Genome-Wide Expression Profiling of Genes Associated with the Lr47-Mediated Wheat Resistance to Leaf Rust (Puccinia triticina)
Abstract
1. Introduction
2. Results
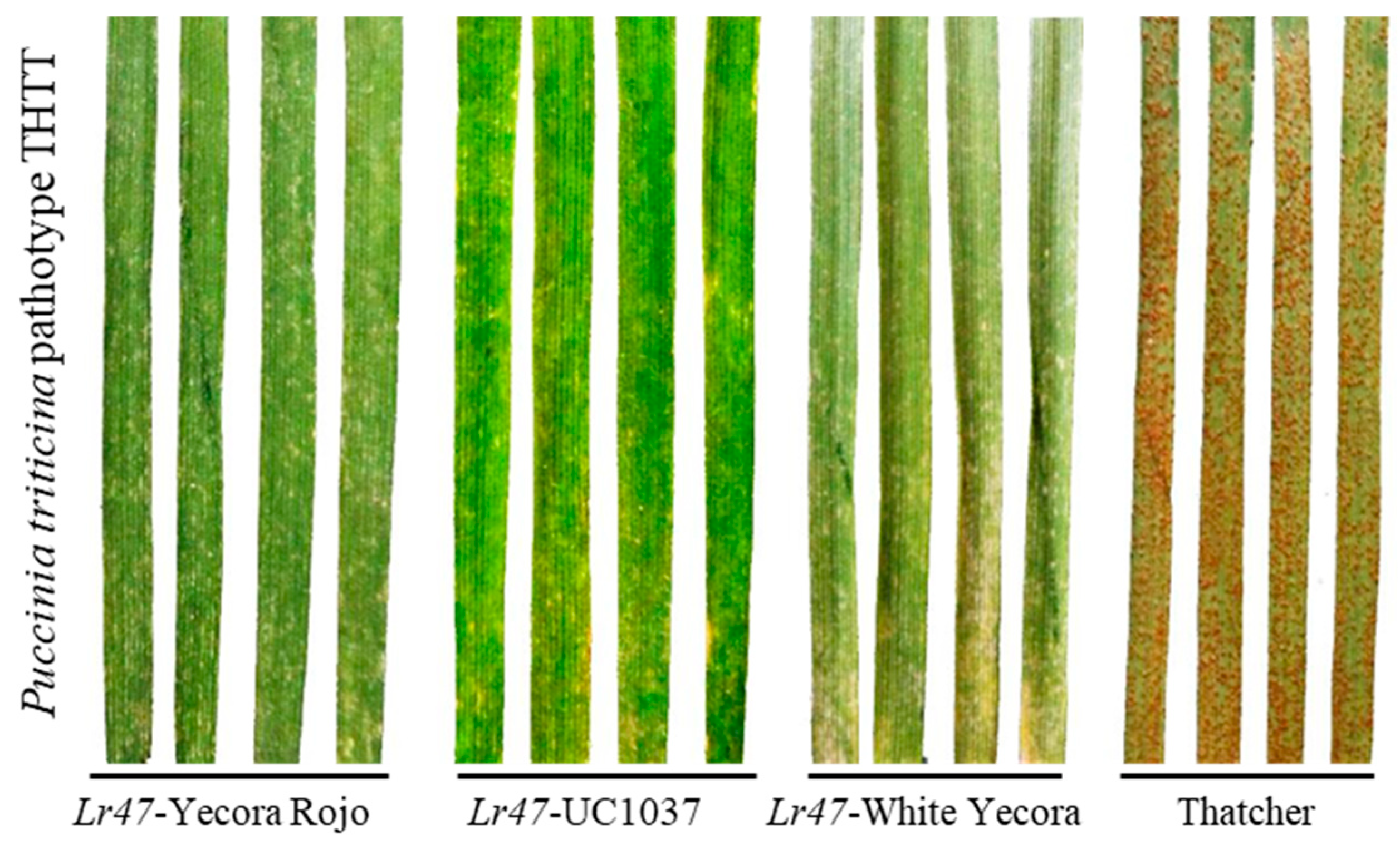
2.1. Wheat Lines Carrying the Lr47 Gene Showed High Resistance to the Epidemic Leaf Rust Race THTT
2.2. The Wheat Lines Carrying the Lr47 Gene were Subjected to RNA-Seq Analysis upon Pt Infection
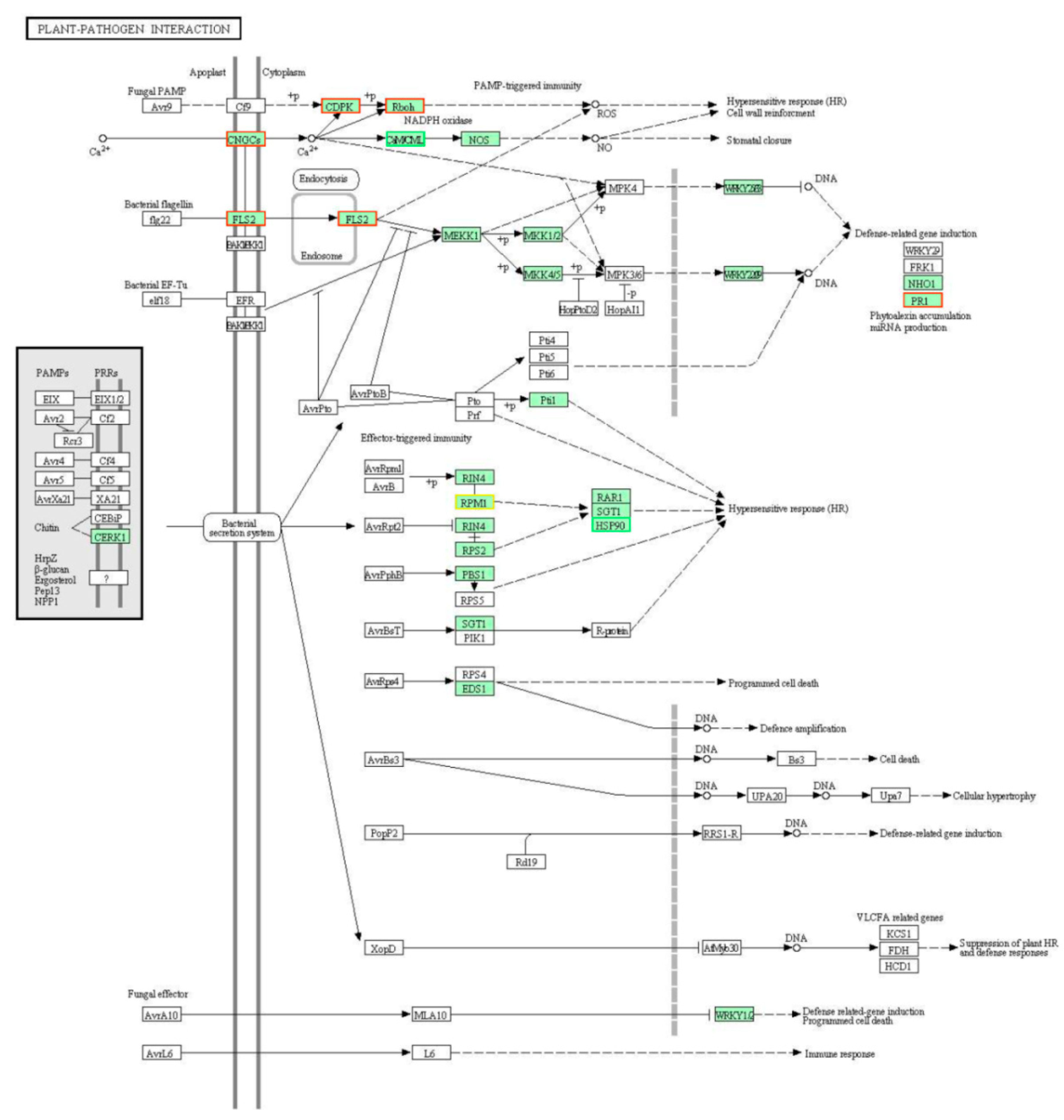
2.3. The Overall DEGs Involved in the Lr47-Mediated Resistance
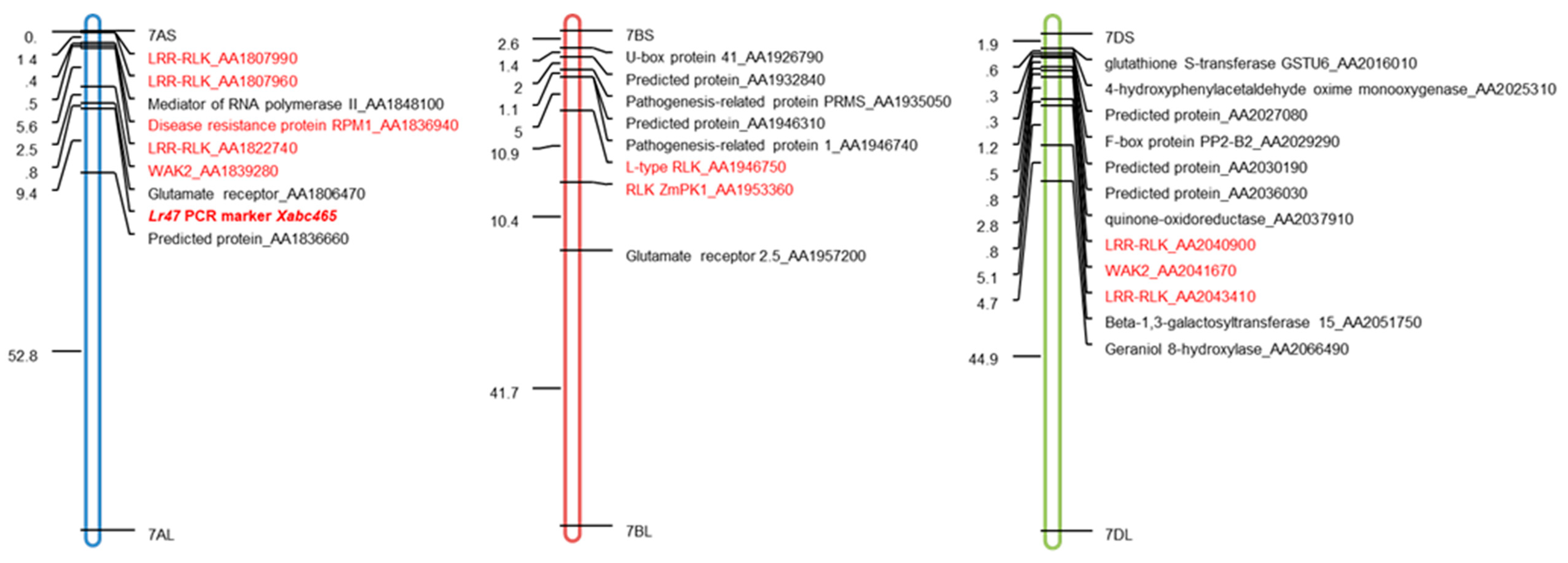
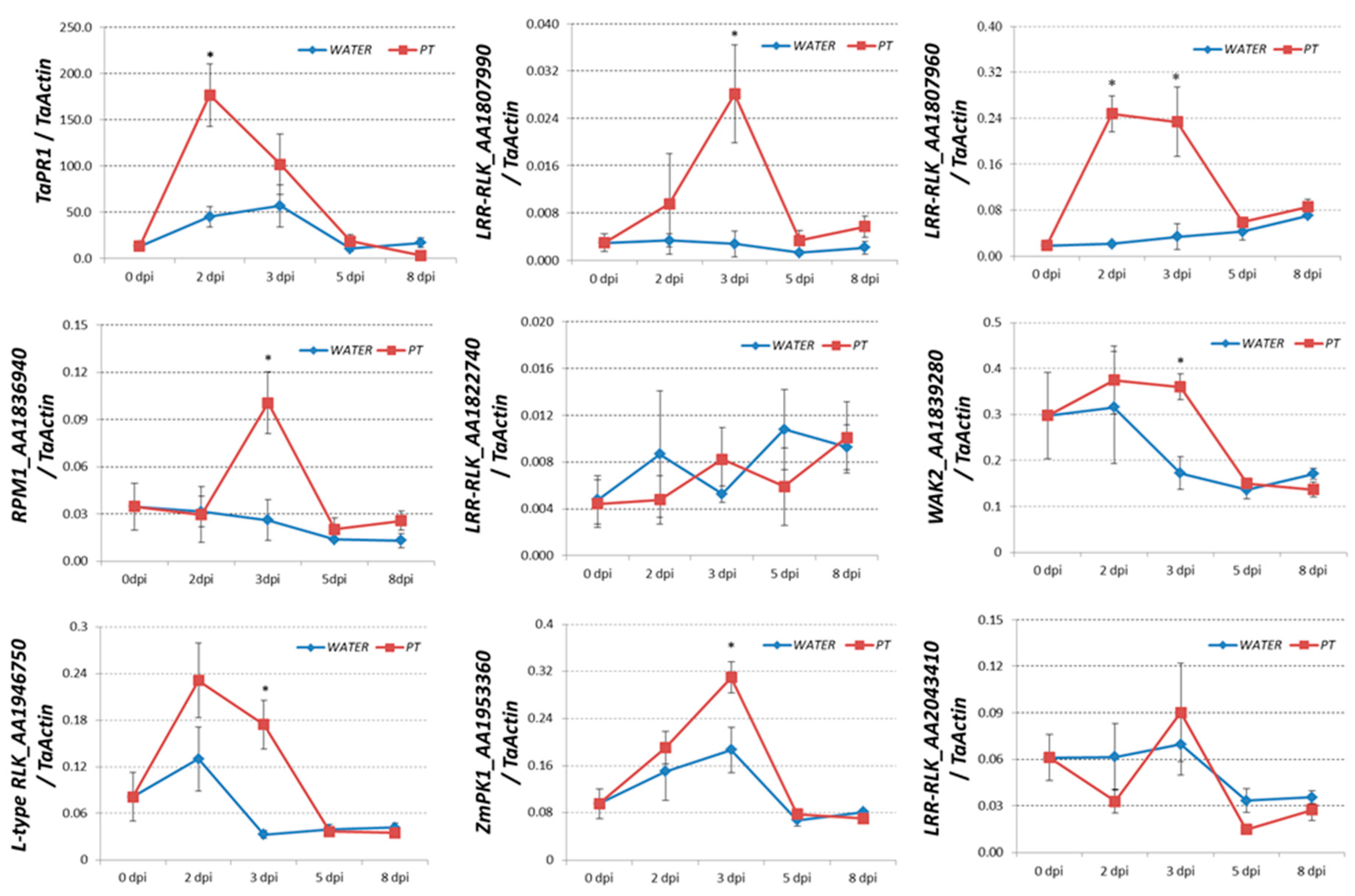
2.4. Transcriptional-Induced Genes Located on Chromosomes 7AS, 7BS, and 7DS were Identified
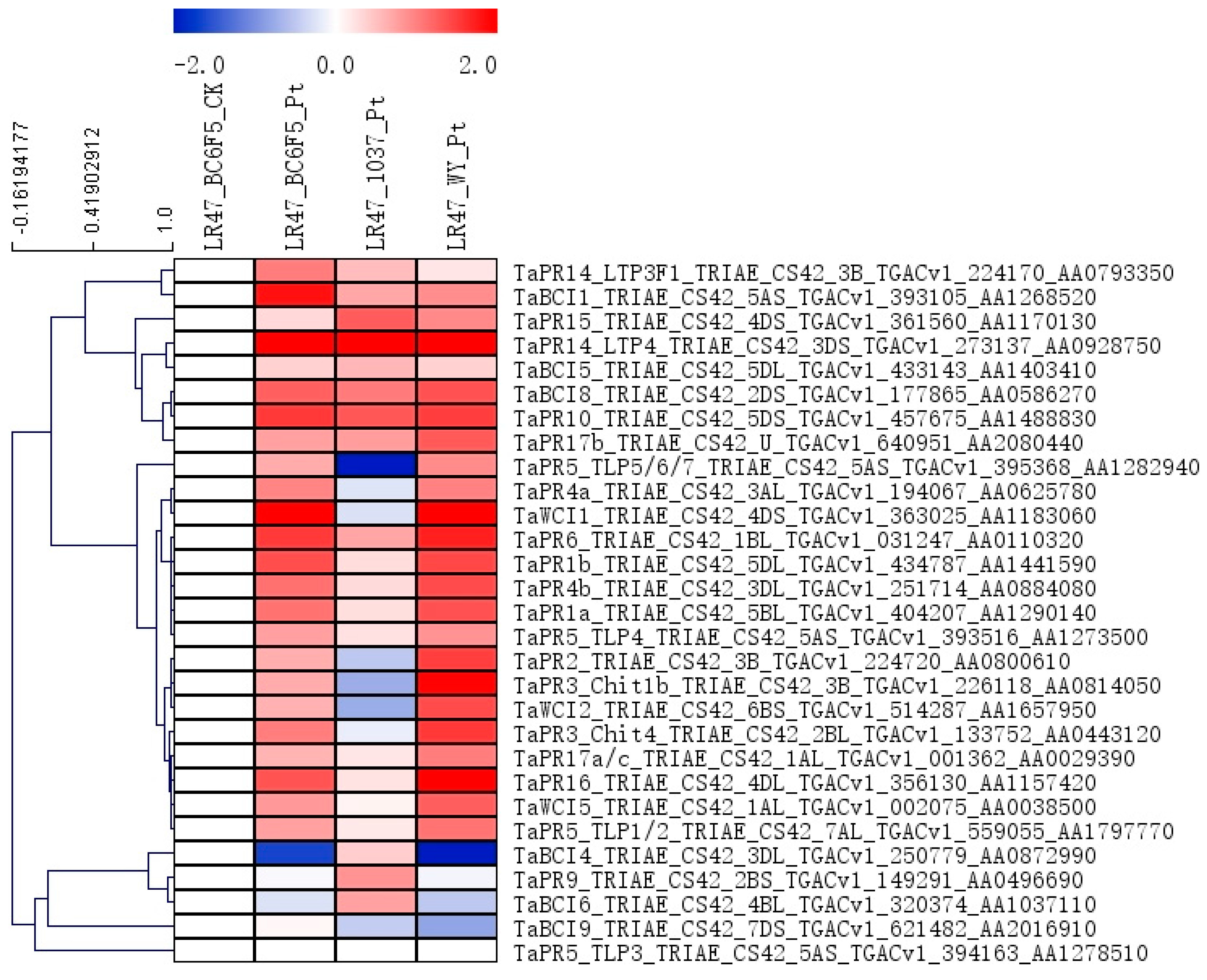
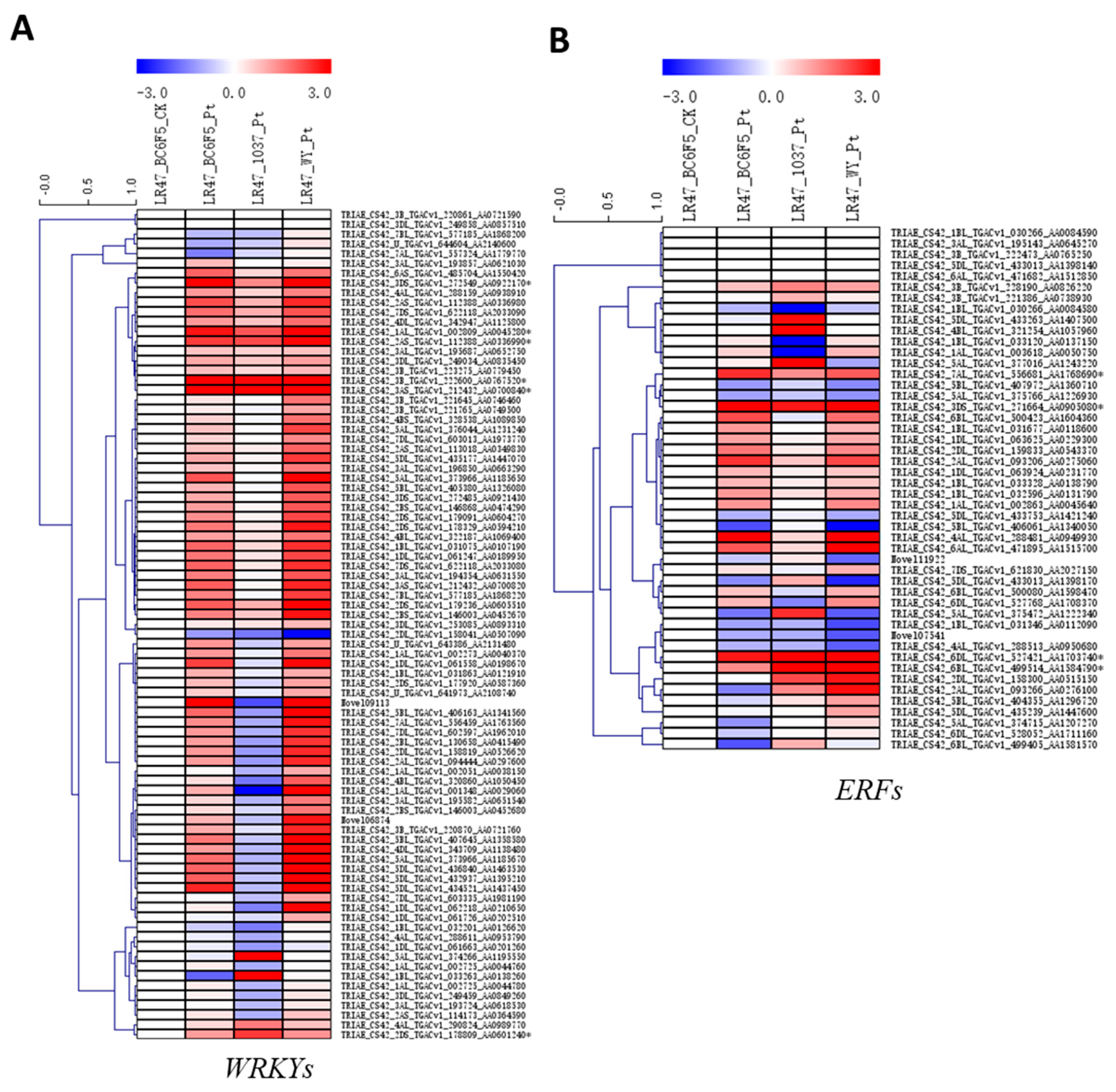
2.5. The Expression Patterns of Genes Encoding PR Proteins and Several Transcription Factor Families during the Lr47-Mediated Resistance were Profiled
3. Discussion
4. Materials and Methods
4.1. Plants Growth and Leaf Rust Inoculation
4.2. RNA-seq and Bioinformatics Analysis
4.3. qRT-PCR Assay
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Bolton, M.D.; Kolmer, J.A.; Garvin, D.F. Wheat leaf rust caused by Puccinia triticina. Mol. Plant Pathol. 2008, 9, 563–575. [Google Scholar] [CrossRef] [PubMed]
- Eversmeyer, M.G.; Kramer, C.L. Epidemiology of wheat leaf and stem rust in the central great plains of the USA. Annu. Rev. Phytopathol. 2000, 38, 491–513. [Google Scholar] [CrossRef] [PubMed]
- Huerta-Espino, J.; Singh, R.; German, S.; McCallum, B.; Park, R.; Chen, W.Q.; Bhardwaj, S.; Goyeau, H. Global status of wheat leaf rust caused by Puccinia triticina. Euphytica 2011, 179, 143–160. [Google Scholar] [CrossRef]
- Helfer, S. Rust fungi and global change. New Phytol. 2014, 201, 770–780. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Meng, Q.; Yan, H.; Liu, D. Virulence and molecular genetic diversity of the Puccinia triticina population in Hebei Province of China in 2008 and 2010. Eur. J. Plant Pathol. 2019, 153, 153–166. [Google Scholar] [CrossRef]
- Zhou, H.; Xia, X.; He, Z.; Li, X.; Wang, C.; Li, Z.; Liu, D. Molecular mapping of leaf rust resistance gene LrNJ97 in Chinese wheat line Neijiang 977671. Theor. Appl. Genet. 2013, 126, 2141–2147. [Google Scholar] [CrossRef] [PubMed]
