Deciphering the Origin and Evolution of the X1X2Y System in Two Closely-Related Oplegnathus Species (Oplegnathidae and Centrarchiformes)
Abstract
1. Introduction
2. Results
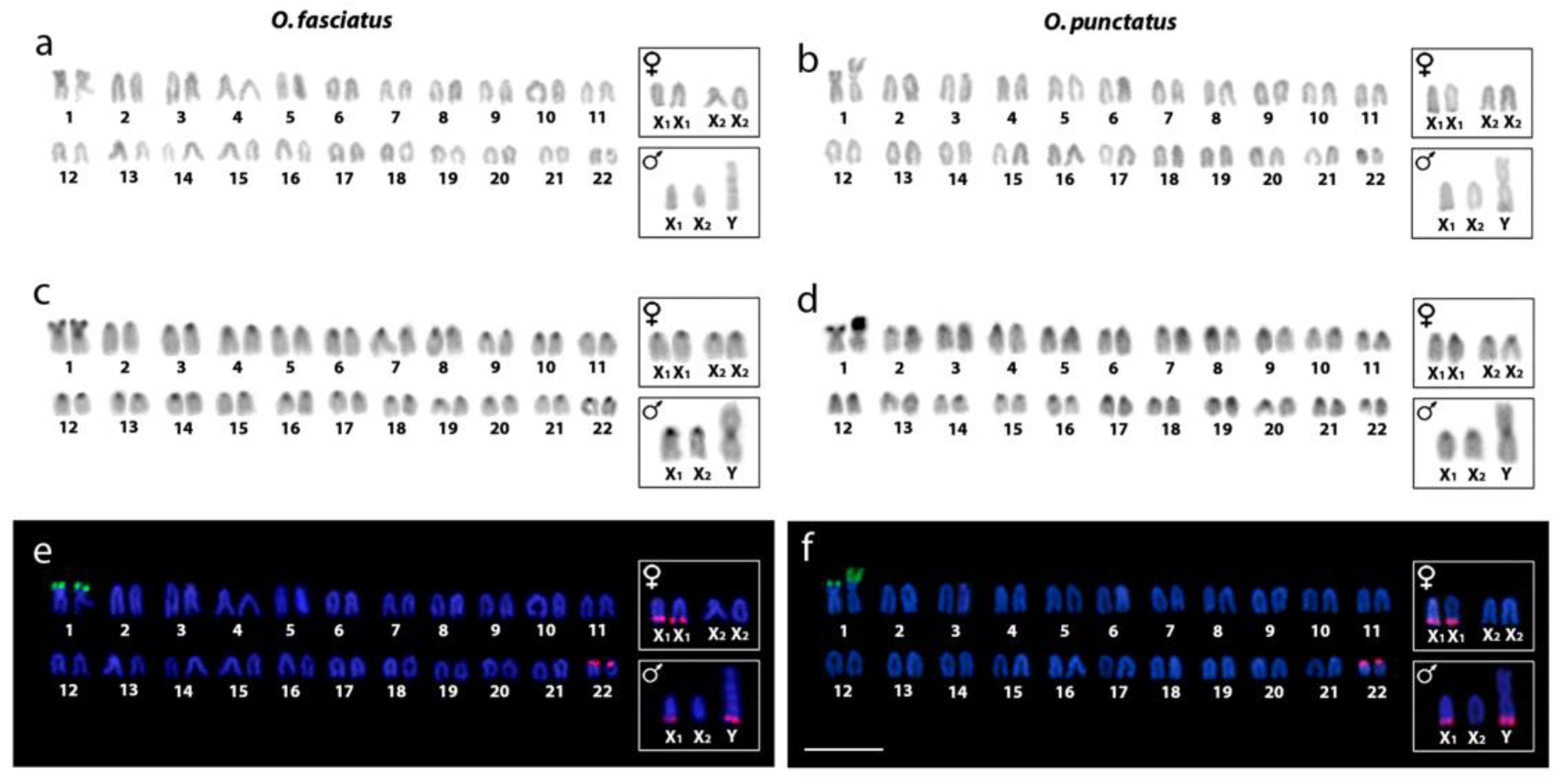
2.1. Karyotype Analysis and Distribution of Constitutive Heterochromatin
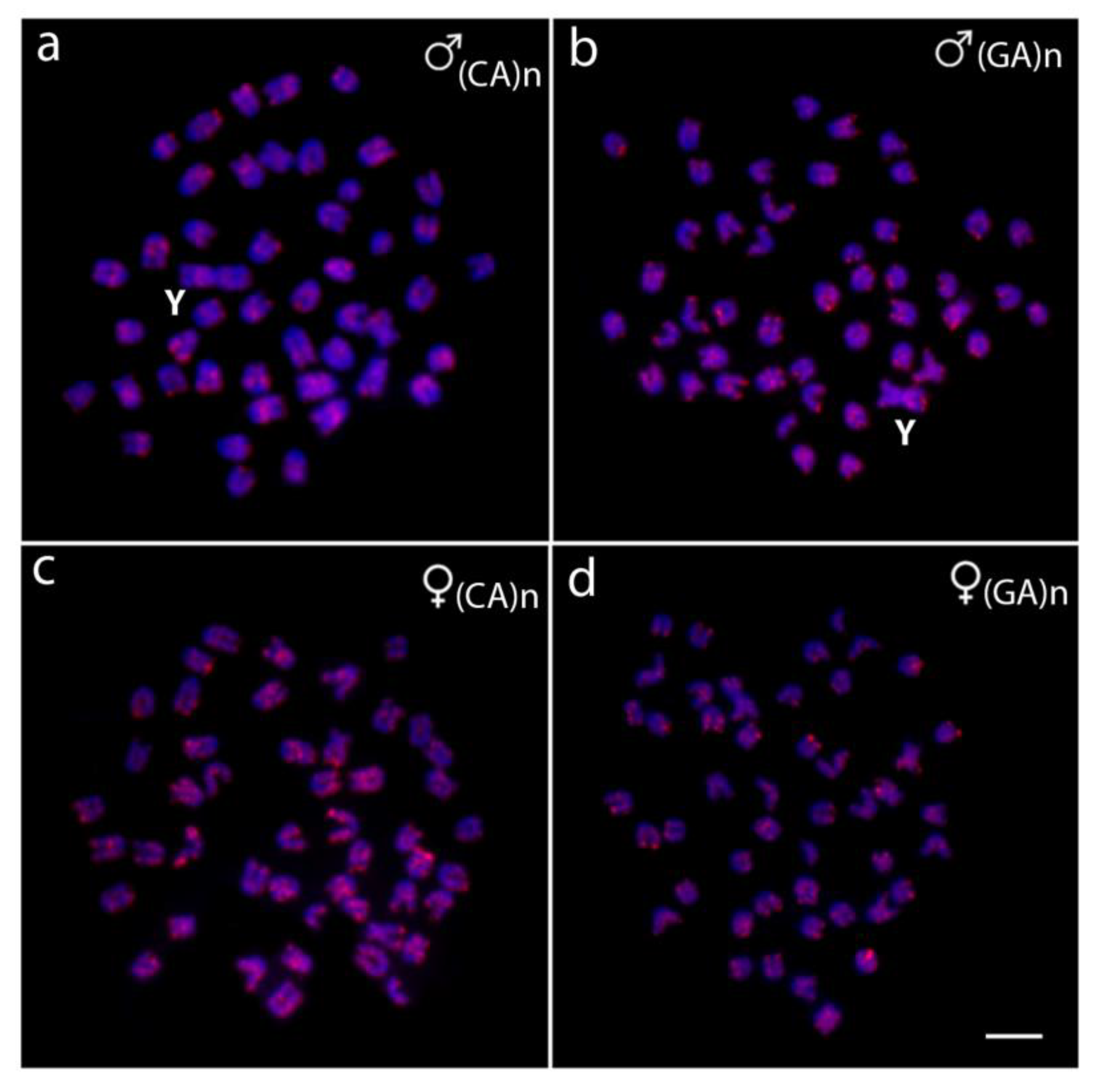
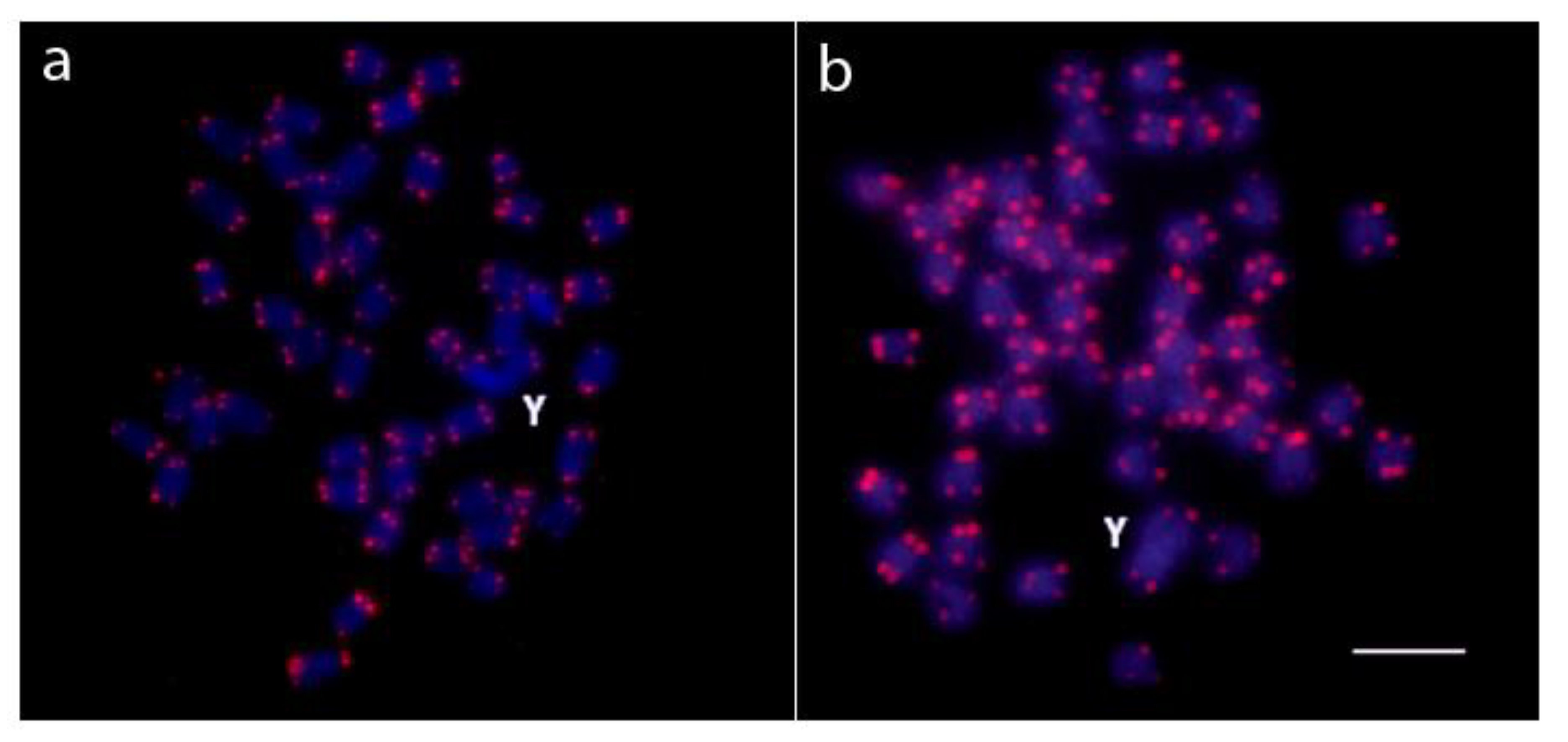
2.2. Chromosomal Mapping of Repetitive DNA Markers
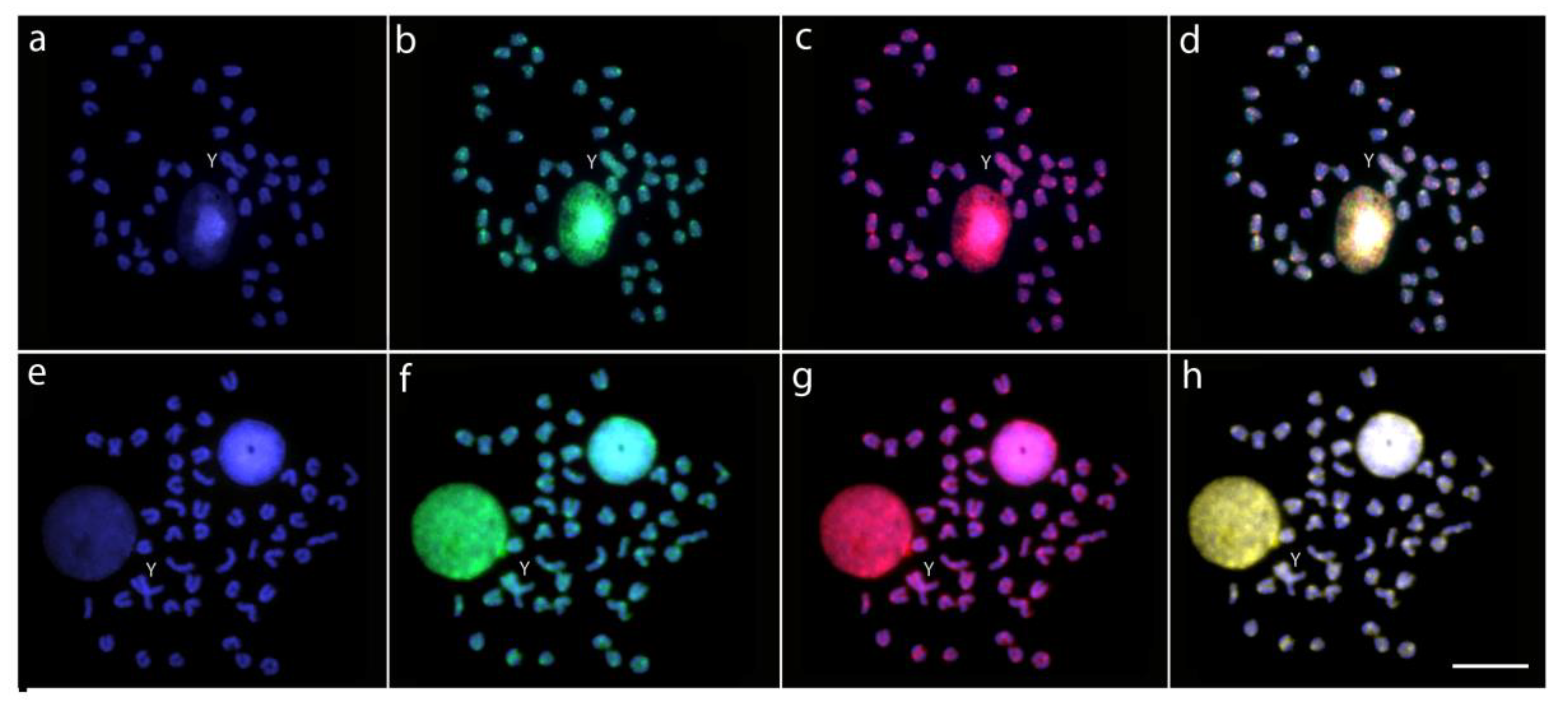
2.3. Characterization of Male vs. Female Genome Differences by CGH
2.4. Detection of Chromosomal Homologies by WCP Experiments
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Animals
5.2. Chromosome Preparation and Analysis of Constitutive Heterochromatin
5.3. FISH with Repetitive DNA Sequences
5.4. Preparation of Probes for Comparative Genomic Hybridization (CGH)
5.5. Chromosome Microdissection, Probe Preparation, and Labeling
5.6. Microscopy and Image Processing
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Fricke, R.; Eschmeyer, W.; van der Laan, R. Eschmeyer’s Catalog of Fishes: Genera, Species, References, California Academy of Sciences; California Academy of Sciences: San Francisco, CA, USA, 2019; Electronic Version; Available online: http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp (accessed on 20 April 2019).
