Potential Functions of Gem-Associated Protein 2-Like Isoform X1 in the Oriental River Prawn Macrobrachium nipponense: Cloning, qPCR, In Situ Hybridization, and RNAi Analysis
Abstract
:1. Introduction
2. Results
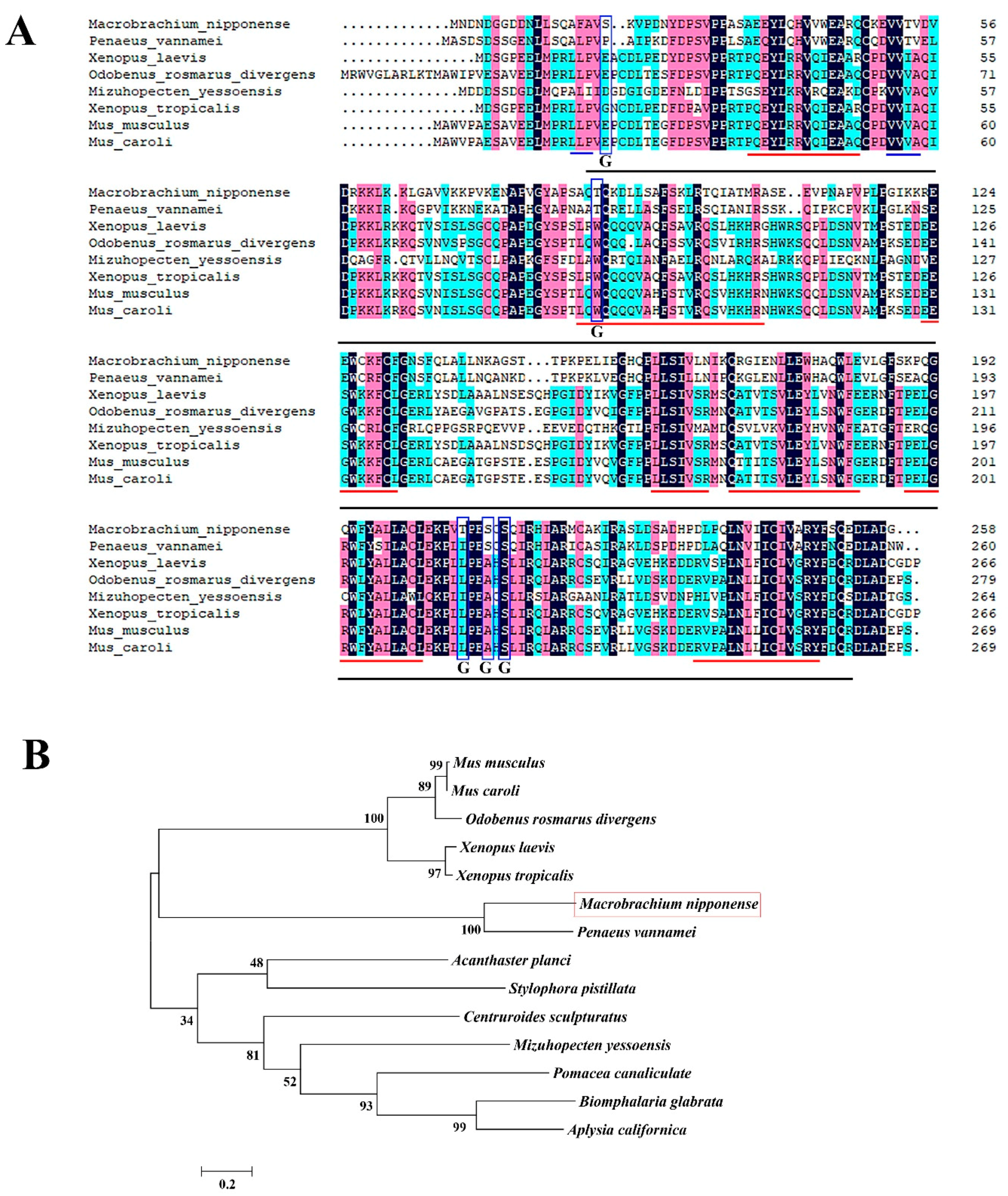
2.1. Sequence Analysis
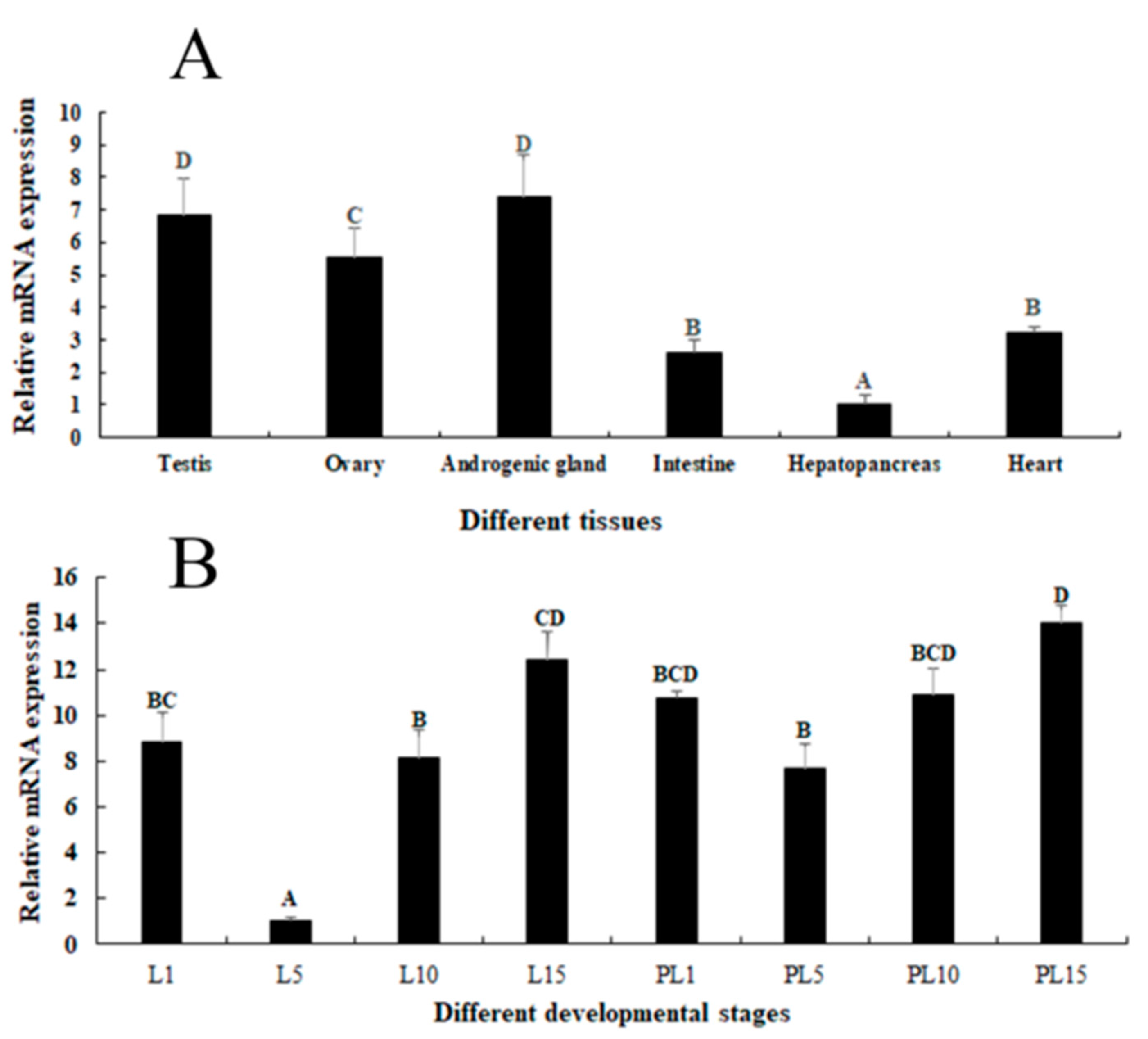
2.2. Expression Analysis in Different Tissues and Developmental Stages
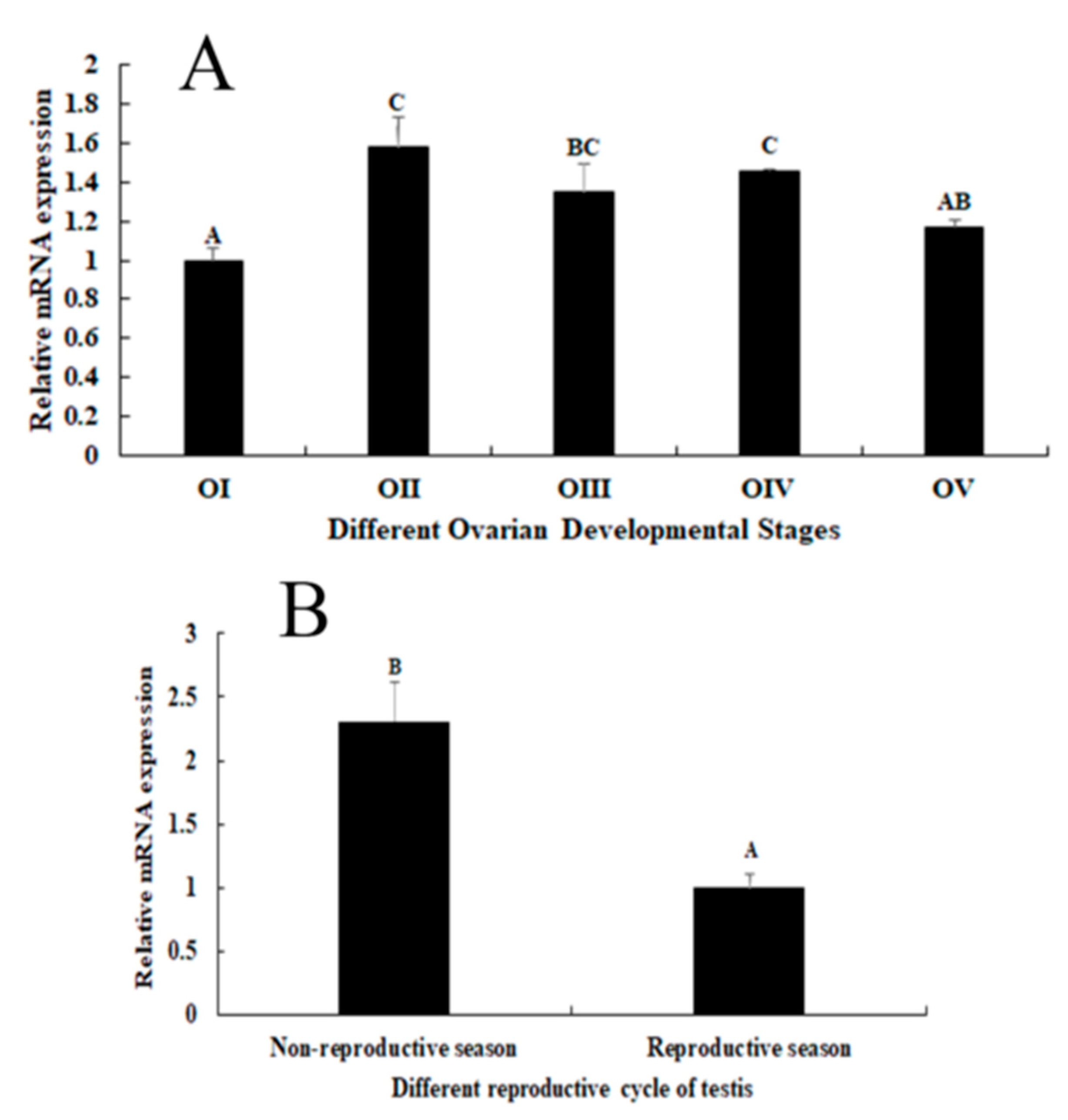
2.3. Expression Analysis during the Reproductive Cycle of Testes and Ovaries
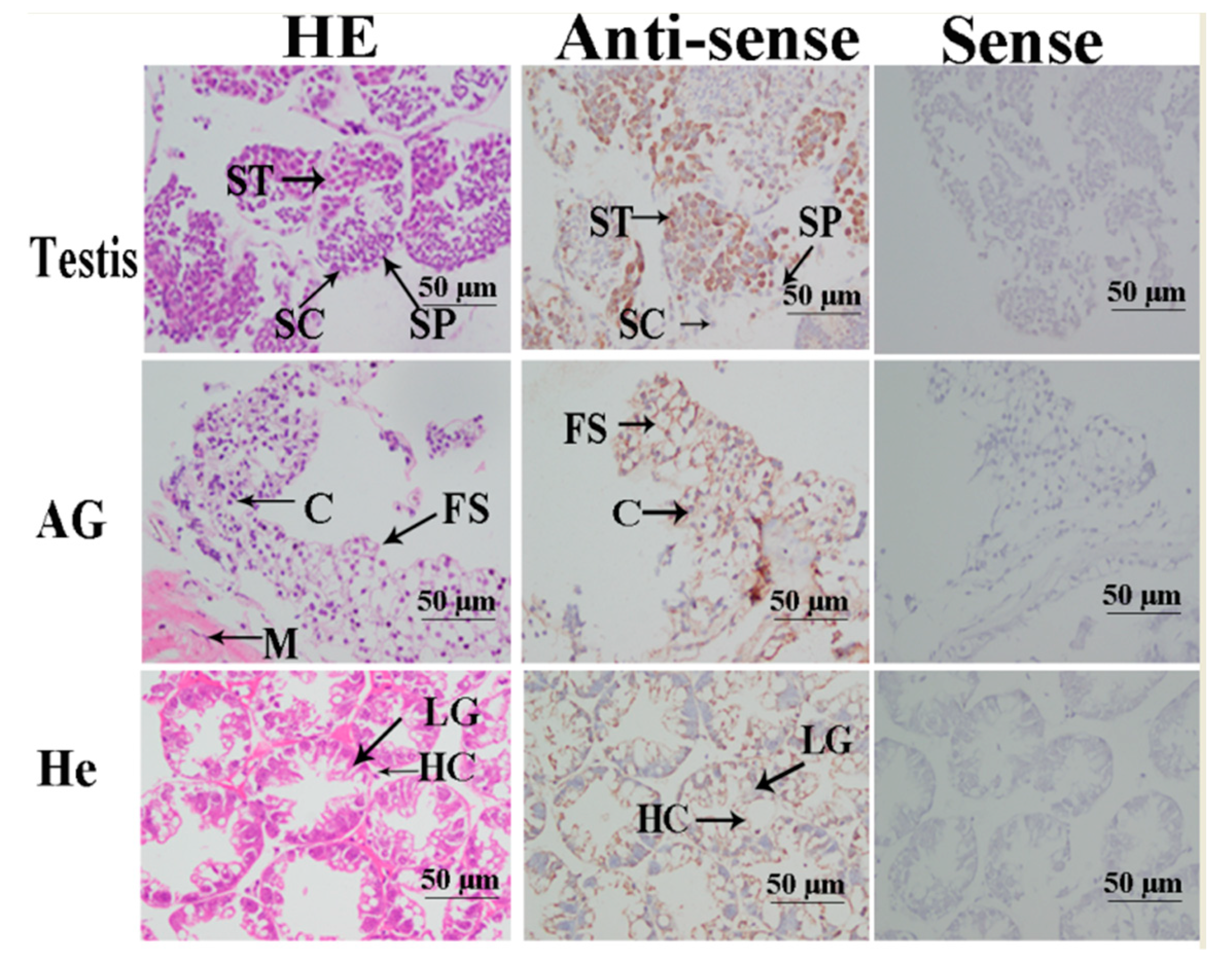
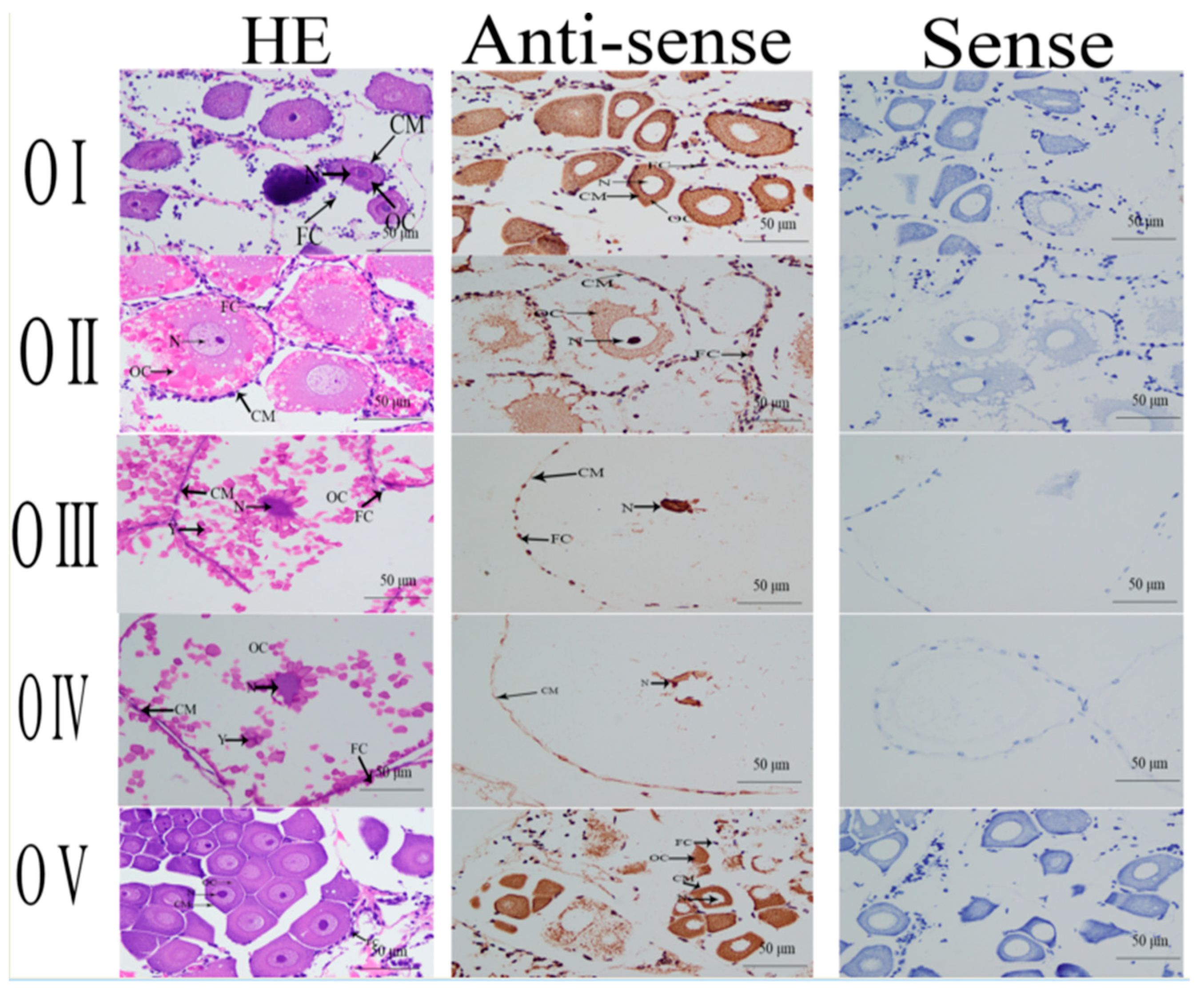
2.4. In situ Hybridization Analysis
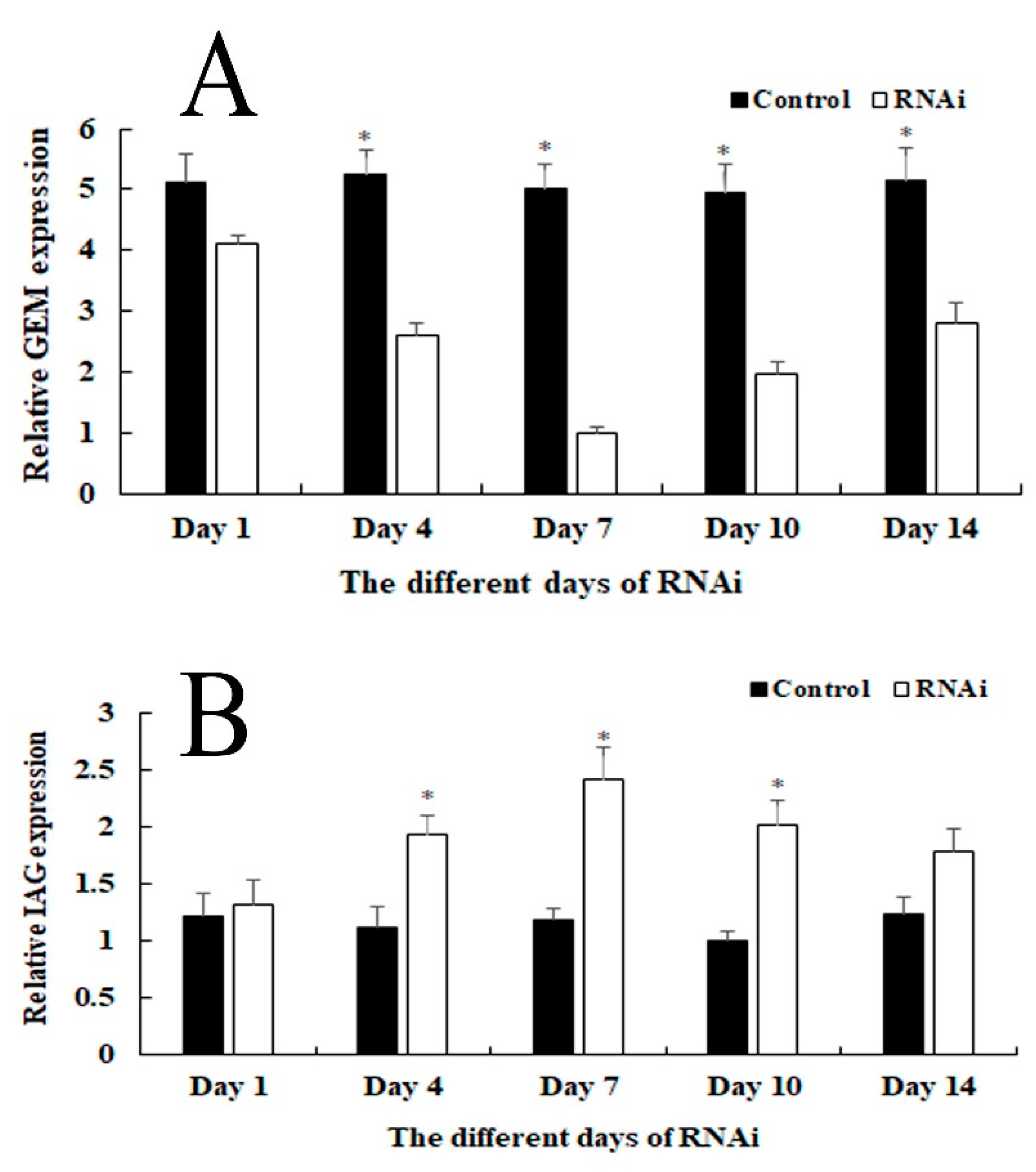
2.5. RNAi Analysis
3. Discussion
4. Materials and Methods
4.1. Sample Preparation
4.2. Rapid Amplification of cDNA Ends (RACE)
4.3. qPCR Analysis
4.4. In situ Hybridization of GEM
4.5. RNA Interference (RNAi) Analysis
4.6. Measurement of The Content of Testosterone
4.7. Statistical Analysis
Author Contributions
Funding
Conflicts of Interest
References
- Cai, Y.; Shokita, S. Report on a collection of freshwater shrimps (Crustacea: Decapoda: Caridea) from the Philippines, with descriptions of four new species. Raffles Bull. Zool. 2006, 54, 245–270. [Google Scholar]
- Grave, S.D.; Ghane, A. The establishment of the oriental river prawn, Macrobrachium nipponense (de Haan, 1849) in Anzali Lagoon, Iran. Aquat. Invasions 2006, 1, 204–208. [Google Scholar] [CrossRef]
- Salman, S.D.; Page, T.J.; Naser, M.D.; Yasser, A.G. The invasion of Macrobrachium nipponense (De Haan, 1849) (Caridea: Palaemonidae) into the southern Iraqi marshes. Aquat. Invasions 2006, 1, 109–115. [Google Scholar] [CrossRef]
- Ma, K.Y.; Feng, J.B.; Lin, J.Y.; Li, J.L. The complete mitochondrial genome of Macrobrachium nipponense. Gene 2011, 487, 160–165. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Miyake, S. Five species of the genus Macrobrachium (Crustacea, Decapoda, Palaemonidae) from Taiwan. Ohmu 1972, 3, 45–55. [Google Scholar]
