Goals and Challenges in Bacterial Phosphoproteomics
Abstract
1. Introduction
2. Bacterial Ser/Thr/Tyr Phosphoproteomics
2.1. Gel-Based Analyses
2.2. LC-MS/MS-Based Phosphoproteomic Analyses
2.2.1. Bacterial Ser/Thr/Tyr Nonquantitative LC-MS/MS-Based Phosphoproteomic Analyses
2.2.2. Bacterial Ser/Thr/Tyr LC-MS/MS-Based Quantitative Phosphoproteomic Analyses
3. Bacterial Proteins and Pathways Modulated by Ser/Thr/Tyr Phosphorylation
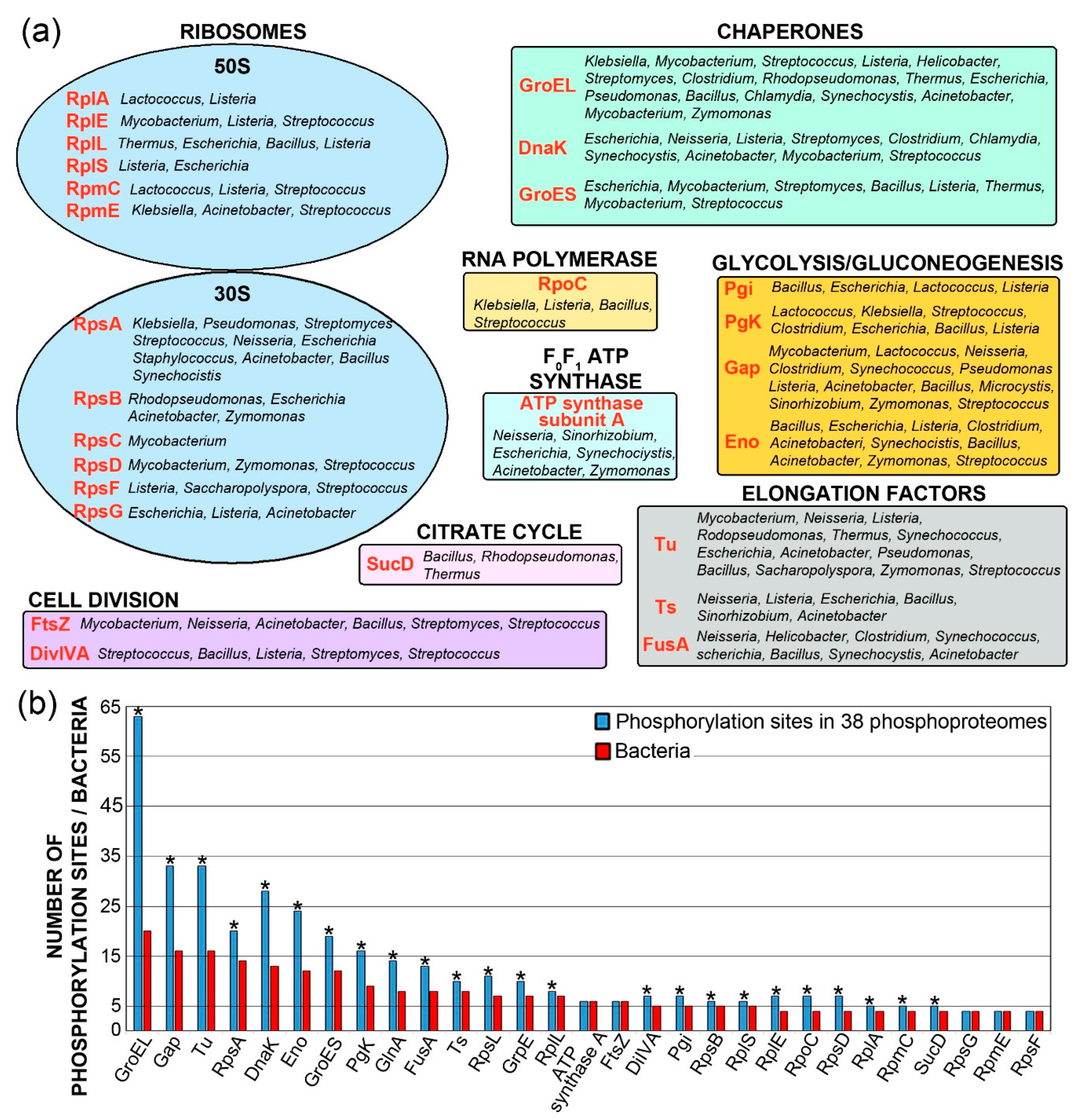
3.1. Bacterial Proteins Identified as Phosphorylated
3.2. Bacterial Ser/Thr/Tyr Phosphorylation Motifs
3.3. Bacterial Processes Demonstrated to be Modulated by Ser/Thr/Tyr Phosphorylation
4. Bacterial Histidine Protein Phosphorylation
4.1. Methodological Challenges
4.2. Bacterial Pathways Modulated by Histidine Phosphorylation
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| LC-MS/MS | liquid chromatography tandem mass spectrometry |
| IMAC | immobilised metal affinity chromatography |
| CPP | calcium phosphate precipitation |
| SILAC | stable isotope labelling by amino acids in cell culture |
| TMT | tandem mass tag |
| sMRM | scheduled multiple reaction monitoring |
References
- Pawson, T.; Scott, J.D. Protein phosphorylation in signaling-50 years and counting. Trends Biochem. Sci. 2005, 30, 286–290. [Google Scholar] [CrossRef] [PubMed]
- Sharma, K.; D’Souza, R.C.; Tyanova, S.; Schaab, C.; Wisniewski, J.R.; Cox, J.; Mann, M. Ultradeep human phosphoproteome reveals a distinct regulatory nature of Tyr and Ser/Thr-based signaling. Cell Rep. 2014, 8, 1583–1594. [Google Scholar] [CrossRef] [PubMed]
- Prisic, S.; Dankwa, S.; Schwartz, D.; Chou, M.F.; Locasale, J.W.; Kang, C.M.; Bemis, G.; Church, G.M.; Steen, H.; Husson, R.N. Extensive phosphorylation with overlapping specificity by Mycobacterium tuberculosis serine/threonine protein kinases. Proc. Natl. Acad. Sci. USA 2010, 107, 7521–7526. [Google Scholar] [CrossRef] [PubMed]
- Adam, K.; Hunter, T. Histidine kinases and the missing phosphoproteome from prokaryotes to eukaryotes. Lab. Invest. 2018, 98, 233–247. [Google Scholar] [CrossRef]
- Lai, S.J.; Tu, I.F.; Wu, W.L.; Yang, J.T.; Luk, L.Y.P.; Lai, M.C.; Tsai, Y.H.; Wu, S.H. Site-specific His/Asp phosphoproteomic analysis of prokaryotes reveals putative targets for drug resistance. BMC Microbiol. 2017, 17, 123. [Google Scholar] [CrossRef]
- Lai, J.H.; Yang, J.T.; Chern, J.; Chen, T.L.; Wu, W.L.; Liao, J.H.; Tsai, S.F.; Liang, S.Y.; Chou, C.C.; Wu, S.H. Comparative Phosphoproteomics Reveals the Role of AmpC beta-lactamase Phosphorylation in the Clinical Imipenem-resistant Strain Acinetobacter baumannii SK17. Mol. Cell Proteom. 2016, 15, 12–25. [Google Scholar] [CrossRef]
- Lin, M.H.; Hsu, T.L.; Lin, S.Y.; Pan, Y.J.; Jan, J.T.; Wang, J.T.; Khoo, K.H.; Wu, S.H. Phosphoproteomics of Klebsiella pneumoniae NTUH-K2044 reveals a tight link between tyrosine phosphorylation and virulence. Mol. Cell Proteom. 2009, 8, 2613–2623. [Google Scholar] [CrossRef]
- Macek, B.; Mijakovic, I.; Olsen, J.V.; Gnad, F.; Kumar, C.; Jensen, P.R.; Mann, M. The serine/threonine/tyrosine phosphoproteome of the model bacterium Bacillus subtilis. Mol. Cell Proteom. 2007, 6, 697–707. [Google Scholar] [CrossRef]
- Macek, B.; Gnad, F.; Soufi, B.; Kumar, C.; Olsen, J.V.; Mijakovic, I.; Mann, M. Phosphoproteome analysis of E. coli reveals evolutionary conservation of bacterial Ser/Thr/Tyr phosphorylation. Mol. Cell Proteom. 2008, 7, 299–307. [Google Scholar] [CrossRef]
