OsMSR3, a Small Heat Shock Protein, Confers Enhanced Tolerance to Copper Stress in Arabidopsis thaliana
Abstract
1. Introduction
2. Results
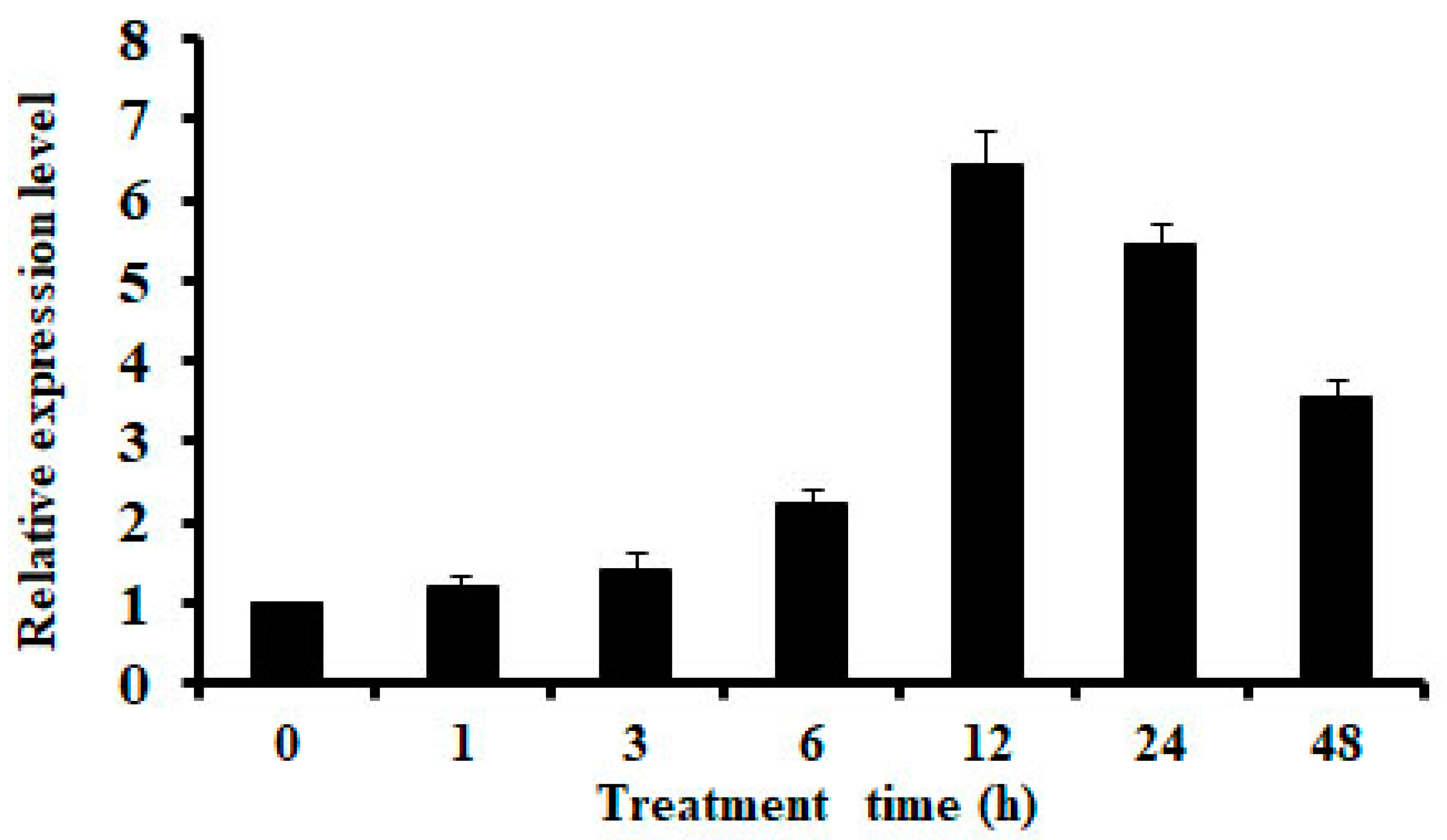
2.1. Expression of OsMSR3 is Induced by Cu Stress
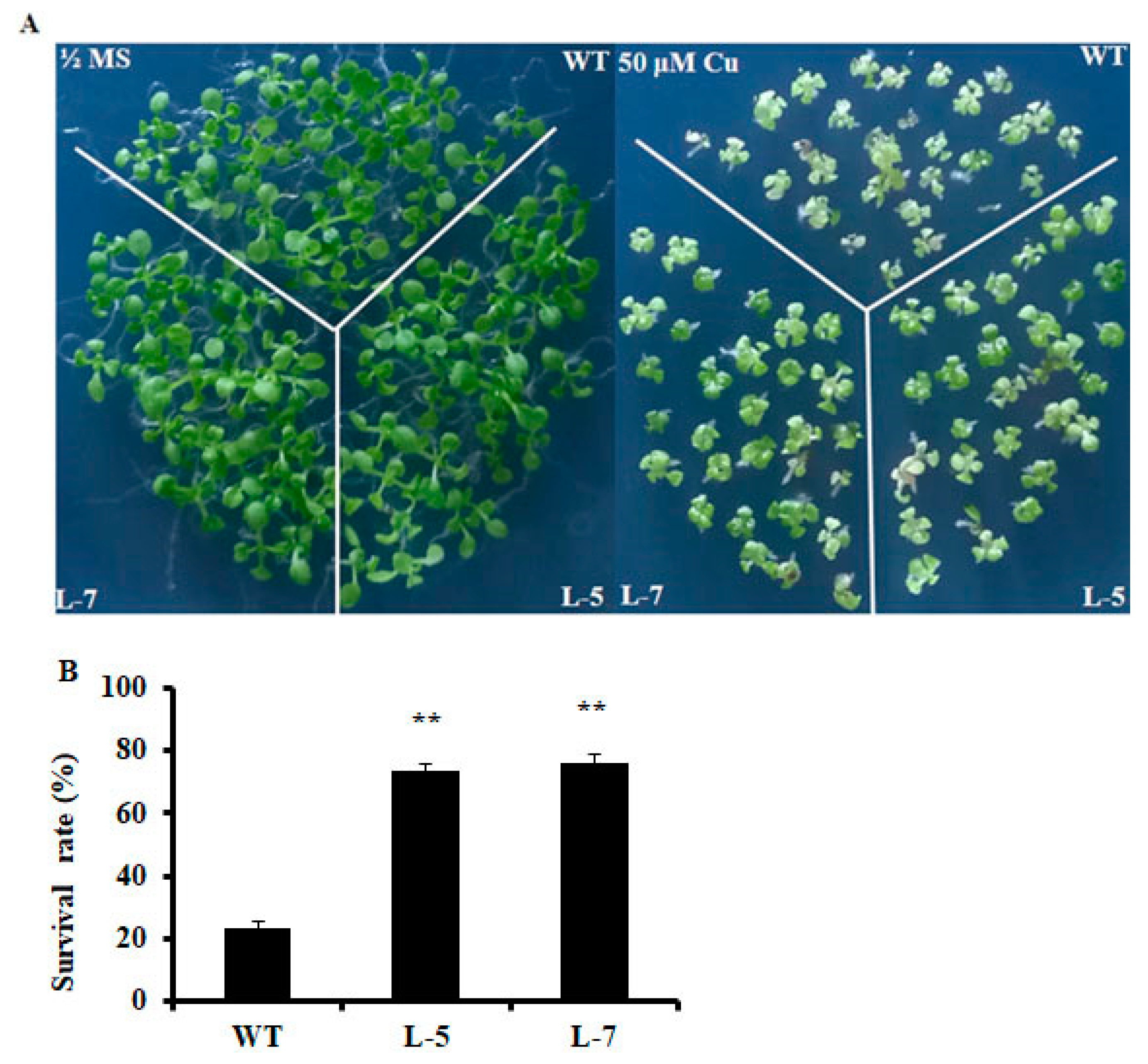
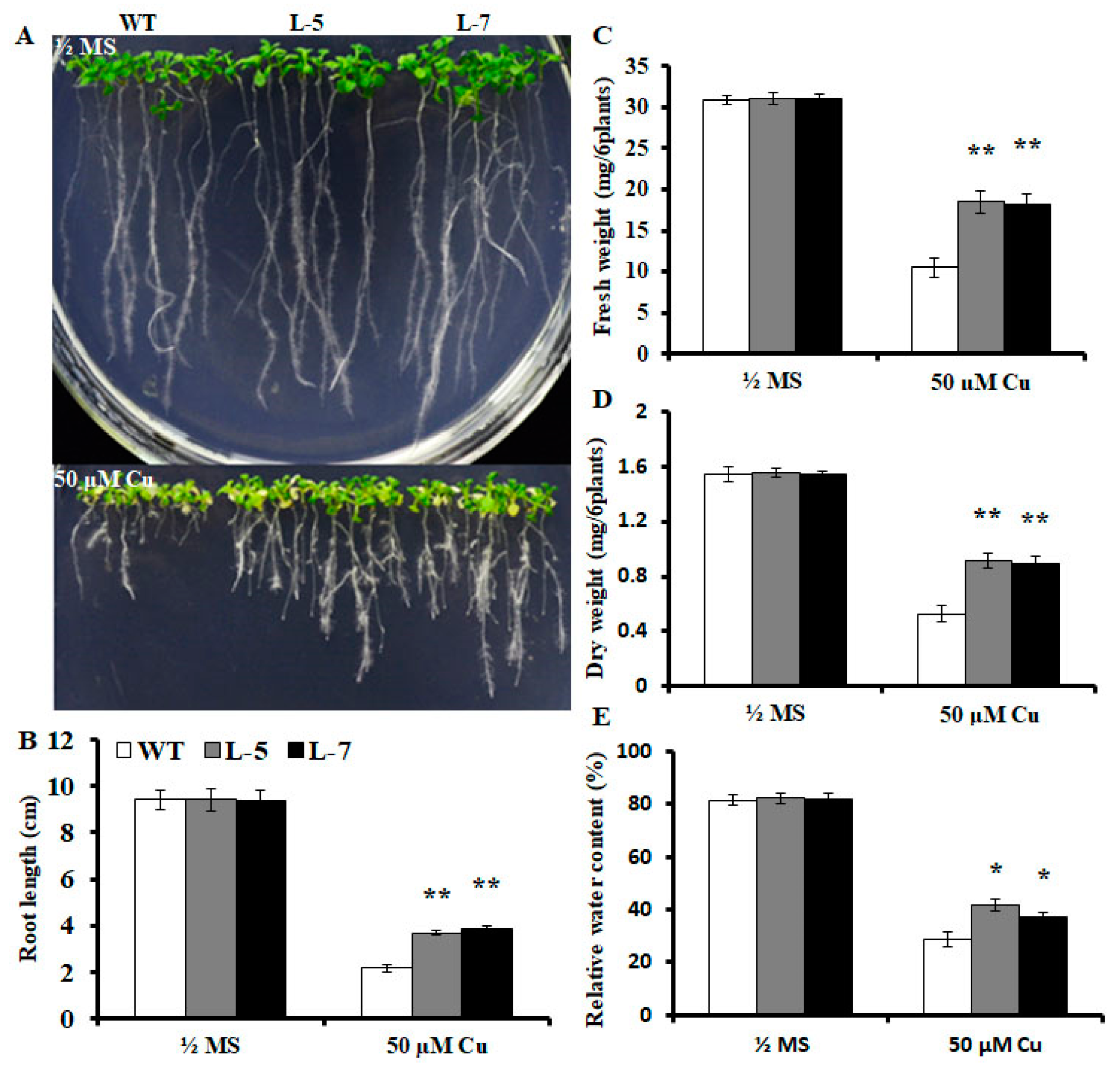
2.2. Expression of OsMSR3 Enhances Cu Tolerance of Transgenic A. thaliana
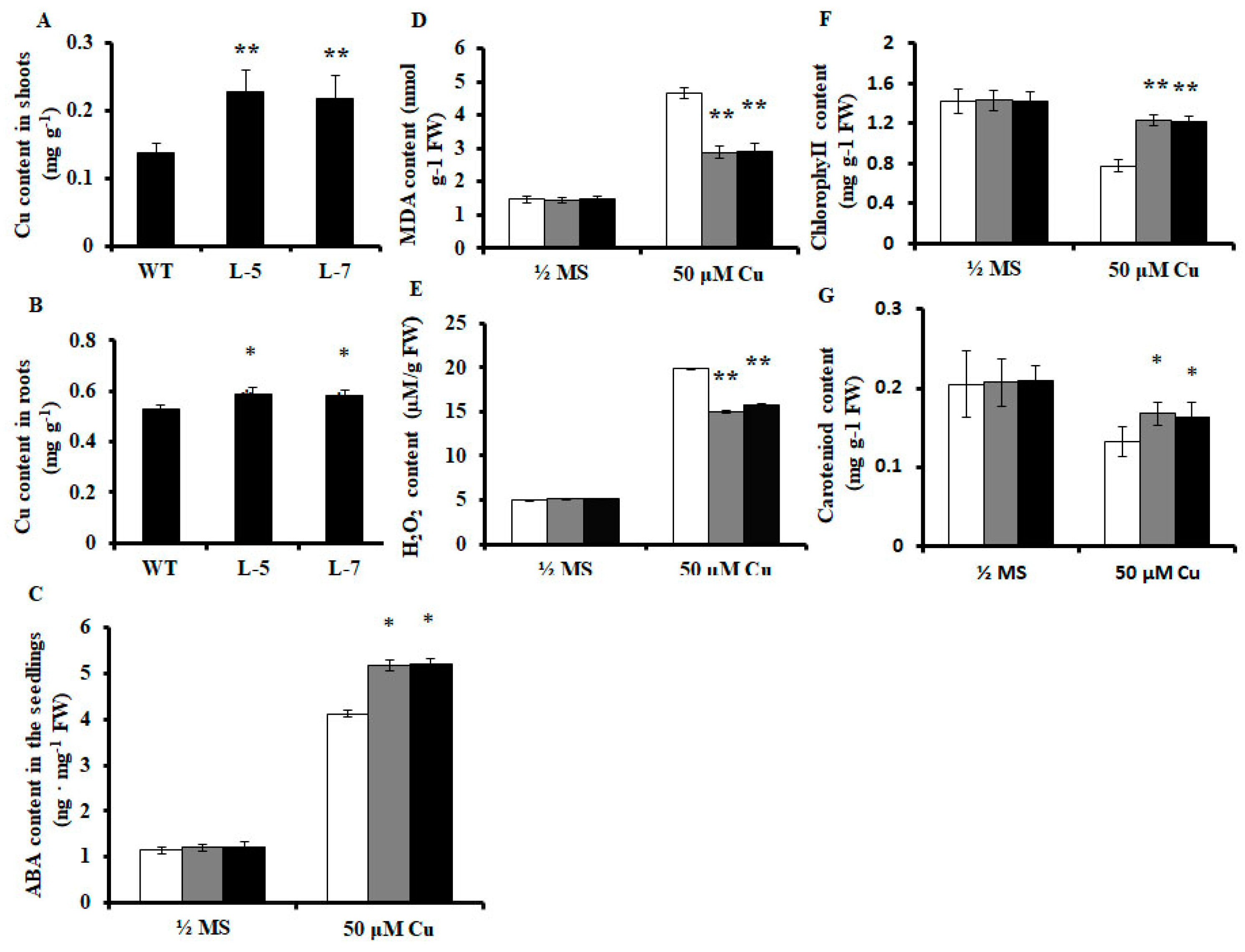
2.3. Expression of OsMSR3 in Arabidopsis Causes Higher Accumulation of Cu
2.4. Effects of OsMSR3 Expression on ABA, Malondialdehyde (MDA), and Hydrogen Peroxide (H2O2) Content in A. thaliana
2.5. Effects of OsMSR3 Expression in A. thaliana on Chlorophyll and Carotenoid Content
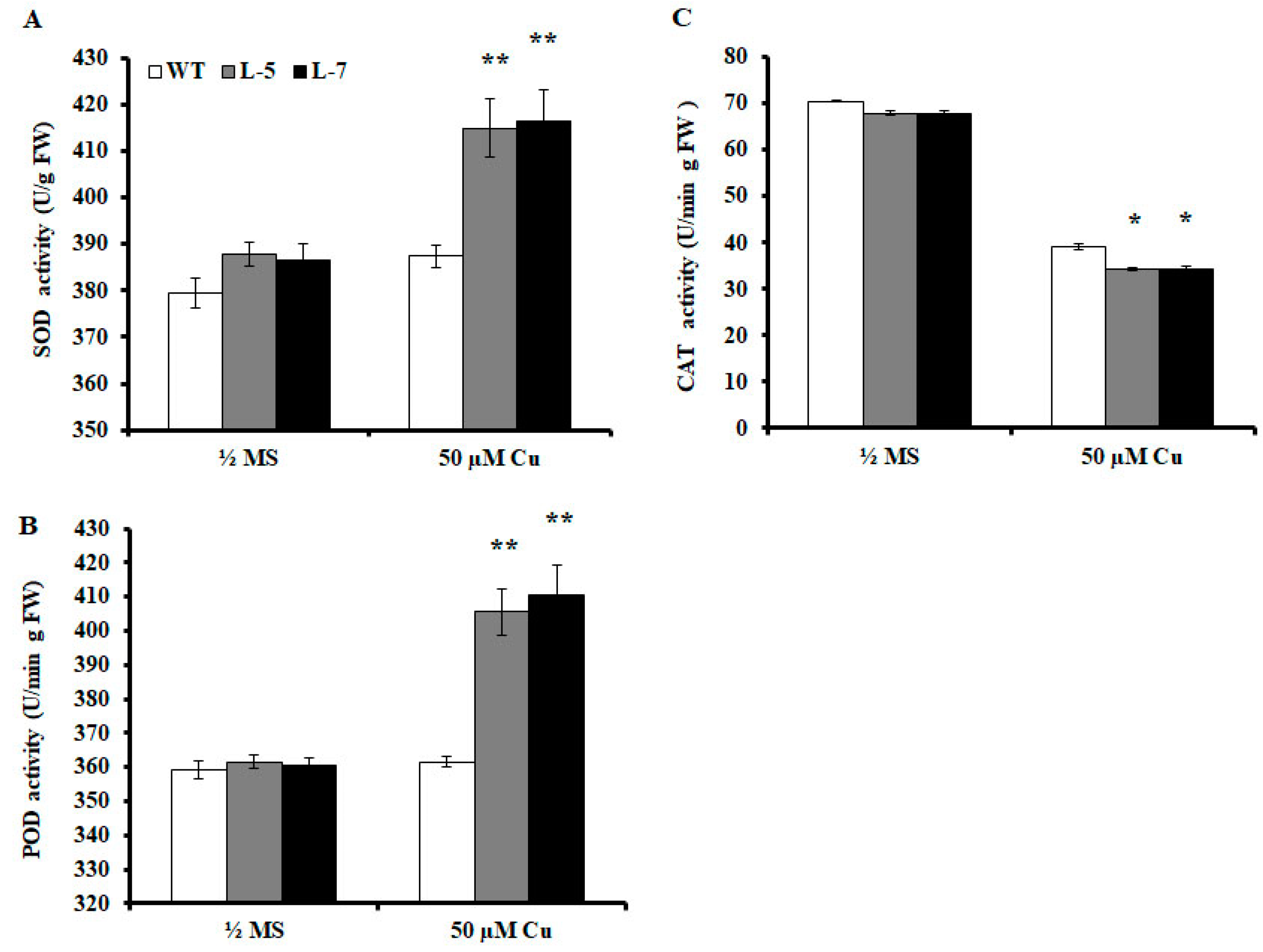
2.6. Antioxidant Enzyme Activities are Altered in Transgenic A. thaliana
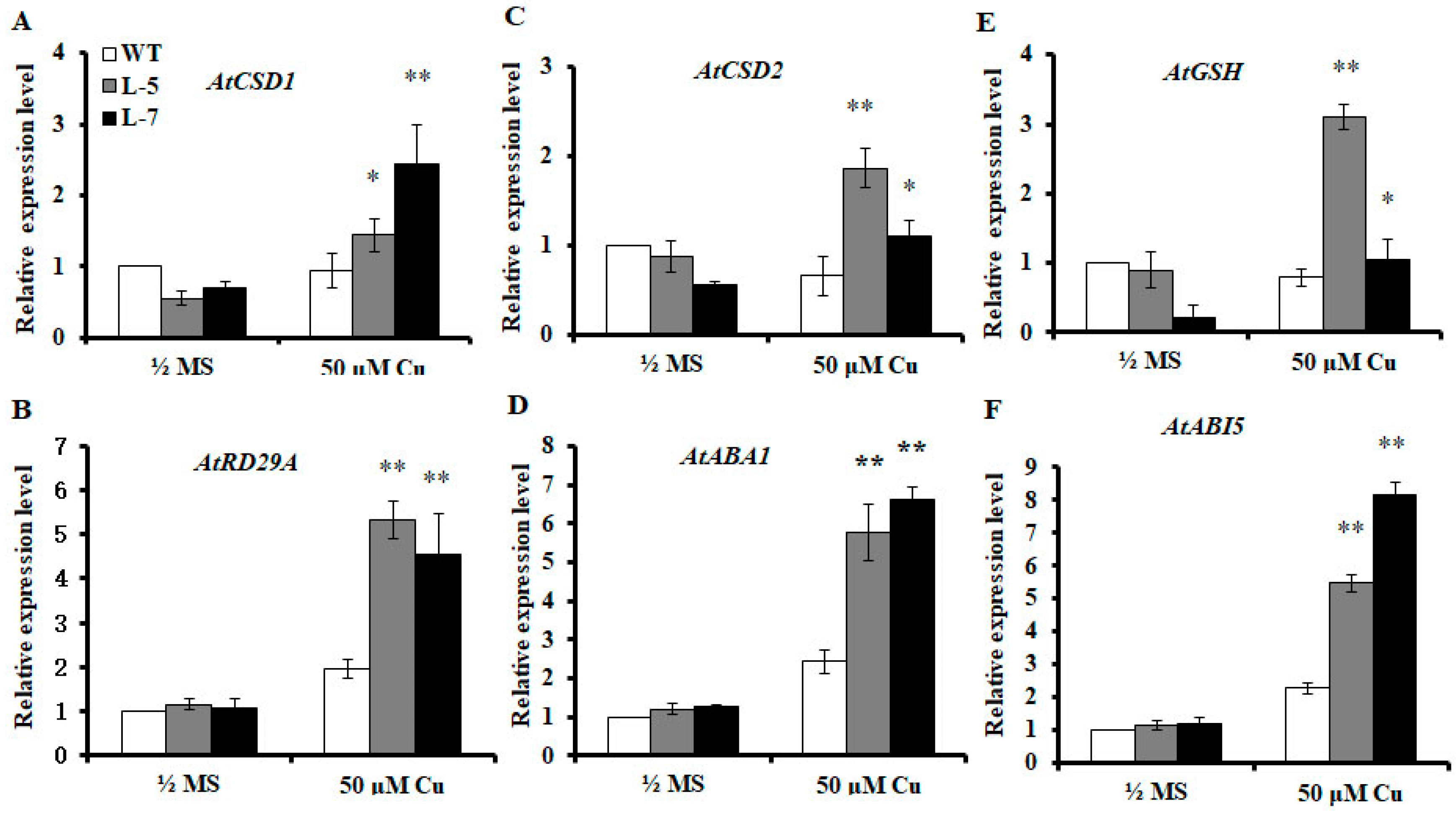
2.7. Expression of OsMSR3 Increases Expression of Antioxidant-Related and ABA-Responsive Genes
3. Discussion
4. Materials and Methods
4.1. Plant Material and Growth Conditions
4.2. Cu Tolerance Assay
4.3. RNA Extraction and qRT-PCR Analysis
4.4. Measurement of Cu Content
4.5. Measurement of MDA Content
4.6. Measurement of ABA Content
4.7. Measurement of Chlorophylls and Carotenoids
4.8. Quantitative Analysis of H2O2
4.9. Assay of Antioxidant Enzyme Activities
4.10. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ABA | abscisic acid |
| SOD | superoxide dismutase |
| POD | peroxidase |
| CAT | catalase |
| ROS | reactive oxygen species |
| ASC | ascorbic acid |
| GSH | glutathione |
| sHSPs | small heat shock protei |
| SD | standard deviation |
References
- Yruela, I.; Pueyo, J.J.; Alonso, P.J.; Picorel, R. Photoinhibition of photosystem II from higher plants effect of copper inhibition. J. Biol. Chem. 1996, 271, 27408–27415. [Google Scholar] [CrossRef]
- Rodriguez, F.I.; Esch, J.J.; Hall, A.E.; Binder, B.M.; Schaller, G.E.; Bleecker, A.B. A copper cofactor for the ethylene receptor ETR1 from Arabidopsis. Science 1999, 283, 996–998. [Google Scholar] [CrossRef]
- Himelblau, E.; Amasino, R.M. Delivering copper within plant cells. Curr. Opin. Plant Biol. 2000, 3, 205–210. [Google Scholar] [CrossRef]
