CpWRKY71, a WRKY Transcription Factor Gene of Wintersweet (Chimonanthus praecox), Promotes Flowering and Leaf Senescence in Arabidopsis
Abstract
:1. Introduction
2. Results
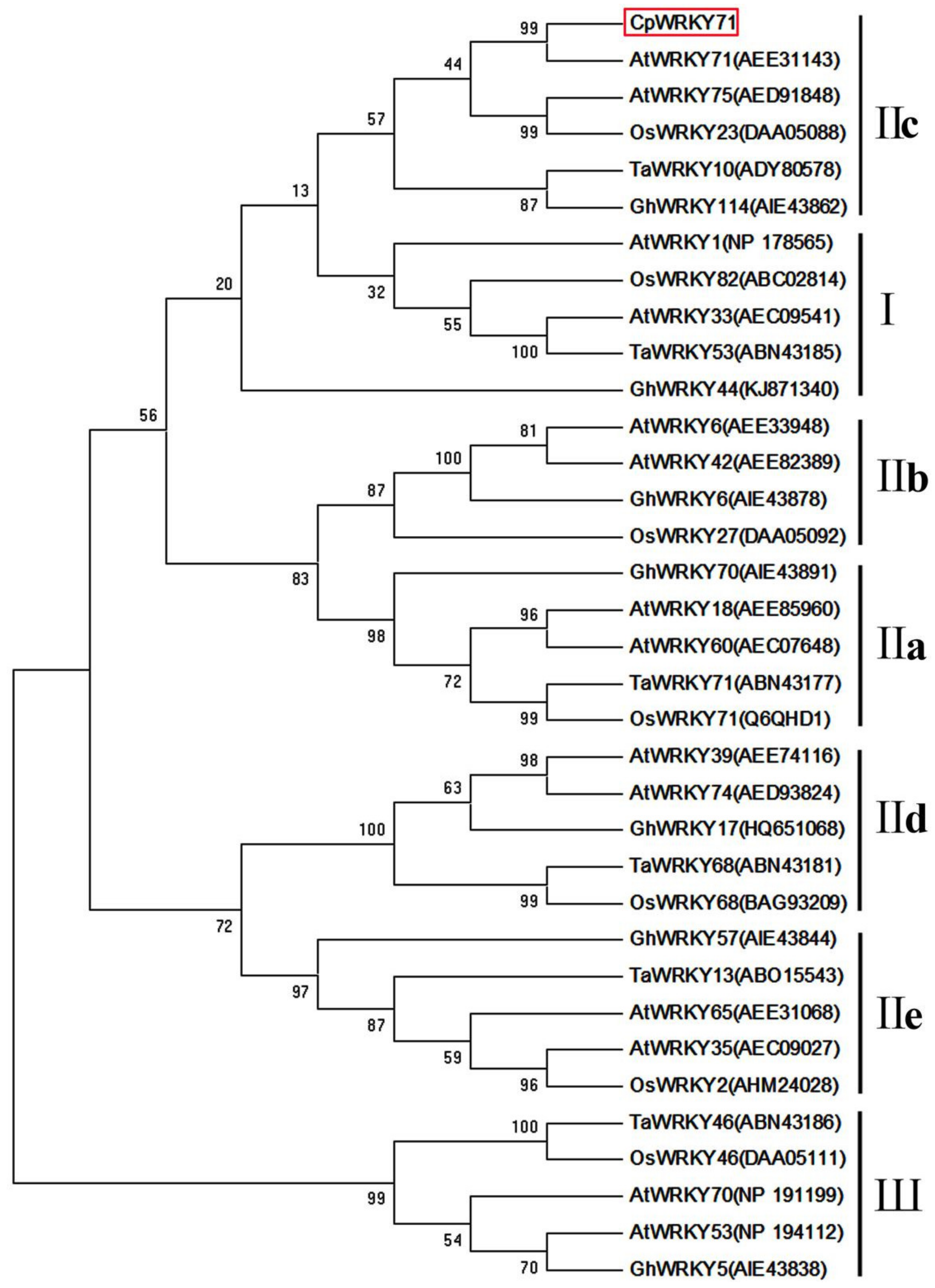
2.1. Isolation and Characterization of CpWRKY71
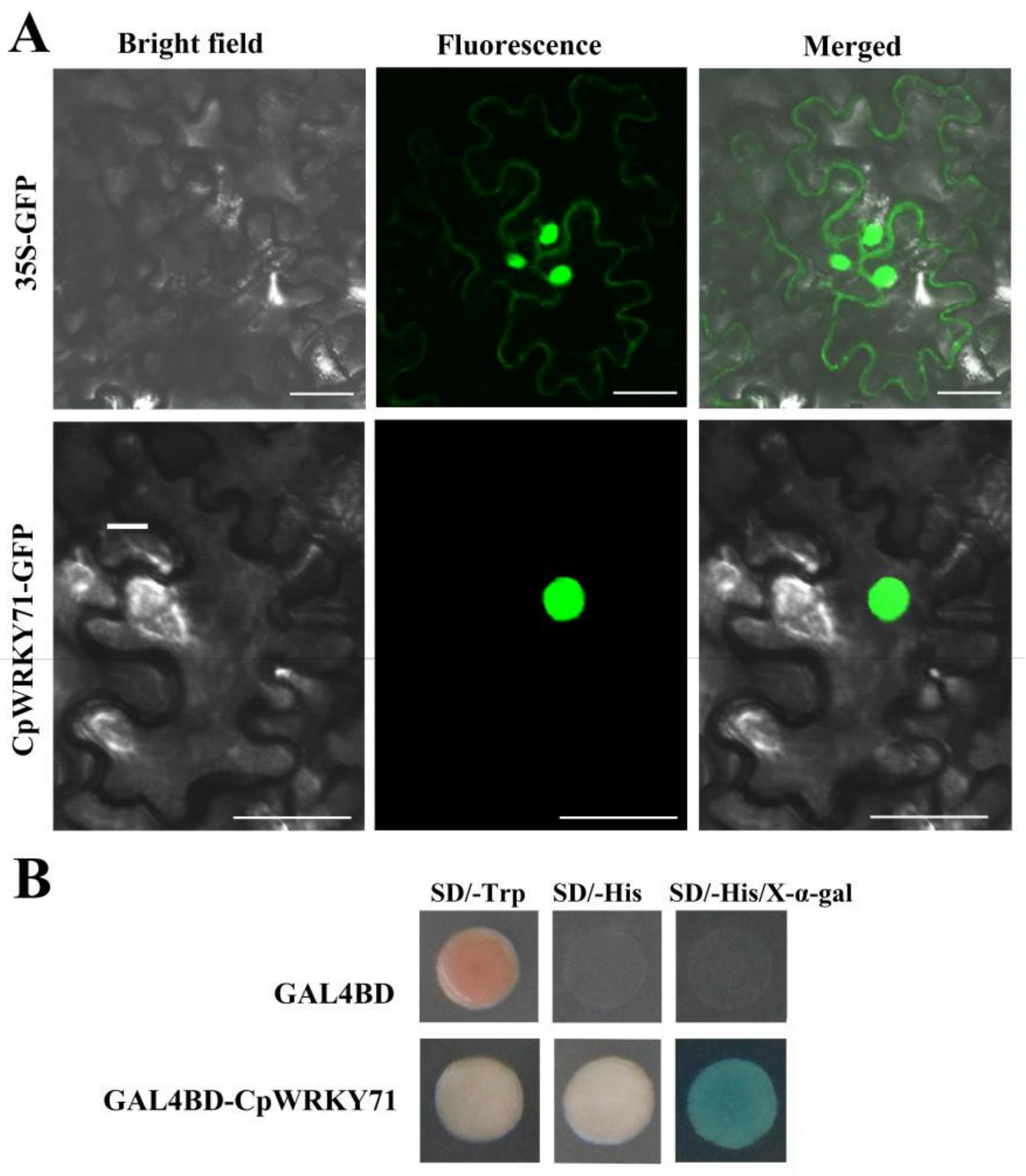
2.2. CpWRKY71 is a Nuclear Protein with Transcriptional Activation Activity in Yeast
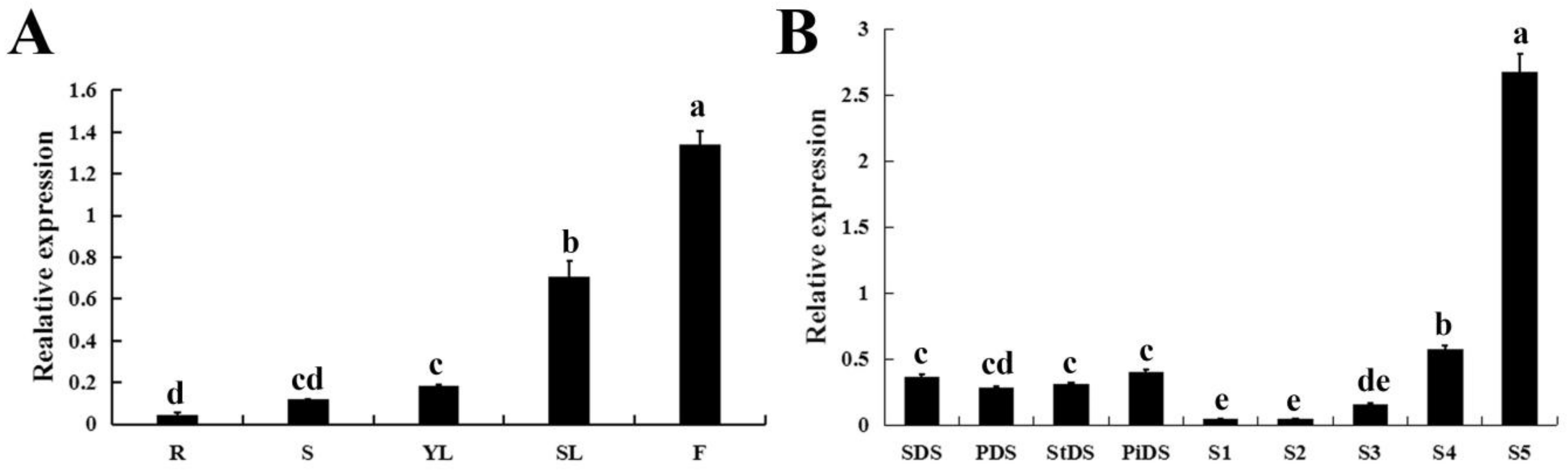
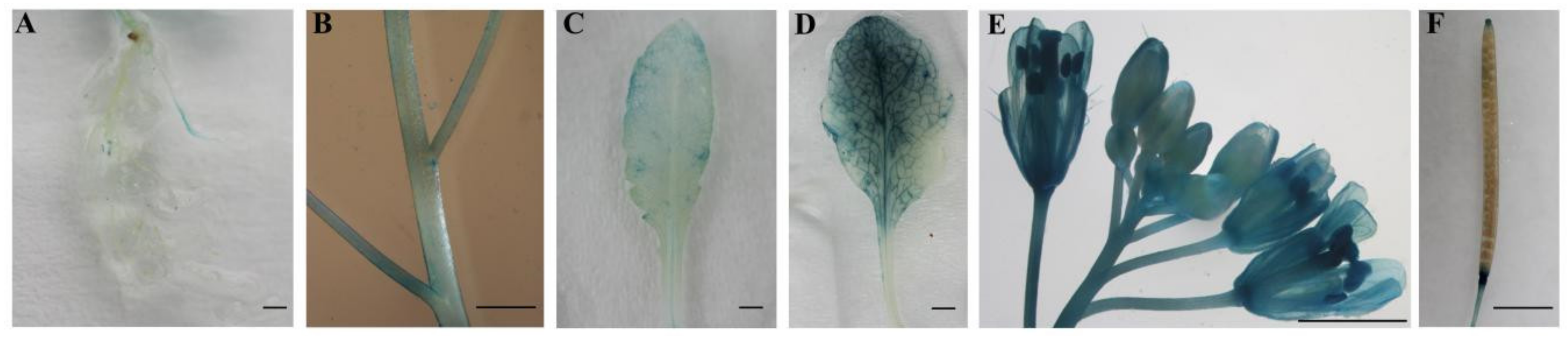
2.3. Tissue-Specific Expression of CpWRKY71
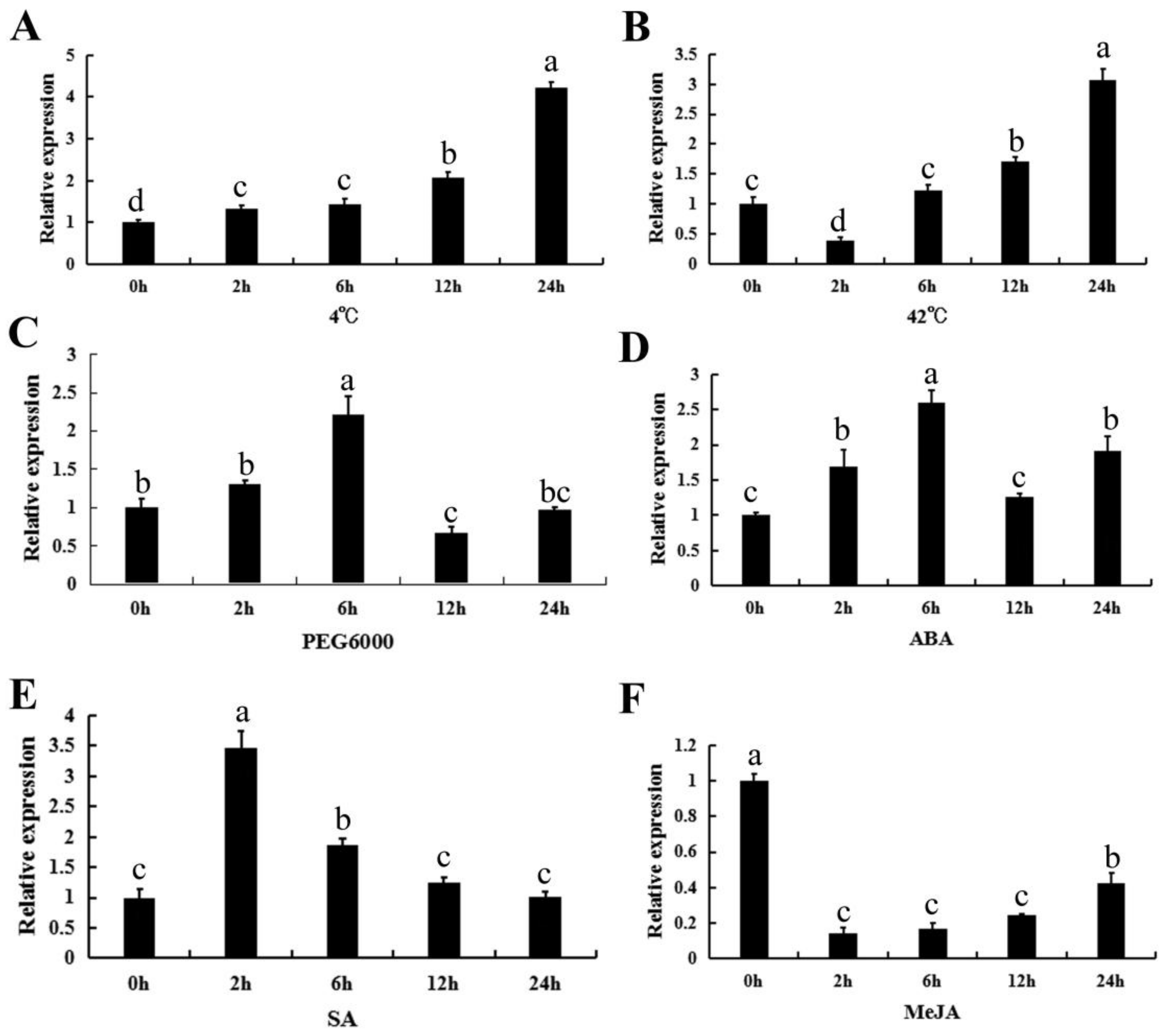
2.4. The Expression Profiles of CpWRKY71 under Abiotic Stress and Hormone Treatments
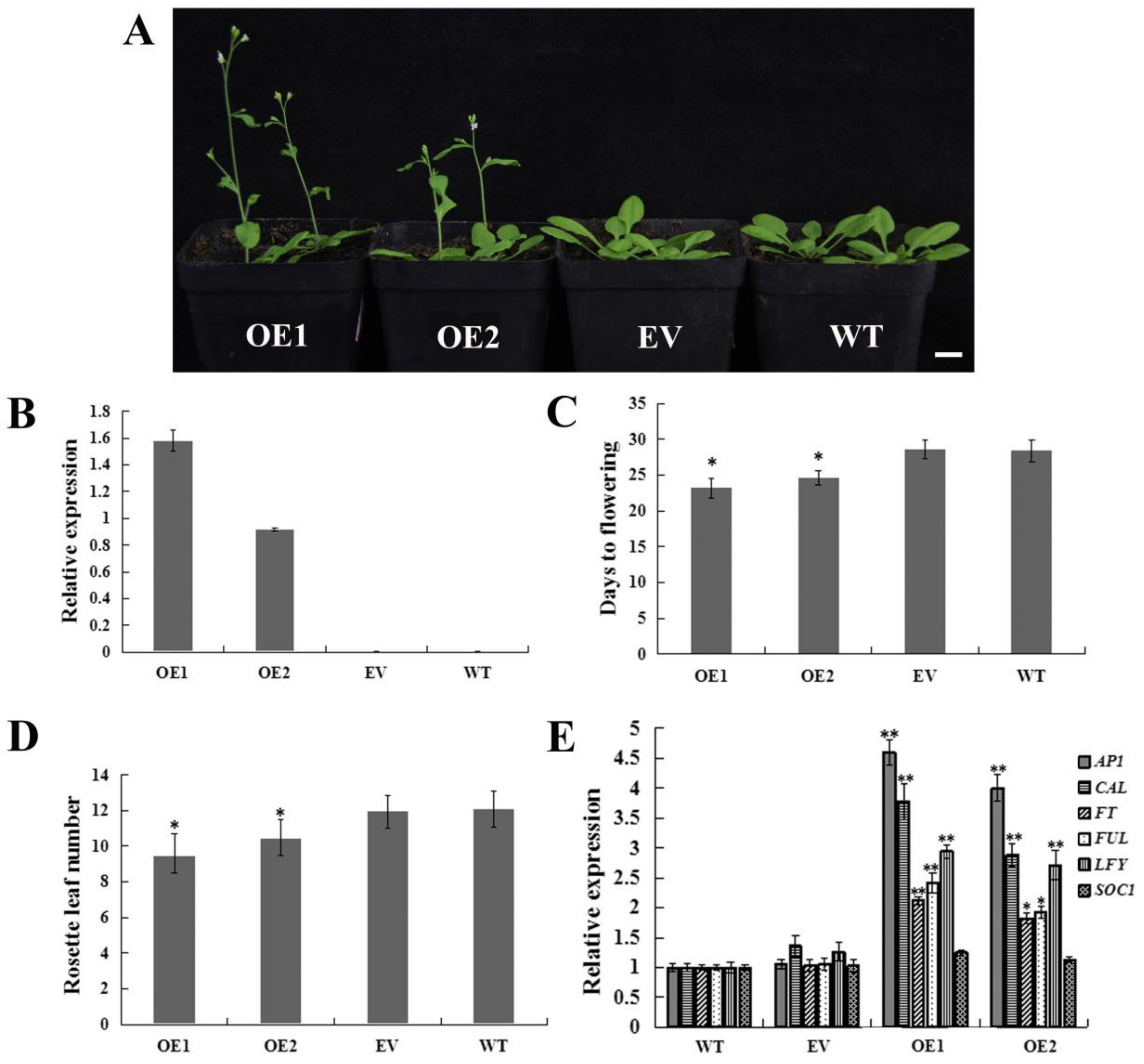
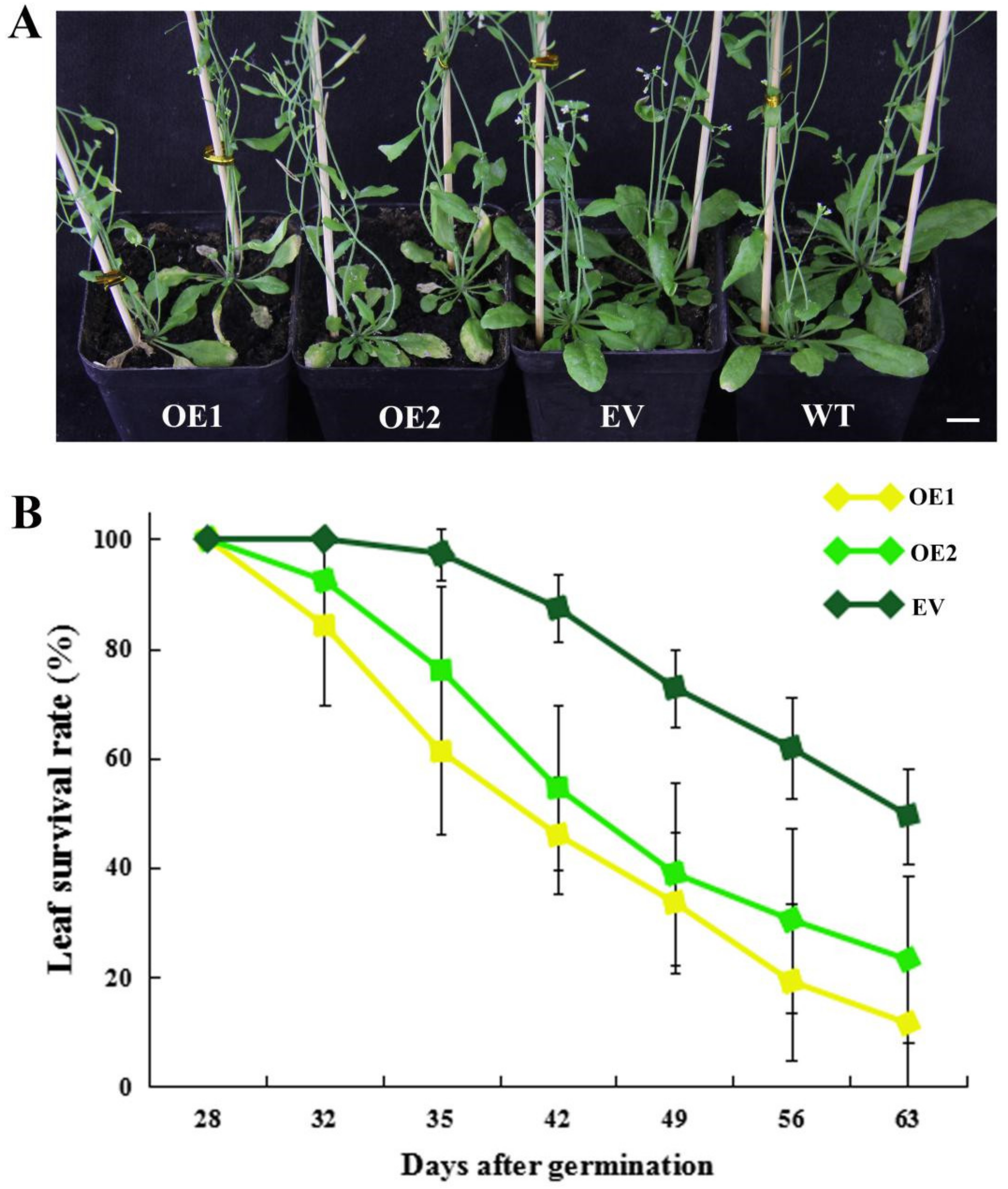
2.5. Heterologous Overexpression of CpWRKY71 in Arabidopsis Promoted Flowering Time and Leaf Senescence
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Gene Cloning
4.3. Plasmid Constructs
4.4. Subcellular Localization and Transactivation Activity Assay of CpWRKY71
4.5. Plant Treatment
4.6. Quantitative Real Time-PCR Analysis
4.7. Arabidopsis Transformation
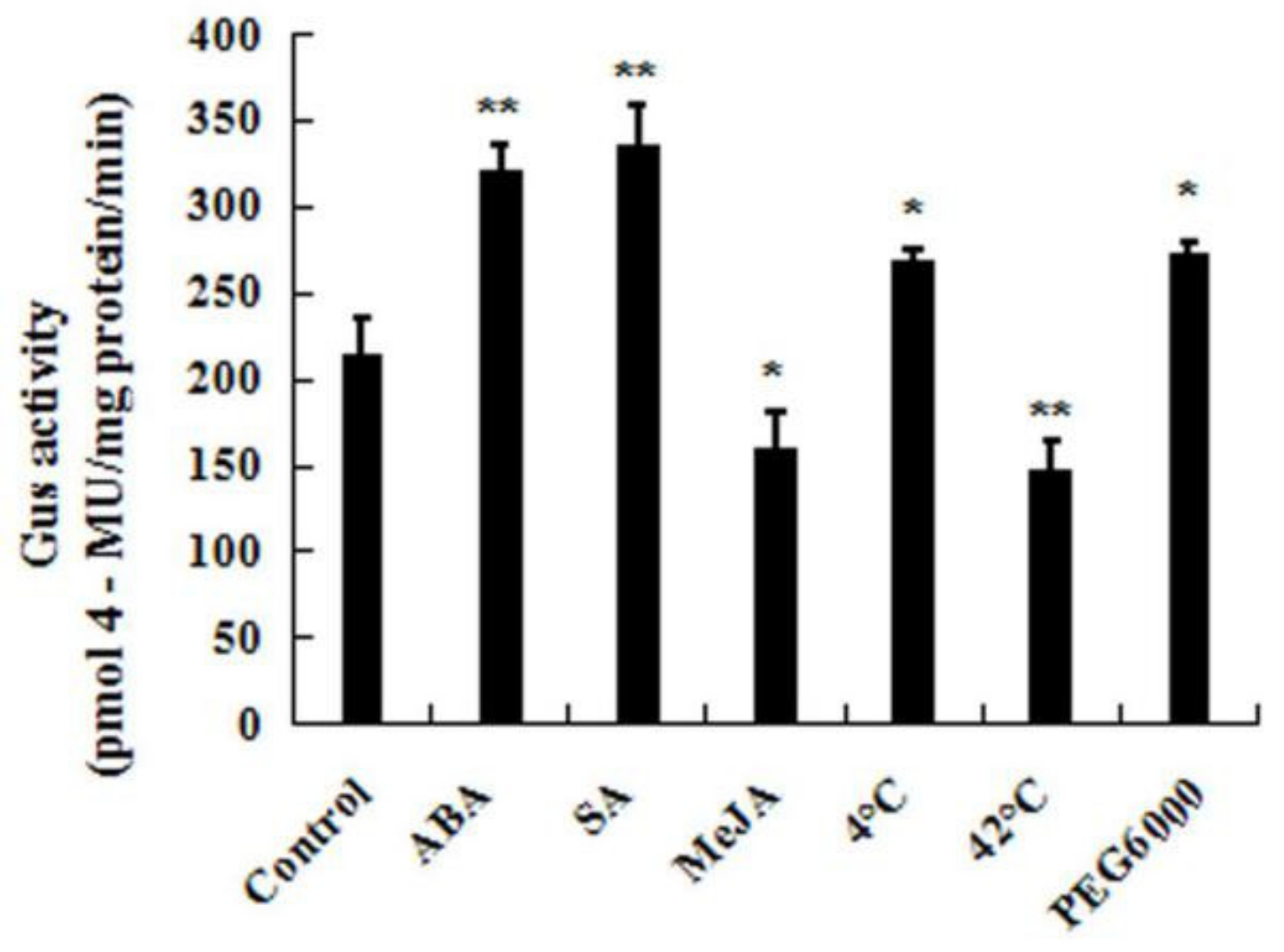
4.8. GUS Histochemical and GUS Activity Assays
4.9. Phenotype Analysis
4.10. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ABA | Abscisic acid |
| CTAB | Cetyltrimethylammonium Bromide |
| GUS | β-glucuronidase |
| MeJA | methyl jasmonate |
| PEG6000 | Polyethylene glycol 6000 |
| SA | Salicylic acid |
| qRT-PCR | Quantitative Real Time-PCR |
References
- Ma, J.; Li, Z.; Wang, B.; Sui, S.Z.; Li, M.Y. Cloning of an expansin gene from Chimonanthus praecox flowers and its expression in flowers treated with ethephon or 1-Methylcyclopropene. HortScience 2012, 47, 1472–1477. [Google Scholar] [CrossRef]
- Sui, S.Z.; Luo, J.H.; Ma, J.; Zhu, Q.L.; Lei, X.H.; Li, M.Y. Generation and analysis of expressed sequence tags from Chimonanthus praecox (Wintersweet) flowers for discovering stress-responsive and floral development-related genes. Comp. Funct. Genom. 2012, 2012, 134596. [Google Scholar] [CrossRef]