- Soufi, B.; Gnad, F.; Jensen, P.R.; Petranovic, D.; Mann, M.; Mijakovic, I.; Macek, B. The Ser/Thr/Tyr phosphoproteome of Lactococcus lactis IL1403 reveals multiply phosphorylated proteins. Proteomics 2008, 8, 3486–3493. [Google Scholar] [CrossRef]
- Ravichandran, A.; Sugiyama, N.; Tomita, M.; Swarup, S.; Ishihama, Y. Ser/Thr/Tyr phosphoproteome analysis of pathogenic and non-pathogenic Pseudomonas species. Proteomics 2009, 9, 2764–2775. [Google Scholar] [CrossRef] [PubMed]
- Aivaliotis, M.; Macek, B.; Gnad, F.; Reichelt, P.; Mann, M.; Oesterhelt, D. Ser/Thr/Tyr protein phosphorylation in the archaeon Halobacterium salinarum--a representative of the third domain of life. PLoS ONE 2009, 4, e4777. [Google Scholar] [CrossRef] [PubMed]
- Parker, J.L.; Jones, A.M.; Serazetdinova, L.; Saalbach, G.; Bibb, M.J.; Naldrett, M.J. Analysis of the phosphoproteome of the multicellular bacterium Streptomyces coelicolor A3(2) by protein/peptide fractionation, phosphopeptide enrichment and high-accuracy mass spectrometry. Proteomics 2010, 10, 2486–2497. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Ge, F.; Xiao, C.L.; Yin, X.F.; Ge, R.; Zhang, L.H.; He, Q.Y. Phosphoproteomic analysis reveals the multiple roles of phosphorylation in pathogenic bacterium Streptococcus pneumoniae. J. Proteome Res. 2010, 9, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Soufi, B.; Kumar, C.; Gnad, F.; Mann, M.; Mijakovic, I.; Macek, B. Stable isotope labeling by amino acids in cell culture (SILAC) applied to quantitative proteomics of Bacillus subtilis. J. Proteome Res. 2010, 9, 3638–3646. [Google Scholar] [CrossRef] [PubMed]
- Bernardini, G.; Laschi, M.; Serchi, T.; Arena, S.; D’Ambrosio, C.; Braconi, D.; Scaloni, A.; Santucci, A. Mapping phosphoproteins in Neisseria meningitidis serogroup A. Proteomics 2011, 11, 1351–1358. [Google Scholar] [CrossRef]
- Manteca, A.; Ye, J.; Sanchez, J.; Jensen, O.N. Phosphoproteome analysis of Streptomyces development reveals extensive protein phosphorylation accompanying bacterial differentiation. J. Proteome Res. 2011, 10, 5481–5492. [Google Scholar] [CrossRef]
- Misra, S.K.; Milohanic, E.; Ake, F.; Mijakovic, I.; Deutscher, J.; Monnet, V.; Henry, C. Analysis of the serine/threonine/tyrosine phosphoproteome of the pathogenic bacterium Listeria monocytogenes reveals phosphorylated proteins related to virulence. Proteomics 2011, 11, 4155–4165. [Google Scholar] [CrossRef]
- Ge, R.; Sun, X.; Xiao, C.; Yin, X.; Shan, W.; Chen, Z.; He, Q.Y. Phosphoproteome analysis of the pathogenic bacterium Helicobacter pylori reveals over-representation of tyrosine phosphorylation and multiply phosphorylated proteins. Proteomics 2011, 11, 1449–1461. [Google Scholar] [CrossRef]
- Bai, X.; Ji, Z. Phosphoproteomic investigation of a solvent producing bacterium Clostridium acetobutylicum. Appl. Microbiol. Biotechnol. 2012, 95, 201–211. [Google Scholar] [CrossRef]
- Hu, C.W.; Lin, M.H.; Huang, H.C.; Ku, W.C.; Yi, T.H.; Tsai, C.F.; Chen, Y.J.; Sugiyama, N.; Ishihama, Y.; Juan, H.F.; et al. Phosphoproteomic analysis of Rhodopseudomonas palustris reveals the role of pyruvate phosphate dikinase phosphorylation in lipid production. J. Proteome Res. 2012, 11, 5362–5375. [Google Scholar] [CrossRef] [PubMed]