- Pilon, M.; Abdel-Ghany, S.E.; Cohu, C.M.; Gogolin, K.A.; Ye, H. Copper cofactor delivery in plant cells. Curr. Opin. Plant Biol. 2006, 9, 256–263. [Google Scholar] [CrossRef]
- Burkhead, J.L.; Reynolds, K.A.; Abdel-Ghany, S.E.; Cohu, C.M.; Pilon, M. Copper homeostasis. New Phytol. 2009, 182, 799–816. [Google Scholar] [CrossRef]
- Bernal, M.; Casero, D.; Singh, V.; Wilson, G.T.; Grande, A.; Yang, H.; Dodani, S.C.; Pellegrini, M.; Huijser, P.; Connolly, E.L.; et al. Transcriptome sequencing identifies SPL7-regulated copper acquisition genes FRO4/FRO5 and the copper dependence of iron homeostasis in Arabidopsis. Plant Cell 2012, 24, 738–761. [Google Scholar] [CrossRef]
- Song, Y.; Zhou, L.; Yang, S.; Wang, C.; Zhang, T.; Wang, J. Dose-dependent sensitivity of Arabidopsis thaliana seedling root to copper is regulated by auxin homeostasis. Environ. Exp. Bot. 2017, 139, 23–30. [Google Scholar] [CrossRef]
- Luna, C.M.; Gonzalez, C.A.; Trippi, V.S. Oxidative damage caused by an excess of copper in oat leaves. Plant Cell Physiol. 1994, 35, 11–15. [Google Scholar] [CrossRef]
- Shen, Z.G.; Zhang, F.Q.; Zhang, F.S. Toxicity of copper and zinc in seedlings of mung bean and inducing accumulation of polyamine. J. Plant Nutr. 1998, 21, 1153–1162. [Google Scholar] [CrossRef]
- Patsikka, E.; Kairavuo, M.; Sersen, F.; Aro, E.M.; Tyystjarvi, E. Excess copper predisposes photosystem II to photoinhibition in vivo by outcompeting iron and causing decrease in leaf chlorophyll. Plant Physiol. 2002, 129, 1359–1367. [Google Scholar] [CrossRef]
- Nielsen, H.D.; Brownlee, C.; Coelho, S.M.; Brown, M.T. Inter-population differences in inherited copper tolerance involve photosynthetic adaptation and exclusion mechanisms in Fucus serratus. New Phytol. 2003, 160, 157–165. [Google Scholar] [CrossRef]
- Demirevska-Kepovaa, K.; Simova-Stoilovaa, L.; Stoyanovaa, Z.; Hölzerb, R.; Feller, U. Biochemical changes in barley plants after excessive supply of copper and manganese. Environ. Exp. Bot. 2004, 52, 253–266. [Google Scholar] [CrossRef]