- Cloutier, S.; McCallum, B.D.; Loutre, C.; Banks, T.W.; Wicker, T.; Feuillet, C.; Keller, B.; Jordan, M.C. Leaf rust resistance gene Lr1, isolated from bread wheat (Triticum aestivum L.) is a member of the large psr567 gene family. Plant Mol. Biol. 2007, 65, 93–106. [Google Scholar] [CrossRef] [PubMed]
- Feuillet, C.; Travella, S.; Stein, N.; Albar, L.; Nublat, A.; Keller, B. Map-based isolation of the leaf rust disease resistance gene Lr10 from the hexaploid wheat (Triticum aestivum L.) genome. Proc. Natl. Acad. Sci. USA 2003, 100, 15253–15258. [Google Scholar] [CrossRef]
- Huang, L.; Brooks, S.A.; Li, W.; Fellers, J.P.; Trick, H.N.; Gill, B.S. Map-based cloning of leaf rust resistance gene Lr21 from the large and polyploid genome of bread wheat. Genetics 2003, 164, 655–664. [Google Scholar]
- Fu, D.; Uauy, C.; Distelfeld, A.; Blechl, A.; Epstein, L.; Chen, X.; Sela, H.; Fahima, T.; Dubcovsky, J. A kinase-START gene confers temperature-dependent resistance to wheat stripe rust. Science 2009, 323, 1357–1360. [Google Scholar] [CrossRef]
- Krattinger, S.G.; Lagudah, E.S.; Spielmeyer, W.; Singh, R.P.; Huerta-Espino, J.; McFadden, H.; Bossolini, E.; Selter, L.L.; Keller, B. A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 2009, 323, 1360–1363. [Google Scholar] [CrossRef] [PubMed]
- Moore, J.W.; Herrera-Foessel, S.; Lan, C.; Schnippenkoetter, W.; Ayliffe, M.; Huerta-Espino, J.; Lillemo, M.; Viccars, L.; Milne, R.; Periyannan, S. A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat. Genet. 2015, 47, 1494. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wang, X.; Kang, Z.; Ren, Z.; Bi, W.; Yang, W.; Liu, D. Suppression subtractive hybridization and microarray analysis reveal differentially expressed genes in the Lr39/41-mediated wheat resistance to Puccinia triticina. Eur. J. Plant Pathol. 2018. [Google Scholar] [CrossRef]
- Yadav, I.S.; Sharma, A.; Kaur, S.; Nahar, N.; Bhardwaj, S.C.; Sharma, T.R.; Chhuneja, P. Comparative temporal transcriptome profiling of wheat near isogenic line carrying Lr57 under compatible and incompatible interactions. Front. Plant Sci. 2016, 7, 1943. [Google Scholar] [CrossRef] [PubMed]