- Meng, Q.W.; Su, J.X.; Miao, X.Z. Fish Taxonomy; China Agriculture Press: Beijing, China, 1995. [Google Scholar]
- Xiao, Z.Z.; Xiao, Y.S.; Ma, D.Y.; Xu, S.H.; Liu, Q.H.; Li, J. Relationship between Oplegnathus fasciatus and Oplegnathus punctatus revealed by mtDNA sequences. Acta Ocean. Sin. 2011, 33, 115–123. [Google Scholar]
- Xiao, Y.; Li, J.; Ren, G.; Ma, D.; Wang, Y.; Xiao, Z.; Xu, S. Pronounced population genetic differentiation in the rock bream Oplegnathus fasciatus inferred from mitochondrial DNA sequences. Mitochondrial DNA A 2016, 27, 2045–2052. [Google Scholar]
- Shin, Y.; Jung, M.; Shin, G.H.; Jung, H.J.; Baek, S.J.; Lee, G.Y.; Kang, B.C.; Shim, J.; Hong, J.M.; Park, J.Y.; et al. First draft genome sequence of the rock bream in the family Oplegnathidae. Sci. Data 2018, 5, 180234. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Xiao, Z.; Ma, D.; Liu, J.; Li, J. Genome sequence of the barred knifejaw Oplegnathus fasciatus (Temminck & Schlegel, 1844): The first chromosome-level draft genome in the family Oplegnathidae. Gigascience 2019, 8, giz013. [Google Scholar] [PubMed]
- Xu, D.; You, F.; Lou, B.; Geng, Z.; Li, J.; Xiao, Z.Z. Comparative analysis of karyotype and C-banding in male and female Oplegnathus fasciatus. Acta Hydrobiol. Sin. 2012, 36, 552–557. (In Chinese) [Google Scholar]
- Xue, R.; An, H.; Liu, Q.H.; Xiao, Z.Z.; Wang, Y.F.; Li, J. Karyotype and Ag-NORs in male and female of Oplegnathus punctatus. Ocean. Limnol. Sin. 2016, 47, 626–632. [Google Scholar]
- Li, P.Z.; Cao, D.D.; Liu, X.B.; Wang, Y.J.; Yu, H.Y.; Li, X.J.; Zhang, Q.Q.; Wang, X.B. Karyotype analysis and ribosomal gene localization of spotted knifejaw Oplegnathus punctatus. Genet. Mol. Res. 2016, 15, gmr15049159. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Lou, B.; Xu, H.; Li, S.; Geng, Z. Isolation and characterization of male-specific DNA markers in the rock bream Oplegnathus fasciatus. Mar. Biotechnol. 2013, 15, 221–229. [Google Scholar] [CrossRef] [PubMed]
- Toder, R.; O’Neill, R.J.; Wienberg, J.; O’Brien, P.C.; Voullaire, L.; Marshall-Graves, J.A. Comparative chromosome painting between two marsupials: Origins of an XX/XY1Y sex chromosome system. Mamm. Genome 1997, 8, 418–422. [Google Scholar] [CrossRef]
- Rens, W.; Grutzner, F.; O’Brien, P.C.M.; Fairclough, H.; Graves, J.A.M.; Ferguson-Smith, M.A. From the cover: Resolution and evolution of the duck-billed platypus karyotype with an X1Y1XYXYXYXY male sex chromosome constitution. Proc. Natl. Acad. Sci. USA 2004, 101, 16257–16261. [Google Scholar] [CrossRef]
- Kitano, J.; Peichel, C.L. Turnover of sex chromosomes and speciation in fishes. Environ. Biol. Fishes 2012, 94, 549–558. [Google Scholar] [CrossRef] [PubMed]
- Moreira-Filho, O.; Bertollo, L.A.C.; Galetti Jr, P.M. Evidences for a multiple sex chromosome system with female heterogamety in Apareiodon affinis (Pisces, Parodontidae). Caryologia 1980, 33, 83–91. [Google Scholar] [CrossRef]
- Král, J.; Kořínková, T.; Krkavcová, L.; Musilová, J.; Forman, M.; Herrera, I.M.Á.; Al, E. Evolution of karyotype, sex chromosomes, and meiosis in mygalomorph spiders (Araneae: Mygalomorphae). Biol. J. Linn. Soc. 2013, 109, 377–408. [Google Scholar] [CrossRef]
- Šíchová, J.; Ohno, M.; Dincă, V.; Watanabe, M.; Sahara, K.; Marec, F. Fissions, fusions, and translocations shaped the karyotype and multiple sex chromosome constitution of the northeast-Asian wood white butterfly, Leptidea amurensis. Biol. J. Linn. Soc. 2016, 118, 457–471. [Google Scholar] [CrossRef]
- Giovannotti, M.; Trifonov, V.A.; Paoletti, A.; Kichigin, I.G.; O’Brien, P.C.; Kasai, F.; Giovagnoli, G.; Ng, B.L.; Ruggeri, P.; Cerioni, P.N.; et al. New insights into sex chromosome evolution in anole lizards (Reptilia, Dactyloidae). Chromosoma 2016, 126, 245–260. [Google Scholar] [CrossRef] [PubMed]
- Howell, E.C.; Armstrong, S.J.; Filatov, D.A. Evolution of neo-sex chromosomes in Silene diclinis. Genetics 2009, 182, 1109–1115. [Google Scholar] [CrossRef] [PubMed]
- Ogata, M.; Hasegawa, Y.; Ohtani, H.; Mineyama, M.; Miura, I. The ZZ/ZW sex-determining mechanism originated twice and independently during evolution of the frog, Rana rugosa. Heredity 2008, 100, 92–99. [Google Scholar] [CrossRef]
- Castillo, E.R.; Marti, D.A.; Bidau, C. Sex and neo-sex chromosomes in Orthoptera: A review. J. Orthoptera Res. 2011, 19, 213–231. [Google Scholar] [CrossRef]
- Nguyen, P.; Sýkorová, M.; Šíchová, J.; Kůta, V.; Dalíková, M.; Frydrychová, R.Č.; Neven, L.G.; Sahara, K.; Marec, F. Neo-sex chromosomes and adaptive potential in tortricid pests. Proc. Natl. Acad. Sci. USA 2013, 110, 6931–6936. [Google Scholar] [CrossRef]
- Palacios-Gimenez, O.M.; Cabral-de-Mello, D.C. Repetitive DNA chromosomal organization in the cricket Cycloptiloides americanus: A case of the unusual XX0 sex chromosome system in Orthoptera. Mol. Genet. Genomics 2015, 290, 623–631. [Google Scholar] [CrossRef]
- Kaiser, V.B.; Bachtrog, D. Evolution of sex chromosomes in insects. Annu. Rev. Genet. 2010, 44, 91–112. [Google Scholar] [CrossRef] [PubMed]
- Ross, J.A.; Urton, J.R.; Boland, J.; Shapiro, M.D.; Peichel, C.L. Turnover of sex chromosomes in the stickleback fishes (Gasterosteidae). PLoS Genet. 2009, 5, e1000391. [Google Scholar] [CrossRef] [PubMed]
