Expression of Osteoblast-Specific Factor 2 (OSF-2, Periostin) Is Associated with Drug Resistance in Ovarian Cancer Cell Lines
Abstract
1. Introduction
2. Results
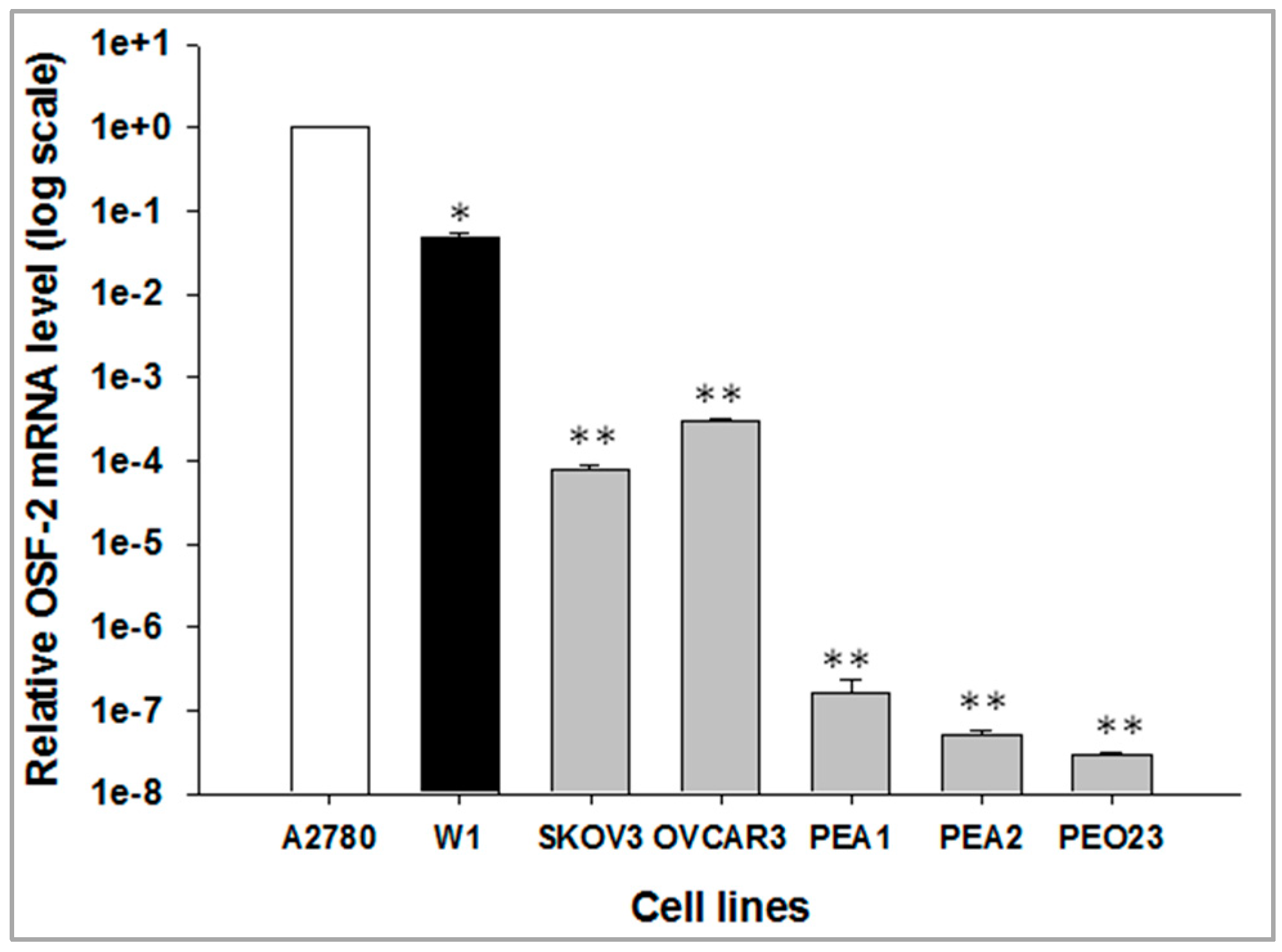
2.1. OSF-2 Gene Expression in Different Ovarian Cancer Cell Lines
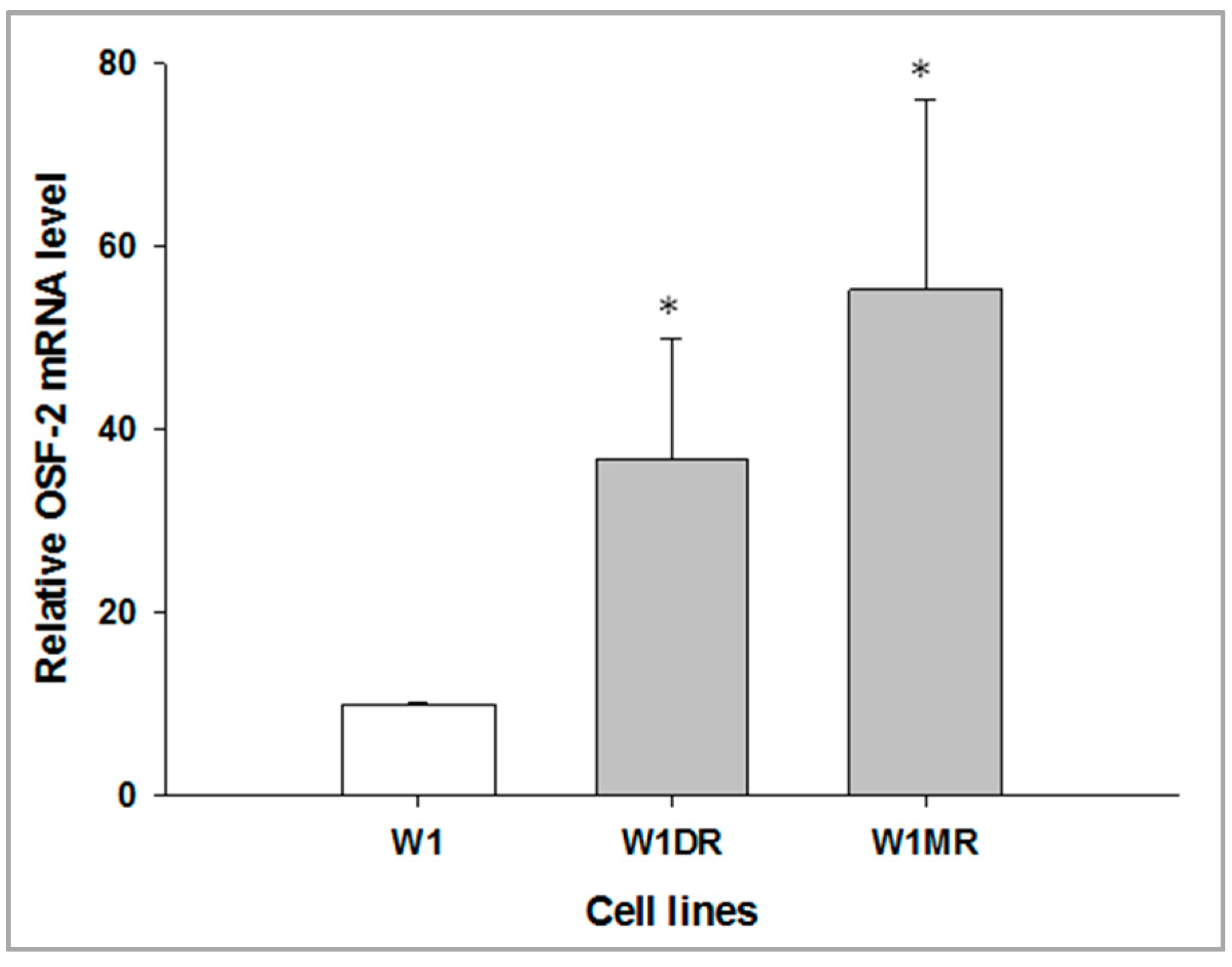
2.2. OSF-2 Gene Expression in Drug-Resistant Ovarian Cancer Cell Lines
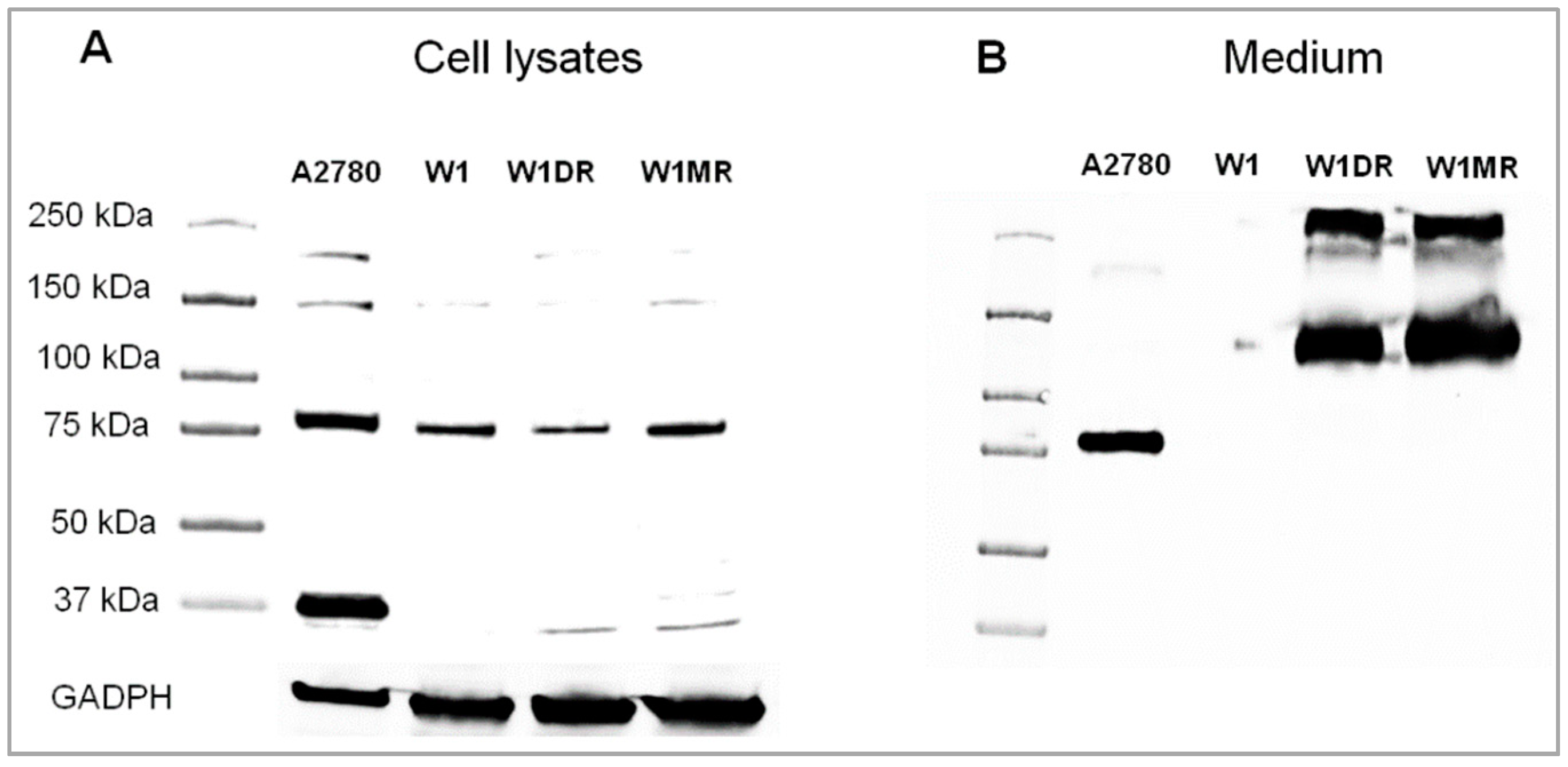
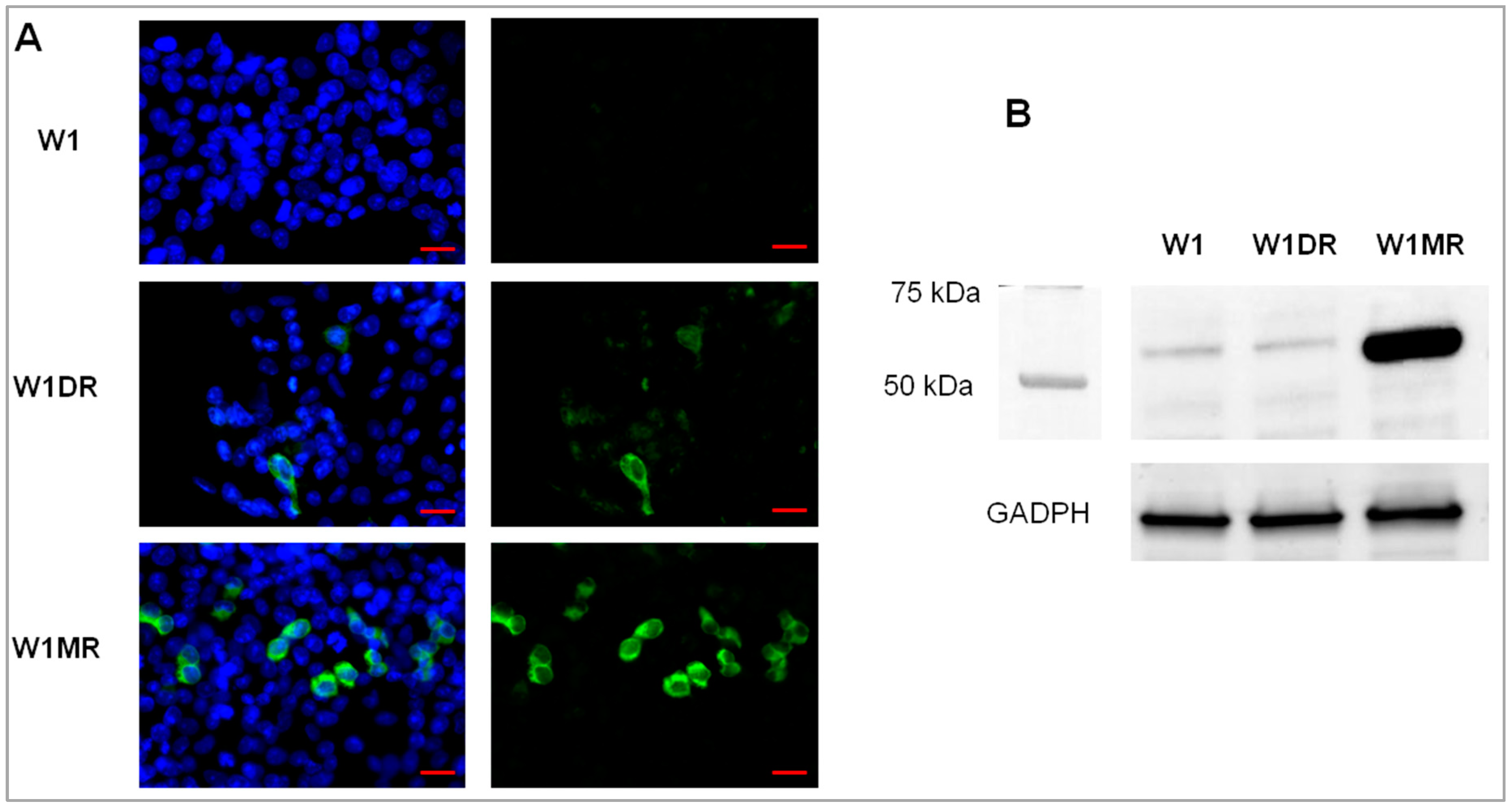
2.3. OSF-2 Protein Expression in Drug-Resistant Ovarian Cancer Cell Lines
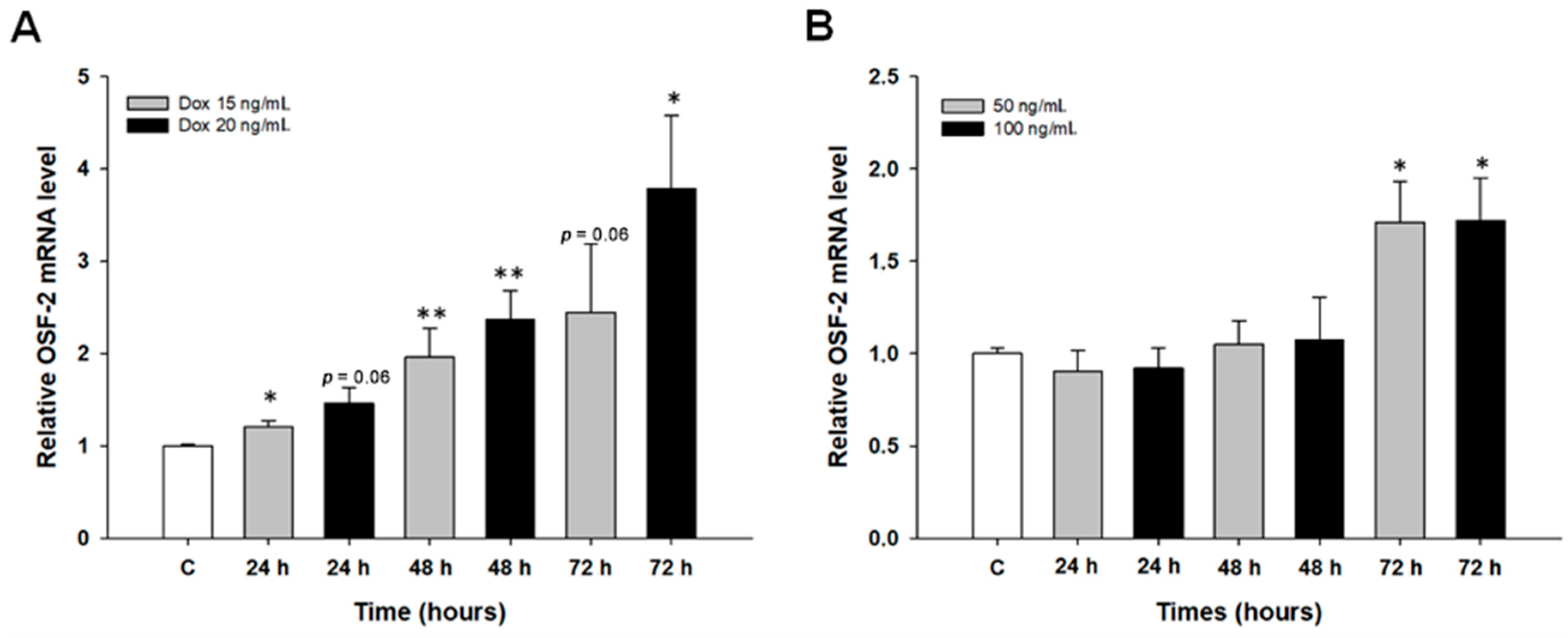
2.4. Early Response to DOX and MTX Treatment in Ovarian Cancer Cell Lines
2.5. ALDH1A1 Expression in Drug-Sensitive and Drug-Resistant Cell Lines
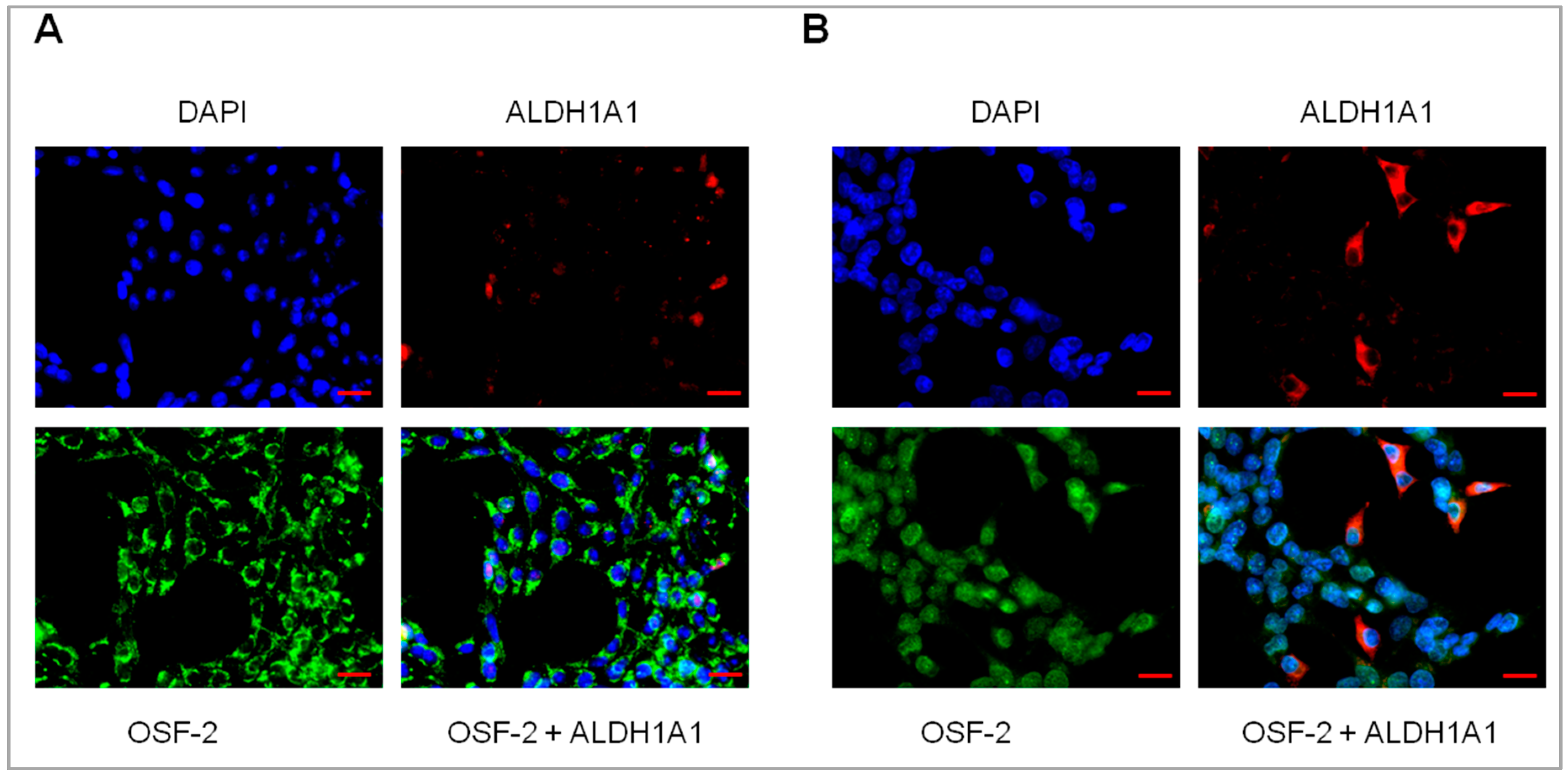
2.6. Coexpression of ALDH1A1 and OSF-2 in Drug-Resistant Cell Lines
3. Discussion
4. Materials and Methods
4.1. Reagents and Antibodies
4.2. Cell Lines and Cell Culture
4.3. QPCR Gene Expression Analysis
4.4. Western Blot Analysis
4.5. Immunofluorescence Analysis
4.6. DOX and MTX Response in Time-Course Experiment
4.7. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Housman, G.; Byler, S.; Heerboth, S.; Lapinska, K.; Longacre, M.; Snyder, N.; Sarkar, S. Drug resistance in cancer: An overview. Cancers 2014, 6, 1769–1792. [Google Scholar] [CrossRef] [PubMed]
- Freimund, A.E.; Beach, J.A.; Christie, E.L.; Bowtell, D.D.L. Mechanisms of Drug Resistance in High-Grade Serous Ovarian Cancer. Hematol. Oncol. Clin. North. Am. 2018, 32, 983–996. [Google Scholar] [CrossRef]
- Yeldag, G.; Rice, A.; Del Río Hernández, A. Chemoresistance and the Self-Maintaining Tumor Microenvironment. Cancers 2018, 10, 471. [Google Scholar] [CrossRef]
- Senthebane, D.A.; Rowe, A.; Thomford, N.E.; Shipanga, H.; Munro, D.; Mazeedi, M.A.M.A.; Almazyadi, H.A.M.; Kallmeyer, K.; Dandara, C.; Pepper, M.S.; et al. The Role of Tumor Microenvironment in Chemoresistance: To Survive, Keep Your Enemies Closer. Int. J. Mol. Sci. 2017, 18, 1586. [Google Scholar] [CrossRef]
- Tannock, I.F.; Lee, C.M.; Tunggal, J.K.; Cowan, D.S.; Egorin, M.J. Limited penetration of anticancer drugs through tumor tissue: A potential cause of resistance of solid tumors to chemotherapy. Clin. Cancer Res. 2002, 8, 878–884. [Google Scholar] [PubMed]