- Singh, D.; Kumar, D.; Satapathy, L.; Pathak, J.; Chandra, S.; Riaz, A.; Bhaganagre, G.; Dhariwal, R.; Kumar, M.; Prabhu, K.V. Insights of Lr28 mediated wheat leaf rust resistance: Transcriptomic approach. Gene 2017, 637, 72–89. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Yang, Y.; Wang, C.; Liu, M.; Li, H.; Fu, Y.; Wang, Y.; Nie, Y.; Liu, X.; Ji, W. Large-scale transcriptome comparison reveals distinct gene activations in wheat responding to stripe rust and powdery mildew. BMC Genom. 2014, 15, 898. [Google Scholar] [CrossRef] [PubMed]
- Xiao, J.; Jin, X.; Jia, X.; Wang, H.; Cao, A.; Zhao, W.; Pei, H.; Xue, Z.; He, L.; Chen, Q. Transcriptome-based discovery of pathways and genes related to resistance against Fusarium head blight in wheat landrace Wangshuibai. BMC Genom. 2013, 14, 197. [Google Scholar] [CrossRef]
- Hao, Y.; Wang, T.; Kang, W.; Wang, X.; Fu, Y.; Huang, L.; Kang, Z. Transcriptome Analysis Provides Insights into the Mechanisms Underlying Wheat Plant Resistance to Stripe Rust at the Adult Plant Stage. PloS ONE 2016, 11, e0150717. [Google Scholar] [CrossRef]
- Dubcovsky, J.; Lukaszewski, A.; Echaide, M.; Antonelli, E.; Porter, D. Molecular characterization of two Triticum speltoides interstitial translocations carrying leaf rust and greenbug resistance genes. Crop Sci. 1998, 38, 1655–1660. [Google Scholar] [CrossRef]
- Helguera, M.; Khan, I.A.; Dubcovsky, J. Development of PCR markers for the wheat leaf rust resistance gene Lr47. Theor. Appl. Genet. 2000, 101, 625–631. [Google Scholar] [CrossRef]
- Vanzetti, L.S.; Brevis, J.C.; Dubcovsky, J.; Helguera, M. Identification of microsatellites linked to Lr47. Electron. J. Biotechnol. 2006, 9, 267–271. [Google Scholar] [CrossRef]
- Brevis, J.C.; Chicaiza, O.; Khan, I.A.; Jackson, L.; Morris, C.F.; Dubcovsky, J. Agronomic and Quality Evaluation of Common Wheat Near-Isogenic Lines Carrying the Leaf Rust Resistance Gene Lr47. Crop Sci. 2008, 48, 1441–1451. [Google Scholar] [CrossRef]
- Gebrewahid, T.W.; Yao, Z.; Yan, X.C.; Gao, P.; Li, Z. Identification of Leaf Rust Resistance Genes in Chinese Common Wheat Cultivars. Plant Dis. 2017, 101. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Bi, W.; Gao, J.; Yu, X.; Wang, H.; Liu, D. Systemic acquired resistance, NPR1, and pathogenesis-related genes in wheat and barley. J. Integr. Agric. 2018, 17, 2468–2477. [Google Scholar] [CrossRef]
