Telomeres and Longevity: A Cause or an Effect?
Abstract
1. Introduction
2. Results
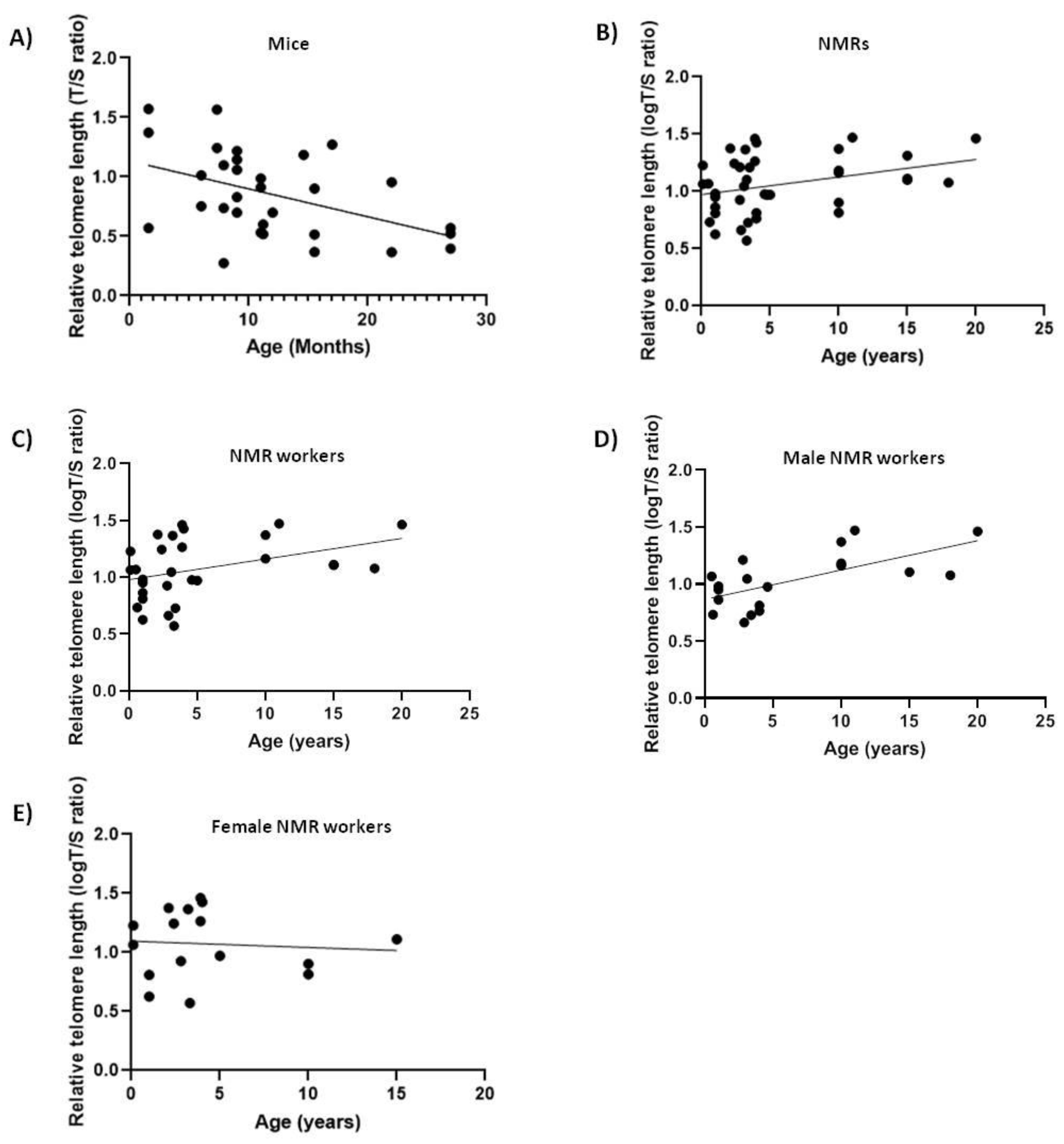
2.1. NMRs and Mice
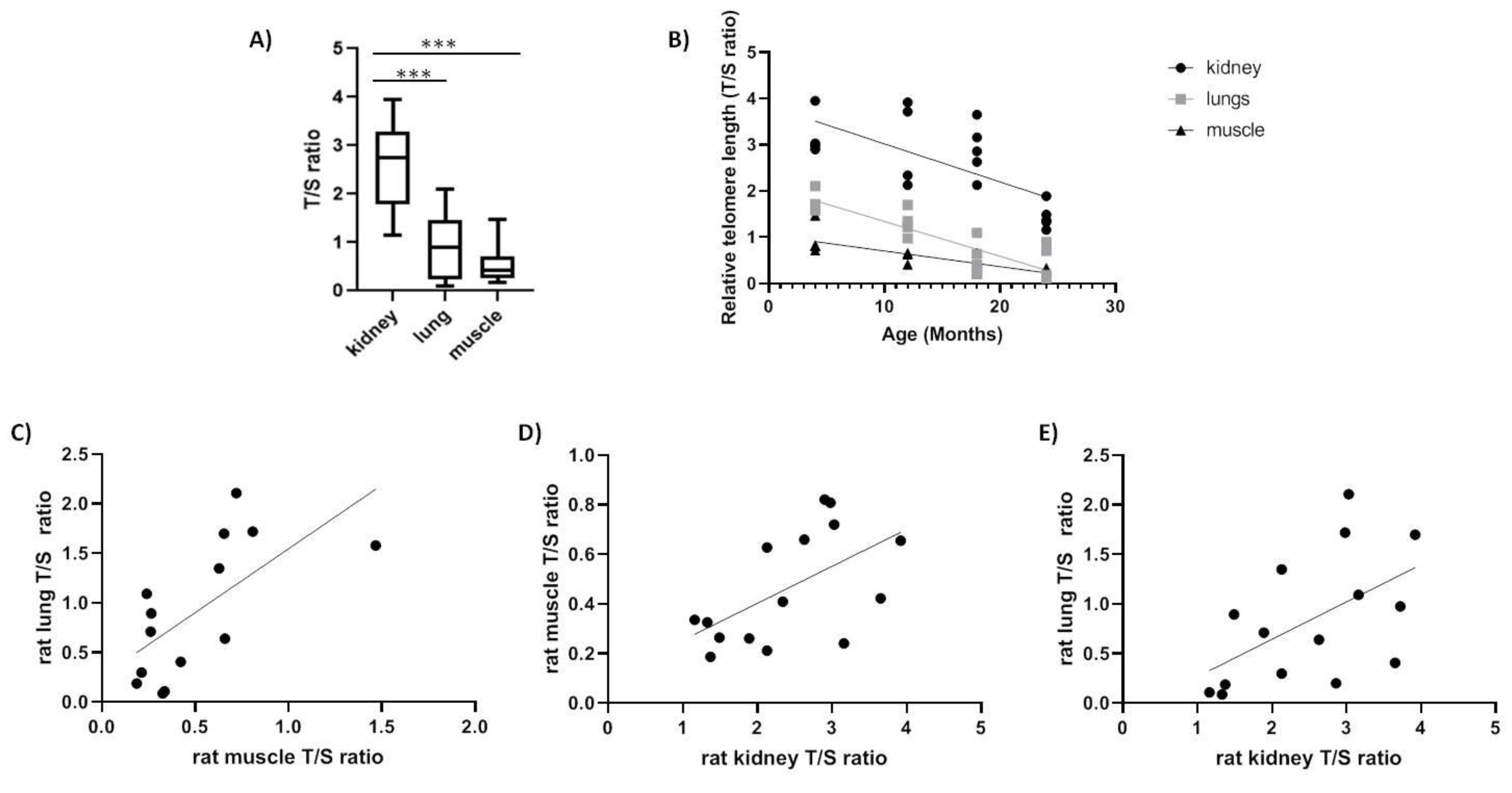
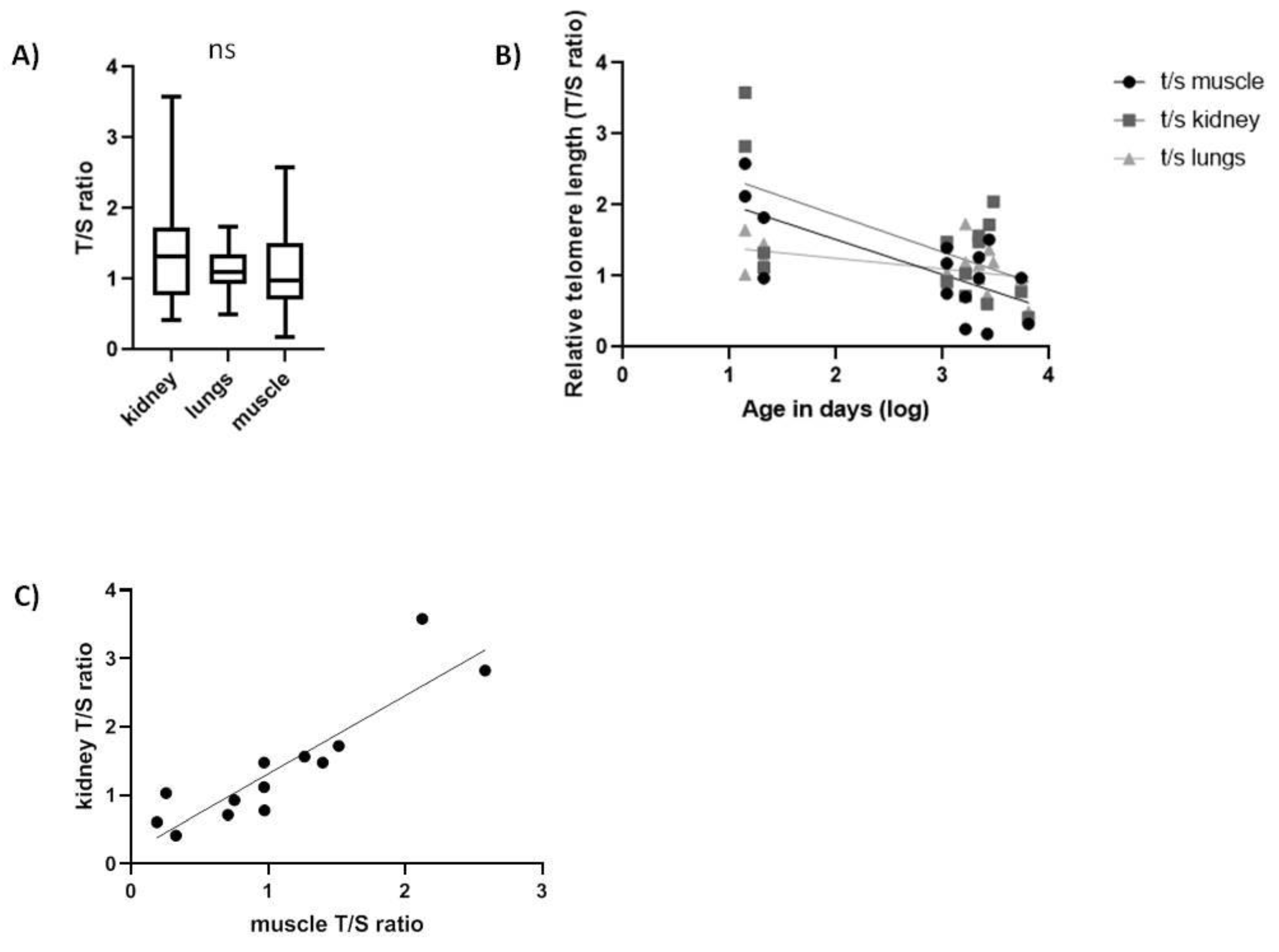
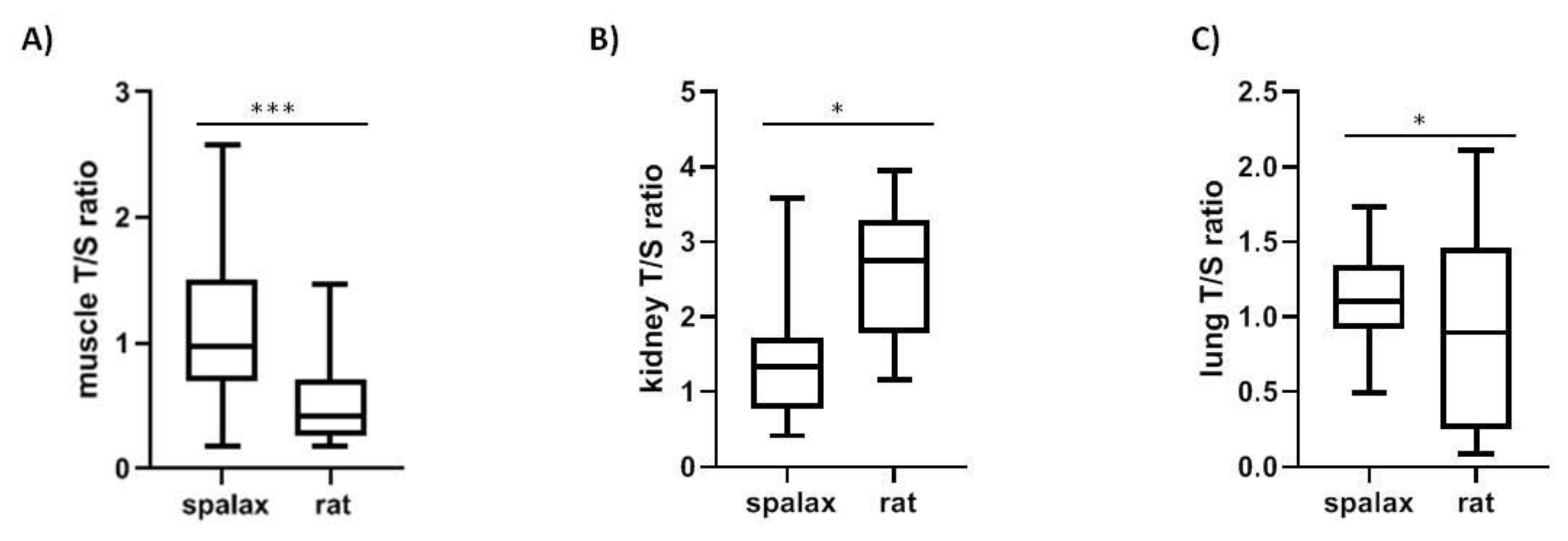
2.2. Spalax and Rats
3. Discussion
4. Materials and Methods
4.1. Animals
4.2. DNA Extraction
4.3. Telomere Measurement
4.4. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| NMR | Naked mole-rat |
| rTL | Relative telomere length |
| ns | Not significant |
| MLSP | Maximum lifespan |
References
- Bize, P.; Criscuolo, F.; Metcalfe, N.B.; Nasir, L.; Monaghan, P. Telomere dynamics rather than age predict life expectancy in the wild. Proc. Biol. Sci. 2009, 276, 1679–1683. [Google Scholar] [CrossRef] [PubMed]
- Steenstrup, T.; Kark, J.D.; Verhulst, S.; Thinggaard, M.; Hjelmborg, J.V.B.; Dalgard, C.; Kyvik, K.O.; Christiansen, L.; Mangino, M.; Spector, T.D.; et al. Telomeres and the natural lifespan limit in humans. Aging (Albany NY) 2017, 9, 1130–1142. [Google Scholar] [CrossRef] [PubMed]
- Armanios, M.; Alder, J.K.; Parry, E.M.; Karim, B.; Strong, M.A.; Greider, C.W. Short telomeres are sufficient to cause the degenerative defects associated with aging. Am. J. Hum. Genet. 2009, 85, 823–832. [Google Scholar] [CrossRef]
- Gomes, N.M.; Ryder, O.A.; Houck, M.L.; Charter, S.J.; Walker, W.; Forsyth, N.R.; Austad, S.N.; Venditti, C.; Pagel, M.; Shay, J.W.; et al. Comparative biology of mammalian telomeres: Hypotheses on ancestral states and the roles of telomeres in longevity determination. Aging Cell 2011, 10, 761–768. [Google Scholar] [CrossRef] [PubMed]