- Di Paolo, A.; Bocci, G. Drug distribution in tumors: Mechanisms, role in drug resistance, and methods for modification. Curr. Oncol. Rep. 2007, 9, 109–114. [Google Scholar] [CrossRef] [PubMed]
- Sethi, T.; Rintoul, R.C.; Moore, S.M.; MacKinnon, A.C.; Salter, D.; Choo, C.; Chilvers, E.R.; Dransfield, I.; Donnelly, S.C.; Strieter, R.; et al. Extracellular matrix proteins protect small cell lung cancer cells against apoptosis: A mechanism for small cell lung cancer growth and drug resistance in vivo. Nat. Med. 1999, 5, 662–668. [Google Scholar] [CrossRef] [PubMed]
- Sherman-Baust, C.A.; Weeraratna, A.T.; Rangel, L.B.; Pizer, E.S.; Cho, K.R.; Schwartz, D.R.; Shock, T.; Morin, P.J. Remodeling of the extracellular matrix through overexpression of collagen VI contributes to cisplatin resistance in ovarian cancer cells. Cancer Cell. 2003, 3, 377–386. [Google Scholar] [CrossRef]
- Işeri, O.D.; Kars, M.D.; Arpaci, F.; Gündüz, U. Gene expression analysis of drug-resistant MCF-7 cells: Implications for relation to extracellular matrix proteins. Cancer Chemother. Pharmacol. 2010, 65, 447–455. [Google Scholar] [CrossRef] [PubMed]
- Klejewski, A.; Sterzyńska, K.; Wojtowicz, K.; Świerczewska, M.; Partyka, M.; Brązert, M.; Nowicki, M.; Zabel, M.; Januchowski, R. The significance of lumican expression in ovarian cancer drug-resistant cell lines. Oncotarget 2017, 8, 74466–74478. [Google Scholar] [CrossRef] [PubMed]
- Sterzyńska, K.; Klejewski, A.; Świerczewska, M.; Nowicki, M.; Brązert, J.; Januchowski, R. Myotilin, a New Topotecan Resistant Protein in Ovarian Cancer Cell Lines. J. Cancer 2018, 9, 4413–4421. [Google Scholar] [CrossRef] [PubMed]
- Sterzyńska, K.; Klejewski, A.; Wojtowicz, K.; Świerczewska, M.; Andrzejewska, M.; Rusek, D.; Sobkowski, M.; Kędzia, W.; Brązert, J.; Nowicki, M.; et al. The Role of Matrix Gla Protein (MGP) Expression in Paclitaxel and Topotecan Resistant Ovarian Cancer Cell Lines. Int. J. Mol. Sci. 2018, 19, 2901. [Google Scholar] [CrossRef]
- Sterzyńska, K.; Klejewski, A.; Wojtowicz, K.; Świerczewska, M.; Nowacka, M.; Kaźmierczak, D.; Andrzejewska, M.; Rusek, D.; Brązert, M.; Brązert, J.; et al. Mutual Expression of ALDH1A1, LOX, and Collagens in Ovarian Cancer Cell Lines as Combined CSCs- and ECM-Related Models of Drug Resistance Development. Int. J. Mol. Sci. 2018, 20, 54. [Google Scholar] [CrossRef] [PubMed]
