Concomitant Expression of Prolactin Receptor and TGFβ Receptors in Breast Cancer: Association with Less Aggressive Phenotype and Favorable Patient Outcome
Abstract
1. Introduction
2. Results
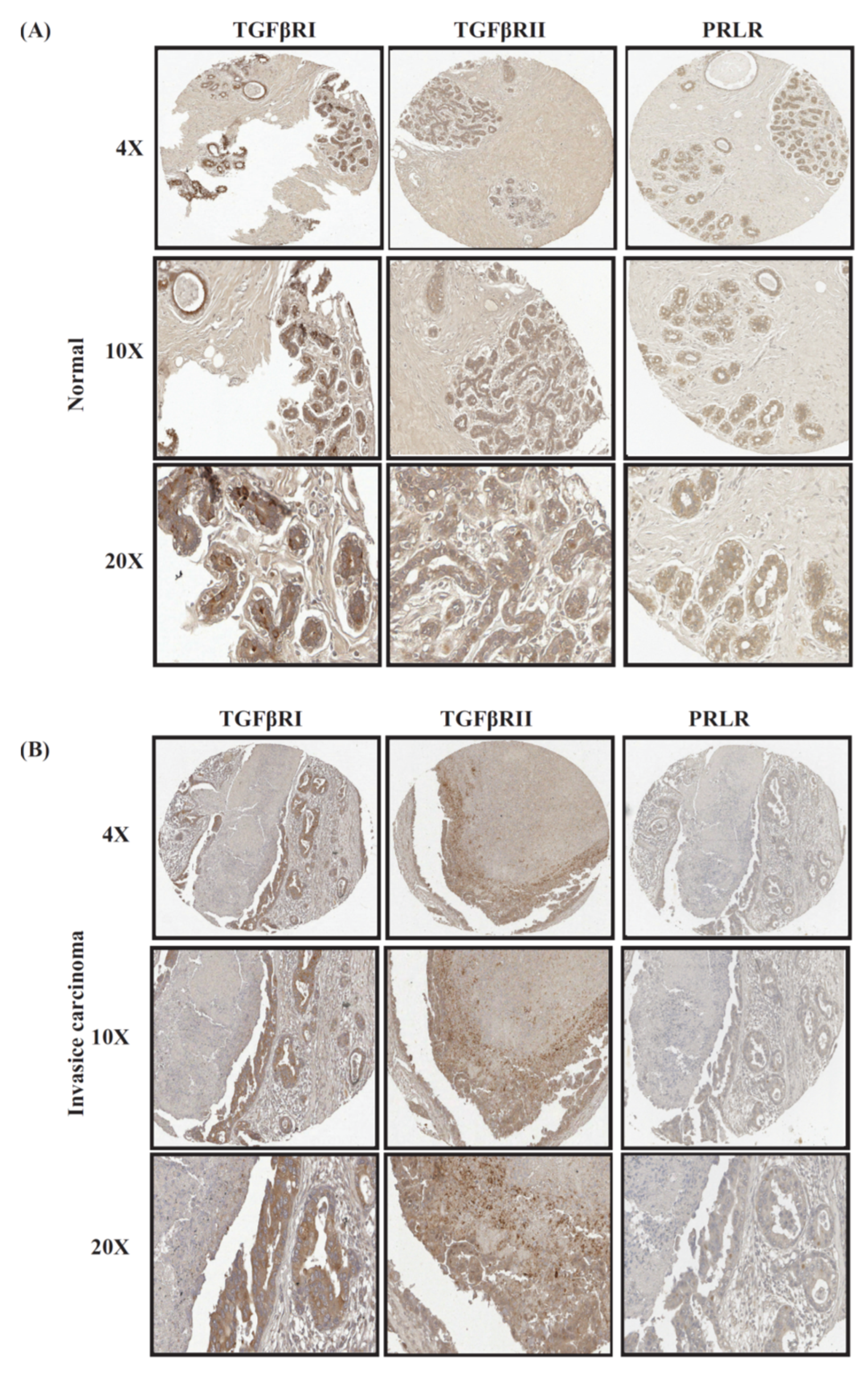
2.1. Immunohistochemical Analyses of Co-Expression of PRLR and TGFβ Receptors in Relation to Breast Cancer Progression
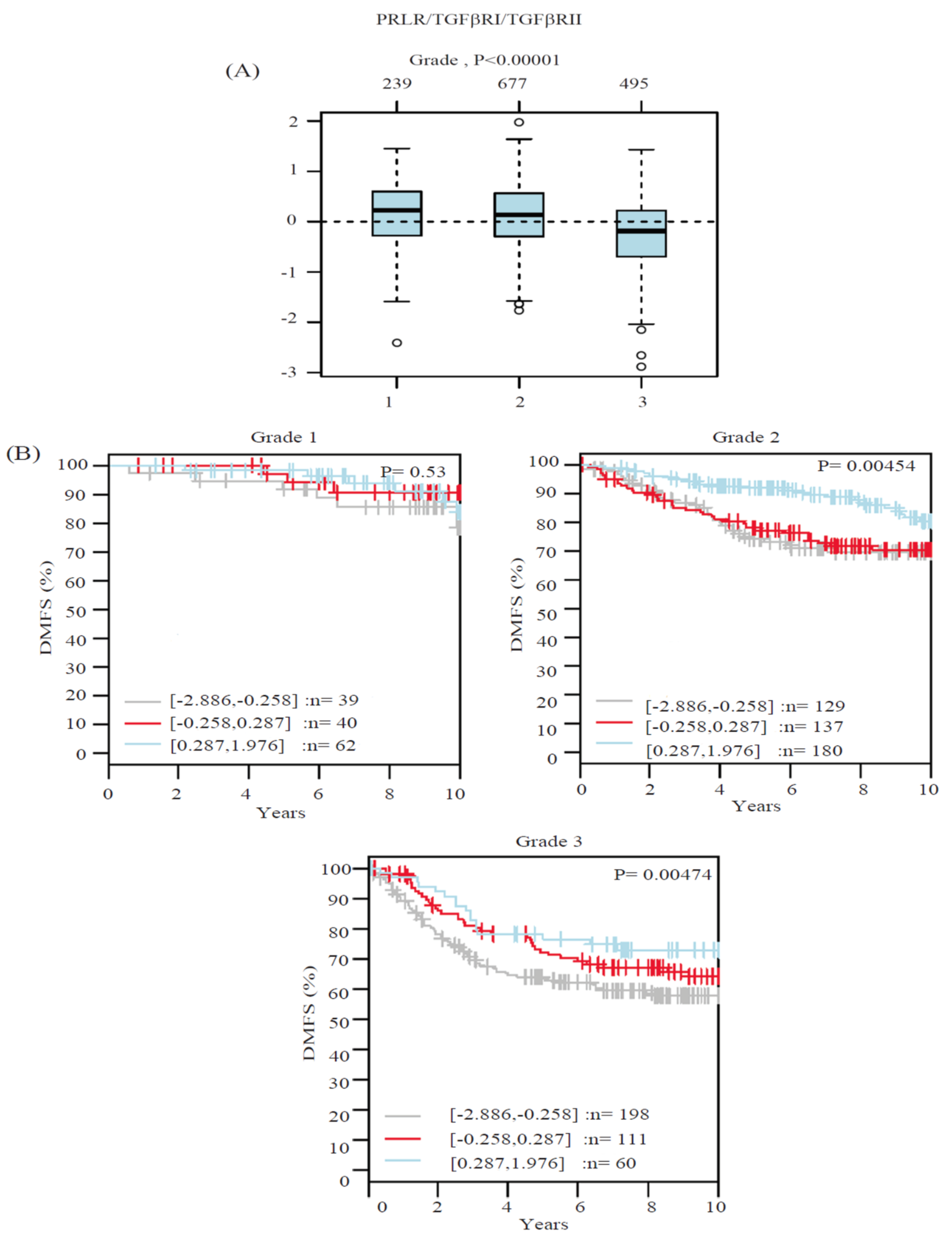
2.2. Co-Expression Levels of PRLR and TGFβ Receptors in Relation to Tumor Grade in Breast Cancer
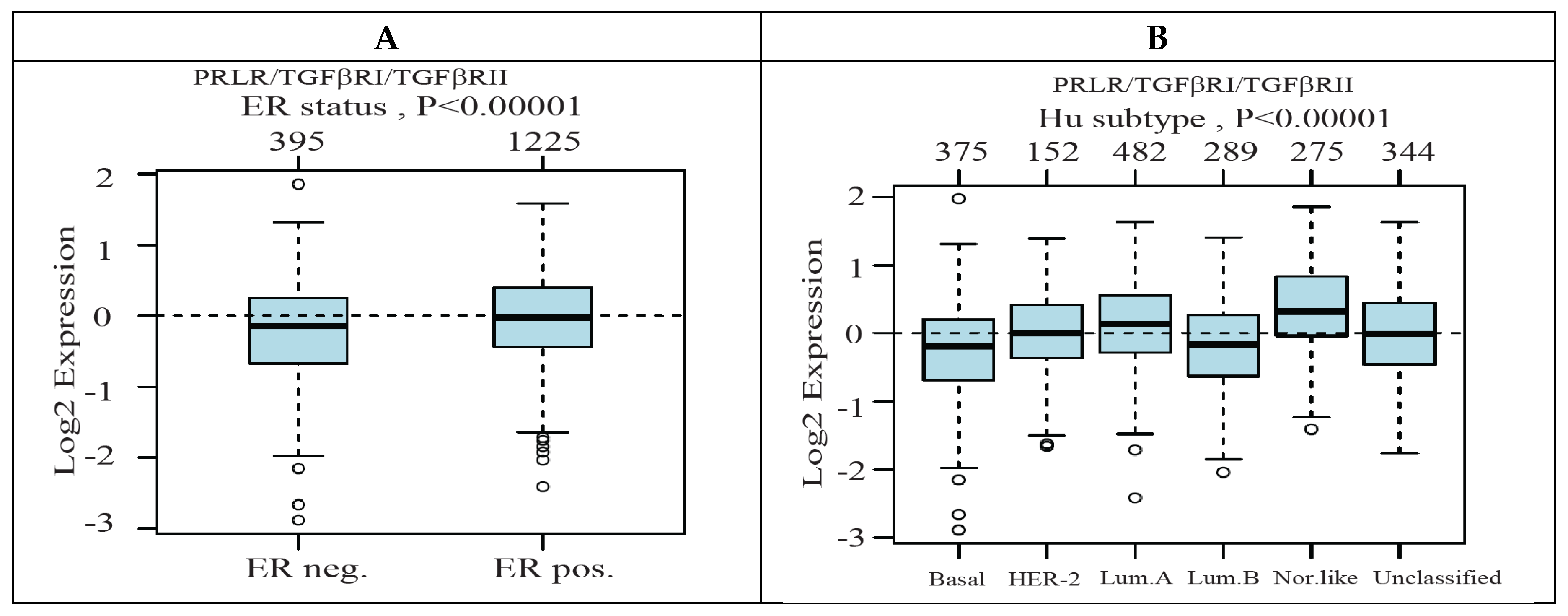
2.3. Co-Expression Levels of PRLR and TGFβ Receptors in Relation to Breast Cancer Molecular Subtypes
2.4. Higher Levels of Expression of PRLR/TGFβRI/TGFβRII Gene Signature Are Associated with Prolonged Survival in Luminal B and HER-2-Enriched Breast Cancer Molecular Subtypes
3. Discussion
4. Materials and Methods
4.1. Tissue Samples
4.2. Immunohistochemistry
4.3. Immunohistochemistry Scoring
4.4. Data Mining
4.5. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| EMT | Epithelial–mesenchymal transition |
| PRLR | Prolactin receptor |
| DMFS | Distant metastasis-free survival |
| TNBC | Triple-negative breast cancer |
| HER-2 | Human epidermal growth factor receptor 2 |
| TGFβ | Transforming growth factor beta |
References
- Da Silva-Diz, V.; Lorenzo-Sanz, L.; Bernat-Peguera, A.; Lopez-Cerda, M.; Muñoz, P. Cancer cell plasticity: Impact on tumor progression and therapy response. Semin Cancer Biol. 2018, 53, 48–58. [Google Scholar] [CrossRef]
- Chaffer, C.L.; San Juan, B.P.; Lim, E.; Weinberg, R.A. EMT, cell plasticity and metastasis. Cancer Metastasis Rev. 2016, 35, 645–654. [Google Scholar] [CrossRef] [PubMed]
- Santamaria, P.G.; Moreno-Bueno, G.; Portillo, F.; Cano, A. EMT: Present and future in clinical oncology. Mol. Oncol. 2017, 11, 718–738. [Google Scholar] [CrossRef]