- O’Sullivan, R.J.; Karlseder, J. Telomeres: Protecting chromosomes against genome instability. Nat. Rev. Mol. Cell Biol. 2010, 11, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Meyne, J.; Ratliff, R.L.; Moyzis, R.K. Conservation of the human telomere sequence (ttaggg)n among vertebrates. Proc. Natl. Acad. Sci. USA 1989, 86, 7049–7053. [Google Scholar] [CrossRef] [PubMed]
- Lansdorp, P.M.; Verwoerd, N.P.; van de Rijke, F.M.; Dragowska, V.; Little, M.T.; Dirks, R.W.; Raap, A.K.; Tanke, H.J. Heterogeneity in telomere length of human chromosomes. Hum. Mol. Genet. 1996, 5, 685–691. [Google Scholar] [CrossRef] [PubMed]
- Oeseburg, H.; de Boer, R.A.; van Gilst, W.H.; van der Harst, P. Telomere biology in healthy aging and disease. Pflug. Arch. 2010, 459, 259–268. [Google Scholar] [CrossRef] [PubMed]
- Lingner, J.; Cooper, J.P.; Cech, T.R. Telomerase and DNA end replication: No longer a lagging strand problem? Science 1995, 269, 1533–1534. [Google Scholar] [CrossRef]
- Greider, C.W. Telomerase is processive. Mol. Cell. Biol. 1991, 11, 4572–4580. [Google Scholar] [CrossRef]
- Cesare, A.J.; Reddel, R.R. Alternative lengthening of telomeres: Models, mechanisms and implications. Nat. Rev. Genet. 2010, 11, 319–330. [Google Scholar] [CrossRef] [PubMed]
- Victorelli, S.; Passos, J.F. Telomeres and cell senescence - size matters not. EBioMedicine 2017, 21, 14–20. [Google Scholar] [CrossRef] [PubMed]
- Cherif, H.; Tarry, J.L.; Ozanne, S.E.; Hales, C.N. Ageing and telomeres: A study into organ- and gender-specific telomere shortening. Nucleic Acids Res. 2003, 31, 1576–1583. [Google Scholar] [CrossRef] [PubMed]
- Haussmann, M.F.; Winkler, D.W.; O’Reilly, K.M.; Huntington, C.E.; Nisbet, I.C.; Vleck, C.M. Telomeres shorten more slowly in long-lived birds and mammals than in short-lived ones. Proc. Biol. Sci. 2003, 270, 1387–1392. [Google Scholar] [CrossRef] [PubMed]