- Zhang, Q.; Wang, B.G.; Duan, K.; Wang, L.G.; Wang, M.; Tang, X.M.; Pan, A.H.; Sui, S.Z.; Wang, G.D. The paleoAP3-type gene CpAP3, an ancestral B-class gene from the basal angiosperm Chimonanthus praecox, can affect stamen and petal development in higher eudicots. Dev. Genes Evol. 2011, 221, 83–93. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.G.; Duan, K.; Zhang, Q.; Pan, A.H.; Sui, S.Z.; Li, M.Y.; Wang, L.G.; Tang, X.M. The AGL6-like gene CpAGL6, a potential regulator of floral time and organ identity in wintersweet (Chimonanthus praecox). Plant Growth Regul. 2011, 30, 343–352. [Google Scholar] [CrossRef]
- Liu, H.M.; Huang, R.W.; Ma, J.; Sui, S.Z.; Guo, Y.L.; Liu, D.F.; Li, Z.N.; Lin, Y.C.; Li, M.Y. Two C3H type zinc finger protein genes, CpCZF1 and CpCZF2, from Chimonanthus praecox affect Stamen development in Arabidopsis. Genes 2017, 8, 199. [Google Scholar] [CrossRef]
- Luo, J.H.; Ma, J.; Liu, D.F.; Yang, J.F.; Men, W.T.; Wan, C.; Sui, S.Z.; Li, M.Y. Effects of Ethylene on Cut Flower Opening and Senescence and Expression of Ethylene Receptor Genes in Wintersweet (Chimonanthus praecox). Plant Physiol. J. 2015, 51, 253–258. [Google Scholar]
- Sui, S.Z.; Luo, J.H.; Liu, D.F.; Ma, J.; Men, W.T.; Fan, L.; Bai, Y.; Li, M.Y. Effects of Hormone Treatments on Cut Flower Opening and Senescence in Wintersweet (Chimonanthus praecox). Hortscience 2015, 50, 1365–1369. [Google Scholar] [CrossRef]
- Peng, H.; Cheng, H.Y.; Chen, C.; Yu, X.W.; Yang, J.N.; Gao, W.R.; Shi, Q.H.; Zhang, H.; Li, J.G.; Ma, H. A NAC transcription factor gene of Chickpea (Cicer arietinum), CarNAC3, is involved in drought stress response and various developmental processes. J. Plant Physiol. 2009, 166, 1934–1945. [Google Scholar] [CrossRef]
- Yu, Y.C.; Hu, R.B.; Wang, H.M.; Cao, Y.P.; He, G.; Fu, C.X.; Zhou, G.K. MlWRKY12, a novel Miscanthus transcription factor, participates in pith secondary cell wall formation and promotes flowering. Plant Sci. 2013, 212, 1–9. [Google Scholar] [CrossRef]