- Ratajczak-Wielgomas, K.; Dziegiel, P. The role of periostin in neoplastic processes. Folia Histochem. Cytobiol. 2015, 53, 120–132. [Google Scholar] [CrossRef] [PubMed]
- Sehra, S.; Yao, W.; Nguyen, E.T.; Ahyi, A.N.; Tuana, F.M.; Ahlfeld, S.K.; Snider, P.; Tepper, R.S.; Petrache, I.; Conway, S.J.; et al. Periostin regulates goblet cell metaplasia in a model of allergic airway inflammation. J. Immunol. 2011, 186, 4959–4966. [Google Scholar] [CrossRef] [PubMed]
- Bornstein, P. Matricellular proteins: An overview. J. Cell Commun. Signal. 2009, 3, 163–165. [Google Scholar] [CrossRef]
- Kii, I.; Nishiyama, T.; Li, M.; Matsumoto, K.; Saito, M.; Amizuka, N.; Kudo, A. Incorporation of tenascin-C into the extracellular matrix by periostin underlies an extracellular meshwork architecture. J. Biol. Chem. 2010, 285, 2028–2039. [Google Scholar] [CrossRef]
- Kii, I.; Ito, H. Periostin and its interacting proteins in the construction of extracellular architectures. Cell Mol. Life Sci. 2017, 74, 4269–4277. [Google Scholar] [CrossRef]
- Norris, R.A.; Damon, B.; Mironov, V.; Kasyanov, V.; Ramamurthi, A.; Moreno-Rodriguez, R.; Trusk, T.; Potts, J.D.; Goodwin, R.L.; Davis, J.; et al. Periostin regulates collagen fibrillogenesis and the biomechanical properties of connective tissues. J. Cell Biochem. 2007, 101, 695–711. [Google Scholar] [CrossRef]
- Kim, B.Y.; Olzmann, J.A.; Choi, S.I.; Ahn, S.Y.; Kim, T.I.; Cho, H.S.; Suh, H.; Kim, E.K. Corneal dystrophy-associated R124H mutation disrupts TGFBI interaction with Periostin and causes mislocalization to the lysosome. J. Biol. Chem. 2009, 284, 19580–19591. [Google Scholar] [CrossRef]
- Gillan, L.; Matei, D.; Fishman, D.A.; Gerbin, C.S.; Karlan, B.Y.; Chang, D.D. Periostin secreted by epithelial ovarian carcinoma is a ligand for alpha(V)beta(3) and alpha(V)beta(5) integrins and promotes cell motility. Cancer Res. 2002, 62, 5358–5364. [Google Scholar] [PubMed]
