Genomic Organization of Repetitive DNA Elements and Extensive Karyotype Diversity of Silurid Catfishes (Teleostei: Siluriformes): A Comparative Cytogenetic Approach
Abstract
1. Introduction
2. Results
2.1. Standard Karyotypes
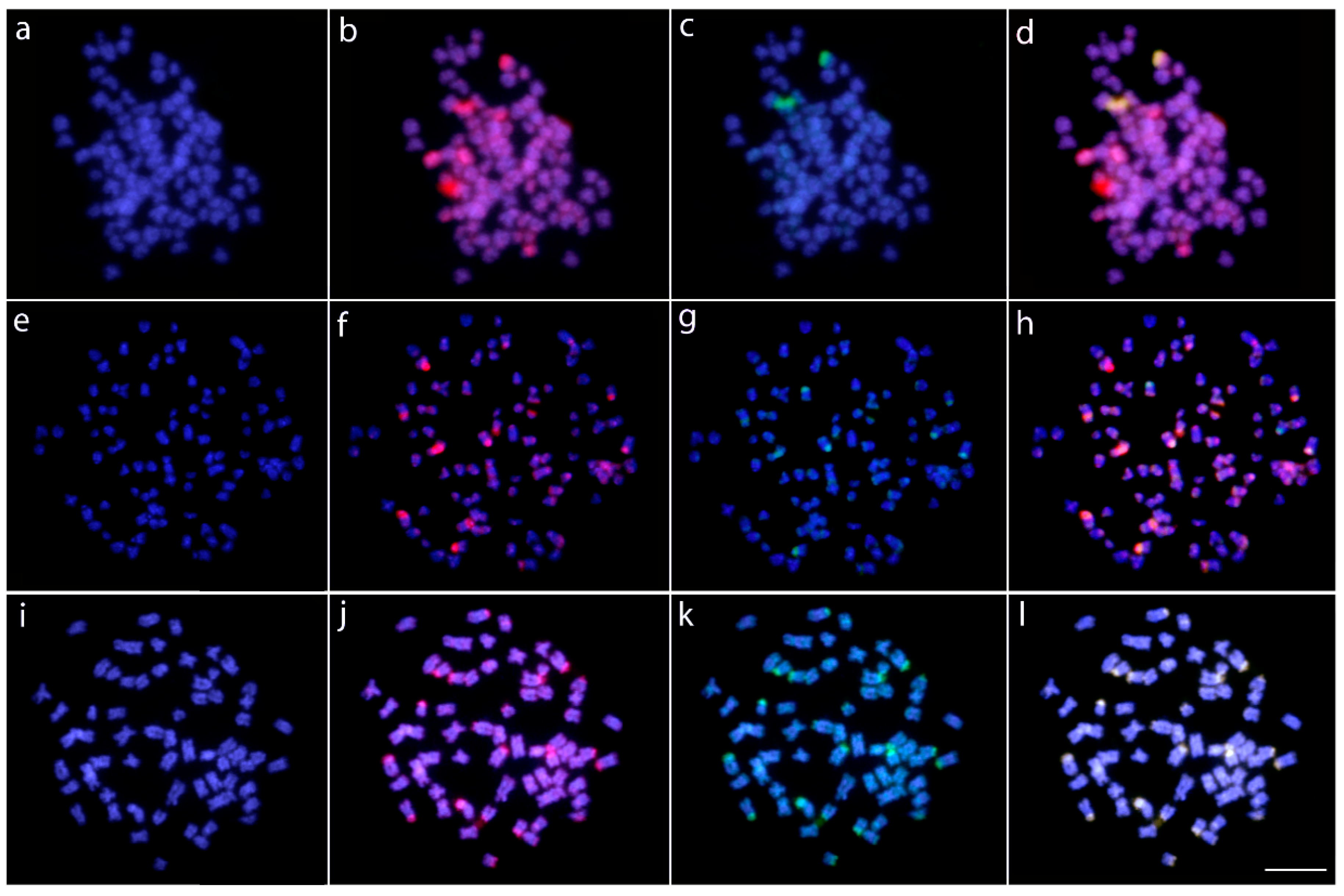
2.2. Fluorescence In Situ Hybridization (FISH)-Mapping
2.3. Comparative Genomic Hybridization (CGH)
3. Discussion
4. Materials and Methods
4.1. Individuals and Mitotic Chromosome Preparation
4.2. Fluorescence In Situ Hybridization (FISH)
4.3. Comparative Genome Hybridization (CGH)
4.4. Cytogenetic Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Kottelat, M. The fishes of the inland waters of Southeast Asia: A catalogue and core bibliography of the fishes known to occur in freshwaters, mangroves and estuaries. Raffles Bull Zool. 2013, 27, 1–663. [Google Scholar]
- Fricke, R.; Eschmeyer, W.; van der Laan, R. Eschmeyer’s Catalog of Fishes: Genera, Species, References; California Academy of Sciences: San Francisco, CA, USA, 2019. [Google Scholar]
- Bornbusch, A.H. Phylogenetic relationships within the Eurasian catfish family Siluridae (Pisces: Siluriformes), with comments on generic validities and biogeography. Zool. J. Linn. Soc. 1995, 115, 1–46. [Google Scholar] [CrossRef]
- Kobayakawa, M. Systematic revision of the catfish genus Silurus, with description of a new species from Thailand and Burma. Jpn. J. Ichthyol. 1989, 36, 155–186. [Google Scholar] [CrossRef]
- Kottelat, M.; Freyhof, J. Handbook of European Freshwater Fishes; Publications Kottelat: Berlin, Germany, 2007. [Google Scholar]
- Bogutskaya, N.G.; Naseka, A.M.; Shedko, S.V.; Vasileva, E.D.; Chereschnev, I.A. The fishes of the Amur River: Updated check-list and zoogeography. Ichtyological Explor. Freshw. 2008, 19, 301–366. [Google Scholar]
- Nelson, J.S.; Grande, T.C.; Wilson, M.V.H. Fishes of the World, 5th ed.; John Wiley & Sons: Hoboken, NJ, USA, 2016. [Google Scholar]
- Bornbusch, A.H. Monophyly of the catfish family Siluridae (Teleostei: Siluriformes), with a critique of previous hypotheses of the family’s relationships. Zool. J. Linn. Soc. 1991, 101, 105–120. [Google Scholar] [CrossRef]
- Hardman, M. The phylogenetic relationships among non-diplomystid catfishes as inferred from mitochondrial cytochrome b sequences; the search for the ictalurid sister taxon (Otophysi: Siluriformes). Mol. Phylogenet. Evol. 2005, 37, 700–720. [Google Scholar] [CrossRef] [PubMed]
- Mo, T. Anatomy, relationships and systematics of the Bagridae (Teleostei: Siluroidei) with a hypothesis of siluroid phylogeny. Theses Zool. 1991, 17, 1–216. [Google Scholar]
- Pinna, M.C.C. Higher-level phylogeny of Siluriformes (Teleostei, Ostariophysi), with a new classification of the order. Ph.D. dissertation, City University of New York, New York, NY, USA, 1993. [Google Scholar]
- Diogo, R. Higher-level phylogeny of Siluriformes: An overview. In Catfishes Science; Arratia, G., Kapoor, B.G., Eds.; Science Publishers: Enfield, UK, 2003; pp. 353–384. [Google Scholar]
- Sullivan, J.P.; Lundberg, J.G.; Hardman, M. A phylogenetic analysis of the major groups of catfishes (Teleostei: Siluriformes) using rag1 and rag2 nuclear gene sequences. Mol. Phylogenet. Evol. 2006, 41, 636–662. [Google Scholar] [CrossRef]
- Stone, R. The last of the Leviathans. Science 2007, 316, 1684–1688. [Google Scholar] [CrossRef]
- Linhart, O.; Šĕtch, L.; Švarc, J.; Rodina, M.; Audebert, J.P.; Grecu, J.; Billard, R. The culture of the European catfish, Silurus glanis, in the Czech Republic and in France. Aquat. Living Resour. 2002, 15, 139–144. [Google Scholar] [CrossRef]
- Saenjundaeng, P. Encyclopedia of Freshwater Fishes in Thailand; Klungnana Vitthaya Press: Khon Kaen, Thailand, 2014. [Google Scholar]
