Analysis of Survival-Related lncRNA Landscape Identifies A Role for LINC01537 in Energy Metabolism and Lung Cancer Progression
Abstract
1. Introduction
2. Results
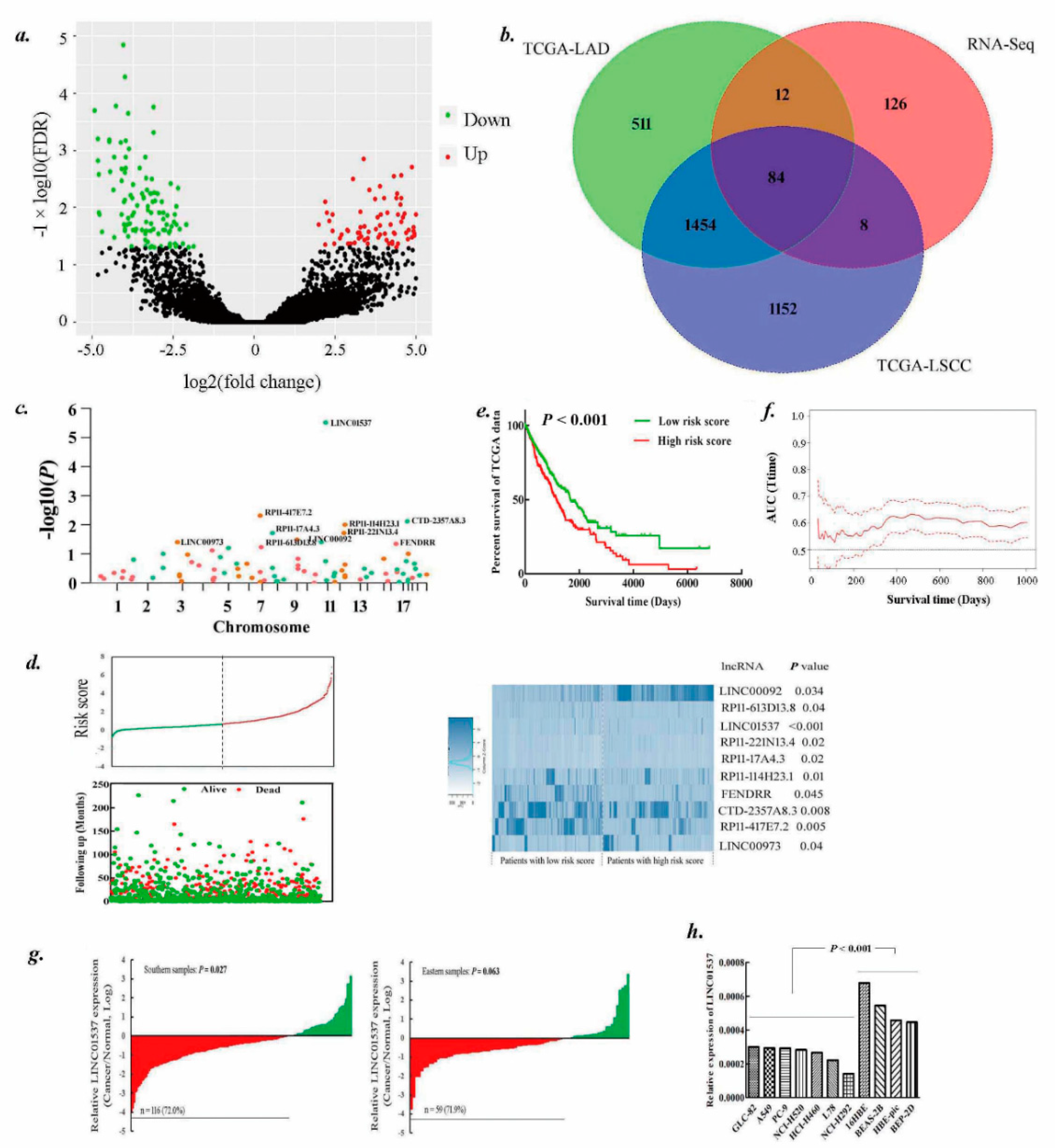
2.1. Identification of lncRNAs Landscape for Lung Cancer Survival
2.2. LINC01537 Is Downregulated in Lung Cancer
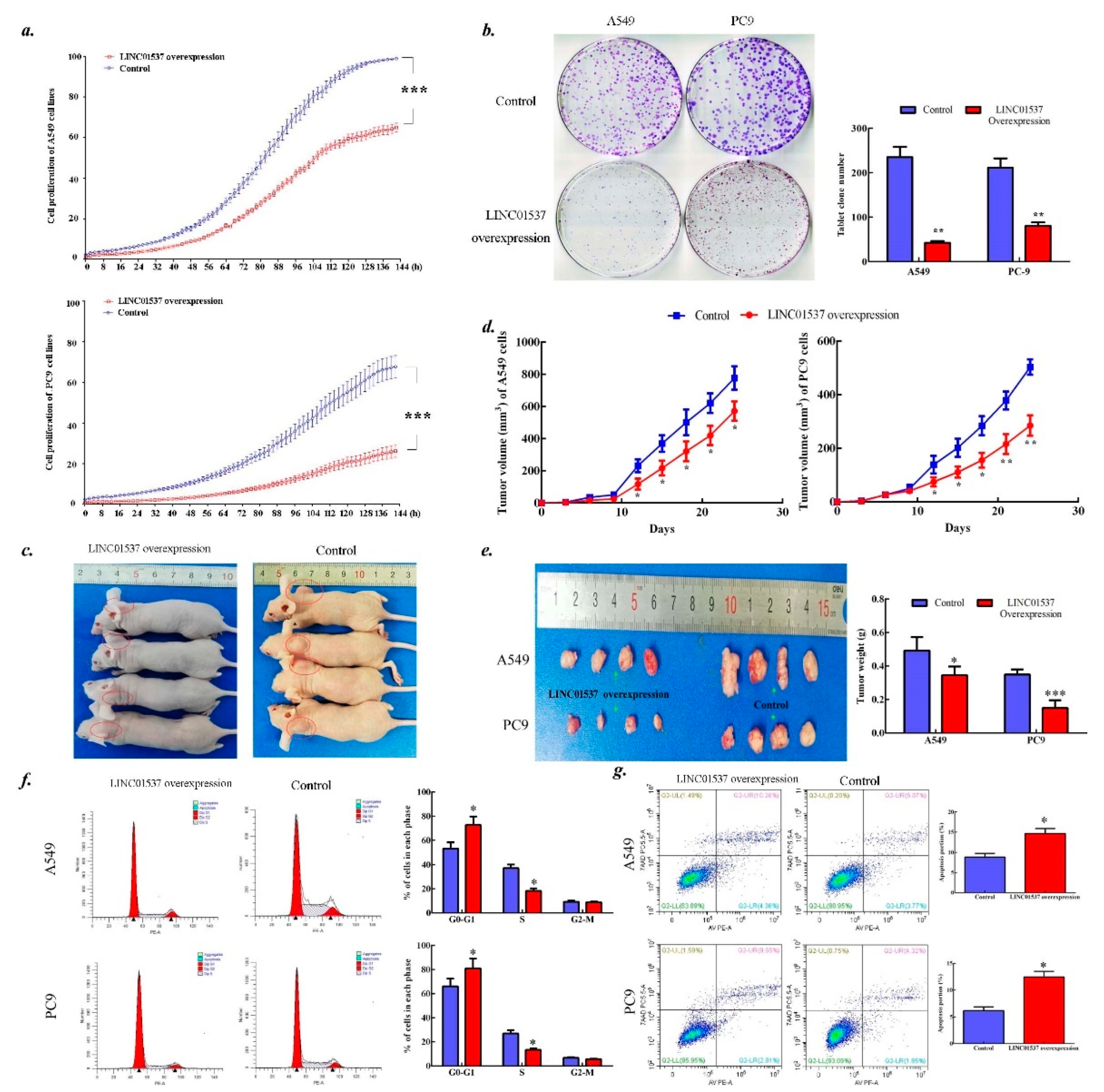
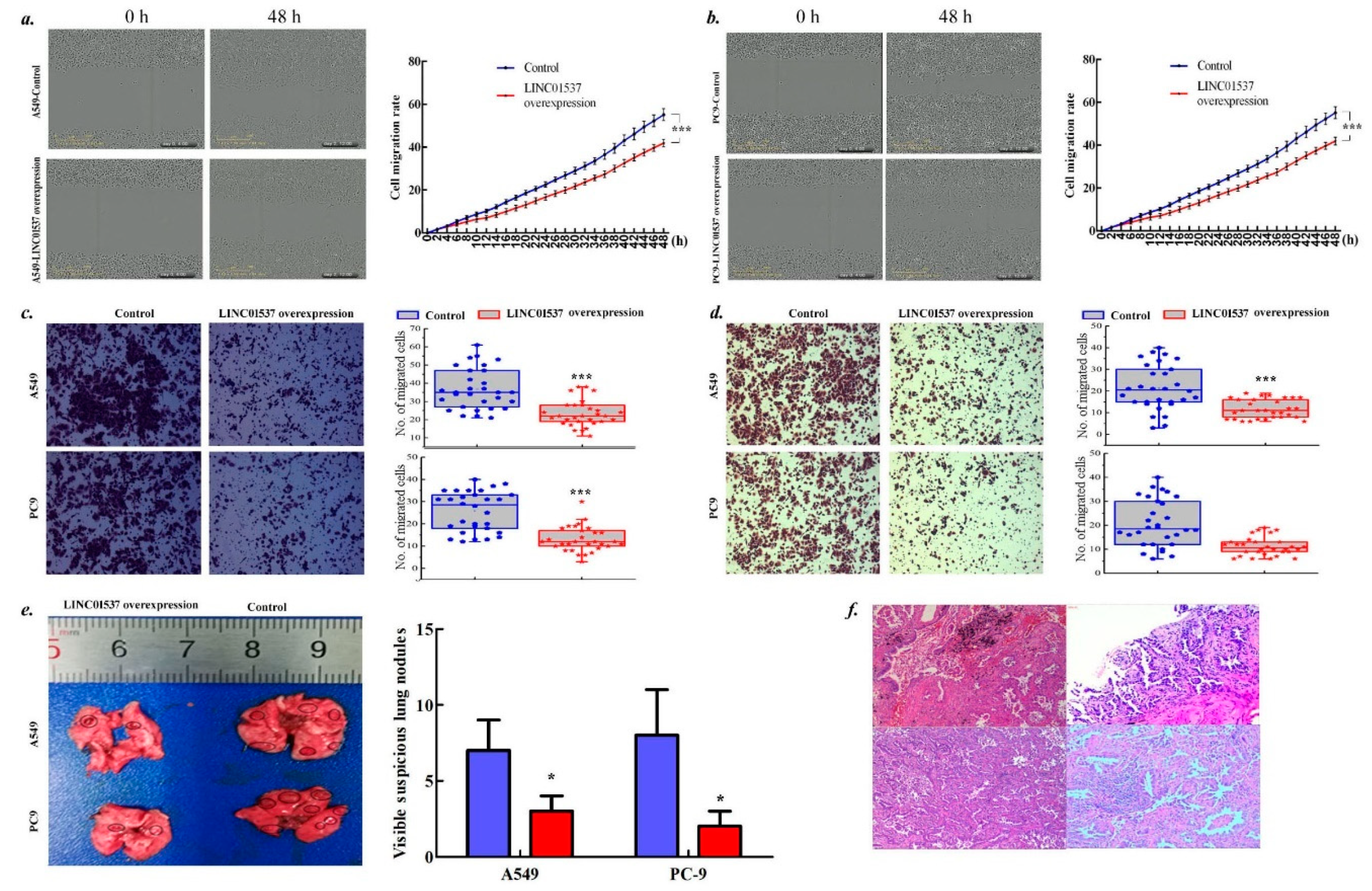
2.3. LINC01537 Inhibits Lung Cancer Development
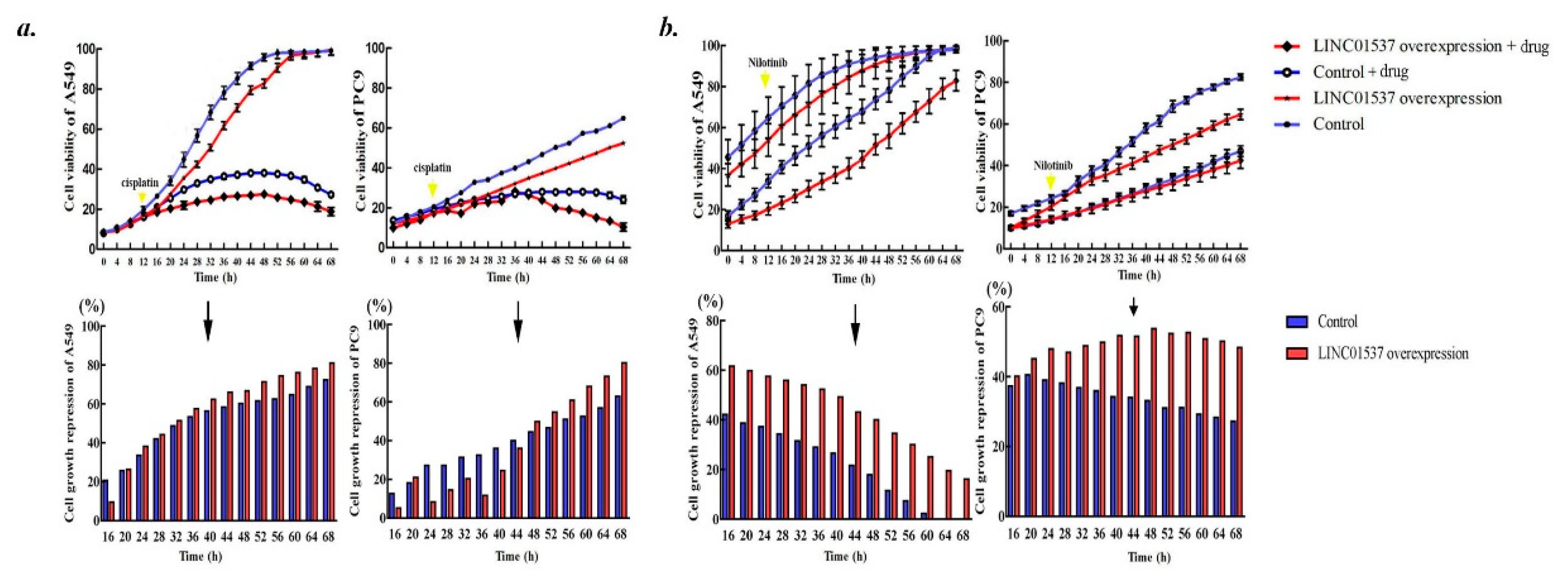
2.4. Overexpressed LINC01537 Benefits Anticancer Medicament
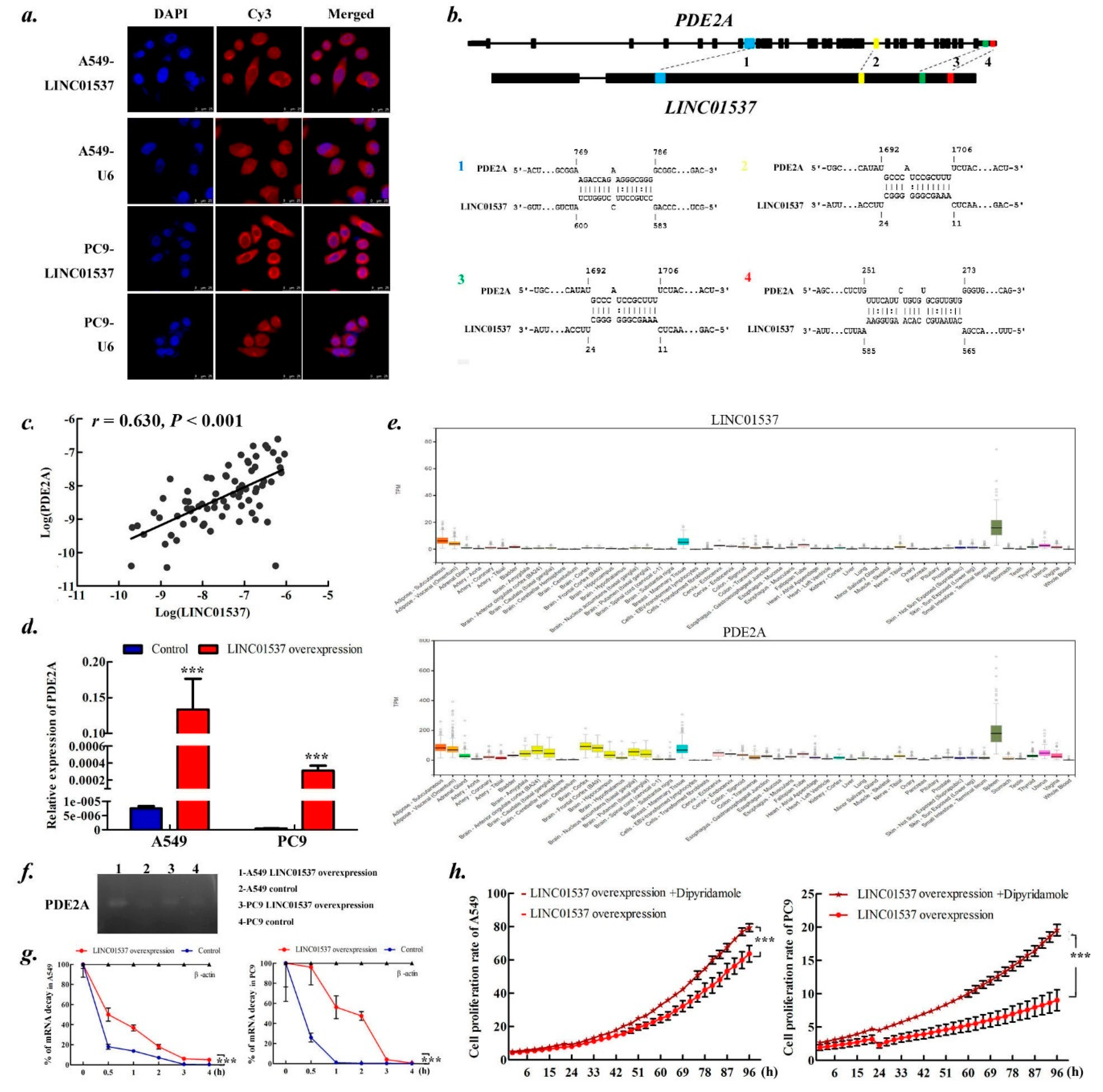
2.5. LINC01537 Regulates PDE2A Expression by Stabling PDE2A mRNA
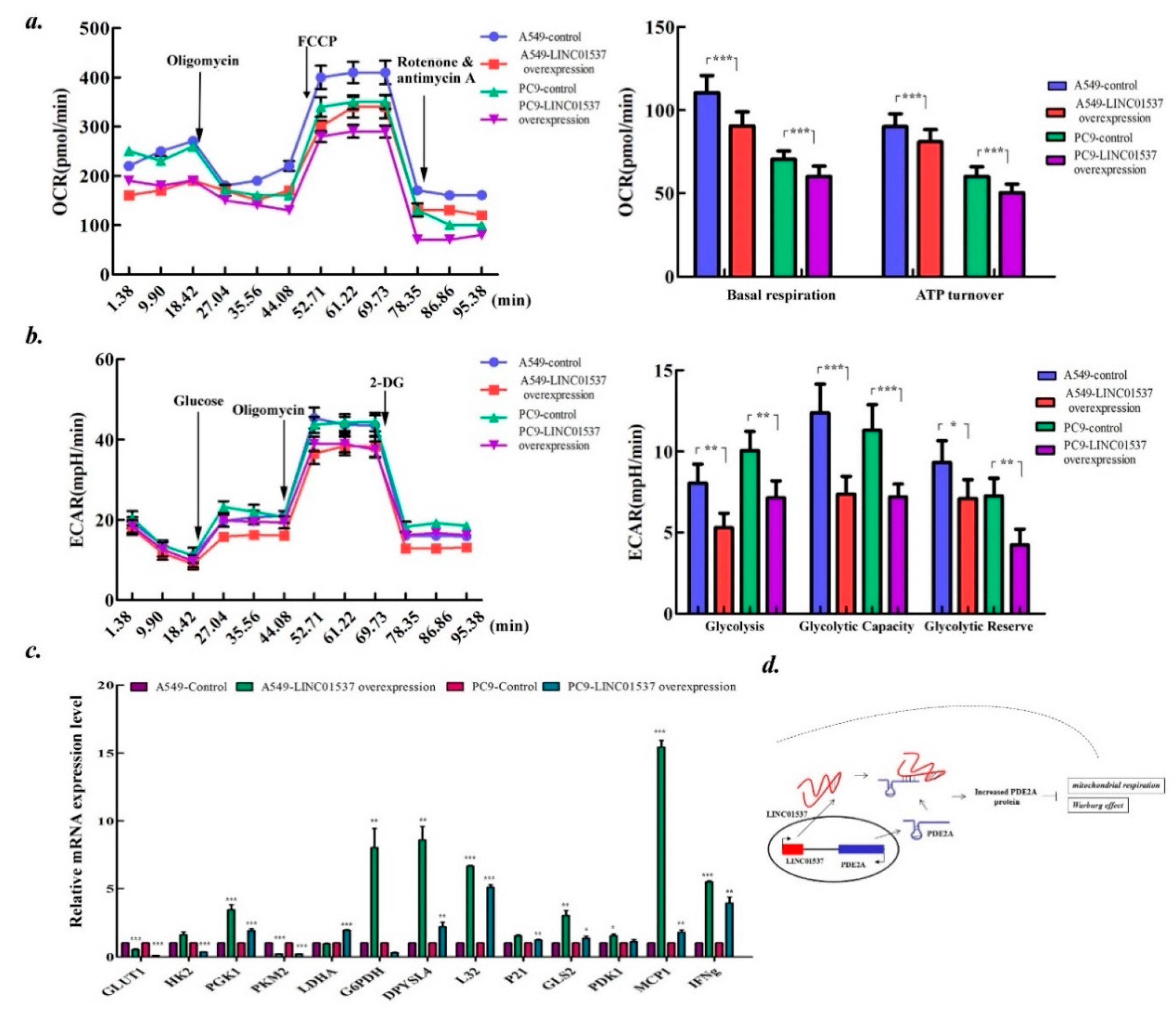
2.6. LINC01537 Affects Both the Mitochondrial Respiration and Warburg Effect
3. Discussion
4. Methods
4.1. Patient Sample Collection
4.2. RNA-seq
4.3. TCGA Data Analysis
4.4. Cell Culture
4.5. RNA Isolation and Quantitative Real-Time RT-PCR (qPCR)
4.6. Construction of LINC01537 Stable Overexpression Cells
4.7. Cell Phenotypic Experiments
4.8. Tumor Xenografts and Neoplasms Metastatic Model in Mice
4.9. Drug Sensitivity Assay
4.10. Subcellular Localization
4.11. Ribonuclease Protection Assay (RPA) and mRNA Stability Test
4.12. PDE2A Interference
4.13. Glycolysis Stress Test and Cell Mitochondria Stress Test
4.14. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Huang, M.; Shen, A.; Ding, J.; Geng, M. Molecularly targeted cancer therapy: Some lessons from the past decade. Trends Pharmacol. Sci. 2014, 35, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Turajlic, S.; Swanton, C. Metastasis as an evolutionary process. Science 2016, 352, 169–175. [Google Scholar] [CrossRef] [PubMed]
- Consortium, E.P.; Birney, E.; Stamatoyannopoulos, J.A.; Dutta, A.; Guigo, R.; Gingeras, T.R.; Margulies, E.H.; Weng, Z.; Snyder, M.; Dermitzakis, E.T.; et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 2007, 447, 799–816. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.; Kapranov, P.; Drenkow, J.; Dike, S.; Brubaker, S.; Patel, S.; Long, J.; Stern, D.; Tammana, H.; Helt, G.; et al. Transcriptional maps of 10 human chromosomes at 5-nucleotide resolution. Science 2005, 308, 1149–1154. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef] [PubMed]
