Transcriptome Analyses Provide Novel Insights into Heat Stress Responses in Chieh-Qua (Benincasa hispida Cogn. var. Chieh-Qua How)
Abstract
1. Introduction
2. Results
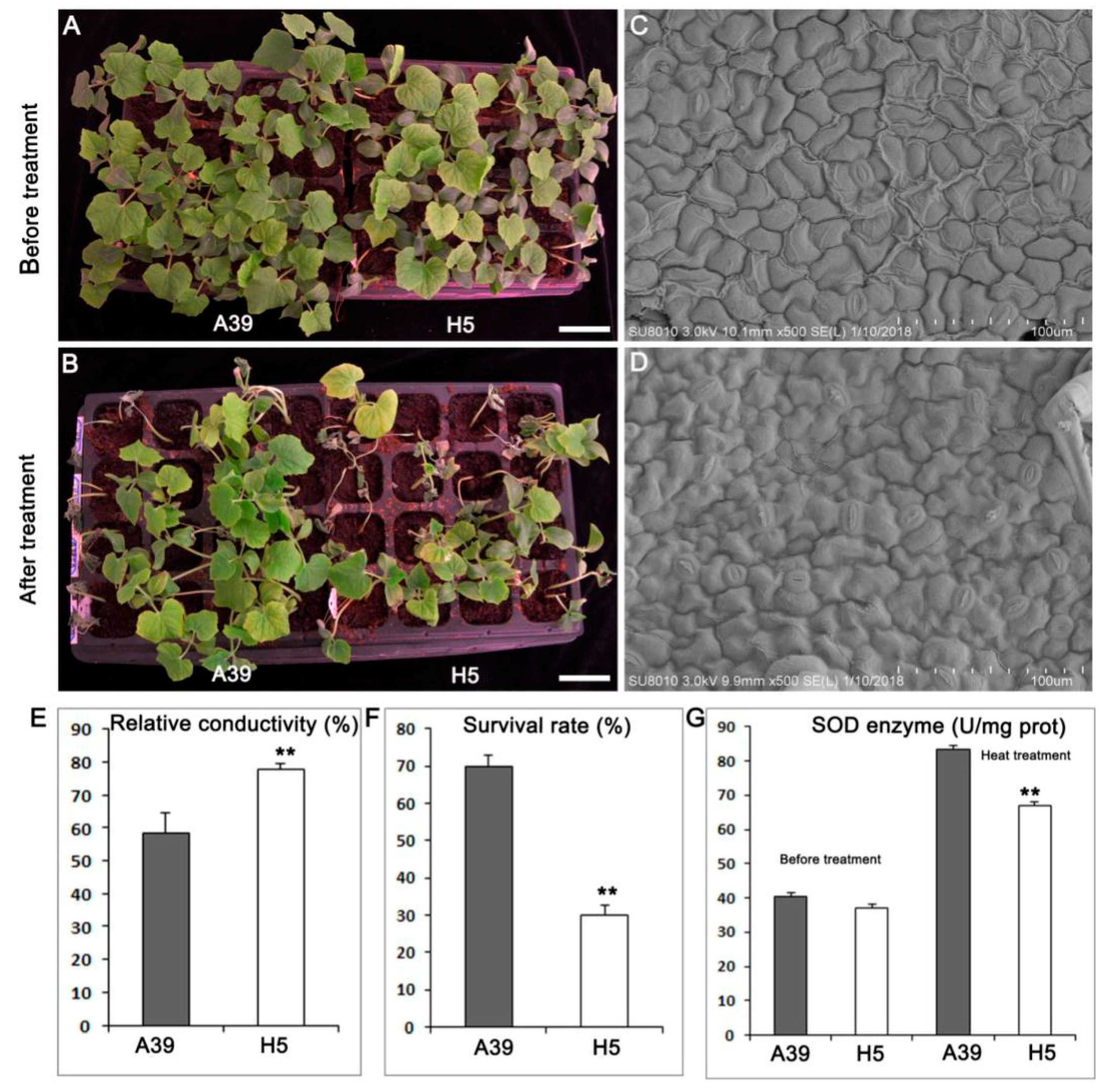
2.1. Phenotypes of A39 and H5 under Normal and Heat Conditions
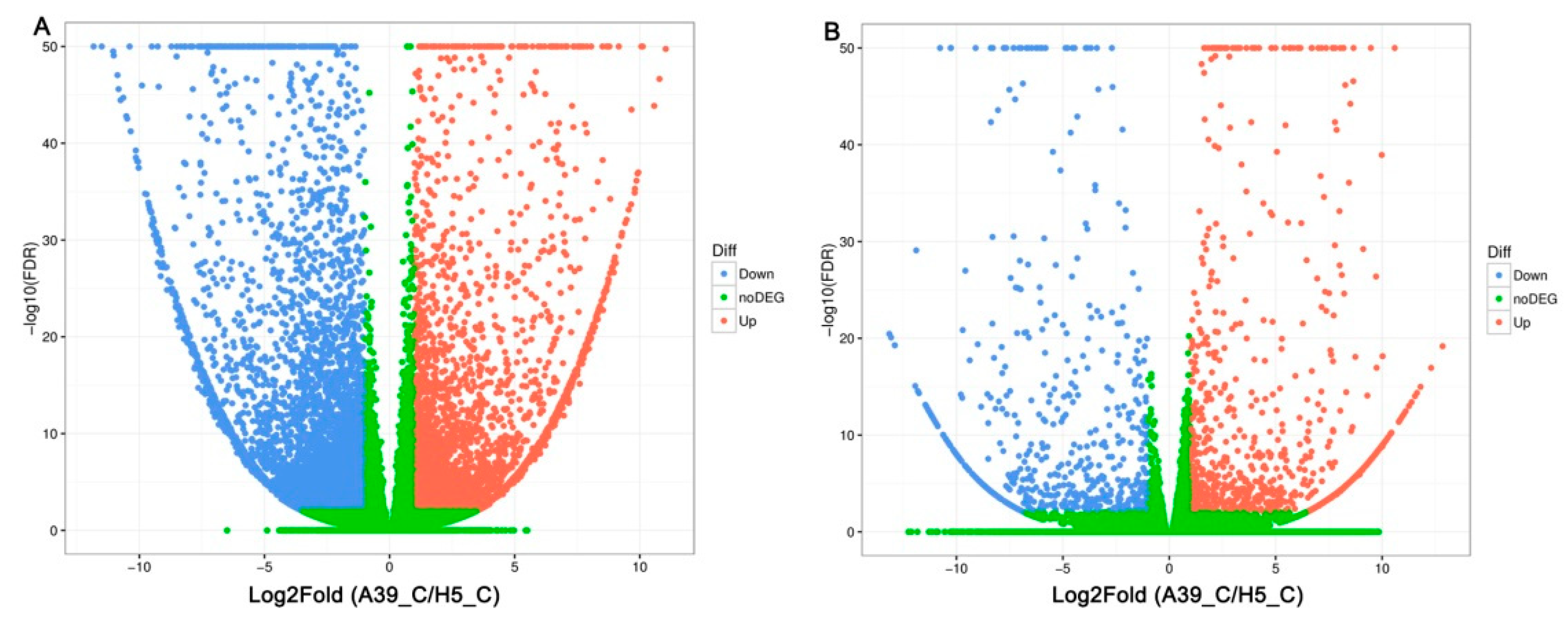
2.2. Transcripts Assembly of A39 and H5
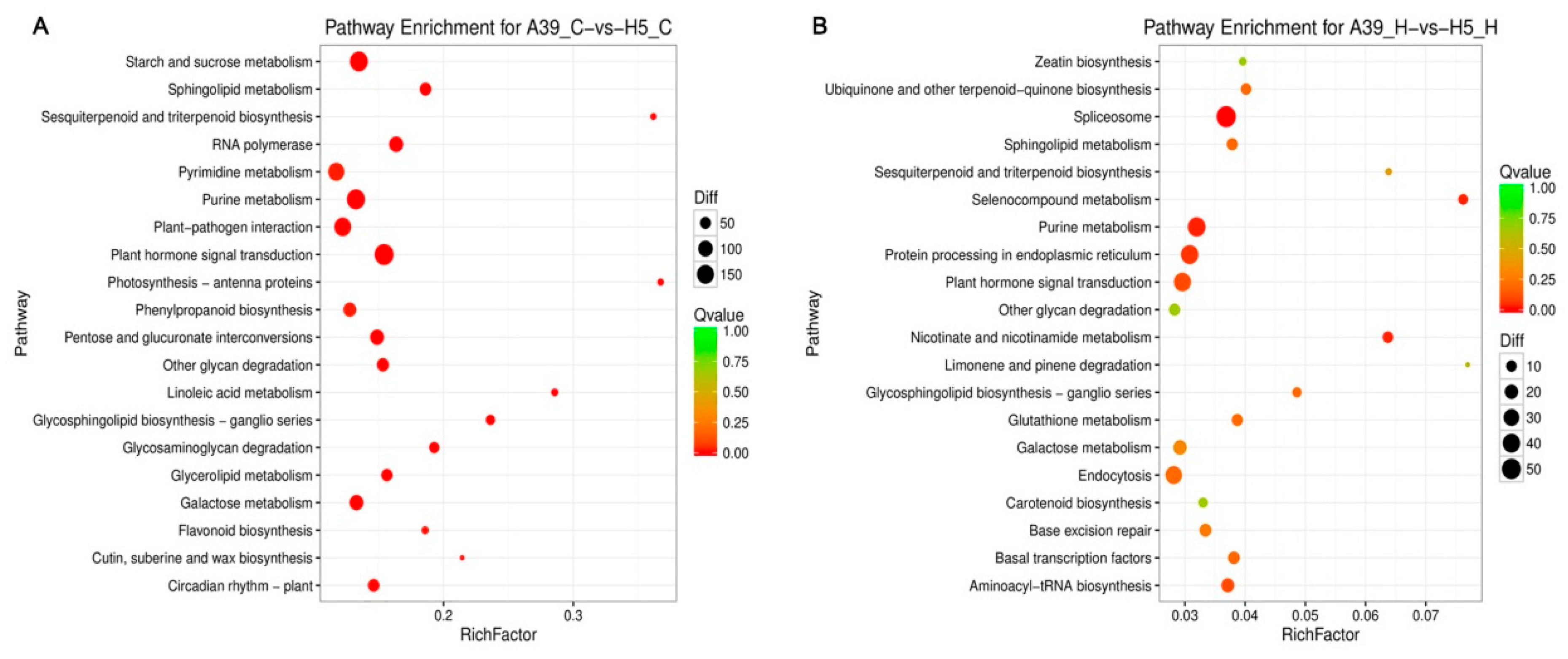
2.3. Functional Classification of Heat Response Genes
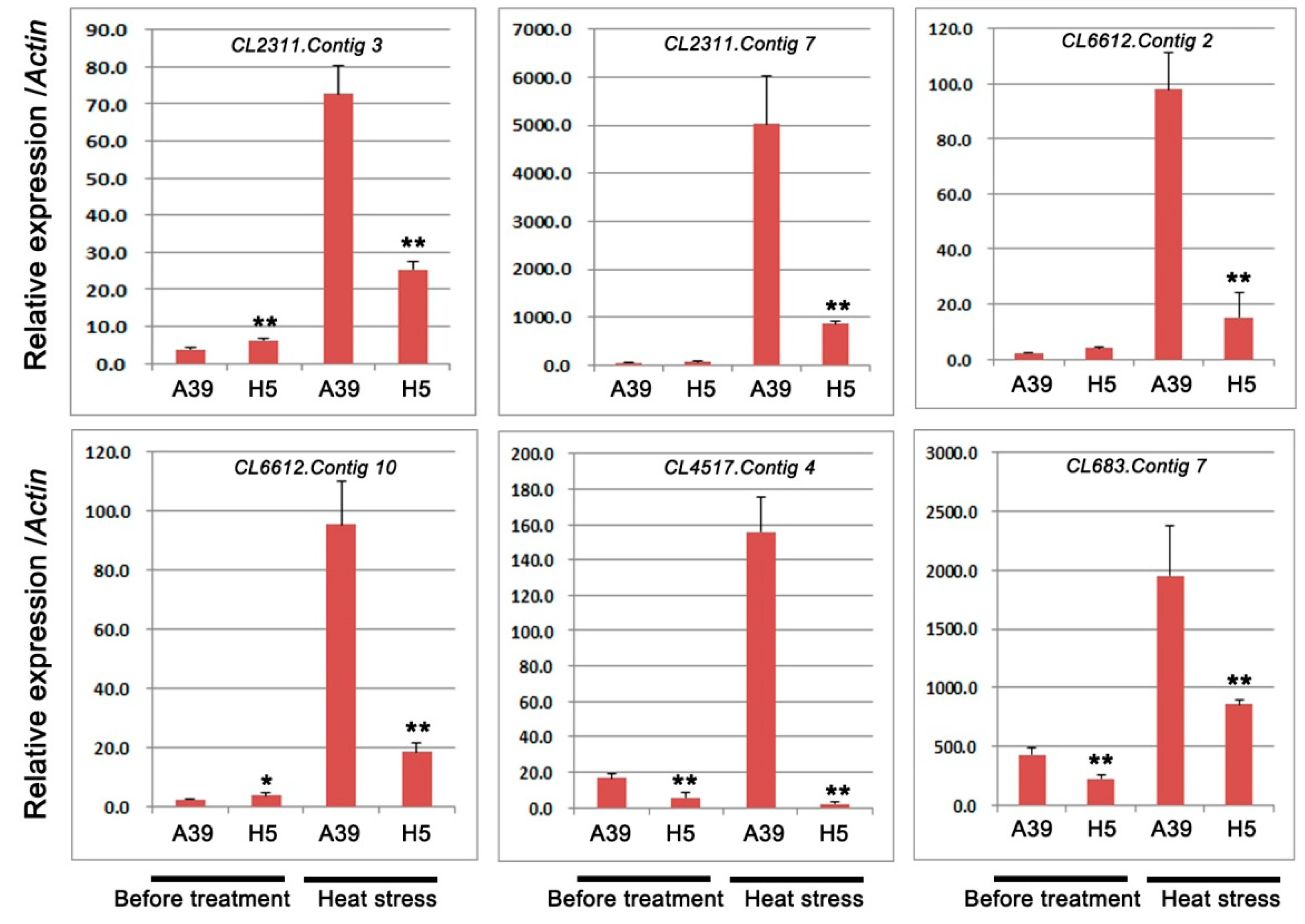
2.4. Expression of Genes Encoding HSPs, HSFs, and Cytochrome P450
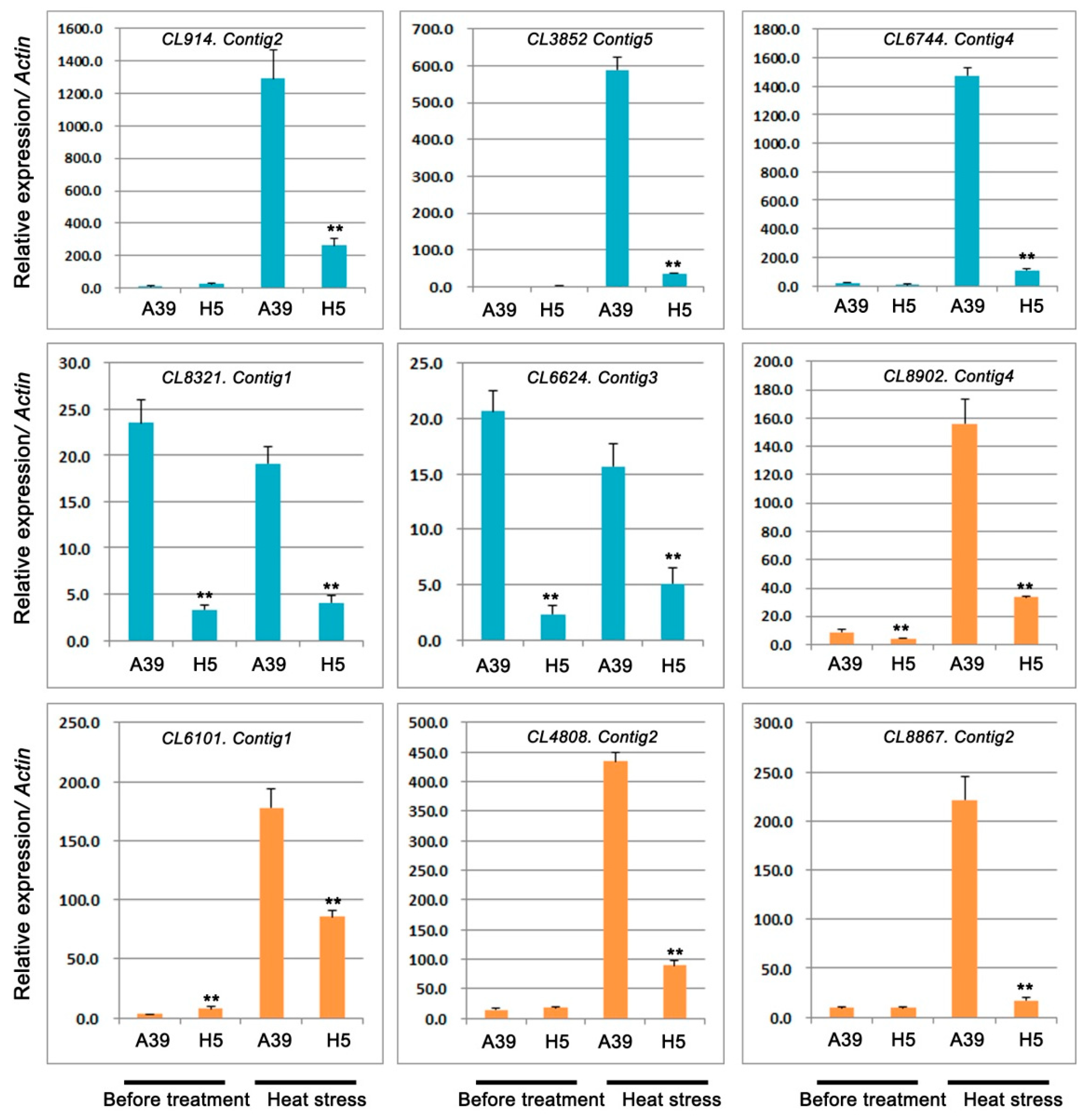
2.5. Expression Analysis of Genes Related to TFs and Pentatricopeptide Repeat-Containing Protein
3. Discussion
3.1. H5 Has More Stomas in the Leaf Than A39
3.2. Analysis of HSPs, HSFs During Heat Stress
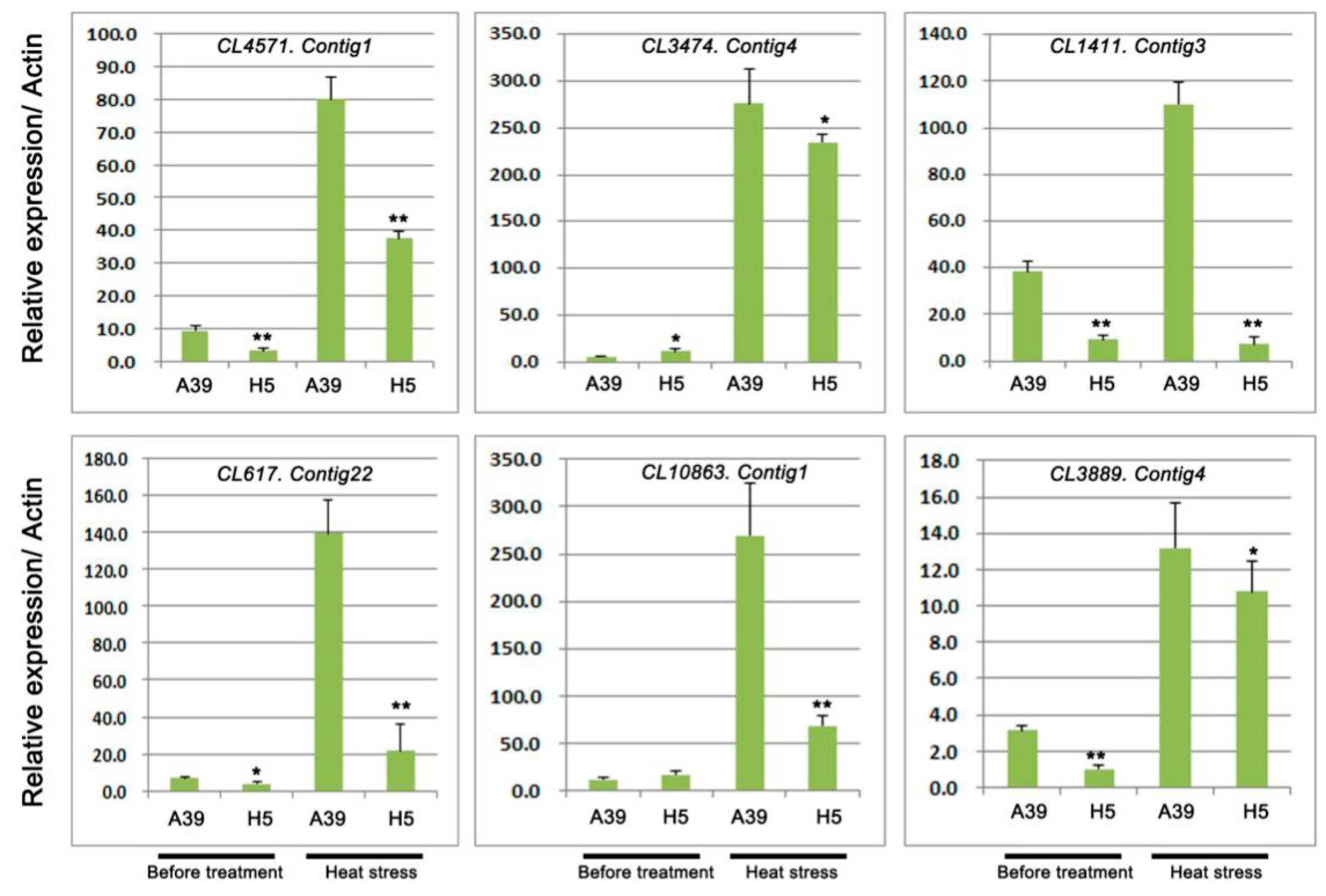
3.3. Analysis of Ubiquitin-Protein Ligase During Heat Stress
3.4. Analysis of TFs During Heat Stress
3.5. Analysis of PPRs During Heat Stress
3.6. Differences of Genes Expression under Control
4. Materials and Methods
4.1. Plant Materials and Heat Stress Treatment
4.2. Measurement of SOD Activity
4.3. Transcriptome Sequencing
4.4. Screening and Significant Test for Differentially Expressed Genes (DEGs)
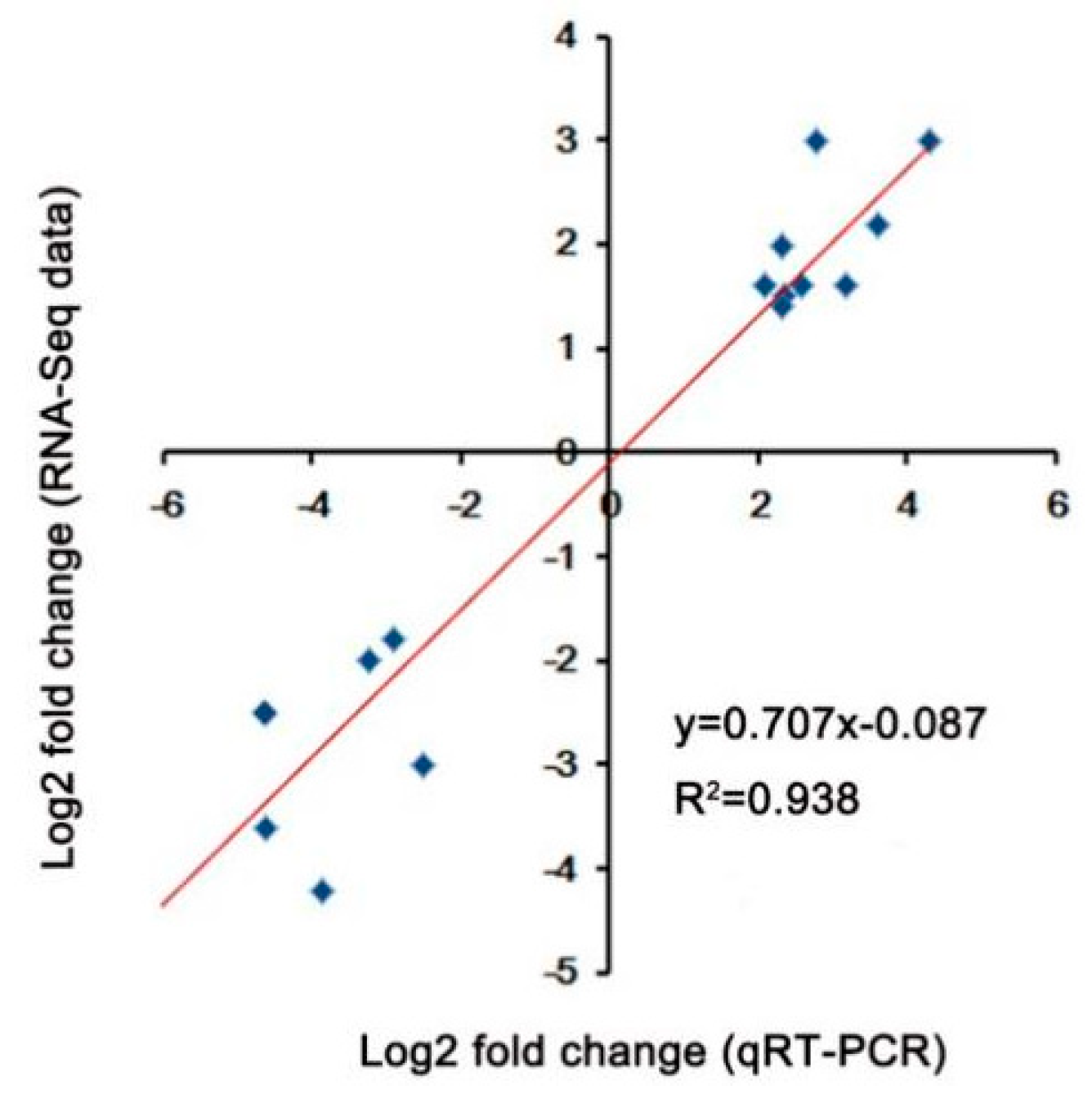
4.5. Quantitative Real-Time PCR Analysis
4.6. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Hall, A.E. Crop Responses to Environment; CRC Press LLC: Boca Raton, FL, USA, 2001. [Google Scholar]
- Ray, D.K.; Gerber, J.S.; MacDonald, G.K.; West, P.C. Climate variation explains a third of global crop yield variability. Nat. Commun. 2015, 6, 5989. [Google Scholar] [CrossRef]
- Wahid, A.; Gelani, S.; Ashraf, M.; Foolad, M.R. Heat tolerance in plants, an overview. Environ. Exp. Bot. 2007, 61, 199–223. [Google Scholar] [CrossRef]
- Liu, J.P.; Zhang, C.; Wei, C.; Liu, X.; Wang, M.G.; Yu, F.; Xie, Q.; Tu, J.M. The RING finger ubiquitin E3 ligase OsHTAS enhances heat tolerance by promoting H2O2-induced stomatal closure in rice. Plant Physiol. 2015, 170, 429. [Google Scholar] [CrossRef]
- Porter, J.R. Rising temperatures are likely to reduce crop yields. Nature 2005, 436, 174. [Google Scholar] [CrossRef]
- McClung, C.R.; Davis, S.J. Ambient thermometers in plants, from physiological outputs towards mechanisms of thermal sensing. Curr. Biol. 2010, 20, R1086–R1092. [Google Scholar] [CrossRef]
