Association with PD-L1 Expression and Clinicopathological Features in 1000 Lung Cancers: A Large Single-Institution Study of Surgically Resected Lung Cancers with a High Prevalence of EGFR Mutation
Abstract
1. Introduction
2. Results
2.1. Clinicopathological Characteristics of 1000 Surgically Resected Lung Cancers
2.2. PD-L1 Expression in Lung Cancer
2.3. Association between PD-L1 Expression Status and Clinicopathological Features in Lung Adenocarcinomas and Squamous Cell Carcinomas
2.4. Association between PD-L1 Expression and Driver Genetic Status in Lung Adenocarcinoma
2.5. Association between PD-L1 Expression Status and Clinicopathological Features in 424 EGFR-Mutated Lung Adenocarcinomas
2.6. Multivariate Logistic Regression Analysis of Clinicopathological Features for PD-L1 Expression in Lung Adenocarcinoma
3. Discussion
4. Materials and Methods
4.1. Patients
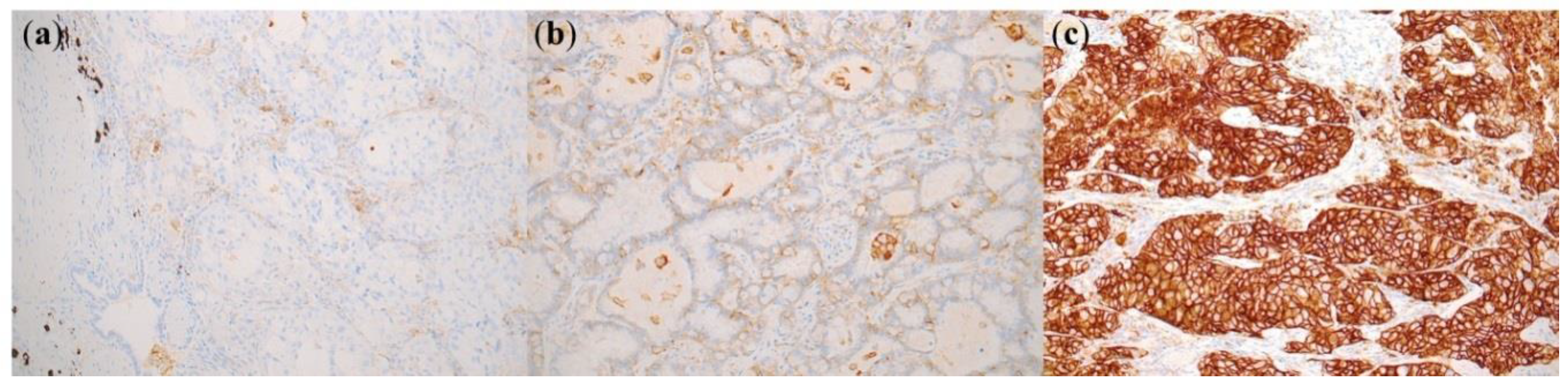
4.2. PD-L1 Immunohistochemistry
4.3. Analysis of EGFR Mutation
4.4. Analysis of ALK Fusion
4.5. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| PD-1 | Programmed cell death 1 |
| PD-L1 | Programmed cell death ligand 1 |
| NSCLC | Non-small cell lung cancer |
Appendix A
| Adenocarcinoma (n = 785) | |||||
|---|---|---|---|---|---|
| PD-L1 Expression | |||||
| Variable | Total Patients No. | <1% (n = 495, 63.1%) | ≥1% (n = 290, 36.9%) | p-Value | |
| Age | 0.063 | ||||
| <65 | 440 (56.1%) | 290 (65.9%) | 150 (34.1%) | ||
| ≥65 | 345 (43.9%) | 205 (59.4%) | 140 (40.6%) | ||
| Sex | <0.001 | ||||
| Male | 377 (48%) | 213 (56.5%) | 164 (43.5%) | ||
| Female | 408 (52%) | 282 (69.1%) | 126 (30.9%) | ||
| Smoking status | <0.001 | ||||
| Never smoker | 438 (55.8%) | 308 (70.3%) | 130 (29.7%) | ||
| Current or Ex smoker | 347 (44.2%) | 187 (53.9%) | 160 (46.1%) | ||
| T_stage | <0.001 | ||||
| 1 | 519 (66.1%) | 354 (68.2%) | 165 (31.8%) | ||
| 2 | 178 (22.7%) | 96 (53.9%) | 82 (46.1%) | ||
| 3 | 68 (8.7%) | 36 (52.9%) | 32 (47.1%) | ||
| 4 | 18 (2.3%) | 8 (44.4%) | 10 (55.6%) | ||
| N_stage | <0.001 | ||||
| 0 | 629 (82.1%) | 427 (67.9%) | 202 (32.1%) | ||
| 1 | 37 (4.8%) | 15 (40.5%) | 22 (59.5%) | ||
| 2 | 100 (13.1%) | 41 (41.0%) | 59 (59.0%) | ||
| M_stage | 0.261 | ||||
| 0 | 752 (96.2%) | 477 (63.4%) | 275 (36.6%) | ||
| 1 | 30 (3.8%) | 16 (53.3%) | 14 (46.7%) | ||
| AJCC stage | <0.001 | ||||
| 1 | 544 (70.3%) | 377 (69.3%) | 167 (30.7%) | ||
| 2 | 99 (12.8%) | 56 (56.6%) | 43 (43.4%) | ||
| 3 | 103 (13.3%) | 41 (39.8%) | 62 (60.2%) | ||
| 4 | 28 (3.6%) | 14 (50.0%) | 14 (50.0%) | ||
| Preop Tx. | 0.15 | ||||
| No | 739 (94.1%) | 471 (63.7%) | 268 (36.3%) | ||
| PreopCCRT | 39 (5.0%) | 19 (48.7%) | 20 (51.3%) | ||
| PreopChemo | 7 (0.9%) | 5 (71.4%) | 2 (28.6%) | ||
| Vascular invasion | 0.001 | ||||
| No | 738 (95.3%) | 475 (64.4%) | 263 (35.6%) | ||
| YES | 36 (4.7%) | 13 (36.1%) | 23 (63.9%) | ||
| Lymphatic invasion | <0.001 | ||||
| No | 608 (78.6%) | 413 (67.9%) | 195 (32.1%) | ||
| YES | 166 (21.4%) | 75 (45.2%) | 91 (54.8%) | ||
| Differentiation | <0.001 | ||||
| W/D | 41 (5.2%) | 37 (90.2%) | 4 (9.8%) | ||
| M/D | 561 (71.5%) | 385 (68.6%) | 176 (31.4%) | ||
| P/D | 147 (18.7%) | 47 (32.0%) | 100 (68.0%) | ||
| Predominant patttern | <0.001 | ||||
| Lepidic | 47 (6.1%) | 40 (85.1%) | 7 (14.9%) | ||
| Acinar | 402 (52.5%) | 283 (70.4%) | 119 (29.6%) | ||
| Papillary | 108 (14.1%) | 69 (63.9%) | 39 (36.1%) | ||
| Micropapillary | 49 (6.4%) | 23 (46.9%) | 26 (53.1%) | ||
| Solid | 90 (11.8%) | 22 (24.4%) | 68 (75.6%) | ||
| Cribriform | 21 (2.7%) | 10 (47.6%) | 11 (52.4%) | ||
| Mucinous | 40 (5.2%) | 33 (82.5%) | 7 (17.5%) | ||
| Pleomorphic | 8 (1.0%) | 1 (12.5%) | 7 (87.5%) | ||
| Squamous Cell Carcinoma (n = 188) | |||||
|---|---|---|---|---|---|
| PD-L1 Expression | |||||
| Variable | Total Patients No. | <1% (n = 52, 27.7%) | ≥1% (n = 136, 72.3%) | p-Value | |
| Age | 0.83 | ||||
| <65 | 71 (37.8%) | 19 (26.8%) | 52 (73.2%) | ||
| ≥65 | 117 (62.2%) | 33 (28.2%) | 84 (71.8%) | ||
| Sex | 0.357 | ||||
| Male | 174 (92.6%) | 50 (28.7%) | 124 (71.3%) | ||
| Female | 14 (7.4%) | 2 (14.3%) | 12 (85.7%) | ||
| Smoking status | 0.756 | ||||
| Never smoker | 13 (6.9%) | 4 (30.8%) | 9 (69.2%) | ||
| Current or Ex smoker | 175 (93.1%) | 48 (27.4%) | 127 (72.6%) | ||
| T_stage | 0.786 | ||||
| 1 | 73 (39.0%) | 20 (27.4%) | 53 (72.6%) | ||
| 2 | 54 (28.9%) | 17 (31.5%) | 37 (68.5%) | ||
| 3 | 45 (24.1%) | 11 (24.4%) | 34 (75.6%) | ||
| 4 | 15 (8.0%) | 3 (20.0%) | 12 (80.0%) | ||
| N_stage | 0.43 | ||||