- Rinn, J.L.; Kertesz, M.; Wang, J.K.; Squazzo, S.L.; Xu, X.; Brugmann, S.A.; Goodnough, L.H.; Helms, J.A.; Farnham, P.J.; Segal, E.; et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007, 129, 1311–1323. [Google Scholar] [CrossRef] [PubMed]
- Pauler, F.M.; Koerner, M.V.; Barlow, D.P. Silencing by imprinted noncoding RNAs: Is transcription the answer? Trends Genet. 2007, 23, 284–292. [Google Scholar] [CrossRef]
- Beltran, M.; Puig, I.; Pena, C.; Garcia, J.M.; Alvarez, A.B.; Pena, R.; Bonilla, F.; de Herreros, A.G. A natural antisense transcript regulates Zeb2/Sip1 gene expression during Snail1-induced epithelial-mesenchymal transition. Genes Dev. 2008, 22, 756–769. [Google Scholar] [CrossRef]
- Zuin, J.; Casa, V.; Pozojevic, J.; Kolovos, P.; van den Hout, M.; van Ijcken, W.F.J.; Parenti, I.; Braunholz, D.; Baron, Y.; Watrin, E.; et al. Regulation of the cohesin-loading factor NIPBL: Role of the lncRNA NIPBL-AS1 and identification of a distal enhancer element. PLoS Genet. 2017, 13, e1007137. [Google Scholar] [CrossRef]
- Sun, Y.; Wei, G.; Luo, H.; Wu, W.; Skogerbo, G.; Luo, J.; Chen, R. The long noncoding RNA SNHG1 promotes tumor growth through regulating transcription of both local and distal genes. Oncogene 2017, 36, 6774–6783. [Google Scholar] [CrossRef]
- Allemani, C.; Weir, H.K.; Carreira, H.; Harewood, R.; Spika, D.; Wang, X.S.; Bannon, F.; Ahn, J.V.; Johnson, C.J.; Bonaventure, A.; et al. Global surveillance of cancer survival 1995–2009: Analysis of individual data for 25,676,887 patients from 279 population-based registries in 67 countries (CONCORD-2). Lancet 2015, 385, 977–1010. