Genome-Wide Analysis of the MADS-Box Transcription Factor Family in Solanum lycopersicum
Abstract
1. Introduction
2. Results
2.1. Identification of MADS-Box Genes in Tomato
2.2. Classification and Phylogenetic Analysis of Tomato MADS-Box Genes
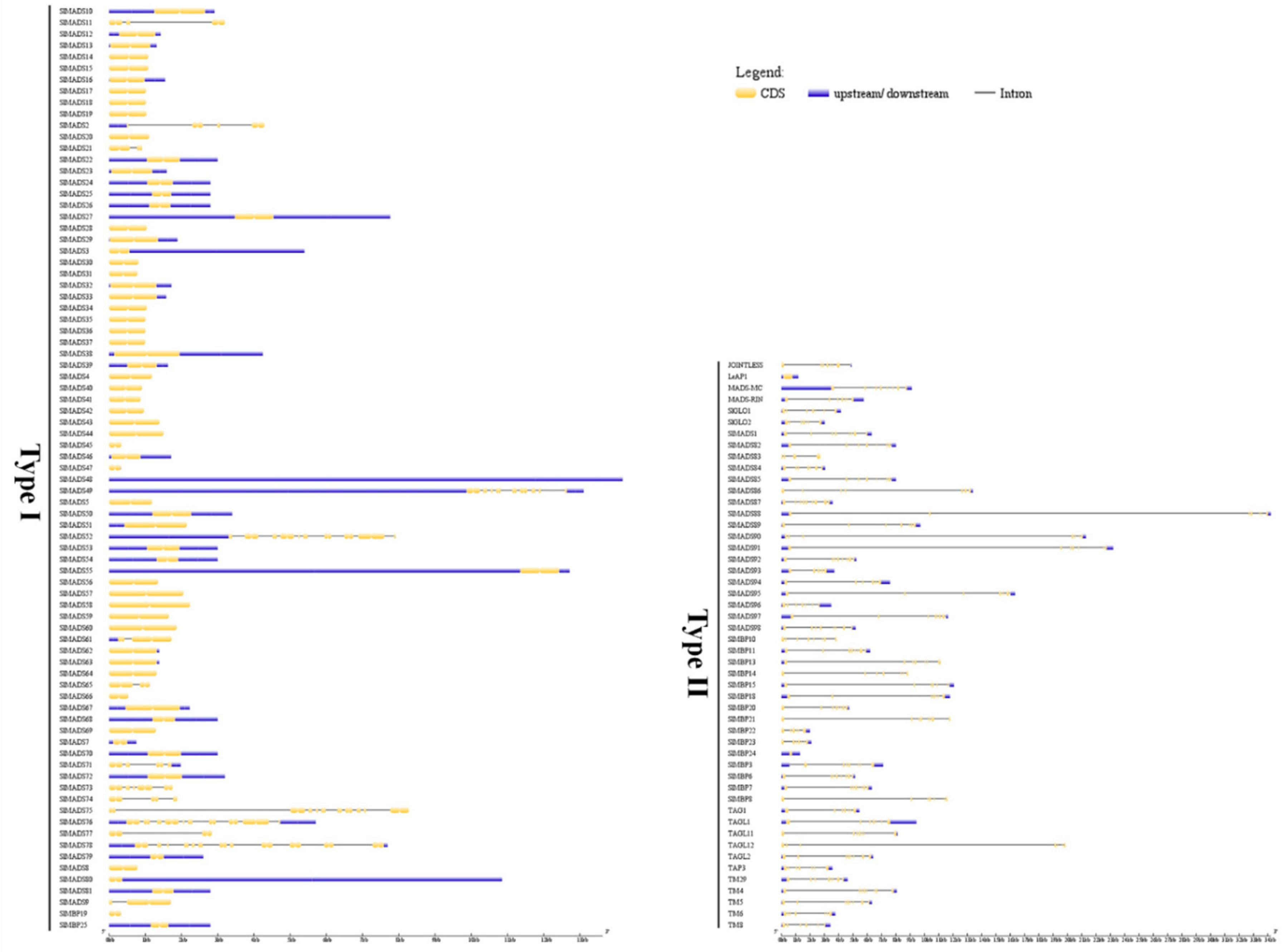
2.3. Conserved Motif and Gene Structure Analysis of Tomato MADS-Box Genes
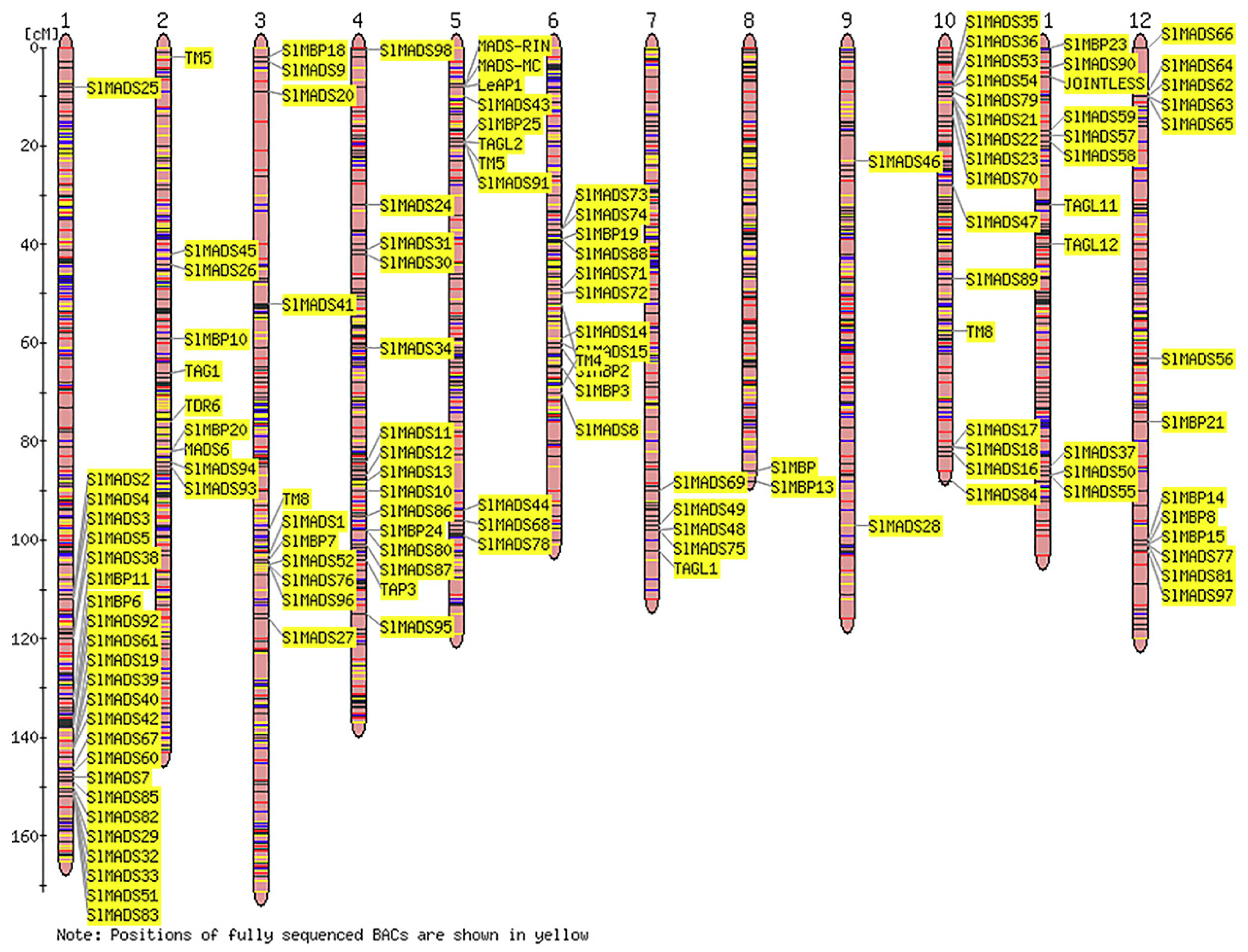
2.4. Chromosomal Locations of Tomato MADS-Box Genes
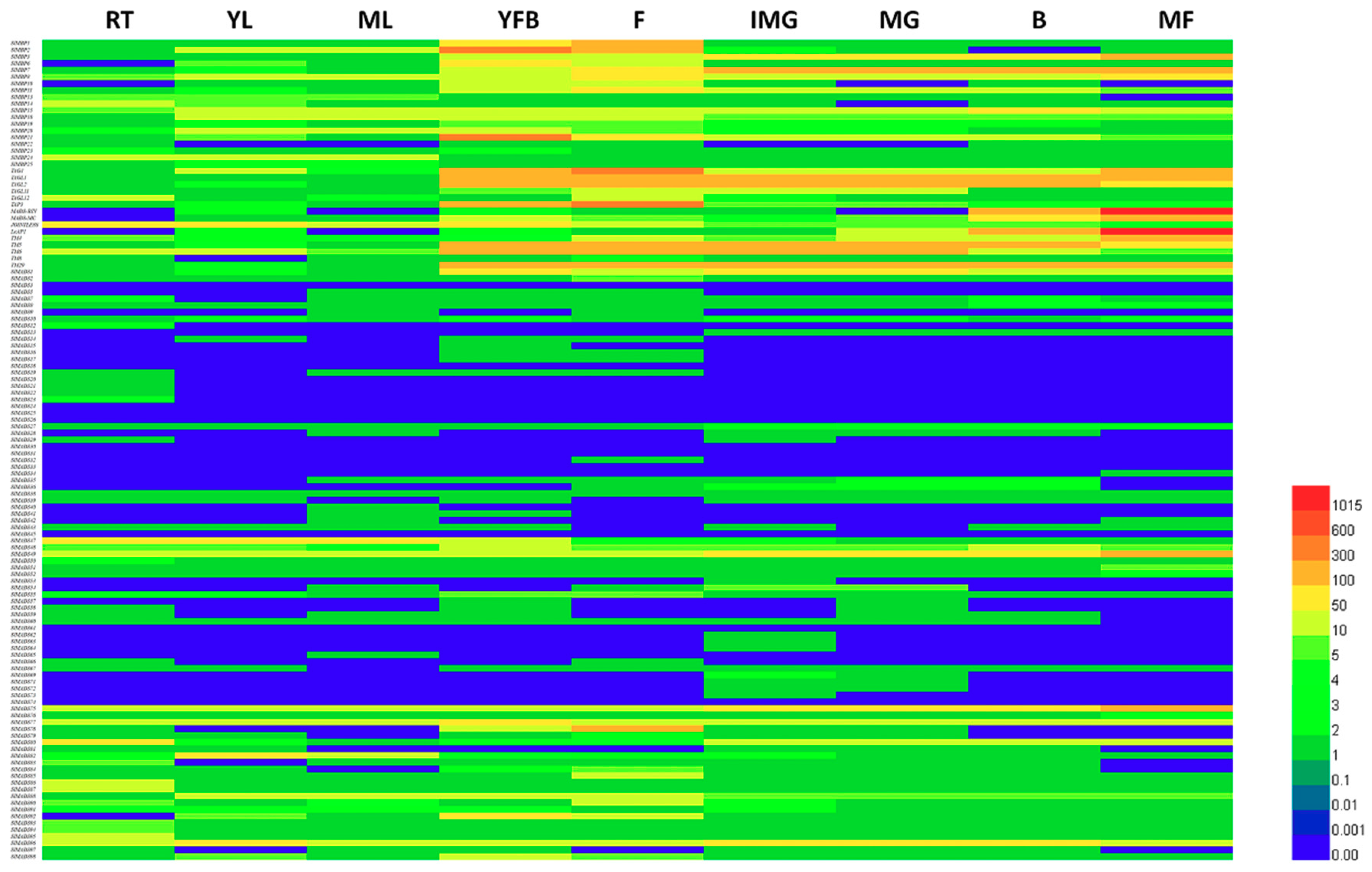
2.5. Predictions of Expression Profiles of Tomato MADS-Box Genes in Different Organs
2.6. The Critical Tomato MADS-Box Genes Involved in Floral Organ Development
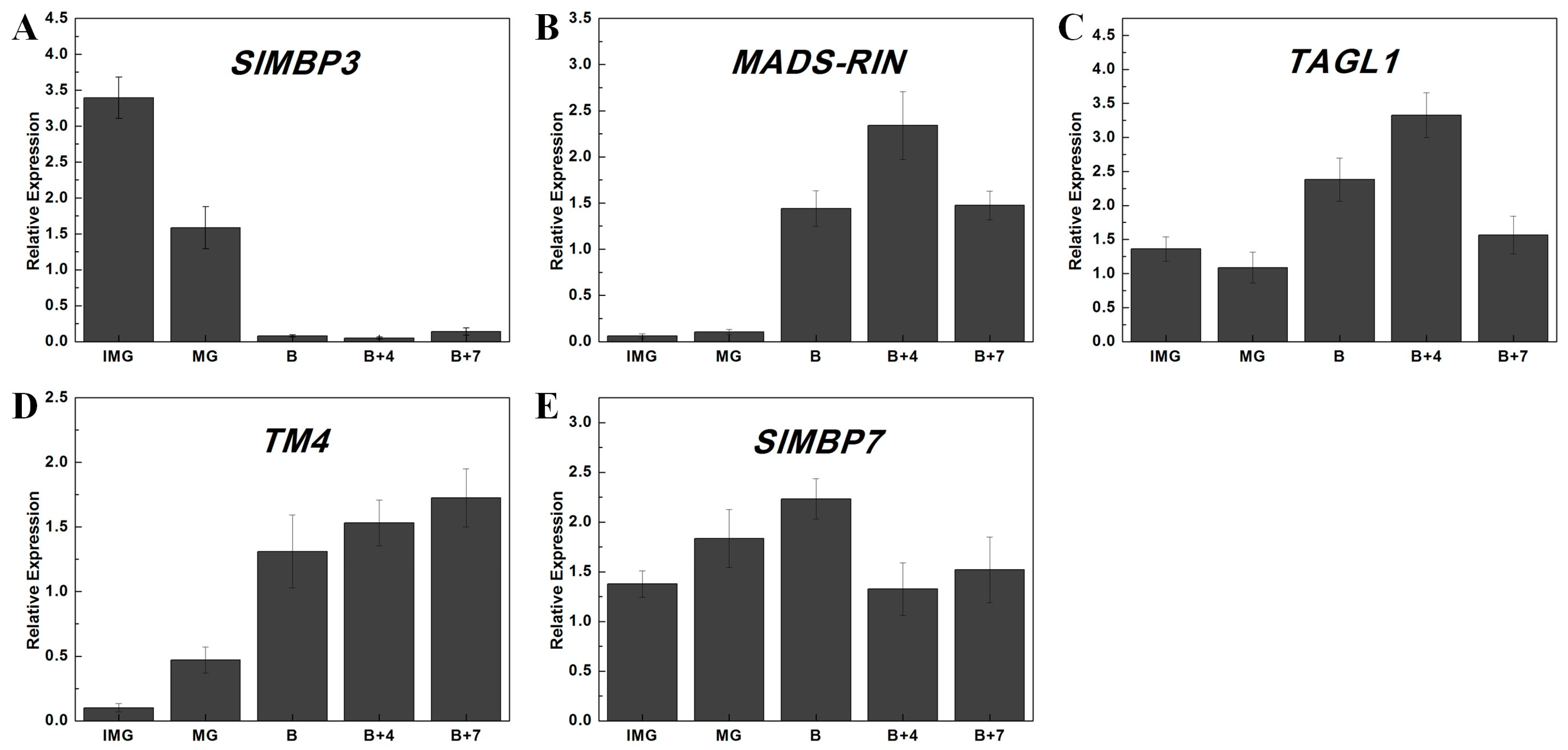
2.7. Differential Expression Analysis of Tomato MADS-Box Genes at Different Stages of Fruit Development and Ripening
3. Discussion
3.1. Characterization of MADS-Box Genes in Tomato
3.2. Prediction of MADS-Box Genes Involved in the Regulation of Flower Development and Floral Organ Identity
3.3. Potential Functions of Tomato MADS-Box Genes during Fruit Development
4. Materials and Methods
4.1. Plant Material and Growth Condition
4.2. Identification of MADS-Box Genes in Tomato
4.3. Phylogenetic Analysis of Tomato MADS-Box Proteins
4.4. The Analysis of Gene Structure and Conserved Motif
4.5. Chromosomal Locations and Identification of Interaction Network
4.6. Digital Gene Expression Analysis of Tomato MADS-Box Genes
4.7. Total RNA Extraction and qPCR Analysis
4.8. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Shore, P.; Sharrocks, A.D. The MADS-box family of transcription factors. Eur. J. Biochem. 1995, 229, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Schwarz-Sommer, Z.; Sommer, H. Genetic Control of Flower Development by Homeotic Genes in Antirrhinum majus. Science 1990, 250, 931–936. [Google Scholar] [CrossRef] [PubMed]
- Alvarezbuylla, E.R.; Pelaz, S.; Liljegren, S.J.; Gold, S.E.; Burgeff, C.; Ditta, G.S.; Ribas, d.P.L.; Martínezcastilla, L.; Yanofsky, M.F. An ancestral MADS-box gene duplication occurred before the divergence of plants and animals. Proc. Natl. Acad. Sci. USA 2000, 97, 5328–5333. [Google Scholar] [CrossRef] [PubMed]
- Smaczniak, C.; Immink, R.G.; Angenent, G.C.; Kaufmann, K. Developmental and evolutionary diversity of plant MADS-domain factors: Insights from recent studies. Development 2012, 139, 3081–3098. [Google Scholar] [CrossRef] [PubMed]
- Henschel, K.; Kofuji, R.; Hasebe, M.; Saedler, H.; Münster, T.; Theissen, G. Two ancient classes of MIKC-type MADS-box genes are present in the moss Physcomitrella patens. Mol. Biol. Evol. 2002, 19, 801–814. [Google Scholar] [CrossRef] [PubMed]
- Parenicová, L.; De, F.S.; Kieffer, M.; Horner, D.S.; Favalli, C.; Busscher, J.; Cook, H.E.; Ingram, R.M.; Kater, M.M.; Davies, B. Molecular and phylogenetic analyses of the complete MADS-box transcription factor family in Arabidopsis: New openings to the MADS world. Plant Cell 2003, 15, 1538–1551. [Google Scholar] [CrossRef] [PubMed]
- Arora, R.; Agarwal, P.; Ray, S.; Singh, A.K.; Singh, V.P.; Tyagi, A.K.; Kapoor, S. MADS-box gene family in rice: Genome-wide identification, organization and expression profiling during reproductive development and stress. BMC Genom. 2007, 8, 242. [Google Scholar] [CrossRef]
- Yanofsky, M.F.; Ma, H.; Bowman, J.L.; Drews, G.N.; Feldmann, K.A.; Meyerowitz, E.M. The protein encoded by the Arabidopsis homeotic gene agamous resembles transcription factors. Nature 1990, 346, 35–39. [Google Scholar] [CrossRef]
- Coen, E.S.; Meyerowitz, E.M. The war of the whorls: Genetic interactions controlling flower development. Nature 1991, 353, 31–37. [Google Scholar] [CrossRef]
- Causier, B.; Schwarz-Sommer, Z.; Davies, B. Floral organ identity: 20 years of ABCs. Semin. Cell Dev. Biol. 2010, 21, 73–79. [Google Scholar] [CrossRef]
- Krizek, B.A.; Meyerowitz, E.M. The Arabidopsis homeotic genes APETALA3 and PISTILLATA are sufficient to provide the B class organ identity function. Development 1996, 122, 11–22. [Google Scholar] [PubMed]
- Theissen, G. Development of floral organ identity: Stories from the MADS house. Curr. Opin. Plant Biol. 2001, 4, 75–85. [Google Scholar] [CrossRef]
- Mandel, M.A.; Gustafson-Brown, C.; Savidge, B.; Yanofsky, M.F. Molecular characterization of the Arabidopsis floral homeotic gene APETALA1. Nature 1992, 360, 273–277. [Google Scholar] [CrossRef] [PubMed]
