Genetic and Physiological Dissection of Photosynthesis in Barley Exposed to Drought Stress
Abstract
1. Introduction
2. Results
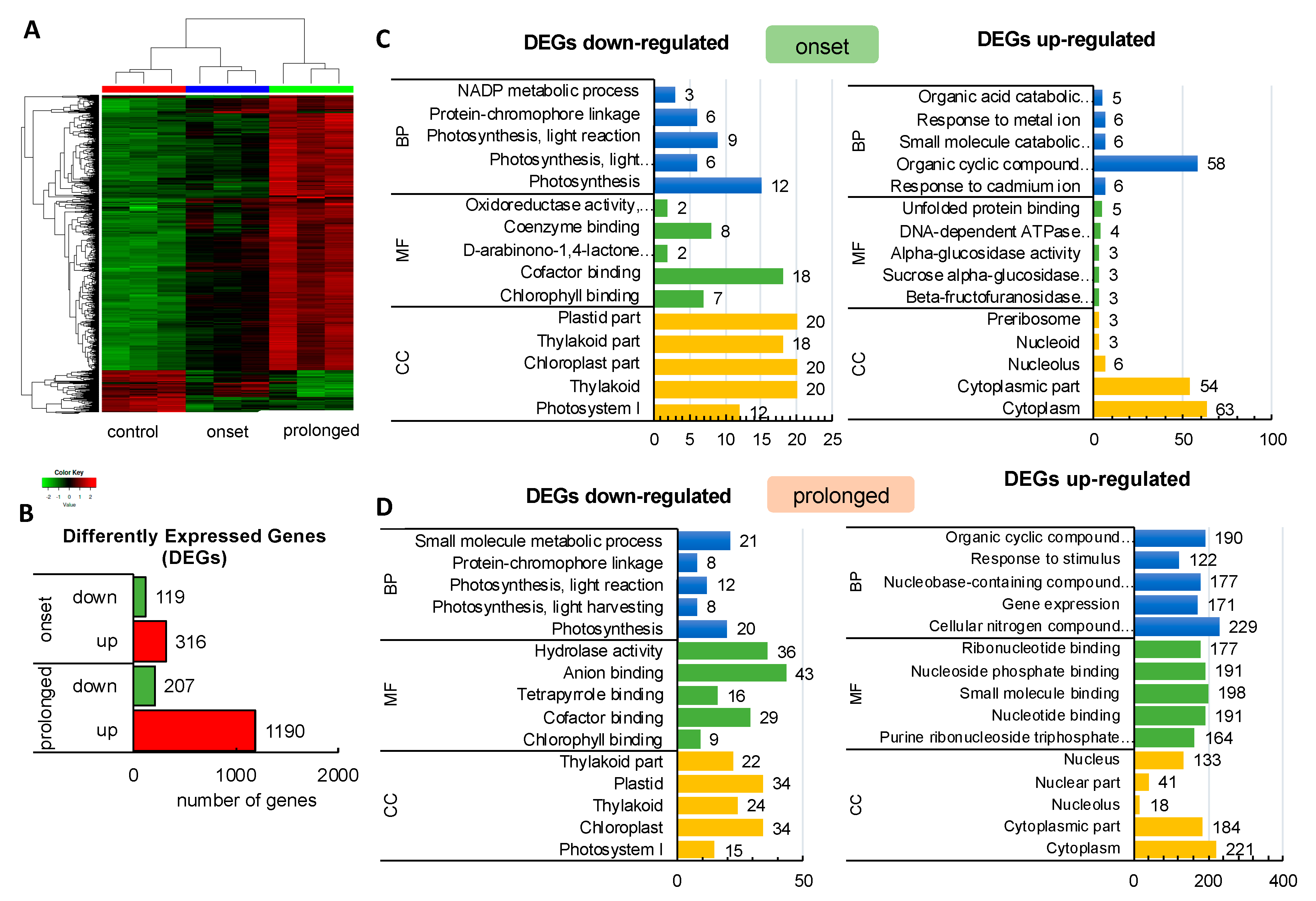
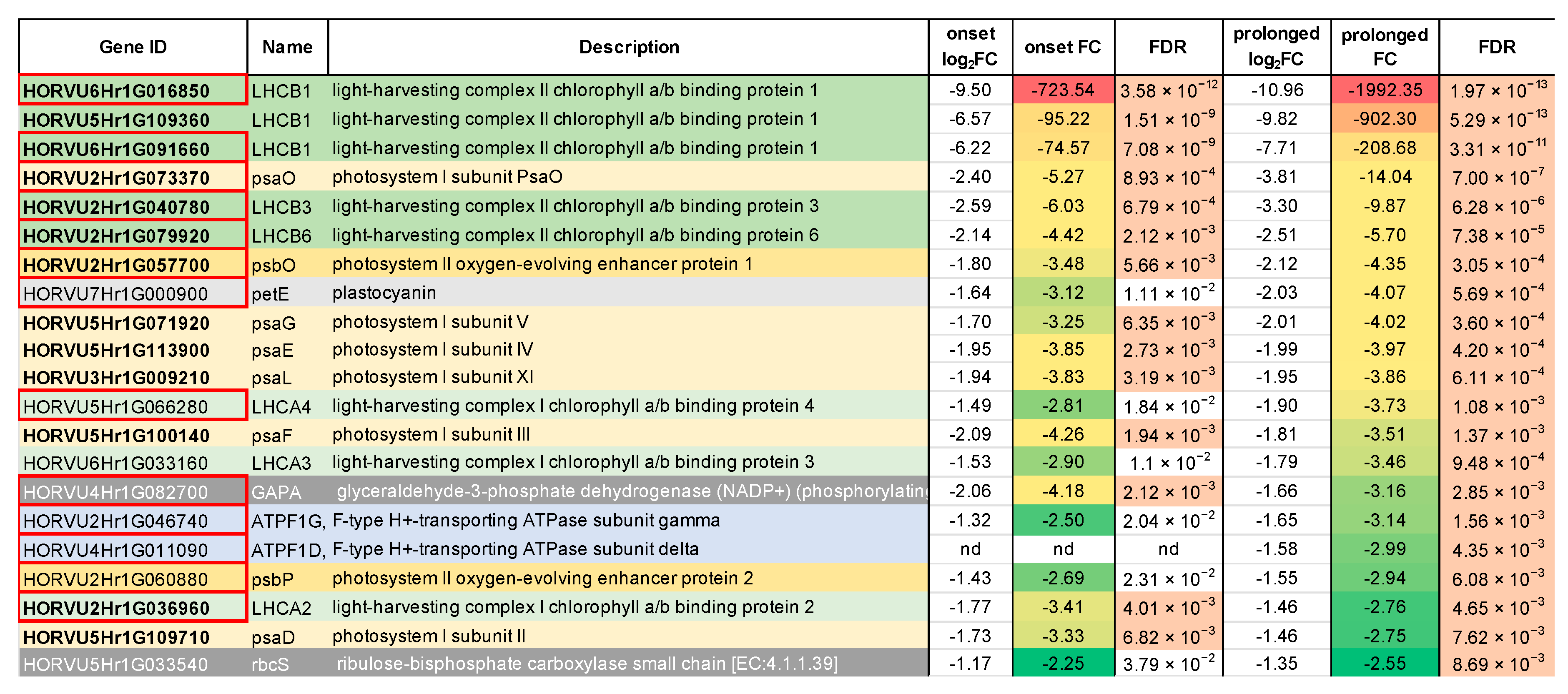
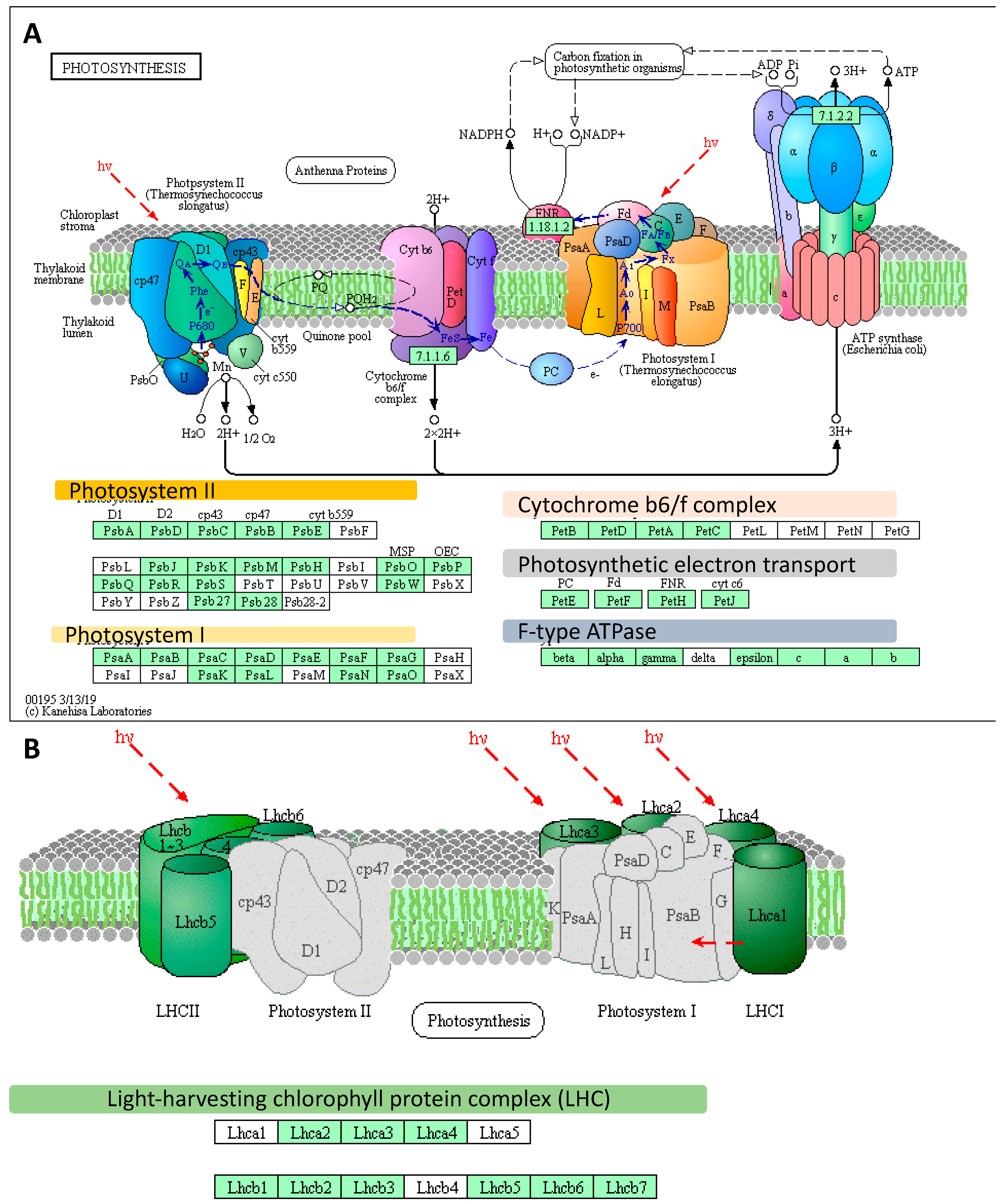
2.1. Transcriptomic Analysis Revealed Expression Changes of Photosynthesis-Related Genes Already Upon the Onset of Drought Stress
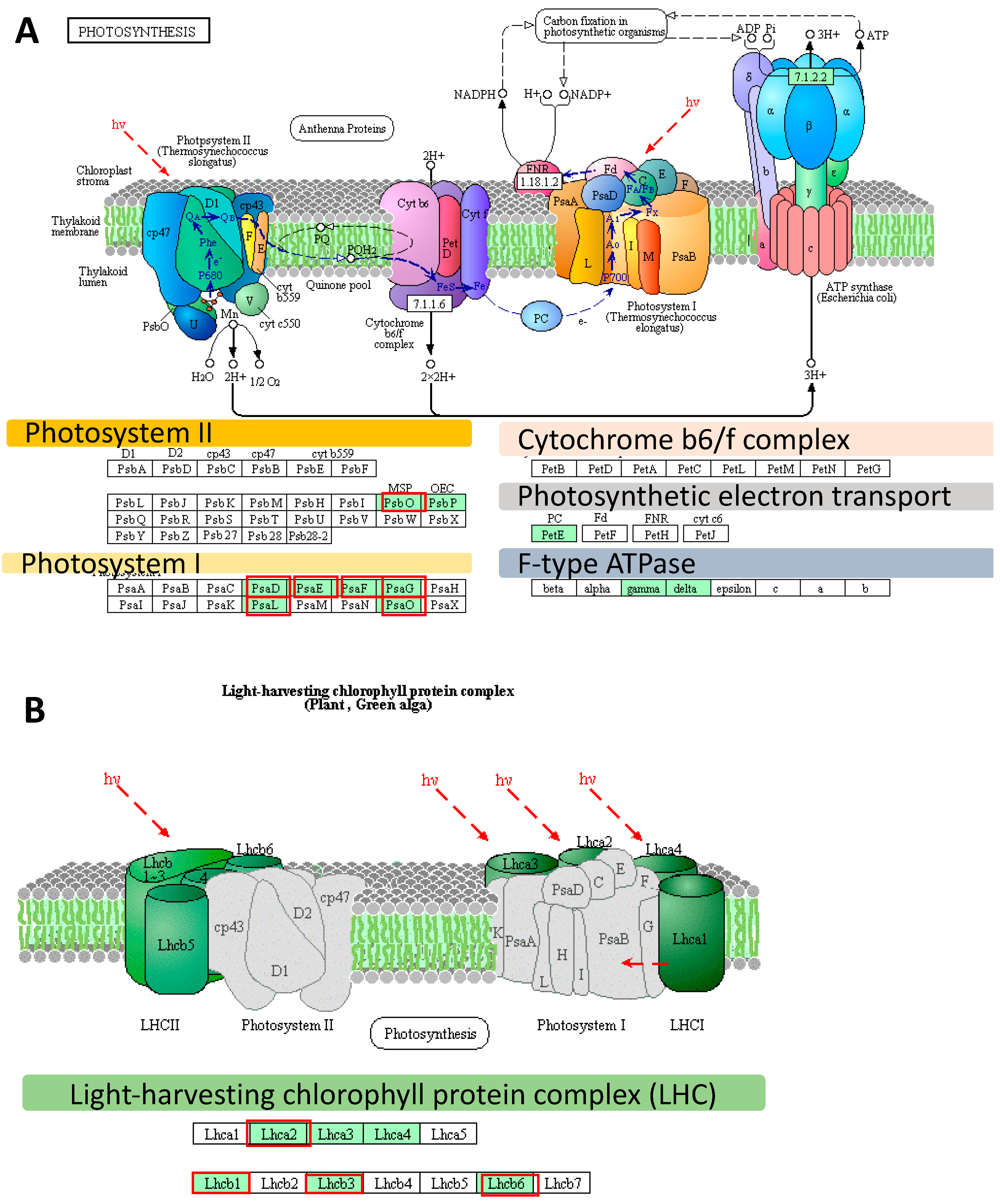
2.2. Deep Transcriptome Sequencing Uncovered a Core Set of Genes Involved in Photosynthesis in Barley
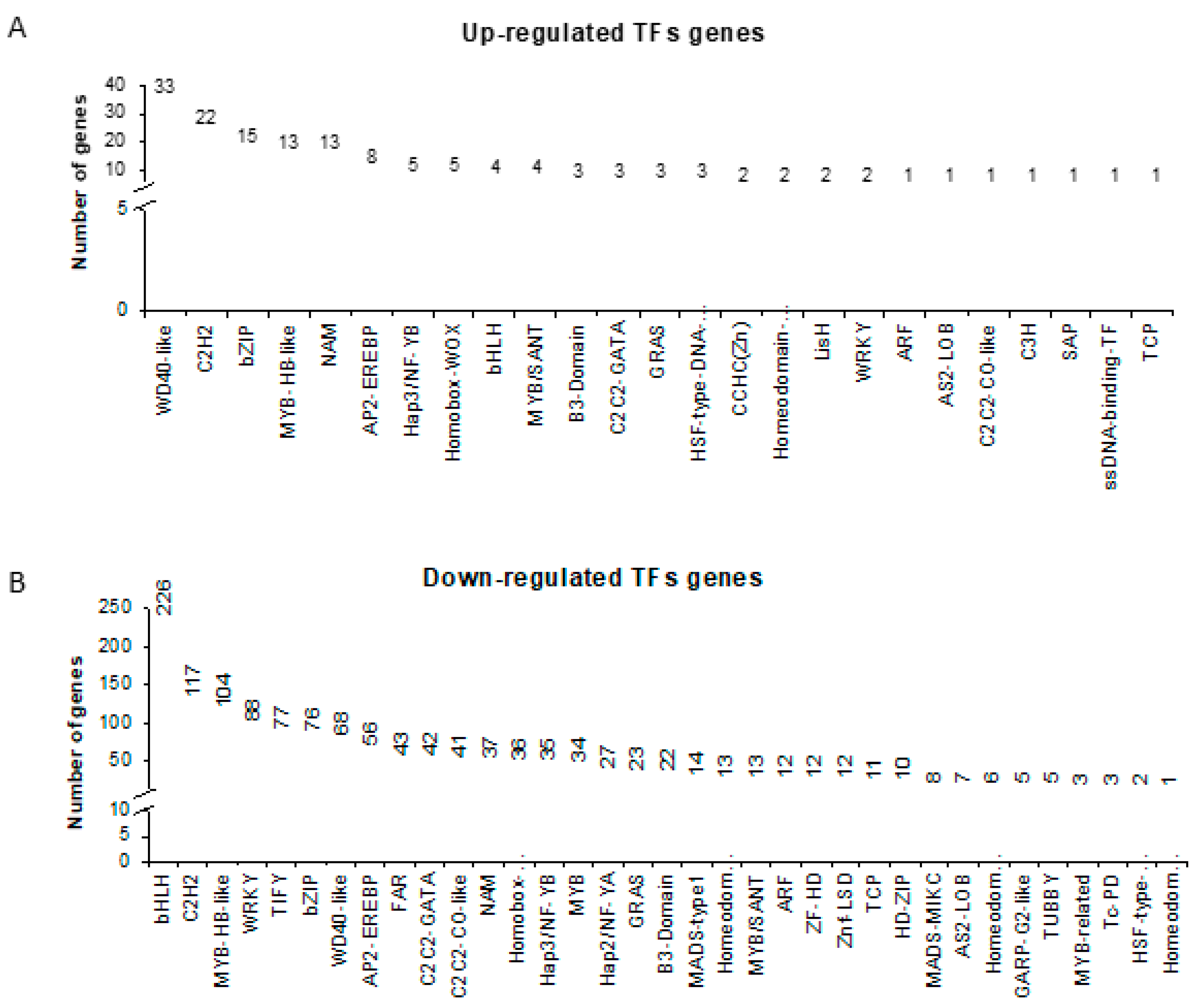
2.3. Promoter Analysis Coupled with Transcription Factors (TFs) Identified in the Transcriptomic Study Predicts New Candidates for Further Studies
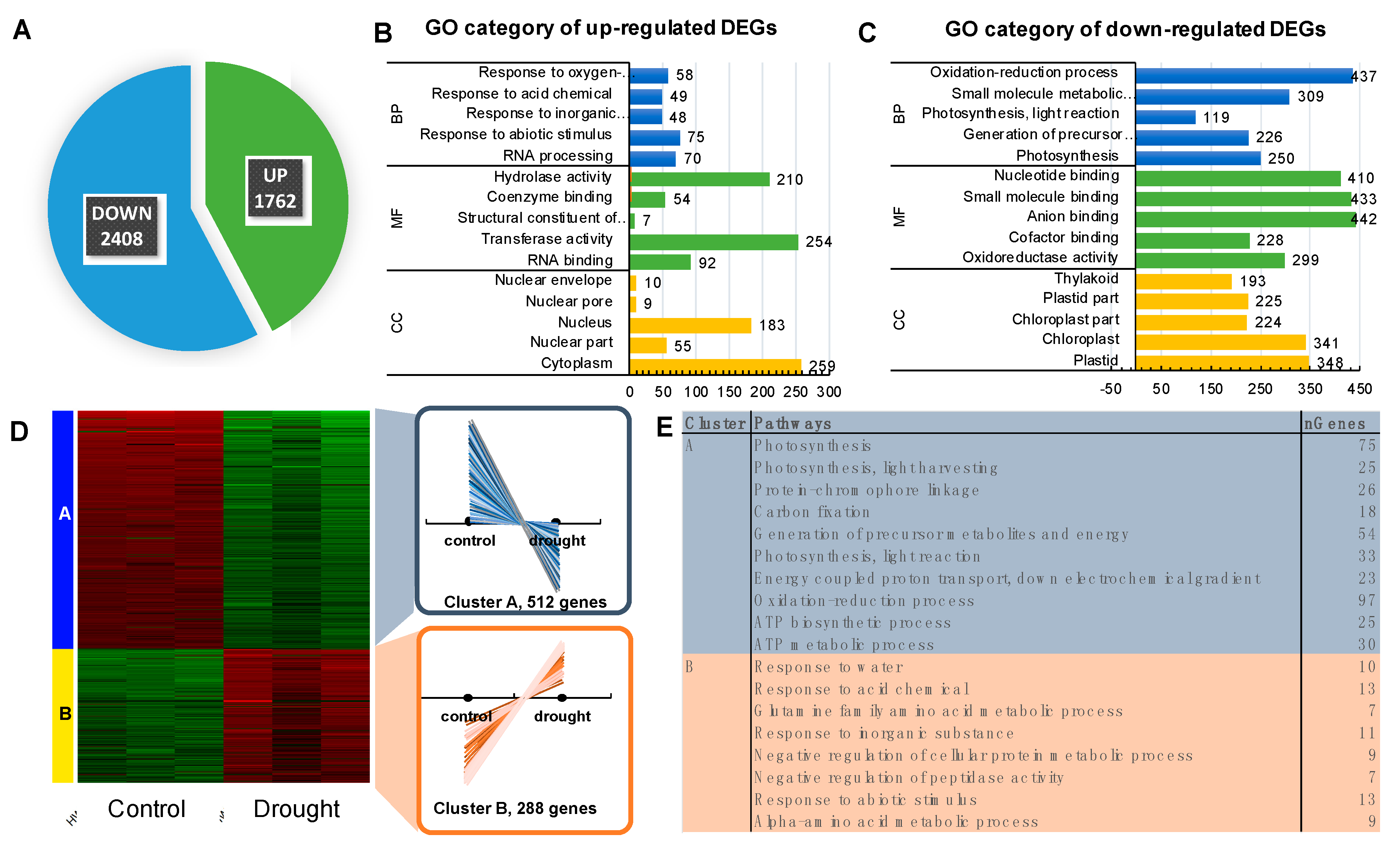
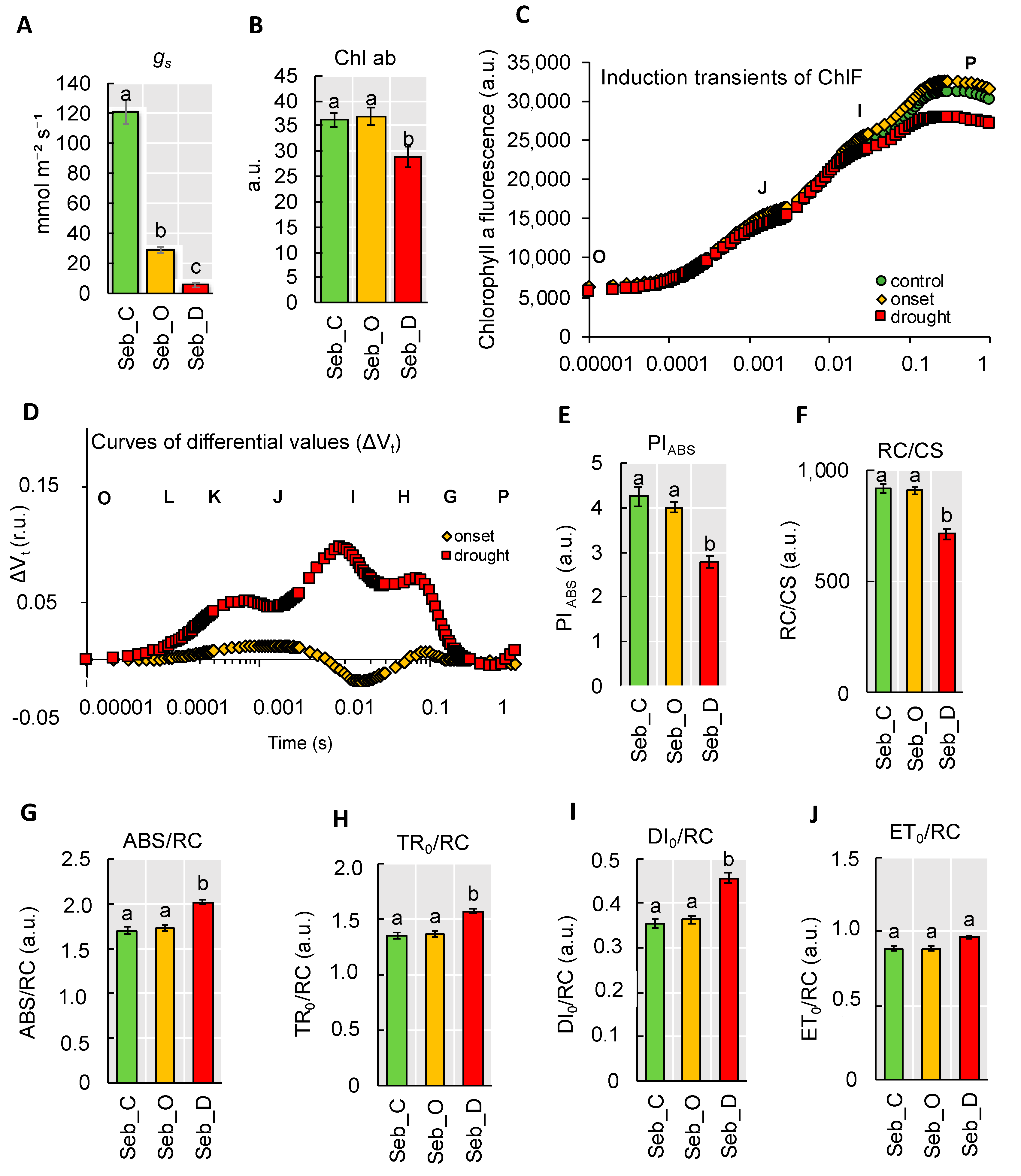
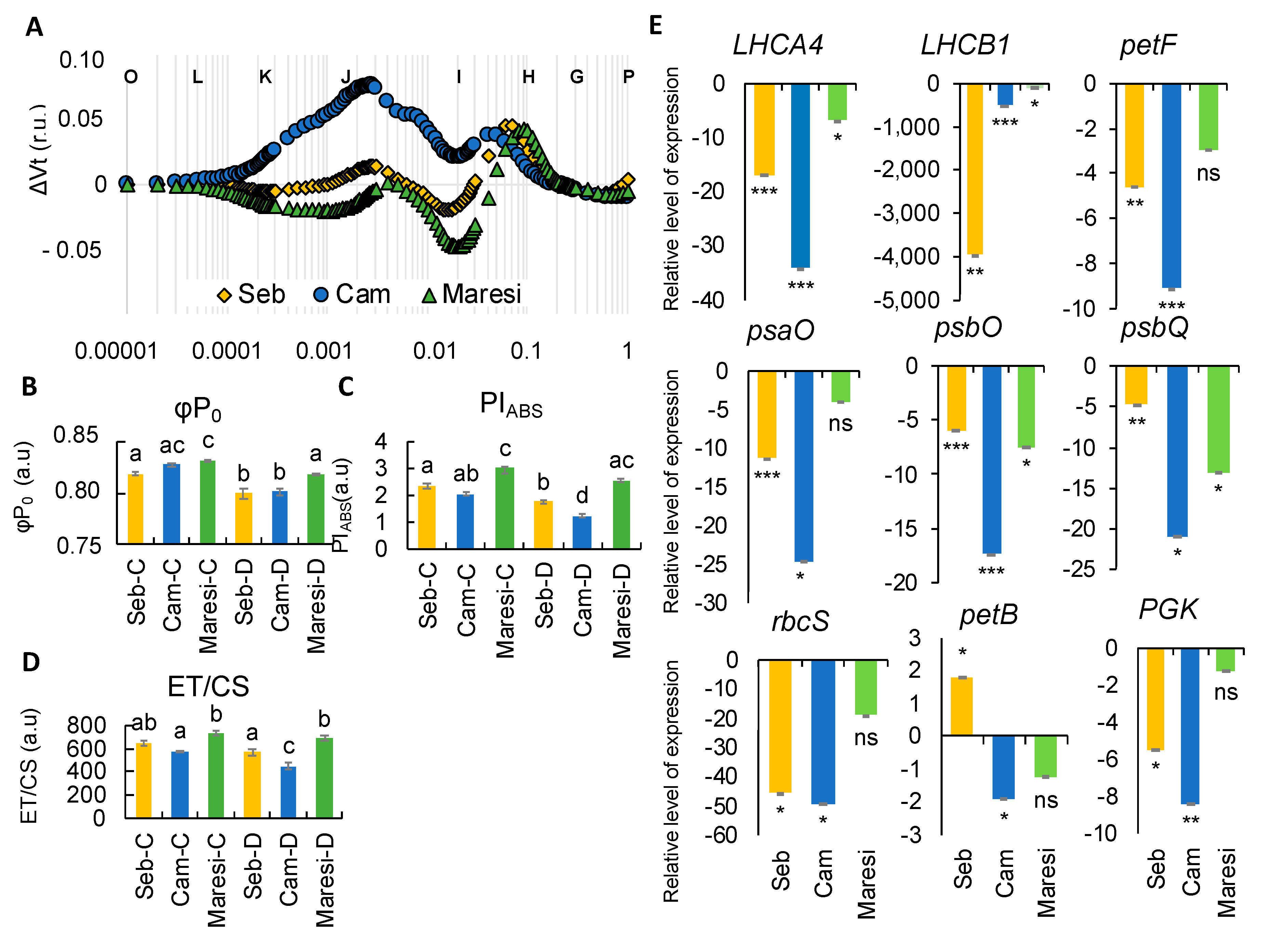
2.4. Physiological Analyses Showed a Pronounced Decline of Photosynthesis Performance only after Prolonged Drought Stress
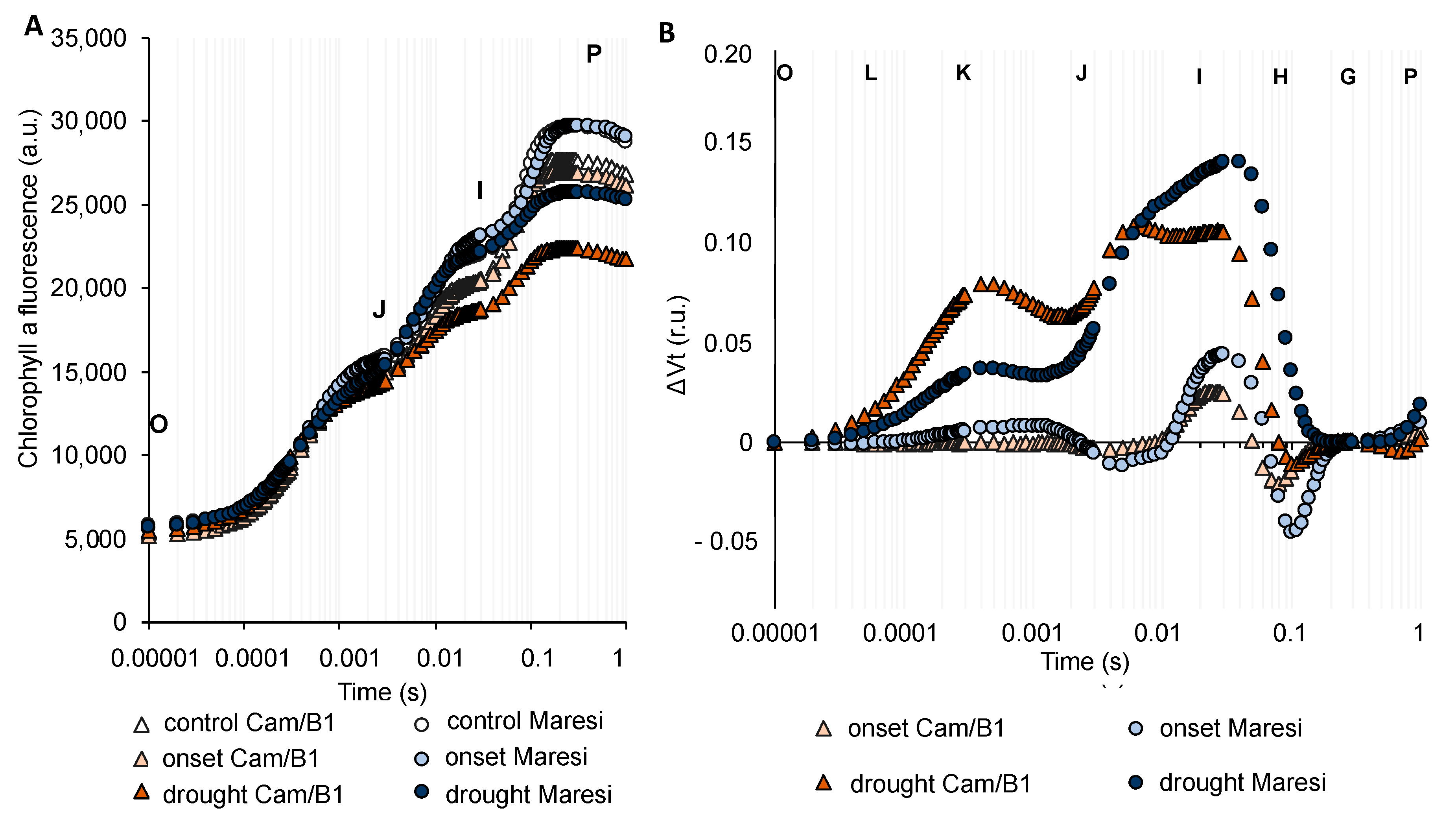
2.5. Drought Stress Influenced Barley Photosynthesis in a Universal, Genotype-Independent Manner
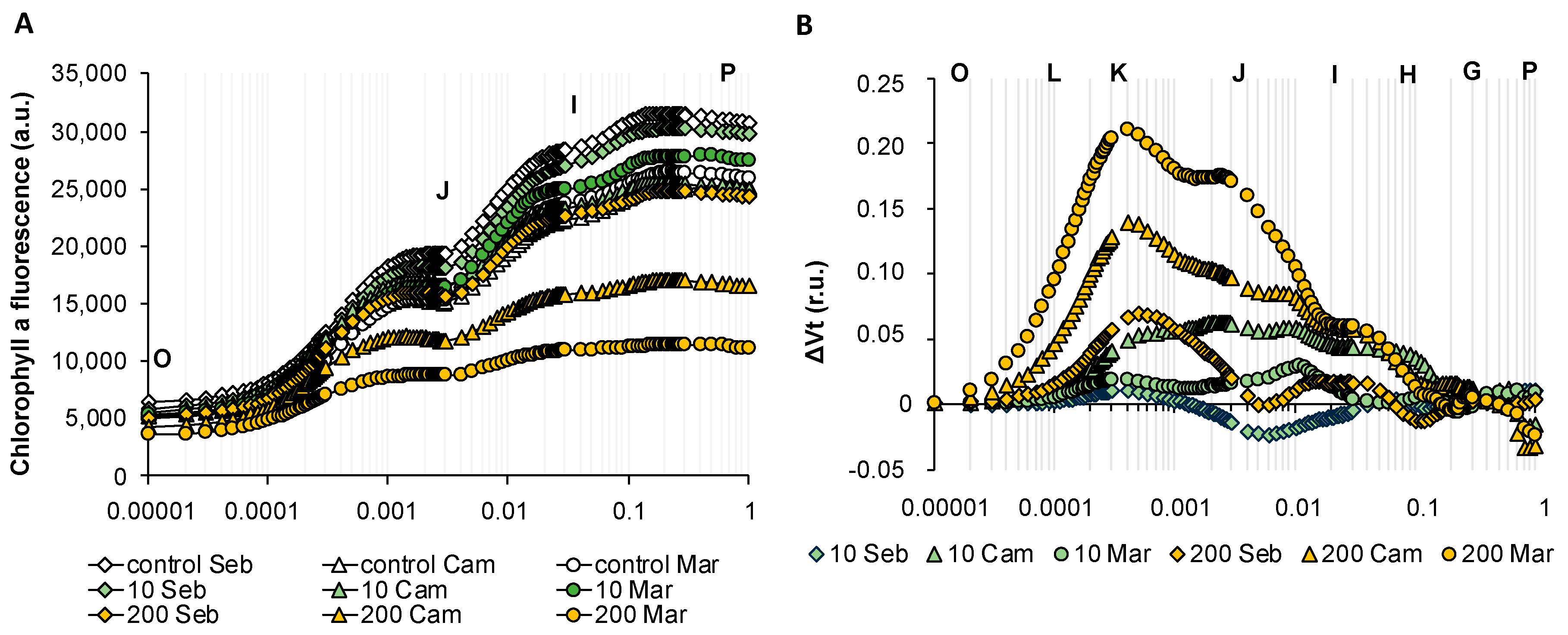
2.6. Rapid Dehydration Disturbed the Balance in Photosynthesis Reactions Probably due to the Disintegration of Components Protecting the Oxygen Evolving Complex
2.7. Exogenously Applied Abscisic Acid (ABA) Mostly Affected the Electron Transport in Barley
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Assays of Seedling Development in the Presence of Abiotic Stresses
4.2.1. Drought Stress Experiment
4.2.2. ABA Treatment
4.2.3. Rapid Dehydration Assay
4.3. Transcriptomic Analyses
4.3.1. Microarray Experiment and Bioinformatics Analysis
4.3.2. RNAseq Experiment and Bioinformatics Data Analysis
4.3.3. Quantitative Real-Time PCR Analysis
4.4. Chlorophyll a Fluorescence and Chlorophyll Content Measurements
4.5. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Daszkowska-Golec, A. Drought Stress Tolerance in Plants. In Drought Stress Tolerance in Plants; Hossain, M.A., Ed.; Springer International Publishing: Cham, Switzerland, 2016; pp. 123–143. ISBN 9783319288970. [Google Scholar]
- Kuromori, T.; Seo, M.; Shinozaki, K. ABA Transport and Plant Water Stress Responses. Trends Plant Sci. 2018, 23, 513–522. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Zhou, G. Responses of leaf stomatal density to water status and its relationship with photosynthesis in a grass. J. Exp. Bot. 2008, 59, 3317–3325. [Google Scholar] [CrossRef]
- Merilo, E.; Yarmolinsky, D.; Jalakas, P.; Parik, H.; Tulva, I.; Rasulov, B.; Kilk, K.; Kollist, H. Stomatal VPD Response: There Is More to the Story Than ABA. Plant Physiol. 2018, 176, 851–864. [Google Scholar] [CrossRef] [PubMed]
- Negin, B.; Yaaran, A.; Kelly, G.; Zait, Y.; Moshelion, M. Mesophyll ABA restrains early growth and flowering but does not directly suppress photosynthesis. Plant Physiol. 2019, 180, 910–925. [Google Scholar] [CrossRef] [PubMed]
- Pinheiro, C.; Chaves, M.M. Photosynthesis and drought: Can we make metabolic connections from available data? J. Exp. Bot. 2011, 62, 869–882. [Google Scholar] [CrossRef]