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2018. CA Cancer J. Clin. 2018, 68, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.Q.; He, C.Y.; Hu, L.; Shi, H.P.; Li, J.F.; Gu, Q.L.; Su, L.P.; Liu, B.Y.; Li, C.; Zhu, Z. Long noncoding RNA UCA1 promotes tumour metastasis by inducing GRK2 degradation in gastric cancer. Cancer Lett. 2017, 408, 10–21. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Wu, D.; Chen, J.; Chen, J.; Qiu, F.; Li, Y.; Liu, L.; Cao, Y.; Yang, B.; Zhou, Y.; et al. A functional CNVR_3425.1 damping lincRNA FENDRR increases lifetime risk of lung cancer and COPD in Chinese. Carcinogenesis 2018, 39, 347–359. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Cui, Y.; Jin, J.; Guo, J.; Wang, G.; Yin, X.; He, Q.Y.; Zhang, G. Translating mRNAs strongly correlate to proteins in a multivariate manner and their translation ratios are phenotype specific. Nucleic Acids Res. 2013, 41, 4743–4754. [Google Scholar] [CrossRef] [PubMed]
- Engreitz, J.M.; Sirokman, K.; McDonel, P.; Shishkin, A.A.; Surka, C.; Russell, P.; Grossman, S.R.; Chow, A.Y.; Guttman, M.; Lander, E.S. RNA-RNA interactions enable specific targeting of noncoding RNAs to nascent Pre-mRNAs and chromatin sites. Cell 2014, 159, 188–199. [Google Scholar] [CrossRef] [PubMed]
- Monterisi, S.; Lobo, M.J.; Livie, C.; Castle, J.C.; Weinberger, M.; Baillie, G.; Surdo, N.C.; Musheshe, N.; Stangherlin, A.; Gottlieb, E.; et al. PDE2A2 regulates mitochondria morphology and apoptotic cell death via local modulation of cAMP/PKA signalling. eLife 2017, 6, e21374. [Google Scholar] [CrossRef] [PubMed]
- Tao, H.; Yang, J.J.; Zhou, X.; Deng, Z.Y.; Shi, K.H.; Li, J. Emerging role of long noncoding RNAs in lung cancer: Current status and future prospects. Respir. Med. 2016, 110, 12–19. [Google Scholar] [CrossRef]
- Zinovieva, O.L.; Grineva, E.N.; Prokofjeva, M.M.; Karpov, D.S.; Zheltukhin, A.O.; Krasnov, G.S.; Snezhkina, A.V.; Kudryavtseva, A.V.; Chumakov, P.M.; Mashkova, T.D.; et al. Expression of long non-coding RNA LINC00973 is consistently increased upon treatment of colon cancer cells with different chemotherapeutic drugs. Biochimie 2018, 151, 67–72. [Google Scholar] [CrossRef]