- Mittler, R.; Finka, A.; Goloubinoff, P. How do plants feel the heat? Trends Biochem. Sci. 2012, 37, 118–125. [Google Scholar] [CrossRef]
- Bita, C.E.; Gerats, T. Plant tolerance to high temperature in a changing environment, scientific fundamentals and production of heat stress-tolerant crops. Front Plant Sci. 2013, 4, 273. [Google Scholar] [CrossRef]
- Mayer, M.P. Recruitment of Hsp70 chaperones, a crucial part of viral survival strategies. Rev. Physiol. Biochem. Pharmacol. 2005, 153, 1–46. [Google Scholar]
- Tkáčová, J.; Angelovičová, M. Heat shock proteins (HSPs), a review. Lucrari Stiintifice Zootehnie Si Biotehnologii 2012, 45, 349–353. [Google Scholar]
- Chakraborty, K.; Bishi, S.K.; Singh, A.L.; Zala, P.V.; Mahatma, M.K.; Kalariya, K.A.; Jat, R.A. Rapid induction of small heat shock proteins improves physiological adaptation to high temperature stress in peanut. J. Agron. Crop Sci. 2018, 204, 285–297. [Google Scholar] [CrossRef]
- Larkindale, J.; Mishkind, M.; Vierling, E. Plant responses to high temperature. In Plant Abiotic Stress; Jenks, M.A., Hasegawa, P.M., Eds.; Blackwell Publishing: Hoboken, NJ, USA, 2005; pp. 100–144. [Google Scholar]
- Kotak, S.; Larkindale, J.; Lee, U.; von Koskull-Döring, P.; Vierling, E.; Scharf, K.D. Complexity of the heat stress response in plants. Current Opinion in Plant Biol. 2007, 10, 310–316. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.T.; Liu, Y.Y.; Pan, Q.H.; Yang, H.R.; Zhan, J.C.; Huang, W.D. Novel interrelationship between salicylic acid, abscisic acid, and pip2-specific phospholipase C in heat acclimation-induced thermotolerance in pea leaves. J. Exp. Bot. 2006, 57, 3337–3347. [Google Scholar] [CrossRef] [PubMed]
- Nakashima, K.; Yamaguchi-Shinozaki, K. ABA signaling in stress-response and seed development. Plant Cell Rep. 2013, 32, 959–970. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Wen, X.; Gong, H.; Lu, Q.; Yang, Z.; Tang, Y.; Liang, Z.; Lu, C. Genetic engineering of the biosynthesis of glycinebetaine enhances thermotolerance of photosystem II in tobacco plants. Planta 2007, 225, 719–733. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Duan, W.; Takabayashi, A.; Endo, T.; Shikanai, T.; Ye, J.Y.; Mi, H. Chloroplastic NAD(P)H dehydrogenase in tobacco leaves functions in alleviation of oxidative damage caused by temperature stress. Plant Physiol. 2006, 141, 465–474. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Lu, M.; Gong, Z.; Chen, R. Cloning and expression of a small heat shock protein gene CaHSP24 from pepper under abiotic stress. Afr. J. Biotechnol. 2011, 1025, 4968–4976. [Google Scholar]