- Zhang, J.Y.; Wang, Q.J.; Guo, Z.R. Progresses on plant AP2/ERF transcription factors. Hereditas 2012, 34, 835–847. [Google Scholar] [CrossRef]
- Eulgem, T.; Rushton, P.J.; Robatzek, S.; Somssich, I.E. The WRKY superfamily of plant transcription factors. Trends Plant Sci. 2000, 5, 199–206. [Google Scholar] [CrossRef]
- Ishiguro, S.; Nakamura, K. Characterization of a cDNA encoding a novel DNA-binding protein, SPF1, that recognizes SP8 sequences in the 50 upstream regions of genes coding for sporamin and beta-amylase from sweet potato. Mol. Gen. Genet. 1994, 244, 563–571. [Google Scholar] [CrossRef] [PubMed]
- Ulker, B.; Somssich, I.E. WRKY transcription factors: From DNA binding towards biological function. Curr. Opin. Plant Biol. 2004, 7, 491–498. [Google Scholar] [CrossRef] [PubMed]
- Wei, K.F.; Chen, J.; Chen, Y.F.; Wu, L.J.; Xie, D.X. Molecular phylogenetic and expression analysis of the complete WRKY transcription factor family in maize. DNA Res. 2012, 19, 153–164. [Google Scholar] [CrossRef] [PubMed]
- Pandey, S.P.; Somssich, I.E. The role of WRKY transcription factors in plant immunity. Plant Physiol. 2009, 150, 1648–1655. [Google Scholar] [CrossRef]
- Yin, G.; Xu, H.; Xiao, S.; Qin, Y.; Li, Y.; Yan, Y.; Hu, Y.K. The large soybean (Glycine max) WRKY TF family expanded by segmental duplication events and subsequent divergent selection among subgroups. BMC Plant Biol. 2013, 13, 148. [Google Scholar] [CrossRef]
- He, H.S.; Dong, Q.; Shao, Y.H.; Jiang, H.Y.; Zhu, S.W.; Cheng, B.J.; Xiang, Y. Genome-wide survey and characterization of the WRKY gene family in Populus trichocarpa. Plant Cell Rep. 2012, 31, 1199–1217. [Google Scholar] [CrossRef]
- Wang, L.; Zhu, W.; Fang, L.C.; Sun, X.M.; Su, L.Y.; Liang, Z.C.; Wang, N.; London, J.P.; Li, S.H.; Xin, H.P. Genome-wide identification of WRKY family genes and their response to cold stress in Vitis vinifera. BMC Plant Biol. 2014, 14, 103. [Google Scholar] [CrossRef]