- Chapman, F.A.; Fitz-Coy, S.A.; Thunberg, E.M.; Adams, C.M. United States of America trade in ornamental fish. J. World Aquac. Soc. 1997, 28, 1–10. [Google Scholar] [CrossRef]
- Oliveira, C.; Almeida-Toledo, L.F.; Foresti, F. Karyotypic evolution in Neotropical fishes. In Fish Cytogenetics; Pisano, E., Ozouf-Costaz, C., Foresti, F., Kapoor, B.G., Eds.; Science Publishers: Enfield, UK, 2007; pp. 111–164. [Google Scholar]
- Cioffi, M.B.; Bertollo, L.A.C. Chromosomal distribution and evolution of repetitive DNAs in fish. In Repetitive DNA; Garrido-Ramos, M.A., Ed.; Karger Publishers: Basel, Switzerland, 2012; Volume 7, pp. 197–221. [Google Scholar]
- Cioffi, M.B.; Moreira-Filho, O.; Ráb, P.; Sember, A.; Molina, W.F.; Bertollo, L.A.C. Conventional cytogenetic approaches—useful and indispensable tools in discovering fish biodiversity. Curr. Genet. Med. Rep. 2018, 6, 176–186. [Google Scholar] [CrossRef]
- Cioffi, M.B.; Bertollo, L.A.C.; Villa, M.A.; de Oliveira, E.A.; Tanomtong, A.; Yano, C.F.; Supiwong, W.; Chaveerach, A. Genomic organization of repetitive DNA elements and its implications for the chromosomal evolution of channid fishes (Actinopterygii, Perciformes). PLoS ONE 2015, 10, e0130199. [Google Scholar] [CrossRef] [PubMed]
- Symonová, R.; Majtánová, Z.; Sember, A.; Staaks, G.B.O.; Bohlen, J.; Freyhof, J. Genome differentiation in a species pair of coregonine fishes: An extremely rapid speciation driven by stress-activated retrotransposons mediating extensive ribosomal DNA multiplications. BMC Evol. Biol. 2013, 13, 42–52. [Google Scholar] [CrossRef] [PubMed]
- Moraes, R.L.R.; Bertollo, L.A.C.; Marinho, M.M.F.; Yano, C.F.; Hatanaka, T.; Barby, F.F.; Troy, W.P.; Cioffi, M.B. Evolutionary relationships and cytotaxonomy considerations in the genus Pyrrhulina (Characiformes, Lebiasinidae). Zebrafish 2017, 14, 536–546. [Google Scholar] [CrossRef]
- Sember, A.; Bertollo, L.A.C.; Ráb, P.; Yano, C.F.; Hatanaka, T.; de Oliveira, E.A.; Cioffi, M.B. Sex chromosome evolution and genomic divergence in the fish Hoplias malabaricus (Characiformes, Erythrinidae). Front. Genet. 2018, 9, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Barby, F.F.; Bertollo, L.A.C.; de Oliveira, E.A.; Yano, C.F.; Hatanaka, T.; Ráb, P.; Sember, A.; Ezaz, T.; Artoni, R.F.; Liehr, T.; et al. Emerging patterns of genome organization in Notopteridae species (Teleostei, Osteoglossiformes) as revealed by zoo-FISH and comparative genomic hybridization (CGH). Sci. Rep. 2019, 9, 1112. [Google Scholar] [CrossRef]
- Verma, J.; Lakra, W.S.; Kushwaha, B.; Sirajuddin, M.; Nagpure, N.S.; Kumar, R. Characterization of two freshwater silurid catfish using conventional and molecular cytogenetic techniques. J. Genet. 2011, 90, 319–322. [Google Scholar] [CrossRef]
- Lee, H.Y.; Yu, C.H.; Jeon, S.K.; Lee, H.S. The karyotype analysis on 29 species of freshwater fish in Korea. Bull. Inst. Basic Sci. Inha Univ 1983, 79–86. [Google Scholar]
- Donsakul, T. Chromosome study on three species of silurid fishes, Kryptopterus cryptopterus, K. bleekeri and Ompok bimaculatus, of Thailand. In Proceedings of the 18th Congress on Science and Technology of Thailand, Bangkok, Thailand, 1992; pp. 398–399. [Google Scholar]
- Donsakul, T. A study of chromosome in freshwater shark (Wallago attu), helicopter catfish (W. leerii), twisted-jaw sheatfish (Belodontichthys dinema) and glass catfish (Kryptopterus bicirrhis) from Thailand. In Proceedings of the 34th Kasetsart University Annual Conference (Fishery Section), Bangkok, Thailand, 1996; pp. 367–377. [Google Scholar]
- Simporn, B. Cytogenetic of Sheatfish (Family Siluridae) in Thailand. Ph.D Dissertation, Khon Kaen University, Khon Kaen, Thailand, 2015. [Google Scholar]
- Hinegardner, R.; Rosen, D.E. Cellular DNA content and the evolution of teleostean fishes. Amer. Natur. 1972, 106, 621–644. [Google Scholar] [CrossRef]