- Görlach, J.; Volrath, S.; Knaufbeiter, G.; Hengy, G.; Beckhove, U.; Kogel, K.H.; Oostendorp, M.; Staub, T.; Ward, E.; Kessmann, H. Benzothiadiazole, a novel class of inducers of systemic acquired resistance, activates gene expression and disease resistance in wheat. Plant Cell 1996, 8, 629–643. [Google Scholar] [PubMed]
- Beßer, K.; Jarosch, B.; Langen, G.; Kogel, K.-H. Expression analysis of genes induced in barley after chemical activation reveals distinct disease resistance pathways. Mol. Plant Pathol. 2000, 1, 277–286. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Xia, X.; He, Z.; Li, X.; Zhang, L.; Wang, H.; Meng, Q.; Yang, W.; Li, G.; Liu, D. Seedling and slow rusting resistance to leaf rust in Chinese wheat cultivars. Plant Dis. 2010, 94, 45–53. [Google Scholar] [CrossRef]
- Li, H.; Mahmood, T.; Antony, G.; Lu, N.; Pumphreys, M.; Gill, B.; Kang, Z.; White, F.F.; Bai, J. The non-host pathogen Puccinia triticina elicits an active transcriptional response in rice. Eur. J. Plant Pathol. 2016, 147, 1–17. [Google Scholar] [CrossRef]
- Morant, M.; Bak, S.; Møller, B.L.; Werck-Reichhart, D. Plant cytochromes P450: Tools for pharmacology, plant protection and phytoremediation. Curr. Opin. Biotechnol. 2003, 14, 151–162. [Google Scholar] [CrossRef]
- Kotak, S.; Larkindale, J.; Lee, U.; von Koskull-Döring, P.; Vierling, E.; Scharf, K.D. Complexity of the heat stress response in plants. Curr. Opin. Plant Biol. 2007, 10, 310–316. [Google Scholar] [CrossRef]
- Mauchmani, B.; Mauch, F. The role of abscisic acid in plant-pathogen interactions. Curr. Opin. Plant Biol. 2005, 8, 409–414. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Islam, M.A.; Lin, H.; Ji, C.; Duan, Y.; Liu, P.; Zeng, Q.; Day, B.; Kang, Z.; Guo, J. Genome-Wide Identification of Cyclic Nucleotide-Gated Ion Channel Gene Family in Wheat and Functional Analyses of TaCNGC14 and TaCNGC16. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Duan, Y.; Liu, C.; Xue, Q.; Guo, J.; Qi, T.; Kang, Z.; Guo, J. The calcium sensor TaCBL4 and its interacting protein TaCIPK5 are required for wheat resistance to stripe rust fungus. J. Exp. Bot. 2018, 69, 4443–4457. [Google Scholar] [CrossRef] [PubMed]
- Steuernagel, B.; Witek, K.; Krattinger, S.G.; Ramirez-Gonzalez, R.H.; Schoonbeek, H.-j.; Yu, G.; Baggs, E.; Witek, A.; Yadav, I.; Krasileva, K.V.; et al. Physical and transcriptional organisation of the bread wheat intracellular immune receptor repertoire. bioRxiv 2018. [Google Scholar]