| 0 | 125 (68.3%) | 32 (25.6%) | 93 (74.4%) | ||
| 1 | 33 (18%) | 7 (21.2%) | 26 (78.8%) | ||
| 2 | 25 (13.7%) | 9 (36.0%) | 16 (64.0%) | ||
| M_stage | 0.062 | ||||
| 0 | 183 (97.9%) | 48 (26.2%) | 135 (73.8%) | ||
| 1 | 4 (2.1%) | 3 (75.0%) | 1 (25.0%) | ||
| AJCC stage | 0.066 | ||||
| 1 | 77 (41.8%) | 23 (29.9%) | 54 (70.1%) | ||
| 2 | 68 (37%) | 14 (20.6%) | 54 (79.4%) | ||
| 3 | 37 (20.1%) | 10 (27.0%) | 27 (73.0%) | ||
| 4 | 2 (1.1%) | 2 (100%) | 0 | ||
| Preop Tx. | 0.29 | ||||
| No | 169 (89.9%) | 45 (26.6%) | 124 (73.4%) | ||
| PreopCCRT | 16 (8.5%) | 5 (31.3%) | 11 (68.8%) | ||
| PreopChemo | 3 (1.6%) | 2 (66.7%) | 1 (33.3%) | ||
| Vascular invasion | 0.342 | ||||
| No | 172 (93%) | 45 (26.2%) | 127 (73.8%) | ||
| YES | 13 (7%) | 5 (38.5%) | 8 (61.5%) | ||
| Lymphatic invasion | 0.05 | ||||
| No | 144 (77.8%) | 34 (23.6%) | 110 (76.4%) | ||
| YES | 41 (22.2%) | 16 (39.0%) | 25 (61.0%) | ||
| Differentiation | 0.016 | ||||
| W/D | 9 (4.8%) | 2 (22.2%) | 7 (77.8%) | ||
| M/D | 129 (68.6%) | 29 (22.5%) | 100 (77.5%) | ||
| P/D | 41 (21.8%) | 15 (36.6%) | 26 (63.4%) | ||
| Adenocarcinoma (n = 785) | ||||||
|---|---|---|---|---|---|---|
| PD-L1 | ||||||
| Variable | Total Patients No. | <1% (n = 495, 63.1%) | ≥1% (n = 290, 36.9%) | p-Value | ||
| EGFR | <0.001 | |||||
| Wild | 361 (46.0%) | 195 (54.0%) | 166 (46.0%) | |||
| Mutant | 424 (54.0%) | 300 (70.8%) | 124 (29.2%) | |||
| Exon19del | 182 (23.2%) | 119 (65.4%) | 63 (34.6%) | |||
| L858R | 202 (25.7%) | 153 (75.7%) | 49 (24.3%) | |||
| Exon20insertion | 17 (2.2%) | 11 (64.7%) | 6 (35.3%) | |||
| Others | 23 (2.9%) | 17 (73.9%) | 6 (26.1%) | |||
| ALK | <0.001 | |||||
| Wild | 758 (96.6%) | 490 (64.6%) | 268 (35.4%) | |||
| Mutant | 27 (3.4%) | 5 (18.5%) | 22 (81.5%) | |||
| EGFR-Mutated Adenocarcinoma (n = 424) | |||||
|---|---|---|---|---|---|
| PD-L1 Expression | |||||
| Variable | Total Patients No. | <1% (n = 300, 70.8%) | ≥1% (n = 124, 29.2%) | P-Value | |
| Age | 0.099 | ||||
| <65 | 255 (60.1%) | 188 (73.7%) | 67 (26.3%) | ||
| ≥65 | 169 (39.9%) | 112 (66.3%) | 57 (33.7%) | ||
| Sex | 0.078 | ||||
| Male | 157 (37.0%) | 103 (65.6%) | 54 (34.4%) | ||
| Female | 267 (63.0%) | 197 (73.8%) | 70 (26.2%) | ||
| Smoking status | 0.01 | ||||
| Never smoker | 279 (65.8%) | 209 (74.9%) | 70 (25.1%) | ||
| Current or Ex smoker | 145 (34.2%) | 91 (62.8%) | 54 (37.2%) | ||
| T_stage | 0.001 | ||||
| 1 | 292 (69.2%) | 223 (76.4%) | 69 (23.6%) | ||
| 2 | 96 (22.7%) | 53 (55.2%) | 43 (44.8%) | ||
| 3 | 30 (7.1%) | 20 (66.7%) | 10 (33.3%) | ||
| 4 | 4 (0.9%) | 3 (75.0%) | 1 (25.0%) | ||
| N_stage | <0.001 | ||||
| 0 | 341 (83.2%) | 256 (75.1%) | 85 (24.9%) | ||
| 1 | 18 (4.4%) | 11 (61.1%) | 7 (38.9%) | ||
| 2 | 51 (12.4%) | 25 (49.0%) | 26 (51.0%) | ||
| M_stage | 0.089 | ||||
| 0 | 406 (96.2%) | 291 (71.7%) | 115 (28.3%) | ||
| 1 | 16 (3.8%) | 8 (50.0%) | 8 (50.0%) | ||
| AJCC stage | <0.001 | ||||
| 1 | 315 (75.7%) | 238 (75.6%) | 77 (24.4%) | ||
| 2 | 37 (8.9%) | 26 (70.3%) | 11 (29.7%) | ||
| 3 | 50 (12.0%) | 25 (50.0%) | 25 (50.0%) | ||
| 4 | 14 (3.4%) | 6 (42.9%) | 8 (57.1%) | ||
| Preop Tx. | 0.091 | ||||
| No | 407 (96.0%) | 284 (69.8%) | 123 (30.2%) | ||
| PreopCCRT | 12 (2.8%) | 11 (91.7%) | 1 (8.3%) | ||
| PreopChemo | 5 (1.2%) | 5 (100%) | 0 | ||
| Vascular invasion | 0.058 | ||||
| No | 398 (96.1%) | 286 (71.9%) | 112 (28.1%) | ||
| YES | 16 (3.9%) | 8 (50.0%) | 8 (50.0%) | ||
| Lymphatic invasion | <0.001 | ||||
| No | 323 (78.0%) | 250 (77.4%) | 73 (22.6%) | ||
| YES | 91 (22.0%) | 44 (48.4%) | 47 (51.6%) | ||
| Differentiation | <0.001 | ||||
| W/D | 20 (4.7%) | 18 (90.0%) | 2 (10.0%) | ||
| M/D | 340 (80.2%) | 251 (73.8%) | 89 (26.2%) | ||
| P/D | 51 (12.0%) | 20 (39.2%) | 31 (60.8%) | ||
| Predominant pattern | <0.001 | ||||
| Lepidic | 26 (6.3%) | 23 (88.5%) | 3 (11.5%) | ||
| Acinar | 273 (65.8%) | 205 (75.1%) | 68 (24.9%) | ||
| Papillary | 58 (14.0%) | 38 (65.5%) | 20 (34.5%) | ||
| Micropapillary | 27 (6.5%) | 13 (48.1%) | 14 (51.9%) | ||
| Solid | 21 (5.1%) | 7 (33.3%) | 14 (66.7%) | ||
| Cribriform | 6 (1.4%) | 3 (50.0%) | 3 (50.0%) | ||
| Mucinous | 4 (1.0%) | 4 (100.0%) | 0 | ||
| EGFR | 0.146 | ||||
| Exon19del | 182 (42.9%) | 119 (65.4%) | 63 (34.6%) | ||
| L858R | 202 (47.6%) | 153 (75.7%) | 49 (24.3%) | ||
| Exon20insertion | 17 (4.0%) | 11 (64.7%) | 6 (35.3%) | ||
| Others | 23 (5.4%) | 17 (73.9%) | 6 (26.1%) | ||
References
