Ceramide Imbalance and Impaired TLR4-Mediated Autophagy in BMDM of an ORMDL3-Overexpressing Mouse Model
Abstract
1. Introduction
2. Results
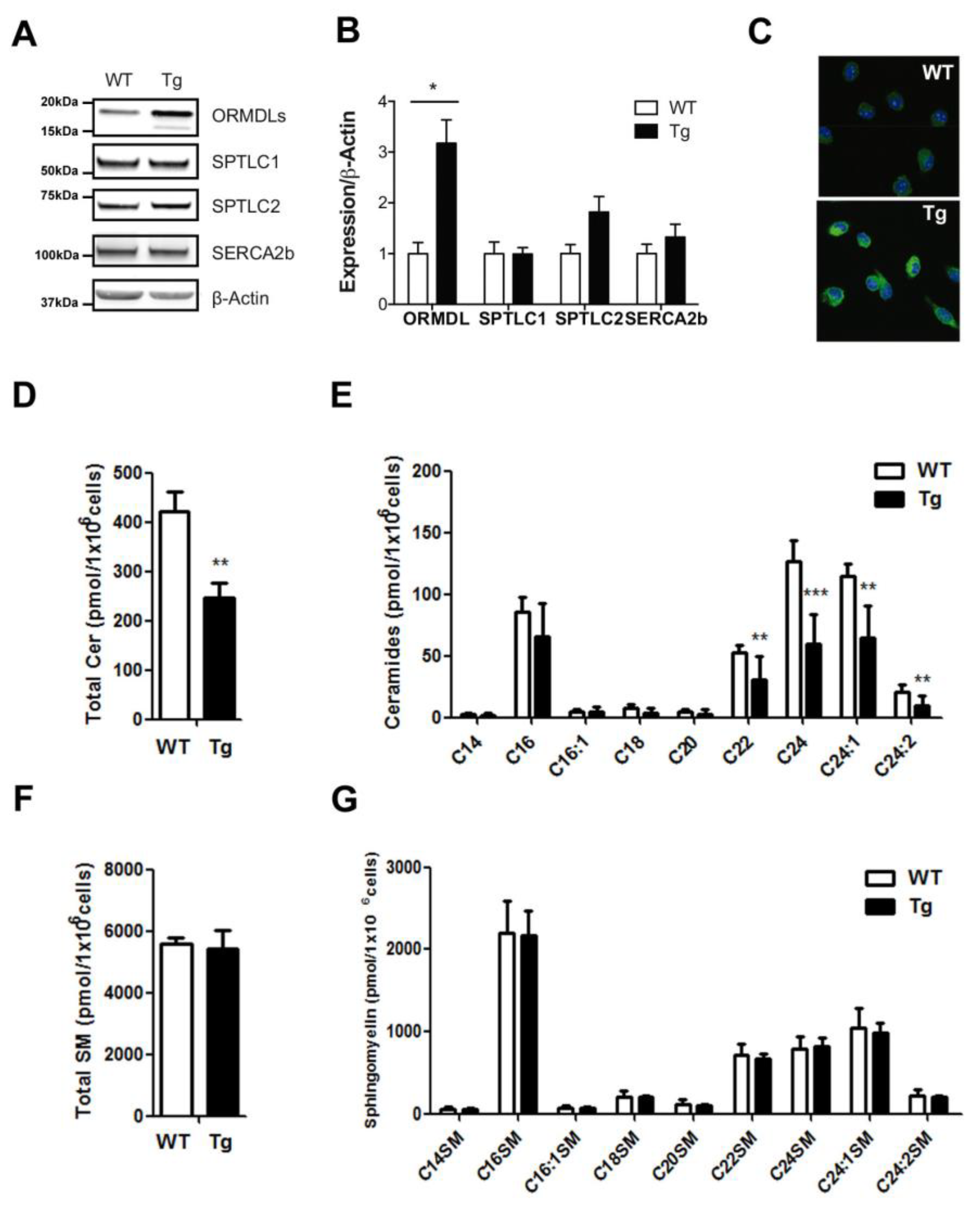
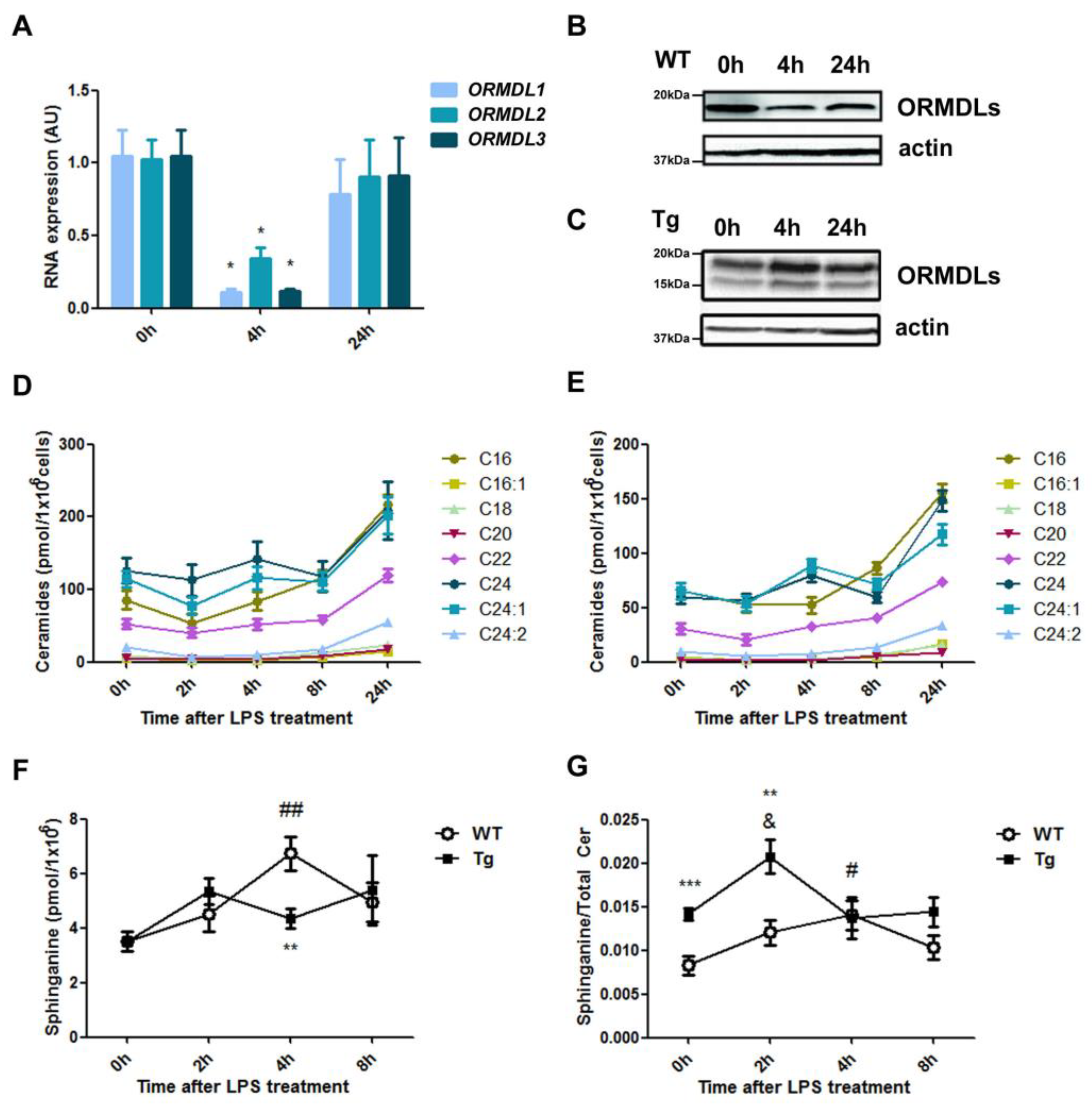
2.1. Ceramide Composition in BMDM from hORMDL3Rosa26 Mice
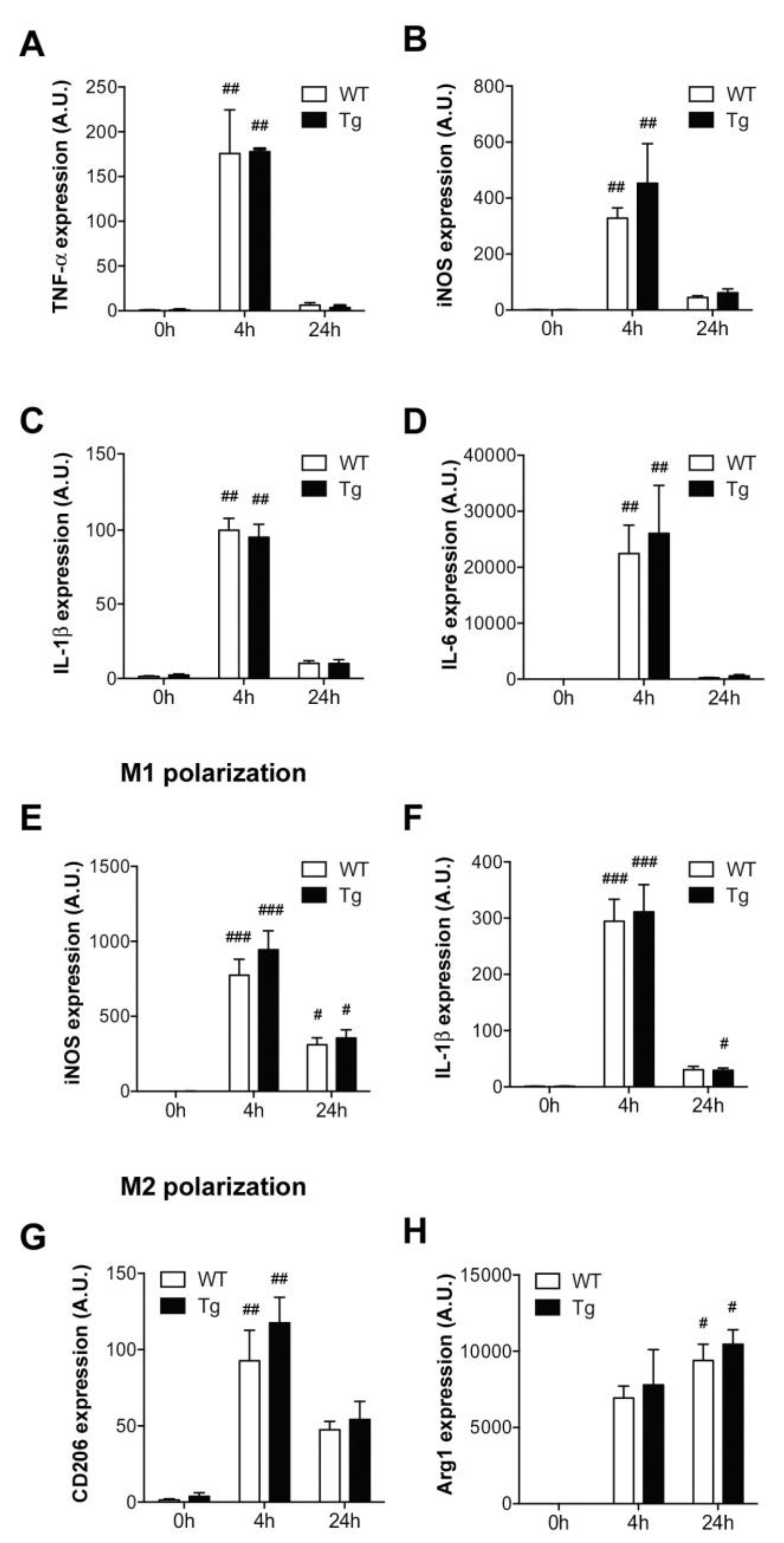
2.2. Macrophage Activation and M1/M2 Polarization in hORMDL3Rosa26 Mice
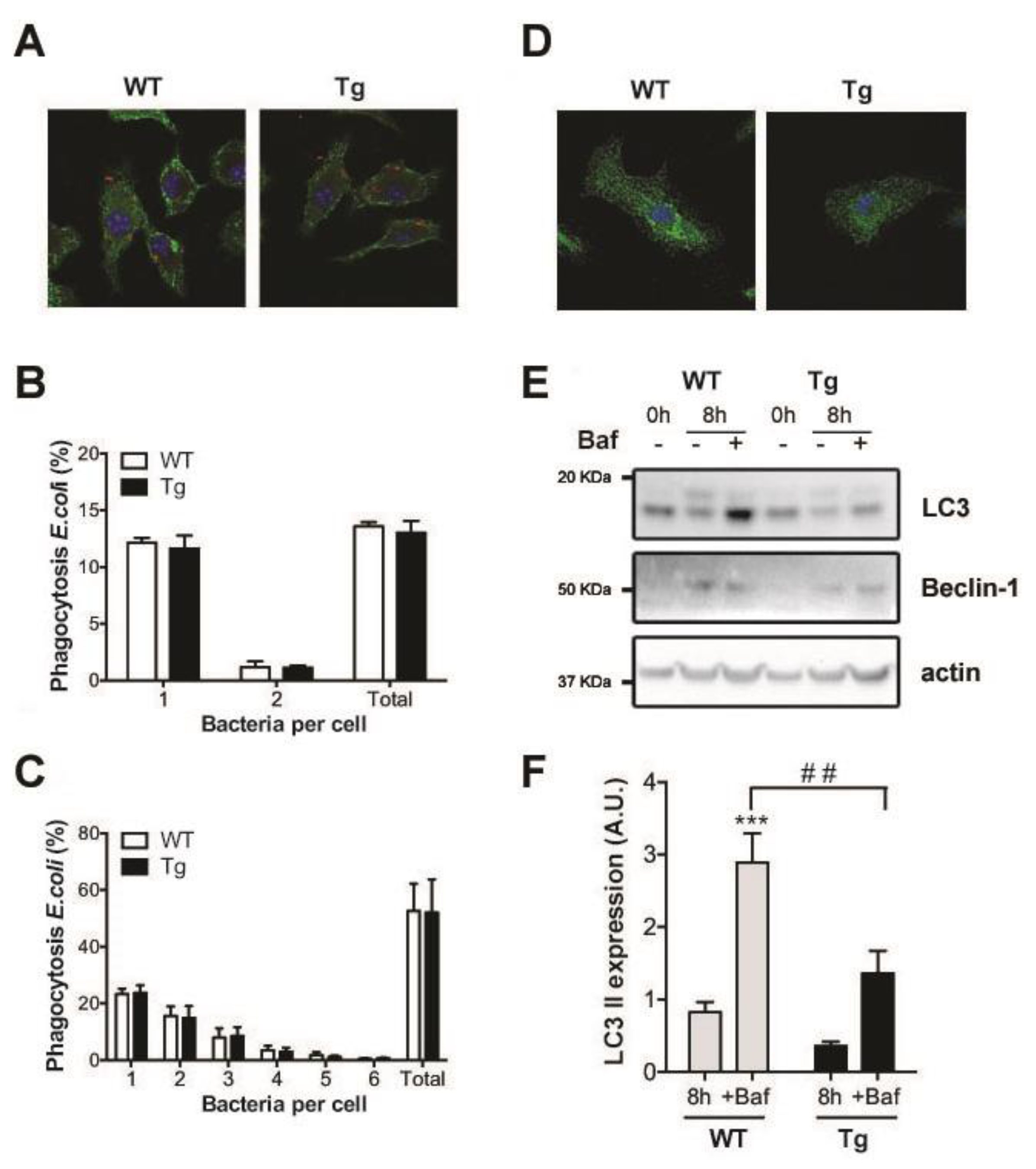
2.3. Phagocytosis and Autophagy in hORMDL3Rosa26 Mice
3. Discussion
4. Materials and Methods
4.1. hORMDL3Rosa26 Mouse Generation
4.2. Murine Bone-Marrow-Derived Macrophages Isolation and Differentiation
4.3. In-Vitro Activation and Polarization
4.4. Quantitative Real-Time (RT) Polymerase Chain Reaction (PCR) Analysis
4.5. Ceramide Quantification
4.6. Western Blot
4.7. Phagocytosis and Fluorescence Microscopy
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Barrett, J.C.; Hansoul, S.; Nicolae, D.L.; Cho, J.H.; Duerr, R.H.; Rioux, J.D.; Brant, S.R.; Silverberg, M.S.; Taylor, K.D.; Barmada, M.M.; et al. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn’s disease. Nat. Genet. 2008, 40, 955–962. [Google Scholar] [CrossRef] [PubMed]
- McGovern, D.P.B.; Gardet, A.; Törkvist, L.; Goyette, P.; Essers, J.; Taylor, K.D.; Neale, B.M.; Ong, R.T.H.; Lagacé, C.; Li, C.; et al. Genome-wide association identifies multiple ulcerative colitis susceptibility loci. Nat. Genet. 2010, 42, 332–337. [Google Scholar] [CrossRef]