- Jack, T.; Brockman, L.L.; Meyerowitz, E.M. The homeotic gene APETALA3 of Arabidopsis thaliana encodes a MADS box and is expressed in petals and stamens. Cell 1992, 68, 683–697. [Google Scholar] [CrossRef]
- Mizukami, Y.; Ma, H. Ectopic expression of the floral homeotic gene AGAMOUS in transgenic Arabidopsis plants alters floral organ identity. Cell 1992, 71, 119–131. [Google Scholar] [CrossRef]
- Pelaz, S.; Ditta, G.S.; Baumann, E.; Wisman, E.; Yanofsky, M.F. B and C floral organ identity functions require SEPALLATA MADS-box genes. Nature 2000, 405, 200–203. [Google Scholar] [CrossRef]
- Angenent, G.C.; Franken, J.; Busscher, M.; Van, D.A.; van Went, J.L.; Dons, H.J.; van Tunen, A.J. A novel class of MADS box genes is involved in ovule development in petunia. Plant Cell 1995, 7, 1569–1582. [Google Scholar] [CrossRef]
- Colombo, L.; Franken, J.; Koetje, E.; Went, J.V.; Dons, H.J.; Angenent, G.C.; Tunen, A.J.V. The petunia MADS box gene FBP11 determines ovule identity. Plant Cell 1995, 7, 1859–1868. [Google Scholar] [CrossRef]
- Vrebalov, J.; Ruezinsky, D.; Padmanabhan, V.; White, R.; Medrano, D.; Drake, R.; Schuch, W.; Giovannoni, J. A MADS-box gene necessary for fruit ripening at the tomato ripening-inhibitor (rin) locus. Science 2002, 296, 343–346. [Google Scholar] [CrossRef]
- Itkin, M.; Seybold, H.; Breitel, D.; Rogachev, I.; Meir, S.; Aharoni, A. TOMATO AGAMOUS-LIKE 1 is a component of the fruit ripening regulatory network. Plant J. Cell Mol. Biol. 2009, 60, 1081–1095. [Google Scholar] [CrossRef]
- Arroyo, A.J.M.; Mcquinn, R.; Poole, M.; Seymour, G.; Giovannoni, J.; Irish, V.F.; Vrebalov, J.; Pan, I.L.; Chung, M.; Rose, J. Fleshy fruit expansion and ripening are regulated by the Tomato SHATTERPROOF gene TAGL1. Plant Cell 2009, 21, 3041–3062. [Google Scholar]
- Bemer, M.; Karlova, R.; Ballester, A.R.; Tikunov, Y.M.; Bovy, A.G.; Wolters-Arts, M.; Rossetto, P.B.; Angenent, G.C.; de Maagd, R.A. The Tomato FRUITFULL Homologs TDR4/FUL1 and MBP7/FUL2 Regulate Ethylene-Independent Aspects of Fruit Ripening. Plant Cell 2012, 24, 4437–4451. [Google Scholar] [CrossRef] [PubMed]
- Shima, Y.; Kitagawa, M.; Fujisawa, M.; Nakano, T.; Kato, H.; Kimbara, J.; Kasumi, T.; Ito, Y. Tomato FRUITFULL homologues act in fruit ripening via forming MADS-box transcription factor complexes with RIN. Plant Mol. Biol. 2013, 82, 427–438. [Google Scholar] [CrossRef] [PubMed]
- Byeong-ha Lee, D.A.H.; Zhu, J.-K. The Arabidopsis Cold-Responsive Transcriptome and Its Regulation by ICE1. Plant Cell 2005, 17, 3155–3175. [Google Scholar] [PubMed]
- Lee, I.; Ronald, P.C. Genetic dissection of the biotic stress response using a genome-scale gene network for rice. Proc. Natl. Acad. Sci. USA 2011, 108, 18548–18553. [Google Scholar] [CrossRef] [PubMed]
- Tardif, G.; Kane, N.A.; Adam, H.; Labrie, L.; Major, G.; Gulick, P.; Sarhan, F.; Laliberté, J.F. Interaction network of proteins associated with abiotic stress response and development in wheat. Plant Mol. Biol. 2007, 63, 703–718. [Google Scholar] [CrossRef] [PubMed]
- Duan, W.; Song, X.; Liu, T.; Huang, Z.; Ren, J.; Hou, X.; Li, Y. Genome-wide analysis of the MADS-box gene family in Brassica rapa (Chinese cabbage). Mol. Genet. Genom. 2015, 290, 239–255. [Google Scholar] [CrossRef]
- Sommer, H.; Beltrán, J.P.; Huijser, P.; Pape, H.; Lönnig, W.E.; Saedler, H.; Schwarz-Sommer, Z. Deficiens, a homeotic gene involved in the control of flower morphogenesis in Antirrhinum majus: The protein shows homology to transcription factors. Embo J. 1990, 9, 605–613. [Google Scholar] [CrossRef]
- Leseberg, C.H.; Li, A.; Kang, H.; Duvall, M.; Mao, L. Genome-wide analysis of the MADS-box gene family in Populus trichocarpa. Gene 2006, 378, 84–94. [Google Scholar] [CrossRef]
- Jian, M.; Yang, Y.; Wei, L.; Yang, C.; Ding, P.; Liu, Y.; Qiao, L.; Chang, Z.; Geng, H.; Wang, P. Genome-wide identification and analysis of the MADS-box gene family in bread wheat (Triticum aestivum L.). PLoS ONE 2017, 12, e0181443. [Google Scholar]
- Liu, J.; Zhang, J.; Zhang, J.; Miao, H.; Wang, J.; Gao, P.; Hu, W.; Jia, C.; Wang, Z.; Xu, B. Genome-wide analysis of banana MADS-box family closely related to fruit development and ripening. Sci. Rep. 2017, 7, 3467. [Google Scholar] [CrossRef] [PubMed]
