psbE-psbL and ndhA Intron, the Promising Plastid DNA Barcode of Fagopyrum
Abstract
1. Introduction
2. Results
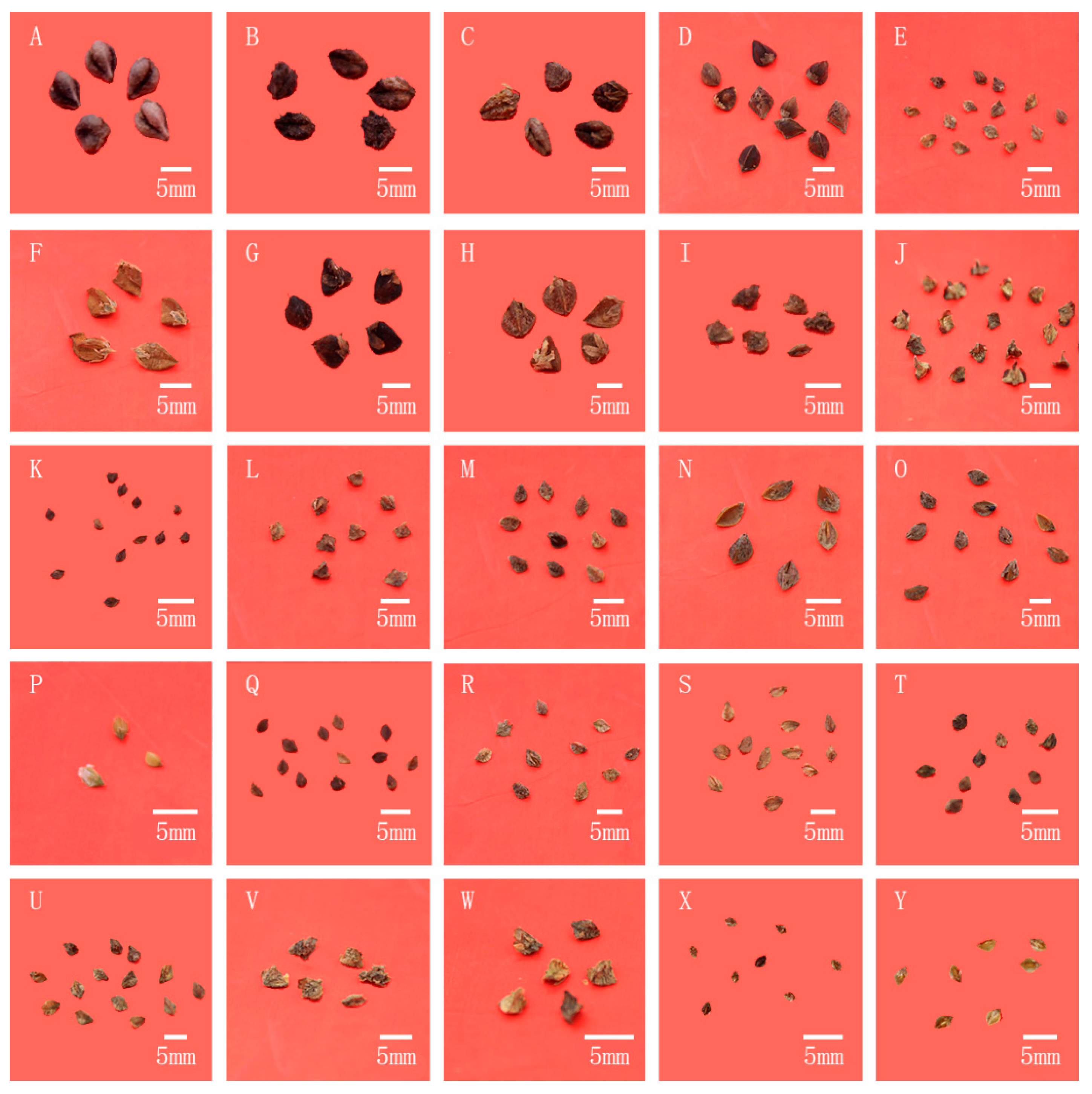
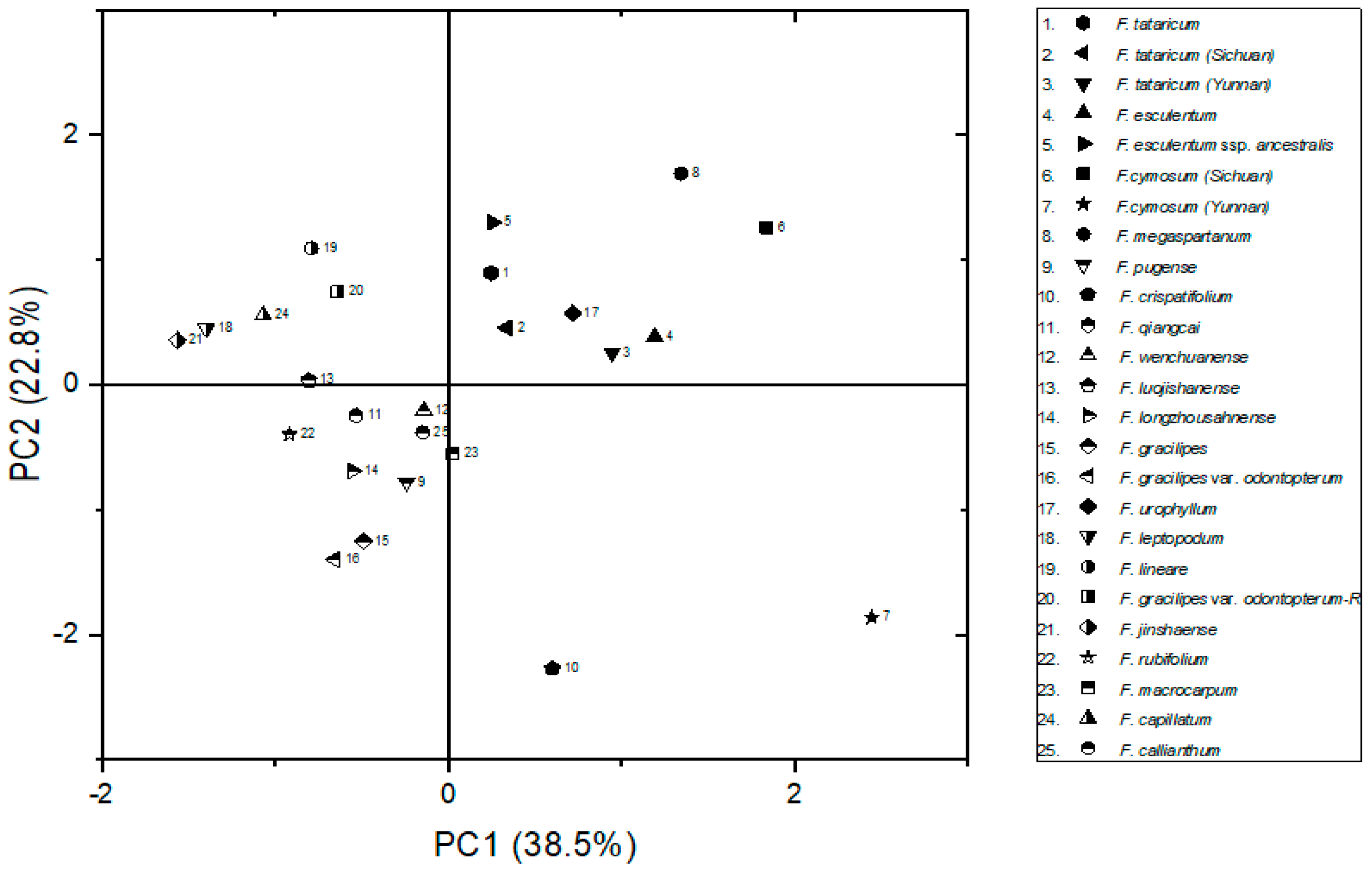
2.1. Analysis Based on Morphological Characteristics of Wild Buckwheat
2.2. psbE-psbL and ndhA Intron Are the Promising Molecular Markers in Fagopyrum
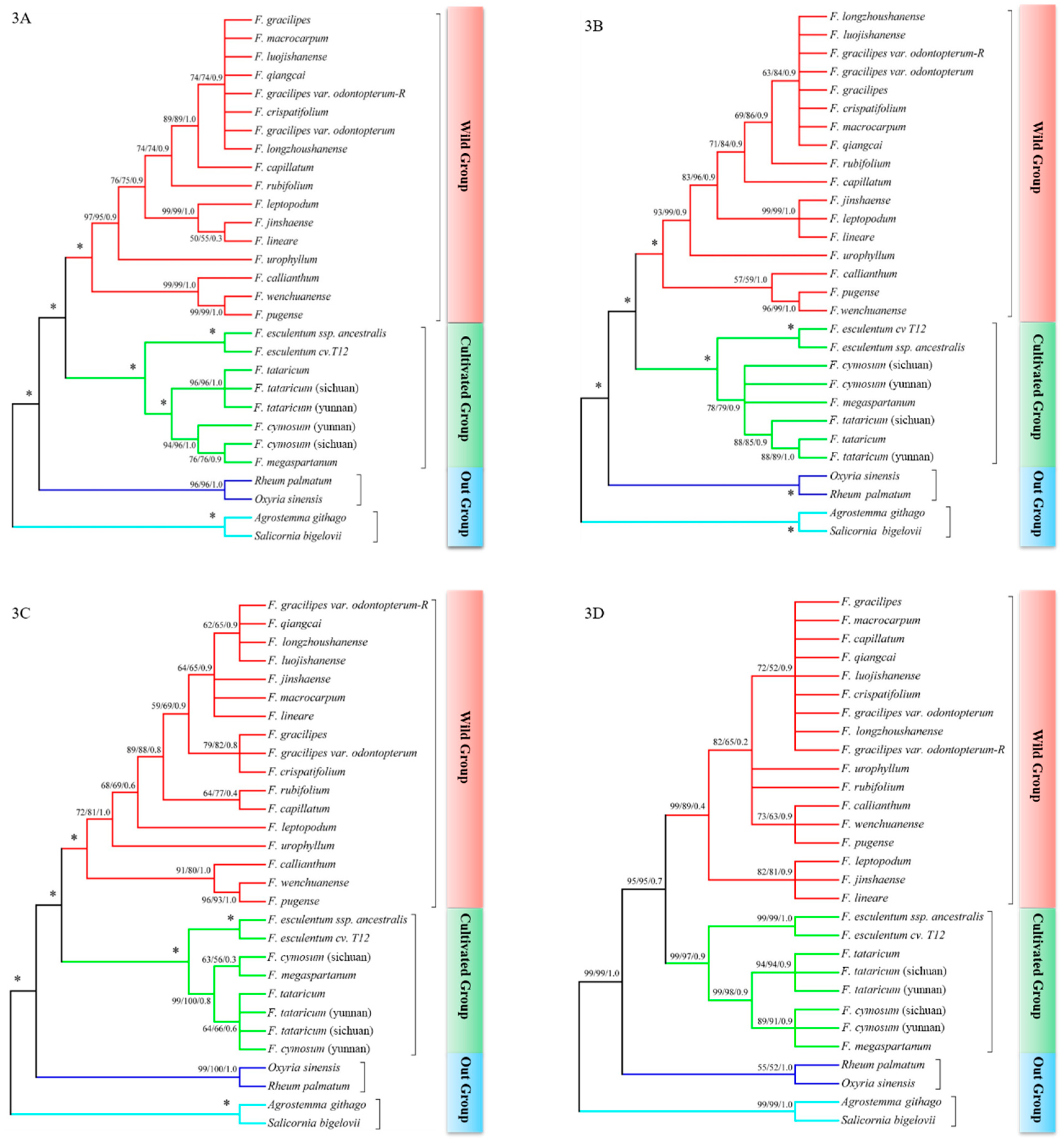
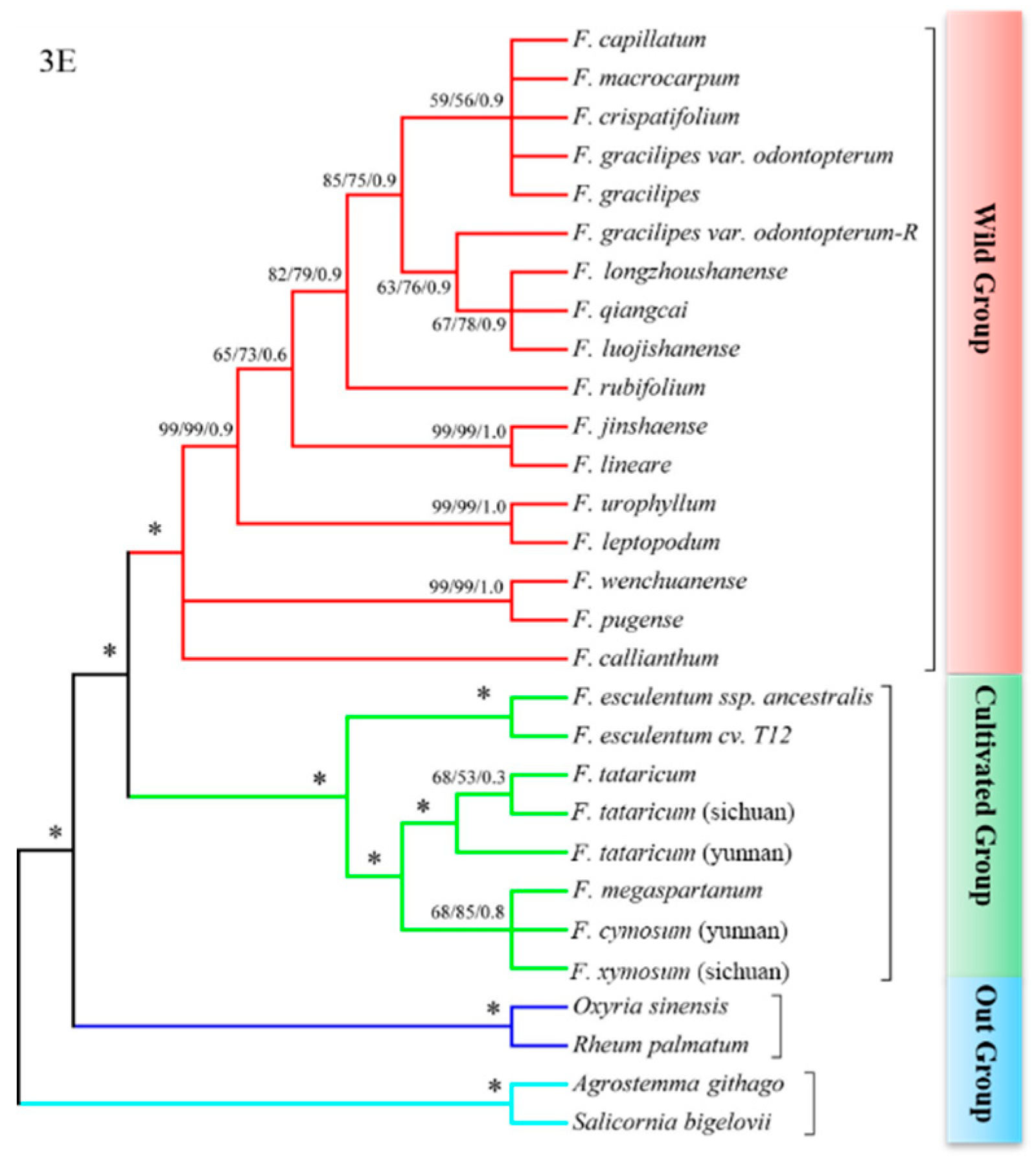
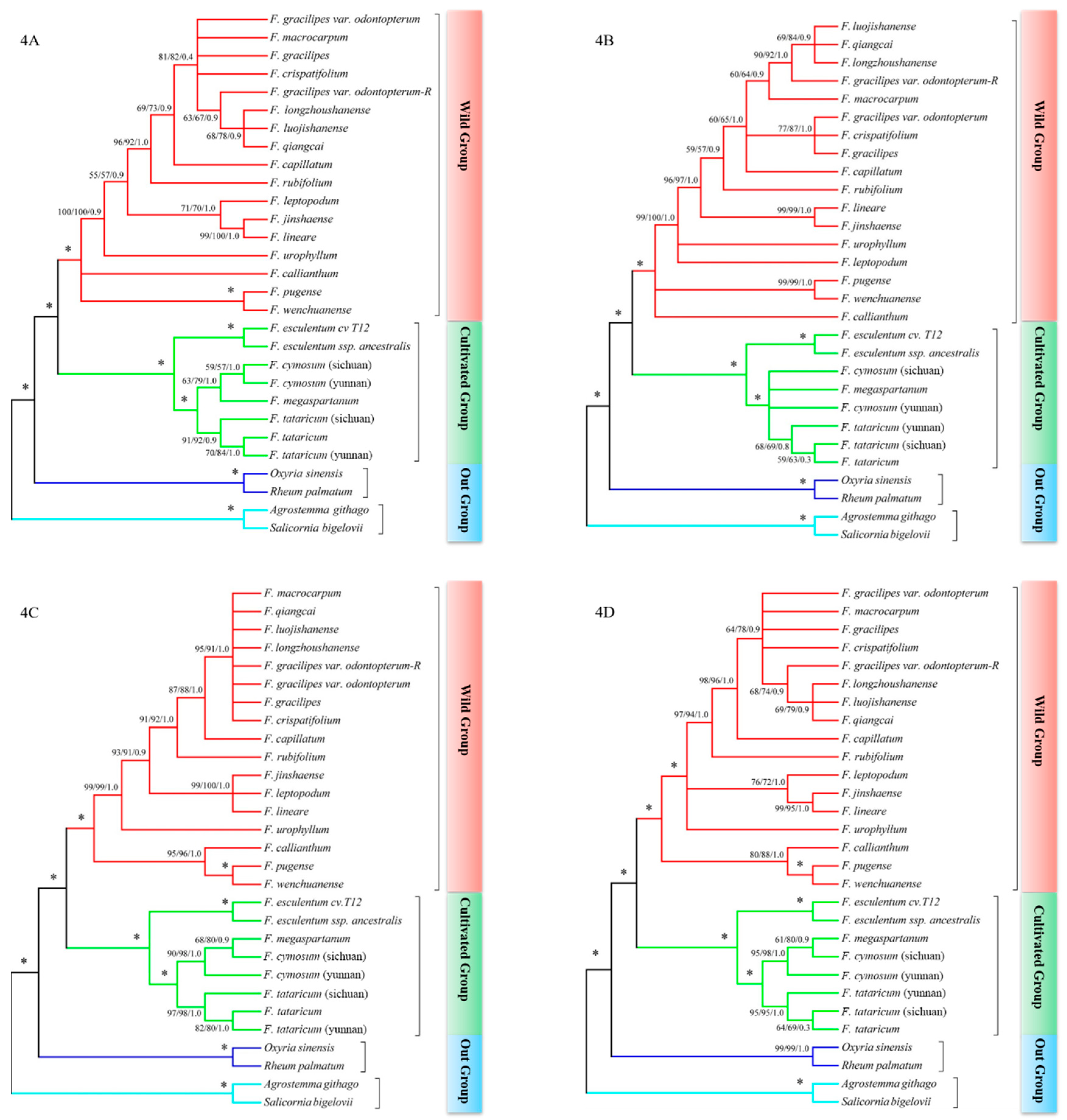
2.3. psbE-psbL + ndhA Intron and psbE-psbL + matK Could Revealed the Relationship between Species Clearly
2.4. The Phylogenetic Relationship between Species in Fagopyrum
3. Discussion
3.1. The Morphological Characteristics Are Not Enough for the Phylogenetic Study in Fagopyrum
3.2. The psbE-psbL and ndhA Intron Could Better Explore the Phylogenetic Relationship in Fagopyrum
3.3. Reconstructing the Phylogenetic Relationship in Fagopyrum Consolidate by Multiple DNA Barcodes
4. Materials and Methods
4.1. Plant Populations
4.2. Observation and Analysis of Morphological Characteristics
4.3. Genome DNA Isolation and Molecular Barcodes Amplification