- Schartl, M.; Schmid, M.; Nanda, I. Dynamics of vertebrate sex chromosome evolution: From equal size to giants and dwarfs. Chromosoma 2016, 125, 553–571. [Google Scholar] [CrossRef] [PubMed]
- Nanda, I.; Schories, S.; Tripathi, N.; Dreyer, C.; Haaf, T.; Schmid, M.; Schartl, M. Sex chromosome polymorphism in guppies. Chromosoma 2014, 123, 373–383. [Google Scholar] [CrossRef] [PubMed]
- Reichwald, K.; Petzold, A.; Koch, P.; Downie, B.R.; Hartmann, N.; Pietsch, S.; Baumgart, M.; Chalopin, D.; Felder, M.; Bens, M.; et al. Insights into sex chromosome evolution and aging from the genome of a short-lived fish. Cell 2015, 163, 1527–1538. [Google Scholar] [CrossRef] [PubMed]
- Gamble, T. Using RAD-seq to recognize sex-specific markers and sex chromosome systems. Mol. Ecol. 2016, 25, 2114–2116. [Google Scholar] [CrossRef]
- Guiguen, Y.; Fostier, A.; Herpin, A. Sex determination and differentiation in fish: Genetic, genomic, and endocrine aspects. In Sex Control in Aquaculture, 1st ed.; Wang, H.-P., Piferrer, F., Chen, S.-L., Eds.; John Wiley & Sons Ltd.: Hoboken, NJ, USA, 2019; Volume 1. [Google Scholar]
- Blanco, D.R.; Vicari, M.R.; Lui, R.L.; Bertollo, L.A.C.; Traldi, J.B.; Moreira-Filho, O. The role of the Robertsonian rearrangements in the origin of the XX/XYY sex chromosome system and in the chromosomal differentiation in Harttia species (Siluriformes, Loricariidae). Rev. Fish Biol. Fish. 2013, 23, 127–134. [Google Scholar] [CrossRef]
- Blanco, D.R.; Vicari, M.R.; Lui, R.L.; Artoni, R.F.; de Almeida, M.C.; Traldi, J.B.; Margarido, V.P.; Moreira-Filho, O. Origin of the XX1XX/XXY sex chromosome system of Harttia punctata (Siluriformes, Loricariidae) inferred from chromosome painting and FISH with ribosomal DNA markers. Genetica 2014, 142, 119–126. [Google Scholar]
- Soares, R.X.; Bertollo, L.A.C.; Cioffi, M.B.; Costa, G.W.W.F.; Molina, W.F. Chromosomal distribution of two multigene families and the unusual occurrence of an X1X1X2X2 /X1X2Y sex chromosome system in the dolphinfish (Coryphaenidae): An evolutionary perspective. Genet. Mol. Res. 2014, 13, 2470–2479. [Google Scholar] [CrossRef]
- Sember, A.; Bohlen, J.; Šlechtová, V.; Altmanová, M.; Symonová, R.; Ráb, P. Karyotype differentiation in 19 species of river loach fishes (Nemacheilidae, Teleostei): Extensive variability associated with rDNA and heterochromatin distribution and its phylogenetic and ecological interpretation. BMC Evol. Biol. 2015, 15, 251–272. [Google Scholar] [CrossRef]
- Krysanov, E.; Demidova, T. Extensive karyotype variability of African fish genus Nothobranchius (Cyprinodontiformes). Comp. Cytogenet. 2018, 12, 387–402. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, M.; Matoso, D.A.; Artoni, R.F.; Feldberg, E. New approach data in electric fish (Teleostei: Gymnotus): Sex chromosome evolution and repetitive DNA. Zebrafish 2014, 11, 528–535. [Google Scholar] [CrossRef] [PubMed]
- Almeida, J.S.; Migues, V.H.; Diniz, D.; Affonso, P.R.A.M. A unique sex chromosome system in the knifefish Gymnotus bahianus with inferences about chromosomal evolution of Gymnotidae. J. Hered. 2015, 106, 177–183. [Google Scholar] [CrossRef]
- Cardoso, A.L.; Pieczarka, J.C.; Nagamachi, C.Y. X1X1X2X2/X1X2Y sex chromosome systems in the Neotropical Gymnotiformes electric fish of the genus Brachyhypopomus. Genet. Mol. Biol. 2015, 38, 213–219. [Google Scholar] [CrossRef]
- Bitencourt, J.A.; Sampaio, I.; Ramos, R.T.C.; Vicari, M.R.; Affonso, P.R.A.M. First report of sex chromosomes in Achiridae (Teleostei: Pleuronectiformes) with inferences about the origin of the multiple X1X1X2X2/X1X2Y system and dispersal of ribosomal genes in Achirus achirus. Zebrafish 2016, 14, 90–95. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Zheng, J.; Zhang, J.; Wang, Z.; Wang, Y.; Cai, M. Cytogenetic characterization and description of an X1X1X2X2/X1X2Y sex chromosome system in Collichthys lucidus (Richardson, 1844). Acta Oceanol. Sin. 2018, 37, 34–3940. [Google Scholar] [CrossRef]
- Cioffi, M.B.; Bertollo, L.A.C. Chromosomal distribution and evolution of repetitive DNAs in fish. In Repetitive DNA; Garrido-Ramos, M.A., Ed.; Karger Publishers: Basel, Switzerland, 2012; Volume 7, pp. 197–221. [Google Scholar]
- Symonová, R.; Howell, W. Vertebrate genome evolution in the light of fish cytogenomics and rDNAomics. Genes 2018, 9, 96. [Google Scholar] [CrossRef]
- Ferguson-Smith, M.A.; Trifonov, V. Mammalian karyotype evolution. Nat. Rev. Genet. 2007, 8, 950–962. [Google Scholar] [CrossRef]
- Traut, W.; Sahara, K.; Otto, T.D.; Marec, F. Molecular differentiation of sex chromosomes probed by comparative genomic hybridization. Chromosoma 1999, 108, 173–180. [Google Scholar] [CrossRef]
- Altmanová, M.; Rovatsos, M.; Kratochvíl, L.; Johnson Pokorná, M. Minute Y chromosomes and karyotype evolution in Madagascan iguanas (Squamata: Iguania: Opluridae). Biol. J. Linn. Soc. 2016, 118, 618–633. [Google Scholar] [CrossRef]
- Sember, A.; Bertollo, L.A.C.; Ráb, P.; Yano, C.F.; Hatanaka, T.; de Oliveira, E.A.; Cioffi, M.B. Sex chromosome evolution and genomic divergence in the fish Hoplias malabaricus (Characiformes, Erythrinidae). Front. Genet. 2018, 9, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Valenzuela, N.; Badenhorst, D.; Montiel, E.E.; Literman, R. Molecular cytogenetic search for cryptic sex chromosomes in painted turtles Chrysemys picta. Cytogenet. Genome Res. 2014, 144, 39–46. [Google Scholar] [CrossRef] [PubMed]