- Bureau of Fisheries; Ministry of Agriculture. Ministry of Agriculture of the People’s Republic of China. In China Fishery Yearbook; China Agricultural Press: Beijing, China, 2016; Volume 236. [Google Scholar]
- Sagi, A.; Cohen, D.; Milner, Y. Effect of androgenic gland ablation on morphotypic differentiation and sexual characteristics of male freshwater prawns, Macrobrachium rosenbergii. Gen. Comp. Endocrinol. 1990, 77, 15–22. [Google Scholar] [CrossRef]
- Sagi, A.; Cohen, D.; Wax, Y. Production of Macrobrachium rosenbetgii in momosex population: Yield characteristes under intensive monoculture conditions in cages. Aquaculture 1986, 51, 265–275. [Google Scholar] [CrossRef]
- Sagi, A.; Cohen, D. Growth, maturation and progeny of sex-reversed Macrobrachium rosenbetgii males. World Aquacult. 1990, 21, 87–90. [Google Scholar]
- Jin, S.B.; Fu, H.T.; Zhou, Q.; Sun, S.M.; Jiang, S.F.; Xiong, Y.W.; Gong, Y.S.; Qiao, H.; Zhang, W.Y. Transcriptome Analysis of Androgenic Gland for Discovery of Novel Genes from the Oriental River Prawn, Macrobrachium nipponense, Using Illumina Hiseq 2000. PLoS ONE 2013, 8, e76840. [Google Scholar] [CrossRef]
- Jin, S.B.; Fu, H.T.; Jiang, S.F.; Xiong, Y.W.; Qiao, H.; Zhang, W.Y.; Gong, Y.S.; Wu, Y. Identification of androgenic gland microRNAs and their targeted genes for discovery of sex-related microRNAs in oriental river prawn, Macrobrachium nipponense. Gen. Mol. Res. 2015, 14, 18396–18406. [Google Scholar] [CrossRef]
- Jin, S.B.; Jiang, S.F.; Xiong, Y.W.; Qiao, H.; Sun, S.M.; Zhang, W.Y.; Gong, Y.S.; Fu, H.T. Molecular cloning of two tropomyosin family genes and expression analysis during development in oriental river prawn, Macrobrachium nipponense. Gene 2014, 546, 390–397. [Google Scholar] [CrossRef] [PubMed]