4.4. Phylogenetic Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Zhu, F. Chemical composition and health effects of Tartary buckwheat. Food Chem. 2016, 203, 231–245. [Google Scholar] [CrossRef] [PubMed]
- Verma, K.C. Biochemical constituents of buckwheat (Fagopyrum esculentum Moench) collected from different geographical regions of Himachal Pradesh. Mol. Biol. Rep. 2018, 45, 2681–2687. [Google Scholar] [CrossRef] [PubMed]
- Sakač, M.; Pestorić, M.; Mišan, A.; Nedeljković, N.; Jambrec, D.; Jovanov, P.; Banjac, V.; Torbica, A.; Hadnađev, M.; Mandić, A. Antioxidant Capacity, Mineral Content and Sensory Properties of Gluten-Free Rice and Buckwheat Cookies. Food Technol. Biotechnol. 2015, 53, 38–47. [Google Scholar] [CrossRef] [PubMed]
- Bae, I.Y.; Hong, I.L.; Ko, A.; Lee, H.G. Substituting whole grain flour for wheat flour: Impact on cake quality and glycemic index. Food Sci. Biotechnol. 2013, 22, 1301–1307. [Google Scholar] [CrossRef]
- Lu, C.L.; Zheng, Q.; Shen, Q.; Song, C.; Zhang, Z.M. Uncovering the relationship and mechanisms of Tartary buckwheat (Fagopyrum tataricum) and Type II diabetes, hypertension, and hyperlipidemia using a network pharmacology approach. Peerj 2017, 5, e4042. [Google Scholar] [CrossRef] [PubMed]
- Hosaka, T.; Sasaga, S.; Yamasaka, Y.; Nii, Y.; Edazawa, K.; Tsutsumi, R.; Shuto, E.; Okahisa, N.; Iwata, S.; Tomotake, H. Treatment with buckwheat bran extract prevents the elevation of serum triglyceride levels and fatty liver in KK-A(y) mice. J. Med. Investig. 2014, 61, 345–352. [Google Scholar] [CrossRef][Green Version]
- Qiu, R.; Wang, J.; Parkin, K.L. Isolation and identification of cytoprotective agents from nonpolar extracts of buckwheat flour. Food Res. Int. 2014, 66, 86–92. [Google Scholar] [CrossRef]
- Tang, Y.; Ding, M.Q.; Tang, Y.X.; Wu, Y.M.; Shao, J.R.; Zhou, M.L. Chapter two—Germplasm Resources of Buckwheat in China. In Molecular Breeding & Nutritional Aspects of Buckwheat; Academic Press: Cambridge, MA, USA, 2016. [Google Scholar]
- Ohnishi, O.; Matsuoka, Y. Search for the wild ancestor of buckwheat II. Taxonomy of Fagopyrum (Polygonaceae) species based on morphology, isozymes and cpDNA variability. Genes Genet. Syst. 1996, 71, 383–390. [Google Scholar] [CrossRef]