- Lawson, T.; Vialet-Chabrand, S. Speedy stomata, photosynthesis and plant water use efficiency. New Phytol. 2019, 221, 93–98. [Google Scholar] [CrossRef]
- Gururani, M.A.; Venkatesh, J.; Tran, L.S.P. Regulation of photosynthesis during abiotic stress-induced photoinhibition. Mol. Plant 2015, 8, 1304–1320. [Google Scholar] [CrossRef]
- Kaiser, E.; Morales, A.; Harbinson, J.; Kromdijk, J.; Heuvelink, E.; Marcelis, L.F.M. Dynamic photosynthesis in different environmental conditions. J. Exp. Bot. 2014, 66, 2415–2426. [Google Scholar] [CrossRef]
- Yamaoka, C.; Suzuki, Y.; Makino, A. Differential expression of genes of the Calvin-Benson cycle and its related genes during leaf development in rice. Plant Cell Physiol. 2016, 57, 115–124. [Google Scholar] [CrossRef]
- Rochaix, J.-D. Fine-Tuning Photosynthesis. Science 2013, 342, 50–51. [Google Scholar] [CrossRef]
- Foyer, C.H.; Bloom, A.J.; Queval, G.; Noctor, G. Photorespiratory Metabolism: Genes, Mutants, Energetics, and Redox Signaling. Ann. Rev. Plant Biol. 2009, 60, 455–484. [Google Scholar] [CrossRef] [PubMed]
- Shen, J. The Structure of Photosystem II and the Mechanism of Water Oxidation in Photosynthesis. Ann. Rev. Plant Biol. 2015, 66, 23–48. [Google Scholar] [CrossRef] [PubMed]
- Kalaji, H.M.; Jajoo, A.; Oukarroum, A.; Brestic, M.; Zivcak, M.; Samborska, I.A.; Cetner, M.D.; Łukasik, I.; Goltsev, V.; Ladle, R.J. Chlorophyll a fluorescence as a tool to monitor physiological status of plants under abiotic stress conditions. Acta Physiol. Plant. 2016, 38, 102. [Google Scholar] [CrossRef]
- Oukarroum, A.; Madidi, S.E.; Schansker, G.; Strasser, R.J. Probing the responses of barley cultivars (Hordeum vulgare L.) by chlorophyll a fluorescence OLKJIP under drought stress and re-watering. Environ. Exp. Bot. 2007, 60, 438–446. [Google Scholar] [CrossRef]
- Jiang, H.-X.; Chen, L.-S.; Zheng, J.-G.; Han, S.; Tang, N.; Smith, B.R. Aluminum-induced effects on Photosystem II photochemistry in citrus leaves assessed by the chlorophyll a fluorescence transient. Tree Physiol. 2008, 28, 1863–1871. [Google Scholar] [CrossRef]
- Stirbet, A.; Govindjee. On the relation between the Kautsky effect (chlorophyll a fluorescence induction) and Photosystem II: Basics and applications of the OJIP fluorescence transient. J. Photochem. Photobiol. B Biol. 2011, 104, 236–257. [Google Scholar] [CrossRef]
- Strasser, R.J.; Srivastava, A.; Tsimili-Michael, M. The fluorescence transient as a tool to characterize and screen photosynthetic samples. In Probing Photosynthesis: Mechanisms, Regulation and Adaptation; Yunus, M., Pathre, U., Mohanty, P., Eds.; Taylor and Francis: London, UK, 2000; pp. 445–483. [Google Scholar]
- Schansker, G.; Tóth, S.Z.; Strasser, R.J. Dark recovery of the Chl a fluorescence transient (OJIP) after light adaptation: The qT-component of non-photochemical quenching is related to an activated photosystem I acceptor side. Biochim. Biophys. Acta Bioenerg. 2006, 1757, 787–797. [Google Scholar] [CrossRef]
- Kalaji, H.M.; Loboda, T. Photosystem II of barley seedlings under cadmium and lead stress. Plant Soil Environ. 2005, 53, 511–516. [Google Scholar] [CrossRef]