- Fehrenbacher, L.; Spira, A.; Ballinger, M.; Kowanetz, M.; Vansteenkiste, J.; Mazieres, J.; Park, K.; Smith, D.; Artal-Cortes, A.; Lewanski, C.; et al. Atezolizumab versus docetaxel for patients with previously treated non-small-cell lung cancer (POPLAR): A multicentre, open-label, phase 2 randomised controlled trial. Lancet 2016, 387, 1837–1846. [Google Scholar] [CrossRef]
- Herbst, R.S.; Baas, P.; Kim, D.W.; Felip, E.; Perez-Gracia, J.L.; Han, J.Y.; Molina, J.; Kim, J.H.; Arvis, C.D.; Ahn, M.J.; et al. Pembrolizumab versus docetaxel for previously treated, PD-L1-positive, advanced non-small-cell lung cancer (KEYNOTE-010): A randomised controlled trial. Lancet 2016, 387, 1540–1550. [Google Scholar] [CrossRef]
- Brahmer, J.; Reckamp, K.L.; Baas, P.; Crino, L.; Eberhardt, W.E.; Poddubskaya, E.; Antonia, S.; Pluzanski, A.; Vokes, E.E.; Holgado, E.; et al. Nivolumab versus Docetaxel in Advanced Squamous-Cell Non-Small-Cell Lung Cancer. N. Engl. J. Med. 2015, 373, 123–135. [Google Scholar] [CrossRef]
- Cooper, W.A.; Tran, T.; Vilain, R.E.; Madore, J.; Selinger, C.I.; Kohonen-Corish, M.; Yip, P.; Yu, B.; O’Toole, S.A.; McCaughan, B.C.; et al. PD-L1 expression is a favorable prognostic factor in early stage non-small cell carcinoma. Lung Cancer 2015, 89, 181–188. [Google Scholar] [CrossRef]
- D’Incecco, A.; Andreozzi, M.; Ludovini, V.; Rossi, E.; Capodanno, A.; Landi, L.; Tibaldi, C.; Minuti, G.; Salvini, J.; Coppi, E.; et al. PD-1 and PD-L1 expression in molecularly selected non-small-cell lung cancer patients. Br. J. Cancer 2015, 112, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Inamura, K.; Yokouchi, Y.; Sakakibara, R.; Kobayashi, M.; Subat, S.; Ninomiya, H.; Nagano, H.; Nomura, K.; Okumura, S.; Ishikawa, Y. Relationship of tumor PD-L1 expression with EGFR wild-type status and poor prognosis in lung adenocarcinoma. Jpn. J. Clin. Oncol. 2016, 46, 935–941. [Google Scholar] [CrossRef]
- Inoue, Y.; Yoshimura, K.; Mori, K.; Kurabe, N.; Kahyo, T.; Mori, H.; Kawase, A.; Tanahashi, M.; Ogawa, H.; Inui, N.; et al. Clinical significance of PD-L1 and PD-L2 copy number gains in non-small-cell lung cancer. Oncotarget 2016, 7, 32113–32128. [Google Scholar] [CrossRef]
- Kim, H.; Kwon, H.J.; Park, S.Y.; Park, Y.; Park, E.; Chung, J.H. Clinicopathological analysis and prognostic significance of programmed cell death-ligand 1 protein and mRNA expression in non-small cell lung cancer. PLoS ONE 2018, 13, e0198634. [Google Scholar] [CrossRef]
- Koh, J.; Go, H.; Keam, B.; Kim, M.Y.; Nam, S.J.; Kim, T.M.; Lee, S.H.; Min, H.S.; Kim, Y.T.; Kim, D.W.; et al. Clinicopathologic analysis of programmed cell death-1 and programmed cell death-ligand 1 and 2 expressions in pulmonary adenocarcinoma: Comparison with histology and driver oncogenic alteration status. Mod. Pathol. 2015, 28, 1154–1166. [Google Scholar] [CrossRef]
- Ota, K.; Azuma, K.; Kawahara, A.; Hattori, S.; Iwama, E.; Tanizaki, J.; Harada, T.; Matsumoto, K.; Takayama, K.; Takamori, S.; et al. Induction of PD-L1 Expression by the EML4-ALK Oncoprotein and Downstream Signaling Pathways in Non-Small Cell Lung Cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2015, 21, 4014–4021. [Google Scholar] [CrossRef]
- Rangachari, D.; VanderLaan, P.A.; Shea, M.; Le, X.; Huberman, M.S.; Kobayashi, S.S.; Costa, D.B. Correlation between Classic Driver Oncogene Mutations in EGFR, ALK, or ROS1 and 22C3-PD-L1 >/=50% Expression in Lung Adenocarcinoma. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 2017, 12, 878–883. [Google Scholar] [CrossRef]
- Song, Z.; Yu, X.; Cheng, G.; Zhang, Y. Programmed death-ligand 1 expression associated with molecular characteristics in surgically resected lung adenocarcinoma. J. Transl. Med. 2016, 14, 188. [Google Scholar] [CrossRef]
- Yang, C.Y.; Lin, M.W.; Chang, Y.L.; Wu, C.T.; Yang, P.C. Programmed cell death-ligand 1 expression in surgically resected stage I pulmonary adenocarcinoma and its correlation with driver mutations and clinical outcomes. Eur. J. Cancer 2014, 50, 1361–1369. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, L.; Li, Y.; Pan, Y.; Wang, R.; Hu, H.; Li, H.; Luo, X.; Ye, T.; Sun, Y.; et al. Protein expression of programmed death 1 ligand 1 and ligand 2 independently predict poor prognosis in surgically resected lung adenocarcinoma. Oncotargets Ther. 2014, 7, 567–573. [Google Scholar] [CrossRef]
- Lan, B.; Ma, C.; Zhang, C.; Chai, S.; Wang, P.; Ding, L.; Wang, K. Association between PD-L1 expression and driver gene status in non-small-cell lung cancer: A meta-analysis. Oncotarget 2018, 9, 7684–7699. [Google Scholar] [CrossRef]