- Drazkiewicz, M.; Skorzynska-Polit, E.; Krupa, Z. Copper-induced oxidative stress and antioxidant defence in Arabidopsis thaliana. Biometals 2004, 17, 379–387. [Google Scholar] [CrossRef]
- Wang, P.; De Schamphelaere, K.A.; Kopittke, P.M.; Zhou, D.M.; Peijnenburg, W.J.; Lock, K. Development of an electrostatic model predicting copper toxicity to plants. J. Exp. Bot. 2012, 63, 659–668. [Google Scholar] [CrossRef][Green Version]
- Navari-Izzo, F.; Cestone, B.; Cavallini, A.; Natali, L.; Giordani, T.; Quartacci, M.F. Copper excess triggers phospholipase D activity in wheat roots. Phytochemistry 2006, 67, 1232–1242. [Google Scholar] [CrossRef]
- Peto, A.; Lehotai, N.; Lozano-Juste, J.; Leon, J.; Tari, I.; Erdei, L.; Kolbert, Z. Involvement of nitric oxide and auxin in signal transduction of copper-induced morphological responses in Arabidopsis seedlings. Ann. Bot. 2011, 108, 449–457. [Google Scholar] [CrossRef]
- Marchand, L.; Nsanganwimana, F.; Lamy, J.B.; Quintela-Sabaris, C.; Gonnelli, C.; Colzi, I.; Fletcher, T.; Oustriere, N.; Kolbas, A.; Kidd, P.; et al. Root biomass production in populations of six rooted macrophytes in response to Cu exposure: Intra-specific variability versus constitutive-like tolerance. Environ. Pollut. 2014, 193, 205–215. [Google Scholar] [CrossRef]
- Liu, D.; Jiang, W.; Meng, Q.; Zou, J.; Gu, J.; Zeng, M. Cytogenetical and ultrastructural effects of copper on root meristem cells of Allium sativum L. Biocell 2009, 33, 25–32. [Google Scholar]
- Madejón, P.; Ramírez-Benítez, J.E.; Corrales, I.; Barceló, J.; Poschenrieder, C. Copper-induced oxidative damage and enhanced antioxidant defenses in the root apex of maize cultivars differing in Cu tolerance. Environ. Exp. Bot. 2009, 67, 415–420. [Google Scholar] [CrossRef]
- Yeh, C.M.; Hung, W.C.; Huang, H.J. Copper treatment activates mitogen-activated protein kinase signalling in rice. Physiol. Plant. 2003, 119, 392–399. [Google Scholar] [CrossRef]
- Zhan, E.; Zhou, H.; Li, S.; Liu, L.; Tan, T.; Lin, H. OTS1-dependent deSUMOylation increases tolerance to high copper levels in Arabidopsis. J. Integr. Plant Biol. 2018, 60, 310–322. [Google Scholar] [CrossRef] [PubMed]