- Monaghan, P. Telomeres and life histories: The long and the short of it. Ann. N. Y. Acad. Sci. 2010, 1206, 130–142. [Google Scholar] [CrossRef] [PubMed]
- Njajou, O.T.; Cawthon, R.M.; Damcott, C.M.; Wu, S.H.; Ott, S.; Garant, M.J.; Blackburn, E.H.; Mitchell, B.D.; Shuldiner, A.R.; Hsueh, W.C. Telomere length is paternally inherited and is associated with parental lifespan. Proc. Natl. Acad. Sci. USA 2007, 104, 12135–12139. [Google Scholar] [CrossRef] [PubMed]
- Cassidy, A.; De Vivo, I.; Liu, Y.; Han, J.; Prescott, J.; Hunter, D.J.; Rimm, E.B. Associations between diet, lifestyle factors, and telomere length in women. Am. J. Clin. Nutr. 2010, 91, 1273–1280. [Google Scholar] [CrossRef] [PubMed]
- Prescott, J.; Kraft, P.; Chasman, D.I.; Savage, S.A.; Mirabello, L.; Berndt, S.I.; Weissfeld, J.L.; Han, J.; Hayes, R.B.; Chanock, S.J.; et al. Genome-wide association study of relative telomere length. PLoS ONE 2011, 6, e19635. [Google Scholar] [CrossRef]
- Song, S.; Johnson, F.B. Epigenetic mechanisms impacting aging: A focus on histone levels and telomeres. Genes (Basel) 2018, 9, 201. [Google Scholar] [CrossRef]
- Koks, S.; Dogan, S.; Tuna, B.G.; Gonzalez-Navarro, H.; Potter, P.; Vandenbroucke, R.E. Mouse models of ageing and their relevance to disease. Mech. Ageing Dev. 2016, 160, 41–53. [Google Scholar] [CrossRef]
- Chen, W.; Kimura, M.; Kim, S.; Cao, X.; Srinivasan, S.R.; Berenson, G.S.; Kark, J.D.; Aviv, A. Longitudinal versus cross-sectional evaluations of leukocyte telomere length dynamics: Age-dependent telomere shortening is the rule. J. Gerontol. A. Biol. Sci. Med. Sci. 2011, 66, 312–319. [Google Scholar] [CrossRef] [PubMed]