- Busscherlange, J. Analysis of the petunia MADS-box transcription factor family. Mol. Genet. Genom. 2003, 268, 598–606. [Google Scholar]
- Velasco, R.; Zharkikh, A.; Affourtit, J.; Dhingra, A.; Cestaro, A.; Kalyanaraman, A.; Fontana, P.; Bhatnagar, S.K.; Troggio, M.; Pruss, D.; et al. The genome of the domesticated apple (Malus x domestica Borkh.). Nat. Genet. 2010, 42, 833–839. [Google Scholar] [CrossRef] [PubMed]
- Leseberg, C.H.; Eissler, C.L.; Wang, X.; Johns, M.A.; Duvall, M.R.; Mao, L. Interaction study of MADS-domain proteins in tomato. J. Exp. Bot. 2008, 59, 2253–2265. [Google Scholar] [CrossRef] [PubMed]
- Hileman, L.C.; Sundstrom, J.F.; Litt, A.; Chen, M.; Shumba, T.; Irish, V.F. Molecular and phylogenetic analyses of the MADS-box gene family in tomato. Mol. Biol. Evol. 2006, 23, 2245–2258. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Hu, Z.; Yin, W.; Yu, X.; Zhu, Z.; Zhang, J.; Chen, G. The tomato floral homeotic protein FBP1-like gene, SlGLO1, plays key roles in petal and stamen development. Sci. Rep. 2016, 6, 20454. [Google Scholar] [CrossRef] [PubMed]
- Martino, G.D.; Pan, I.; Emmanuel, E.; Levy, A.; Irish, V.F. Functional Analyses of Two Tomato APETALA3 Genes Demonstrate Diversification in Their Roles in Regulating Floral Development. Plant Cell 2006, 18, 1833–1845. [Google Scholar] [CrossRef]
- Huang, B.; Routaboul, J.M.; Liu, M.; Deng, W.; Maza, E.; Mila, I.; Hu, G.; Zouine, M.; Frasse, P.; Vrebalov, J.T. Overexpression of the class D MADS-box gene Sl-AGL11 impacts fleshy tissue differentiation and structure in tomato fruits. J. Exp. Bot. 2017, 68, 4869–4884. [Google Scholar] [CrossRef]
- Ampomahdwamena, C.; Morris, B.A.; Sutherland, P.; Veit, B.; Yao, J.L. Down-regulation of TM29, a tomato SEPALLATA homolog, causes parthenocarpic fruit development and floral reversion. Plant Physiol. 2002, 130, 605–617. [Google Scholar] [CrossRef]
- Litt, A.; Irish, V.F. Duplication and Diversification in the APETALA1/FRUITFULL Floral Homeotic Gene Lineage: Implications for the Evolution of Floral Development. Genetics 2003, 165, 821–833. [Google Scholar]
- Yao, J.L.; Ampomah-Dwamena, C. The tomato SEPALLATA homolog TM29 has diverse functions in regulating flower and fruit development. Plant Physiol. 2003, 35, 28–33. [Google Scholar]
- Wang, S.; Gang, L.; Zheng, H.; Luo, Z.; Wang, T.; Li, H.; Zhang, J.; Ye, Z. Members of the tomato FRUITFULL MADS-box family regulate style abscission and fruit ripening. J. Exp. Bot. 2014, 65, 3005–3014. [Google Scholar] [CrossRef] [PubMed]
- Shima, Y.; Fujisawa, M.; Kitagawa, M.; Nakano, T.; Kimbara, J.; Nakamura, N.; Shiina, T.; Sugiyama, J.; Nakamura, T.; Kasumi, T. Tomato FRUITFULL homologs regulate fruit ripening via ethylene biosynthesis. J. Agric. Chem. Soc. Jpn. 2014, 78, 231–237. [Google Scholar]
- Burko, Y.; Shleizer-Burko, S.; Yanai, O.; Shwartz, I.; Zelnik, I.D.; Jacob-Hirsch, J.; Kela, I.; Eshed-Williams, L.; Ori, N. A role for APETALA1/fruitfull transcription factors in tomato leaf development. Plant Cell 2013, 25, 2070–2083. [Google Scholar] [CrossRef] [PubMed]
- Yin, W.; Hu, Z.; Hu, J.; Zhu, Z.; Yu, X.; Cui, B.; Chen, G. Tomato (Solanum lycopersicum) MADS-box transcription factor SlMBP8 regulates drought, salt tolerance and stress-related genes. Plant Growth Regul. 2017, 83, 55–68. [Google Scholar] [CrossRef]
- Yin, W.; Hu, Z.; Cui, B.; Guo, X.; Hu, J.; Zhu, Z.; Chen, G. Suppression of the MADS-box gene SlMBP8 accelerates fruit ripening of tomato (Solanum lycopersicum). Plant Physiol. Biochem. 2017, 118, 235–244. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Chen, G.; Cui, B.; Gao, Q.; Guo, J.E.; Li, A.; Zhang, L.; Hu, Z. Solanum lycopersicum agamous-like MADS-box protein AGL15-like gene, SlMBP11, confers salt stress tolerance. Mol. Breed. 2016, 36, 125. [Google Scholar] [CrossRef]
- Xie, Q.; Hu, Z.; Zhu, Z.; Dong, T.; Zhao, Z.; Cui, B.; Chen, G. Overexpression of a novel MADS-box gene SlFYFL delays senescence, fruit ripening and abscission in tomato. Sci. Rep. 2014, 4, 4367. [Google Scholar] [CrossRef]
- Li, N.; Huang, B.; Tang, N.; Jian, W.; Zou, J.; Chen, J.; Cao, H.; Habib, S.; Dong, X.; Wei, W. The MADS-box Gene SlMBP21 Regulates Sepal Size Mediated by Ethylene and Auxin in Tomato. Plant Cell Physiol. 2017, 58, 2241–2256. [Google Scholar] [CrossRef]