- Magtoon, W.; Donsakul, T. Karyotypes of Silurichthys phaiosoma, S. schneideri, Ompok fumidus and Kryptopterus macrocephalus (Family Siluridae) from Thailand. In Proceedings of the 35th Congress on Science and Technology of Thailand, Bangkok, Thailand, 2009; p. B0084. [Google Scholar]
- Pinthong, K.; Supiwong, W.; Simporn, B.; Chooseangjaew, S.; Kakampuy, W.; Tanomtong, A. A First chromosomal and nucleolar organizer regions (NORs) Analyses of Chevey’s sheetfish, Micronema cheveyi (Siluriformes, Siluridae). Cytologia (Tokyo) 2017, 82, 435–441. [Google Scholar] [CrossRef]
- Rishi, K.K. Mitotic and meiotic chromosomes of a teleost, Callichrous bimaculatus (Bloch) with indications of male heterogamety. Cienc. Cult. 1976, 28, 1171–1173. [Google Scholar]
- Khuda-Bukhsh, A.R. Cytogenetic analyses in eight species of teleostean fishes (Pisces): Karyotypes, multiple Ag-NORs, sex chromosomes. Res. Rev. BioSci. 2007, 1, 47–52. [Google Scholar]
- Sharma, R. Chromosomal stuides on two endangered catfishes Ompok bimaculatus (Bloch) and Ompak pabda (Hamilton-Buchanan). Indian J. Anim. Sci. 2008. [Google Scholar]
- Gomonteir, B.; Tanomtong, A.; Supiwong, W.; Sikhruadong, S.; Neeratanaphan, L.; Sanoamuang, L. Standardized karyotype and idiogram of two-spot glass catfish, Ompok bimaculatus (Siluriformes, Siluridae) in Thailand by conventional and Ag-NOR staining techniques. Cytologia (Tokyo). 2012, 77, 459–464. [Google Scholar] [CrossRef][Green Version]
- Das, R.K.; Kar, R.N. Somatic chromosome analysis of a siluroid fish, Rita chrysea Day. Caryologia 1977, 30, 247–253. [Google Scholar] [CrossRef]
- Datta, S.; Rahman, M.M.; Haque, M.A. Chromosomal studies on Ompok pabda (Hamilton). Online J. Biol. Sci. 2003, 3, 460–465. [Google Scholar]
- Khuda Bukhsh, A.R.; Chanda, T. Somatic chromosomes of three species of hill stream fishes from Assam. In Fish Genetics in India; Das, P., Jhingran, A.G., Eds.; Today Tomorrow’s Printers and Publishers: New Delhi, India, 1989; pp. 69–73. [Google Scholar]
- Supiwong, W.; Tanomtong, A.; Jumrusthanasan, S.; Khakhong, S.; Leelarasamee, K.; Sanoamuang, L. A First karyotype analysis and chromosomal characteristic of nucleolar organizer regions (NORs) of common sheathfish, Micronema apogon (Siluriformes, Siluridae) in Thailand. Cytologia (Tokyo). 2012, 77, 53–58. [Google Scholar] [CrossRef]
- Iliadou, K.; Rackham, B.D. The chromosomes of a catfish Parasilurus aristotelis from Greece. Jpn. J. Ichthyol. 1990, 37, 144–148. [Google Scholar]
- Rab, P.; Karakousis, Y.; Peios, C. Karyotype of Silurus aristotelis with reference to the cytotaxonomy of the genus Silurus (Piscus, Siluridae). Folia Zool. (Czech Republic) 1994, 43, 75–81. [Google Scholar]
- Nogusa, S. Chromosome studies in Pisces, II. On the chromosomes of Sillago sihama (Sillaginidae) and Parasilurus asotus (Siluridae). Jpn. J. Genet 1951, 26, 153–155. [Google Scholar] [CrossRef]
- Muramoto, J. On the chromosomes of Parasilurus asotus (Linne). Chrom. Inf. Serv. 1969, 10, 18–19. [Google Scholar]
- Fujioka, Y. A comparative study of the chromosomes in Japanese freshwater fishes I. A study of the somatic chromosome of common catfish Parasilurus asotus (Linne) and forktailed bullhead Paleobagrus nudiceps (Sauvage). Bull. Fac. Educ. Yamaguchi Univ. 1973, 23, 191–195. [Google Scholar]
- Arai, R.; Katsuyama, I. A chromosome study on four species of Japanese catfishes (Pisces, Siluriformes). Bull. Nat. Sci. Museum 1974, 187–192. [Google Scholar]
- Ojima, Y. Cellular DNA contents of fishes determined by flow cytometry. La Kromosomo II 1990, 57, 1871–1888. [Google Scholar]
- Kim, D.S. Karyotypes of nine species of the Korean catfishes (Teleostomi: Siluriformes). Korean J. Genet. 1982, 4, 57–68. [Google Scholar]
- Yunhan, H.; Tun, Z. Karyotypic study of two species of Chinese catfishes (Siluridae). J. Wuhan Univ. Nat. Sci. Ed. 1983, 3. [Google Scholar]