- Kurreeman, F.A.S.; Stahl, E.A.; Okada, Y.; Liao, K.; Diogo, D.; Raychaudhuri, S.; Freudenberg, J.; Kochi, Y.; Patsopoulos, N.A.; Gupta, N.; et al. Use of a multiethnic approach to identify rheumatoid- arthritis-susceptibility loci, 1p36 and 17q12. Am. J. Hum. Genet. 2012, 90, 524–532. [Google Scholar] [CrossRef] [PubMed]
- Moffatt, M.F.; Kabesch, M.; Liang, L.; Dixon, A.L.; Strachan, D.; Heath, S.; Depner, M.; von Berg, A.; Bufe, A.; Rietschel, E.; et al. Genetic variants regulating ORMDL3 expression contribute to the risk of childhood asthma. Nature 2007, 448, 470–473. [Google Scholar] [CrossRef] [PubMed]
- Cantero-Recasens, G.; Fandos, C.; Rubio-Moscardo, F.; Valverde, M.A.; Vicente, R. The asthma-associated ORMDL3 gene product regulates endoplasmic reticulum-mediated calcium signaling and cellular stress. Hum. Mol. Genet. 2010, 19, 111–121. [Google Scholar] [CrossRef] [PubMed]
- Carreras-Sureda, A.; Cantero-Recasens, G.; Rubio-Moscardo, F.; Kiefer, K.; Peinelt, C.; Niemeyer, B.A.; Valverde, M.A.; Vicente, R. ORMDL3 modulates store-operated calcium entry and lymphocyte activation. Hum. Mol. Genet. 2013, 22, 519–530. [Google Scholar] [CrossRef]
- Carreras-Sureda, A.; Rubio-Moscardo, F.; Olvera, A.; Argilaguet, J.; Kiefer, K.; Mothe, B.; Meyerhans, A.; Brander, C.; Vicente, R. Lymphocyte Activation Dynamics Is Shaped by Hereditary Components at Chromosome Region 17q12-q21. PLoS ONE 2016, 11, e0166414. [Google Scholar] [CrossRef] [PubMed]
- Han, S.; Lone, M.A.; Schneiter, R.; Chang, A. Orm1 and Orm2 are conserved endoplasmic reticulum membrane proteins regulating lipid homeostasis and protein quality control. Proc. Natl. Acad. Sci. USA 2010, 107, 5851–5856. [Google Scholar] [CrossRef]