- Liu, D.; Wang, D.; Qin, Z.; Zhang, D.; Yin, L.; Wu, L.; Colasanti, J.; Li, A.; Mao, L. The SEPALLATA MADS-box protein SLMBP21 forms protein complexes with JOINTLESS and MACROCALYX as a transcription activator for development of the tomato flower abscission zone. Plant J. 2014, 77, 284–296. [Google Scholar] [CrossRef]
- Roldan, M.V.G.; Périlleux, C.; Morin, H.; Huerga-Fernandez, S.; Latrasse, D.; Benhamed, M.; Bendahmane, A. Natural and induced loss of function mutations inSlMBP21MADS-box gene led tojointless-2phenotype in tomato. Sci. Rep. 2017, 7, 4402. [Google Scholar] [CrossRef] [PubMed]
- Gimenez, E.; Castañeda, L.; Pineda, B.; Pan, I.L.; Moreno, V.; Angosto, T.; Lozano, R. TOMATO AGAMOUS1 and ARLEQUIN/TOMATO AGAMOUS—LIKE1 MADS-box genes have redundant and divergent functions required for tomato reproductive development. Plant Mol. Biol. 2016, 91, 513–531. [Google Scholar] [CrossRef] [PubMed]
- Pnueli, L.; Hareven, D.; Rounsley, S.D.; Yanofsky, M.F.; Lifschitz, E. Isolation of the Tomato AGAMOUS Gene TAG1 and Analysis of Its Homeotic Role in Transgenic Plants. Plant Cell 1994, 6, 163–173. [Google Scholar] [CrossRef] [PubMed]
- Giménez, E.; Pineda, B.; Capel, J.; Antón, M.T.; Atarés, A.; Pérezmartín, F.; Garcíasogo, B.; Angosto, T.; Moreno, V.; Lozano, R. Functional analysis of the Arlequin mutant corroborates the essential role of the ARLEQUIN/TAGL1 gene during reproductive development of tomato. PLoS ONE 2010, 5, e14427. [Google Scholar] [CrossRef] [PubMed]
- Gaffe, J.; Lemercier, C.; Alcaraz, J.P.; Kuntz, M. Identification of three tomato flower and fruit MADS-box proteins with a putative histone deacetylase binding domain. Gene 2011, 471, 19–26. [Google Scholar] [CrossRef] [PubMed]
- Busi, M.V.; Bustamante, C.; D’Angelo, C.; Hidalgocuevas, M.; Boggio, S.B.; Valle, E.M.; Zabaleta, E. MADS-box genes expressed during tomato seed and fruit development. Plant Mol. Biol. 2003, 52, 801–815. [Google Scholar] [PubMed]
- Kramer, E.M.; Dorit, R.L.; Irish, V.F. Molecular evolution of genes controlling petal and stamen development: Duplication and divergence within the APETALA3 and PISTILLATA MADS-box gene lineages. Genetics 1998, 149, 765–783. [Google Scholar]
- Pracros, P.; Renaudin, J.; Eveillard, S.; Mouras, A.; Hernould, M. Tomato flower abnormalities induced by stolbur phytoplasma infection are associated with changes of expression of floral development genes. Mol. Plant Microbe Interact. 2006, 19, 62–68. [Google Scholar] [CrossRef]
- Fujisawa, M.; Shima, Y.; Nakagawa, H.; Kitagawa, M.; Kimbara, J.; Nakano, T.; Kasumi, T.; Ito, Y. Transcriptional regulation of fruit ripening by tomato FRUITFULL homologs and associated MADS box proteins. Plant Cell 2014, 26, 89–101. [Google Scholar] [CrossRef]
- Ito, Y. Regulation of Tomato Fruit Ripening by MADS-Box Transcription Factors. Jpn. Agric. Res. Q. 2016, 50, 33–38. [Google Scholar] [CrossRef][Green Version]
- Martel, C.; Vrebalov, J.; Tafelmeyer, P.; Giovannoni, J.J. The tomato (Solanum lycopersicum) MADS-box transcription factor RIN interacts with promoters involved in numerous ripening processes in a CNR dependent manner. Plant Physiol. 2011, 157, 1568–1579. [Google Scholar] [CrossRef] [PubMed]
- Ito, Y.; Kitagawa, M.; Ihashi, N.; Yabe, K.; Kimbara, J.; Yasuda, J.; Ito, H.; Inakuma, T.; Hiroi, S.; Kasumi, T. DNA-binding specificity, transcriptional activation potential, and the rin mutation effect for the tomato fruit-ripening regulator RIN. Plant J. 2008, 55, 212–223. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Zhu, B.; Fu, D.; Luo, Y. RIN transcription factor plays an important role in ethylene biosynthesis of tomato fruit ripening. J. Sci. Food Agric. 2011, 91, 2308–2314. [Google Scholar] [CrossRef] [PubMed]
- Nakano, T.; Kimbara, J.; Fujisawa, M.; Kitagawa, M.; Ihashi, N.; Maeda, H.; Kasumi, T.; Ito, Y. MACROCALYX and JOINTLESS interact in the transcriptional regulation of tomato fruit abscission zone development. Plant Physiol. 2012, 158, 439–450. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yin, J. Functional Studies of JOINTLESS and Its Interacting MADS-domain Proteins in Tomato. Acta Hortic. Sin. 2011, 38, 701–708. [Google Scholar]