- Horiuchi, K.; Amizuka, N.; Takeshita, S.; Takamatsu, H.; Katsuura, M.; Ozawa, H.; Toyama, Y.; Bonewald, L.F.; Kudo, A. Identification and characterization of a novel protein, periostin, with restricted expression to periosteum and periodontal ligament and increased expression by transforming growth factor beta. J. Bone Min. Res. 1999, 14, 1239–1249. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, B.; del Monte, F.; Hajjar, R.J.; Chang, Y.S.; Lebeche, D.; Arab, S.; Keating, M.T. Periostin induces proliferation of differentiated cardiomyocytes and promotes cardiac repair. Nat. Med. 2007, 13, 962–969. [Google Scholar] [CrossRef] [PubMed]
- Shao, R.; Bao, S.; Bai, X.; Blanchette, C.; Anderson, R.M.; Dang, T.; Gishizky, M.L.; Marks, J.R.; Wang, X.F. Acquired expression of periostin by human breast cancers promotes tumor angiogenesis through up-regulation of vascular endothelial growth factor receptor 2 expression. Mol. Cell Biol. 2004, 24, 3992–4003. [Google Scholar] [CrossRef]
- Puglisi, F.; Puppin, C.; Pegolo, E.; Andreetta, C.; Pascoletti, G.; D’Aurizio, F.; Pandolfi, M.; Fasola, G.; Piga, A.; Damante, G.; et al. Expression of periostin in human breast cancer. J. Clin. Pathol. 2008, 61, 494–498. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, G.; Li, J.; Tao, Q.; Tang, W. The expression analysis of periostin in human breast cancer. J. Surg. Res. 2010, 160, 102–106. [Google Scholar] [CrossRef] [PubMed]
- Morra, L.; Rechsteiner, M.; Casagrande, S.; von Teichman, A.; Schraml, P.; Moch, H.; Soltermann, A. Characterization of periostin isoform pattern in non-small cell lung cancer. Lung Cancer. 2012, 76, 183–190. [Google Scholar] [CrossRef] [PubMed]
- Takanami, I.; Abiko, T.; Koizumi, S. Expression of periostin in patients with non-small cell lung cancer: Correlation with angiogenesis and lymphangiogenesis. Int. J. Biol. Markers. 2008, 23, 182–186. [Google Scholar] [CrossRef]
- Bao, S.; Ouyang, G.; Bai, X.; Huang, Z.; Ma, C.; Liu, M.; Shao, R.; Anderson, R.M.; Rich, J.N.; Wang, X.F. Periostin potently promotes metastatic growth of colon cancer by augmenting cell survival via the Akt/PKB pathway. Cancer Cell. 2004, 5, 329–339. [Google Scholar] [CrossRef]
- Cheon, D.J.; Tong, Y.; Sim, M.S.; Dering, J.; Berel, D.; Cui, X.; Lester, J.; Beach, J.A.; Tighiouart, M.; Walts, A.E.; et al. A collagen-remodeling gene signature regulated by TGF-β signaling is associated with metastasis and poor survival in serous ovarian cancer. Clin. Cancer Res. 2014, 20, 711–723. [Google Scholar] [CrossRef]