- Yang, X.H.; Liang, Z.; Lu, C.M. Genetic engineering of the biosynthesis of glycinebetaine enhances photosynthesis against high temperature stress in transgenic tobacco plants. Plant Physiol. 2005, 138, 2299–2309. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Liu, S.S.; Yi, C.Y.; Wang, F.; Zhou, J.; Xia, X.J.; Shi, K.; Zhou, Y.H.; Yu, J.Q. Hydrogen peroxide mediates abscisic acid-induced HSP70 accumulation and heat tolerance in grafted cucumber plants. Plant Cell Environ. 2015, 37, 2768–2780. [Google Scholar] [CrossRef] [PubMed]
- Garg, R.; Shankar, R.; Thakkar, B.; Kudapa, H.; Krishnamurthy, L.; Mantri, N.; Varshney, R.K.; Bhatia, S.; Jain, M. Transcriptome analyses reveal genotype-and developmental stage-specific molecular responses to drought and salinity stresses in chickpea. Sci. Rep. 2016, 6, 19228. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Yang, P.; Cui, F.; Zhang, F.T.; Luo, X.D.; Xie, J.K. Transcriptome analysis of salt stress responsiveness in the seedlings of Dongxiang wild rice (Oryza rufipogon Griff.). PLoS ONE 2016, 11, e0146242. [Google Scholar] [CrossRef] [PubMed]
- Pawełkowicz, M.; Zieliński, K.; Zielińska, D.; Plader, W.; Yagi, K.; Wojcieszek, M.; Siedlecka, E.; Bartoszewski, G.; Skarzynska, A.; Przybecki, Z. Next generation sequencing and omics in cucumber (Cucumissativus L.) breeding directed research. Plant Sci. Int. J. Exp. Plant Biol. 2016, 242, 77–88. [Google Scholar]
- Grassi, S.; Piro, G.; Lee, J.M.; Zheng, Y.; Fei, Z.; Dalessandro, G.; Giovannoni, J.J.; Lenucci, M.S. Comparative genomics reveals candidate carotenoid pathway regulators of ripening watermelon fruit. BMC Genomics 2013, 14, 781. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.L.; Chen, B.H.; Chen, X.J.; Guo, Y.Y.; Yang, H.L.; Li, X.Z.; Wang, G.Y. Transcriptome profiling of pumpkin (CucurbitamoschataDuch.) leaves infected with powdery mildew. PLoS ONE 2018, 13, e0190175. [Google Scholar]
- Behera, T.K.; Rao, A.R.; Amarnath, R.; Kumar, R.R. Comparative transcriptome analysis of female and hermaphrodite flower buds in bitter gourd (Momordica charantia L.) by RNA sequencing. J. Pomol. Hortic. Sci. 2016, 91, 250–257. [Google Scholar] [CrossRef]