- Jin, S.B.; Fu, H.T.; Jiang, S.F.; Xiong, Y.W.; Sun, S.M.; Qiao, H.; Zhang, W.Y.; Gong, Y.S.; Wu, Y. Molecular Cloning, Expression, and in situ Hybridization Analysis of Forkhead Box Protein L2 during Development in Macrobrachium nipponense. J. World Aquacult. Soc. 2018, 49, 429–440. [Google Scholar] [CrossRef]
- Li, F.J.; Bai, H.K.; Xiong, Y.W.; Fu, H.T.; Jiang, S.F.; Jin, S.B.; Sun, S.M.; Qiao, H.; Zhang, W.Y. Molecular characterization of insulin-like androgenic gland hormone-binding protein gene from the oriental river prawn Macrobrachium nipponense and investigation of its transcriptional relationship with the insulin-like androgenic gland hormone gene. Gen. Comp. Endocrinol. 2015, 216, 152–160. [Google Scholar] [CrossRef] [PubMed]
- Ma, K.Y.; Li, J.L.; Qiu, G.F. Identification of putative regulatory region of insulin-like androgenic gland hormone gene (IAG) in the prawn Macrobrachium nipponense and proteins that interact with IAG by using yeast two-hybrid system. Gen. Comp. Endocrinol. 2016, 229, 112–118. [Google Scholar] [CrossRef] [PubMed]
- Jin, S.B.; Fu, H.T.; Sun, S.M.; Jiang, S.F.; Xiong, Y.W.; Gong, Y.S.; Qiao, H.; Zhang, W.Y.; Wu, Y. iTRAQ-based quantitative proteomic analysis of the androgenic glands of the oriental river prawn, Macrobrachium nipponense, during nonreproductive and reproductive seasons. Comp. Biochem. Physiol. D 2018, 26, 50–57. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J.P.; Liu, H.T.; Du, Y.G. O-GlcNAcylation and related research progress in biological function. Acta Microbiol. Sin. 2017, 57, 1141–1151. [Google Scholar]
- Zhang, Y.P.; Qiao, H.; Zhang, W.Y.; Sun, S.M.; Jiang, S.F.; Gong, Y.S.; Xiong, Y.W.; Jin, S.B.; Fu, H.T. Molecular cloning and expression analysis two sex-lethal homolog genesduring development in Oriental river prawn, Macrobrachium nipponense. Genet. Mol. Res. 2013, 12, 4698–4711. [Google Scholar] [CrossRef]