- Nakano, T.; Kato, H.; Shima, Y.; Ito, Y. Apple SVP family MADS-box proteins and the tomato pedicel abscission zone regulator JOINTLESS have similar molecular activities. Plant Cell Physiol. 2015, 56, 1097–1106. [Google Scholar] [CrossRef] [PubMed]
- Szymkowiak, E.J.; Irish, E.E. JOINTLESS suppresses sympodial identity in inflorescence meristems of tomato. Planta 2006, 223, 646–658. [Google Scholar] [CrossRef]
- Mao, L.; Begum, D.; Chuang, H.W.; Budiman, M.A.; Szymkowiak, E.J.; Irish, E.E.; Wing, R.A. JOINTLESS is a MADS-box gene controlling tomato flower abscission zone development. Nature 2000, 406, 910–913. [Google Scholar] [CrossRef]
- Pnueli, L.; Abuabeid, M.; Zamir, D.; Nacken, W.; Schwarzsommer, Z.; Lifschitz, E. The MADS box gene family in tomato: Temporal expression during floral development, conserved secondary structures and homology with homeotic genes from Antirrhinum and Arabidopsis. Plant J. 1991, 1, 255–266. [Google Scholar] [CrossRef]
- Daminato, M.; Masiero, S.; Resentini, F.; Lovisetto, A.; Casadoro, G. Characterization of TM8, a MADS-box gene expressed in tomato flowers. BMC Plant Biol. 2014, 14, 319. [Google Scholar] [CrossRef]
- Dong, T.; Hu, Z.; Deng, L.; Wang, Y.; Zhu, M.; Zhang, J.; Chen, G. A Tomato MADS-Box Transcription Factor, SlMADS1, Acts as a Negative Regulator of Fruit Ripening. Plant Physiol. 2013, 163, 1026–1036. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Hu, Z.; Yao, Q.; Guo, X.; Nguyen, V.; Li, F.; Chen, G. A tomato MADS-box protein, SlCMB1, regulates ethylene biosynthesis and carotenoid accumulation during fruit ripening. Sci. Rep. 2018, 8, 3413. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Ming, M.; Li, J.; Shi, D.; Qiao, X.; Li, L.; Zhang, S.; Wu, J. Genome-wide identification of the MADS-box transcription factor family in pear (Pyrus bretschneideri) reveals evolution and functional divergence. Peerj 2017, 5, e3776. [Google Scholar] [CrossRef] [PubMed]
- Hughes, A.L. The evolution of functionally novel proteins after gene duplication. Proc. Biol. Sci. 1994, 256, 119–124. [Google Scholar] [PubMed]
- Alvarez-Buylla, E.R.; Liljegren, S.J.; Pelaz, S.; Gold, S.E.; Burgeff, C.; Ditta, G.S.; Vergara-Silva, F.; Yanofsky, M.F. MADS-box gene evolution beyond flowers: Expression in pollen, endosperm, guard cells, roots and trichomes. Plant J. 2000, 24, 457–466. [Google Scholar] [CrossRef] [PubMed]
- Causier, B.; Kieffer, M.; Davies, B. Plant biology. MADS-box genes reach maturity. Science 2002, 296, 275–276. [Google Scholar] [CrossRef] [PubMed]
- Rijpkema, A.; Gerats, T.; Vandenbussche, M. Genetics of Floral Development in Petunia. Adv. Bot. Res. 2006, 44, 237–278. [Google Scholar]
- Becker, A.; Theißen, G. The major clades of MADS-box genes and their role in the development and evolution of flowering plants. Mol. Phylogenet. Evol. 2003, 29, 464–489. [Google Scholar] [CrossRef]
- Rounsley, S.D.; Ditta, G.S.; Yanofsky, M.F. Diverse roles for MADS box genes in Arabidopsis development. Plant Cell 1995, 7, 1259–1269. [Google Scholar] [CrossRef]
- Airoldi, C.A.; Davies, B. Gene Duplication and the Evolution of Plant MADS-box Transcription Factors. J. Genet. Genom. 2012, 39, 157–165. [Google Scholar] [CrossRef]
- Liu, Y.; Jiang, H.Y.; Chen, W.; Qian, Y.; Ma, Q.; Cheng, B.; Zhu, S. Genome-wide analysis of the auxin response factor (ARF) gene family in maize (Zea mays). Plant Growth Regul. 2011, 63, 225–234. [Google Scholar] [CrossRef]
- Ma, H. The unfolding drama of flower development: Recent results from genetic and molecular analyses. Genes Dev. 1994, 8, 745–756. [Google Scholar] [CrossRef] [PubMed]
- Pelaz, S.; Tapialópez, R.; Alvarezbuylla, E.R.; Yanofsky, M.F. Conversion of leaves into petals in Arabidopsis. Curr. Biol. 2001, 11, 182–184. [Google Scholar] [CrossRef]
- Davies, B.; Egeacortines, M.; De, E.A.S.; Saedler, H.; Sommer, H. Multiple interactions amongst floral homeotic MADS box proteins. Embo J. 1996, 15, 4330–4343. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Koh, J.; Yoo, M.J.; Kong, H.; Hu, Y.; Ma, H.; Soltis, P.S.; Soltis, D.E. Expression of floral MADS-box genes in basal angiosperms: Implications for the evolution of floral regulators. Plant J. Cell Mol. Biol. 2005, 43, 724–744. [Google Scholar] [CrossRef] [PubMed]