- Ding, Z.J.; Yan, J.Y.; Li, X.G.; Wu, Z.C.; Zhang, S.Q.; Zheng, S.J. WRKY41 controls Arabidopsis seed dormancy via direct regulation of ABI3 transcript levels not downstream of ABA. Plant J. 2014, 79, 810–823. [Google Scholar] [CrossRef]
- Gu, L.J.; Li, L.B.; Wei, H.L.; Wang, H.T.; Su, J.J.; Guo, Y.N.; Yu, S.X. Identification of the group IIa WRKY subfamily and the functional analysis of GhWRKY17 in upland cotton (Gossypium hirsutum L.). PLoS ONE 2018, 13, e0191681. [Google Scholar] [CrossRef]
- Cai, Y.H.; Chen, X.J.; Xie, K.; Xing, Q.K.; Wu, Y.W.; Li, J.; Du, C.H.; Sun, Z.X. Dlf1, a WRKY transcription factor, is involved in the control of flowering time and plant height in rice. PLoS ONE 2014, 9, e102529. [Google Scholar] [CrossRef] [PubMed]
- Li, P.L.; Song, A.P.; Gao, C.Y.; Jiang, J.F.; Chen, S.M.; Fang, W.M.; Zhang, F.; Chen, F.D. The over-expression of a chrysanthemum WRKY transcription factor enhances aphid resistance. Plant Physiol. Biochem. 2015, 95, 26–34. [Google Scholar] [CrossRef] [PubMed]
- Qin, Y.X.; Tian, Y.C.; Liu, X.Z. A wheat salinity-induced WRKY transcription factor TaWRKY93 confers multiple abiotic stress tolerance in Arabidopsis thaliana. Biochem. Biophys. Res. Commun. 2015, 464, 428–433. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.F.; Sui, S.Z.; Ma, J.; Li, Z.N.; Guo, Y.L.; Luo, D.P.; Yang, J.F.; Li, M.Y. Transcriptomic analysis of flower development in wintersweet (Chimonanthus praecox). PLoS ONE 2014, 9, e86976. [Google Scholar] [CrossRef]
- Yu, Y.C.; Liu, Z.H.; Wang, L.; Kim, S.G.; Seo, P.J.; Qiao, M.; Wang, N.; Li, S.; Cao, X.F.; Park, C.M.; et al. WRKY71 accelerates flowering via the direct activation of FLOWERING LOCUS T and LEAFY in Arabidopsis thaliana. Plant J. 2016, 85, 96–106. [Google Scholar] [CrossRef]
- Zhang, L.P.; Chen, L.G.; Yu, D.Q. Transcription factor WRKY75 interacts with DELLA proteins to affect flowering. Plant Physiol. 2018, 176, 790–803. [Google Scholar] [CrossRef]