- Breslow, D.K.; Collins, S.R.; Bodenmiller, B.; Aebersold, R.; Simons, K.; Shevchenko, A.; Ejsing, C.S.; Weissman, J.S. Orm family proteins mediate sphingolipid homeostasis. Nature 2010, 463, 1048–1053. [Google Scholar] [CrossRef]
- Maceyka, M.; Spiegel, S. Sphingolipid metabolites in inflammatory disease. Nature 2014, 510, 58–67. [Google Scholar] [CrossRef]
- Siow, D.L.; Wattenberg, B.W. Mammalian ORMDL Proteins Mediate the Feedback Response in Ceramide Biosynthesis. J. Biol. Chem. 2012, 287, 40198–40204. [Google Scholar] [CrossRef] [PubMed]
- Kiefer, K.; Carreras-Sureda, A.; García-López, R.; Rubio-Moscardó, F.; Casas, J.; Fabriàs, G.; Vicente, R. Coordinated Regulation of the Orosomucoid-like Gene Family Expression Controls de Novo Ceramide Synthesis in Mammalian Cells. J. Biol. Chem. 2015, 290, 2822–2830. [Google Scholar] [CrossRef] [PubMed]
- Zhakupova, A.; Debeuf, N.; Krols, M.; Toussaint, W.; Vanhoutte, L.; Alecu, I.; Kutalik, Z.; Vollenweider, P.; Ernst, D.; von Eckardstein, A.; et al. ORMDL3 expression levels have no influence on the activity of serine palmitoyltransferase. FASEB J. 2016, 30, 4289–4300. [Google Scholar] [CrossRef] [PubMed]
- Oyeniran, C.; Sturgill, J.L.; Hait, N.C.; Huang, W.-C.; Avni, D.; Maceyka, M.; Newton, J.; Allegood, J.C.; Montpetit, A.; Conrad, D.H.; et al. Aberrant ORM (yeast)-like protein isoform 3 (ORMDL3) expression dysregulates ceramide homeostasis in cells and ceramide exacerbates allergic asthma in mice. J. Allergy Clin. Immunol. 2015, 136, 1035–1046. [Google Scholar] [CrossRef] [PubMed]