- Tsuji, K.; Ohnishi, O. Phylogenetic relationships among wild and cultivated Tartary buckwheat (Fagopyrum tataricum Gaert.) populations revealed by AFLP analyses. Genes Genet. Syst. 2001, 76, 47–52. [Google Scholar] [CrossRef]
- Tsuji, K.; Ohnishi, O. Phylogenetic position of east Tibetan natural populations in Tartary buckwheat (Fagopyrum tataricum Gaert.) revealed by RAPD analyses. Genet. Resour. Crop Evol. 2001, 48, 63–67. [Google Scholar] [CrossRef]
- Suvorova, G.; Zhou, M. Chapter Three—Distribution of Cultivated Buckwheat Resources in the World. In Buckwheat Germplasm in the World; Zhou, M., Kreft, I., Suvorova, G., Tang, Y., Woo, S.H., Eds.; Academic Press: Cambridge, MA, USA, 2018; pp. 21–35. [Google Scholar]
- Zhou, M.; Tang, Y.; Deng, X.; Ruan, C.; Kreft, I.; Tang, Y.; Wu, Y. Chapter One—Overview of Buckwheat Resources in the World. In Buckwheat Germplasm in the World; Zhou, M., Kreft, I., Suvorova, G., Tang, Y., Woo, S.H., Eds.; Academic Press: Cambridge, MA, USA, 2018; pp. 1–7. [Google Scholar]