- Clemens, S. Molecular mechanisms of plant metal tolerance and homeostasis. Planta 2001, 212, 475–486. [Google Scholar] [CrossRef] [PubMed]
- Sandalio, L.M.; Dalurzo, H.C.; Gomez, M.; Romero-Puertas, M.C.; Del Rio, L.A. Cadmium-induced changes in the growth and oxidative metabolism of pea plants. J. Exp. Bot. 2001, 52, 2115–2126. [Google Scholar] [CrossRef] [PubMed]
- Li, M.J.; Xu, G.Y.; Xia, X.J.; Wang, M.L.; Yin, X.M.; Zhang, B.; Zhang, X.; Cui, Y.C. Deciphering the physiological and molecular mechanisms for copper tolerance in autotetraploid Arabidopsis. Plant Cell Rep. 2017, 36, 1585–1597. [Google Scholar] [CrossRef] [PubMed]
- Jaleel, C.A.; Riadh, K.; Gopi, R.; Manivannan, P.; Ine’s, J.; Al-Juburi, H.J.; Zhao, C.X.; Shao, H.B.; Rajaram, P. Antioxidant defense responses: Physiological plasticity in higher plants under abiotic constraints. Acta Physiol. Plant. 2009, 31, 427–436. [Google Scholar] [CrossRef]
- Maksymiec, W.; Krupa, Z. The effects of short-term exposition to Cd, excess Cu ions and jasmonate on oxidative stress appearing in Arabidopsis thaliana. Environ. Exp. Bot. 2006, 57, 187–194. [Google Scholar] [CrossRef]
- Hollenbach, B.; Schreiber, L.; Hartung, W.; Dietz, K.J. Cadmium leads to stimulated expression of the lipid transfer protein genes in barley: Implications for the involvement of lipid transfer proteins in wax assembly. Planta 1997, 203, 9–19. [Google Scholar] [CrossRef]
- Bücker-Neto, L.; Paiva, A.L.S.; Machado, R.D.; Arenhart, R.; Margis-Pinheiro, M. Interactions between plant hormones and HMs responses. Genet. Mol. Biol. 2017, 40, 373–386. [Google Scholar] [CrossRef]
- Fediuc, E.; Lips, S.H.; Erdei, L. O-acetylserine (thiol) lyase activity in Phragmites and Typha plants under cadmium and NaCl stress conditions and the involvement of ABA in the stress response. J. Plant Physiol. 2005, 162, 865–872. [Google Scholar] [CrossRef]
- Stroinski, A.; Chadzinikolau, T.; Gizewska, K.; Zielezinska, M. ABA or cadmium induced phytochelatin synthesis in potato tubers. Biol. Plant. 2010, 54, 117–120. [Google Scholar] [CrossRef]