- Cai, L.; Oyeniran, C.; Biswas, D.D.; Allegood, J.; Milstien, S.; Kordula, T.; Maceyka, M.; Spiegel, S. ORMDL proteins regulate ceramide levels during sterile inflammation. J. Lipid Res. 2016, 57, 1412–1422. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Willis-Owen, S.A.G.; Spiegel, S.; Lloyd, C.M.; Moffatt, M.F.; Cookson, W.O.C.M. The ORMDL3 Asthma Gene Regulates ICAM1 and Has Multiple Effects on Cellular Inflammation. Am. J. Respir. Crit. Care Med. 2019, 199, 478–488. [Google Scholar] [CrossRef] [PubMed]
- Worgall, T.S.; Veerappan, A.; Sung, B.; Kim, B.I.; Weiner, E.; Bholah, R.; Silver, R.B.; Jiang, X.-C.; Worgall, S. Impaired sphingolipid synthesis in the respiratory tract induces airway hyperreactivity. Sci. Transl. Med. 2013, 5, 186ra67. [Google Scholar] [CrossRef]
- Miller, M.; Rosenthal, P.; Beppu, A.; Mueller, J.L.; Hoffman, H.M.; Tam, A.B.; Doherty, T.A.; McGeough, M.D.; Pena, C.A.; Suzukawa, M.; et al. ORMDL3 Transgenic Mice Have Increased Airway Remodeling and Airway Responsiveness Characteristic of Asthma. J. Immunol. 2014, 192, 3475–3487. [Google Scholar] [CrossRef]
- Schilling, J.D.; Machkovech, H.M.; He, L.; Sidhu, R.; Fujiwara, H.; Weber, K.; Ory, D.S.; Schaffer, J.E. Palmitate and lipopolysaccharide trigger synergistic ceramide production in primary macrophages. J. Biol. Chem. 2013, 288, 2923–2932. [Google Scholar] [CrossRef]
- Andreyev, A.Y.; Fahy, E.; Guan, Z.; Kelly, S.; Li, X.; McDonald, J.G.; Milne, S.; Myers, D.; Park, H.; Ryan, A.; et al. Subcellular organelle lipidomics in TLR-4-activated macrophages. J. Lipid Res. 2010, 51, 2785–2797. [Google Scholar] [CrossRef]
