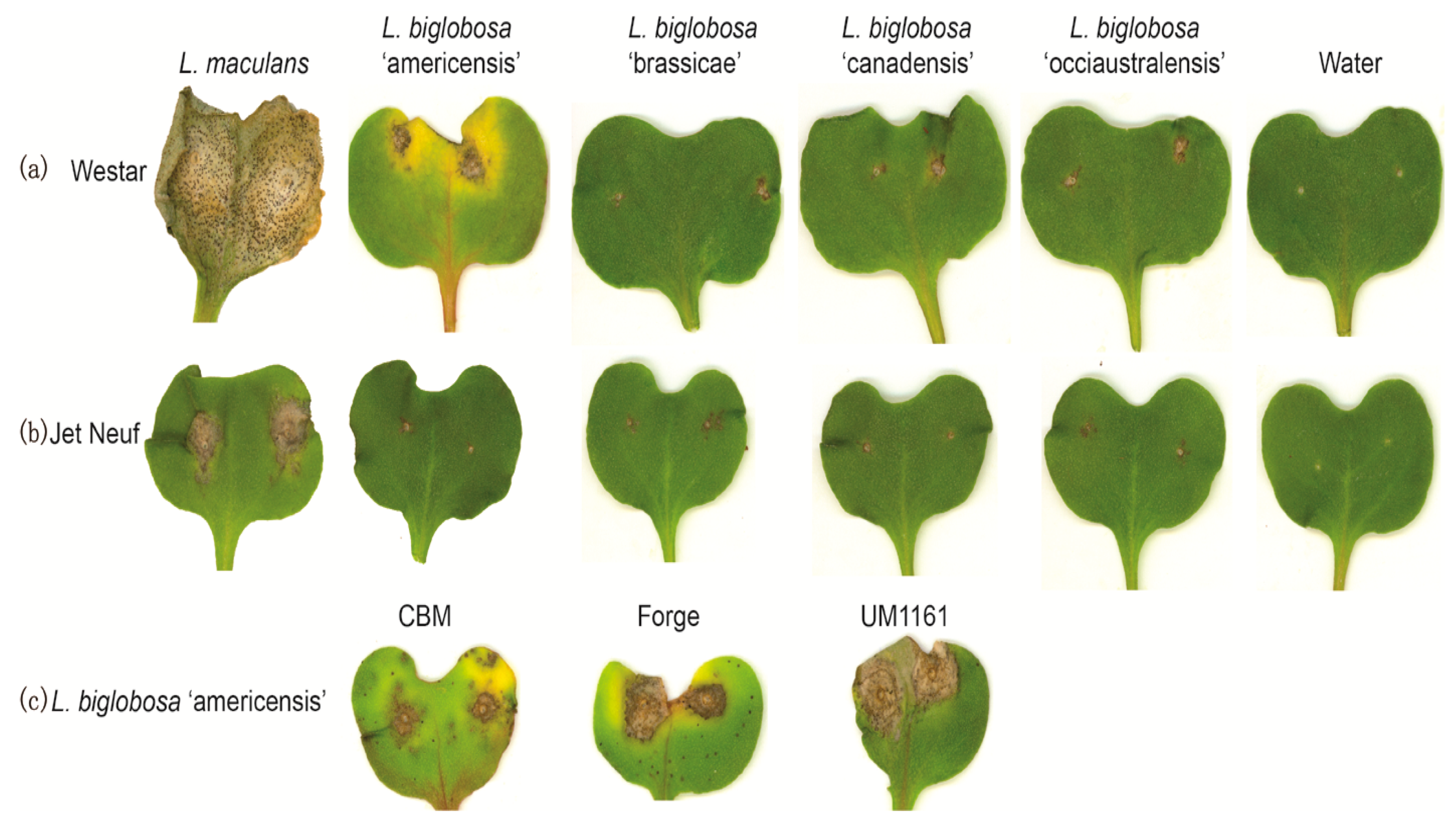
2.1. Pathogenicity Tests
2.2. Characterization of B. rapa-Derived Isolates from the Willamette Valley of Oregon
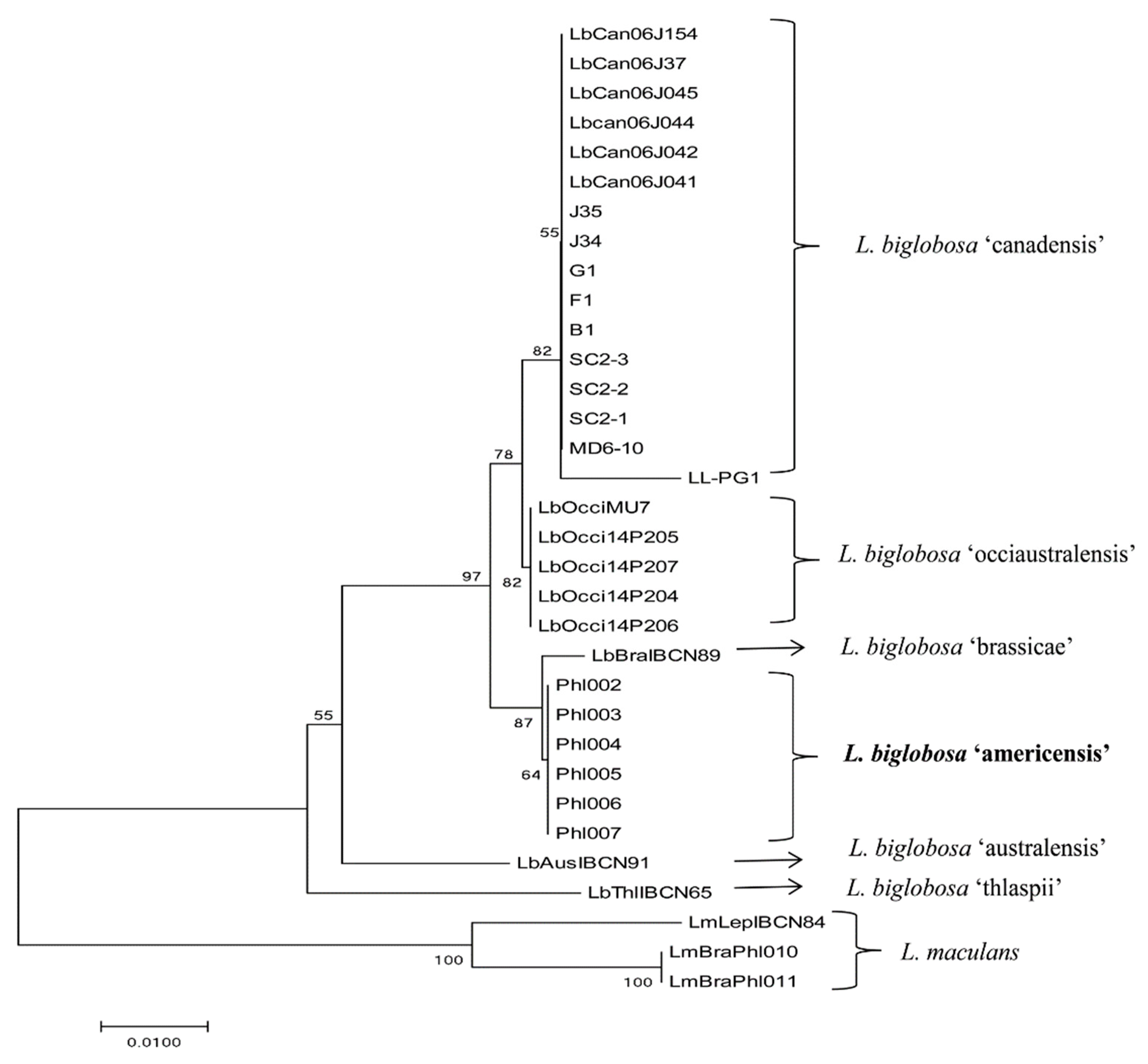
2.3. Identification of a New Subclade of L. biglobosa by Phylogenetic Analysis
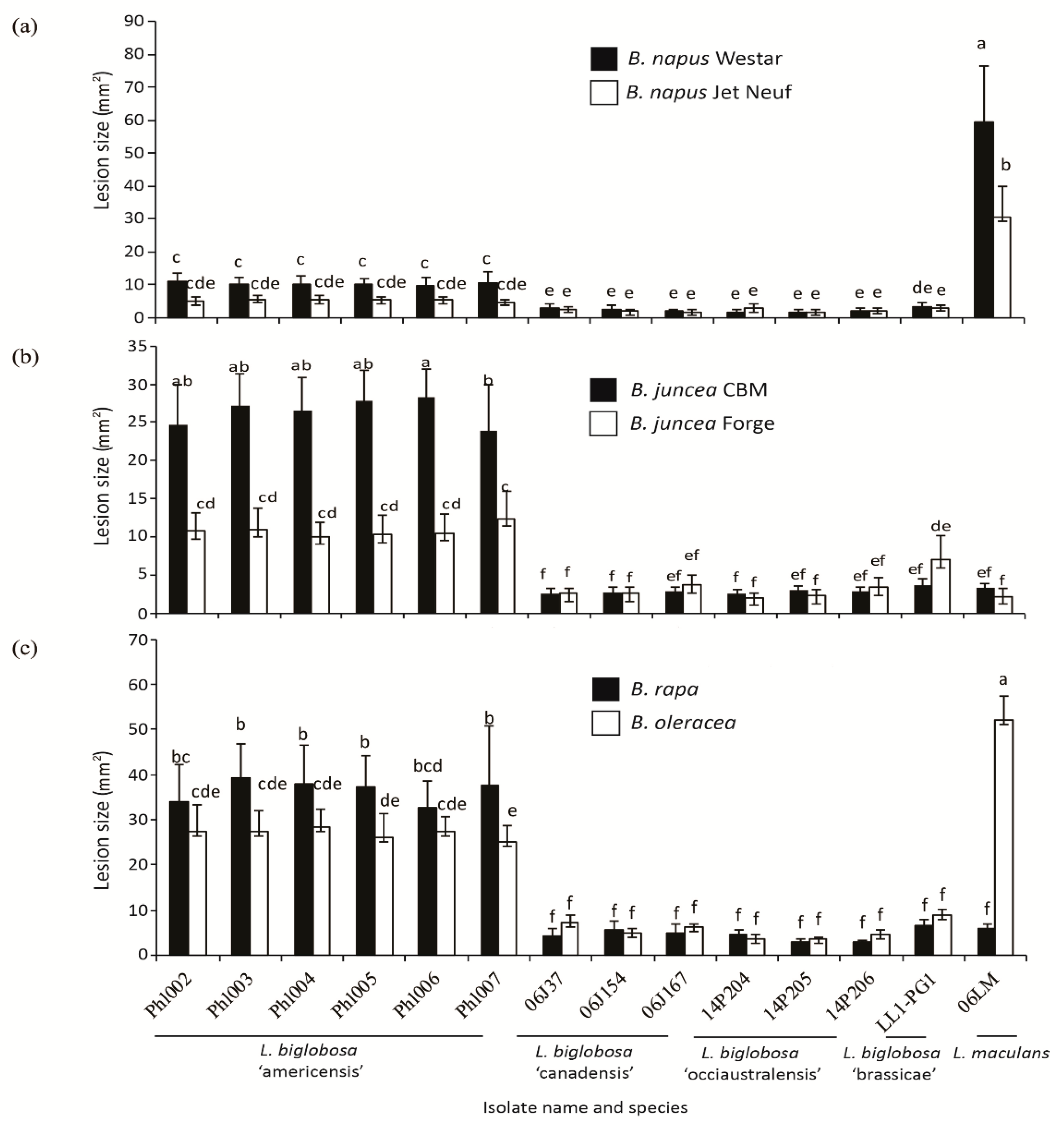
2.4. Differences in the Pathogenicity of Isolates of L. biglobosa ‘americensis’ and Other L. biglobosa Subspecies
3. Discussion
4. Materials and Methods
4.1. Isolate Collection
4.2. Initial Pathogenicity Testing of the Oregon B. rapa-Derived Leptosphaeria Isolates on Brassica Species
4.3. PCR Identification of Isolates
4.4. Conserved Gene Sequencing and Sequence Alignment
4.5. Phylogenetic Analysis
4.6. Pathogenicity of L. biglobosa Subclade Isolates on Brassica Species
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Brun, H.; Levivier, S.; Eber, F.; Renard, M.; Chèvre, A.M. Electrophoretic analysis of natural populations of Leptosphaeria maculans directly from leaf lesions. Plant Pathol. 1997, 46, 147–154. [Google Scholar] [CrossRef]
- West, J.S.; Kharbanda, P.D.; Barbetti, M.J.; Fitt, B.D.L. Epidemiology and management of Leptosphaeria maculans (Phoma stem canker) on oilseed rape in Australia, Canada and Europe. Plant Pathol. 2001, 50, 10–27. [Google Scholar] [CrossRef]
- Fitt, B.D.L.; Brun, H.; Barbetti, M.J.; Rimmer, S.R. World-wide importance of Phoma stem canker (Leptosphaeria maculans and L. biglobosa) on oilseed rape (Brassica napus). Eur. J. Plant Pathol. 2006, 114, 3–15. [Google Scholar] [CrossRef]