- Neel, J.-C.; Humbert, L.; Lebrun, J.-J. The Dual Role of TGF in Human Cancer: From Tumor Suppression to Cancer Metastasis. ISRN Mol. Biol. 2012, 2012, 28. [Google Scholar] [CrossRef]
- Muraoka-Cook, R.S.; Dumont, N.; Arteaga, C.L. Dual role of transforming growth factor beta in mammary tumorigenesis and metastatic progression. Clin. Cancer Res. 2005, 11, 937s–943s. [Google Scholar] [PubMed]
- Derynck, R.; Akhurst, R.J.; Balmain, A. TGF-β signaling in tumor suppression and cancer progression. Nat. Genet. 2001, 29, 117–129. [Google Scholar] [CrossRef] [PubMed]
- Wakefield, L.M.; Roberts, A.B. TGF-β signaling: Positive and negative effects on tumorigenesis. Curr. Opin. Genet. Dev. 2002, 12, 22–29. [Google Scholar] [CrossRef]
- Derynck, R.; Goeddel, D.V.; Ullrich, A.; Gutterman, J.U.; Williams, R.D.; Bringman, T.S.; Berger, W.H. Synthesis of messenger RNAs for transforming growth factors α and β and the epidermal growth factor receptor by human tumors. Cancer Res. 1987, 47, 707–712. [Google Scholar] [PubMed]
- Dickson, R.B.; Kasid, A.; Huff, K.K.; Bates, S.E.; Knabbe, C.; Bronzert, D.; Gelmann, E.P.; Lippman, M.E. Activation of growth factor secretion in tumorigenic states of breast cancer induced by 17 β-estradiol or v-Ha-ras oncogene. Proc. Natl. Acad. Sci. USA 1987, 84, 837–841. [Google Scholar] [CrossRef]
- Hachim, I.Y.; Hachim, M.Y.; López-Ozuna, V.M.; Ali, S.; Lebrun, J.J. A dual prognostic role for the TGFβ receptors in human breast cancer. Hum. Pathol. 2016, 57, 140–151. [Google Scholar] [CrossRef]
- Hennighausen, L.; Robinson, G.W. Information networks in the mammary gland. Nat. Rev. Mol. Cell. Biol. 2005, 6, 715–725. [Google Scholar] [CrossRef] [PubMed]
