The Role of Ubiquitination in Regulating Embryonic Stem Cell Maintenance and Cancer Development
Abstract
1. Introduction
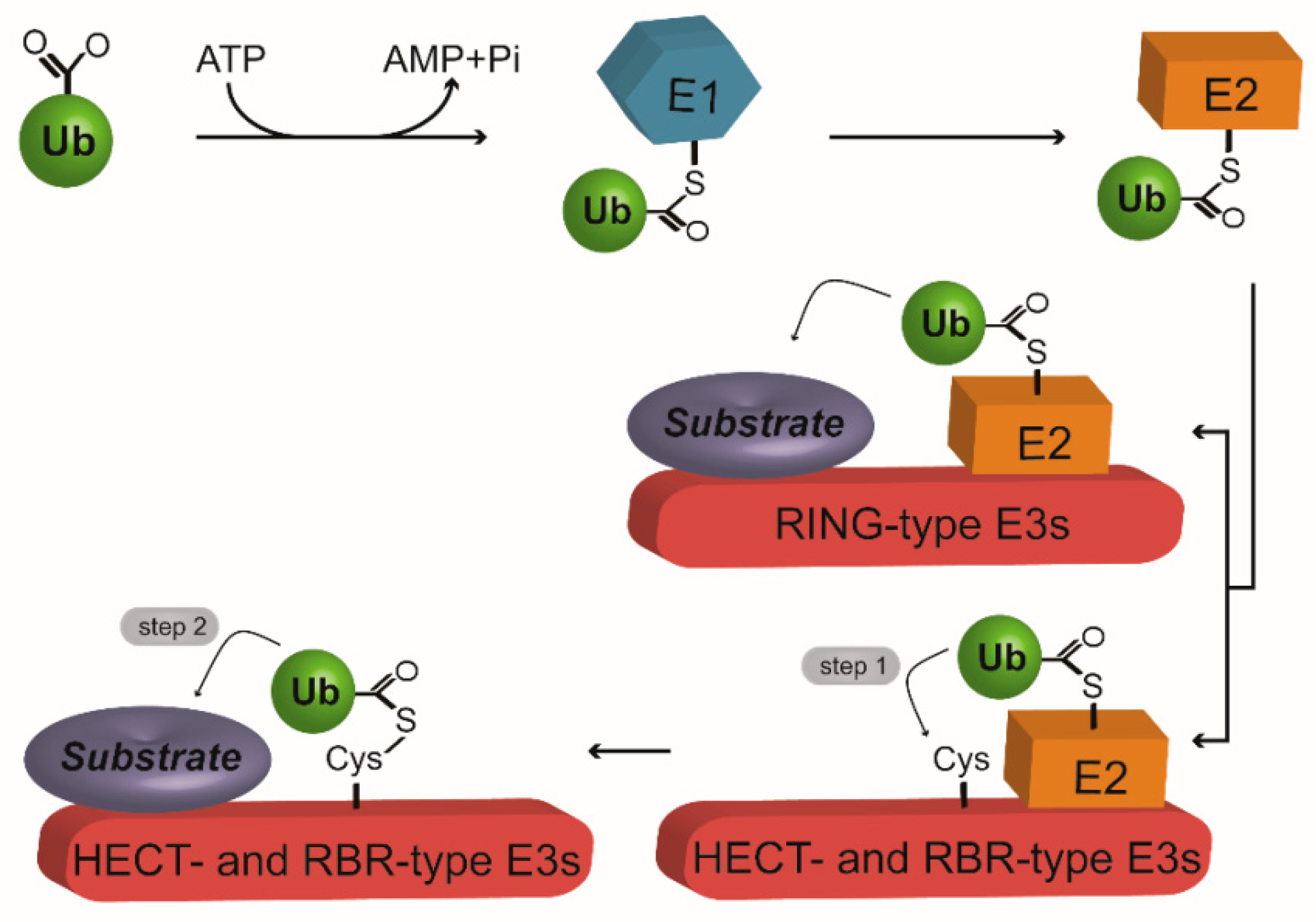
2. The Ubiquitination Machinery
3. The Types of Ubiquitination
4. Ubiquitination and Embryonic Stem Cells
4.1. Regulation of Stemness-Related TFs by Ubiquitination in ES Cells
4.2. Regulation of Signal Transduction Pathways by Ubiquitination in ES Cells
4.3. Regulation of ES Cell-Related Epigenetic Regulators by Ubiquitination
5. Ubiquitination-Mediated Regulation of Cancer Development
6. Concluding Remarks and Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| APC | Adenomatous polyposis coli |
| APC/C | Anaphase-promoting complex/cyclosome |
| BMP | Bone morphogenetic protein |
| CDK | Cyclin-dependent kinase |
| CHIP | Hsc70-interacting protein |
| CK1 | Casein kinase 1 |
| CRC | Colorectal cancer |
| CRLs | Cullin-RING ligases |
| DCAF13 | DDB1- and CUL4-associated factor 13 |
| DDB1 | DNA damage binding protein 1 |
| DTT | DNA damage tolerance |
| E3 | Ub ligase |
| EC cells | embryonic carcinoma cells |
| EG cells | Embryonic germ cells |
| EMT | Epithelial–mesenchymal transition |
| ENK | Arly embryo specific NK |
| ES cells | Embryonic stem cells |
| FBXW8 | F-box and WD40 domain-containing protein 8 |
| FGF2 | Fibroblast growth factor |
| FZD | ubiquitinates Frizzled |
| GSK3 | Glycogen synthase kinase 3 |
| H2B-K120Ub1 | K120 residue of Histone H2B can be monoubiquitylated |
| HACE1 | HECT domain and ankyrin-repeat-containing E3 ubiquitin-protein ligase 1 |
| HECT | homologous to E6-AP COOH terminus |
| hESCs | Human ES cells |
| HIF-1α | Hypoxia inducible factor 1α |
| hUREB | Human upstream regulatory element binding protein |
| ICM | Inner cell mass |
| IDs | Inhibitors of differentiation |
| IKK | Inhibitors of I-κB kinase |
| iPSCs | Induced pluripotent stem cells |
| LIF | Leukemia inhibitory factor |
| LRP5/6 | low-density-lipoprotein-related protein 5/6 |
| MCL1 | Myeloid cell leukemia 1 |
| MDM2 | Mouse double minute 2 |
| mESCs | Mouse ES cells |
| MMR | Mismatch repair |
| MSCs | Mesenchymal stem cells |
| MSI | Microsatellite instability |