- Guo, P.K.; Li, Z.H.; Huang, P.X.; Li, B.S.; Shuang, F.; Chu, J.F.; Guo, H.W. A tripartite amplification loop involving the transcription factor WRKY75, salicylic acid, and reactive oxygen species accelerates leaf senescence. Plant Cell 2017, 29, 2854–2870. [Google Scholar] [CrossRef]
- Kiranmai, K.; Rao, G.L.; Pandurangaiah, M.; Nareshkumar, A.; Reddy, V.A.; Lokesh, U.; Venkatesh, B.; Johnson, A.M.A.; Sudhakar, C. A novel WRKY transcription factor, MuWRKY3 (Macrotyloma uniflorum Lam. Verdc.) enhances drought stress tolerance in transgenic groundnut (Arachis hypogaea L.) plants. Front. Plant Sci. 2017, 9, 346. [Google Scholar] [CrossRef]
- Yang, L.; Zhao, X.; Yang, F.; Fan, D.; Jiang, Y.Z.; Luo, K.M. PtrWRKY19, a novel WRKY transcription factor, contributes to the regulation of pith secondary wall formation in Populus trichocarpa. Sci. Rep. 2016, 6, 18643. [Google Scholar] [CrossRef]
- Gu, L.J.; Dou, L.L.; Guo, Y.N.; Wang, H.T.; Li, L.B.; Wang, C.C.; Ma, L.; Wei, H.L.; Yu, S.X. The WRKY transcription factor GhWRKY27 coordinates the senescence regulatory pathway in upland cotton (Gossypium hirsutum L.). BMC Plant Biol. 2019, 19, 116. [Google Scholar] [CrossRef]
- Johnson, C.S.; Kolevski, B.; Smyth, D.R. TRANSPARENT TESTA GLABRA2, a trichome and seed coat development gene of Arabidopsis, encodes a WRKY transcription factor. Plant Cell 2002, 14, 1359–1375. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Wang, H.P.; Yu, D.Q. Arabidopsis WRKY transcription factors WRKY12 and WRKY13 oppositely regulate flowering under Short-day conditions. Mol. Plant 2016, 9, 1492–1503. [Google Scholar] [CrossRef] [PubMed]
- Miao, Y.; Laun, T.; Zimmermann, P.; Zentgraf, U. Targets of the WRKY53 transcription factor and its role during leaf senescence in Arabidopsis. Plant Mol. Biol. 2004, 55, 853–867. [Google Scholar] [CrossRef] [PubMed]
- Robatzek, S.; Somssich, I.E. Targets of AtWRKY6 regulation during plant senescence and pathogen defense. Genes Dev. 2002, 16, 1139–1149. [Google Scholar] [CrossRef] [PubMed]