- Ghanbarnia, K.; Fudal, I.; Larkan, N.J.; Links, M.G.; Balesdent, M.H.; Profotova, B.; Fernando, W.G.D.; Borhan, M.H. Rapid identification of the Leptosphaeria maculans avirulence gene AvrLm2 using an intraspecific comparative genomics approach. Mol. Plant Pathol. 2015, 16, 699–709. [Google Scholar] [CrossRef] [PubMed]
- Lowe, R.G.; Cassin, A.; Grandaubert, J.; Clark, B.L.; Van de Wouw, A.P.; Rouxel, T.; Howlett, B.J. Genomes and transcriptomes of partners in plant-fungal-interactions between canola (Brassica napus) and two Leptosphaeria species. PLoS ONE 2014, 9, e103098. [Google Scholar] [CrossRef]
- Mendes-Pereira, E.; Balesdent, M.H.; Hortense, B.; Rouxel, T. Molecular phylogeny of the Leptosphaeria maculans–L. biglobosa species complex. Mycol. Res. 2003, 107, 1287–1304. [Google Scholar] [CrossRef]
- Vincenot, L.; Balesdent, M.H.; Li, H.; Barbetti, M.J.; Sivasithamparam, K.; Gout, L.; Rouxel, T. Occurrence of a new subclade of Leptosphaeria biglobosa in Western Australia. Phytopathology 2008, 98, 321–329. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Latunde-Dada, A.O.; Hall, A.M.; Fitt, B.D.L. Phoma stem canker disease on oilseed rape (Brassica napus) in China is caused by Leptosphaeria biglobosa ‘brassicae’. Eur. J. Plant Pathol. 2014, 140, 841–857. [Google Scholar] [CrossRef]