- He, X.M.; Xie, D.S.; Chen, Q.H.; Peng, Q.W. Chieh-qua biotechnology, progress and prospects. Asia and Aust. J. Sci. Biotechnol. 2007, 1, 19–22. [Google Scholar]
- Lim, C.W.; Baek, W.; Jung, J.; Kim, J.H.; Lee, S.C. Function of ABA in Stomatal Defense against Biotic and Drought Stresses. Int. J. Mol. Sci. 2015, 16, 15251–15270. [Google Scholar] [CrossRef]
- Sheng, P.; Tan, J.; Jin, M.; Wu, F.; Zhou, K.; Ma, W.; Heng, Y.; Wang, J.; Guo, X.; Zhang, X.; et al. Albino midrib 1, encoding a putative potassium efflux antiporter, affects chloroplast development and drought tolerance in rice. Plant Cell Rep. 2014, 33, 1581–1594. [Google Scholar] [CrossRef]
- Cui, L.G.; Shan, J.X.; Shi, M.; Gao, J.P.; Lin, H.X. DCA1 acts as a transcriptional co-activator of DST and contributes to drought and salt tolerance in rice. PLoS Genet. 2015, 11, e1005617. [Google Scholar] [CrossRef]
- Queitsch, C.; Hong, S.W.; Vierling, E.; Lindquist, S. Heat shock protein 101 plays a crucial role in thermotolerance in Arabidopsis. Plant Cell 2000, 12, 479–492. [Google Scholar] [CrossRef]
- Murakami, T.; Matsuba, S.H.; Kawaguchi, K.; Saruyama, H.; Tanida, M.; Sato, Y. Over-expression of a small heat shock protein, sHSP17.7, confers both heat tolerance and UV-B resistance to rice plants. Mol. Breed. 2004, 13, 165–175. [Google Scholar] [CrossRef]
- Liu, H.C.; Liao, H.T.; Charng, Y.Y. The role of class A1 heat shock factors (HSFA1s) in response to heat and other stresses in Arabidopsis. Plant Cell Environ. 2011, 34, 738–751. [Google Scholar] [CrossRef] [PubMed]
- Schramm, F.; Larkindale, J.; Kiehlmann, E.; Ganguli, A.; Englich, G.; Vierling, E.; von Koskull-Doring, P. A cascade of transcription factor DREB2A and heat stress transcription factor HsfA3 regulates the heat stress response of Arabidopsis. Plant J. 2008, 53, 264–274. [Google Scholar] [CrossRef] [PubMed]
- Smalle, J.; Vierstra, R.D. The ubiquitin 26S proteasome proteolytic pathway. Annu. Rev. Plant Biol. 2004, 55, 555–590. [Google Scholar] [CrossRef]
- Santner, A.; Estelle, M. The ubiquitin-proteasome system regulates plant hormone signaling. Plant J. 2010, 61, 1029–1040. [Google Scholar] [CrossRef] [PubMed]
- Dreher, K.; Callis, J. Ubiquitin, hormones and biotic stress in plants. Ann. Bot. (Lond.) 2007, 99, 787–822. [Google Scholar] [CrossRef] [PubMed]