- Kim, Y.H.; Khan, A.L.; Kim, D.H.; Lee, S.Y.; Kim, K.M.; Waqas, M.; Jung, H.Y.; Shin, J.H.; Kim, J.G.; Lee, I.J. Silicon mitigates heavy metal stress by regulating P-type heavy metal ATPases, Oryza sativa low silicon genes, and endogenous phytohormones. BMC Plant Biol. 2014, 14, 13. [Google Scholar] [CrossRef] [PubMed]
- Munzuro, Ö.; Fikriye, K.Z.; Yahyagil, Z. The abscisic acid levels of wheat (Triticum aestivum L. cv. Çakmak 79) seeds that were germinated under heavy metal (Hg++, Cd++, Cu++) stress. Gazi Univ. J. Sci. 2008, 21, 1–7. [Google Scholar]
- Rauser, W.E.; Dumbroff, E.B. Effects of excess cobalt, nickel and zinc on the water relations of Phaseolus vulgaris. Environ. Exp. Bot. 1981, 21, 249–255. [Google Scholar] [CrossRef]
- Poschenrieder, C.; Gunsé, B.; Barceló, J. Influence of cadmium on water relations, stomatal resistance, and abscisic acid content in expanding bean leaves. Plant Physiol. 1989, 90, 1365–1371. [Google Scholar] [CrossRef]
- Sarkar, N.K.; Kim, Y.K.; Grover, A. Rice sHsp genes: Genomic organization and expression profiling under stress and development. BMC Genom. 2009, 10, 393. [Google Scholar] [CrossRef]
- Siddique, M.; Gernhard, S.; von Koskull-Doring, P.; Vierling, E.; Scharf, K.D. The plant sHSP superfamily: Five new members in Arabidopsis thaliana with unexpected properties. Cell Stress Chaperon. 2008, 13, 183–197. [Google Scholar] [CrossRef]
- Kim, K.H.; Alam, I.; Kim, Y.G.; Sharmin, S.A.; Lee, K.W.; Lee, S.H.; Lee, B.H. Overexpression of a chloroplast-localized small heat shock protein OsHSP26 confers enhanced tolerance against oxidative and heat stresses in tall fescue. Biotechnol. Lett. 2012, 34, 371–377. [Google Scholar] [CrossRef]
- Sato, Y.; Yokoya, S. Enhanced tolerance to drought stress in transgenic rice plants overexpressing a small heat-shock protein, sHSP17.7. Plant Cell Rep. 2008, 27, 329–334. [Google Scholar] [CrossRef]
- Ju, Y.; Tian, H.; Zhang, R.; Zuo, L.; Chu, Z. Overexpression of oshsp18.0-CI enhances resistance to bacterial leaf streak in rice. Rice 2017, 10, 12. [Google Scholar] [CrossRef]
- Cui, Y.C.; Xu, M.L.; Li, L.Y.; Wang, M.L.; Xu, G.Y.; Xia, X.J. Expression and cloning of a multiple stress responsive gene (OsMSR3) in rice. J. Wuhan. Bot. Res. 2009, 6, 574–581. [Google Scholar]