- Takahata, Y.; Inoue, M.; Kim, K.; Iio, Y.; Miyamoto, M.; Masui, R.; Ishihama, Y.; Kuramitsu, S. Close proximity of phosphorylation sites to ligand in the phosphoproteome of the extreme thermophile Thermus thermophilus HB8. Proteomics 2012, 12, 1414–1430. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.L.; Liao, J.H.; Lin, G.H.; Lin, M.H.; Chang, Y.C.; Liang, S.Y.; Yang, F.L.; Khoo, K.H.; Wu, S.H. Phosphoproteomic analysis reveals the effects of PilF phosphorylation on type IV pilus and biofilm formation in Thermus thermophilus HB27. Mol. Cell Proteom. 2013, 12, 2701–2713. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.K.; Qiao, Z.X.; Zhang, W.Y.; Xiong, Q.; Zhang, J.; Li, T.; Ge, F.; Zhao, J.D. Global phosphoproteomic analysis reveals diverse functions of serine/threonine/tyrosine phosphorylation in the model cyanobacterium Synechococcus sp. strain PCC 7002. J. Proteome Res. 2013, 12, 1909–1923. [Google Scholar] [CrossRef] [PubMed]
- Soares, N.C.; Spat, P.; Krug, K.; Macek, B. Global dynamics of the Escherichia coli proteome and phosphoproteome during growth in minimal medium. J. Proteome Res. 2013, 12, 2611–2621. [Google Scholar] [CrossRef]
- Basell, K.; Otto, A.; Junker, S.; Zuhlke, D.; Rappen, G.M.; Schmidt, S.; Hentschker, C.; Macek, B.; Ohlsen, K.; Hecker, M.; et al. The phosphoproteome and its physiological dynamics in Staphylococcus aureus. Int. J. Med. Microbiol. 2014, 304, 121–132. [Google Scholar] [CrossRef]
- Soares, N.C.; Spat, P.; Mendez, J.A.; Nakedi, K.; Aranda, J.; Bou, G. Ser/Thr/Tyr phosphoproteome characterization of Acinetobacter baumannii: Comparison between a reference strain and a highly invasive multidrug-resistant clinical isolate. J. Proteom. 2014, 102, 113–124. [Google Scholar] [CrossRef]
- Ouidir, T.; Jarnier, F.; Cosette, P.; Jouenne, T.; Hardouin, J. Extracellular Ser/Thr/Tyr phosphorylated proteins of Pseudomonas aeruginosa PA14 strain. Proteomics 2014, 14, 2017–2030. [Google Scholar] [CrossRef]
- Misra, S.K.; Moussan Desiree Ake, F.; Wu, Z.; Milohanic, E.; Cao, T.N.; Cossart, P.; Deutscher, J.; Monnet, V.; Archambaud, C.; Henry, C. Quantitative proteome analyses identify PrfA-responsive proteins and phosphoproteins in Listeria monocytogenes. J. Proteome Res. 2014, 13, 6046–6057. [Google Scholar] [CrossRef]
- Licona-Cassani, C.; Lim, S.; Marcellin, E.; Nielsen, L.K. Temporal dynamics of the Saccharopolyspora erythraea phosphoproteome. Mol. Cell Proteom. 2014, 13, 1219–1230. [Google Scholar] [CrossRef]
- Ravikumar, V.; Shi, L.; Krug, K.; Derouiche, A.; Jers, C.; Cousin, C.; Kobir, A.; Mijakovic, I.; Macek, B. Quantitative phosphoproteome analysis of Bacillus subtilis reveals novel substrates of the kinase PrkC and phosphatase PrpC. Mol. Cell Proteom. 2014, 13, 1965–1978. [Google Scholar] [CrossRef] [PubMed]
- Fisher, D.J.; Adams, N.E.; Maurelli, A.T. Phosphoproteomic analysis of the Chlamydia caviae elementary body and reticulate body forms. Microbiology 2015, 161, 1648–1658. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Tian, C.F.; Chen, W.X. Site-Specific Ser/Thr/Tyr Phosphoproteome of Sinorhizobium meliloti at Stationary Phase. PLoS ONE 2015, 10, e0139143. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.; Marcellin, E.; Jacob, S.; Nielsen, L.K. Global dynamics of Escherichia coli phosphoproteome in central carbon metabolism under changing culture conditions. J. Proteom. 2015, 126, 24–33. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, A.; Soufi, B.; Ravikumar, V.; Soares, N.C.; Krug, K.; Smith, Y.; Macek, B.; Ben-Yehuda, S. Phosphoproteome dynamics mediate revival of bacterial spores. BMC Biol. 2015, 13, 76. [Google Scholar] [CrossRef] [PubMed]
- Spat, P.; Macek, B.; Forchhammer, K. Phosphoproteome of the cyanobacterium Synechocystis sp. PCC 6803 and its dynamics during nitrogen starvation. Front. Microbiol. 2015, 6, 248. [Google Scholar] [CrossRef]
- Albeldas, C.; Ganief, N.; Calder, B.; Nakedi, K.C.; Garnett, S.; Nel, A.J.M.; Blackburn, J.M.; Soares, N.C. Global proteome and phosphoproteome dynamics indicate novel mechanisms of vitamin C induced dormancy in Mycobacterium smegmatis. J. Proteom. 2018, 180, 1–10. [Google Scholar] [CrossRef]