- Vierstra, R.D. The ubiquitin-26S proteasome system at the nexus of plant biology. Nat. Rev. Mol. Cell Biol. 2009, 10, 385–397. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Wang, J.; Hwang, J.R.; Patterson, C.; Zhang, H. AtCHIP, a U-Box-containing E3 ubiquitin ligase, plays a critical role in temperature stress tolerance in Arabidopsis. Plant Physiol. 2003, 13, 861–869. [Google Scholar] [CrossRef]
- Zhang, Y.; Yang, C.; Li, Y.; Zheng, N.; Chen, H.; Zhao, Q.; Gao, T.; Guo, H.; Xie, Q. SDIR1 is a RING finger E3 ligase that positively regulates stress-responsive abscisic acid signaling in Arabidopsis. Plant Cell 2007, 19, 1912–1929. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Kim, H.Y. Functional analysis of a calcium-binding transcription factor involved in plant salt stress signaling. FEBS Lett. 2006, 580, 5251–5256. [Google Scholar] [CrossRef] [PubMed]
- George, C.; Witmer, J.A.; Cryer, J.D. Corrigendum to “Possible roles of basic helix-loop-helix transcription factors in adaptation to drought”. Plant Sci. 2014, 223, 1–7. [Google Scholar]
- Wang, Y.J.; Zhang, Z.G.; He, X.J.; Zhou, H.L.; Wen, Y.X.; Dai, J.X.; Zhang, J.S.; Chen, S.Y. A rice transcription factor OsbHLH1 is involved in cold stress response. Theor. Appl. Genet. 2003, 107, 1402–1409. [Google Scholar] [CrossRef] [PubMed]
- Kiribuchi, K.; Jikumaru, Y.; Kaku, H.; Minami, E.; Hasegawa, M.; Kodama, O.; Seto, H.; Okada, K.; Nojiri, H.; Yamane, H. Involvement of the basic helix-loophelix transcription factor RERJ1 in wounding and drought stress responses in rice plants. Biosci. Biotechnol. Biochem. 2005, 69, 1042–1044. [Google Scholar] [CrossRef] [PubMed]
- Yi, K.; Wu, Z.; Zhou, J.; Du, L.; Guo, L.; Wu, Y.; Wu, P. OsPTF1, a novel transcription factor involved in tolerance to phosphate starvation in rice. Plant Physiol. 2005, 138, 2087–2096. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Li, F.; Wang, J.L.; Ma, Y.; Chong, K.; Xu, Y. Basic helix-loop-helix transcription factor from wild rice (OrbHLH2) improves tolerance to salt and osmotic stress in Arabidopsis. J. Plant Physiol. 2009, 166, 1296–1306. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Jiang, W.; Dai, Y.; Xiao, N.; Zhang, C.; Li, H.; Lu, Y.; Wu, M.Q.; Tao, X.Y.; Deng, D.X.; Chen, J.M. A maize phytochrome-interacting factor 3 improves drought and salt stress tolerance in rice. Plant Mol. Biol. 2015, 87, 413–428. [Google Scholar] [CrossRef] [PubMed]