- Tang, D.; Wang, G.; Zhou, J.-M. Receptor kinases in plant-pathogen interactions: More than pattern recognition. Plant Cell 2017, 29, 618–637. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.D.; Dangl, J.L. The plant immune system. Nature 2006, 444, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Swiderski, M.R.; Innes, R.W. The Arabidopsis PBS1 resistance gene encodes a member of a novel protein kinase subfamily. Plant J. 2001, 26, 101–112. [Google Scholar] [CrossRef] [PubMed]
- Singh, K.; Foley, R.C.; Oñatesánchez, L. Transcription factors in plant defense and stress responses. Curr. Opin. Plant Biol. 2002, 5, 430–436. [Google Scholar] [CrossRef]
- Alves, M.S.; Dadalto, S.P.; Gonçalves, A.B.; Souza, G.B.D.; Barros, V.A.; Fietto, L.G. Plant bZIP Transcription Factors Responsive to Pathogens: A Review. Int. J. Mol. Sci. 2013, 14, 7815–7828. [Google Scholar] [CrossRef]
- Ambawat, S.; Sharma, P.; Yadav, N.R.; Yadav, R.C. MYB transcription factor genes as regulators for plant responses: An overview. Physiol. Mol. Biol. Plants 2013, 19, 307. [Google Scholar] [CrossRef]
- Gao, J.; Bi, W.; Li, H.; Wu, J.; Yu, X.; Liu, D.; Wang, X. WRKY Transcription Factors Associated With NPR1-Mediated Acquired Resistance in Barley Are Potential Resources to Improve Wheat Resistance to Puccinia triticina. Front. Plant Sci. 2018, 9, 1486. [Google Scholar] [CrossRef] [PubMed]
- Clavijo, B.J.; Venturini, L.; Schudoma, C.; Accinelli, G.G.; Kaithakottil, G.; Wright, J.; Borrill, P.; Kettleborough, G.; Heavens, D.; Chapman, H. An improved assembly and annotation of the allohexaploid wheat genome identifies complete families of agronomic genes and provides genomic evidence for chromosomal translocations. Genome Res. 2017, 27, 885–896. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Pachter, L.; Salzberg, S.L. TopHat: Discovering splice junctions with RNA-Seq. Bioinformatics 2009, 25, 1105–1111. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; Van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Young, M.D.; Wakefield, M.J.; Smyth, G.K.; Oshlack, A. Gene ontology analysis for RNA-seq: Accounting for selection bias. Genome Biol. 2010, 11, R14. [Google Scholar] [CrossRef] [PubMed]