- Verma, R.; Pinto, S.M.; Patil, A.H.; Advani, J.; Subba, P.; Kumar, M.; Sharma, J.; Dey, G.; Ravikumar, R.; Buggi, S.; et al. Quantitative Proteomic and Phosphoproteomic Analysis of H37Ra and H37Rv Strains of Mycobacterium tuberculosis. J. Proteome Res. 2017, 16, 1632–1645. [Google Scholar] [CrossRef]
- Qu, J.; Shen, L.; Zhao, M.; Li, W.; Jia, C.; Zhu, H.; Zhang, Q. Determination of the Role of Microcystis aeruginosa in Toxin Generation Based on Phosphoproteomic Profiles. Toxins 2018, 10, 304. [Google Scholar] [CrossRef]
- Rioseras, B.; Shliaha, P.V.; Gorshkov, V.; Yague, P.; Lopez-Garcia, M.T.; Gonzalez-Quinonez, N.; Kovalchuk, S.; Rogowska-Wrzesinska, A.; Jensen, O.N.; Manteca, A. Quantitative Proteome and Phosphoproteome Analyses of Streptomyces coelicolor Reveal Proteins and Phosphoproteins Modulating Differentiation and Secondary Metabolism. Mol. Cell Proteom. 2018, 17, 1591–1611. [Google Scholar] [CrossRef]
- Tatli, M.; Hebert, A.S.; Coon, J.J.; Amador-Noguez, D. Genome Wide Phosphoproteome Analysis of Zymomonas mobilis Under Anaerobic, Aerobic, and N2-Fixing Conditions. Front. Microbiol. 2019, 10, 1986. [Google Scholar] [CrossRef] [PubMed]
- Henry, C.; Haller, L.; Blein-Nicolas, M.; Zivy, M.; Canette, A.; Verbrugghe, M.; Mezange, C.; Boulay, M.; Gardan, R.; Samson, S.; et al. Identification of Hanks-Type Kinase PknB-Specific Targets in the Streptococcus thermophilus Phosphoproteome. Front. Microbiol. 2019, 10, 1329. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, G.K.; Thelen, J.J. A high-resolution two dimensional Gel- and Pro-Q DPS-based proteomics workflow for phosphoprotein identification and quantitative profiling. Methods Mol. Biol. 2009, 527, 3–19, ix. [Google Scholar] [CrossRef] [PubMed]
- Simpson, K.; Browning, M. Antibodies That Work Again and Again and Again. In Signal Transduction Immunohistochemistry: Methods and Protocols; Kalyuzhny, A.E., Ed.; Springer: New York, NY, USA, 2017; pp. 41–59. [Google Scholar] [CrossRef]
- Rogowska-Wrzesinska, A.; Le Bihan, M.C.; Thaysen-Andersen, M.; Roepstorff, P. 2D gels still have a niche in proteomics. J. Proteom. 2013, 88, 4–13. [Google Scholar] [CrossRef] [PubMed]
- Thingholm, T.E.; Larsen, M.R. Phosphopeptide Enrichment by Immobilized Metal Affinity Chromatography. Methods Mol. Biol. 2016, 1355, 123–133. [Google Scholar] [CrossRef]
- Zhang, X.; Ye, J.; Jensen, O.N.; Roepstorff, P. Highly Efficient Phosphopeptide Enrichment by Calcium Phosphate Precipitation Combined with Subsequent IMAC Enrichment. Mol. Cell Proteom. 2007, 6, 2032–2042. [Google Scholar] [CrossRef]
- Xia, Q.; Cheng, D.; Duong, D.M.; Gearing, M.; Lah, J.J.; Levey, A.I.; Peng, J. Phosphoproteomic analysis of human brain by calcium phosphate precipitation and mass spectrometry. J. Proteome Res. 2008, 7, 2845–2851. [Google Scholar] [CrossRef]
- Zhao, X.; Leon, I.R.; Bak, S.; Mogensen, M.; Wrzesinski, K.; Hojlund, K.; Jensen, O.N. Phosphoproteome analysis of functional mitochondria isolated from resting human muscle reveals extensive phosphorylation of inner membrane protein complexes and enzymes. Mol. Cell Proteom. 2011, 10, M110 000299. [Google Scholar] [CrossRef]
- Schindler, J.; Ye, J.; Jensen, O.N.; Nothwang, H.G. Monitoring the native phosphorylation state of plasma membrane proteins from a single mouse cerebellum. J. Neurosci. Methods 2013, 213, 153–164. [Google Scholar] [CrossRef]