- Chang, Z.Q.; Lee, S.Y.; Kim, H.J.; Kim, J.R.; Kim, S.J.; Hong, I.K.; Oh, B.C.; Choi, C.S.; Goldberg, I.J.; Park, T.S. Endotoxin activates de novo sphingolipid biosynthesis via nuclear factor κB-mediated upregulation of Sptlc2. Prostaglandins Other Lipid Mediat. 2011, 94, 44–52. [Google Scholar] [CrossRef] [PubMed]
- Sims, K.; Haynes, C.A.; Kelly, S.; Allegood, J.C.; Wang, E.; Momin, A.; Leipelt, M.; Reichart, D.; Glass, C.K.; Sullards, M.C.; et al. Kdo2-lipid A, a TLR4-specific agonist, induces de novo sphingolipid biosynthesis in RAW264.7 macrophages, which is essential for induction of autophagy. J. Biol. Chem. 2010, 285, 38568–38579. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Valerio, M.; Bielawski, J. Fenretinide inhibited de novo ceramide synthesis and proinflammatory cytokines induced by Aggregatibacter actinomycetemcomitans. J. Lipid Res. 2013, 54, 189–201. [Google Scholar] [CrossRef] [PubMed]
- Tafesse, F.G.; Rashidfarrokhi, A.; Schmidt, F.I.; Freinkman, E.; Dougan, S.; Dougan, M.; Esteban, A.; Maruyama, T.; Strijbis, K.; Ploegh, H.L. Disruption of Sphingolipid Biosynthesis Blocks Phagocytosis of Candida albicans. PLOS Pathog. 2015, 11, e1005188. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Qiu, R.; Dang, J.; Li, J.; Hu, Q.; Shan, S.; Xin, Q.; Pan, W.; Bian, X.; Yuan, Q. ORMDL3 contributes to the risk of atherosclerosis in Chinese Han population and mediates oxidized low-density lipoprotein-induced autophagy in endothelial cells. Sci. Rep. 2015, 5, 17194. [Google Scholar] [CrossRef] [PubMed]