- Rapacz, M.; Wójcik-Jagła, M.; Fiust, A.; Kalaji, H.M.; Kościelniak, J. Genome-Wide Associations of Chlorophyll Fluorescence OJIP Transient Parameters Connected With Soil Drought Response in Barley. Front. Plant. Sci. 2019, 10, 78. [Google Scholar] [CrossRef]
- Daszkowska-Golec, A.; Skubacz, A.; Sitko, K.; Słota, M.; Kurowska, M.; Szarejko, I. Mutation in barley ERA1 (Enhanced Response to ABA1) gene confers better photosynthesis efficiency in response to drought as revealed by transcriptomic and physiological analysis. Environ. Exp. Bot. 2018, 148, 12–26. [Google Scholar] [CrossRef]
- Daszkowska-Golec, A.; Skubacz, A.; Marzec, M.; Slota, M.; Kurowska, M.; Gajecka, M.; Gajewska, P.; Płociniczak, T.; Sitko, K.; Pacak, A.; et al. Mutation in HvCBP20 (Cap binding protein 20) adapts barley to drought stress at phenotypic and transcriptomic levels. Front. Plant. Sci. 2017, 8, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Gudys, K.; Guzy-wrobelska, J.; Janiak, A.; Dxiurka, M.A.; Ostrowska, A.; Hura, K.; Jurczyk, B.; Zmuda, K.; Grzybkowska, D.; Srobka, J.; et al. Prioritization of Candidate Genes in QTL Regions for Physiological and Biochemical Traits Underlying Drought Response in Barley (Hordeum vulgare L.). Front. Plant. Sci. 2018, 9, 1–26. [Google Scholar] [CrossRef] [PubMed]
- Dawson, I.K.; Russell, J.; Powell, W.; Steffenson, B.; Thomas, W.T.B.; Waugh, R. Tansley review Barley: A translational model for adaptation to climate change. New Phytol. 2015, 206, 913–931. [Google Scholar] [CrossRef] [PubMed]
- Mascher, M.; Gundlach, H.; Himmelbach, A.; Beier, S.; Twardziok, S.O.; Wicker, T.; Radchuk, V.; Dockter, C.; Hedley, P.E.; Russell, J.; et al. A chromosome conformation capture ordered sequence of the barley genome. Nature 2017, 544, 427–433. [Google Scholar] [CrossRef]
- Neumann, K.; Zhao, Y.; Chu, J.; Keilwagen, J.; Reif, J.C.; Kilian, B.; Graner, A. Genetic architecture and temporal patterns of biomass accumulation in spring barley revealed by image analysis. BMC Plant Biol. 2017, 17, 1–12. [Google Scholar] [CrossRef]
- Janiak, A.; Kwasniewski, M.; Sowa, M.; Gajek, K.; Żmuda, K.; Kościelniak, J.; Szarejko, I. No Time to Waste: Transcriptome Study Reveals that Drought Tolerance in Barley May Be Attributed to Stressed-Like Expression Patterns that Exist before the Occurrence of Stress. Front. Plant Sci. 2018, 8, 2212. [Google Scholar] [CrossRef]
- Filek, M.; Łabanowska, M.; Kościelniak, J.; Biesaga-Kościelniak, J.; Kurdziel, M.; Szarejko, I.; Hartikainen, H. Characterization of Barley Leaf Tolerance to Drought Stress by Chlorophyll Fluorescence and Electron Paramagnetic Resonance Studies. J. Agron. Crop Sci. 2015, 201, 228–240. [Google Scholar] [CrossRef]
- Daszkowska-Golec, A.; Skubacz, A.; Kurowska, M.; Słota, M.; Swiergolik, D.; Szarejko, I. Methods for the Simple and Reliable Assessment of Barley Sensitivity to Abiotic Stresses During Early Development. In Barley: Methods and Protocols; Harwood, W.A., Ed.; Springer: Berlin, Germany, 2019; pp. 127–151. [Google Scholar]