- Zhu, M.; Fejzo, M.S.; Anderson, L.; Dering, J.; Ginther, C.; Ramos, L.; Gasson, J.C.; Karlan, B.Y.; Slamon, D.J. Periostin promotes ovarian cancer angiogenesis and metastasis. Gynecol. Oncol. 2010, 119, 337–344. [Google Scholar] [CrossRef] [PubMed]
- Ryner, L.; Guan, Y.; Firestein, R.; Xiao, Y.; Choi, Y.; Rabe, C.; Lu, S.; Fuentes, E.; Huw, L.Y.; Lackner, M.R.; et al. Upregulation of Periostin and Reactive Stroma Is Associated with Primary Chemoresistance and Predicts Clinical Outcomes in Epithelial Ovarian Cancer. Clin Cancer Res. 2015, 21, 2941–2951. [Google Scholar] [CrossRef] [PubMed]
- Sung, P.L.; Jan, Y.H.; Lin, S.C.; Huang, C.C.; Lin, H.; Wen, K.C.; Chao, K.C.; Lai, C.R.; Wang, P.H.; Chuang, C.M.; et al. Periostin in tumor microenvironment is associated with poor prognosis and platinum resistance in epithelial ovarian carcinoma. Oncotarge 2016, 7, 4036–4047. [Google Scholar] [CrossRef]
- Hennessy, B.T.; Coleman, R.L.; Markman, M. Ovarian cancer. Lancet. 2009, 374, 1371–1382. [Google Scholar] [CrossRef]
- Mutch, D.G.; Orlando, M.; Goss, T.; Teneriello, M.G.; Gordon, A.N.; McMeekin, S.D.; Wang, Y.; Scribner, D.R., Jr.; Marciniack, M.; Naumann, R.W.; et al. Randomized phase III trial of gemcitabine compared with pegylated liposomal doxorubicin in patients with platinum-resistant ovarian cancer. J. Clin. Oncol. 2007, 25, 2811–2818. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.C. DNA topoisomerases. Annu. Rev. Biochem. 1996, 65, 635–692. [Google Scholar] [CrossRef] [PubMed]
- Lindgren, M.; Rosenthal-Aizman, K.; Saar, K.; Eiríksdóttir, E.; Jiang, Y.; Sassian, M.; Ostlund, P.; Hällbrink, M.; Langel, U. Overcoming methotrexate resistance in breast cancer tumour cells by the use of a new cell-penetrating peptide. Biochem. Pharm. 2006, 71, 416–425. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, J.I.; Haber, M.; Henderson, M.J.; Norris, M.D. ABC transporters in cancer: More than just drug efflux pumps. Nat. Rev. Cancer. 2010, 10, 147–156. [Google Scholar] [CrossRef] [PubMed]