- Keeble-Gagnère, G.; Rigault, P.; Tibbits, J.; Pasam, R.; Hayden, M.; Forrest, K.; Frenkel, Z.; Korol, A.; Huang, B.E.; Cavanagh, C. Optical and physical mapping with local finishing enables megabase-scale resolution of agronomically important regions in the wheat genome. Genome Biol. 2018, 19, 112. [Google Scholar] [CrossRef] [PubMed]
- Paolacci, A.R.; Tanzarella, O.A.; Porceddu, E.; Ciaffi, M. Identification and validation of reference genes for quantitative RT-PCR normalization in wheat. BMC Mol. Biol. 2009, 10, 11. [Google Scholar] [CrossRef]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative C T method. Nat. Protoc. 2008, 3, 1101. [Google Scholar] [CrossRef]
- Chen, A.; Li, C.; Hu, W.; Lau, M.Y.; Lin, H.; Rockwell, N.C.; Martin, S.S.; Jernstedt, J.A.; Lagarias, J.C.; Dubcovsky, J. PHYTOCHROME C plays a major role in the acceleration of wheat flowering under long-day photoperiod. Proc. Natl. Acad. Sci. USA 2014, 111, 10037–10044. [Google Scholar] [CrossRef]

| KEGG Annotation | Gene ID | R47_CK FPKM | R47_PT FPKM | R47_1037 FPKM | R47_WY FPKM | R47_PT vs R47_CK P Adjust | Gene Annotation |
|---|---|---|---|---|---|---|---|
| CNGCs | TRIAE_CS42_4BL_TGACv1_322377_AA1070960 | 6.93 | 10.86 | 0.85 | 0.66 | 1.1 × 10−5 | Zinc finger RING/FYVE/PHD-type protein |
| CDPK | TRIAE_CS42_5AL_TGACv1_374737_AA1207890 | 0.18 | 0.80 | 8.13 | 10.42 | 2.0 × 10−2 | EF-Hand calcium-binding protein |
| TRIAE_CS42_2AS_TGACv1_112475_AA0338720 | 195.08 | 99.02 | 7.65 | 7.89 | 3.2 × 10−2 | TRIAE calcium-binding protein | |
| Rboh | TRIAE_CS42_3AL_TGACv1_195736_AA0653240 | 0.45 | 3.18 | 0.16 | 0.37 | 6.0 × 10−4 | Respiratory burst oxidase homologs |
| CaM | TRIAE_CS42_5AL_TGACv1_375077_AA1215670 | 0.98 | 0.45 | 2.18 | 1.52 | 2.5 × 10−2 | TRIAE calcium-binding protein |
| TRIAE_CS42_2BS_TGACv1_146898_AA0475060 | 25.78 | 13.30 | 4.35 | 1.66 | 3.8 × 10−3 | TRIAE calcium-binding protein | |
| TRIAE_CS42_1DS_TGACv1_080174_AA0242230 | 184.29 | 92.24 | 7.04 | 6.37 | 3.0 × 10−2 | TRIAE calcium-binding protein | |
| TRIAE_CS42_1BS_TGACv1_050118_AA0167620 | 0.17 | 0.98 | 4.70 | 2.70 | 4.7 × 10−2 | TRIAE calcium-binding protein | |
| FLS2 | TRIAE_CS42_2AL_TGACv1_094921_AA0304880 | 0.59 | 2.02 | 0.34 | 0.62 | 5.0 × 10−4 | LRR receptor-like serine/threonine-protein kinase FLS2 |