- Dilmaghani, A.; Balesdent, M.H.; Didier, J.P.; Wu, C.; Davey, J.; Barbetti, M.J.; Li, H.; Moreno-Rico, O.; Phillips, D.; Despeghel, J.P.; et al. The Leptosphaeria maculans–Leptosphaeria biglobosa species complex in the American continent. Plant Pathol. 2009, 58, 1044–1058. [Google Scholar] [CrossRef]
- Van de Wouw, A.P.; Thomas, V.L.; Cozijnsen, A.J.; Marcroft, S.J.; Salisbury, P.A.; Howlett, B.J. Identification of Leptosphaeria biglobosa ‘canadensis’ on Brassica juncea stubble from northern New South Wales, Australia. Australas. Plant Dis. Notes 2008, 3, 124–128. [Google Scholar] [CrossRef]
- Voigt, K.; Cozijnsen, A.J.; Kroymann, J.; Pöggeler, S.; Howlett, B.J. Phylogenetic relationships between members of the crucifer pathogenic Leptosphaeria maculans species complex as shown by mating type (MAT1-2), actin, and β-tubulin sequences. Mol. Phylogenetics Evol. 2005, 37, 541–557. [Google Scholar] [CrossRef] [PubMed]
- Plummer, K.M.; Dunse, K.; Howlett, B.J. Non-aggressive strains of the blackleg fungus, Leptosphaeria maculans, are present in Australia and can be distinguished from aggressive strains by molecular analysis. Aust. J. Bot. 1994, 42, 1–8. [Google Scholar] [CrossRef]
- O’Donnell, K.; Ward, T.J.; Geiser, D.M.; Kistler, H.C.; Aoki, T. Genealogical concordance between the mating type locus and seven other nuclear genes supports formal recognition of nine phylogenetically distinct species within the Fusarium graminearum clade. Fungal Genet. Biol. 2004, 41, 600–623. [Google Scholar] [CrossRef]
- Leslie, J.F.; Summerell, B.A. The Fusarium Laboratory Manual; Wiley-Blackwell: Ames, IA, USA, 2006; pp. 81–96. [Google Scholar]