- Sun, J.M.; Zhou, W.; Choi, Y.L.; Choi, S.J.; Kim, S.E.; Wang, Z.; Dolled-Filhart, M.; Emancipator, K.; Wu, D.; Weiner, R.; et al. Prognostic Significance of PD-L1 in Patients with Non-Small Cell Lung Cancer: A Large Cohort Study of Surgically Resected Cases. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 2016, 11, 1003–1011. [Google Scholar] [CrossRef]
- Jiang, L.; Su, X.; Zhang, T.; Yin, X.; Zhang, M.; Fu, H.; Han, H.; Sun, Y.; Dong, L.; Qian, J.; et al. PD-L1 expression and its relationship with oncogenic drivers in non-small cell lung cancer (NSCLC). Oncotarget 2017, 8, 26845–26857. [Google Scholar] [CrossRef]
- Pawelczyk, K.; Piotrowska, A.; Ciesielska, U.; Jablonska, K.; Gletzel-Plucinska, N.; Grzegrzolka, J.; Podhorska-Okolow, M.; Dziegiel, P.; Nowinska, K. Role of PD-L1 Expression in Non-Small Cell Lung Cancer and Their Prognostic Significance according to Clinicopathological Factors and Diagnostic Markers. Int. J. Mol. Sci. 2019, 20, 824. [Google Scholar] [CrossRef]
- Shimoji, M.; Shimizu, S.; Sato, K.; Suda, K.; Kobayashi, Y.; Tomizawa, K.; Takemoto, T.; Mitsudomi, T. Clinical and pathologic features of lung cancer expressing programmed cell death ligand 1 (PD-L1). Lung Cancer 2016, 98, 69–75. [Google Scholar] [CrossRef]
- Dong, Z.Y.; Zhang, C.; Li, Y.F.; Su, J.; Xie, Z.; Liu, S.Y.; Yan, L.X.; Chen, Z.H.; Yang, X.N.; Lin, J.T.; et al. Genetic and Immune Profiles of Solid Predominant Lung Adenocarcinoma Reveal Potential Immunotherapeutic Strategies. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 2018, 13, 85–96. [Google Scholar] [CrossRef]
- Yoshida, A.; Kohno, T.; Tsuta, K.; Wakai, S.; Arai, Y.; Shimada, Y.; Asamura, H.; Furuta, K.; Shibata, T.; Tsuda, H. ROS1-rearranged lung cancer: A clinicopathologic and molecular study of 15 surgical cases. Am. J. Surg. Pathol. 2013, 37, 554–562. [Google Scholar] [CrossRef]
- Takeuchi, K.; Soda, M.; Togashi, Y.; Suzuki, R.; Sakata, S.; Hatano, S.; Asaka, R.; Hamanaka, W.; Ninomiya, H.; Uehara, H.; et al. RET, ROS1 and ALK fusions in lung cancer. Nat. Med. 2012, 18, 378–381. [Google Scholar] [CrossRef]
- Lee, S.E.; Lee, B.; Hong, M.; Song, J.Y.; Jung, K.; Lira, M.E.; Mao, M.; Han, J.; Kim, J.; Choi, Y.L. Comprehensive analysis of RET and ROS1 rearrangement in lung adenocarcinoma. Mod. Pathol. 2015, 28, 468–479. [Google Scholar] [CrossRef]
- Cho, J.H.; Zhou, W.; Choi, Y.L.; Sun, J.M.; Choi, H.; Kim, T.E.; Dolled-Filhart, M.; Emancipator, K.; Rutkowski, M.A.; Kim, J. Retrospective Molecular Epidemiology Study of PD-L1 Expression in Patients with EGFR-Mutant Non-small Cell Lung Cancer. Cancer Res. Treat. Off. J. Korean Cancer Assoc. 2018, 50, 95–102. [Google Scholar] [CrossRef]
- Song, P.; Wu, S.; Zhang, L.; Zeng, X.; Wang, J. Correlation Between PD-L1 Expression and Clinicopathologic Features in 404 Patients with Lung Adenocarcinoma. Interdiscip. Sci. Comput. Life Sci. 2019, 11, 258–265. [Google Scholar] [CrossRef]
- Lynch, T.J.; Bell, D.W.; Sordella, R.; Gurubhagavatula, S.; Okimoto, R.A.; Brannigan, B.W.; Harris, P.L.; Haserlat, S.M.; Supko, J.G.; Haluska, F.G.; et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N. Engl. J. Med. 2004, 350, 2129–2139. [Google Scholar] [CrossRef]
- Fukui, T.; Otani, S.; Hataishi, R.; Jiang, S.X.; Nishii, Y.; Igawa, S.; Mitsufuji, H.; Kubota, M.; Katagiri, M.; Masuda, N. Successful rechallenge with erlotinib in a patient with EGFR-mutant lung adenocarcinoma who developed gefitinib-related interstitial lung disease. Cancer Chemother. Pharmacol. 2010, 65, 803–806. [Google Scholar] [CrossRef]
- Sun, P.L.; Seol, H.; Lee, H.J.; Yoo, S.B.; Kim, H.; Xu, X.; Jheon, S.; Lee, C.T.; Lee, J.S.; Chung, J.H. High incidence of EGFR mutations in Korean men smokers with no intratumoral heterogeneity of lung adenocarcinomas: Correlation with histologic subtypes, EGFR/TTF-1 expressions, and clinical features. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 2012, 7, 323–330. [Google Scholar] [CrossRef]
- Gainor, J.F.; Shaw, A.T.; Sequist, L.V.; Fu, X.; Azzoli, C.G.; Piotrowska, Z.; Huynh, T.G.; Zhao, L.; Fulton, L.; Schultz, K.R.; et al. EGFR Mutations and ALK Rearrangements Are Associated with Low Response Rates to PD-1 Pathway Blockade in Non-Small Cell Lung Cancer: A Retrospective Analysis. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2016, 22, 4585–4593. [Google Scholar] [CrossRef]