- Ort, D.R.; Merchant, S.S.; Alric, J.; Barkan, A.; Blankenship, R.E.; Bock, R.; Croce, R.; Hanson, M.R.; Hibberd, J.M.; Long, S.P.; et al. Redesigning photosynthesis to sustainably meet global food and bioenergy demand. Proc. Natl. Acad. Sci. USA 2015, 112, 8529–8536. [Google Scholar] [CrossRef]
- Koochak, H.; Puthiyaveetil, S.; Mullendore, D.L.; Li, M.; Kirchhoff, H. The structural and functional domains of plant thylakoid membranes. Plant J. 2019, 97, 412–429. [Google Scholar] [CrossRef]
- Albertsson, P.Å. The structure and function of the chloroplast photosynthetic membrane—A model for the domain organization. Photosynth. Res. 1995, 46, 141–149. [Google Scholar] [CrossRef]
- Jansson, S. A guide to the Lhc genes and their relatives in Arabidopsis. Trends Plant Sci. 1999, 4, 236–240. [Google Scholar] [CrossRef]
- Andersson, J.; Wentworth, M.; Walters, R.G.; Howard, C.A.; Ruban, A.V.; Horton, P.; Jansson, S. Absence of the Lhcb1 and Lhcb2 proteins of the light-harvesting complex of photosystem II-Effects on photosynthesis, grana stacking and fitness. Plant J. 2003, 35, 350–361. [Google Scholar] [PubMed]
- Maxwell, K.; Johnson, G.N. Chlorophyll fluorescence-A practical guide. J. Exp. Bot. 2000, 51, 659–668. [Google Scholar] [CrossRef] [PubMed]
- Jedmowski, C.; Ashoub, A.; Brüggemann, W. Reactions of Egyptian landraces of Hordeum vulgare and Sorghum bicolor to drought stress, evaluated by the OJIP fluorescence transient analysis. Acta Physiol. Plant. 2013, 35, 345–354. [Google Scholar] [CrossRef]
- Kalaji, H.M.; Rastogi, A.; Živčák, M.; Brestic, M.; Daszkowska-Golec, A.; Sitko, K.; Alsharafa, K.Y.; Lotfi, R.; Stypiński, P.; Samborska, I.A.; et al. Prompt chlorophyll fluorescence as a tool for crop phenotyping: An example of barley landraces exposed to various abiotic stress factors. Photosynthetica 2018, 56, 953–961. [Google Scholar] [CrossRef]
- Chmielewska, K.; Rodziewicz, P.; Swarcewicz, B.; Sawikowska, A.; Krajewski, P.; Marczak, Ł.; Ciesiołka, D.; Kuczyńska, A.; Mikołajczak, K.; Ogrodowicz, P.; et al. Analysis of Drought-Induced Proteomic and Metabolomic Changes in Barley (Hordeum vulgare L.) Leaves and Roots Unravels Some Aspects of Biochemical Mechanisms Involved in Drought Tolerance. Front. Plant Sci. 2016, 7, 1–14. [Google Scholar] [CrossRef]
- Filek, M.; Łabanowska, M.; Kurdziel, M.; Wesełucha-Birczyńska, A.; Bednarska-Kozakiewicz, E. Structural and biochemical response of chloroplasts in tolerant and sensitive barley genotypes to drought stress. J. Plant Physiol. 2016, 207, 61–72. [Google Scholar] [CrossRef]
- Piasecka, A.; Sawikowska, A.; Kuczyńska, A.; Ogrodowicz, P.; Mikołajczak, K.; Krystkowiak, K.; Gudyś, K.; Guzy-Wróbelska, J.; Krajewski, P.; Kachlicki, P. Drought-related secondary metabolites of barley (Hordeum vulgare L.) leaves and their metabolomic quantitative trait loci. Plant J. 2017, 89, 898–913. [Google Scholar] [CrossRef]
- Ogrodowicz, P.; Adamski, T.; Mikołajczak, K.; Kuczyńska, A.; Surma, M.; Krajewski, P.; Sawikowska, A.; Górny, A.G.; Gudyś, K.; Szarejko, I.; et al. QTLs for earliness and yield-forming traits in the Lubuski × CamB barley RIL population under various water regimes. J. Appl. Genet. 2017, 58, 49–65. [Google Scholar] [CrossRef]