- Ormandy, C.J.; Binart, N.; Kelly, P.A. Mammary gland development in prolactin receptor knockout mice. J. Mammary Gland Biol. Neoplasia 1997, 2, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Nouhi, Z.; Chughtai, N.; Hartley, S.; Cocolakis, E.; Lebrun, J.J.; Ali, S. Defining the role of prolactin as an invasion suppressor hormone in breast cancer cells. Cancer Res. 2006, 66, 1824–1832. [Google Scholar] [CrossRef]
- Sultan, A.S.; Xie, J.; LeBaron, M.J.; Ealley, E.L.; Nevalainen, M.T.; Rui, H. Stat5 promotes homotypic adhesion and inhibits invasive characteristics of human breast cancer cells. Oncogene 2005, 24, 746–760. [Google Scholar] [CrossRef] [PubMed]
- Sultan, A.S.; Brim, H.; Sherif, Z.A. Co-overexpression of Janus kinase 2 and signal transducer and activator of transcription 5a promotes differentiation of mammary cancer cells through reversal of epithelial-mesenchymal transition. Cancer Sci. 2008, 99, 272–279. [Google Scholar] [CrossRef]
- Hachim, I.Y.; Hachim, M.Y.; Lopez, V.M.; Lebrun, J.J.; Ali, S. Prolactin Receptor Expression is an Independent Favorable Prognostic Marker in Human Breast Cancer. Appl. Immunohistochem. Mol. Morphol. 2016, 24, 238–245. [Google Scholar] [CrossRef] [PubMed]
- Galsgaard, E.D.; Rasmussen, B.B.; Folkesson, C.G.; Rasmussen, L.M.; Berchtold, M.W.; Christensen, L.; Panina, S. Re-evaluation of the prolactin receptor expression in human breast cancer. J. Endocrinol. 2009, 201, 115–128. [Google Scholar] [CrossRef] [PubMed]
- Hachim, I.Y.; Shams, A.; Lebrun, J.J.; Ali, S. A Favorable Role of Prolactin in Human Breast Cancer Reveals Novel Pathway Based Gene Signatures Indicative of Tumor Differentiation and Favorable Patient Outcome. Hum. Pathol. 2016, 53. [Google Scholar] [CrossRef]