| RPM1 | TRIAE_CS42_7AS_TGACv1_570514_AA1836940 | 1.02 | 1.73 | 0.67 | 1.16 | 2.5 × 10−4 | NB-ARC, P-loop containing nucleoside triphosphate hydrolase |
| TRIAE_CS42_4AL_TGACv1_290009_AA0980370 | 4.04 | 1.83 | 0.49 | 0.35 | 3.2 × 10−2 | NB-ARC, P-loop containing nucleoside triphosphate hydrolase | |
| TRIAE_CS42_2AS_TGACv1_114436_AA0367230 | 0.47 | 1.08 | 0.09 | 0.41 | 2.7 × 10−2 | NB-ARC, P-loop containing nucleoside triphosphate hydrolase | |
| TRIAE_CS42_1DS_TGACv1_080256_AA0244390 | 5.26 | 13.59 | 1.22 | 2.83 | 3.3 × 10−2 | NB-ARC, P-loop containing nucleoside triphosphate hydrolase | |
| PR1 | TRIAE_CS42_U_TGACv1_709776_AA2166170 | 5.52 | 2.89 | 3.06 | 0.87 | 1.9 × 10−2 | Cysteine-rich secretory protein |
| TRIAE_CS42_7BS_TGACv1_592960_AA1946740 | 1.52 | 2.46 | 5.40 | 8.16 | 4.5 × 10−2 | Cysteine-rich secretory protein | |
| TRIAE_CS42_7BS_TGACv1_592288_AA1935050 | 0.26 | 0.69 | 1.75 | 1.96 | 4.1 × 10−2 | Cysteine-rich secretory protein | |
| TRIAE_CS42_5BL_TGACv1_405770_AA1335010 | 0.74 | 0.25 | 22.06 | 37.50 | 3.1 × 10−2 | Cysteine-rich secretory protein | |
| TRIAE_CS42_5BL_TGACv1_405157_AA1321310 | 0.13 | 0.39 | 3.83 | 3.28 | 8.1 × 10−14 | Cysteine-rich secretory protein | |
| HSP90 | TRIAE_CS42_7BS_TGACv1_593752_AA1954160 | 10.16 | 5.17 | 97.98 | 135.18 | 7.4 × 10−3 | Heat shock protein Hsp90 |
| TRIAE_CS42_7AS_TGACv1_571863_AA1850400 | 6.02 | 9.48 | 105.27 | 154.30 | 1.1 × 10−2 | Heat shock protein Hsp90 | |
| TRIAE_CS42_5BS_TGACv1_423847_AA1384070 | 5.20 | 2.04 | 14.29 | 14.90 | 1.6 × 10−2 | Heat shock protein Hsp90 | |
| TRIAE_CS42_2DS_TGACv1_177184_AA0568130 | 24.49 | 61.73 | 0.81 | 0.31 | 8.9 × 10−3 | Heat shock protein Hsp90 |
| Gene ID | R47_CK FPKM Value | R47_PT FPKM Value | R47_1037 FPKM Value | R47_WY FPKM Value | R47_PT vs R47_CK P Adjust Value | Gene Annotation |
|---|---|---|---|---|---|---|
| TRIAE_CS42_7AS_TGACv1_569126_AA1807990 | 1.44 | 3.37 | 1.94 | 3.63 | 8.4 × 10−4 | Putative LRR receptor-like serine/threonine-protein kinase |
| TRIAE_CS42_7AS_TGACv1_569126_AA1807960 | 0.44 | 1.33 | 0.78 | 1.31 | 5.7 × 10−3 | Putative LRR receptor-like serine/threonine-protein kinase |
| TRIAE_CS42_7AS_TGACv1_570514_AA1836940 | 0.47 | 1.08 | 0.67 | 1.16 | 2.5 × 10−4 | Putative LRR receptor-like serine/threonine-protein kinase |