- Huynh, T.G.; Morales-Oyarvide, V.; Campo, M.J.; Gainor, J.F.; Bozkurtlar, E.; Uruga, H.; Zhao, L.; Gomez-Caraballo, M.; Hata, A.N.; Mark, E.J.; et al. Programmed Cell Death Ligand 1 Expression in Resected Lung Adenocarcinomas: Association with Immune Microenvironment. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 2016, 11, 1869–1878. [Google Scholar] [CrossRef]
- Kim, H.; Chung, J.H. PD-L1 Testing in Non-small Cell Lung Cancer: Past, Present, and Future. J. Pathol. Transl. Med. 2019, 53, 199–206. [Google Scholar] [CrossRef]
- McLaughlin, J.; Han, G.; Schalper, K.A.; Carvajal-Hausdorf, D.; Pelekanou, V.; Rehman, J.; Velcheti, V.; Herbst, R.; LoRusso, P.; Rimm, D.L. Quantitative Assessment of the Heterogeneity of PD-L1 Expression in Non-Small-Cell Lung Cancer. JAMA Oncol. 2016, 2, 46–54. [Google Scholar] [CrossRef]
- Ilie, M.; Long-Mira, E.; Bence, C.; Butori, C.; Lassalle, S.; Bouhlel, L.; Fazzalari, L.; Zahaf, K.; Lalvee, S.; Washetine, K.; et al. Comparative study of the PD-L1 status between surgically resected specimens and matched biopsies of NSCLC patients reveal major discordances: A potential issue for anti-PD-L1 therapeutic strategies. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2016, 27, 147–153. [Google Scholar] [CrossRef]
- Zhang, Y.H.; Zhao, Y.; Rajapaksa, U.S.; Lawrence, T.M.; Peng, Y.C.; Liu, J.; Xu, K.; Hu, K.; Qin, L.; Liu, N.; et al. A Comprehensive Analysis of the Impact of HIV on HCV Immune Responses and Its Association with Liver Disease Progression in a Unique Plasma Donor Cohort. PLoS ONE 2016, 11, e0158037. [Google Scholar] [CrossRef]
- Jia, X.; Zhang, L.; Wu, W.; Zhang, W.; Wu, C. Driver Mutation Analysis and PD-L1 Expression in Synchronous Double Primary Lung Cancer. Appl. Immunohistochem. Mol. Morphol. 2018, 26, 246–253. [Google Scholar] [CrossRef]
- Garon, E.B.; Rizvi, N.A.; Hui, R.; Leighl, N.; Balmanoukian, A.S.; Eder, J.P.; Patnaik, A.; Aggarwal, C.; Gubens, M.; Horn, L.; et al. Pembrolizumab for the treatment of non-small-cell lung cancer. N. Engl. J. Med. 2015, 372, 2018–2028. [Google Scholar] [CrossRef]
- Roach, C.; Zhang, N.; Corigliano, E.; Jansson, M.; Toland, G.; Ponto, G.; Dolled-Filhart, M.; Emancipator, K.; Stanforth, D.; Kulangara, K. Development of a Companion Diagnostic PD-L1 Immunohistochemistry Assay for Pembrolizumab Therapy in Non-Small-cell Lung Cancer. Appl. Immunohistochem. Mol. Morphol. 2016, 24, 392–397. [Google Scholar] [CrossRef]
- Park, H.Y.; Oh, I.J.; Kho, B.G.; Kim, T.O.; Shin, H.J.; Park, C.K.; Kwon, Y.S.; Kim, Y.I.; Lim, S.C.; Kim, Y.C.; et al. Clinical Characteristics of Korean Patients with Lung Cancer Who Have Programmed Death-Ligand 1 Expression. Tuberc. Respir. Dis. 2019, 82, 227–233. [Google Scholar] [CrossRef]
- Kim, H.J.; Lee, K.Y.; Kim, Y.C.; Kim, K.S.; Lee, S.Y.; Jang, T.W.; Lee, M.K.; Shin, K.C.; Lee, G.H.; Lee, J.C.; et al. Detection and comparison of peptide nucleic acid-mediated real-time polymerase chain reaction clamping and direct gene sequencing for epidermal growth factor receptor mutations in patients with non-small cell lung cancer. Lung Cancer 2012, 75, 321–325. [Google Scholar] [CrossRef]
- Lee, B.; Han, G.; Kwon, M.J.; Han, J.; Choi, Y.L. KRAS Mutation Detection in Non-small Cell Lung Cancer Using a Peptide Nucleic Acid-Mediated Polymerase Chain Reaction Clamping Method and Comparative Validation with Next-Generation Sequencing. Korean J. Pathol. 2014, 48, 100–107. [Google Scholar] [CrossRef]
| Adenocarcinoma (n = 785) | ||||||
|---|---|---|---|---|---|---|
| PD-L1 Expression | ||||||
| Variable | Total Patients No. | <1% (n = 495, 63.1%) | 1–49% (n = 177, 22.5%) | ≥50% (n = 113, 14.4%) | p-Value | |
| Age | 0.173 | |||||
| <65 year | 440 (56.1%) | 290 (65.9%) | 91 (20.7%) | 59 (13.4%) | ||
| ≥65 year | 345 (43.9%) | 205 (59.4%) | 86 (24.9%) | 54 (15.7%) | ||
| Sex | <0.001 | |||||
| Male | 377 (48%) | 213 (56.5%) | 92 (24.4%) | 72 (19.1%) | ||
| Female | 408 (52%) | 282 (69.1%) | 85 (20.8%) | 41 (10.0%) | ||
| Smoking status | <0.001 | |||||
| Never smoker | 438 (55.8%) | 308 (70.3%) | 87 (19.9%) | 43 (9.8%) | ||
| Current or Ex smoker | 347 (44.2%) | 187 (53.9%) | 90 (25.9%) | 70 (20.2%) | ||
| T_stage | 0.001 | |||||
| 1 | 519 (66.3%) | 354 (68.2%) | 109 (21.0%) | 56 (10.8%) | ||
| 2 | 178 (22.7%) | 96 (53.9%) | 48 (27.0%) | 34 (19.1%) | ||
| 3 | 68 (8.7%) | 36 (52.9%) | 15 (22.1%) | 17 (25.0%) | ||
| 4 | 18 (2.3%) | 8 (44.4%) | 4 (22.2%) | 6 (33.3%) | ||
| N_stage | <0.001 | |||||
| 0 | 629 (82.1%) | 427 (67.9%) | 125 (19.9%) | 77 (12.2%) | ||
| 1 | 37 (4.8%) | 15 (40.5%) | 15 (40.5%) | 7 (18.9%) | ||