- Gale, L.R.; Harrison, S.A.; Ward, T.J.; O’Donnell, K.; Milus, E.A.; Gale, S.W.; Kistler, H.C. Nivalenol-type populations of Fusarium graminearum and F. asiaticum are prevalent on wheat in Southern Louisiana. Phytopathology 2011, 101, 124–134. [Google Scholar] [CrossRef]
- Sarver, B.A.; Ward, T.J.; Gale, L.R.; Broz, K.; Kistler, H.C.; Aoki, T.; Nicholson, P.; Carter, J.; O’Donnell, K. Novel Fusarium head blight pathogens from Nepal and Louisiana revealed by multilocus genealogical concordance. Fungal Genet. Biol. 2011, 48, 1096–1107. [Google Scholar] [CrossRef]
- Wang, J.; Ndoye, M.; Zhang, J.; Li, H.; Liao, Y. Population structure and genetic diversity of Fusarium graminearum species complex. Toxin 2011, 3, 1020–1037. [Google Scholar] [CrossRef] [PubMed]
- Rouxel, T.; Balesdent, M.H. The stem canker (blackleg) fungus, Leptosphaeria maculans, enters the genomic era. Mol. Plant Pathol. 2005, 6, 225–241. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.J.; Fitt, B.D.L.; Jedryczka, M.; Dakowska, S.; West, J.S.; Gladders, P.; Steed, J.M.; Li, Z. Patterns of ascospore release in relation to Phoma stem canker epidemiology in England (Leptosphaeria maculans) and Poland (Leptosphaeria biglobosa). Eur. J. Plant Pathol. 2005, 111, 263–277. [Google Scholar] [CrossRef]
- El Hadrami, A.; Fernando, W.G.D.; Daayf, F. Variations in relative humidity modulate Leptosphaeria spp. pathogenicity and interfere with canola mechanisms of defence. Eur. J. Plant Pathol. 2010, 126, 187–202. [Google Scholar] [CrossRef]