- López-Ozuna, V.M.; Hachim, I.Y.; Hachim, M.Y.; Lebrun, J.-J.; Ali, S. Prolactin Pro-Differentiation Pathway in Triple Negative Breast Cancer: Impact on Prognosis and Potential Therapy. Sci. Rep. 2016, 6, 30934. [Google Scholar] [CrossRef]
- Peck, A.R.; Witkiewicz, A.K.; Liu, C.; Klimowicz, A.C.; Stringer, G.A.; Pequignot, E.; Freydin, B.; Yang, N.; Ertel, A.; Tran, T.H.; et al. Low levels of Stat5a protein in breast cancer are associated with tumor progression and unfavorable clinical outcomes. Breast Cancer Res. 2012, 14, R130. [Google Scholar] [CrossRef]
- Nevalainen, M.T.; Xie, J.; Torhorst, J.; Bubendorf, L.; Haas, P.; Kononen, J.; Sauter, G.; Rui, H. Signal transducer and activator of transcription-5 activation and breast cancer prognosis. J. Clin. Oncol. 2004, 22, 2053–2060. [Google Scholar] [CrossRef]
- Cocolakis, E.; Dai, M.; Drevet, L.; Ho, J.; Haines, E.; Ali, S.; Lebrun, J.J. Smad signaling antagonizes STAT5-mediated gene transcription and mammary epithelial cell differentiation. J. Biol. Chem. 2008, 283, 1293–1307. [Google Scholar] [CrossRef]
- Hachim, I.Y.; Villatoro, M.; Canaff, L.; Hachim, M.Y.; Boudreault, J.; Haiub, H.; Ali, S.; Lebrun, J. Transforming Growth Factor-β Regulation of Ephrin Type-A Receptor 4 Signaling in Breast Cancer Cellular Migration. Sci. Rep. 2017, 7, 14976. [Google Scholar] [CrossRef] [PubMed]
- Brabletz, T. To differentiate or not—routes towards metastasis. Nat. Rev. Cancer 2012, 12, 425–436. [Google Scholar] [CrossRef]
- Nieto, M.A.; Huang, R.Y.; Jackson, R.A.; Thiery, J.P. Emt: 2016. Cell 2016, 166, 21–45. [Google Scholar] [CrossRef]