- Wientjes, E.; Oostergetel, G.T.; Jansson, S.; Boekema, E.J.; Croce, R. The role of Lhca complexes in the supramolecular organization of higher plant photosystem I. J. Biol. Chem. 2009, 284, 7803–7810. [Google Scholar] [CrossRef]
- Paunov, M.; Koleva, L.; Vassilev, A.; Vangronsveld, J.; Goltsev, V. Effects of different metals on photosynthesis: Cadmium and zinc affect chlorophyll fluorescence in durum wheat. Int. J. Mol. Sci. 2018, 19, 787. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, K.N.; Iverson, T.M.; Maghlaoui, K.; Barber, J.; Iwata, S. Architecture of the Photosynthetic Oxygen-Evolving Center. Science 2004, 303, 1831–1838. [Google Scholar] [CrossRef] [PubMed]
- Bricker, T.M.; Frankel, L.K. The structure and function of CP47 and CP43 in Photosystem II. Photosynth Res. 2002, 72, 131–146. [Google Scholar] [CrossRef] [PubMed]
- Morse, M.; Rafudeen, M.S.; Farrant, J.M. An Overview of the Current Understanding of Desiccation Tolerance in the Vegetative Tissues of Higher Plants, 1st ed.; Elsevier Ltd.: Amsterdam, The Netherlands, 2011; Volume 57, ISBN 9780123876928. [Google Scholar]
- Rossini, L.; Cribb, L.; Martin, D.J.; Langdale, J.A. The Maize Golden2 Gene Defines a Novel Class of Transcriptional Regulators in Plants. Plant Cell 2007, 13, 1231. [Google Scholar] [CrossRef] [PubMed]
- Fitter, D.W.; Martin, D.J.; Copley, M.J.; Scotland, R.W.; Langdale, J.A. GLK gene pairs regulate chloroplast development in diverse plant species. Plant J. 2002, 31, 713–727. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, H.; Muramatsu, M.; Hakata, M.; Ueno, O.; Nagamura, Y.; Hirochika, H.; Takano, M.; Ichikawa, H. Ectopic overexpression of the transcription factor osglk1 induces chloroplast development in non-green rice cells. Plant Cell Physiol. 2009, 50, 1933–1949. [Google Scholar] [CrossRef]
- Waters, M.T.; Moylan, E.C.; Langdale, J.A. GLK transcription factors regulate chloroplast development in a cell-autonomous manner. Plant J. 2008, 56, 432–444. [Google Scholar] [CrossRef]
- Ahmad, R.; Liu, Y.; Wang, T.-J.; Meng, Q.; Yin, H.; Wang, X.; Wu, Y.; Nan, N.; Liu, B.; Xu, Z.-Y. GOLDEN 2-LIKE Transcription Factors Regulate WRKY40 Expression in Response to Abscisic Acid. Plant Physiol. 2019, 179, 1844–1860. [Google Scholar] [CrossRef]
- Van Aken, O.; Zhang, B.; Law, S.; Narsai, R.; Whelan, J. AtWRKY40 and AtWRKY63 Modulate the Expression of Stress-Responsive Nuclear Genes Encoding Mitochondrial and Chloroplast Proteins. Plant Physiol. 2013, 162, 254–271. [Google Scholar] [CrossRef]
- Li, S.; Fu, Q.; Chen, L.; Huang, W.; Yu, D. Arabidopsis thaliana WRKY25, WRKY26, and WRKY33 coordinate induction of plant thermotolerance. Planta 2011, 233, 1237–1252. [Google Scholar] [CrossRef]
- Fey, V.; Wagner, R.; Bräutigam, K.; Pfannschmidt, T. Photosynthetic redox control of nuclear gene expression. J. Exp. Bot. 2005, 56, 1491–1498. [Google Scholar] [CrossRef] [PubMed]