- Januchowski, R.; Zawierucha, P.; Ruciński, M.; Zabel, M. Microarray-based detection and expression analysis of extracellular matrix proteins in drug-resistant ovarian cancer cell lines. Oncol. Rep. 2014, 32, 1981–1990. [Google Scholar] [CrossRef] [PubMed]
- Januchowski, R.; Wojtowicz, K.; Sujka-Kordowska, P.; Andrzejewska, M.; Zabel, M. MDR gene expression analysis of six drug-resistant ovarian cancer cell lines. Biomed. Res. Int. 2013, 241763. [Google Scholar] [CrossRef] [PubMed]
- Januchowski, R.; Zawierucha, P.; Ruciński, M.; Andrzejewska, M.; Wojtowicz, K.; Nowicki, M.; Zabel, M. Drug transporter expression profiling in chemoresistant variants of the A2780 ovarian cancer cell line. Biomed. Pharm. 2014, 68, 447–453. [Google Scholar] [CrossRef]
- Januchowski, R.; Wojtowicz, K.; Sterzyńska, K.; Sosińska, P.; Andrzejewska, M.; Zawierucha, P.; Nowicki, M.; Zabel, M. Inhibition of ALDH1A1 activity decreases expression of drug transporters and reduces chemotherapy resistance in ovarian cancer cell lines. Int. J. Biochem. Cell Biol. 2016, 78, 248–259. [Google Scholar] [CrossRef]
- Januchowski, R.; Świerczewska, M.; Sterzyńska, K.; Wojtowicz, K.; Nowicki, M.; Zabel, M. Increased expression of several collagen genes is associated with drug resistance in ovarian cancer cell lines. J. Cancer 2016, 25, 1295–1310. [Google Scholar] [CrossRef] [PubMed]
- Correia, A.L.; Bissell, M.J. The tumor microenvironment is a dominant force in multidrug resistance. Drug Resist. Updat. 2012, 15, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Morin, P.J. Drug resistance and the microenvironment: Nature and nurture. Drug Resist. Updat. 2003, 6, 169–172. [Google Scholar] [CrossRef]
- Januchowski, R.; Zawierucha, P.; Ruciński, M.; Nowicki, M.; Zabel, M. Extracellular matrix proteins expression profiling in chemoresistant variants of the A2780 ovarian cancer cell line. Biomed. Res. Int. 2014, 365867. [Google Scholar] [CrossRef]
- Sidhu, S.S.; Yuan, S.; Innes, A.L.; Kerr, S.; Woodruff, P.G.; Hou, L.; Muller, S.J.; Fahy, J.V. Roles of epithelial cell-derived periostin in TGF-beta activation, collagen production, and collagen gel elasticity in asthma. Proc. Natl. Acad. Sci. USA 2010, 107, 14170–14175. [Google Scholar] [CrossRef]