- Theissen, G.; Becker, A.; Rosa, A.D.; Kanno, A.; Kim, J.T.; Münster, T.; Winter, K.U.; Saedler, H. A short history of MADS-box genes in plants. Plant Mol. Biol. 2000, 42, 115–149. [Google Scholar] [CrossRef] [PubMed]
- Angenent, G.C.; Franken, J.; Busscher, M.; Colombo, L.; van Tunen, A.J. Petal and stamen formation in petunia is regulated by the homeotic gene fbp1. Plant J. 1993, 4, 101–112. [Google Scholar] [CrossRef] [PubMed]
- Bowman, J.L.; Alvarez, J.; Weigel, D.; Meyerowitz, E.M.; Smyth, D.R. Control of flower development in Arabidopsis thaliana by APETALA1 and interacting genes. Development 1993, 119, 721–743. [Google Scholar]
- Poupin, M.; Federici, F.; Medina, C.; Mattis, J.; Timmermann, T.J.P. Isolation of the three grape sub-lineages of B-class MADS-box TM6, PISTILLATA and APETALA3 genes which are differentially expressed during flower and fruit development. Gene 2007, 404, 10–24. [Google Scholar] [CrossRef]
- Cheng, X.F.; Wittich, P.E.; Kieft, H.; Angenent, G.; Xuhan, X.; Lammeren, A.A.M.V. Temporal and Spatial Expression of MADS Box Genes, FBP7 and FBP11, During Initiation and Early Development of Ovules in Wild Type and Mutant Petunia hybrida. Plant Biol. 2000, 2, 693–702. [Google Scholar] [CrossRef]
- Fujisawa, M.; Nakano, T.; Shima, Y.; Ito, Y. A large-scale identification of direct targets of the tomato MADS box transcription factor RIPENING INHIBITOR reveals the regulation of fruit ripening. Plant Cell 2013, 25, 371–386. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. Molecular Evolutionary Genetics Analysis using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol. Biol. Evol. 2014, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Guo, A.Y.; Zhu, Q.H.; Chen, X.; Luo, J.C. GSDS: A gene structure display server. Hereditas 2007, 29, 1023–1026. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.J.; Zhang, M. Protein-Protein Interaction Networks; Springer: New York, NY, USA, 2013; pp. 141–181. [Google Scholar]
- Deng, W.; Wang, Y.; Liu, Z.; Cheng, H.; Xue, Y. HemI: A toolkit for illustrating heatmaps. PLoS ONE 2014, 9, e111988. [Google Scholar] [CrossRef] [PubMed]
- Expósito-Rodríguez, M.; Borges, A.A.; Borges-Pérez, A.; Pérez, J.A. Selection of internal control genes for quantitative real-time RT-PCR studies during tomato development process. BMC Plant Biol. 2008, 8, 131. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]



| Gene Name | Gene Locus | Protein | Type | Reference | ||
|---|---|---|---|---|---|---|
| Length (aa) | MW (Da) | pI | ||||
| SlMBP1/SlGLO1/PI/LePI-B | Solyc08g067230.2.1 | 210 | 24740.2 | 8.69 | Type II | [36] |
| SlMBP2/SlGLO2/LePI/TPI | Solyc06g059970.2.1 | 214 | 24867.4 | 9.49 | Type II | [37] |
| SlMBP3/ SlAGL11 | Solyc06g064840.3.1 | 237 | 27469.6 | 9.25 | Type II | [38] |
| SlMBP6/SlAGL6 | Solyc01g093960.2.1 | 252 | 28608.5 | 8.37 | Type II | [39,40,41] |
| SlMBP7/LeFUL2 | Solyc03g114830.2.1 | 247 | 28614.3 | 9.31 | Type II | [22,40,42,43,44] |
| SlMBP8 | Solyc12g087830.1.1 | 198 | 22643.3 | 8.97 | Type II | [45,46] |
| SlMBP10 | Solyc02g065730.1.1 | 234 | 27368.1 | 10.12 | Type II | |
| SlMBP11/AGL15-like | Solyc01g087990.2.1 | 271 | 30847 | 6.01 | Type II | [47] |
| SlMBP13 | Solyc08g080100.2.1 | 224 | 25839 | 9.76 | Type II | |
| SlMBP14 | Solyc12g056460.1.1 | 206 | 23968.4 | 6.57 | Type II | |
| SlMBP15 | Solyc12g087830.2.1 | 204 | 23667.2 | 8.35 | Type II | |
| SlMBP18/SlFYFL | Solyc03g006830.2.1 | 222 | 25369.2 | 9.46 | Type II | [48] |
| SlMBP19 | Solyc06g035570.1.1 | 54 | 6224.26 | 11.29 | Type I | |
| SlMBP20 | Solyc02g089210.2.1 | 250 | 28589.2 | 9.87 | Type II | [44] |
| SlMBP21 | Solyc12g038510.1.1 | 250 | 28442.4 | 9.26 | Type II | [49,50,51] |
| SlMBP22 | Solyc11g005120.1.1 | 238 | 27791.6 | 6.96 | Type II | |
| SlMBP23/TDR3 | Solyc10g017630.2.1 | 166 | 19196.3 | 9.49 | Type II | |
| SlMBP24 | Solyc04g076280.3.1 | 235 | 26483.6 | 7.67 | Type I | |
| SlMBP25 | Solyc05g015730.1.1 | 80 | 9311.97 | 11.01 | Type II | |
| TAG1 | Solyc02g071730.2.1 | 248 | 28723.6 | 9.97 | Type II | [52,53] |
| TAGL1 | Solyc07g055920.2.1 | 267 | 29940.5 | 9.56 | Type II | [20,21,52,54] |
| TAGL2 | Solyc05g015750.2.1 | 241 | 27579.3 | 9.07 | Type II | [55,56] |
| TAGL11 | Solyc11g028020.1.1 | 223 | 26051.7 | 9.76 | Type II | [38,56] |