- Yu, X.J.; Zhou, T.; Li, Y.C.; Li, K.; Zhou, M. Chromosomes of Chinese Freshwater Fishes; Science Press: Beijing, China, 1989. [Google Scholar]
- Cui, J.X.; Ren, X.H.; Yu, Q.X. Nuclear DNA content variation in fishes. Cytologia. 1991, 56, 425–429. [Google Scholar]
- Zhang, S.M. Nucleolar organizer regions in four species of fishes. Chromosom. Inf. Serv. 1992, 53, 6–8. [Google Scholar]
- Li, Y.; Li, K.; Zhou, T. Cellular DNA content of fourteen species of freshwater fishes. Acta Genet. Sin. 1983, 10, 384–389. [Google Scholar]
- Kim, D.S.; Kim, I.B.; Huh, H.T.; Park, I. Cytogenetic analysis of catfish, Silurus asotus (Teleostomi: Siluriformes). Ocean Polar Res. 1988, 10, 33–37. [Google Scholar]
- Yin, H.B.; Sun, Z.W.; Pan, W. The karyotype study on triploid catfish (Silurus asotus). J. Fish. China 1996, 20, 178–182. [Google Scholar]
- Meszáros, B.; Bozhko, S.; Marian, T.; Krasznai, Z. About karyotype of Silurus glanis. Halászat 1975, 21, 110–111. [Google Scholar]
- Krasznai, Z. Silurus glanis kariólogial és szerologiai vizsgálatának eredményei. Results of karyological and serological studies in catfish Silurus glanis. Halászat. 1978, 24, 2–4. [Google Scholar]
- Ráb, P. Karyotype of European catfish Silurus glanis (Siluridae, Pisces), with remarks on cytogenetics of siluroid fishes. Folia Zool. 1981, 32, 271–286. [Google Scholar]
- Sofradzija, A. Chromosomes of the species Silurus glanis L. 1758 (Siluridae, Pisces). Genetics 1982, 14, 103–110. [Google Scholar]
- Vujosevic, M.; Zivkovic, C.; Desanka, R.; Jurisic, S.; Cakic, P. The chromosomes of 9 fish species from Dunav basin in Yugoslavia. Ichthyologia 1983, 15, 29–40. [Google Scholar]
- Vasilev, V.P. Evolutionary Karyology of Fishes; Nauka Publishers: Moscow, Russia, 1985. [Google Scholar]
- Al-Sabti, K. Cytogenetic studies on five species of Pisces from Yugoslavia. Cytobios 1987, 49, 175–188. [Google Scholar]
- Ráb, P.; Mayr, B.; Roth, P. Chromosome banding study of European catfish, Silurus glanis (Pisces, Siluridae). Genetica 1991, 83, 153–157. [Google Scholar]
- Hu, G.; Sun, G.; Zhang, Y.; Gu, Q.; Li, G.; Tang, H.; Fan, L. Karyotype analyses of northern sheatfish (Silurus soldatovi). J. Dalian Fish. Coll. 2005, 20, 318–321. [Google Scholar]
- Rishi, K.K.; Singh, J. Chromosomal analysis of the Indian silurid, Wallago attu (Schneider) (Family: Siluridae). Chrom Inf. Serv. 1983, 34, 10–11. [Google Scholar]
- Sharma, O.P.; Tripathi, N.K. Somatic chromosome analysis of Wallago attu, a siluroid fish from Jammu (J & K), India. Perspect. Cytol. Genet. 1984, 4, 437–439. [Google Scholar]
- Oliveira, C.; Gosztonyi, A.E. A cytogenetic study of Diplomystes mesembrinus (Teleostei, Siluriformes, Diplomystidae) with a discussion of chromosome evolution in Siluriforms. Caryologia. 2000, 53, 31–37. [Google Scholar] [CrossRef]
- Kappas, I.; Vittas, S.; Pantzartzi, C.N.; Drosopoulou, E.; Scouras, Z.G. A time-calibrated mitogenome phylogeny of catfish (Teleostei: Siluriformes). PLoS ONE 2016, 11, e0166988. [Google Scholar] [CrossRef] [PubMed]
- White, M.J.D. Chain processes in chromosomal speciation. Syst. Zool. 1978, 27, 285–298. [Google Scholar] [CrossRef]
- King, M. Species Evolution: The Role of Chromosome Change; Cambridge University Press: Cambridge, UK, 1995; ISBN 0521484545. [Google Scholar]
- Potter, S.; Bragg, J.G.; Blom, M.P.K.; Deakin, J.E.; Kirkpatrick, M.; Eldridge, M.D.B.; Moritz, C. Chromosomal speciation in the genomics era: Disentangling phylogenetic evolution of rock-wallabies. Front. Genet. 2017, 8, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Biémont, C.; Vieira, C. Genetics: Junk DNA as an evolutionary force. Nature 2006, 443, 521–524. [Google Scholar] [CrossRef]