- Kudo, A.; Kii, I. Periostin function in communication with extracellular matrices. J. Cell Commun. Signal. 2018, 12, 301–308. [Google Scholar] [CrossRef]
- Xu, D.; Xu, H.; Ren, Y.; Liu, C.; Wang, X.; Zhang, H.; Lu, P. Cancer Stem Cell-Related Gene Periostin: A Novel Prognostic Marker for Breast Cancer. PLoS ONE. 2012, 7, 46670. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Z.M.; Wang, X.Y.; Wang, A.M. Periostin induces chemoresistance in colon cancer cells through activation of the PI3K/Akt/survivin pathway. Biotechnol. Appl. Biochem. 2015, 62, 401–406. [Google Scholar] [CrossRef]
- Świerczewska, M.; Klejewski, A.; Brązert, M.; Kaźmierczak, D.; Iżycki, D.; Nowicki, M.; Zabel, M.; Januchowski, R. New and Old Genes Associated with Primary and Established Responses to Paclitaxel Treatment in Ovarian Cancer Cell Lines. Molecules 2018, 23, 891. [Google Scholar] [CrossRef] [PubMed]
- Klejewski, A.; Świerczewska, M.; Zaorska, K.; Brązert, M.; Nowicki, M.; Zabel, M.; Januchowski, R. New and Old Genes Associated with Topotecan Resistance Development in Ovarian Cancer Cell Lines. Anticancer Res. 2017, 37, 1625–1636. [Google Scholar] [CrossRef] [PubMed]
- Świerczewska, M.; Klejewski, A.; Wojtowicz, K.; Brązert, M.; Iżycki, D.; Nowicki, M.; Zabel, M.; Januchowski, R. New and Old Genes Associated with Primary and Established Responses to Cisplatin and Topotecan Treatment in Ovarian Cancer Cell Lines. Molecules 2017, 22, 1717. [Google Scholar] [CrossRef] [PubMed]
- Clevers, H. The cancer stem cell: Premises, promises and challenges. Nat. Med. 2011, 17, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Alison, M.R.; Guppy, N.J.; Lim, S.M.; Nicholson, L.J. Finding cancer stem cells: Are aldehyde dehydrogenases fit for purpose? J. Pathol. 2010, 222, 335–344. [Google Scholar] [CrossRef] [PubMed]
- Silva, I.A.; Bai, S.; McLean, K.; Yang, K.; Griffith, K.; Thomas, D.; Ginestier, C.; Johnston, C.; Kueck, A.; Reynolds, R.K.; et al. Aldehyde dehydrogenase in combination with CD133 defines angiogenic ovarian cancer stem cells that portend poor patient survival. Cancer Res. 2011, 71, 3991–4001. [Google Scholar] [CrossRef]