- Fan, Q.; Liu, B. Identification of a RNA-Seq Based 8-Long Non-Coding RNA Signature Predicting Survival in Esophageal Cancer. Med. Sci. Monitor 2016, 22, 5163–5172. [Google Scholar] [CrossRef]
- Sun, R.; Liu, Z.; Tong, D.; Yang, Y.; Guo, B.; Wang, X.; Zhao, L.; Huang, C. miR-491-5p, mediated by Foxi1, functions as a tumor suppressor by targeting Wnt3a/beta-catenin signaling in the development of gastric cancer. Cell Death Dis. 2017, 8, e2714. [Google Scholar]
- Zhao, L.; Ji, G.; Le, X.; Wang, C.; Xu, L.; Feng, M.; Zhang, Y.; Yang, H.; Xuan, Y.; Yang, Y.; et al. Long Noncoding RNA LINC00092 Acts in Cancer-Associated Fibroblasts to Drive Glycolysis and Progression of Ovarian Cancer. Cancer Res. 2017, 77, 1369–1382. [Google Scholar] [CrossRef] [PubMed]
- Tang, R.X.; Chen, W.J.; He, R.Q.; Zeng, J.H.; Liang, L.; Li, S.K.; Ma, J.; Luo, D.Z.; Chen, G. Identification of a RNA-Seq based prognostic signature with five lncRNAs for lung squamous cell carcinoma. Oncotarget 2017, 8, 50761–50773. [Google Scholar] [CrossRef] [PubMed]
- Shibata, T.; Uryu, S.; Kokubu, A.; Hosoda, F.; Ohki, M.; Sakiyama, T.; Matsuno, Y.; Tsuchiya, R.; Kanai, Y.; Kondo, T.; et al. Genetic classification of lung adenocarcinoma based on array-based comparative genomic hybridization analysis: Its association with clinicopathologic features. Clin. Cancer Res. 2005, 11, 6177–6185. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, Y.; Pitchiaya, S.; Cieslik, M.; Niknafs, Y.S.; Tien, J.C.; Hosono, Y.; Iyer, M.K.; Yazdani, S.; Subramaniam, S.; Shukla, S.K.; et al. Analysis of the androgen receptor-regulated lncRNA landscape identifies a role for ARLNC1 in prostate cancer progression. Nature Genet. 2018, 50, 814–824. [Google Scholar] [CrossRef]
- Faghihi, M.A.; Modarresi, F.; Khalil, A.M.; Wood, D.E.; Sahagan, B.G.; Morgan, T.E.; Finch, C.E.; St Laurent, G.; Kenny, P.J.; Wahlestedt, C. Expression of a noncoding RNA is elevated in Alzheimer’s disease and drives rapid feed-forward regulation of beta-secretase. Nature Med. 2008, 14, 723–730. [Google Scholar] [CrossRef]