| TAGL12 | Solyc11g032100.1.1 | 201 | 23104.8 | 6.94 | Type II | [56] |
| TAP3/LeAP3/LeDEF | Solyc04g081000.2.1 | 228 | 26478.2 | 9.76 | Type II | [54,57,58,59] |
| MADS-RIN | Solyc05g012020.2.1 | 242 | 27968.4 | 8.00 | Type II | [19,60,61,62,63] |
| MADS-MC | Solyc05g056620.1.1 | 244 | 28660.6 | 8.44 | Type II | [64] |
| JOINTLESS | Solyc11g010570.1.1 | 265 | 30426.3 | 7.39 | Type II | [64,65,66,67,68] |
| LeAP1 | Solyc05g012020.3.1 | 213 | 25077.3 | 5.47 | Type II | [40] |
| TM4/TDR4/LeFUL1 | Solyc06g069430.2.1 | 245 | 28290 | 9.57 | Type II | [22,42,43,59] |
| TM5/TDR5/LeSEP3 | Solyc05g015750.3.1 | 224 | 25999.3 | 9.79 | Type II | [40] |
| TM6/TDR6 | Solyc02g084630.2.1 | 225 | 26092.7 | 9.71 | Type II | [69] |
| TM8/TDR8 | Solyc03g019710.2.1 | 177 | 20703.9 | 10.44 | Type II | [70] |
| SlMADS1 | Solyc03g114840.2.1 | 246 | 28398.2 | 8.88 | Type II | [55,71] |
| SlMADS2 | Solyc01g060300.1.1 | 211 | 23122.6 | 9.36 | Type I | |
| SlMADS3 | Solyc01g060310.1.1 | 139 | 15117.5 | 8.99 | Type I | |
| SlMADS4 | Solyc01g060284.1.1 | 197 | 21565.5 | 8.63 | Type I | |
| SlMADS5 | Solyc01g066500.1.1 | 309 | 34544.4 | 4.75 | Type I | |
| SlMADS6/TM29/LeSEP1 | Solyc02g089200.2.1 | 246 | 28481.3 | 8.29 | Type II | [39,41] |
| SlMADS7 | Solyc01g103870.1.1 | 125 | 14435.6 | 9.54 | Type I | |
| SlMADS8 | Solyc06g071300.1.1 | 130 | 15009.1 | 5.14 | Type I | |
| SlMADS9 | Solyc03g007020.1.1 | 215 | 24702 | 9.21 | Type I | |
| SlMADS10 | Solyc04g064860.1.1 | 233 | 25871.3 | 7.92 | Type I | |
| SlMADS11 | Solyc04g054517.1.1 | 138 | 15663.1 | 7.75 | Type I | |
| SlMADS12 | Solyc04g056550.1.1 | 182 | 20838.7 | 5.14 | Type I | |
| SlMADS13 | Solyc04g056740.1.1 | 180 | 20521.5 | 7.44 | Type I | |
| SlMADS14 | Solyc06g054680.1.1 | 180 | 21152.8 | 7.19 | Type I | |
| SlMADS15 | Solyc06g059780.1.1 | 180 | 21057.6 | 6.37 | Type I | |
| SlMADS16 | Solyc10g050950.1.1 | 161 | 18602.2 | 6.80 | Type I | |
| SlMADS17 | Solyc10g050900.1.1 | 169 | 19796.5 | 7.83 | Type I | |
| SlMADS18 | Solyc10g050940.1.1 | 169 | 19623.3 | 7.77 | Type I | |
| SlMADS19 | Solyc01g097850.1.1 | 172 | 20073.9 | 7.35 | Type I | |
| SlMADS20 | Solyc03g034260.1.1 | 184 | 21441 | 6.29 | Type I | |
| SlMADS21 | Solyc10g018070.1.1 | 119 | 13063.9 | 10.14 | Type I | |
| SlMADS22 | Solyc10g018080.1.1 | 151 | 17218.5 | 7.04 | Type I | |
| SlMADS23 | Solyc10g018110.1.1 | 184 | 20762.4 | 6.55 | Type I | |
| SlMADS24 | Solyc04g025970.1.1 | 117 | 13333.4 | 10.16 | Type I | |
| SlMADS25 | Solyc01g010300.1.1 | 88 | 10052.7 | 10.79 | Type I | |
| SlMADS26 | Solyc02g032000.1.1 | 97 | 10944.7 | 10.32 | Type I | |
| SlMADS27 | Solyc03g119680.1.1 | 177 | 20293.2 | 7.26 | Type I | |
| SlMADS28 | Solyc09g061950.1.1 | 173 | 20138.1 | 9.33 | Type I | |
| SlMADS29 | Solyc01g106710.1.1 | 221 | 25105.7 | 10.23 | Type I | |
| SlMADS30 | Solyc04g025030.1.1 | 135 | 15411 | 10.37 | Type I | |
| SlMADS31 | Solyc04g025110.1.1 | 130 | 14313.7 | 10.26 | Type I | |
| SlMADS32 | Solyc01g106720.1.1 | 211 | 23531.2 | 10.35 | Type I | |
| SlMADS33 | Solyc01g106730.1.1 | 221 | 23888.6 | 10.14 | Type I | |
| SlMADS34 | Solyc04g047870.1.1 | 190 | 21695.2 | 10.70 | Type I | |
| SlMADS35 | Solyc10g012180.1.1 | 167 | 18824.5 | 5.38 | Type I | |
| SlMADS36 | Solyc10g012200.1.1 | 167 | 19113.1 | 8.98 | Type I | |
| SlMADS37 | Solyc11g067163.1.1 | 166 | 18919.9 | 7.67 | Type I | |
| SlMADS38 | Solyc01g066730.2.1 | 300 | 33023.4 | 5.04 | Type I | |
| SlMADS39 | Solyc01g098070.1.1 | 155 | 17200.8 | 10.09 | Type I | |
| SlMADS40 | Solyc01g098060.1.1 | 152 | 17191.5 | 9.84 | Type I | |
| SlMADS41 | Solyc03g062820.1.1 | 145 | 16350.8 | 9.23 | Type I | |
| SlMADS42 | Solyc01g098050.1.1 | 160 | 18178 | 10.45 | Type I | |
| SlMADS43 | Solyc05g013370.1.1 | 232 | 26476.7 | 5.46 | Type I | |
| SlMADS44 | Solyc05g046345.1.1 | 250 | 28489.8 | 5.47 | Type I | |
| SlMADS45 | Solyc04g025050.1.1 | 56 | 6259.52 | 11.07 | Type I | |
| SlMADS46 | Solyc07g017343.1.1 | 226 | 26532.7 | 5.2 | Type I | |
| SlMADS47 | Solyc00g179240.1.1 | 55 | 6310.11 | 11.16 | Type I | |
| SlMADS48 | Solyc07g052707.1.1 | 195 | 21878.8 | 4.86 | Type I | |
| SlMADS49 | Solyc07g052700.2.1 | 190 | 21334.2 | 7.41 | Type I | |
| SlMADS50 | Solyc11g069770.2.1 | 175 | 20171.9 | 5.45 | Type I | |
| SlMADS51 | Solyc01g106700.2.1 | 213 | 24279.5 | 7.35 | Type I | |