| 2 | 100 (13.1%) | 41 (41.0%) | 31 (31.0%) | 28 (28.0%) | ||
| M_stage | 0.049 | |||||
| 0 | 752 (96.2%) | 477 (63.4%) | 164 (21.8%) | 111 (14.8%) | ||
| 1 | 30 (3.8%) | 16 (53.3%) | 12 (40.0%) | 2 (6.7%) | ||
| AJCC stage | <0.001 | |||||
| 1 | 544 (70.3%) | 377 (69.3%) | 108 (19.9%) | 59 (10.8%) | ||
| 2 | 99 (12.8%) | 56 (56.6%) | 23 (23.2%) | 20 (20.2%) | ||
| 3 | 103 (13.3%) | 41 (39.8%) | 31 (30.1%) | 31 (30.1%) | ||
| 4 | 28 (3.6%) | 14 (50.0%) | 12 (42.9%) | 2 (7.1%) | ||
| Preop Tx. | 0.011 | |||||
| No | 739 (94.1%) | 471 (63.7%) | 168 (22.7%) | 100 (13.5%) | ||
| PreopCCRT | 39 (5.0%) | 19 (48.7%) | 7 (17.9%) | 13 (33.3%) | ||
| PreopChemo | 7 (0.9%) | 5 (71.4%) | 2 (28.6%) | 0 | ||
| Vascular invasion | 0.002 | |||||
| No | 738 (95.3%) | 475 (64.4%) | 161 (21.8%) | 102 (13.8%) | ||
| YES | 36 (4.7%) | 13 (36.1%) | 12 (33.3%) | 11 (30.6%) | ||
| Lymphatic invasion | <0.001 | |||||
| No | 608 (78.6%) | 413 (67.9%) | 120 (19.7%) | 75 (12.3%) | ||
| YES | 166 (21.4%) | 75 (45.2%) | 53 (31.9%) | 38 (22.9%) | ||
| Differentiation | <0.001 | |||||
| W/D | 41 (5.2%) | 37 (90.2%) | 4 (9.8%) | 0 | ||
| M/D | 561 (71.5%) | 385 (68.6%) | 126 (22.5%) | 50 (8.9%) | ||
| P/D | 147 (18.7%) | 47 (32.0%) | 40 (27.2%) | 60 (40.8%) | ||
| Predominant pattern | <0.001 | |||||
| Lepidic | 47 (6.1%) | 40 (85.1%) | 7 (14.9%) | 0 | ||
| Acinar | 402 (52.5%) | 283 (70.4%) | 84 (20.9%) | 35 (8.7%) | ||
| Papillary | 108 (14.1%) | 69 (63.9%) | 29 (26.9%) | 10 (9.3%) | ||
| Micropapillary | 49 (6.4%) | 23 (46.9%) | 17 (34.7%) | 9 (18.4%) | ||
| Solid | 90 (11.8%) | 22 (24.4%) | 22 (24.4%) | 46 (51.1%) | ||
| Cribriform | 21 (2.7%) | 10 (47.6%) | 6 (28.6%) | 5 (23.8%) | ||
| Mucinous | 40 (5.2%) | 33 (82.5%) | 7 (17.5%) | 0 | ||
| Pleomorphic | 8 (1.0%) | 1 (12.5%) | 0 | 7 (87.5%) | ||
| Squamous Cell Carcinoma (n = 188) | ||||||
|---|---|---|---|---|---|---|
| PD-L1 Expression | ||||||
| Variable | Total Patients No. | <1% (n = 52, 27.7%) | 1–49% (n = 50, 26.6%) | ≥50% (n = 86, 45.7%) | p-Value | |
| Age | 0.927 | |||||
| <65 | 71 (37.8%) | 19 (26.8%) | 20 (28.2%) | 32 (45.1%) | ||
| ≥65 | 117 (62.2%) | 33 (28.2%) | 30 (25.6%) | 54 (46.2%) | ||
| Sex | ||||||
| Male | 174 (92.6%) | 50 (28.7%) | 47 (27.0%) | 77 (44.3%) | ||
| Female | 14 (7.4%) | 2 (14.3%) | 3 (21.4%) | 9 (64.3%) | ||
| Smoking status | 0.944 | |||||
| Never smoker | 13 (6.9%) | 4 (30.8%) | 3 (23.1%) | 6 (46.2%) | ||
| Current or Ex smoker | 175 (93.1%) | 48 (27.4%) | 47 (26.9%) | 80 (45.7%) | ||
| T_stage | 0.698 | |||||
| 1 | 73 (38.9%) | 20 (27.4%) | 20 (27.4%) | 33 (45.2%) | ||
| 2 | 54 (28.7%) | 17 (31.5%) | 14 (25.9%) | 23 (42.6%) | ||
| 3 | 45 (23.9%) | 11 (24.4%) | 10 (22.2%) | 24 (53.3%) | ||
| 4 | 15 (8.0%) | 3 (20.0%) | 6 (40.0%) | 6 (40.0%) | ||
| N_stage | 0.449 | |||||
| 0 | 125 (68.3%) | 32 (25.6%) | 38 (30.4%) | 55 (44.0%) | ||
| 1 | 33 (18%) | 7 (21.2%) | 7 (21.2%) | 19 (57.6%) | ||
| 2 | 25 (13.7%) | 9 (36.0%) | 5 (20.0%) | 11 (44.0%) | ||
| M_stage | 0.086 | |||||
| 0 | 183 (97.9%) | 48 (26.2%) | 50 (27.3%) | 85 (46.4%) | ||
| 1 | 4 (2.1%) | 3 (75.0%) | 0 | 1 (25.0%) | ||
| AJCC stage | 0.063 | |||||
| 1 | 77 (41.8%) | 23 (29.9%) | 26 (33.8%) | 28 (36.4%) | ||
| 2 | 68 (37%) | 14 (20.6%) | 16 (23.5%) | 38 (55.9%) | ||
| 3 | 37 (20.1%) | 10 (27.0%) | 8 (21.6%) | 19 (51.4%) | ||
| 4 | 2 (1.1%) | 2 (100%) | 0 | 0 | ||
| Preop Tx. | 0.343 | |||||
| No | 169 (89.9%) | 45 (26.6%) | 48 (28.4%) | 76 (45.0%) | ||
| PreopCCRT | 16 (8.5%) | 5 (31.3%) | 2 (12.5%) | 9 (56.3%) | ||
| PreopChemo | 3 (1.6%) | 2 (66.7%) | 0 | 1 (33.3%) | ||
| Vascular invasion | 0.247 | |||||
| No | 172 (93%) | 45 (26.2%) | 49 (28.5%) | 78 (45.3%) | ||
| YES | 13 (7%) | 5 (38.5%) | 1 (7.7%) | 7 (53.8%) | ||
| Lymphatic invasion | 0.139 | |||||
| No | 144 (77.8%) | 34 (23.6%) | 40 (27.8%) | 70 (48.6%) | ||
| YES | 41 (22.2%) | 16 (39.0%) | 10 (24.4%) | 15 (36.6%) | ||
| Differentiation | 0.044 | |||||
| W/D | 9 (4.8%) | 2 (22.2%) | 4 (44.4%) | 3 (33.3%) | ||
| M/D | 129 (68.6%) | 29 (22.5%) | 35 (27.1%) | 65 (50.4%) | ||
| P/D | 41 (21.8%) | 15 (36.6%) | 11 (26.8%) | 15 (36.6%) | ||
| Adenocarcinoma (n = 785) | |||||||
|---|---|---|---|---|---|---|---|
| PD-L1 | |||||||
| Driver Gene | Total Patients No. | <1% (n = 495) | 1–49% (n = 177) | ≥50% (n = 113) | p-Value | ||
| EGFR | <0.001 | ||||||
| Wild | 361 (46.0%) | 195 (54.0%) | 87 (24.1%) | 79 (21.9%) | |||
| Mutant | 424 (54.0%) | 300 (70.8%) | 90 (21.2%) | 34 (8.0%) | |||
| Exon19del | 182 (42.9%) | 119 (65.4%) | 42 (23.1%) | 21 (11.5%) | |||