- Landen, C.N., Jr.; Goodman, B.; Katre, A.A.; Steg, A.D.; Nick, A.M.; Stone, R.L.; Miller, L.D.; Mejia, P.V.; Jennings, N.B.; Gershenson, D.M.; et al. Targeting aldehyde dehydrogenase cancer stem cells in ovarian cancer. Mol. Cancer Ther. 2010, 9, 3186–3199. [Google Scholar] [CrossRef]
| Transcript | Sequence (5’-3’ Direction) | ENST Number (Available online: http://www.ensembl.org) | Product Size (bp) |
|---|---|---|---|
| OSF-2 | TTCTGACGCCTCAAAACTGA CATTCACGTTGCTCTCCAAA | 00000379742 | 128 bp |
| GADPH | GAAGGTGAAGGTCGGAGTCA GACAAGCTTCCCGTTCTCAG | 00000229239 | 199 bp |
| β-actin | TCTGGCACCACACCTTCTAC GATAGCACAGCCTGGATAGC | 00000331789 | 169 bp |
| HRPT1 | CTGAGGATTTGGAAAGGGTG AATCCAGCAGGTCAGCAAAG | 00000298556 | 156 bp |
| β2M | CGCTACTCTCTCTTTCTGGC ATGTCGGATGGATGAAACCC | 00000558401 | 133 bp |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sterzyńska, K.; Kaźmierczak, D.; Klejewski, A.; Świerczewska, M.; Wojtowicz, K.; Nowacka, M.; Brązert, J.; Nowicki, M.; Januchowski, R. Expression of Osteoblast-Specific Factor 2 (OSF-2, Periostin) Is Associated with Drug Resistance in Ovarian Cancer Cell Lines. Int. J. Mol. Sci. 2019, 20, 3927. https://doi.org/10.3390/ijms20163927
Sterzyńska K, Kaźmierczak D, Klejewski A, Świerczewska M, Wojtowicz K, Nowacka M, Brązert J, Nowicki M, Januchowski R. Expression of Osteoblast-Specific Factor 2 (OSF-2, Periostin) Is Associated with Drug Resistance in Ovarian Cancer Cell Lines. International Journal of Molecular Sciences. 2019; 20(16):3927. https://doi.org/10.3390/ijms20163927
Chicago/Turabian StyleSterzyńska, Karolina, Dominika Kaźmierczak, Andrzej Klejewski, Monika Świerczewska, Karolina Wojtowicz, Marta Nowacka, Jacek Brązert, Michał Nowicki, and Radosław Januchowski. 2019. "Expression of Osteoblast-Specific Factor 2 (OSF-2, Periostin) Is Associated with Drug Resistance in Ovarian Cancer Cell Lines" International Journal of Molecular Sciences 20, no. 16: 3927. https://doi.org/10.3390/ijms20163927
APA StyleSterzyńska, K., Kaźmierczak, D., Klejewski, A., Świerczewska, M., Wojtowicz, K., Nowacka, M., Brązert, J., Nowicki, M., & Januchowski, R. (2019). Expression of Osteoblast-Specific Factor 2 (OSF-2, Periostin) Is Associated with Drug Resistance in Ovarian Cancer Cell Lines. International Journal of Molecular Sciences, 20(16), 3927. https://doi.org/10.3390/ijms20163927