| NADH | Nicotinamide adenine dinucleotide |
| OC | Osteocalcin |
| Oct4 | Octamer-binding transcription factor-4 |
| OPN/SPP1 | Osteopontin |
| OS | Osteosarcoma |
| PCNA | Proliferating cell nuclear antigen |
| POUh | POU homeo-domain |
| POUs | POU-specific domain |
| PRC1 | Polycomb repressive complex 1 |
| PTEN | Phosphatase and tensin homolog deleted on chromosome 10 |
| pVHL | von Hippel-Lindau protein |
| RA | Retinoic acid |
| RARα | Retinoic acid receptor alpha |
| RBR | RING between RING |
| RBX1 | RING-box protein 1 |
| RhoGDI2 | Rho guanosine diphosphate dissociation inhibitor 2 |
| RING | Really Interesting New Gene |
| RIP1 | Receptor interacting protein 1 |
| RTK | Receptor tyrosine kinase |
| RNF6 | RING-type E3 ring finger protein 6 |
| RNF43 | Ubiquitin E3 ligase ring finger 43 |
| SATB1 | Special AT-rich sequence-binding protein-1 |
| SCF | Skp1/Cul1/F-box |
| SHN3 | Zinc finger-containing adaptor Schnurri-3 |
| Smurf1 | Smad ubiquitination regulatory factor 11 |
| Sox2 | SRY-related HMG Box 2 |
| SPOP | Speckle-type POZ protein |
| Tcf/Lef | T-cell factor/lymphoid enhancer factors |
| TLS | Translesion DNA synthesis |
| TRAF6 | NF receptor-associated factor 6 |
| Ub | Ubiquitin |
| Uba/E1 | Ub-activating enzyme |
| UBC/E2 | Ub-conjugating enzyme |
| Ube2s | E2 ubiquitin-conjugating Enzyme E2S |
| WSB1 | WD repeat and SOCS box containing 1 |
References
- Hicke, L.; Dunn, R. Regulation of membrane protein transport by ubiquitin and ubiquitin-binding proteins. Annu. Rev. Cell Dev. Biol. 2003, 19, 141–172. [Google Scholar] [CrossRef]
- Vijay-Kumar, S.; Bugg, C.E.; Wilkinson, K.; Vierstra, R.D.; Hatfield, P.; Cook, W.J. Comparison of the three-dimensional structures of human, yeast, and oat ubiquitin. J. Biol. Chem. 1987, 262, 6396–6399. [Google Scholar] [PubMed]
- Callis, J.; Vierstra, R.D. Ubiquitin and ubiquitin genes in higher plants. Oxford Surv. Plant Mol. Cell. Biol. 1989, 6, 1–30. [Google Scholar]
- Hershko, A.; Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 1998, 67, 425–479. [Google Scholar] [CrossRef]
- Peng, J.; Schwartz, D.; Elias, J.E.; Thoreen, C.C.; Cheng, D.; Marsischky, G.; Roelofs, J.; Finley, D.; Gygi, S.P. A proteomics approach to understanding protein ubiquitination. Nat. Biotechnol. 2003, 21, 921–926. [Google Scholar] [CrossRef]