- Cui, Y.C.; Xu, G.Y.; Wang, M.L.; Yu, Y.; Li, M.J.; da Rocha, P.S.C.F.; Xia, X.J. Expression of OsMSR3 in Arabidopsis enhances tolerance to cadmium stress. Plant Cell Tiss. Org. Cult. 2013, 113, 331–340. [Google Scholar] [CrossRef]
- Kumar, G.; Kushwaha, H.R.; Panjabi-Sabharwal, V.; Kumari, S.; Joshi, R.; Karan, R.; Mittal, S.; Singla Pareek, S.L.; Pareek, A. Clustered metallothionein genes are co-regulated in rice and ectopic expression of OsMT1e-P confers multiple abiotic stress tolerance in tobacco via ROS scavenging. BMC Plant Biol. 2012, 12, 107. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Shafi, M.; Ma, J.; Zhong, B.; Guo, J.; Hu, X.; Xu, W.; Yang, Y.; Ruan, Z.; Wang, Y.; et al. Effect of amendments on contaminated soil of multiple heavy metals and accumulation of heavy metals in plants. Environ. Sci. Pollut. Res. 2018, 25, 28695–28704. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Li, S.S.; Li, X.Y.; Zhao, Z.Q.; Chen, S.B. An overview of current status of copper pollution in soil and remediation efforts in China. Earth Sci. Front. 2018, 25, 305–313. [Google Scholar]
- De Vos, C.H.R.; Ten Bookum, W.M.; Vooijs, R.; Schat, H.; Dekok, L.J. Effect of copper on fatty acid composition and peroxidation of lipids in the roots of copper tolerant and sensitive Silenecucubalus. Plant Physiol. Biochem. 1993, 31, 151–158. [Google Scholar]
- Heath, R.L.; Packer, L. Photoperoxidation in isolated chloroplasts: I. Kinetics and stoichiometry of fatty acid peroxidation. Arch. Biochem. Biophys. 1968, 125, 189–198. [Google Scholar] [CrossRef]
- Nishiyama, R.; Watanabe, Y.; Leyva-Gonzalez, M.A.; Van Ha, C.; Fujita, Y.; Tanaka, M.; Seki, M.; Yamaguchi-Shinozaki, K.; Shinozaki, K.; Herrera-Estrella, L.; et al. Arabidopsis AHP2, AHP3, and AHP5 histidine phosphotransfer proteins function as redundant negative regulators of drought stress response. Proc. Natl. Acad. Sci. USA 2013, 110, 4840–4845. [Google Scholar] [CrossRef]
- Demidchik, V.; Straltsova, D.; Medvedev, S.S.; Pozhvanov, G.A.; Sokolik, A.; Yurin, V. Stress-induced electrolyte leakage: The role of K+-permeable channels and involvement in programmed cell death and metabolic adjustment. J. Exp. Bot. 2014, 65, 1259–1270. [Google Scholar] [CrossRef]