- Rishmawi, L.; Pesch, M.; Juengst, C.; Schauss, A.C.; Schrader, A.; Hulskamp, M. Non-cell-autonomous regulation of root hair patterning genes by WRKY75 in Arabidopsis. Plant Physiol. 2014, 165, 186–195. [Google Scholar] [CrossRef] [PubMed]
- Encinas-Villarejo, S.; Maldonado, A.M.; Amil-Ruiz, F.; de los Santos, B.; Romero, F.; Pliego-Alfaro, F.; Mucloz-Blanco, J.; Caballero, J.L. Evidence for a positive regulatory role of strawberry (Fragaria x ananassa) FaWRKY1 and Arabidopsis AtWRKY75 proteins in resistance. J. Exp. Bot 2009, 60, 3043–3065. [Google Scholar] [CrossRef]
- Zhang, C.Q.; Xu, Y.; Lu, Y.; Yu, H.X.; Gu, M.H.; Liu, Q.Q. The WRKY transcription factor OsWRKY78 regulates stem elongation and seed development in rice. Planta 2011, 234, 541–554. [Google Scholar] [CrossRef]
- Besseau, S.; Li, J.; Palva, E.T. WRKY54 and WRKY70 co-operate as negative regulators of leaf senescence in Arabidopsis thaliana. J. Exp. Bot. 2012, 63, 2667–2679. [Google Scholar] [CrossRef]
- Chen, L.G.; Xiang, S.Y.; Chen, Y.L.; Li, D.B.; Yu, D.Q. Arabidopsis WRKY45 interacts with the DELLA protein RGL1 to positively regulate age-triggered leaf senescence. Mol. Plant. 2017, 10, 1174–1189. [Google Scholar] [CrossRef]
- Wu, A.H.; Allu, A.D.; Garapati, P.; Siddiqui, H.; Dortay, H.; Zanor, M.; Asensi-Fabado, M.A.; Munné-Bosch, S.; Antoni, C.; Tohge, T.; et al. JUNGBRUNNEN1, a reactive oxygen species-responsive NAC transcription factor, regulates longevity in Arabidopsis. Plant Cell 2012, 24, 482–506. [Google Scholar] [CrossRef]
- Zhang, K.W.; Halitschke, R.; Yin, C.X.; Liu, C.J.; Gan, S.S. Salicylic acid 3-hydroxylase regulates Arabidopsis leaf longevity by mediating salicylic acid catabolism. Proc. Natl. Acad. Sci. USA 2013, 110, 14807–14812. [Google Scholar] [CrossRef] [PubMed]