- Gururani, M.A.; Mohanta, T.K.; Bae, H. Current understanding of the interplay between phytohormones and photosynthesis under environmental stress. Int. J. Mol. Sci. 2015, 16, 19055–19085. [Google Scholar] [CrossRef] [PubMed]
- Maslenkova, L.T.; Zanev, Y.; Popova, L.P. Effect of abscisic acid on the photosynthetic oxygen evolution in barley chloroplasts. Photosynth. Res. 1989, 21, 45–50. [Google Scholar] [PubMed]
- Du, H.; Wang, N.; Cui, F.; Li, X.; Xiao, J.; Xiong, L. Characterization of the Carotene Hydroxylase Gene DSM2 Conferring Drought and Oxidative Stress Resistance by Increasing Xanthophylls and Abscisic Acid Synthesis in Rice. Plant Physiol. 2010, 154, 1304–1318. [Google Scholar] [CrossRef]
- Humplík, J.F.; Bergougnoux, V.; Van Volkenburgh, E. To Stimulate or Inhibit? That Is the Question for the Function of Abscisic Acid. Trends Plant Sci. 2017, 22, 830–841. [Google Scholar] [CrossRef]
- Szurman-Zubrzycka, M.E.; Zbieszczyk, J.; Marzec, M.; Jelonek, J.; Chmielewska, B.; Kurowska, M.M.; Krok, M.; Daszkowska-Golec, A.; Guzy-Wrobelska, J.; Gruszka, D.; et al. HorTILLUS—A Rich and Renewable Source of Induced Mutations for Forward/Reverse Genetics and Pre-breeding Programs in Barley (Hordeum vulgare L.). Front. Plant Sci. 2018, 9, 1–16. [Google Scholar] [CrossRef]
- Strasser, R.J.; Tsimilli-Michael, A.S. Analysis of the Fluorescence Transient—Summary II. The Theoretical Background. In Chlorophyll A Fluorescence: A Signature of Photosynthesis; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2004; pp. 1–47. [Google Scholar]
- Kwasniewski, M.; Daszkowska-Golec, A.; Janiak, A.; Chwialkowska, K.; Nowakowska, U.; Sablok, G.; Szarejko, I. Transcriptome analysis reveals the role of the root hairs as environmental sensors to maintain plant functions under water-deficiency conditions. J. Exp. Bot. 2016, 67, 1079–1094. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2009, 26, 139–140. [Google Scholar] [CrossRef]
- Bray, N.L.; Pimentel, H.; Melsted, P.; Pachter, L. Near-optimal RNA-Seq quantification. Nat. Biotechnol. 2016, 34, 525–527. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Tian, T.; Liu, Y.; Yan, H.; You, Q.; Yi, X.; Du, Z.; Xu, W.; Su, Z. AgriGO v2.0: A GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res. 2017, 45, W122–W129. [Google Scholar] [CrossRef] [PubMed]
- Dai, X.; Sinharoy, S.; Udvardi, M.; Zhao, P.X. PlantTFcat: An online plant transcription factor and transcriptional regulator categorization and analysis tool. BMC Bioinform. 2013, 14, 321. [Google Scholar] [CrossRef] [PubMed]
- Kozera, B.; Rapacz, M. Reference genes in real-time PCR. J. Appl. Genet. 2013, 54, 391–406. [Google Scholar] [CrossRef] [PubMed]
- Ramakers, C.; Ruijter, J.M.; Lekanne Deprez, R.H.; Moorman, A.F.M. Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci. Lett. 2003, 339, 62–66. [Google Scholar] [CrossRef]
- Chomczynski, P.; Sacchi, N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 1987, 162, 156–159. [Google Scholar] [CrossRef]
- Kalaji, H.M.; Oukarroum, A.; Alexandrov, V.; Kouzmanova, M.; Brestic, M.; Zivcak, M.; Samborska, I.A.; Cetner, M.D.; Allakhverdiev, S.I.; Goltsev, V. Identification of nutrient deficiency in maize and tomato plants by invivo chlorophyll a fluorescence measurements. Plant Physiol. Biochem. 2014, 81, 16–25. [Google Scholar] [CrossRef]