- Reales-Calderon, J.A.; Sylvester, M.; Strijbis, K.; Jensen, O.N.; Nombela, C.; Molero, G.; Gil, C. Candida albicans induces pro-inflammatory and anti-apoptotic signals in macrophages as revealed by quantitative proteomics and phosphoproteomics. J. Proteom. 2013, 91, 106–135. [Google Scholar] [CrossRef]
- Fischbach, M.A.; Walsh, C.T. Antibiotics for emerging pathogens. Science 2009, 325, 1089–1093. [Google Scholar] [CrossRef] [PubMed]
- Shah, I.M.; Laaberki, M.H.; Popham, D.L.; Dworkin, J. A eukaryotic-like Ser/Thr kinase signals bacteria to exit dormancy in response to peptidoglycan fragments. Cell 2008, 135, 486–496. [Google Scholar] [CrossRef] [PubMed]
- Shah, I.M.; Dworkin, J. Induction and regulation of a secreted peptidoglycan hydrolase by a membrane Ser/Thr kinase that detects muropeptides. Mol. Microbiol. 2010, 75, 1232–1243. [Google Scholar] [CrossRef] [PubMed]
- Maurya, G.K.; Modi, K.; Banerjee, M.; Chaudhary, R.; Rajpurohit, Y.S.; Misra, H.S. Phosphorylation of FtsZ and FtsA by a DNA Damage-Responsive Ser/Thr Protein Kinase Affects Their Functional Interactions in Deinococcus radiodurans. Msphere 2018, 3. [Google Scholar] [CrossRef]
- Hempel, A.M.; Cantlay, S.; Molle, V.; Wang, S.B.; Naldrett, M.J.; Parker, J.L.; Richards, D.M.; Jung, Y.G.; Buttner, M.J.; Flardh, K. The Ser/Thr protein kinase AfsK regulates polar growth and hyphal branching in the filamentous bacteria Streptomyces. Proc. Natl. Acad. Sci. USA 2012, 109, E2371–E2379. [Google Scholar] [CrossRef]
- Ni, H.; Fan, W.; Li, C.; Wu, Q.; Hou, H.; Hu, D.; Zheng, F.; Zhu, X.; Wang, C.; Cao, X.; et al. Streptococcus suis DivIVA Protein Is a Substrate of Ser/Thr Kinase STK and Involved in Cell Division Regulation. Front. Cell Infect. Microbiol. 2018, 8, 85. [Google Scholar] [CrossRef]
- Yang, Z.; Wang, X.; Xu, W.; Zhou, M.; Zhang, Y.; Ma, Y.; Wang, Q. Phosphorylation of PppA at threonine 253 controls T6SS2 expression and bacterial killing capacity in the marine pathogen Vibrio alginolyticus. Microbiol. Res. 2018, 209, 70–78. [Google Scholar] [CrossRef]
- Yang, Z.; Zhou, X.; Ma, Y.; Zhou, M.; Waldor, M.K.; Zhang, Y.; Wang, Q. Serine/threonine kinase PpkA coordinates the interplay between T6SS2 activation and quorum sensing in the marine pathogen Vibrio alginolyticus. Environ. Microbiol. 2018, 20, 903–919. [Google Scholar] [CrossRef]
- Zhang, C.; Sun, W.; Tan, M.; Dong, M.; Liu, W.; Gao, T.; Li, L.; Xu, Z.; Zhou, R. The Eukaryote-Like Serine/Threonine Kinase STK Regulates the Growth and Metabolism of Zoonotic Streptococcus suis. Front. Cell Infect. Microbiol. 2017, 7, 66. [Google Scholar] [CrossRef]
- Falord, M.; Mader, U.; Hiron, A.; Debarbouille, M.; Msadek, T. Investigation of the Staphylococcus aureus GraSR regulon reveals novel links to virulence, stress response and cell wall signal transduction pathways. PLoS ONE 2011, 6, e21323. [Google Scholar] [CrossRef]
- Lin, W.J.; Walthers, D.; Connelly, J.E.; Burnside, K.; Jewell, K.A.; Kenney, L.J.; Rajagopal, L. Threonine phosphorylation prevents promoter DNA binding of the Group B Streptococcus response regulator CovR. Mol. Microbiol. 2009, 71, 1477–1495. [Google Scholar] [CrossRef] [PubMed]
- Sajid, A.; Arora, G.; Gupta, M.; Singhal, A.; Chakraborty, K.; Nandicoori, V.K.; Singh, Y. Interaction of Mycobacterium tuberculosis elongation factor Tu with GTP is regulated by phosphorylation. J. Bacteriol. 2011, 193, 5347–5358. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Zhan, J.; Chen, Y.; Yang, M.; He, C.; Ge, F.; Wang, Q. Effects of Phosphorylation of beta Subunits of Phycocyanins on State Transition in the Model Cyanobacterium Synechocystis sp. PCC 6803. Plant. Cell Physiol. 2015, 56, 1997–2013. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Teper, D.; Andrade, M.O.; Zhang, T.; Chen, S.; Song, W.Y.; Wang, N. A Phosphorylation Switch on Lon Protease Regulates Bacterial Type III Secretion System in Host. MBio 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Spudich, J.L.; Stoeckenius, W. Light-regulated retinal-dependent reversible phosphorylation of Halobacterium proteins. J. Biol. Chem. 1980, 255, 5501–5503. [Google Scholar]