| L858R | 202 (47.6%) | 153 (75.7%) | 39 (19.3%) | 10 (5.0%) | |||
| Exon20ins | 17 (4.0%) | 11 (64.7%) | 4 (23.5%) | 2 (11.8%) | |||
| Others | 23 (5.4%) | 17 (73.9%) | 5 (21.7%) | 1 (4.3%) | |||
| ALK | <0.001 | ||||||
| Wild | 758 (96.6%) | 490 (64.6%) | 168 (22.2%) | 100 (13.2%) | |||
| Mutant | 27 (3.4%) | 5 (18.5%) | 9 (33.3%) | 13 (48.1%) | |||
| EGFR-Mutated Adenocarcinoma (n = 424) | ||||||
|---|---|---|---|---|---|---|
| PD-L1 Expression | ||||||
| Variable | Total Patients No. | <1% (n = 300, 70.8%) | 1–49% (n = 90, 21.2%) | ≥50% (n = 34, 8.0%) | p-Value | |
| Age | 0.159 | |||||
| <65 | 255 (60.1%) | 188 (73.7%) | 51 (20.0%) | 16 (6.3%) | ||
| ≥65 | 169 (39.9%) | 112 (66.3.%) | 39 (23.1%) | 18 (10.7%) | ||
| Sex | 0.202 | |||||
| Male | 157 (37.0%) | 103 (65.6%) | 39 (24.8%) | 15 (9.6%) | ||
| Female | 267 (63.0%) | 197 (73.8%) | 51 (19.1%) | 19 (7.1%) | ||
| Smoking status | 0.025 | |||||
| Never smoker | 279 (65.8%) | 209 (74.9%) | 49 (17.6%) | 21 (7.5%) | ||
| Current or Ex smoker | 145 (34.2%) | 91 (62.8%) | 41 (28.3%) | 13 (9.0%) | ||
| T_stage | 0.003 | |||||
| 1 | 292 (69.2%) | 223 (76.4%) | 53 (18.2%) | 16 (5.5%) | ||
| 2 | 96 (22.7%) | 53 (55.2%) | 30 (31.3%) | 13 (13.5%) | ||
| 3 | 30 (7.1%) | 20 (66.7%) | 5 (16.7%) | 5 (16.7%) | ||
| 4 | 4 (0.9%) | 3 (75.0%) | 1 (25.0%) | 0.00% | ||
| N_stage | 0.001 | |||||
| 0 | 341 (83.2%) | 256 (75.1%) | 64 (18.8%) | 21 (6.2%) | ||
| 1 | 18 (4.4%) | 11 (61.1%) | 5 (27.8%) | 2 (11.1%) | ||
| 2 | 51 (12.4%) | 25 (49.0%) | 16 (31.4%) | 10 (19.6%) | ||
| M_stage | 0.011 | |||||
| 0 | 406 (96.2%) | 291 (71.7%) | 81 (20.0%) | 34 (8.4%) | ||
| 1 | 16 (3.8%) | 8 (50.0%) | 8 (50.0%) | 0 | ||
| AJCC stage | <0.001 | |||||
| 1 | 315 (75.7%) | 238 (75.6%) | 58 (18.4%) | 19 (6.0%) | ||
| 2 | 37 (8.9%) | 26 (70.3%) | 8 (21.6%) | 3 (8.1%) | ||
| 3 | 50 (12.0%) | 25 (50.0%) | 14 (28.0%) | 11 (22.0%) | ||
| 4 | 14 (3.4%) | 6 (42.9%) | 8 (57.1%) | 0 | ||
| Preop Tx. | 0.298 | |||||
| No | 407 (96.0%) | 284 (69.8%) | 89 (21.9%) | 34 (8.4%) | ||
| PreopCCRT | 12 (2.8%) | 11 (91.7%) | 1 (8.3%) | 0 | ||
| PreopChemo | 5 (1.2%) | 5 (100%) | 0 | 0 | ||
| Vascular invasion | 0.161 | |||||
| No | 398 (96.1%) | 286 (71.9%) | 80 (20.1%) | 32 (8.0%) | ||
| YES | 16 (3.9%) | 8 (50.0%) | 6 (37.5%) | 2 (12.5%) | ||
| Lymphatic invasion | <0.001 | |||||
| No | 323 (78.0%) | 250 (77.4%) | 57 (17.6%) | 16 (5.0%) | ||
| YES | 91 (22.0%) | 44 (48.4%) | 29 (31.9%) | 18 (19.8%) | ||
| Differentiation | <0.001 | |||||
| W/D | 20 (4.7%) | 18 (90.0%) | 2 (10.0%) | 0 | ||
| M/D | 340 (80.2%) | 251 (73.8%) | 70 (20.6%) | 19 (5.6%) | ||
| P/D | 51 (12.0%) | 20 (39.2%) | 16 (31.4%) | 15 (29.4%) | ||
| Predominant pattern | <0.001 | |||||
| Lepidic | 26 (6.3%) | 23 (88.5%) | 3 (11.5%) | 0 | ||
| Acinar | 273 (65.8%) | 205 (75.1%) | 54 (19.8%) | 14 (5.1%) | ||
| Papillary | 58 (14.0%) | 38 (65.5%) | 16 (27.6%) | 4 (6.9%) | ||
| Micropapillary | 27 (6.5%) | 13 (48.1%) | 9 (33.3%) | 5 (18.5%) | ||
| Solid | 21 (5.1%) | 7 (33.3%) | 5 (23.8%) | 9 (42.9%) | ||
| Cribriform | 6 (1.4%) | 3 (50.0%) | 1 (16.7%) | 2 (33.3%) | ||
| Mucinous | 4 (1.0%) | 4 (100.0%) | 0 | 0 | ||
| EGFR mutation genotyping | 0.229 | |||||
| Exon19del | 182 (42.9%) | 119 (65.4%) | 42 (23.1%) | 21 (11.5%) | ||
| L858R | 202 (47.6%) | 153 (75.7%) | 39 (19.3%) | 10 (5.0%) | ||
| Exon20insertion | 17 (4.0%) | 11 (64.7%) | 4 (23.5%) | 2 (11.8%) | ||
| Others | 23 (5.4%) | 17 (73.9%) | 5 (21.7%) | 1 (4.3%) | ||
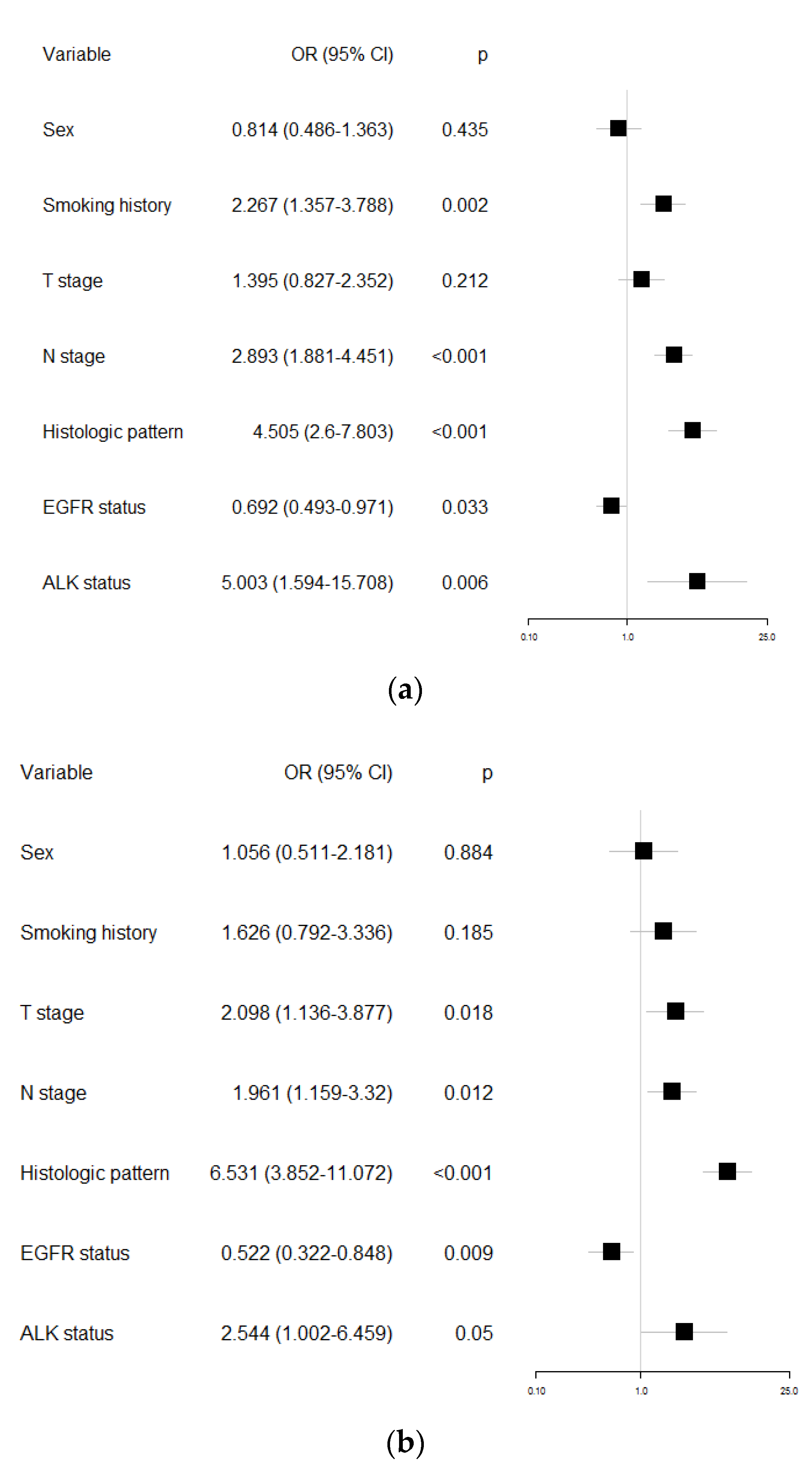
| Factors to Predict PD-L1 Positivity in Lung Adenocarcinoma | Univariable Analysis | Multivariable Analysis | |||
|---|---|---|---|---|---|
| (1% Cutoff Value for PD-L1 Positivity) | OR (95% CI) | p-Value | OR (95% CI) | p-Value | |
| Sex | Male (vs. Female) | 1.723 (1.286–2.309) | <0.001 | 0.814 (0.486–1.363) | 0.435 |
| Age | ≥65 (vs. < 65) | 1.320 (0.986–1.767) | 0.062 | ||
| Smoking history | Ever (vs. Never) smoker | 2.027 (1.510–2.721) | <0.001 | 2.267 (1.357–3.788) | 0.002 |
| T stage | T stage 3/4 (vs. Stage 1/2) | 1.739 (1.108–2.728) | 0.016 | 1.395 (0.827–2.352) | 0.212 |
| N stage | N stage 1/2 (vs. N stage 0) | 3.058 (2.092–14.69) | <0.001 | 2.893 (1.881–4.451) | <0.001 |
| M stage | M1 stage (vs. M0 stage) | 1.518 (0.730–3.157) | 0.264 | ||
| Histologic pattern | Solid (vs. Non-solid) variant | 6.568 (3.955–10.907) | <0.001 | 4.505 (2.6–7.803) | <0.001 |
| EGFR status | Mutant (vs. Wild) type | 0.521 (0.388–0.698) | <0.001 | 0.692 (0.493–0.971) | 0.033 |