- Sochorová, J.; Garcia, S.; Gálvez, F.; Symonová, R.; Kovařík, A. Evolutionary trends in animal ribosomal DNA loci: Introduction to a new online database. Chromosoma 2018, 127, 141–150. [Google Scholar] [CrossRef]
- Gornung, E. Twenty years of physical mapping of major ribosomal RNA genes across the teleosts: A review of research. Cytogenet. Genome Res. 2013, 141, 90–102. [Google Scholar] [CrossRef]
- Rebordinos, L.; Cross, I.; Merlo, A. High evolutionary dynamism in 5S rDNA of fish: State of the art. Cytogenet Genome Res. 2013, 141, 103–113. [Google Scholar] [CrossRef]
- Pereira, C.S.A.; Aboim, M.A.; Ráb, P.; Collares-Pereira, M.J. Introgressive hybridization as a promoter of genome reshuffling in natural homoploid fish hybrids (Cyprinidae, Leuciscinae). Heredity 2014, 112, 343–350. [Google Scholar] [CrossRef]
- Sember, A.; Bohlen, J.; Šlechtová, V.; Altmanová, M.; Symonová, R.; Ráb, P. Karyotype differentiation in 19 species of river loach fishes (Nemacheilidae, Teleostei): Extensive variability associated with rDNA and heterochromatin distribution and its phylogenetic and ecological interpretation. BMC Evol. Biol. 2015, 15, 251–272. [Google Scholar] [CrossRef]
- Maneechot, N.; Yano, C.F.; Bertollo, L.A.C.; Getlekha, N.; Molina, W.F.; Ditcharoen, S.; Tengjaroenkul, B.; Supiwong, W.; Tanomtong, A.; De Bello Cioffi, M. Genomic organization of repetitive DNAs highlights chromosomal evolution in the genus Clarias (Clariidae, Siluriformes). Mol. Cytogenet. 2016. [Google Scholar] [CrossRef]
- Symonová, R.; Howell, W. Vertebrate genome evolution in the light of fish cytogenomics and rDNAomics. Genes 2018, 9, 96. [Google Scholar] [CrossRef]
- Boron, A.; Porycka, K.; Ito, D.; Abe, S.; Kirtiklis, L. Comparative molecular cytogenetic analysis of three Leuciscus species (Pisces, Cyprinidae) using chromosome banding and FISH with rDNA. Genetica 2009, 135, 199. [Google Scholar] [CrossRef]
- Cioffi, M.B.; Martins, C.; Bertollo, L.A. Chromosome spreading of associated transposable elements and ribosomal DNA in the fish Erythrinus erythrinus. Implications for genome change and karyoevolution in fish. BMC Evol. Biol. 2010, 10, 271–279. [Google Scholar] [CrossRef]
- Raskina, O.; Belyayev, A.; Nevo, E. Activity of the En/Spm-like transposons in meiosis as a base for chromosome repatterning in a small, isolated, peripheral population of Aegilops speltoides Tausch. Chromosom. Res. 2004, 12, 153–161. [Google Scholar] [CrossRef]
- Raskina, O.; Barber, J.C.; Nevo, E.; Belyayev, A. Repetitive DNA and chromosomal rearrangements: Speciation-related events in plant genomes. Cytogenet. Genome Res. 2008, 120, 351–357. [Google Scholar] [CrossRef]
- Eickbush, T.H.; Eickbush, D.G. Finely orchestrated movements: Evolution of the ribosomal RNA genes. Genetics 2007, 175, 477–485. [Google Scholar] [CrossRef]
- Glugoski, L.; Giuliano-Caetano, L.; Moreira-Filho, O.; Vicari, M.R.; Nogaroto, V. Co-located hAT transposable element and 5S rDNA in an interstitial telomeric sequence suggest the formation of Robertsonian fusion in armored catfish. Gene 2018, 650, 49–54. [Google Scholar] [CrossRef]
- Slijepcevic, P. Telomeres and mechanisms of Robertsonian fusion. Chromosoma 1998, 107, 136–140. [Google Scholar] [CrossRef]
- Ruiz-Herrera, A.; Nergadze, S.G.; Santagostino, M.; Giulotto, E. Telomeric repeats far from the ends: Mechanisms of origin and role in evolution. Cytogenet. Genome Res. 2008, 122, 219–228. [Google Scholar] [CrossRef]
- Ocalewicz, K. Telomeres in fishes. Cytogenet. Genome Res. 2013, 141, 114–125. [Google Scholar] [CrossRef]
- Charlesworth, B.; Snlegowski, P.; Stephan, W. The evolutionary dynamics of repetitive DNA in eukaryotes. Nature 1994, 371. [Google Scholar] [CrossRef]
- López-Flores, I.; Garrido-Ramos, M.A. The repetitive DNA content of eukaryotic genomes. In Repetitive DNA.; Garrido-Ramos, M.A., Ed.; Karger Publishers: Basel, Switzerland, 2012; Volume 7, pp. 1–28. [Google Scholar]