- Flanary, B.E.; Kletetschka, G. Analysis of telomere length and telomerase activity in tree species of various life-spans, and with age in the bristlecone pine pinus longaeva. Biogerontology 2005, 6, 101–111. [Google Scholar] [CrossRef] [PubMed]
- Olsson, M.; Pauliny, A.; Wapstra, E.; Blomqvist, D. Proximate determinants of telomere length in sand lizards (lacerta agilis). Biol. Lett. 2010, 6, 651–653. [Google Scholar] [CrossRef] [PubMed]
- Atzmon, G.; Cho, M.; Cawthon, R.M.; Budagov, T.; Katz, M.; Yang, X.; Siegel, G.; Bergman, A.; Huffman, D.M.; Schechter, C.B.; et al. Evolution in health and medicine sackler colloquium: Genetic variation in human telomerase is associated with telomere length in ashkenazi centenarians. Proc. Natl. Acad. Sci. USA 2010, 107, 1710–1717. [Google Scholar] [CrossRef] [PubMed]
- Dammann, P. Slow aging in mammals-lessons from african mole-rats and bats. Semin. Cell Dev. Biol. 2017, 70, 154–163. [Google Scholar] [CrossRef]
- Valenzano, D.R.; Aboobaker, A.; Seluanov, A.; Gorbunova, V. Non-canonical aging model systems and why we need them. EMBO J. 2017, 36, 959–963. [Google Scholar] [CrossRef] [PubMed]
- Edrey, Y.H.; Hanes, M.; Pinto, M.; Mele, J.; Buffenstein, R. Successful aging and sustained good health in the naked mole rat: A long-lived mammalian model for biogerontology and biomedical research. Ilar J. 2011, 52, 41–53. [Google Scholar] [CrossRef]
- Lagunas-Rangel, F.A. Cancer-free aging: Insights from spalax ehrenbergi superspecies. Ageing Res. Rev. 2018, 47, 18–23. [Google Scholar] [CrossRef]
- Grimes, K.M.; Reddy, A.K.; Lindsey, M.L.; Buffenstein, R. And the beat goes on: Maintained cardiovascular function during aging in the longest-lived rodent, the naked mole-rat. Am. J. Physiol. Heart Circ. Physiol. 2014, 307, H284–H291. [Google Scholar] [CrossRef]