- Yano, C.F.; Bertollo, L.A.C.; Ezaz, T.; Trifonov, V.; Sember, A.; Liehr, T.; Cioffi, M.B. Highly conserved Z and molecularly diverged W chromosomes in the fish genus Triportheus (Characiformes, Triportheidae). Heredity 2017, 118, 276–283. [Google Scholar] [CrossRef] [PubMed]
- Montiel, E.E.; Badenhorst, D.; Tamplin, J.; Burke, R.L.; Valenzuela, N. Discovery of the youngest sex chromosomes reveals first case of convergent co-option of ancestral autosomes in turtles. Chromosoma 2017, 126, 105–113. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, E.A.; Sember, A.; Bertollo, L.A.C.; Yano, C.F.; Ezaz, T.; Moreira-Filho, O.; Hatanaka, T.; Trifonov, V.; Liehr, T.; Al-Rikabi, A.B.H.; et al. Tracking the evolutionary pathway of sex chromosomes among fishes: Characterizing the unique XX/XY1Y2 system in Hoplias malabaricus (Teleostei, Characiformes). Chromosoma 2018, 127, 115–128. [Google Scholar] [CrossRef] [PubMed]
- Zrzavá, M.; Hladová, I.; Dalíková, M.; Šíchová, J.; Õunap, E.; Kubíčková, S.; Marec, F. Sex chromosomes of the iconic moth Abraxas grossulariata (Lepidoptera, Geometridae) and its congener A. sylvata. Genes 2018, 9, 279. [Google Scholar] [CrossRef] [PubMed]
- Pokorná, M.; Rens, W.; Rovatsos, M.; Kratochvíl, L. A ZZ/ZW sex chromosome system in the thick-tailed gecko (Underwoodisaurus milii; Squamata: Gekkota: Carphodactylidae), a member of the ancient gecko lineage. Cytogenet. Genome Res. 2014, 142, 190–196. [Google Scholar] [CrossRef]
- Pokorná, M.; Giovannotti, M.; Kratochvíl, L.; Caputo, V.; Olmo, E.; Ferguson-Smith, M.A.; Rens, W. Conservation of chromosomes syntenic with avian autosomes in squamate reptiles revealed by comparative chromosome painting. Chromosoma 2012, 121, 409–418. [Google Scholar] [CrossRef]
- Henning, F.; Moysés, C.B.; Calcagnotto, D.; Meyer, A.; Almeida-Toledo, L.F. Independent fusions and recent origins of sex chromosomes in the evolution and diversification of glass knife fishes (Eigenmannia). Heredity 2011, 106, 391–400. [Google Scholar] [CrossRef]
- Carvalho, P.C.; de Oliveira, E.A.; Bertollo, L.A.C.; Yano, C.F.; Oliveira, C.; Decru, E.; Jegede, O.I.; Hatanaka, T.; Liehr, T.; Al-Rikabi, A.B.H.; et al. First chromosomal analysis in Hepsetidae (Actinopterygii, Characiformes): Insights into relationship between African and Neotropical fish groups. Front. Genet. 2017, 8, 203. [Google Scholar] [CrossRef]
- Noronha, R.C.R.; Nagamachi, C.Y.; O’Brien, P.C.M.; Ferguson-Smith, M.A.; Pieczarka, J.C. Neo-XY body: An analysis of XY1Y2 meiotic behavior in Carollia (Chiroptera, Phyllostomidae) by chromosome painting. Cytogenet. Genome Res. 2009, 124, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Cioffi, M.B.; Sánchez, A.; Marchal, J.A.; Kosyakova, N.; Liehr, T.; Trifonov, V.; Bertollo, L.A.C. Whole chromosome painting reveals independent origin of sex chromosomes in closely related forms of a fish species. Genetica 2011, 139, 1065–1072. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, W.O.; da Costa, M.J.; Pieczarka, J.C.; Rissino, J.; Pereira, J.C.; Ferguson-Smith, M.A.; Nagamachi, C.Y. Identification of two independent X-autosome translocations in closely related mammalian (Proechimys) species. Sci. Rep. 2019, 9, 1–11. [Google Scholar]
- Xu, D.; Lou, B.; Bertollo, L.A.C.; Cioffi, M.D.B. Chromosomal mapping of microsatellite repeats in the rock bream fish Oplegnathus fasciatus, with emphasis of their distribution in the neo-Y chromosome. Mol. Cytogenet. 2013, 6, 12. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, M.; Garcia, C.; Matoso, D.A.; de Jesus, I.S.; Feldberg, E. A new multiple sex chromosome system X1X1X2X2/X1Y1X2Y2 in Siluriformes: Cytogenetic characterization of Bunocephalus coracoideus (Aspredinidae). Genetica 2016, 144, 591–599. [Google Scholar] [CrossRef] [PubMed]
- Cioffi, M.B.; Martins, C.; Vicari, M.R.; Rebordinos, L.; Bertollo, L.A.C. Differentiation of the XY sex chromosomes in the fish Hoplias malabaricus (Characiformes, Erythrinidae): Unusual accumulation of repetitive sequences on the X chromosome. Sex Dev. 2010, 4, 176–185. [Google Scholar] [CrossRef] [PubMed]
- Cioffi, M.B.; Martins, C.; Bertollo, L.A.C. Chromosome spreading of associated transposable elements and ribosomal DNA in the fish Erythrinus erythrinus. Implications for genome change and karyoevolution in fish. BMC Evol. Biol. 2010, 10, 271–279. [Google Scholar] [CrossRef] [PubMed]
- Martins, N.F.; Bertollo, L.A.C.; Troy, W.P.; Feldberg, E.; de Souza Valentin, F.C.; Cioffi, M.B. Differentiation and evolutionary relationships in Erythrinus erythrinus (Characiformes, Erythrinidae): Comparative chromosome mapping of repetitive sequences. Rev. Fish Biol. Fish. 2013, 23, 261–269. [Google Scholar] [CrossRef]
- Yano, C.F.; Bertollo, L.A.C.; Molina, W.; Liehr, T.; Cioffi, M. Genomic organization of repetitive DNAs and its implications for male karyotype and the neo-Y chromosome differentiation in Erythrinus erythrinus (Characiformes, Erythrinidae). Comp. Cytogenet. 2014, 8, 139–151. [Google Scholar]
- Parise-Maltempi, P.P.; Martins, C.; Oliveira, C.; Foresti, F. Identification of a new repetitive element in the sex chromosomes of Leporinus elongatus (Teleostei: Characiformes: Anostomidae): New insights into the sex chromosomes of Leporinus. Cytogenet. Genome Res. 2007, 116, 218–223. [Google Scholar] [CrossRef]