- Gong, C.; Maquat, L.E. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3’ UTRs via Alu elements. Nature 2011, 470, 284–288. [Google Scholar] [CrossRef]
- Kretz, M.; Siprashvili, Z.; Chu, C.; Webster, D.E.; Zehnder, A.; Qu, K.; Lee, C.S.; Flockhart, R.J.; Groff, A.F.; Chow, J.; et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 2013, 493, 231–235. [Google Scholar] [CrossRef]
- Pusztai, L.; Zhen, J.H.; Arun, B.; Rivera, E.; Whitehead, C.; Thompson, W.J.; Nealy, K.M.; Gibbs, A.; Symmans, W.F.; Esteva, F.J.; et al. Phase I and II study of exisulind in combination with capecitabine in patients with metastatic breast cancer. J. Clin. Oncol. 2003, 21, 3454–3461. [Google Scholar] [CrossRef]
- Yang, B.; Zhang, L.; Cao, Y.; Chen, S.; Cao, J.; Wu, D.; Chen, J.; Xiong, H.; Pan, Z.; Qiu, F.; et al. Overexpression of lncRNA IGFBP4-1 reprograms energy metabolism to promote lung cancer progression. Mol. Cancer 2017, 16, 154. [Google Scholar] [CrossRef]
- Liberti, M.V.; Locasale, J.W. The Warburg Effect: How Does it Benefit Cancer Cells? Trends Biochem. Sci. 2016, 41, 211–218. [Google Scholar] [CrossRef]
- Vyas, S.; Zaganjor, E.; Haigis, M.C. Mitochondria and Cancer. Cell 2016, 166, 555–566. [Google Scholar] [CrossRef] [PubMed]
- Romero-Garcia, S.; Lopez-Gonzalez, J.S.; Baez-Viveros, J.L.; Aguilar-Cazares, D.; Prado-Garcia, H. Tumor cell metabolism: An integral view. Cancer Biol. Ther. 2011, 12, 939–948. [Google Scholar] [CrossRef]
- De Paepe, B.; Lefever, S.; Mestdagh, P. How long noncoding RNAs enforce their will on mitochondrial activity: Regulation of mitochondrial respiration, reactive oxygen species production, apoptosis, and metabolic reprogramming in cancer. Current Genet. 2018, 64, 163–172. [Google Scholar] [CrossRef] [PubMed]
- Pan, Z.; Liu, L.; Nie, W.; Miggin, S.; Qiu, F.; Cao, Y.; Chen, J.; Yang, B.; Zhou, Y.; Lu, J.; et al. Long non-coding RNA AGER-1 functionally upregulates the innate immunity gene AGER and approximates its anti-tumor effect in lung cancer. Mol. Carcinog 2018, 57, 305–318. [Google Scholar] [CrossRef] [PubMed]