- Kudo, M.; Kidokoro, S.; Yoshida, T.; Mizoi, J.; Todaka, D.; Fernie, A.R.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Double overexpression of DREB and PIF transcription factors improves drought stress tolerance and cell elongation in transgenic plants. Plant Biotechnol. J. 2016, 15, 458–471. [Google Scholar] [CrossRef] [PubMed]
- Yu, Q.B.; Jiang, Y.; Chong, K.; Yang, Z.N. AtECB2, a pentatricopeptide repeat protein, is required for chloroplast transcript accD RNA editing and early chloroplast biogenesis in Arabidopsis thaliana. Plant J. 2009, 59, 1011–1023. [Google Scholar] [CrossRef]
- Ichinose, M.; Tasaki, E.; Sugita, C.; Sugita, M. A PPR-DYW protein is required for splicing of a group II intron of cox1 pre-mRNA in Physcomitrella patens. Plant Journal 2012, 70, 271–278. [Google Scholar] [CrossRef]
- Liu, X.; Lan, J.; Huang, Y.S.; Cao, P.H.; Zhou, C.L.; Ren, Y.; He, N.Q.; Liu, S.J.; Tian, Y.L.; Nguyen, T.; Jiang, L.; Wan, J.M. WSL5, a pentatricopeptide repeat protein, is essential for chloroplast biogenesis in rice under cold stress. J. Exp. Bot. 2018, 69, 3949–3961. [Google Scholar] [CrossRef]
- Oliveira, H.; Costa, A.; Santos, C. NaCl and Phaeomoniella chlamydospora affect differently starch and; sucrose metabolism in grapevines. Acta Sci. Agron. 2013, 35, 153–159. [Google Scholar] [CrossRef]
- Liao, W.B.; Li, Y.Y.; Lu, C.; Peng, M. Expression of sucrose metabolism and transport genes in cassava petiole abscission zones in response to water stress. Biol. Plant. 2017, 61, 219–226. [Google Scholar] [CrossRef]
- Chang, C.W.; Ryan, R.D. Effects of water stress on starch and sucrose metabolism in cotton leaves. Starch 1987, 39, 84–87. [Google Scholar] [CrossRef]
- Netrphan, S.; Tungngoen, K.; Suksangpanomrung, M.; Boonseng, O.; Narangajavana, J. Differential expression of genes involved in sucrose synthesis in source and sink organs of cassava plants undergoing seasonal drought stress. J. Agric. Sci. 2012, 4, 171–185. [Google Scholar]
- Giannopolitis, C.N.; Ries, S.K. Superoxide dismutase. Plant Physiol. 1977, 59, 309–314. [Google Scholar] [CrossRef] [PubMed]
- Paranidharan, V.; Palaniswami, A.; Vidhyasekaran, P.; Velazhahan, R. A host-specific toxin of Rhizoctonia solani triggers superoxide dismutase (SOD) activity in rice. Archiv für Pflanzenschutz 2005, 38, 151–156. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM, Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Fang, L.; Zheng, H.; Zhang, Y.; Chen, J.; Zhang, Z.; Wang, J.; Li, S.; Li, R.; Bolund, L.; et al. WEGO, A web tool for plotting GO annotations. Nucleic Acids Res. 2006, 34, W293. [Google Scholar] [CrossRef]