| TRIAE_CS42_7AS_TGACv1_569722_AA1822740 | 0.09 | 0.81 | 0.16 | 0.40 | 7.2 × 10−5 | Putative LRR receptor-like serine/threonine-protein kinase |
| TRIAE_CS42_7AS_TGACv1_570678_AA1839280 | 0.17 | 0.50 | 0.37 | 0.74 | 9.15 × 10−6 | Wall-associated receptor kinase 2 |
| TRIAE_CS42_7BS_TGACv1_592960_AA1946750 | 3.39 | 7.68 | 5.04 | 8.88 | 5.91 × 10−4 | L-type receptor-like protein kinase IX.1 |
| TRIAE_CS42_7BS_TGACv1_593644_AA1953360 | 1.25 | 2.54 | 2.87 | 3.11 | 1.86 × 10−5 | Receptor-like protein kinase ZmPK1 |
| TRIAE_CS42_7DS_TGACv1_622500_AA2040900 | 0.00 | 0.11 | 0.04 | 0.03 | 5.84 × 10−5 | Putative LRR receptor-like serine/threonine-protein kinase |
| TRIAE_CS42_7DS_TGACv1_622551_AA2041670 | 0.77 | 1.58 | 1.18 | 1.85 | 2.4 × 10−2 | Wall-associated receptor kinase 2 |
| TRIAE_CS42_7DS_TGACv1_622658_AA2043410 | 0.71 | 1.96 | 2.83 | 1.82 | 1.8 × 10−4 | Putative LRR receptor-like serine/threonine-protein kinase |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, J.; Gao, J.; Bi, W.; Zhao, J.; Yu, X.; Li, Z.; Liu, D.; Liu, B.; Wang, X. Genome-Wide Expression Profiling of Genes Associated with the Lr47-Mediated Wheat Resistance to Leaf Rust (Puccinia triticina). Int. J. Mol. Sci. 2019, 20, 4498. https://doi.org/10.3390/ijms20184498
Wu J, Gao J, Bi W, Zhao J, Yu X, Li Z, Liu D, Liu B, Wang X. Genome-Wide Expression Profiling of Genes Associated with the Lr47-Mediated Wheat Resistance to Leaf Rust (Puccinia triticina). International Journal of Molecular Sciences. 2019; 20(18):4498. https://doi.org/10.3390/ijms20184498
Chicago/Turabian StyleWu, Jiaojiao, Jing Gao, Weishuai Bi, Jiaojie Zhao, Xiumei Yu, Zaifeng Li, Daqun Liu, Bo Liu, and Xiaodong Wang. 2019. "Genome-Wide Expression Profiling of Genes Associated with the Lr47-Mediated Wheat Resistance to Leaf Rust (Puccinia triticina)" International Journal of Molecular Sciences 20, no. 18: 4498. https://doi.org/10.3390/ijms20184498
APA StyleWu, J., Gao, J., Bi, W., Zhao, J., Yu, X., Li, Z., Liu, D., Liu, B., & Wang, X. (2019). Genome-Wide Expression Profiling of Genes Associated with the Lr47-Mediated Wheat Resistance to Leaf Rust (Puccinia triticina). International Journal of Molecular Sciences, 20(18), 4498. https://doi.org/10.3390/ijms20184498