- Wagner, P.D.; Steeg, P.S.; Vu, N.D. Two-component kinase-like activity of nm23 correlates with its motility-suppressing activity. Proc. Natl. Acad. Sci. USA 1997, 94, 9000–9005. [Google Scholar] [CrossRef]
- Sickmann, A.; Meyer, H.E. Phosphoamino acid analysis. Proteomics 2001, 1, 200–206. [Google Scholar] [CrossRef]
- Fuhs, S.R.; Meisenhelder, J.; Aslanian, A.; Ma, L.; Zagorska, A.; Stankova, M.; Binnie, A.; Al-Obeidi, F.; Mauger, J.; Lemke, G.; et al. Monoclonal 1- and 3-Phosphohistidine Antibodies: New Tools to Study Histidine Phosphorylation. Cell 2015, 162, 198–210. [Google Scholar] [CrossRef]
- Kee, J.M.; Oslund, R.C.; Perlman, D.H.; Muir, T.W. A pan-specific antibody for direct detection of protein histidine phosphorylation. Nat. Chem. Biol. 2013, 9, 416–421. [Google Scholar] [CrossRef]
- Kleinnijenhuis, A.J.; Kjeldsen, F.; Kallipolitis, B.; Haselmann, K.F.; Jensen, O.N. Analysis of histidine phosphorylation using tandem MS and ion-electron reactions. Anal. Chem. 2007, 79, 7450–7456. [Google Scholar] [CrossRef]
- Bednarz, B.; Kotowska, M.; Pawlik, K.J. Multi-level regulation of coelimycin synthesis in Streptomyces coelicolor A3(2). Appl. Microbiol. Biotechnol. 2019, 103, 6423–6434. [Google Scholar] [CrossRef] [PubMed]
- Moglich, A. Signal transduction in photoreceptor histidine kinases. Protein Sci. 2019. [Google Scholar] [CrossRef] [PubMed]
- Jung, K.; Brameyer, S.; Fabiani, F.; Gasperotti, A.; Hoyer, E. Phenotypic Heterogeneity Generated by Histidine Kinase-Based Signaling Networks. J. Mol. Biol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Rao, X.; Zhang, K. Nucleoside diphosphate kinase (Ndk): A pleiotropic effector manipulating bacterial virulence and adaptive responses. Microbiol. Res. 2017, 205, 125–134. [Google Scholar] [CrossRef] [PubMed]
- Borland, S.; Prigent-Combaret, C.; Wisniewski-Dye, F. Bacterial hybrid histidine kinases in plant-bacteria interactions. Microbiology 2016, 162, 1715–1734. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Pan, X.; Xu, N.; Guo, M. Bacterial chemotaxis coupling protein: Structure, function and diversity. Microbiol. Res. 2019, 219, 40–48. [Google Scholar] [CrossRef]
- Humphrey, S.J.; Karayel, O.; James, D.E.; Mann, M. High-throughput and high-sensitivity phosphoproteomics with the EasyPhos platform. Nat. Protoc. 2018, 13, 1897–1916. [Google Scholar] [CrossRef]
| Bacterium | Year | pSer (%) | pThr (%) | pTyr (%) | Reference |
|---|---|---|---|---|---|
| Bacillus subtillis | 2007 | 69.2 | 20.5 | 10.3 | [8] |
| Escherichia coli (E. coli) | 2008 | 68 | 23 | 9 | [9] |
| Lactococcus lactis | 2008 | 46.5 | 50.6 | 2.7 | [10] |
| Klebsiella pneumoniae | 2009 | 31.2 | 15.1 | 25.8 | [7] |
| Pseudomonas aeruginosa/putida | 2009 | 52.8 | 36.1 | 11.1 | [11] |
| Halobacterium salinarum | 2009 | 84 | 16 | 0 | [12] |
| Mycobacterium tuberculosis | 2010 | 40 | 60 | 0 | [3] |
| Streptomyces coelicolor | 2010 | 34 | 52 | 14 | [13] |
| Streptococcus pneumoniae | 2010 | 47 | 44 | 9 | [14] |
| Bacillus subtilis | 2010 | n.r. | n.r. | n.r. | [15] |
| Neisseria meningitidis | 2011 | n.r. | n.r. | n.r. | [16] |
| Streptomyces coelicolor | 2011 | 46.8 | 48 | 5.2 | [17] |