- Skulachev, V.P.; Holtze, S.; Vyssokikh, M.Y.; Bakeeva, L.E.; Skulachev, M.V.; Markov, A.V.; Hildebrandt, T.B.; Sadovnichii, V.A. Neoteny, prolongation of youth: From naked mole rats to “naked apes” (humans). Physiol. Rev. 2017, 97, 699–720. [Google Scholar] [CrossRef]
- Buffenstein, R. Negligible senescence in the longest living rodent, the naked mole-rat: Insights from a successfully aging species. J. Comp. Physiol. B 2008, 178, 439–445. [Google Scholar] [CrossRef] [PubMed]
- Chiu, C.L.; Hearn, N.L.; Paine, D.; Steiner, N.; Lind, J.M. Does telomere shortening precede the onset of hypertension in spontaneously hypertensive mice? Twin Res. Hum. Genet. 2016, 19, 422–429. [Google Scholar] [CrossRef] [PubMed]
- Heidinger, B.J.; Blount, J.D.; Boner, W.; Griffiths, K.; Metcalfe, N.B.; Monaghan, P. Telomere length in early life predicts lifespan. Proc. Natl. Acad. Sci. USA 2012, 109, 1743–1748. [Google Scholar] [CrossRef] [PubMed]
- Eastwood, J.R.; Hall, M.L.; Teunissen, N.; Kingma, S.A.; Hidalgo Aranzamendi, N.; Fan, M.; Roast, M.; Verhulst, S.; Peters, A. Early-life telomere length predicts lifespan and lifetime reproductive success in a wild bird. Mol. Ecol. 2019, 28, 1127–1137. [Google Scholar] [CrossRef] [PubMed]
- Haussmann, M.F.; Winkler, D.W.; Huntington, C.E.; Nisbet, I.C.; Vleck, C.M. Telomerase activity is maintained throughout the lifespan of long-lived birds. Exp. Gerontol. 2007, 42, 610–618. [Google Scholar] [CrossRef]
- Vera, E.; Bernardes de Jesus, B.; Foronda, M.; Flores, J.M.; Blasco, M.A. The rate of increase of short telomeres predicts longevity in mammals. Cell Rep. 2012, 2, 732–737. [Google Scholar] [CrossRef]
- Gorbunova, V.; Hine, C.; Tian, X.; Ablaeva, J.; Gudkov, A.V.; Nevo, E.; Seluanov, A. Cancer resistance in the blind mole rat is mediated by concerted necrotic cell death mechanism. Proc. Natl. Acad. Sci. USA 2012, 109, 19392–19396. [Google Scholar] [CrossRef]