| SlMADS52 | Solyc03g115910.1.1 | 417 | 47275.1 | 5.82 | Type I | |
| SlMADS53 | Solyc10g012380.1.1 | 149 | 17086.7 | 5.29 | Type I | |
| SlMADS54 | Solyc10g012390.1.1 | 149 | 17086.7 | 5.29 | Type I | |
| SlMADS55 | Solyc11g069770.1.1 | 181 | 20857.8 | 5.20 | Type I | |
| SlMADS56 | Solyc12g042967.1.1 | 225 | 25552.3 | 9.05 | Type I | |
| SlMADS57 | Solyc11g020620.1.1 | 342 | 38747.9 | 5.42 | Type I | |
| SlMADS58 | Solyc11g020660.1.1 | 312 | 35558.7 | 4.61 | Type I | |
| SlMADS59 | Solyc11g020320.1.1 | 275 | 31757.7 | 6.68 | Type I | |
| SlMADS60 | Solyc01g103550.1.1 | 311 | 35839.7 | 7.87 | Type I | |
| SlMADS61 | Solyc01g060310.2.1 | 136 | 15230.6 | 6.16 | Type I | |
| SlMADS62 | Solyc12g016170.1.1 | 219 | 25087.7 | 6.57 | Type I | |
| SlMADS63 | Solyc12g016180.1.1 | 219 | 25207.8 | 6.57 | Type I | |
| SlMADS64 | Solyc12g016150.1.1 | 154 | 17645.1 | 5.38 | Type I | |
| SlMADS65 | Solyc12g017300.1.1 | 154 | 17740.1 | 5.11 | Type I | |
| SlMADS66 | Solyc12g005210.1.1 | 88 | 9985.65 | 10.41 | Type I | |
| SlMADS67 | Solyc01g102260.2.1 | 249 | 27882.8 | 9.66 | Type I | |
| SlMADS68 | Solyc05g047712.1.1 | 156 | 18199.1 | 8.89 | Type I | |
| SlMADS69 | Solyc07g043080.1.1 | 215 | 24384.8 | 6.86 | Type I | |
| SlMADS70 | Solyc06g034317.1.1 | 153 | 16949.3 | 4.41 | Type I | |
| SlMADS71 | Solyc06g048380.1.1 | 176 | 20038.6 | 4.90 | Type I | |
| SlMADS72 | Solyc06g048380.2.1 | 132 | 15027 | 4.82 | Type I | |
| SlMADS73 | Solyc06g033820.1.1 | 196 | 22657.4 | 7.25 | Type I | |
| SlMADS74 | Solyc06g033830.1.1 | 113 | 13058.8 | 8.44 | Type I | |
| SlMADS75 | Solyc07g052700.3.1 | 197 | 22133.2 | 8.44 | Type I | |
| SlMADS76 | Solyc03g115910.2.1 | 434 | 49552.6 | 5.6 | Type I | |
| SlMADS77 | Solyc12g087820.1.1 | 106 | 11969.2 | 11.03 | Type I | |
| SlMADS78 | Solyc05g051830.2.1 | 389 | 44560 | 5.91 | Type I | |
| SlMADS79 | Solyc10g017640.1.1 | 61 | 7032.22 | 11.24 | Type I | |
| SlMADS80 | Solyc04g076680.2.1 | 63 | 70492.3 | 11.03 | Type I | |
| SlMADS81 | Solyc12g088080.1.1 | 97 | 11295.2 | 10.59 | Type I | |
| SlMADS82 | Solyc01g105800.2.1 | 279 | 32104.7 | 9.11 | Type II | |
| SlMADS83 | Solyc01g106170.2.1 | 148 | 17032.2 | 10.98 | Type II | |
| SlMADS84 | Solyc10g080030.1.1 | 229 | 25791.3 | 6.37 | Type II | |
| SlMADS85 | Solyc01g105810.3.1 | 215 | 24518.2 | 9.22 | Type II | |
| SlMADS86 | Solyc04g078300.3.1 | 145 | 16547.2 | 9.9 | Type II | |
| SlMADS87 | Solyc04g078300.2.1 | 334 | 38130.7 | 5.96 | Type II | |
| SlMADS88 | Solyc03g006830.3.1 | 209 | 24310.2 | 9.43 | Type II | |
| SlMADS89 | Solyc10g044967.1.1 | 226 | 26172 | 6.14 | Type II | |
| SlMADS90 | Solyc11g032100.2.1 | 201 | 23104.8 | 6.45 | Type II | |
| SlMADS91 | Solyc05g015720.2.1 | 204 | 23433.5 | 5.36 | Type II | |
| SlMADS92 | Solyc01g093960.3.1 | 242 | 27359.1 | 8.19 | Type II | |
| SlMADS93 | Solyc02g091550.2.1 | 249 | 28342.2 | 9.07 | Type II | |
| SlMADS94 | Solyc02g091550.1.1 | 235 | 27198.1 | 9.65 | Type II | |
| SlMADS95 | Solyc04g076700.3.1 | 226 | 25933.3 | 8.59 | Type II | |
| SlMADS96 | Solyc03g019720.3.1 | 193 | 22631.1 | 9.88 | Type II | |
| SlMADS97 | Solyc12g088090.2.1 | 239 | 27695.8 | 9.84 | Type II | |
| SlMADS98/SlCMB1 | Solyc04g005320.2.1 | 238 | 27522.1 | 8.60 | Type II | [72] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Y.; Zhang, J.; Hu, Z.; Guo, X.; Tian, S.; Chen, G. Genome-Wide Analysis of the MADS-Box Transcription Factor Family in Solanum lycopersicum. Int. J. Mol. Sci. 2019, 20, 2961. https://doi.org/10.3390/ijms20122961
Wang Y, Zhang J, Hu Z, Guo X, Tian S, Chen G. Genome-Wide Analysis of the MADS-Box Transcription Factor Family in Solanum lycopersicum. International Journal of Molecular Sciences. 2019; 20(12):2961. https://doi.org/10.3390/ijms20122961
Chicago/Turabian StyleWang, Yunshu, Jianling Zhang, Zongli Hu, Xuhu Guo, Shibing Tian, and Guoping Chen. 2019. "Genome-Wide Analysis of the MADS-Box Transcription Factor Family in Solanum lycopersicum" International Journal of Molecular Sciences 20, no. 12: 2961. https://doi.org/10.3390/ijms20122961
APA StyleWang, Y., Zhang, J., Hu, Z., Guo, X., Tian, S., & Chen, G. (2019). Genome-Wide Analysis of the MADS-Box Transcription Factor Family in Solanum lycopersicum. International Journal of Molecular Sciences, 20(12), 2961. https://doi.org/10.3390/ijms20122961