| ALK status | Positive (vs. Negative) | 8.05 (3.012–21.486) | <0.001 | 5.003 (1.594–15.708) | 0.006 |
| Factors to Predict PD-L1 Positivity in Lung Adenocarcinoma | Univariable Analysis | Multivariable Analysis | |||
|---|---|---|---|---|---|
| (50% Cutoff Value for PD-L1 Positivity) | OR (95% CI) | p-Value | OR (95% CI) | p-Value | |
| Sex | Male (vs. Female) | 2.135 (1.413–3.226) | <0.001 | 1.056 (0.511–2.181) | 0.884 |
| Age | ≥65 (vs. < 65) | 1.204 (0.808–1.796) | 0.362 | ||
| Smoking history | Ever (vs. Never) smoker | 2.335 (1.550–3.518) | <0.001 | 1.626 (0.792–3.336) | 0.185 |
| T stage | T stage 3/4 (vs. Stage 1/2) | 2.490 (1.469–4.218) | 0.001 | 2.098 (1.136–3.877) | 0.018 |
| N stage | N stage 1/2 (vs. N stage 0) | 2.500 (1.590–3.932) | <0.001 | 1.961 (1.159–3.320) | 0.012 |
| M stage | M1 stage (vs. M0 stage) | 0.461 (0.108–1.972) | 0.296 | ||
| Histologic pattern | Solid (vs. Non-solid) variant | 9.839 (6.044–16.015) | <0.001 | 6.531 (3.852–11.072) | <0.001 |
| EGFR status | Mutant (vs. Wild) type | 0.346 (0.227–0.527) | <0.001 | 0.522 (0.322–0.848) | 0.009 |
| ALK status | Positive (vs. Negative) | 6.064 (2.769–13.277) | <0.001 | 2.544 (1.002–6.459) | 0.05 |
| No. of Study | Authors | Years | Ethnicity | Tissue Type | PD-L1 Antibody | Cutoff Value | ALK Positive Cases | PD-L1 Positive Cases | p-Value |
|---|---|---|---|---|---|---|---|---|---|
| 1 [34] | Zhang | 2014 | East-Asian | Whole tissue section | SAB2900365 | 40% | 9/143 (6.3%) | 3/9 (33.3%) | 0.494 |
| 2 [13] | Yang | 2014 | East-Asian | Whole tissue section | Proteintech Group Inc. | 5% | 3/163 (1.84%) | 2/3 (66.7%) | 0.564 |
| 3 [4] | Cooper | 2015 | Non-East-Asian | TMA | 22C3 | 50% | 3/678 (0.44%) | 0/3 (0%) | 1 |
| 4 [5] | Incecco | 2015 | Non-East-Asian | Whole tissue section | ab58810 | 5% | 10/125 (8%) | 3/10 (30%) | 1 |
| 5 [9] | Koh | 2015 | East-Asian | TMA | E1L3N | 10% | 23/497 (4.63%) | 18/23 (78.3%) | 0.054 |
| 6 [10] | Ota | 2015 | East-Asian | Whole tissue section | Lifespan Biosciences | H-score | 11/134 (8.21%) | N/A | <0.001 |
| 7 [12] | Song | 2016 | East-Asian | Whole tissue section | Proteintech Group Inc. | 5% | 18/385 (4.68%) | 10/18 (55.6%) | 0.53 |
| 8 [6] | Inamura | 2016 | East-Asian | TMA | E1L3N | 5% | 10/268 (3.73%) | 1/10 (10%) | 1 |
| 9 [7] | Inoue | 2016 | East-Asian | TMA | E1L3N | 5% | 10/654 (1.53%) | 5/10 (50%) | 0.169 |
| 10 [30] | Huynh | 2016 | Non-East-Asian | TMA | E1L3N | 5% | 4/261 (1.53%) | 1/4 (25%) | N/A |
| 11 [11] | Rangachari | 2017 | Non-East-Asian | Whole tissue section | 22C3 | 50% | 4/71 (5.63%) | 1/3 (33.3%) | N/A |
| 12 [35] | Jia | 2018 | East-Asian | Whole tissue section | E1L3N | 10% | 5/55 (9.09%) | 1/5 (20%) | 0.822 |
| 13 [8] | Kim | 2018 | East-Asian | TMA | 22C3 | 1%, 50% | 24/429 (5.59%) | 3/24 (12.5%) | >0.05 |
| 14 | This study | 2019 | East-Asian | Whole tissue section | 22C3 | 1%, 50% | 27/994 (2.72%) | 22/27 (81.5%) | <0.001 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, S.E.; Kim, Y.J.; Sung, M.; Lee, M.-S.; Han, J.; Kim, H.K.; Choi, Y.-L. Association with PD-L1 Expression and Clinicopathological Features in 1000 Lung Cancers: A Large Single-Institution Study of Surgically Resected Lung Cancers with a High Prevalence of EGFR Mutation. Int. J. Mol. Sci. 2019, 20, 4794. https://doi.org/10.3390/ijms20194794
Lee SE, Kim YJ, Sung M, Lee M-S, Han J, Kim HK, Choi Y-L. Association with PD-L1 Expression and Clinicopathological Features in 1000 Lung Cancers: A Large Single-Institution Study of Surgically Resected Lung Cancers with a High Prevalence of EGFR Mutation. International Journal of Molecular Sciences. 2019; 20(19):4794. https://doi.org/10.3390/ijms20194794
Chicago/Turabian StyleLee, Seung Eun, Yu Jin Kim, Minjung Sung, Mi-Sook Lee, Joungho Han, Hong Kwan Kim, and Yoon-La Choi. 2019. "Association with PD-L1 Expression and Clinicopathological Features in 1000 Lung Cancers: A Large Single-Institution Study of Surgically Resected Lung Cancers with a High Prevalence of EGFR Mutation" International Journal of Molecular Sciences 20, no. 19: 4794. https://doi.org/10.3390/ijms20194794
APA StyleLee, S. E., Kim, Y. J., Sung, M., Lee, M.-S., Han, J., Kim, H. K., & Choi, Y.-L. (2019). Association with PD-L1 Expression and Clinicopathological Features in 1000 Lung Cancers: A Large Single-Institution Study of Surgically Resected Lung Cancers with a High Prevalence of EGFR Mutation. International Journal of Molecular Sciences, 20(19), 4794. https://doi.org/10.3390/ijms20194794