| Listeria monocytogenes | 2011 | 93 | 43 | 7 | [18] |
| Helicobacter pylori | 2011 | 42.8 | 38.7 | 18.5 | [19] |
| Clostridium acetobutylicum | 2012 | 40 | 50 | 10 | [20] |
| Rhodopseudomonas palustris (Ch) | 2012 | 63.3 | 16.1 | 19.4 | [21] |
| Thermus thermophilus | 2012 | 65.3 | 26 | 8.7 | [22] |
| Thermus thermophilus | 2013 | 57 | 36 | 7 | [23] |
| Synechococcus sp. | 2013 | 43.9 | 42.44 | 13.66 | [24] |
| E. coli | 2013 | 75.9 | 16.7 | 7.4 | [25] |
| Staphylococcus aureus | 2014 | n.r. | n.r. | n.r. | [26] |
| Acinetobacter baumanii Abh12O-A2 | 2014 | 71.8 | 25.2 | 3.8 | [27] |
| Acinetobacter baumanii ATCC 17879 | 2014 | 68.9 | 24.1 | 5.2 | [27] |
| Pseudomonas aeruginosa | 2014 | 49 | 24 | 27 | [28] |
| Listeria monocytogenes | 2014 | 64 | 31 | 5 | [29] |
| Saccharopolyspora erythraea | 2014 | 47 | 45 | 8 | [30] |
| Bacillus subtilis | 2014 | 74.6 | 18.6 | 7.3 | [31] |
| Chlamydia caviae | 2015 | n.r. | n.r. | n.r. | [32] |
| Sinorhizobium meliloti | 2015 | 63 | 28 | 5 | [33] |
| E. coli | 2015 | n.r. | n.r. | n.r. | [34] |
| Bacillus subtilis | 2015 | n.r. | n.r. | 22.6 | [35] |
| Synechocystis sp. | 2015 | n.r. | n.r. | n.r. | [36] |
| Acinetobacter baumannii SK17-S | 2016 | 47 | 27.6 | 12.4 | [6] |
| Acinetobacter baumannii SK17-R | 2016 | 41.4 | 29.5 | 17.5 | [6] |
| Mycobacterium smegmatis | 2017 | 27.79 | 73.97 | 1.24 | [37] |
| Mycobacterium tuberculosis | 2017 | 68 | 29 | 3 | [38] |
| Microcystis aeruginosa | 2018 | n.r | n.r. | n.r. | [39] |
| Streptomyces coelicolor | 2018 | 50.6 | 47.4 | 2 | [40] |
| Zymomonas mobilis | 2019 | 73 | 21 | 6 | [41] |
| Streptococcus thermophilus | 2019 | 43 | 33 | 23 | [42] |
| Average | 55.9 | 34.1 | 9.9 |
| Bacterium | Year | Phosphoproteins | Phosphorylation Sites | Phosphoproteome | Reference |
|---|---|---|---|---|---|
| Neisseria meningitidis | 2011 | 51 | n.r. | Many biological processes | [16] |
| Staphylococcus aureus | 2014 | 103 | 76 | Pathogenicity and virulence | [26] |
| Chlamydia caviae (elementary body) | 2015 | 42 | n.r. | Virulence | [32] |
| Chlamydia caviae (reticulate body) | 2015 | 34 | n.r. | Virulence | [32] |
| Bacterium | Year | Phosphoproteins | Phosphorylation Sites | Phosphoproteome | Reference |
|---|---|---|---|---|---|
| Bacillus subtilis | 2007 | 78 | 78 | Carbohydrate metabolism | [8] |
| E. coli | 2008 | 79 | 81 | Similar to Bacillus | [9] |
| Lactococcus lactis | 2008 | 63 | 79 | Over-representation of phosphothreonines | [10] |
| Klebsiella pneumoniae | 2009 | 81 | 93 | Capsular biosynthesis | [7] |
| Pseudomonas aeruginosa | 2009 | 39 | 61 | Motility, transport and pathogenicity | [11] |
| Pseudomonas putida | 2009 | 59 | 55 | Several biochemical pathways | [11] |
| Halobacterium salinarum | 2009 | 26 | 31 | Phosphoproteome in Archaea | [12] |
| Mycobacterium tuberculosis | 2010 | 301 | 500 | Several biochemical pathways | [3] |
| Streptomyces coelicolor | 2010 | 40 | 46 | Housekeeping proteins | [13] |
| Streptococcus pneumoniae | 2010 | 84 | 163 | Carbon/protein/nucleotide metabolisms, cell cycle and division | [14] |
| Listeria monocytogenes | 2011 | 112 | 143 | Virulence, translation, carbohydrate metabolism and stress response | [18] |
| Helicobacter pylori | 2011 | 67 | 126 | Virulence | [19] |
| Clostridium acetobutylicum | 2012 | 61 | 107 | Carbon metabolism | [20] |
| Rhodopseudomonas palustris (Ch) | 2012 | 54 | 63 | Carbon metabolism | [21] |
| Rhodopseudomonas palustris (Ph) | 2012 | 42 | 59 | Carbon metabolism | [21] |
| Thermus thermophilus | 2012 | 48 | 46 | Wide variety of cellular processes | [22] |