- Rehman, M.; Liu, L.; Wang, Q.; Saleem, M.H.; Bashir, S.; Ullah, S.; Peng, D. Copper environmental toxicology, recent advances, and future outlook: A review. Environ. Sci. Pollut. Res. Int. 2019, 26, 18003–18016. [Google Scholar] [CrossRef]
- Sui, N.; Tian, S.; Wang, W.; Wang, M.; Fan, H. Overexpression of glycerol-3-phosphate acyltransferase from Suaeda salsa improves salt tolerance in Arabidopsis. Front. Plant Sci. 2017, 8, 1337. [Google Scholar] [CrossRef]
- Polívka, T.; Frank, H.A. Molecular factors controlling photosynthetic light harvesting by carotenoids. Acc. Chem. Res. 2010, 43, 1125–1134. [Google Scholar] [CrossRef] [PubMed]
- Polívka, T.; Pullerits, T.; Frank, H.A.; Cogdell, R.J.; Sundström, V. Ultrafast formation of a carotenoid radical in LH2 antenna complexes of purple bacteria. J. Phys. Chem. B 2004, 108, 15398–15407. [Google Scholar] [CrossRef]
- Saglam, A.; Yetiddin, F.; Demiralay, M.; Terzi, R. Copper stress and responses in plants. In Plant Metal Interaction: Emerging Remediation Techniques; Ahmad, P., Ed.; Elsevier: Amsterdam, The Netherlands, 2016; pp. 21–40. [Google Scholar] [CrossRef]
- Zabalza, A.; Galvez, L.; Marino, D.; Royuela, M.; Arrese-Igor, C.; Gonzalez, E.M. The application of ascorbate or its immediate precursor, galactono-1, 4-lactone, does not affect the response of nitrogen-fixing pea nodules to water stress. J. Plant Physiol. 2008, 165, 805–812. [Google Scholar] [CrossRef]
- Thounaojam, T.C.; Panda, P.; Mazumdar, P.; Kumar, D.; Sharma, G.; Sahoo, L.; Panda, S. Excess copper induced oxidative stress and response of antioxidants in rice. Plant Physiol. Biochem. 2012, 53, 33–39. [Google Scholar] [CrossRef] [PubMed]
- Gu, C.S.; Liu, L.Q.; Deng, Y.M.; Zhu, X.D.; Huang, S.Z.; Lu, X.Q. The heterologous expression of the Iris lactea var. chinensis type 2 metallothionein IlMT2b gene enhances copper tolerance in Arabidopsis thaliana. Bull. Environ. Contam. Toxicol. 2015, 94, 247–253. [Google Scholar] [CrossRef] [PubMed]
- Mittler, R. Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci. 2002, 7, 405–410. [Google Scholar] [CrossRef]