- Vicari, M.R.; Artoni, R.F.; Bertollo, L.A.C. Comparative cytogenetics of Hoplias malabaricus (Pisces, Erythrinidae). A population analysis in adjacent hydrographic basins. Genet. Mol. Biol. 2005, 28, 103–110. [Google Scholar] [CrossRef]
- Da Rosa, R.; Laforga Vanzela, A.L.; Rubert, M.; Martins-Santos, I.C.; Giuliano-Caetano, L. Differentiation of Y chromosome in the X1 × 1X 2 × 2/X1 × 2Y sex chromosome system of Hoplias malabaricus (Characiformes, Erythrinidae). Cytogenet. Genome Res. 2010, 127, 54–60. [Google Scholar] [CrossRef]
- Cioffi, M.B.; Martins, C.; Bertollo, L.A.C. Comparative chromosome mapping of repetitive sequences. Implications for genomic evolution in the fish, Hoplias malabaricus. BMC Genet. 2009, 10, 34. [Google Scholar]
- De Oliveira, E.A.; Sember, A.; Bertollo, L.A.C.; Yano, C.F.; Ezaz, T.; Moreira-Filho, O.; Hatanaka, T.; Trifonov, V.; Liehr, T.; Al-Rikabi, A.B.H.; et al. Tracking the evolutionary pathway of sex chromosomes among fishes: Characterizing the unique XX/XY1Y2 system in Hoplias malabaricus (Teleostei, Characiformes). Chromosoma 2018, 127, 115–128. [Google Scholar] [CrossRef]
- Eyo, J.E. Cytogenetic variations in Clarias species (Clariidae: Siluriformes) of the Anambra river using leucocytes culture tecniques. Anim. Res. Int. 2005, 2, 275–286. [Google Scholar]
- Arai, R. Fish Karyotype A Check List; Springer Japan: Tokio, Japan, 2011. [Google Scholar]
- Bertollo, L.A.C.; Cioffi, M.B.; Moreira-Filho, O. Direct chromosome preparation from freshwater teleost fishes. In Fish Cytogenetic Techniques (Chondrichthyans and Teleosts); Ozouf-Costaz, C., Pisano, E., Foresti, F., Almeida Toledo, L.F., Eds.; CRC Press: Boca Raton, FL, USA, 2015; pp. 21–26. [Google Scholar]
- Yano, C.F.; Bertollo, L.A.C.; Cioffi, M.B. Fish-FISH: Molecular cytogenetics in fish species. In Fluorescence in Situ Hybridization (FISH)—Application Guide; Liehr, T., Ed.; Springer: Berlin, Germany, 2017; pp. 429–444. [Google Scholar]
- Pendás, A.M.; Móran, P.; Freije, J.P.; Garcia-Vásquez, E. Chromosomal location and nucleotide sequence of two tandem repeats of the Atlantic salmon 5S rDNA. Cytogenet. Cell. Genet. 1994, 67, 31–36. [Google Scholar] [CrossRef]
- Cioffi, M.B.; Martins, C.; Centofante, L.; Jacobina, U.; Bertollo, L.A.C. Chromosomal variability among allopatric populations of Erythrinidae fish Hoplias malabaricus: Mapping of three classes of repetitive DNAs. Cytogenet. Genome Res. 2009, 125, 132–141. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2001. [Google Scholar]
- Zwick, M.S.; Hanson, R.E.; Mcknight, T.D.; Islam-Faridi, M.H.; Stelly, D.M.; Wing, R.A.; Price, H.J. A rapid procedure for the isolation of C 0 t-1 DNA from plants. Genome 1997, 40, 138–142. [Google Scholar] [CrossRef]
- Levan, A.; Fredga, K.; Sandberg, A.A. Nomenclature for centromeric position on chromosomes. Hereditas 1964, 52, 201–220. [Google Scholar] [CrossRef]
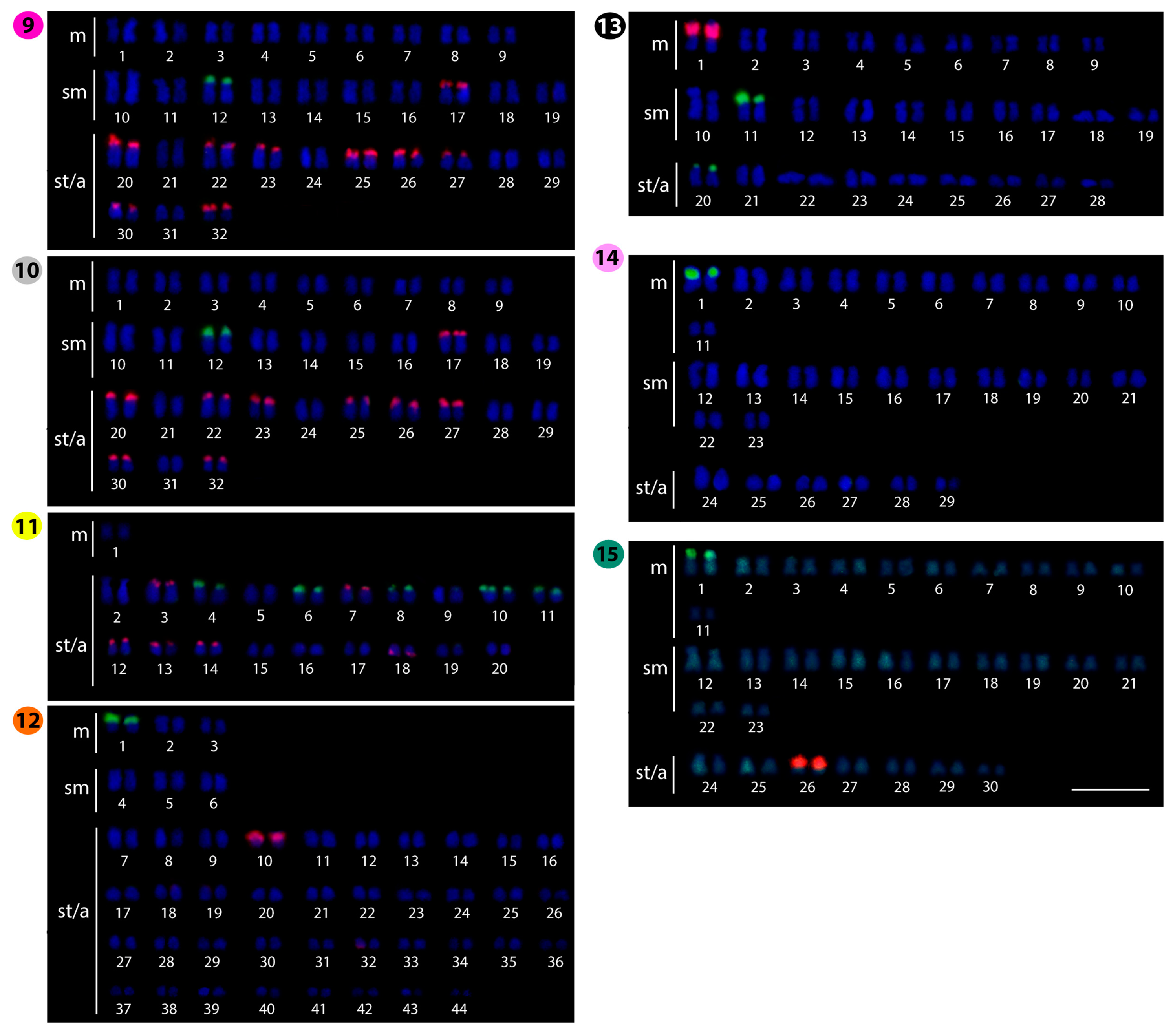
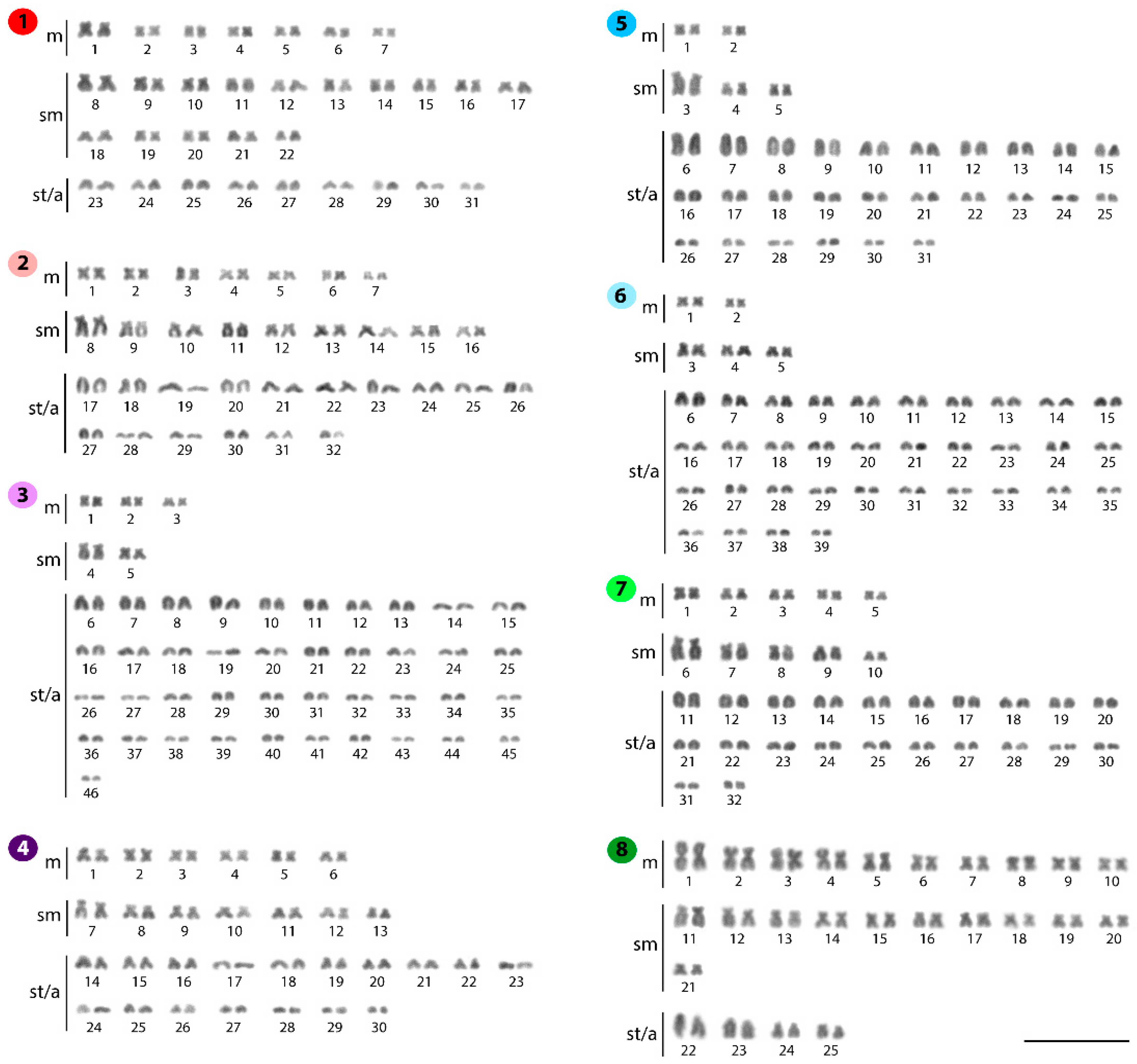
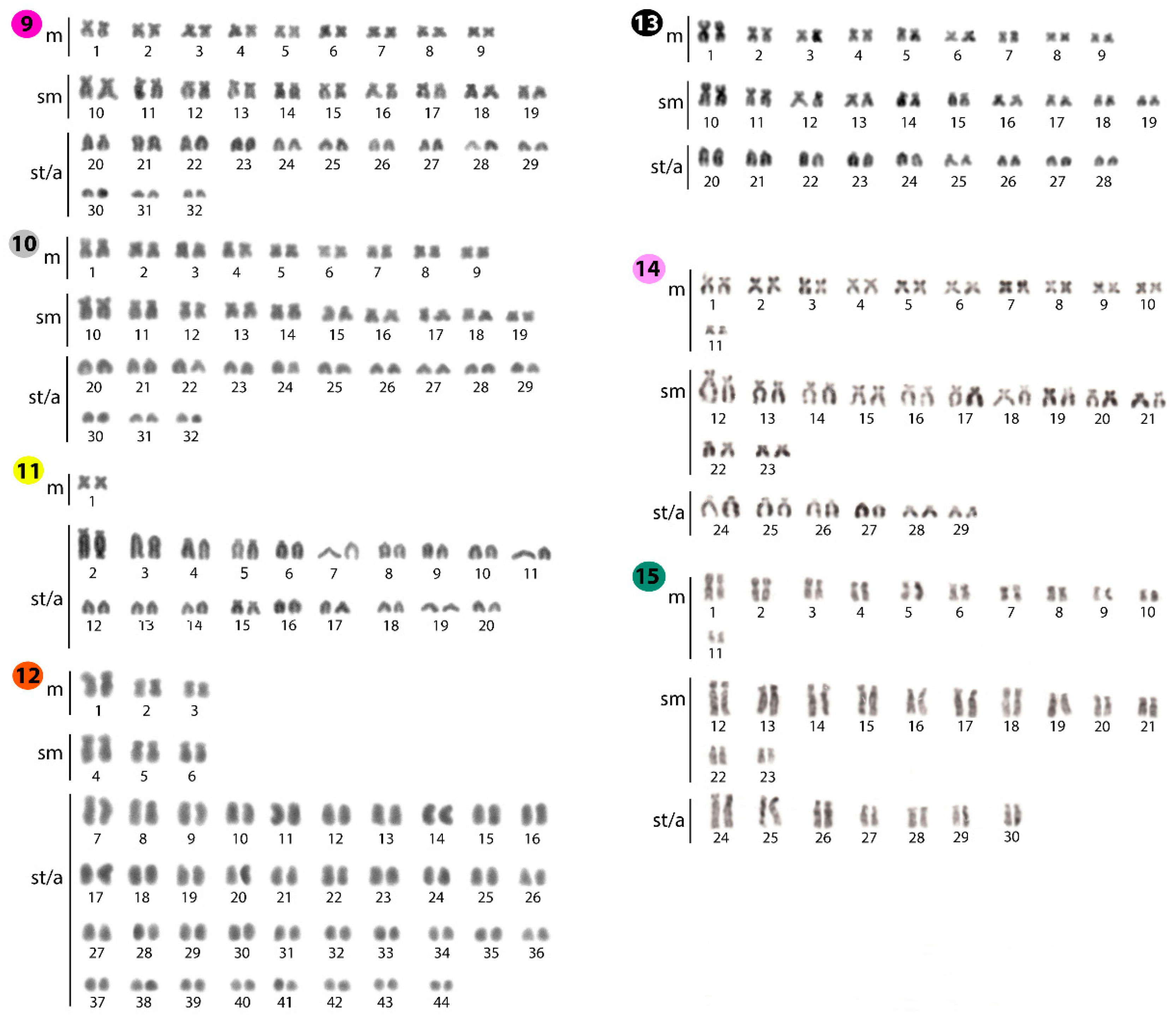



| Species | Locality | 2n | NF | Karyotype | NORs/18S rDNA | Reference |
|---|---|---|---|---|---|---|
| Belodontichthys truncatus | Thailand | 62 | 100 | 20m+10sm+8st+24a | - | [29] |
| B. truncatus | Thailand | 62 | 112 | 14m+30sm+6a+12t | 2 | [30] |
| B. truncatus | Thailand | 62 | 106 | 14m+30sm+18st/a | 2 | Present work |
| Kryptopterus bicirrhis | SE Asia | 60 | - | - | - | [31] |
| K. bicirrhis | Thailand | 64 | 98 | 20m+10sm+4st+30a | - | [29] |
| K. bicirrhis | Thailand | 64 | 96 | 14m+18sm+32st/a | 2 | Present work |
| K. cryptopterus | Thailand | 92 | 110 | 8m+10st+74a | - | [28] |
| K. geminus | Thailand | 92 | 102 | 6m+4sm+82st/a | 4 | Present work |
| K. limpok | Thailand | 60 | 86 | 12m+14sm+34st/a | 2 | Present work |