| Transcription Factor Gene ID | FC of TF Gene | FDR | TF Family | Description of TF Gene | Target Gene ID | Target Description | FC of Target Gene | FDR |
|---|---|---|---|---|---|---|---|---|
| HORVU4Hr1G078410 | −20.36 | 2.25 × 10−15 | HD-ZIP | Homeobox-leucine zipper protein family | HORVU2Hr1G010630 | ribulose-bisphosphate carboxylase small chain [EC:4.1.1.39] | −432.2 | 1.56 × 10−4 |
| HORVU1Hr1G073300 | −12.71 | 7.62 × 10−5 | MYB_related | myb-like transcription factor family protein | HORVU1Hr1G035720 | ribulose-bisphosphate carboxylase small chain [EC:4.1.1.39] | −37.5 | 1.26 × 10−84 |
| HORVU6Hr1G066000 | −8.18 | 9.07 × 10−5 | MYB_related | myb-like transcription factor family protein | HORVU1Hr1G067300 | phosphoglycerate kinase [EC:2.7.2.3] | −52.7 | 2.27 × 10−32 |
| HORVU6Hr1G019650 | −7.20 | 8.16 × 10−6 | G2-like | myb-like transcription factor family protein | HORVU5Hr1G109260 | light-harvesting complex II chlorophyll a/b binding protein 1 | −1357 | 5.10 × 10−37 |
| HORVU5Hr1G014170 | −5.41 | 5.13 × 10−4 | bZIP | Basic-leucine zipper (bZIP) transcription factor family protein | HORVU2Hr1G060480 | photosystem I subunit X | −54.7 | 2.56 × 10−27 |
| HORVU3Hr1G088200 | −3.78 | 3.03 × 10−3 | WRKY | WRKY DNA-binding protein 33 | HORVU5Hr1G087250 | light-harvesting complex II chlorophyll a/b binding protein 1 | −13.1 | 1.86 × 10−6 |
| HORVU3Hr1G032440 | −3.26 | 8.28 × 10−4 | G2-like | Two-component response regulator-like APRR2 | HORVU2Hr1G075200 | F-type H+-transporting ATPase subunit b | −52.7 | 2.84 × 10−16 |
| HORVU2Hr1G060680 | −2.62 | 6.33 × 10−4 | bHLH | Transcription factor PIF5 | HORVU6Hr1G033160 | light-harvesting complex I chlorophyll a/b binding protein 3 | −30.2 | 5.16 × 10−11 |
| HORVU6Hr1G074970 | −2.37 | 2.93 × 10−3 | ERF | Ethylene-responsive transcription factor 1 | HORVU3Hr1G013350 | triosephosphate isomerase (TIM) [EC:5.3.1.1] | −6.7 | 8.10 × 10−16 |
| HORVU4Hr1G052330 | 2.46 | 3.42 × 10−3 | bZIP | transcription factor-related | HORVU2Hr1G063740 | glutamate--glyoxylate aminotransferase [EC:2.6.1.4 2.6.1.2 2.6.1.44] | −8.0 | 2.03 × 10−11 |
| HORVU1Hr1G074960 | 2.80 | 4.37 × 10−8 | bZIP | G-box binding factor 4 | HORVU1Hr1G021830 | cytochrome b6-f complex subunit 4 | −4.3 | 1.49 × 10−3 |
| HORVU5Hr1G036330 | 3.05 | 7.45 × 10−4 | GATA | GATA transcription factor 9 | HORVU6Hr1G051650 | photosystem II oxygen-evolving enhancer protein 3 | −202.0 | 7.00 × 10−4 |
| HORVU1Hr1G090030 | 3.06 | 8.67 × 10−8 | bZIP | G-box binding factor 2 | HORVU2Hr1G062090 | cytochrome b6 | −6.7 | 6.65 × 10−9 |
| HORVU4Hr1G020540 | 3.12 | 2.20 × 10−4 | bZIP | Transcription factor VIP1 | HORVU2Hr1G043240 | photosystem II P680 reaction center D1 protein [EC:1.10.3.9] | −5.9 | 5.99 × 10−13 |
| HORVU2Hr1G119610 | 3.53 | 4.62 × 10−4 | MYB_related | myb-like transcription factor family protein | HORVU2Hr1G043240 | photosystem II P680 reaction center D1 protein [EC:1.10.3.9] | −5.9 | 5.99 × 10−13 |