- Rouxel, T.; Grandaubert, J.; Hane, J.K.; Hoede, C.; Van de Wouw, A.P.; Couloux, A.; Dominguez, V.; Anthouard, V.; Bally, P.; Bourras, S.; et al. Effector diversification within compartments of the Leptosphaeria maculans genome affected by repeat-induced point mutations. Nat. Commun. 2011, 2, 202. [Google Scholar] [CrossRef]
- Balesdent, M.; Barbetti, M.; Li, H.; Sivasithamparam, K.; Gout, L.; Rouxel, T. Analysis of Leptosphaeria maculans race structure in a worldwide collection of isolates. Phytopathology 2005, 95, 1061–1071. [Google Scholar] [CrossRef]
- Van de Wouw, A.P.; Elliott, V.L.; Ware, A.; Lindbeck, K.; Howlett, B.J.; Marcroft, S.J. Infection of canola pods by Leptosphaeria maculans and subsequent seed contamination. Eur. J. Plant Pathol. 2016, 145, 687–695. [Google Scholar] [CrossRef]
- Zhang, X.; Peng, G.; Kutcher, H.R.; Balesdent, M.H.; Delourme, R.; Fernando, W.G.D. Breakdown of Rlm3 resistance in the Brassica napus–Leptosphaeria maculans pathosystem in western Canada. Eur. J. Plant Pathol. 2016, 1–16. [Google Scholar] [CrossRef]
- Williams, P.H.; Delwiche, P.A. Screening for resistance to blackleg of crucifers in the seedling stage. In Eucarpia Conf. Breed. Cruciferous Crops 1979; Foundation for Agricultural Plant Breeding: Wageningen, The Netherlands, 1980; pp. 164–170. [Google Scholar]