| K. macrocephalus | Thailand | 62 | 98 | 24m+12sm+26a | - | [32] |
| K. macrocephalus | Thailand | 62 | 72 | 4m+6sm+52st/a | 2 | Present work |
| Micronema cheveyi | Thailand | 78 | 96 | 4m+6sm+10a+58t | 2 | [30] |
| M. cheveyi | Thailand | 78 | 96 | 4m+6sm+10a+58t | 2 | [33] |
| M. cheveyi | Thailand | 78 | 88 | 4m+6sm+68st/a | 2 | Present work |
| Ompok bimaculatus | India | 42 | 72 | 18m+12sm+12a (F) XX | - | [34] |
| O. bimaculatus | India | 41 | 70 | 17m+12sm+12a (M) XY | - | [34] |
| O. bimaculatus | India | 42 | 72 | 6m+24sm+12a (F) | 2 | [35] |
| O. bimaculatus | India | 41 | 70 | 5m+24sm+12a (M) | 2 | [35] |
| O. bimaculatus | India | 42 | 68 | 12m+14sm+16a/t | - | [36] |
| O. bimaculatus | India | 54 | 102 | 16m+26sm+6a+6t | 2 | [26] |
| O. bimaculatus | Thailand | 50 | 90 | 14m+20sm+6a+10t | 2 | [37] |
| O. fumidus | Thailand | 60 | 82 | 20m+2sm+2st+36a | - | [32] |
| O. fumidus | Thailand | 64 | 88 | 10m+10sm+44st/a | 2 | Present work |
| O. pabda | India | 54 | 100 | 28m+10sm+8st+8a | - | [38] |
| O. pabda | India | 42 | - | - | - | [39] |
| O. pabda | India | 42 | 68 | 12m+14sm+16a/t | - | [36] |
| O. pabda | India | 42 | 84 | 26m+10sm+6a | 2 | [26] |
| O. pabo | India | 54 | 102 | 36m+12sm+6a | - | [40] |
| O. siluriodes | Thailand | 50 | 88 | 34m+2sm+2st+12a | - | [28] |
| O. siluriodes | Thailand | 50 | 96 | 20m+22sm+4a+4t | 2 | [30] |
| O. siluriodes | Thailand | 50 | 92 | 20m+22sm+8st/a | 4 | Present work |
| Phalacronotus apogon | Thailand | 64 | 108 | 18m+20sm+6a+20t | 2 | [41] |
| P. apogon | Thailand | 64 | 102 | 18m+20sm+26st/a | 2 | Present work |
| P. bleekeri | Thailand | 64 | 92 | 20m+6sm+2st+36a | - | [28] |
| P. bleekeri | Thailand | 64 | 106 | 14m+22sm+6a+22t | 2 | [33] |
| P. bleekeri | Thailand | 64 | 102 | 18m+20sm+26st/a | 2 | Present work |
| Silurichthys phaiosoma | Thailand | 40 | 46 | 2m+4sm+8st+26a | - | [32] |
| S. phaiosoma | Thailand | 40 | 42 | 2m+38st/a | 10 | Present work |
| S. schneideri | Thailand | 40 | 50 | 6m+4sm+4st+26a | - | [32] |
| Silurus aristotelis | Greece | 58 | 116 | 30m+20sm+8st | - | [42] |
| Si. aristotelis | Greece | 58 | 102 | 20m+24sm+14st/a | 2 | [43] |
| Si. aristotelis | Czech | 58 | 104 | 22m+24sm+12st/a | 2 | Present work |
| Si. asotus | - | 58 | - | 58t | - | [44] |
| Si. asotus | Japan | 58 | 104 | 38m/sm+8st+12a | - | [45] |
| Si. asotus | Japan | 58 | 104 | 46m/sm+12st/a | 2 | [46] |
| Si. asotus | Japan | 58 | 102 | 44m/sm+14st/a | - | [47,48] |
| Si. asotus | Korea | 58 | 106 | 24m+24sm+10st/a | - | [49] |
| Si. asotus | China | 58 | 102 | 20m+24sm+10st+4a | - | [50,51,52] |
| Si. asotus | China | 58 | 98 | 20m+14sm+6st+18a | 2 | [53,54] |
| Si. asotus | Mongolia | 58 | - | 42m/sm+16st/a | - | [27] |
| Si. asotus | Korea | 58 | 106 | - | - | [55] |
| Si. asotus | China | 58 | 112 | 20m+24sm+10st+4a | - | [56] |
| Si. biwaensis | Japan | 58 | 102 | 44m/sm+14st/a | - | [47] |
| Si. glanis | - | 60 | 100 | 40m/sm/st+20a | - | [57] |
| Si. glanis | - | 60 | 100 | 40m/sm+20a | - | [58] |
| Si. glanis | Czech | 60 | 120 | 28m+26sm+6st | - | [59] |
| Si. glanis | - | 60 | 98 | 38m/sm+22a | - | [60] |
| Si. glanis | Serbia | 60 | 94 | 16m+18sm+14st+12a | - | [61] |
| Si. glanis | Russia | 60 | 110 | 18m+32sm/st+10a | - | [62] |
| Si. glanis | Serbia | 48 | 78 | 30m/sm+18st/a | - | [63] |
| Si. glanis | Czech | 60 | 120 | 22m+38sm/st | 2 | [64] |
| Si. glanis | Czech | 60 | 106 | 22m+24sm+14st/a | 2 | Present work |
| Si. lithophilus | Japan | 58 | 102 | 44m/sm+14st/a | - | [47] |
| Si. lithophilus | China | 58 | 98 | 20m+20sm+18st/a | - | [50] |
| Si. meridionalis | China | 58 | 112 | 20m+20sm+14st+4a | - | [50,51,52] |
| Si. meridionalis | Korea | 60 | 106 | 22m+24sm+12st/a+2 microchromosomes | - | [49] |
| Si. microdorsalis | Korea | 28 | 56 | 12m+14sm+2st | - | [27] |