- Zhang, Y.P.; Fu, H.T.; Qiao, H.; Jin, S.B.; Gong, Y.S.; Jiang, S.F.; Xiong, Y.W. cDNA cloning, characterization and expression analysis of a transformer-2 gene in the oriental river prawn, Macrobrachium nipponense. J. World Aquacult. Soc. 2013, 44, 338–349. [Google Scholar] [CrossRef]
- Jin, S.B.; Zhang, Y.; Guan, H.H.; Fu, H.T.; Jiang, S.F.; Xiong, Y.W.; Qiao, H.; Zhang, W.Y.; Gong, Y.S.; Wu, Y. Histological observation of gonadal development during post-larva in oriental river prawn, Macrobrachium nipponense. Chin. J. Fish. 2016, 29, 11–16. [Google Scholar]
- Tautz, D.; Pfeifle, C. A non-radioactive in situ hybridization method for the localization of specific RNAs in Drosophila embryos reveals translational control of the segmentation gene hunchback. Chromosoma 1989, 98, 81–85. [Google Scholar] [CrossRef]
- Li, F.; Qiao, H.; Fu, H.T.; Sun, S.M.; Zhang, W.Y.; Jin, S.B.; Jiang, S.F.; Gong, Y.S.; Xiong, Y.W.; Wu, Y.; et al. Identification and characterization of opsin gene and its role in ovarian maturation in the oriental river prawn Macrobrachium nipponense. Comp. Biochem. Physiol. B 2018, 218, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Jones, H.D. Wheat transformation: Current technology and applications to grain development and composition. J. Cereal Sci. 2005, 41, 137–147. [Google Scholar] [CrossRef]
- Kusaba, M. RNA interference in crop plants. Curr. Opin. Biol. 2004, 15, 139–143. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.A.; Roeszler, K.N.; Ohnesorg, T.; Cummins, D.M.; Farlie, P.G.; Doran, T.J.; Sinclair, A.H. The avian Z-linked gene DMRT1 is required for male sex determination in the chicken. Nature 2009, 461, 267–271. [Google Scholar] [CrossRef] [PubMed]