- Wang, C.L.; Li, Z.Q.; Ding, M.Q.; Tang, Y.; Zhu, X.M.; Liu, J.L.; Shao, J.R.; Zhou, M.L. Fagopyrum longzhoushanense, a new species of Polygonaceae from Sichuan, China. Phytotaxa 2017, 291, 73–80. [Google Scholar] [CrossRef]
- Zhou, M.L.; Wang, C.L.; Wang, D.Z.; Zheng, Y.D.; Li, F.L.; Zhu, X.M.; Tang, Y.X.; Shao, J.R.; Tang, Y.; Wu, Y.M. Phylogenetic relationship of four new species related to southwestern Sichuan Fagopyrum based on morphological and molecular characterization. Biochem. Syst. Ecol. 2014, 57, 403–409. [Google Scholar] [CrossRef]
- Wang, C.L.; Ding, M.Q.; Zou, C.Y.; Zhu, X.M.; Tang, Y.; Zhou, M.L.; Shao, J.R. Comparative Analysis of Four Buckwheat Species Based on Morphology and Complete Chloroplast Genome Sequences. Sci. Rep. 2017, 7, 6514. [Google Scholar] [CrossRef] [PubMed]
- Takanori, O.; Kyoko, Y.; Ohmi, O. Two new Fagopyrum (Polygonaceae) species, F. gracilipedoides and F. jinshaense from Yunnan, China. Jpn. J. Genet. 2002, 77, 399–408. [Google Scholar]
- Yuriko, N.; Ohmi, O.; Masami, H. Topological incongruence between nuclear and chloroplast DNA trees suggesting hybridization in the urophyllum group of the genus Fagopyrum (Polygonaceae). Genes Genet. Syst. 2003, 78, 139–153. [Google Scholar]
- Yasui, Y.; Ohnishi, O. Interspecific relationships in Fagopyrum (Polygonaceae) revealed by the nucleotide sequences of the rbcL and accD genes and their intergenic region. Am. J. Bot. 1998, 85, 1134–1142. [Google Scholar] [CrossRef] [PubMed]
- Ohnishi, O. Search for the Wild Ancestor of Buckwheat III. The Wild Ancestor of Cultivated Common Buckwheat, and of Tatary Buckwheat. Econ. Bot. 1998, 52, 123–133. [Google Scholar] [CrossRef]
- Wu, L.Y.; Wang, B.; Schoen, D.J.; Huang, S.Q. Transitions from distyly to homostyly are associated with floral evolution in the buckwheat genus (Fagopyrum). Am. J. Bot. 2017, 104, 1232–1240. [Google Scholar] [CrossRef]
- Chen, Q.F. Discussion on the Origin of Cultivated Buckwheat in Genus Fagopyrum (Polygonaceae). In Proceedings of the Eighth ISB, Chunchon, Korea, 30 Aug–2 September 2001; pp. 206–213. [Google Scholar]
- The Plant List. Available online: http://www.theplantlist.org/tpl1.1/search?q=fagopyrum (accessed on 14 July 2019).