- Ren, T.T.; Wang, J.W.; Zhao, M.M.; Gong, X.M.; Wang, S.X.; Wang, G.; Zhou, C.J. Involvement of NAC transcription factor SiNAC1 in a positive feedback loop via ABA biosynthesis and leaf senescence in foxtail millet. Planta 2018, 247, 53–68. [Google Scholar] [CrossRef] [PubMed]
- Siegel, C.S.; Stevenson, F.O.; Zimmer, E.A. Evaluation and comparison of FTA card and CTAB DNA extraction methods for non-agricultural taxa. Appl. Plant Sci. 2017, 5, 1600109. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Qin, R.Y.; Xu, R.F.; Li, H.; Yang, Y.C.; Li, L.; Wei, P.C.; Yang, J.B. Isolation and identifcation of five cold-inducible promoters from Oryza sativa. Planta 2018, 247, 99–111. [Google Scholar] [CrossRef]
- Ma, J.; Men, W.T.; Chen, X.L.; Sui, S.Z.; Li, M.Y. Cloning and expression analysis of a small GTP-binding protein gene (CpRAC1) in Chimonanthus praecox. Acta Hortic. Sin. 2018, 45, 2177–2187. [Google Scholar]
- Yan, H.R.; Jia, H.H.; Chen, X.B.; Hao, L.L.; An, H.L.; Guo, X.Q. The cotton WRKY transcription factor GhWRKY17 functions in drought and salt stress in transgenic Nicotiana benthamiana through ABA signaling and the modulation of reactive oxygen species production. Plant Cell Physiol. 2014, 55, 2060–2076. [Google Scholar] [CrossRef]
- Li, J.; Wang, J.; Wang, N.X.; Guo, X.Q.; Gao, Z. GhWRKY44, a WRKY transcription factor of cotton, mediates defense responses to pathogen infection in transgenic Nicotiana benthamiana. Plant Cell Tissue Org. 2015, 121, 127–140. [Google Scholar] [CrossRef]
- Wang, F.Q.; Yang, J.; Dai, C.; Wu, M.Z.; Zhang, Y.H.; Shen, W.B. Characterization of the Arabidopsis thaliana heme oxygenase 1 promoter in response to salinity, iron deficiency, and mercury exposure. Biol. Plant. 2017, 61, 35–47. [Google Scholar] [CrossRef]
- Niu, G.L.; Gou, W.; Han, X.L.; Qin, C.; Zhang, L.X.; Abomohra, A.E.; Ashraf, M. Cloning and functional analysis of phosphoethanolamine methyltransferase promoter from Maize (Zea mays L.). Int. J. Mol. Sci. 2018, 19, 191. [Google Scholar] [CrossRef]