- Seluanov, A.; Chen, Z.; Hine, C.; Sasahara, T.H.; Ribeiro, A.A.; Catania, K.C.; Presgraves, D.C.; Gorbunova, V. Telomerase activity coevolves with body mass not lifespan. Aging Cell 2007, 6, 45–52. [Google Scholar] [CrossRef]
- Vaiserman, A.M.A.M. Epigenetics of aging and longevity. In Translational Epigenetics, 1st ed.; Alexey Moskalev, A.M.V., Ed.; Academic Press: Cambridge, MA, USA, 2017; Volume 4, p. 544. [Google Scholar]
- Lowe, D.; Horvath, S.; Raj, K. Epigenetic clock analyses of cellular senescence and ageing. Oncotarget 2016, 7, 8524–8531. [Google Scholar] [CrossRef]
- Kabacik, S.; Horvath, S.; Cohen, H.; Raj, K. Epigenetic ageing is distinct from senescence-mediated ageing and is not prevented by telomerase expression. Aging (Albany NY) 2018, 10, 2800–2815. [Google Scholar] [CrossRef]
- Reichert, S.; Criscuolo, F.; Verinaud, E.; Zahn, S.; Massemin, S. Telomere length correlations among somatic tissues in adult zebra finches. PLoS ONE 2013, 8, e81496. [Google Scholar] [CrossRef] [PubMed]
- Gramatges, M.M.; Bertuch, A.A. Measuring relative telomere length: Is tissue an issue? Aging 2010, 2, 756–757. [Google Scholar] [CrossRef] [PubMed]
- Dlouha, D.; Maluskova, J.; Kralova Lesna, I.; Lanska, V.; Hubacek, J.A. Comparison of the relative telomere length measured in leukocytes and eleven different human tissues. Physiol. Res. 2014, 63, S343–S350. [Google Scholar] [PubMed]
- Yu, C.F.; Li, Y.; Holmes, A.; Szafranski, K.; Faulkes, C.G.; Coen, C.W.; Buffenstein, R.; Platzer, M.; de Magalhaes, J.P.; Church, G.M. Rna sequencing reveals differential expression of mitochondrial and oxidation reduction genes in the long-lived naked mole-rat when compared to mice. PLoS ONE 2011, 6, e26729. [Google Scholar] [CrossRef] [PubMed]