- Alizadeh, A.A.; Gentles, A.J.; Alencar, A.J.; Liu, C.L.; Kohrt, H.E.; Houot, R.; Goldstein, M.J.; Zhao, S.; Natkunam, Y.; Advani, R.H.; et al. Prediction of survival in diffuse large B-cell lymphoma based on the expression of 2 genes reflecting tumor and microenvironment. Blood 2011, 118, 1350–1358. [Google Scholar] [CrossRef] [PubMed]
- Sundaram, G.M.; Common, J.E.; Gopal, F.E.; Srikanta, S.; Lakshman, K.; Lunny, D.P.; Lim, T.C.; Tanavde, V.; Lane, E.B.; Sampath, P. ‘See-saw’ expression of microRNA-198 and FSTL1 from a single transcript in wound healing. Nature 2013, 495, 103–106. [Google Scholar] [CrossRef]
- Cullis, J.; Meiri, D.; Sandi, M.J.; Radulovich, N.; Kent, O.A.; Medrano, M.; Mokady, D.; Normand, J.; Larose, J.; Marcotte, R.; et al. The RhoGEF GEF-H1 is required for oncogenic RAS signaling via KSR-1. Cancer Cell 2014, 25, 181–195. [Google Scholar] [CrossRef]
- Blay, J.Y.; Shen, L.; Kang, Y.K.; Rutkowski, P.; Qin, S.; Nosov, D.; Wan, D.; Trent, J.; Srimuninnimit, V.; Papai, Z.; et al. Nilotinib versus imatinib as first-line therapy for patients with unresectable or metastatic gastrointestinal stromal tumours (ENESTg1): A randomised phase 3 trial. Lancet Oncol. 2015, 16, 550–560. [Google Scholar] [CrossRef]
- Guo, W.; Liu, S.; Cheng, Y.; Lu, L.; Shi, J.; Xu, G.; Li, N.; Cheng, K.; Wu, M.; Cheng, S.; et al. ICAM-1-Related Noncoding RNA in Cancer Stem Cells Maintains ICAM-1 Expression in Hepatocellular Carcinoma. Clin. Cancer Res. 2016, 22, 2041–2050. [Google Scholar] [CrossRef]
- Wunder, F.; Gnoth, M.J.; Geerts, A.; Barufe, D. A novel PDE2A reporter cell line: Characterization of the cellular activity of PDE inhibitors. Mol. Pharm. 2009, 6, 326–336. [Google Scholar] [CrossRef] [PubMed]
- Heagerty, P.J.; Lumley, T.; Pepe, M.S. Time-dependent ROC curves for censored survival data and a diagnostic marker. Biometrics 2000, 56, 337–344. [Google Scholar] [CrossRef] [PubMed]