- Khan, M.A.; Meng, Y.L.; Liu, D.F.; Tang, H.S.; Lü, S.H.; Imtiaz, M.; Jiang, G.M.; Lü, P.T.; Ji, Y.Q.; Gao, J.P.; et al. Responses of rose RhACS1 and RhACS2 promoters to abiotic stresses in transgenic Arabidopsis thaliana. Plant Cell Rep. 2015, 34, 795–804. [Google Scholar] [CrossRef]
- Ma, J.; Li, Z.; Chen, X.L.; Zhang, S.Y.; Sui, S.Z.; Li, M.Y. Cloning and activity analysis of CpEXP1 gene promoter from Chimonanthus praecox. Sci. Silvae Sin. 2018, 54, 61–72. [Google Scholar]
- Tao, Y.; Wang, F.T.; Jia, D.M.; Li, J.T.; Zhang, Y.M.; Jia, C.G.; Wang, D.P.; Pan, H.Y. Cloning and Functional Analysis of the Promoter of a Stress-inducible Gene (ZmRXO1) in Maize. Plant Mol. Biol. Rep. 2015, 33, 200–208. [Google Scholar] [CrossRef]
- Clough, S.J.; Ben, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, R.; Liu, D.; Huang, M.; Ma, J.; Li, Z.; Li, M.; Sui, S. CpWRKY71, a WRKY Transcription Factor Gene of Wintersweet (Chimonanthus praecox), Promotes Flowering and Leaf Senescence in Arabidopsis. Int. J. Mol. Sci. 2019, 20, 5325. https://doi.org/10.3390/ijms20215325
Huang R, Liu D, Huang M, Ma J, Li Z, Li M, Sui S. CpWRKY71, a WRKY Transcription Factor Gene of Wintersweet (Chimonanthus praecox), Promotes Flowering and Leaf Senescence in Arabidopsis. International Journal of Molecular Sciences. 2019; 20(21):5325. https://doi.org/10.3390/ijms20215325
Chicago/Turabian StyleHuang, Renwei, Daofeng Liu, Min Huang, Jing Ma, Zhineng Li, Mingyang Li, and Shunzhao Sui. 2019. "CpWRKY71, a WRKY Transcription Factor Gene of Wintersweet (Chimonanthus praecox), Promotes Flowering and Leaf Senescence in Arabidopsis" International Journal of Molecular Sciences 20, no. 21: 5325. https://doi.org/10.3390/ijms20215325
APA StyleHuang, R., Liu, D., Huang, M., Ma, J., Li, Z., Li, M., & Sui, S. (2019). CpWRKY71, a WRKY Transcription Factor Gene of Wintersweet (Chimonanthus praecox), Promotes Flowering and Leaf Senescence in Arabidopsis. International Journal of Molecular Sciences, 20(21), 5325. https://doi.org/10.3390/ijms20215325