- Kanehisa, M.; Araki, M.; Goto, S.; Hattori, M.; Hirakawa, M.; Itoh, M.; Katayama, T.; Kawashima, S.; Okuda, S.; Tokimatsu, T.; et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 2008, 36, 480–484. [Google Scholar] [CrossRef]
- Vandesompele, J.; de Preter, K.; Pattyn, F.; Poppe, B.; van Roy, N.; de Paepe, A.; Speleman, F. Accuratenormalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal controlgenes. Genome Biol. 2002, 3, research0034.1. [Google Scholar] [CrossRef] [PubMed]
| Type | A39_C | H5_C | A39_H | H5_H |
|---|---|---|---|---|
| Read Length | 150 | 150 | 150 | 150 |
| Total Raw Reads | 45,345,693.33 | 54,886,327.3 | 47,159,128 | 50,066,083.33 |
| Total Raw Bases | 6,801,854,000 | 8,232,949,100 | 7,073,869,200 | 7,509,912,500 |
| Total Clean Reads | 45,233,028 | 54,676,526.67 | 47,003,104 | 49,889,418 |
| Total Clean Reads Ratio (%) | 99.75 | 99.63 | 99.67 | 99.65 |
| Total Clean Bases | 6,784,954,200 | 8,201,479,000 | 7,050,465,600 | 7,483,412,700 |
| Total Clean Bases Ratio (%) | 99.75 | 99.63 | 99.67 | 99.65 |
| Total Adapter Reads | 89,349.3 | 181,258.67 | 126,836.7 | 150,814 |
| Total Adapter Reads Ratio (%) | 0.2 | 0.32 | 0.27 | 0.3 |
| Total Low-Quality Reads | 23,316 | 28,542 | 29,187.3 | 25,851.33 |
| Total Low-Quality Reads Ratio (%) | 0.05 | 0.05 | 0.06 | 0.05 |
| Clean Reads GC (%) | 45.49 | 44.67 | 44.82 | 44.29 |
| Clean Reads Q20 (%) | 97.17 | 97.25 | 97.02 | 97.21 |
| Clean Reads Q30 (%) | 93.15 | 93.31 | 92.89 | 93.27 |
| GeneID | log2 Fold | p-Value | Diff | Nr-Annotation |
|---|---|---|---|---|
| CL2311.Contig3 | −10.26 | 9.62 × 10−267 | Down | heat shock protein (HSPs) |
| CL2311.Contig7 | −3.92 | 2.92 × 10−239 | Down | heat shock protein(HSPs) |
| CL6612.Contig2 | −11.46 | 7.01 × 10−16 | Down | heat stress transcription factor(HSFs) |
| CL6612.Contig10 | −4.79 | 5.23 × 10−6 | Down | heat stress transcription factor (HSFs) |
| CL4517.Contig4 | −1.02 | 2.8121 × 10−4 | Down | cytochrome P450 |
| CL683.Contig7 | −1.10 | 3.19 × 10−16 | Down | cytochrome P450 |
| GeneID | log2 Fold | p Value | Diff | Nr-Annotation |
|---|---|---|---|---|
| CL4571.Contig1 | −2.41 | 9.88 × 10−8 | Down | ubiquitin carboxyl-terminal hydrolase 2 |
| CL3474.Contig4 | −8.74 | 3.91 × 10−7 | Down | ubiquitin carboxyl-terminal hydrolase 14 |
| CL1411.Contig3 | −1.42 | 1.07 × 10−22 | Down | ubiquitin domain-containing protein DSK2a-like |