- von Zglinicki, T. Oxidative stress shortens telomeres. Trends Biochem. Sci. 2002, 27, 339–344. [Google Scholar] [CrossRef]
- Gardner, M.; Bann, D.; Wiley, L.; Cooper, R.; Hardy, R.; Nitsch, D.; Martin-Ruiz, C.; Shiels, P.; Sayer, A.A.; Barbieri, M.; et al. Gender and telomere length: Systematic review and meta-analysis. Exp. Gerontol. 2014, 51, 15–27. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Xie, Y.; Zhang, Z.; Mao, P.; Liu, J.; Ma, W.; Zhao, Y. Telomeric recombination induced by DNA damage results in telomere extension and length heterogeneity. Neoplasia 2018, 20, 905–916. [Google Scholar] [CrossRef] [PubMed]
- Bernadotte, A.; Mikhelson, V.M.; Spivak, I.M. Markers of cellular senescence. Telomere shortening as a marker of cellular senescence. Aging (Albany NY) 2016, 8, 3–11. [Google Scholar] [CrossRef]
- Cawthon, R.M. Telomere length measurement by a novel monochrome multiplex quantitative pcr method. Nucleic Acids Res. 2009, 37, e21. [Google Scholar] [CrossRef]
- Axelrad, M.D.; Budagov, T.; Atzmon, G. Telomere length and telomerase activity; a yin and yang of cell senescence. J. Vis. Exp. 2013, 75, e50246. [Google Scholar] [CrossRef]
| Species | MLSP * (Years) | Body Size (g) | DNA Source | Average Telomere Length (kb) | Age Range | Number of Samples |
|---|---|---|---|---|---|---|
| Naked Mole-Rat (NMR) | 31 | 40 | EDTA whole blood | 34 | 0–20 years | 40 |
| Mice | 4 | 30 | EDTA whole blood | 72 | 0–27 months | 31 |
| Blind Mole-Rat (Spalax) | 20.2 | 120 | Kidney, lung, muscle | 50 | 0–17.5 years | 18 |
| Rats | 5 | 400 | Kidney, lung, muscle | 60 | 4–24 months | 20 |
| Animal | DNA Source | Correlation Coefficient |