| Gene Symbol | Lung Adenocarcinoma | Lung Squamous Cell Carcinoma | ||
|---|---|---|---|---|
| Correlation Coefficient | p Value | Correlation Coefficient | p Value | |
| PDE2A | 0.636 | 2.14 × 10−41 | 0.688 | 3.07 × 10−32 |
| CXorf36 | 0.5 | 1.12 × 10−23 | 0.563 | 8.78 × 10−20 |
| PEAR1 | 0.467 | 1.55 × 10−20 | 0.525 | 5.37 × 10−17 |
| RHOJ | 0.454 | 2.38 × 10−19 | 0.515 | 2.48 × 10−16 |
| LAMA4 | 0.452 | 3.34 × 10−19 | 0.494 | 6.33 × 10−15 |
| TIE1 | 0.451 | 4.46 × 10−19 | 0.481 | 3.82 × 10−14 |
| CD93 | 0.448 | 8.47 × 10−19 | 0.508 | 7.57 × 10−16 |
| FLT4 | 0.446 | 1.27 × 10−18 | 0.402 | 5.7 × 10−10 |
| MYCT1 | 0.441 | 3.06 × 10−18 | 0.459 | 6.91 × 10−13 |
| MMRN2 | 0.438 | 5.20 × 10−18 | 0.522 | 9.02 × 10−17 |
| VGLL3 | 0.437 | 6.24 × 10−18 | 0.433 | 1.87 × 10−11 |
| GIPC3 | 0.432 | 1.60 × 10−17 | 0.497 | 3.95 × 10−15 |
| GPR124 | 0.432 | 1.89 × 10−17 | 0.405 | 4.21 × 10−10 |
| BNC2 | 0.431 | 2.31 × 10−17 | 0.414 | 1.63 × 10−10 |
| LDB2 | 0.431 | 2.27 × 10−17 | 0.438 | 9.76 × 10−12 |
| CD34 | 0.43 | 2.42 × 10−17 | 0.476 | 7.32 × 10−14 |
| ROBO4 | 0.43 | 2.60 × 10−17 | 0.513 | 3.77 × 10−16 |
| SYNPO | 0.428 | 3.99 × 10−17 | 0.486 | 2 × 10−14 |
| ELTD1 | 0.422 | 1.04 × 10−16 | 0.489 | 1.19 × 10−14 |
| BCL6B | 0.417 | 2.98 × 10−16 | 0.465 | 3.14 × 10−13 |
| TIMP3 | 0.411 | 8.14 × 10−16 | 0.421 | 7.16 × 10−11 |
| PDGFRB | 0.41 | 1.02 × 10−15 | 0.435 | 1.39 × 10−11 |
| ITGA5 | 0.409 | 1.05 × 10−15 | 0.517 | 2.03 × 10−16 |
| SOX18 | 0.409 | 1.16 × 10−15 | 0.421 | 7.41 × 10−11 |
| ARHGEF15 | 0.408 | 1.33 × 10−15 | 0.438 | 9.87 × 10−12 |
| SPON1 | 0.408 | 1.31 × 10−15 | 0.418 | 1.01 × 10−10 |
| CACNA1C | 0.406 | 1.87 × 10−15 | 0.42 | 8.64 × 10−11 |
| PALMD | 0.405 | 2.35 × 10−15 | 0.411 | 2.26 × 10−10 |
| MMP2 | 0.401 | 4.86 × 10−15 | 0.414 | 1.62 × 10−10 |
| Characteristic | Southern Samples N (%) | Eastern Samples N (%) | Pearson χ2 | p Value |
|---|---|---|---|---|
| Total | 161 (66.3) | 82 (33.7) | ||
| Age | ||||
| <60 | 92 (57.1) | 43 (52.4) | 0.487 | 0.485 |