- Rosen, O.; Manor, R.; Weil, S.; Gafni, O.; Linial, A.; Aflalo, E.D.; Ventura, T.; Sagi, A. A sexual shift induced by silencing of a single insulin-like gene in crayfish: Ovarian upregulation and testicular degeneration. PLoS ONE 2010, 5, e15281. [Google Scholar] [CrossRef] [PubMed]
- Ventura, T.; Manor, R.; Aflalo, E.D.; Weil, S.; Raviv, S.; Glazer, L.; Sagi, A. Temporal silencing of an androgenic gland-specific insulin-like gene affecting phenotypical gender differences and spermatogenesis. Endocrinology 2009, 150, 1278–1286. [Google Scholar] [CrossRef] [PubMed]
- Ventura, T.; Manor, R.; Aflalo, E.D.; Weil, S.; Khalaila, I.; Rosen, O.; Sagi, A. Expression of an androgenic gland-specific insulin-like peptide during the course of prawn sexual and morphotypic differentiation. ISRN Endocrinol. 2011, 2011, 476283. [Google Scholar] [CrossRef]
- Ventura, T.; Manor, R.; Aflalo, E.D.; Weil, S.; Rosen, O.; Sagi, A. Timing sexual differentiation: Full functional sex reversal achieved through silencing of a single insulin-like gene in the prawn, Macrobrachium rosenbergii. Biol. Reprod. 2012, 86, 90. [Google Scholar] [CrossRef]
- Södergård, R.; Bäckström, T.; Shanbhag, V.; Carstensen, H. Calculation of free and bound fractions of testosterone and estradiol-17 beta to human plasma proteins at body temperature. J. Steroid Biochem. 1982, 16, 801–810. [Google Scholar] [CrossRef]
- Page, S.T.; Amory, J.K.; Bowman, F.D.; Anawalt, B.D.; Matsumoto, A.M.; Bremner, W.J.; Tenover, J.L. Exogenous testosterone (T) alone or with finasteride increases physical performance, grip strength, and lean body mass in older men with low serum T. J. Clin. Endocrinol. Metab. 2005, 90, 1502–1510. [Google Scholar] [CrossRef]
- Steidle, C.; Schwartz, S.; Jacoby, K.; Sebree, T.; Smith, T.; Bachand, R. North American AA2500 T Gel Study Group. AA2500 testosterone gel normalizes androgen levels in aging males with improvements in body composition and sexual function. J. Clin. Endocrinol. Metab. 2003, 88, 2673–2681. [Google Scholar] [CrossRef] [PubMed]