| Thermus thermophilus | 2013 | 53 | 67 | Central metabolic pathways and protein/cell envelope biosynthesis | [23] |
| Synechococcus sp. | 2013 | 245 | 410 | Two-component signalling pathway and photosynthesis | [24] |
| Acinetobacter baumanii Abh12O-A2 | 2014 | 70 | 80 | Pathogenicity and drug resistance | [27] |
| Acinetobacter baumanii ATCC 17879 | 2014 | 41 | 48 | Several biochemical pathways | [27] |
| Pseudomonas aeruginosa | 2014 | 28 | 59 | Extracellular virulence factors | [28] |
| Sinorhizobium meliloti | 2015 | 77 | 96 | Rhizobial adaptation | [33] |
| Microcystis aeruginosa (nontoxic) | 2018 | 37 | n.r. | Several biochemical pathways | [39] |
| Microcystis aeruginosa (toxic) | 2018 | 18 | n.r. | Regulation of toxin generation | [39] |
| Bacterium | Year | Phosphoproteins | Phosphorylation Sites | Phosphoproteome | Method | Reference |
|---|---|---|---|---|---|---|
| Bacillus subtilis | 2010 | 27 | 45 | Phosphoproteome changes in different media | SILAC | [15] |
| Streptomyces coelicolor | 2011 | 127 | 289 | Sporulation factors, transcriptional regulators, protein kinases and other regulatory proteins | Label-free | [17] |
| E. coli | 2013 | 133 | 108 | Stationary phase | SILAC | [25] |
| Bacillus subtilis | 2014 | 141 | 177 | Stationary phase | SILAC | [31] |
| Listeria monocytogenes | 2014 | 191 | 242 | Purine biosynthesis regulated by PrfA phosphorylation | SILAC | [29] |
| Saccharopolyspora erythraea | 2014 | 88 | 109 | Carbon metabolism, environmental stress and protein synthesis affected by phosphorylation | SRM | [30] |
| E. coli | 2015 | 71 | n.r. | Phosphorylation varied during development | SRM | [34] |
| Bacillus subtilis | 2015 | 124 | 155 | Spore-specific determinants | Label-free | [35] |
| Synechocystis sp. | 2015 | 188 | 262 | Increased phosphorylation during nitrogen limitation | Dimethyl | [36] |
| Acinetobacter baumannii SK17-S | 2016 | 248 | 410 | Antibiotic resistance | Label-free | [6] |
| Acinetobacter baumannii SK17-R | 2016 | 211 | 285 | Antibiotic resistance | Label-free | [6] |
| Mycobacterium smegmatis | 2017 | 154 | 224 | Transmembrane proteins | Label-free | [37] |
| Mycobacterium tuberculosis | 2017 | 257 | 512 | Virulence | Tandem mass tag (TMT) | [38] |
| Streptomyces coelicolor | 2018 | 48 | 85 | Regulatory proteins | TMT | [40] |
| Zymomonas mobilis | 2019 | 125 | 177 | N2 fixing regulated by phosphorylation | Label-free | [41] |
| Streptococcus thermophilus | 2019 | 106 | 161 | Divisome proteins phosphorylated by the PknB kinase | Dimethyl | [42] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yagüe, P.; Gonzalez-Quiñonez, N.; Fernández-García, G.; Alonso-Fernández, S.; Manteca, A. Goals and Challenges in Bacterial Phosphoproteomics. Int. J. Mol. Sci. 2019, 20, 5678. https://doi.org/10.3390/ijms20225678
Yagüe P, Gonzalez-Quiñonez N, Fernández-García G, Alonso-Fernández S, Manteca A. Goals and Challenges in Bacterial Phosphoproteomics. International Journal of Molecular Sciences. 2019; 20(22):5678. https://doi.org/10.3390/ijms20225678
Chicago/Turabian StyleYagüe, Paula, Nathaly Gonzalez-Quiñonez, Gemma Fernández-García, Sergio Alonso-Fernández, and Angel Manteca. 2019. "Goals and Challenges in Bacterial Phosphoproteomics" International Journal of Molecular Sciences 20, no. 22: 5678. https://doi.org/10.3390/ijms20225678
APA StyleYagüe, P., Gonzalez-Quiñonez, N., Fernández-García, G., Alonso-Fernández, S., & Manteca, A. (2019). Goals and Challenges in Bacterial Phosphoproteomics. International Journal of Molecular Sciences, 20(22), 5678. https://doi.org/10.3390/ijms20225678