| ≥60 | 69 (42.9) | 39(47.6) | ||
| Gender | ||||
| Female | 49 (30.4) | 23 (28.0) | 0.148 | 0.700 |
| Male | 112 (69.6) | 59 (72.0) | ||
| Family tumor history | ||||
| No | 141 (87.6) | 73 (89.0) | 0.108 | 0.742 |
| Yes | 20 (12.4) | 9 (11.0) | ||
| Smoking | ||||
| No | 62 (38.5) | 26 (31.7) | 1.088 | 0.297 |
| Yes | 99 (61.5) | 56 (68.3) | ||
| Stages | ||||
| I + II | 61(37.9) | 25(30.5) | 1.301 | 0.254 |
| III + IV | 100(62.1) | 57(69.5) | ||
| Pathology | ||||
| Adenocarcinoma | 77(47.8) | 35(42.7) | 0.899 | 0.638 |
| Squamous carcinoma | 44(27.3) | 27(32.9) | ||
| Other types a | 40(24.8) | 20(24.4) | ||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gong, W.; Yang, L.; Wang, Y.; Xian, J.; Qiu, F.; Liu, L.; Lin, M.; Feng, Y.; Zhou, Y.; Lu, J. Analysis of Survival-Related lncRNA Landscape Identifies A Role for LINC01537 in Energy Metabolism and Lung Cancer Progression. Int. J. Mol. Sci. 2019, 20, 3713. https://doi.org/10.3390/ijms20153713
Gong W, Yang L, Wang Y, Xian J, Qiu F, Liu L, Lin M, Feng Y, Zhou Y, Lu J. Analysis of Survival-Related lncRNA Landscape Identifies A Role for LINC01537 in Energy Metabolism and Lung Cancer Progression. International Journal of Molecular Sciences. 2019; 20(15):3713. https://doi.org/10.3390/ijms20153713
Chicago/Turabian StyleGong, Wei, Lei Yang, Yuanyuan Wang, Jianfeng Xian, Fuman Qiu, Li Liu, Mingzhu Lin, Yingyi Feng, Yifeng Zhou, and Jiachun Lu. 2019. "Analysis of Survival-Related lncRNA Landscape Identifies A Role for LINC01537 in Energy Metabolism and Lung Cancer Progression" International Journal of Molecular Sciences 20, no. 15: 3713. https://doi.org/10.3390/ijms20153713
APA StyleGong, W., Yang, L., Wang, Y., Xian, J., Qiu, F., Liu, L., Lin, M., Feng, Y., Zhou, Y., & Lu, J. (2019). Analysis of Survival-Related lncRNA Landscape Identifies A Role for LINC01537 in Energy Metabolism and Lung Cancer Progression. International Journal of Molecular Sciences, 20(15), 3713. https://doi.org/10.3390/ijms20153713