|---|---|---|
| NMRs | Blood | 0.334 * |
| Workers blood | 0.384 * | |
| Female blood | 0.1096 ns | |
| Male blood | 0.6843 ** | |
| Mice | Blood | −0.459 ** |
| Spalax | Kidney | −0.5168 * |
| Muscle | −0.538 * | |
| Lung | −0.5398 * | |
| Rats | Kidney | −0.68 ** |
| Muscle | −0.848 *** | |
| Lung | −0.808 *** |
| Kidney | Muscle | ||
|---|---|---|---|
| Spalax | Kidney | 0.894 *** | |
| Lung | 0.1739 ns | 0.306 ns | |
| Rat | Kidney | 0.566 * | |
| Lung | 0.535 * | 0.659 * |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Adwan Shekhidem, H.; Sharvit, L.; Leman, E.; Manov, I.; Roichman, A.; Holtze, S.; M. Huffman, D.; Y. Cohen, H.; Bernd Hildebrandt, T.; Shams, I.; et al. Telomeres and Longevity: A Cause or an Effect? Int. J. Mol. Sci. 2019, 20, 3233. https://doi.org/10.3390/ijms20133233
Adwan Shekhidem H, Sharvit L, Leman E, Manov I, Roichman A, Holtze S, M. Huffman D, Y. Cohen H, Bernd Hildebrandt T, Shams I, et al. Telomeres and Longevity: A Cause or an Effect? International Journal of Molecular Sciences. 2019; 20(13):3233. https://doi.org/10.3390/ijms20133233
Chicago/Turabian StyleAdwan Shekhidem, Huda, Lital Sharvit, Eva Leman, Irena Manov, Asael Roichman, Susanne Holtze, Derek M. Huffman, Haim Y. Cohen, Thomas Bernd Hildebrandt, Imad Shams, and et al. 2019. "Telomeres and Longevity: A Cause or an Effect?" International Journal of Molecular Sciences 20, no. 13: 3233. https://doi.org/10.3390/ijms20133233
APA StyleAdwan Shekhidem, H., Sharvit, L., Leman, E., Manov, I., Roichman, A., Holtze, S., M. Huffman, D., Y. Cohen, H., Bernd Hildebrandt, T., Shams, I., & Atzmon, G. (2019). Telomeres and Longevity: A Cause or an Effect? International Journal of Molecular Sciences, 20(13), 3233. https://doi.org/10.3390/ijms20133233