- Xu, G.Y.; Rocha, P.S.C.F.; Wang, M.L.; Xu, M.L.; Cui, Y.C.; Li, L.Y.; Zhu, Y.X.; Xia, X.J. A novel rice calmodulin-like gene, OsMSR2, enhances drought and salt tolerance and increases ABA sensitivity in Arabidopsis. Planta 2011, 234, 47–59. [Google Scholar] [CrossRef]
- Xu, M.L.; Chen, R.J.; Rocha, P.S.C.F.; Wang, M.L.; Xu, G.Y.; Xia, X.J. Expression and cloning of a novel stress responsive gene (OsMsr1) in rice. Acta Agron. Sin. 2008, 10, 1712–1718. [Google Scholar] [CrossRef]
- Cui, Y.C.; Li, M.J.; Yin, X.M.; Song, S.F.; Xu, G.Y.; Wang, M.L.; Li, C.Y.; Peng, C.; Xia, X.J. OsDSSR1, a novel small peptide, enhances drought tolerance in transgenic rice. Plant Sci. 2018, 270, 85–96. [Google Scholar] [CrossRef]
- Lichtenthaler, H.K.; Wellburn, A.R. Determinations of total carotenoids and chlorophylls a and b of leaf extracts in different solvents. Biochem. Soc. Trans. 1983, 11, 591e592. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cui, Y.; Wang, M.; Yin, X.; Xu, G.; Song, S.; Li, M.; Liu, K.; Xia, X. OsMSR3, a Small Heat Shock Protein, Confers Enhanced Tolerance to Copper Stress in Arabidopsis thaliana. Int. J. Mol. Sci. 2019, 20, 6096. https://doi.org/10.3390/ijms20236096
Cui Y, Wang M, Yin X, Xu G, Song S, Li M, Liu K, Xia X. OsMSR3, a Small Heat Shock Protein, Confers Enhanced Tolerance to Copper Stress in Arabidopsis thaliana. International Journal of Molecular Sciences. 2019; 20(23):6096. https://doi.org/10.3390/ijms20236096
Chicago/Turabian StyleCui, Yanchun, Manling Wang, Xuming Yin, Guoyun Xu, Shufeng Song, Mingjuan Li, Kai Liu, and Xinjie Xia. 2019. "OsMSR3, a Small Heat Shock Protein, Confers Enhanced Tolerance to Copper Stress in Arabidopsis thaliana" International Journal of Molecular Sciences 20, no. 23: 6096. https://doi.org/10.3390/ijms20236096
APA StyleCui, Y., Wang, M., Yin, X., Xu, G., Song, S., Li, M., Liu, K., & Xia, X. (2019). OsMSR3, a Small Heat Shock Protein, Confers Enhanced Tolerance to Copper Stress in Arabidopsis thaliana. International Journal of Molecular Sciences, 20(23), 6096. https://doi.org/10.3390/ijms20236096