- Reed, K.M.; Phillips, R.B. Polymorphism of the nucleolus organizer region (NOR) on the putative sex chromosomes of Arctic char (Salvelinus alpinus) is not sex related. Chromosome Res. 1997, 5, 221–227. [Google Scholar] [CrossRef] [PubMed]
- Palacios-Gimenez, O.M.; Castillo, E.R.; Marti, D.A.; Cabral-de-Mello, D.C. Tracking the evolution of sex chromosome systems in Melanoplinae grasshoppers through chromosomal mapping of repetitive DNA sequences. BMC Evol. Biol. 2013, 13, 167. [Google Scholar] [CrossRef] [PubMed]
- McKee, B.D.; Karpen, G.H. Drosophila ribosomal RNA genes function as an X-Y pairing site during male meiosis. Cell 1990, 61, 61–72. [Google Scholar] [CrossRef]
- Ota, K.; Tateno, Y.; Gojobori, T. Highly differentiated and conserved sex chromosome in fish species (Aulopus japonicus: Teleostei, Aulopidae). Gene 2003, 317, 187–193. [Google Scholar] [CrossRef]
- Guerrero, R.F.; Kirkpatrick, M. Local adaptation and the evolution of chromosome fusions. Evolution 2014, 68, 2747–2756. [Google Scholar] [CrossRef] [PubMed]
- Galián, J.; Proença, S.J.R.; Vogler, A.P. Evolutionary dynamics of autosomal-heterosomal rearrangements in a multiple-X chromosome system of tiger beetles (Cicindelidae). BMC Evol. Biol. 2007, 7, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Ocalewicz, K. Telomeres in Fishes. Cytogenet. Genome Res. 2013, 141, 114–125. [Google Scholar] [CrossRef] [PubMed]
- Meyne, J.; Baker, R.J.; Hobart, H.H.; Hsu, T.C.; Ryder, O.A.; Ward, O.G.; Wiley, J.E.; Wurster-Hill, D.H.; Yates, T.L.; Moyzis, R.K. Distribution of non-telomeric sites of the (TTAGGG)n telomeric sequence in vertebrate chromosomes. Chromosoma 1990, 99, 3–10. [Google Scholar] [CrossRef]
- Rovatsos, M.; Kratochvíl, L.; Altmanová, M.; Pokorná, M.J. Interstitial telomeric motifs in squamate reptiles: When the exceptions outnumber the rule. PLoS ONE 2015, 10, e0134985. [Google Scholar] [CrossRef]
- Matsubara, K.; Uno, Y.; Srikulnath, K.; Matsuda, Y.; Miller, E.; Olsson, M. No interstitial telomeres on autosomes but remarkable amplification of telomeric repeats on the W sex chromosome in the sand lizard (Lacerta agilis). J. Hered. 2015, 106, 753–757. [Google Scholar] [CrossRef]
- Cioffi, M.B.; Bertollo, L.A.C. Initial steps in XY chromosome differentiation in Hoplias malabaricus and the origin of an X1X2Y sex chromosome system in this fish group. Heredity 2010, 105, 554–561. [Google Scholar] [CrossRef] [PubMed]
- Favarato, R.M.; Silva, M.; de Oliveira, R.R.; Matoso, D. Cytogenetic diversity and the evolutionary dynamics of rDNA genes and telomeric sequences in the Ancistrus genus (Loricariidae: Ancistrini). Zebrafish 2016, 13, 103–111. [Google Scholar] [CrossRef] [PubMed]
- Schemberger, M.O.; Bellafronte, E.; Nogaroto, V.; Almeida, M.C.; Schühli, G.S.; Artoni, R.F.; Moreira-Filho, O.; Vicari, M.R. Differentiation of repetitive DNA sites and sex chromosome systems reveal closely related group in Parodontidae (Actinopterygii: Characiformes). Genetica 2011, 139, 1499–1508. [Google Scholar] [CrossRef] [PubMed]
- Slijepcevic, P. Telomeres and mechanisms of Robertsonian fusion. Chromosoma 1998, 107, 136–140. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Russell, D.W. Molecular Cloning, A Laboratory Manual; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2001. [Google Scholar]
- Basset, P.; Yannic, G.; Yang, F.; O’Brien, P.C.M.; Graphodatsky, A.S.; Ferguson-Smith, M.A.; Balmus, G.; Volobouev, V.T.; Hausser, J. Chromosome localization of microsatellite markers in the shrews of the Sorex araneus group. Chromosome Res. 2006, 14, 253–262. [Google Scholar] [CrossRef] [PubMed]
- Martins, C. Chromosomes and repetitive DNAs: A contribution to the knowledge of fish genome. In Fish Cytogenetics; Pisano, E., Ozouf-Costaz, C., Foresti, F., Kapoor, B.G., Eds.; Science Publisher: Enfield, UK, 2007; pp. 421–453. [Google Scholar]
- Kejnovský, E.; Hobza, R.; Čermák, T.; Kubát, Z.; Vyskot, B. The role of repetitive DNA in structure and evolution of sex chromosomes in plants. Heredity 2009, 102, 533–541. [Google Scholar] [CrossRef] [PubMed]
- Pokorná, M.; Kratochvíl, L.; Kejnovský, E. Microsatellite distribution on sex chromosomes at different stages of heteromorphism and heterochromatinization in two lizard species (Squamata: Eublepharidae: Coleonyx elegans and Lacertidae: Eremias velox). BMC Genet. 2011, 12, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Poltronieri, J.; Marquioni, V.; Bertollo, L.A.C.; Kejnovský, E.; Molina, W.F.; Liehr, T.; Cioffi, M.B. Comparative chromosomal mapping of microsatellites in Leporinus species (Characiformes, Anostomidae): Unequal accumulation on the W chromosomes. Cytogenet. Genome Res. 2014, 142, 40–45. [Google Scholar] [CrossRef] [PubMed]
- Gunski, R.; Kretschmer, R.; de Souza, M.; Furo, I.; Barcellos, S.; Costa, A.; Cioffi, M.B.; de Oliveira, E.; Garnero, A. Evolution of bird sex chromosomes narrated by repetitive sequences: Unusual W-chromosome enlargement in Gallinula melanops (Aves: Gruiformes: Rallidae). Cytogenet. Genome Res. 2019, in press. [Google Scholar] [CrossRef]