- Behre, H.M.; Abshagen, K.; Oettel, M.; Hübler, D.; Nieschlag, E. Intramuscular injection of testosterone undecanoate for the treatment of male hypogonadism: Phase I studies. Eur. J. Endocrinol. 1999, 140, 414–419. [Google Scholar] [CrossRef] [PubMed]
- Cao, L.Q.; Tao, Y.G.; Tan, H.Z. Studies on unisexualization in Red Tipapia. J. Fish. China 1994, 18, 75–80. [Google Scholar]
- Lou, Y.D.; Song, T.F.; Wang, Y.M.; Wei, H.; Wu, M.C.; Xu, H.M.; Wang, Z.Q. Studies on sex control in allogynogenetic Crucian Carp. J. Fish. China 1994, 3, 169–176. [Google Scholar]
- Zhang, P.Y.; Hu, W.; Wang, Y.P.; Zhu, Z.Y. Methyltestosterone Induction of Sex Reversal in Allogynogenetic Crucian Carp. Hereditas 2000, 22, 25–27. [Google Scholar]
- Li, G.L.; Liu, X.C.; Lin, H.R. Effects of 17α-methyltestosterone on sex reversal in red-spotted grouper, Epinephelus akaara. J. Fish. China 2006, 30, 145–150. [Google Scholar]
- Kang, X.J.; Liu, Z.M.; Zhou, K.X.; Wang, S.A. Primary study of exogenous steroid hormone effect on the testis development of the Chinese Mitten-Handed Crab, Eriocheir sinensis (Crustcea, Decapoda). Donghai Mar. Sci. 1998, 16, 39–44. [Google Scholar]
- Qiao, H.; Xiong, Y.W.; Zhang, W.Y.; Fu, H.T.; Jiang, S.F.; Sun, S.M.; Bai, H.K.; Jin, S.B.; Gong, Y.S. Characterization, expression, and function analysis of gonad-inhibiting hormone in Oriental River prawn, Macrobrachium nipponense and its induced expression by temperature. Comp. Biochem. Physiol. A 2015, 185, 1–8. [Google Scholar] [CrossRef]
- Hu, Y.N.; Fu, H.T.; Qiao, H.; Sun, S.M.; Zhang, W.Y.; Jin, S.B.; Jiang, S.F.; Gong, Y.S.; Xiong, Y.W.; Wu, Y. Validation and evaluation of reference genes for Quantitative real-time PCR in Macrobrachium nipponense. Int. J. Mol. Sci. 2018, 19, 2258. [Google Scholar] [CrossRef]