- Dang, J.; Bian, X.; Ma, X.; Li, J.; Long, F.; Shan, S.; Yuan, Q.; Xin, Q.; Li, Y.; Gao, F. ORMDL3 Facilitates the Survival of Splenic B Cells via an ATF6α–Endoplasmic Reticulum Stress–Beclin1 Autophagy Regulatory Pathway. J. Immunol. 2017, 199, 1647–1659. [Google Scholar] [CrossRef] [PubMed]
- Levine, B.; Mizushima, N.; Virgin, H.W. Autophagy in immunity and inflammation. Nature 2011, 469, 323–335. [Google Scholar] [CrossRef]
- Cadwell, K.; Liu, J.Y.; Brown, S.L.; Miyoshi, H.; Loh, J.; Lennerz, J.K.; Kishi, C.; Carrero, W.K.J.; Hunt, S.; Stone, C.D. A key role for autophagy and the autophagy gene Atg16l1 in mouse and human intestinal Paneth cells. Nature 2008, 456, 259–263. [Google Scholar] [CrossRef]
- Thachil, E.; Hugot, J.-P.; Arbeille, B.; Paris, R.; Grodet, A.; Peuchmaur, M.; Codogno, P.; Barreau, F.; Ogier-Denis, E.; Berrebi, D.; et al. Abnormal activation of autophagy-induced crinophagy in Paneth cells from patients with Crohn’s disease. Gastroenterology 2012, 142, 1097–1099. [Google Scholar] [CrossRef]
- Franke, A.; McGovern, D.P.B.; Barrett, J.C.; Wang, K.; Radford-Smith, G.L.; Ahmad, T.; Lees, C.W.; Balschun, T.; Lee, J.; Roberts, R.; et al. Genome-wide meta-analysis increases to 71 the number of confirmed Crohn’s disease susceptibility loci. Nat. Genet. 2010, 42, 1118–1125. [Google Scholar] [CrossRef]
- Matalonga, J.; Glaria, E.; Bresque, M.; Escande, C.; Carbó, J.M.; Kiefer, K.; Vicente, R.; León, T.E.; Beceiro, S.; Pascual-García, M.; et al. The Nuclear Receptor LXR Limits Bacterial Infection of Host Macrophages through a Mechanism that Impacts Cellular NAD Metabolism. Cell Rep. 2017, 18, 1241–1255. [Google Scholar] [CrossRef]