| HORVU4Hr1G051290 | 10.72 | 1.46 × 10−9 | NAC | NAC domain protein. | HORVU6Hr1G037020 | photosystem I P700 chlorophyll a apoprotein A1 | −5.9 | 4.15 × 10−7 |
| HORVU2Hr1G021080 | 13.16 | 5.84 × 10−7 | bZIP | Basic-leucine zipper (bZIP) transcription factor family protein | HORVU2Hr1G062090 | cytochrome b6 | −6.7 | 6.65 × 10−9 |
| Quantum Yields and Probabilities | |
| ϕP0 = 1 − F0/FM | Maximum quantum yield of primary photochemistry in PSII (at t = 0) |
| ϕE0 = (1 − F0/FM)(1 − VJ) | Quantum yield of electron transport (at t = 0) beyond QA− |
| ϕR0 = (1 − F0/FM)(1 − VI) | Quantum yield for reduction of the end of electron acceptors at PSI acceptor side (RE) |
| ϕD0 = F0/FM | Quantum yield (at t = 0) of energy dissipation |
| δR0 = (1 − VI)/(1 − VJ) | The efficiency with which an electron can move from the reduced intersystem electron acceptors to the PSI end electron acceptors |
| ψE0 = 1 − VJ | Probability (at t = 0) that a trapped exciton will move an electron into electron transport chain beyond QA− |
| Specific Energy fluxes expressed per active PSII reaction center (RC) | |
| ABS/RC = (1 − γRC)/ γRC = M0 × (1/VJ)/[1 − (F0/FM)] | Apparent antenna size of active PSII RC |
| TR0/RC = Mo(1/Vj) | Trapping flux (leading to QA reduction) per RC |
| DI0/RC = (ABS/RC − TR0/RC) | Dissipated energy flux per RC |
| ET0/RC = M0(1/Vj) ψ0 | Electron transport flux per RC (further than QA−) |
| RC/CS0 = FO φP0 Vj/M0 | Density of RCs (QA− reducing PSII reaction centers) |
| Performance index per absorption | |
| PIABS = 1 − (F0/FM)/Mo/VJ × FM − F0/F0 × 1 − VJ/VJ | Performance index (potential) for energy conservation from exciton to the reduction of intersystem electron acceptors |
| Phenomenological energy fluxes per excited cross-section (CS) | |
| ABS/CS | Absorption energy flux per CS |
| TR0/CS | Trapped energy flux per CS |
| ET0/CS | Electron transport flux per CS |
| DI0/CS | Dissipation energy flux per CS |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Daszkowska-Golec, A.; Collin, A.; Sitko, K.; Janiak, A.; Kalaji, H.M.; Szarejko, I. Genetic and Physiological Dissection of Photosynthesis in Barley Exposed to Drought Stress. Int. J. Mol. Sci. 2019, 20, 6341. https://doi.org/10.3390/ijms20246341
Daszkowska-Golec A, Collin A, Sitko K, Janiak A, Kalaji HM, Szarejko I. Genetic and Physiological Dissection of Photosynthesis in Barley Exposed to Drought Stress. International Journal of Molecular Sciences. 2019; 20(24):6341. https://doi.org/10.3390/ijms20246341
Chicago/Turabian StyleDaszkowska-Golec, Agata, Anna Collin, Krzysztof Sitko, Agnieszka Janiak, Hazem M. Kalaji, and Iwona Szarejko. 2019. "Genetic and Physiological Dissection of Photosynthesis in Barley Exposed to Drought Stress" International Journal of Molecular Sciences 20, no. 24: 6341. https://doi.org/10.3390/ijms20246341
APA StyleDaszkowska-Golec, A., Collin, A., Sitko, K., Janiak, A., Kalaji, H. M., & Szarejko, I. (2019). Genetic and Physiological Dissection of Photosynthesis in Barley Exposed to Drought Stress. International Journal of Molecular Sciences, 20(24), 6341. https://doi.org/10.3390/ijms20246341