- Jiang, F.W.; Fu, H.T.; Qiao, H.; Zhang, W.Y.; Jiang, S.F.; Xiong, Y.W.; Sun, S.M.; Gong, Y.S.; Jin, S.B. The RNA Interference Regularity of Transformer-2 Gene of Oriental River Prawn Macrobrachium nipponense. Chin. Agric. Sci. Bull. 2014, 30, 32–37. [Google Scholar]


| Species | Accession Number |
|---|---|
| Macrobrachium nipponense | |
| Penaeus vannamei | XP_027229945.1 |
| Acanthaster planci | XP_022103928.1 |
| Xenopus laevis | XP_018087535.1 |
| Stylophora pistillata | XP_022804116.1 |
| Odobenus rosmarus divergens | XP_004399834.1 |
| Centruroides sculpturatus | XP_023219163.1 |
| Biomphalaria glabrata | XP_013071285.1 |
| Aplysia californica | XP_005105416.1 |
| Mizuhopecten yessoensis | XP_021367489.1 |
| Xenopus tropicalis | NP_001096228.1 |
| Xenopus laevis | NP_001087945.2 |
| Mus musculus | NP_079932.2 |
| Mus caroli | XP_021034811.1 |
| Primer Name | Nucleotide Sequence (5′→3′) | Purpose |
|---|---|---|
| GEM-3GSP1 | AGTATGGGAGGCTCGTCAGT | First forward primer for GEM 3′ RACE |
| GEM-3GSP2 | ATGCCCAGTGGCTAGAAGTG | Second forward primer for GEM 3′ RACE |
| GEM-5GSP1 | AATGCTCCAGTTCCTTTGCCT | First reverse primer for GEM 5′ RACE |
| GEM-5GSP2 | CCTGTCGGATATGCTCCGTC | Second reverse primer for GEM 5′ RACE |
| 3′ RACE OUT | TACCGTCGTTCCACTAGTGATTT | RVS first primer for 3′ RACE |
| 3′ RACE IN | CGCGGATCCTCCACTAGTGATTTCACTATAGG | RVS second primer for 3′ RACE |
| 5′ RACE OUT | CATGGCTACATGCTGACAGCCTA | FWD first primer for 5′ RACE |
| 5′ RACE IN | CGCGGATCCACAGCCTACTGATGATCAGTCGATG | FWD second primer for 5′ RACE |
| GEM-RTF | ATGCCCAGTGGCTAGAAGTG | FWD primer for GEM expression |
| GEM-RTR | GCAGAATCCAGCGATGCTCT | RVS primer for GEM expression |
| EIF-F | CATGGATGTACCTGTGGTGAAAC | FWD primer for β-actin expression |
| EIF-R | CTGTCAGCAGAAGGTCCTCATTA | RVS primer for β-actin expression |