- Calderon, C.; Ward, E.; Freeman, J.; Foster, S.J.; McCartney, H.A. Detection of airborne inoculum of Leptosphaeria maculans and Pyrenopeziza brassicae in oilseed rape crops by polymerase chain reaction (PCR) assays. Plant Pathol. 2002, 51, 303–310. [Google Scholar] [CrossRef]
- Cozijnsen, A.J.; Howlett, B.J. Characterisation of the mating-type locus of the plant pathogenic ascomycete Leptosphaeria maculans. Curr. Genet. 2003, 43, 351–357. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic. Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Söding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef]
- Nicholas, K.B.; Nicholas, H.B.J.; Deerfield, D.W. GeneDoc: Analysis and visualization of genetic variation. Embnew 1997, 4, 14. [Google Scholar]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
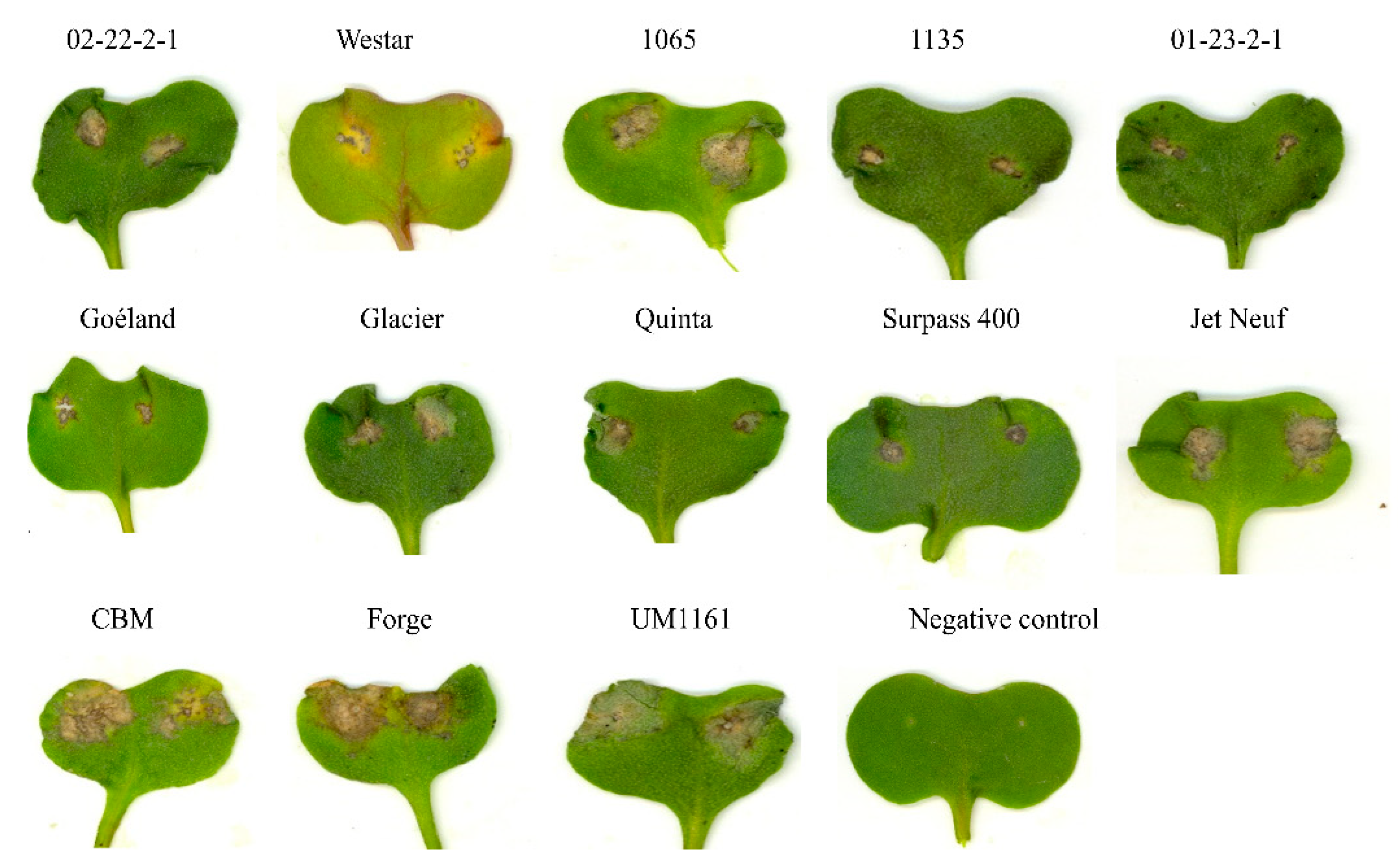

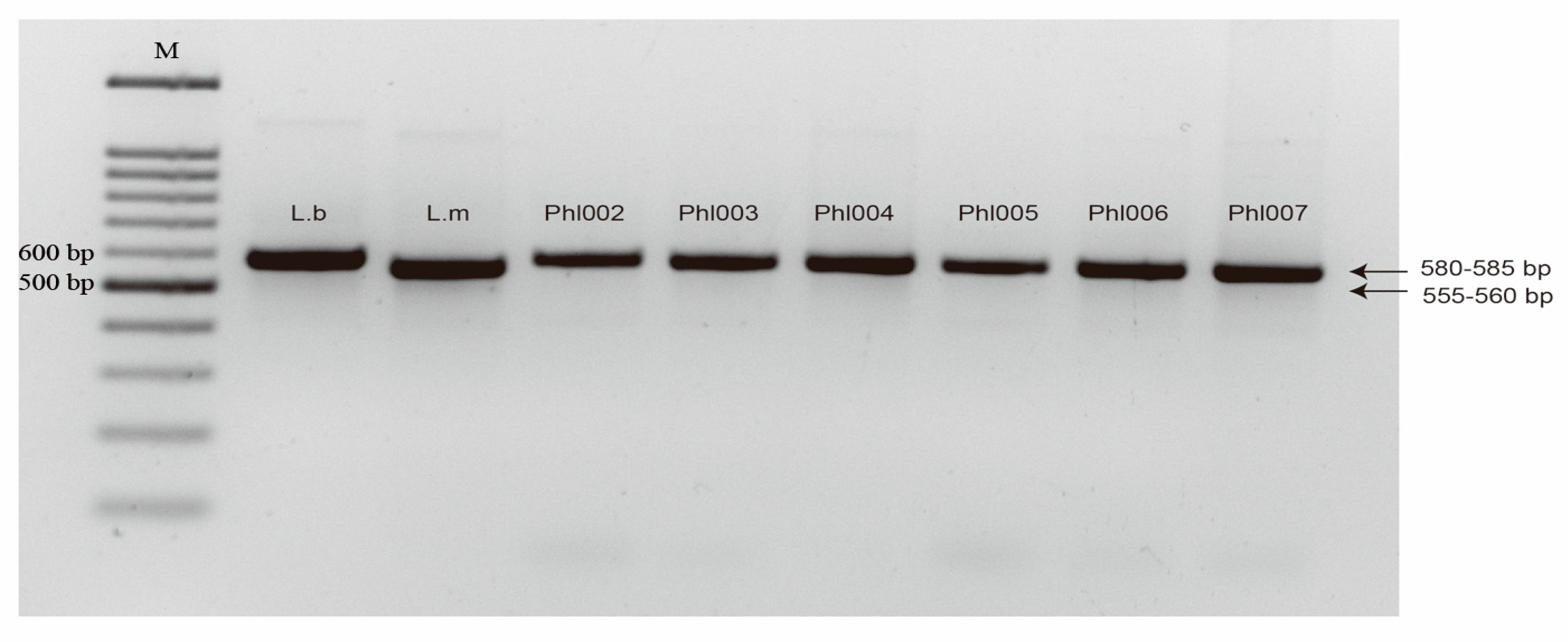
| Cultivar/Line | Brassica Species | B. rapa-Derived Isolates | Inferred Phenotype | |||||
|---|---|---|---|---|---|---|---|---|
| Phl002 | Phl003 | Phl004 | Phl005 | Phl006 | Phl007 | |||
| Westar | Brassica napus | 4.50 a,b | 4.40 | 4.57 | 5.29 | 4.83 | 4.29 | IR c |
| Jet Neuf | Brassica napus | 3.00 | 4.33 | 4.86 | 4.86 | 3.29 | 4.14 | R |
| Surpass 400 | Brassica napus | 3.71 | 3.57 | 2.69 | 2.43 | 2.71 | 2.71 | R |
| 01-23-2-1 | Brassica napus | 5.33 | 4.86 | 3.83 | 3.43 | 2.83 | 2.57 | R |
| Quinta | Brassica napus | 3.67 | 5.00 | 3.00 | 2.60 | 2.33 | 2.79 | R |
| 1065 | Brassica napus | 4.50 | 6.33 | 5.33 | 4.57 | 4.00 | 4.67 | IR |