- Chen, Q.F.; Hsam, S.L.K.; Zeller, F.J. A Study of Cytology, Isozyme, and Interspecific Hybridization on the Big-Achene Group of Buckwheat Species (Polygonaceae). Crop Sci. 2004, 44, 1511–1518. [Google Scholar] [CrossRef]
- Chen, Q.F. Hybridization between Fagopyrum (Polygonaceae) species native to China. Bot. J. Linn. Soc. 2010, 131, 177–185. [Google Scholar] [CrossRef]
- Hu, Y.; Zhang, Z.; Wu, B.; Gao, J.; Li, Y. Genetic relationships of buckwheat species based on the sequence analysis of ITS and ndhF-rpl32. Biodivers. Sci. 2016, 24, 296–303. [Google Scholar] [CrossRef]
- Ohsako, T.; Ohnishi, O. Nucleotide sequence variation of the chloroplast trnK/matK region in two wild Fagopyrum (Polygonaceae) species, F. leptopodum and F. statice. Genes Genet. Syst. 2001, 76, 39–46. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yasui, Y.; Ohnishi, O. Phylogenetic relationships among Fagopyrum species revealed by the nucleotide sequences of the ITS region of the nuclear rRNA gene. Genes Genet. Syst. 1998, 73, 201–210. [Google Scholar] [CrossRef] [PubMed]
- Jansen, R.K.; Ruhlman, T.A. Plastid Genomes of Seed Plants; Springer: Dordrecht, The Netherlands, 2012; pp. 103–126. [Google Scholar]
- Awad, M.; Fahmy, R.M.; Mosa, K.A.; Helmy, M.; El-Feky, F.A.; Awad, M.; Fahmy, R.M.; Mosa, K.A.; Helmy, M.; El-Feky, F.A. Identification of Effective DNA Barcodes for Triticum Plants through Chloroplast Genome-wide Analysis. Comput. Biol. Chem. 2017, 71, 20–31. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, C.; Kano, C.H.; Tate, J.A.; Esteves, G.L. Molecular Phylogenetics and Character Evolution of Gaya and Related Genera (Malvoideae, Malvaceae). Syst. Bot. 2018, 43, 676–688. [Google Scholar] [CrossRef]
- Wenpan, D.; Jing, L.; Jing, Y.; Ling, W.; Shiliang, Z. Highly variable chloroplast markers for evaluating plant phylogeny at low taxonomic levels and for DNA barcoding. PLoS ONE 2012, 7, e35071. [Google Scholar]
- Yun, S.; Yan, C.; Lv, J.; Jin, X.; Zhu, S.; Li, M.F.; Chen, N. Development of Chloroplast Genomic Resources for Oryza Species Discrimination. Front. Plant Sci. 2017, 8, 1854. [Google Scholar]
- Silva, G.A.R.; Jojima, C.L.; Moraes, E.M.; Antonelli, A.; Manfrin, M.H.; Franco, F.F. Intra and interspecific sequence variation in closely related species of Cereus (CACTACEAE). Biochem. Syst. Ecol. 2016, 65, 137–142. [Google Scholar] [CrossRef]
- Steele, P.R.; Pires, J.C. Biodiversity assessment: State-of-the-art techniques in phylogenomics and species identification. Am. J. Bot. 2011, 98, 415–425. [Google Scholar] [CrossRef]
- Yuan, J.; Cheng, F.; Zhou, S. Hybrid origin of Paeonia × yananensis revealed by microsatellite markers, chloroplast gene sequences, and morphological characteristics. Int. J. Plant Sci. 2010, 171, 409–420. [Google Scholar] [CrossRef]
- Hahn, C.; Müller, A.; Kuhnert, N.; Albach, D. The diversity of kale (Brassica oleracea var. sabellica): Glucosinolate content and phylogenetic relationships. J. Agric. Food Chem. 2016, 64, 3215–3325. [Google Scholar] [CrossRef] [PubMed]