- Rakha, E.A.; Reis-Filho, J.S.; Baehner, F.; Dabbs, D.J.; Decker, T.; Eusebi, V.; Fox, S.B.; Ichihara, S.; Jacquemier, J.; Lakhani, S.R.; et al. Breast cancer prognostic classification in the molecular era: The role of histological grade. Breast Cancer Res. 2010, 12, 207. [Google Scholar] [CrossRef]
- Wang, H.; Unternaehrer, J.J. Epithelial-mesenchymal Transition and Cancer Stem Cells: At the Crossroads of Differentiation and Dedifferentiation. Dev. Dyn. 2019, 248, 10–20. [Google Scholar] [CrossRef]
- Goldhirsch, A.; Wood, W.C.; Coates, A.S.; Gelber, R.D.; Thürlimann, B.; Senn, H.J. Panel members. Strategies for subtypes—dealing with the diversity of breast cancer: Highlights of the St. Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann. Oncol. 2011, 22, 1736–1747. [Google Scholar] [CrossRef]
- Ringnér, M.; Fredlund, E.; Häkkinen, J.; Borg, Å.; Staaf, J. GOBO: Gene expression-based outcome for breast cancer online. PLoS ONE 2011, 6, e17911. [Google Scholar] [CrossRef]


| PRLR | PRLR | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| TGFβRII-positive cases | Negative | Positive | % co-expression | p-value | TGFβRI-positive cases | Negative | Positive | % co-expression | p-value |
| Normal vs. malignant cases | |||||||||
| Normal and benign | 1 | 3 | 75% | 0.049 | 1 | 4 | 80% | 0.021 | |
| Invasive Ca | 46 | 18 | 28.12% | 43 | 18 | 29.50% | |||
| Histological grade | |||||||||
| Grade I | 7 | 3 | 30% | 0.67 | 8 | 2 | 20% | 0.72 | |
| Grade II | 29 | 10 | 25.64% | 29 | 10 | 25.6% | |||
| Grade III | 11 | 2 | 15.38% | 11 | 2 | 15.38% | |||
| LN involvement | |||||||||
| LN-negative | 23 | 13 | 36.11% | 0.17 | 21 | 13 | 38.23% | 0.38 | |
| LN-positive | 26 | 7 | 21.21% | 23 | 9 | 28.12% | |||
| Tumor size | |||||||||
| T1 and T2 | 22 | 9 | 29.03% | 0.76 | 21 | 9 | 30% | 0.71 | |
| T3 and T4 | 26 | 9 | 25.71% | 23 | 8 | 25.80% | |||