| Glacier | Brassica napus | 3.00 | 3.80 | 3.60 | 2.25 | 2.60 | 3.20 | R |
| 1135 | Brassica napus | 2.14 | 2.29 | 1.57 | 1.57 | 2.14 | 1.57 | R |
| Goéland | Brassica napus | 1.50 | 2.67 | 2.40 | 1.71 | 1.10 | 1.50 | R |
| 02-22-2-1 | Brassica napus | 4.17 | 4.29 | 4.43 | 4.43 | 4.86 | 4.71 | R |
| Varox | Brassica juncea | 3.00 | 2.67 | 3.00 | 3.83 | 2.67 | 3.00 | R |
| Estilin | Brassica juncea | 2.67 | 3.83 | 3.83 | 3.83 | 2.17 | 1.83 | R |
| UM3309 | Brassica juncea | 3.00 | 2.33 | 1.67 | 2.67 | 1.83 | 2.67 | R |
| Forge | Brassica juncea | 5.17 | 6.17 | 6.17 | 7.33 | 6.17 | 6.50 | S |
| Vox-0 | Brassica juncea | 4.57 | 2.67 | 3.00 | 3.83 | 2.67 | 3.00 | R |
| Dohirda | Brassica juncea | 3.83 | 3.00 | 2.33 | 1.83 | 2.67 | 2.67 | R |
| CBM | Brassica juncea | 6.17 | 7.00 | 6.17 | 6.17 | 6.83 | 6.67 | S |
| UM1112 | Brassica rapa | 6.67 | 6.34 | 6.34 | 6.67 | 6.83 | 6.17 | S |
| UM1161 | Brassica rapa | 7.33 | 6.17 | 6.17 | 6.50 | 6.34 | 6.67 | S |
| UM1402 | Brassica rapa | 2.67 | 3.83 | 2.33 | 3.00 | 2.67 | 2.67 | R |
| UM1113 | Brassica rapa | 6.17 | 6.34 | 6.67 | 7.33 | 6.00 | 5.67 | S |
| UM1147 | Brassica rapa | 2.67 | 3.00 | 1.83 | 2.33 | 2.33 | 2.67 | R |
| UM1154 | Brassica rapa | 3.00 | 3.00 | 2.67 | 1.83 | 2.67 | 2.33 | R |
| Primer Name a | Sequence (5′–3′) | Annealing Temperature (°C) | Reference |
|---|---|---|---|
| ITS F (PN3) | CCGTTGGTGAACCAGCGGAGGGATC | 58 | Mendes-Pereira et al. (2003) [6] |
| ITS R (PN10) | TCCGCTTATTGATATGCTTAAG | ||