- Vicente, R.; Escalada, A.; Coma, M.; Fuster, G.; Sánchez-Tilló, E.; López-Iglesias, C.; Soler, C.; Solsona, C.; Celada, A.; Felipe, A. Differential Voltage-dependent K+ Channel Responses during Proliferation and Activation in Macrophages. J. Biol. Chem. 2003, 278, 46307–46320. [Google Scholar] [CrossRef]
- Munoz-Olaya, J.M.; Matabosch, X.; Bedia, C.; Egido-Gabás, M.; Casas, J.; Llebaria, A.; Delgado, A.; Fabriàs, G. Synthesis and biological activity of a novel inhibitor of dihydroceramide desaturase. ChemMedChem. 2008, 3, 946–953. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kiefer, K.; Casas, J.; García-López, R.; Vicente, R. Ceramide Imbalance and Impaired TLR4-Mediated Autophagy in BMDM of an ORMDL3-Overexpressing Mouse Model. Int. J. Mol. Sci. 2019, 20, 1391. https://doi.org/10.3390/ijms20061391
Kiefer K, Casas J, García-López R, Vicente R. Ceramide Imbalance and Impaired TLR4-Mediated Autophagy in BMDM of an ORMDL3-Overexpressing Mouse Model. International Journal of Molecular Sciences. 2019; 20(6):1391. https://doi.org/10.3390/ijms20061391
Chicago/Turabian StyleKiefer, Kerstin, Josefina Casas, Roberto García-López, and Rubén Vicente. 2019. "Ceramide Imbalance and Impaired TLR4-Mediated Autophagy in BMDM of an ORMDL3-Overexpressing Mouse Model" International Journal of Molecular Sciences 20, no. 6: 1391. https://doi.org/10.3390/ijms20061391
APA StyleKiefer, K., Casas, J., García-López, R., & Vicente, R. (2019). Ceramide Imbalance and Impaired TLR4-Mediated Autophagy in BMDM of an ORMDL3-Overexpressing Mouse Model. International Journal of Molecular Sciences, 20(6), 1391. https://doi.org/10.3390/ijms20061391