- Ezaz, T.; Quinn, A.E.; Miura, I.; Sarre, S.D.; Georges, A.; Marshall Graves, J.A. The dragon lizard Pogona vitticeps has ZZ/ZW micro-sex chromosomes. Chromosome Res. 2005, 13, 763–776. [Google Scholar] [CrossRef]
- de Freitas, N.L.; Al-Rikabi, A.B.H.; Bertollo, L.A.C.; Ezaz, T.; Yano, C.F.; de Oliveira, E.A.; Hatanaka, T.; Cioffi, M.B. Early stages of XY sex chromosomes differentiation in the fish Hoplias malabaricus (Characiformes, Erythrinidae) revealed by DNA repeats accumulation. Curr. Genomics 2018, 19, 216–226. [Google Scholar] [CrossRef] [PubMed]
- Keinath, M.C.; Timoshevskaya, N.; Timoshevskiy, V.A.; Voss, R.; Smith, J.J. Miniscule differences between sex chromosomes in the giant genome of a salamander. Sci. Rep. 2018, 8, 17882. [Google Scholar] [CrossRef] [PubMed]
- Marková, M.; Vyskot, B. New horizons of genomic in situ hybridization. Cytogenet. Genome Res. 2010, 126, 368–375. [Google Scholar] [CrossRef] [PubMed]
- Chester, M.; Leitch, A.R.; Soltis, P.S.; Soltis, D.E. Review of the application of modern cytogenetic methods (FISH/GISH) to the study of reticulation (polyploidy/hybridisation). Genes 2010, 1, 166–192. [Google Scholar] [CrossRef] [PubMed]
- Mariotti, B.; Manzano, S.; Kejnovský, E.; Vyskot, B.; Jamilena, M. Accumulation of Y-specific satellite DNAs during the evolution of Rumex acetosa sex chromosomes. Mol. Genet. Genomics 2009, 281, 249–259. [Google Scholar] [CrossRef]
- Palácios-Gimenez, O.M.; Dias, G.B.; De Lima, L.G.; Kuhn, G.C.E.S.; Ramos, É.; Martins, C.; Cabral-De-Mello, D.C. High-throughput analysis of the satellitome revealed enormous diversity of satellite DNAs in the neo-Y chromosome of the cricket Eneoptera surinamensis. Sci. Rep. 2017, 7, 6422. [Google Scholar] [CrossRef] [PubMed]
- Bachtrog, D. Y-chromosome evolution: Emerging insights into processes of Y-chromosome degeneration. Nat. Rev. Genet. 2013, 14, 113124. [Google Scholar] [CrossRef]
- Schoumans, J.; Nordgren, A.; Ruivenkamp, C.; Brondum-Nielsen, K.; The, B.T.; Annéren, G.; Holmberg, E.; Nordenskjold, M.; Anderlid, B.M. Genome-wide screening using array-CGH does not reveal microdeletions/microduplications in children with Kabuki syndrome. Eur. J. Hum. Genet. 2005, 13, 260–263. [Google Scholar] [CrossRef]
- Liu, H.; Pang, M.; Yu, X.; Zhou, Y.; Tong, J.; Fu, B. Sex-specific markers developed by next-generation sequencing confirmed an XX/XY sex determination system in bighead carp (Hypophthalmichehys nobilis) and silver carp (Hypophthalmichthys molitrix). DNA Res. 2018, 25, 257–264. [Google Scholar] [CrossRef]
- Morris, J.; Darolti, I.; Bloch, N.I.; Wright, A.E.; Mank, J.E. Shared and species-specific patterns of nascent Y chromosome evolution in two guppy species. Genes 2018, 9, 238. [Google Scholar] [CrossRef]
- Kirubakaran, T.G.; Andersen, Ø.; De Rosa, M.C.; Andersstuen, T.; Hallan, K.; Kent, M.P.; Lien, S. Characterization of a male specific region containing a candidate sex determining gene in Atlantic cod. Sci. Rep. 2019, 9, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Pan, Q.; Feron, R.; Yano, A.; Guyomard, R.; Jouanno, E.; Vigouroux, E.; Wen, M.; Busnel, J.M.; Bobe, J.; Concordet, J.P.; et al. Identification of the master sex determining gene in Northern pike (Esox lucius) reveals restricted sex chromosome differentiation. bioRxiv 2019. Preprint. [Google Scholar]
- Kamiya, T.; Kai, W.; Tasumi, S.; Oka, A.; Matsunaga, T.; Mizuno, N.; Fujita, M.; Suetake, H.; Suzuki, S.; Hosoya, S.; et al. A trans-species missense SNP in Amhr2 is associated with sex determination in the tiger Pufferfish, Takifugu rubripes (Fugu). PLoS Genet. 2012, 8, e1002798. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, K.; Makino, T.; Yamaguchi, K.; Shigenobu, S.; Hasebe, M.; Kawata, M.; Kume, M.; Mori, S.; Peichel, C.L.; Toyoda, A.; et al. Sex chromosome turnover contributes to genomic divergence between incipient stickleback species. PLoS Genet. 2014, 10, e1004223. [Google Scholar] [CrossRef] [PubMed]
- Natri, H.M.; Shikano, T.; Merilä, J. Progressive recombination suppression and differentiation in recently evolved neo-sex chromosomes. Mol. Biol. Evol. 2013, 30, 1131–1144. [Google Scholar] [CrossRef] [PubMed]
- Moraes, R.L.R.; Sember, A.; Bertollo, L.A.C.; Oliveira, E.A.; Ráb, P.; Hatanaka, T.; Marinho, M.M.F.; Liehr, T.; Al-Rikabi, A.B.H.; Feldberg, E.; et al. Comparative cytogenetics and neo-Y formation in small-sized fish species of the genus Pyrrhulina (Characiformes, Lebiasinidae). Front. Genet 2019, in press. [Google Scholar] [CrossRef]
- Moreira-Filho, O.; Bertollo, L.A.C.; Galetti, P.M., Jr. Distribution of sex chromosome mechanisms in Neotropical fish and description of a ZZ/ZW system in Parodon hilarii (Parodontidae). Caryologia 1993, 46, 115–125. [Google Scholar] [CrossRef][Green Version]
- Charlesworth, B.; Wall, J.D. Inbreeding, heterozygote advantage and the evolution of neo-X and neo-Y sex chromosomes. Proc. Biol. Sci. 1999, 266, 51–56. [Google Scholar] [CrossRef]
- Kitano, J.; Ross, J.A.; Mori, S.; Kume, M.; Jones, F.C.; Chan, Y.F.; Absher, D.M.; Grimwood, J.; Schmutz, J.; Myers, R.M.; et al. A role for a neo-sex chromosome in stickleback speciation. Nature 2009, 461, 1079–1083. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.A.S.; Gordon, I.J.; Traut, W.; Herren, J.; Collins, S.; Martins, D.J.; Saitoti, K.; Ireri, P. A neo-W chromosome in a tropical butterfly links colour pattern, male-killing, and speciation. Proc. R. Soc. B 2016, 283, 20160821. [Google Scholar] [CrossRef]