| Actin F | GAGCAGGAGATCCAGACTGC | 56 | Van de Wouw et al. (2008) [10] |
| Actin R | TTCGAGATCCACATCTGCTG | ||
| β-tubulin F | GTCGAGAACTCCGACGAGAC | 55 | Van de Wouw et al. (2008) [10] |
| β-tubulin R | ATCTGGTCCTCGACCTCCTT | ||
| MAT1.1-F | CTCGATGCAATGTACTTGG | 56 | Cozijinsen and Howlett (2003) [27] |
| MAT1.2-F | AGCCGGAGGTGAAGTTGAAGCCG | ||
| MAT-R | TGGCGAATTAAGGGATTGCTG |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zou, Z.; Zhang, X.; Parks, P.; du Toit, L.J.; Van de Wouw, A.P.; Fernando, W.G.D. A New Subclade of Leptosphaeria biglobosa Identified from Brassica rapa. Int. J. Mol. Sci. 2019, 20, 1668. https://doi.org/10.3390/ijms20071668
Zou Z, Zhang X, Parks P, du Toit LJ, Van de Wouw AP, Fernando WGD. A New Subclade of Leptosphaeria biglobosa Identified from Brassica rapa. International Journal of Molecular Sciences. 2019; 20(7):1668. https://doi.org/10.3390/ijms20071668
Chicago/Turabian StyleZou, Zhongwei, Xuehua Zhang, Paula Parks, Lindsey J. du Toit, Angela P. Van de Wouw, and W. G. Dilantha Fernando. 2019. "A New Subclade of Leptosphaeria biglobosa Identified from Brassica rapa" International Journal of Molecular Sciences 20, no. 7: 1668. https://doi.org/10.3390/ijms20071668
APA StyleZou, Z., Zhang, X., Parks, P., du Toit, L. J., Van de Wouw, A. P., & Fernando, W. G. D. (2019). A New Subclade of Leptosphaeria biglobosa Identified from Brassica rapa. International Journal of Molecular Sciences, 20(7), 1668. https://doi.org/10.3390/ijms20071668