| CL617.Contig22 | −10.09 | 8.83 × 10−11 | Down | E3 ubiquitin-protein ligase UPL7 |
| CL10863.Contig1 | −8.26 | 3.16 × 10−6 | Down | E3 ubiquitin-protein ligase COP1 |
| CL3889.Contig4 | −8.03 | 9.51 × 10−6 | Down | ubiquitin carboxyl-terminal hydrolase 22-like |
| GeneID | log2 Fold | p Value | Diff | Nr-Annotation |
|---|---|---|---|---|
| CL914.Contig2 | −3.97 | 8.54 × 10−5 | Down | transcription factor bHLH128 |
| CL3852.Contig5 | −8.00 | 1.09 × 10−4 | Down | transcription factor PCL1-like |
| CL6744.Contig4 | −5.72 | 2.52 × 10−11 | Down | transcription factor PIF1-like |
| CL8321.Contig1 | −7.73 | 4.47 × 10−100 | Down | transcription factor bHLH143-like |
| CL6624.Contig3 | −8.54 | 5.01 × 10−17 | Down | transcription factor PIF1-like |
| CL8902.Contig4 | −8.49 | 1.33 × 10−5 | Down | pentatricopeptide repeat-containing protein |
| CL6101.Contig1 | −7.47 | 6.24 × 10−8 | Down | pentatricopeptide repeat-containing protein |
| CL4808.Contig2 | −3.06 | 7.6 × 10−12 | Down | pentatricopeptide repeat-containing protein |
| CL8867.Contig2 | −9.98 | 1.95 × 10−10 | Down | pentatricopeptide repeat-containing protein |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, M.; Jiang, B.; Liu, W.; Lin, Y.; Liang, Z.; He, X.; Peng, Q. Transcriptome Analyses Provide Novel Insights into Heat Stress Responses in Chieh-Qua (Benincasa hispida Cogn. var. Chieh-Qua How). Int. J. Mol. Sci. 2019, 20, 883. https://doi.org/10.3390/ijms20040883
Wang M, Jiang B, Liu W, Lin Y, Liang Z, He X, Peng Q. Transcriptome Analyses Provide Novel Insights into Heat Stress Responses in Chieh-Qua (Benincasa hispida Cogn. var. Chieh-Qua How). International Journal of Molecular Sciences. 2019; 20(4):883. https://doi.org/10.3390/ijms20040883
Chicago/Turabian StyleWang, Min, Biao Jiang, Wenrui Liu, Yu’e Lin, Zhaojun Liang, Xiaoming He, and Qingwu Peng. 2019. "Transcriptome Analyses Provide Novel Insights into Heat Stress Responses in Chieh-Qua (Benincasa hispida Cogn. var. Chieh-Qua How)" International Journal of Molecular Sciences 20, no. 4: 883. https://doi.org/10.3390/ijms20040883
APA StyleWang, M., Jiang, B., Liu, W., Lin, Y., Liang, Z., He, X., & Peng, Q. (2019). Transcriptome Analyses Provide Novel Insights into Heat Stress Responses in Chieh-Qua (Benincasa hispida Cogn. var. Chieh-Qua How). International Journal of Molecular Sciences, 20(4), 883. https://doi.org/10.3390/ijms20040883