| Si. soldatovi | China | 58 | 112 | 24m+16sm+14st+4a | - | [65] |
| Wallago attu | India | 86 | 106 | 12m+6sm+2st+66a | - | [66] |
| W. attu | India | 86 | 116 | 10m+12sm+8st+56a | - | [67] |
| W. attu | Thailand | 88 | 110 | 16m+2sm+4st+66a | - | [29] |
| W. attu | Thailand | 88 | 108 | 6m+6sm+8a+68t | 2 | [30] |
| W. attu | Thailand | 88 | 100 | 6m+6sm+76st/a | 2 | Present work |
| W. micropogon | Thailand | 56 | 86 | 26m+4sm+26a | - | [29] |
| W. micropogon | Thailand | 56 | 100 | 18m+20sm+6a+12t | 2 | [30] |
| W. micropogon | Thailand | 56 | 94 | 18m+20sm+18st/a | 4 | Present work |
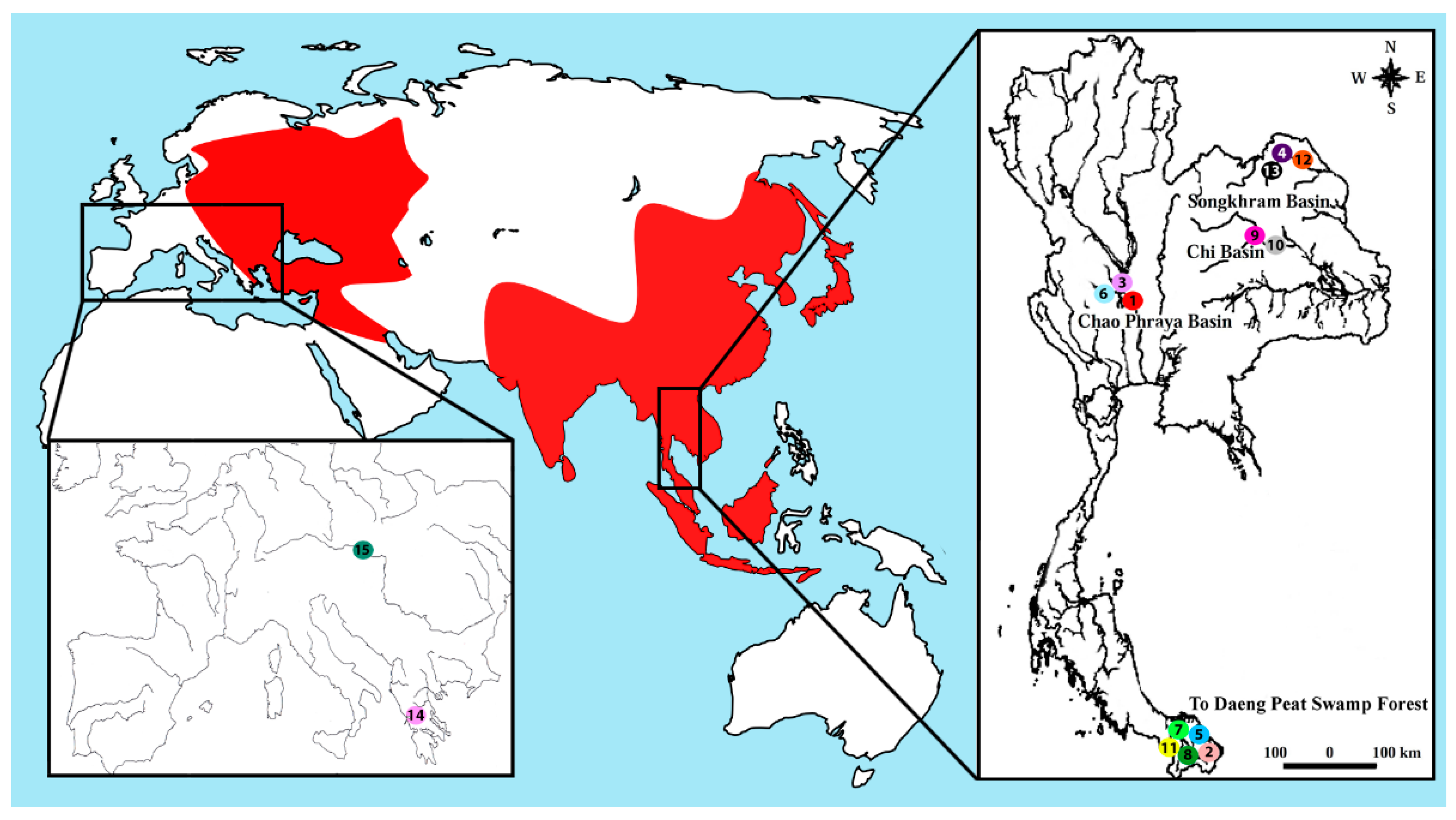
| Species | Locality | No. of Individuals |
|---|---|---|
| Belodontichthys truncatus | Chao Phraya Basin (Thailand) (site 1) | (04 ♀; 04 ♂) |
| Kryptopterus bicirrhis | To Daeng peat swamp forest (Thailand) (site 2) | (07 ♀; 08 ♂) |
| Kryptopterus geminus | Chao Phraya Basin (Thailand) (site 3) | (08 ♀; 11 ♂) |
| Kryptopterus limpok | Songkhram Basin (Thailand) (site 4) | (07 ♀; 10 ♂) |
| Kryptopterus macrocephalus | To Daeng peat swamp forest (Thailand) (site 5) | (06 ♀; 06 ♂) |
| Micronema cheveyi | Chao Phraya Basin (Thailand) (site 6) | (09 ♀; 10 ♂) |
| Ompok fumidus | To Daeng peat swamp forest (Thailand) (site 7) | (05 ♀; 07 ♂) |
| Ompok siluroides | To Daeng peat swamp forest (Thailand) (site 8) | (04 ♀; 05 ♂) |
| Phalacronotus apogon | Chi Basin (Thailand) (site 9) | (06 ♀; 05 ♂) |
| Phalacronotus bleekeri | Chi Basin (Thailand) (site 10) | (07 ♀; 04 ♂) |
| Silurichthys phaiosoma | To Daeng peat swamp forest (Thailand) (site 11) | (04 ♀; 06 ♂) |
| Wallago attu | Songkhram Basin (Thailand) (site 12) | (03 ♀; 04 ♂) |
| Wallago micropogon | Songkhram Basin (Thailand) (site 13) | (04 ♀; 04 ♂) |
| Silurus aristotelis | Trichonida Lake (Greece) (site 14) | (03 ♀; 05 ♂) |
| Silurus glanis | Dyje River, Danube basin (Czech republic) (site 15) | (08 ♀; 06 ♂) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ditcharoen, S.; Antonio Carlos Bertollo, L.; Ráb, P.; Hnátková, E.; Franco Molina, W.; Liehr, T.; Tanomtong, A.; Triantaphyllidis, C.; Ozouf-Costaz, C.; Tongnunui, S.; et al. Genomic Organization of Repetitive DNA Elements and Extensive Karyotype Diversity of Silurid Catfishes (Teleostei: Siluriformes): A Comparative Cytogenetic Approach. Int. J. Mol. Sci. 2019, 20, 3545. https://doi.org/10.3390/ijms20143545
Ditcharoen S, Antonio Carlos Bertollo L, Ráb P, Hnátková E, Franco Molina W, Liehr T, Tanomtong A, Triantaphyllidis C, Ozouf-Costaz C, Tongnunui S, et al. Genomic Organization of Repetitive DNA Elements and Extensive Karyotype Diversity of Silurid Catfishes (Teleostei: Siluriformes): A Comparative Cytogenetic Approach. International Journal of Molecular Sciences. 2019; 20(14):3545. https://doi.org/10.3390/ijms20143545
Chicago/Turabian StyleDitcharoen, Sukhonthip, Luiz Antonio Carlos Bertollo, Petr Ráb, Eva Hnátková, Wagner Franco Molina, Thomas Liehr, Alongklod Tanomtong, Costas Triantaphyllidis, Catherine Ozouf-Costaz, Sampan Tongnunui, and et al. 2019. "Genomic Organization of Repetitive DNA Elements and Extensive Karyotype Diversity of Silurid Catfishes (Teleostei: Siluriformes): A Comparative Cytogenetic Approach" International Journal of Molecular Sciences 20, no. 14: 3545. https://doi.org/10.3390/ijms20143545
APA StyleDitcharoen, S., Antonio Carlos Bertollo, L., Ráb, P., Hnátková, E., Franco Molina, W., Liehr, T., Tanomtong, A., Triantaphyllidis, C., Ozouf-Costaz, C., Tongnunui, S., Pengseng, P., Supiwong, W., Aroutiounian, R., & de Bello Cioffi, M. (2019). Genomic Organization of Repetitive DNA Elements and Extensive Karyotype Diversity of Silurid Catfishes (Teleostei: Siluriformes): A Comparative Cytogenetic Approach. International Journal of Molecular Sciences, 20(14), 3545. https://doi.org/10.3390/ijms20143545