- Marc, J.M.; Christophe, L.; Bautista, J.M.; JoL, F.; Monique, E. Molecular phylogeny and species identification of sardines. J. Agric. Food Chem. 2003, 51, 43–50. [Google Scholar]
- Kyoko, Y.; Yasuo, Y.; Ohmi, O. Intraspecific cpDNA variations of diploid and tetraploid perennial buckwheat, Fagopyrum cymosum (Polygonaceae). Am. J. Bot. 2003, 90, 339–346. [Google Scholar]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Cummings, M.P. PAUP* [Phylogenetic Analysis Using Parsimony (and Other Methods)]; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2004. [Google Scholar]
- Fredrik, R.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar]


| Number | Category | Species | Locations |
|---|---|---|---|
| 1 | Cultivated species | F. esculentum cv. T12 | Chengdu, Sichuan |
| 2 | F. tataricum | Chengdu, Sichuan | |
| 3 | Wild species | F. esculentum ssp. ancestralis | Diqing state, Yunnan |
| 4 | F. tataricum (sichuan) | Aba state, Sichuan | |
| 5 | F. tataricum (yunnan) | Yuxi city, Yunnan | |
| 6 | F.cymosum (sichuan) | Liangshan state, Sichuan | |
| 7 | F.cymosum (yunnan) | Dali state, Yunnan | |
| 8 | F. megaspartanum | Diqing state, Yunnan | |
| 9 | F. pugense | Liangshan state, Sichuan | |
| 10 | F. crispatifolium | Liangshan state, Sichuan | |
| 11 | F. qiangcai | Aba state, Sichuan | |
| 12 | F. wenchuanense | Aba state, Sichuan | |
| 13 | F. luojishanense | Liangshan state, Sichuan | |
| 14 | F. longzhousahnense | Liangshan state, Sichuan | |
| 15 | F. gracilipes | Dali state, Yunnan | |
| 16 | F. urophyllum | Dali state, Yunnan | |
| 17 | F. leptopodum | Yaan city, Sichuan | |
| 18 | F. jinshaense | Lijiang city, Yunnan | |
| 19 | F. rubifolium | Aba state, Sichuan | |
| 20 | F. callianthum | Aba state, Sichuan | |
| 21 | F. capillatum | Lijiang city, Yunnan | |
| 22 | F. lineare | Dali state, Yunnan | |
| 23 | F. macrocarpum | Aba state, Sichuan | |
| 24 | Variation | F. gracilipes var. odontopterum | Lijiang city, Yunnan |
| 25 | F. gracilipes var. odontopterum-R | Liangshan state, Sichuan |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, Y.; Li, Z.; Wang, C.; Zou, C.; Wen, W.; Shao, J.; Zhu, X. psbE-psbL and ndhA Intron, the Promising Plastid DNA Barcode of Fagopyrum. Int. J. Mol. Sci. 2019, 20, 3455. https://doi.org/10.3390/ijms20143455
Huang Y, Li Z, Wang C, Zou C, Wen W, Shao J, Zhu X. psbE-psbL and ndhA Intron, the Promising Plastid DNA Barcode of Fagopyrum. International Journal of Molecular Sciences. 2019; 20(14):3455. https://doi.org/10.3390/ijms20143455
Chicago/Turabian StyleHuang, Yue, Zhiqiang Li, Chenglong Wang, Chenyan Zou, Wen Wen, Jirong Shao, and Xuemei Zhu. 2019. "psbE-psbL and ndhA Intron, the Promising Plastid DNA Barcode of Fagopyrum" International Journal of Molecular Sciences 20, no. 14: 3455. https://doi.org/10.3390/ijms20143455
APA StyleHuang, Y., Li, Z., Wang, C., Zou, C., Wen, W., Shao, J., & Zhu, X. (2019). psbE-psbL and ndhA Intron, the Promising Plastid DNA Barcode of Fagopyrum. International Journal of Molecular Sciences, 20(14), 3455. https://doi.org/10.3390/ijms20143455