- Yasukochi, Y.; Miura, N.; Nakano, R.; Sahara, K.; Ishikawa, Y. Sex-linked pheromone receptor genes of the European corn borer, Ostrinia nubilalis, are in tandem arrays. PLoS ONE 2011, 6, e18843. [Google Scholar] [CrossRef] [PubMed]
- Devlin, R.H.; Nagahama, Y. Sex determination and sex differentiation in fish: An overview of genetic, physiological, and environmental influences. Aquaculture 2002, 208, 191–364. [Google Scholar] [CrossRef]
- Mank, J.E.; Avise, J.C. Phylogenetic conservation of chromosome numbers in Actinopterygiian fishes. Genetica 2006, 127, 321–327. [Google Scholar] [CrossRef]
- Cioffi, M.B.; Liehr, T.; Trifonov, V.; Molina, W.F.; Bertollo, L.A.C. Independent sex chromosome evolution in lower vertebrates: A molecular cytogenetic overview in the erythrinidae fish family. Cytogenet. Genome Res. 2013, 141, 186–194. [Google Scholar] [CrossRef]
- Woram, R.A.; Gharbi, K.; Sakamoto, T.; Hoyheim, B.; Holm, L.E.; Naish, K.; McGowan, C.; Ferguson, M.M.; Phillips, R.B.; Stein, J.; et al. Comparative genome analysis of the primary sex-determining locus in salmonid fishes. Genome Res. 2003, 13, 272–280. [Google Scholar] [CrossRef]
- Gammerdinger, W.J.; Kocher, T.D. Unusual diversity of sex chromosomes in African cichlid fishes. Genes 2018, 9, 480. [Google Scholar] [CrossRef] [PubMed]
- Henning, F.; Trifonov, V.; Ferguson-Smith, M.A.; Almeida-Toledo, L.F. Non-homologous sex chromosomes in two species of the genus Eigenmannia (Teleostei: Gymnotiformes). Cytogenet. Genome Res. 2008, 121, 391–400. [Google Scholar] [CrossRef]
- Cioffi, M.B.; Camacho, J.P.M.; Bertollo, L.A.C. Repetitive DNAs and differentiation of sex chromosomes in neotropical fishes. Cytogenet. Genome Res. 2011, 132, 188–194. [Google Scholar] [CrossRef]
- Bertollo, L.A.C.; Cioffi, M.B.; Moreira-Filho, O. Direct chromosome preparation from Freshwater Teleost Fishes. In Fish Cytogenetic Techniques (Chondrichthyans and Teleosts); Ozouf-Costaz, C., Pisano, E., Foresti, F., Almeida Toledo, L.F., Eds.; CRC Press: Enfield, CT, USA, 2015; pp. 21–26. [Google Scholar]
- Sumner, A.T. A simple technique for demonstrating centromeric heterochromatin. Exp. Cell Res. 1972, 75, 304–306. [Google Scholar] [CrossRef]
- Pendás, A.M.; Móran, P.; Freije, J.P.; Garcia-Vásquez, E. Chromosomal location and nucleotide sequence of two tandem repeats of the Atlantic salmon 5S rDNA. Cytogenet. Cell Genet. 1994, 67, 31–36. [Google Scholar] [CrossRef]
- Cioffi, M.B.; Martins, C.; Centofante, L.; Jacobina, U.; Bertollo, L.A.C. Chromosomal variability among allopatric populations of Erythrinidae fish Hoplias malabaricus: Mapping of three classes of repetitive DNAs. Cytogenet. Genome Res. 2009, 125, 132–141. [Google Scholar] [CrossRef] [PubMed]
- Kubát, Z.; Hobza, R.; Vyskot, B.; Kejnovský, E. Microsatellite accumulation in the Y chromosome of Silene latifolia. Genome 2008, 51, 350–356. [Google Scholar] [CrossRef] [PubMed]
- Zwick, M.S.; Hanson, R.E.; Mcknight, T.D.; Islam-Faridi, M.H.; Stelly, D.M.; Wing, R.A.; Price, H.J. A rapid procedure for the isolation of C0t-1 DNA from plants. Genome 1997, 40, 138–142. [Google Scholar] [CrossRef] [PubMed]
- Symonová, R.; Sember, A.; Majtánová, Z.; Ráb, P. Characterization of fish genomes by GISH and CGH. In Fish Cytogenet. Tech. Ray-Fin Fishes Chondrichthyans; CCR Press: Boca Raton, FL, USA, 2015; pp. 118–131. [Google Scholar]
- Yang, F.; Trifonov, V.; Ng, B.L.; Kosyakova, N.; Carter, N.P. Generation of paint probes by flow-sorted and microdissected chromosomes BT. In Fluorescence In Situ Hybridization (FISH)—Application Guide; Liehr, T., Ed.; Springer: Berlin/Heidelberg, Germany, 2009; pp. 35–52. [Google Scholar]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, D.; Sember, A.; Zhu, Q.; Oliveira, E.A.d.; Liehr, T.; Al-Rikabi, A.B.H.; Xiao, Z.; Song, H.; Cioffi, M.d.B. Deciphering the Origin and Evolution of the X1X2Y System in Two Closely-Related Oplegnathus Species (Oplegnathidae and Centrarchiformes). Int. J. Mol. Sci. 2019, 20, 3571. https://doi.org/10.3390/ijms20143571
Xu D, Sember A, Zhu Q, Oliveira EAd, Liehr T, Al-Rikabi ABH, Xiao Z, Song H, Cioffi MdB. Deciphering the Origin and Evolution of the X1X2Y System in Two Closely-Related Oplegnathus Species (Oplegnathidae and Centrarchiformes). International Journal of Molecular Sciences. 2019; 20(14):3571. https://doi.org/10.3390/ijms20143571
Chicago/Turabian StyleXu, Dongdong, Alexandr Sember, Qihui Zhu, Ezequiel Aguiar de Oliveira, Thomas Liehr, Ahmed B. H. Al-Rikabi, Zhizhong Xiao, Hongbin Song, and Marcelo de Bello Cioffi. 2019. "Deciphering the Origin and Evolution of the X1X2Y System in Two Closely-Related Oplegnathus Species (Oplegnathidae and Centrarchiformes)" International Journal of Molecular Sciences 20, no. 14: 3571. https://doi.org/10.3390/ijms20143571
APA StyleXu, D., Sember, A., Zhu, Q., Oliveira, E. A. d., Liehr, T., Al-Rikabi, A. B. H., Xiao, Z., Song, H., & Cioffi, M. d. B. (2019). Deciphering the Origin and Evolution of the X1X2Y System in Two Closely-Related Oplegnathus Species (Oplegnathidae and Centrarchiformes). International Journal of Molecular Sciences, 20(14), 3571. https://doi.org/10.3390/ijms20143571