| Tumor stage | |||||||||
| Stages 1 and 2 | 26 | 12 | 31.57% | 0.31 | 25 | 13 | 34.21% | 0.38 | |
| Stages 3 and 4 | 23 | 6 | 20.68% | 19 | 6 | 24% | |||
| Breast cancer subtype | |||||||||
| Luminal | 23 | 10 | 30.3% | 0.29 | 19 | 10 | 34.48% | 0.34 | |
| HER-2 | 13 | 5 | 27.77% | 13 | 5 | 27.77% | |||
| TNBC | 6 | 0 | 0% | 6 | 0 | 0% | |||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hachim, I.Y.; López-Ozuna, V.M.; Hachim, M.Y.; Lebrun, J.-J.; Ali, S. Concomitant Expression of Prolactin Receptor and TGFβ Receptors in Breast Cancer: Association with Less Aggressive Phenotype and Favorable Patient Outcome. Int. J. Mol. Sci. 2019, 20, 1640. https://doi.org/10.3390/ijms20071640
Hachim IY, López-Ozuna VM, Hachim MY, Lebrun J-J, Ali S. Concomitant Expression of Prolactin Receptor and TGFβ Receptors in Breast Cancer: Association with Less Aggressive Phenotype and Favorable Patient Outcome. International Journal of Molecular Sciences. 2019; 20(7):1640. https://doi.org/10.3390/ijms20071640
Chicago/Turabian StyleHachim, Ibrahim Y., Vanessa M. López-Ozuna, Mahmood Y. Hachim, Jean-Jacques Lebrun, and Suhad Ali. 2019. "Concomitant Expression of Prolactin Receptor and TGFβ Receptors in Breast Cancer: Association with Less Aggressive Phenotype and Favorable Patient Outcome" International Journal of Molecular Sciences 20, no. 7: 1640. https://doi.org/10.3390/ijms20071640
APA StyleHachim, I. Y., López-Ozuna, V. M., Hachim, M. Y., Lebrun, J.-J., & Ali, S. (2019). Concomitant Expression of Prolactin Receptor and TGFβ Receptors in Breast Cancer: Association with Less Aggressive Phenotype and Favorable Patient Outcome. International Journal of Molecular Sciences, 20(7), 1640. https://doi.org/10.3390/ijms20071640