| GEM anti-sense Probe | GCACTGACGAGCCTCCCATACTACATGCTGAAGATAC | Probe for GEM in situ hybridization analysis |
| GEM sense Probe | GTATCTTCAGCATGTAGTATGGGAGGCTCGTCAGTGC | Probe for GEM in situ hybridization analysis |
| GEM RNAi-F | TAATACGACTCACTATAGGGGATATGCTCCGTCTGCACAA | FWD primer for RNAi analysis |
| GEM RNAi-R | TAATACGACTCACTATAGGGCGAATCTCATCCATCAGCAA | RVS primer for RNAi analysis |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jin, S.; Hu, Y.; Fu, H.; Jiang, S.; Xiong, Y.; Qiao, H.; Zhang, W.; Gong, Y.; Wu, Y. Potential Functions of Gem-Associated Protein 2-Like Isoform X1 in the Oriental River Prawn Macrobrachium nipponense: Cloning, qPCR, In Situ Hybridization, and RNAi Analysis. Int. J. Mol. Sci. 2019, 20, 3995. https://doi.org/10.3390/ijms20163995
Jin S, Hu Y, Fu H, Jiang S, Xiong Y, Qiao H, Zhang W, Gong Y, Wu Y. Potential Functions of Gem-Associated Protein 2-Like Isoform X1 in the Oriental River Prawn Macrobrachium nipponense: Cloning, qPCR, In Situ Hybridization, and RNAi Analysis. International Journal of Molecular Sciences. 2019; 20(16):3995. https://doi.org/10.3390/ijms20163995
Chicago/Turabian StyleJin, Shubo, Yuning Hu, Hongtuo Fu, Sufei Jiang, Yiwei Xiong, Hui Qiao, Wenyi Zhang, Yongsheng Gong, and Yan Wu. 2019. "Potential Functions of Gem-Associated Protein 2-Like Isoform X1 in the Oriental River Prawn Macrobrachium nipponense: Cloning, qPCR, In Situ Hybridization, and RNAi Analysis" International Journal of Molecular Sciences 20, no. 16: 3995. https://doi.org/10.3390/ijms20163995
APA StyleJin, S., Hu, Y., Fu, H., Jiang, S., Xiong, Y., Qiao, H., Zhang, W., Gong, Y., & Wu, Y. (2019). Potential Functions of Gem-Associated Protein 2-Like Isoform X1 in the Oriental River Prawn Macrobrachium nipponense: Cloning, qPCR, In Situ Hybridization, and RNAi Analysis. International Journal of Molecular Sciences, 20(16), 3995. https://doi.org/10.3390/ijms20163995