- Kim, W.; Bennett, E.J.; Huttlin, E.L.; Guo, A.; Li, J.; Possemato, A.; Sowa, M.E.; Rad, R.; Rush, J.; Comb, M.J.; et al. Systematic and quantitative assessment of the ubiquitin-modified proteome. Mol. Cell 2011, 44, 325–340. [Google Scholar] [CrossRef]
- Wagner, S.A.; Beli, P.; Weinert, B.T.; Nielsen, M.L.; Cox, J.; Mann, M.; Choudhary, C. A proteome-wide, quantitative survey of in vivo ubiquitylation sites reveals widespread regulatory roles. Mol. Cell Proteomics 2011, 10, M111.013284. [Google Scholar] [CrossRef]
- Fukasawa, H.; Fujigaki, Y.; Yamamoto, T.; Hishida, A.; Kitagawa, M. Protein Degradation by the Ubiquitin-Proteasome Pathway and Organ Fibrosis. Curr. Med. Chem. 2012, 19, 893–900. [Google Scholar] [CrossRef]
- Aviel, S.; Winberg, G.; Massucci, M.; Ciechanover, A. Degradation of the epstein-barr virus latent membrane protein 1 (LMP1) by the ubiquitin-proteasome pathway. Targeting via ubiquitination of the N-terminal residue. J. Biol. Chem. 2000, 275, 23491–23499. [Google Scholar] [CrossRef] [PubMed]
- Pan, G.J.; Pei, D.Q. Identification of two distinct transactivation domains in the pluripotency sustaining factor nanog. Cell Res. 2003, 13, 499–502. [Google Scholar] [CrossRef] [PubMed]
- Ciechanover, A. N-terminal ubiquitination: More protein substrates join in. Trends Cell Biol. 2004, 14, 103–106. [Google Scholar] [CrossRef] [PubMed]
- Breitschopf, K.; Bengal, E.; Ziv, T.; Admon, A.; Ciechanover, A. A novel site for ubiquitination: The N-terminal residue, and not internal lysines of MyoD, is essential for conjugation and degradation of the protein. EMBO J. 1998, 17, 5964–5973. [Google Scholar] [CrossRef] [PubMed]
- Cadwell, K.; Coscoy, L. Ubiquitination on nonlysine residues by a viral E3 ubiquitin ligase. Science 2005, 309, 127–130. [Google Scholar] [CrossRef]
- Bhogaraju, S.; Kalayil, S.; Liu, Y.; Bonn, F.; Colby, T.; Matic, I.; Dikic, I. Phosphoribosylation of Ubiquitin Promotes Serine Ubiquitination and Impairs Conventional Ubiquitination. Cell 2016, 167, 1636–1649. [Google Scholar] [CrossRef]
- Bhogaraju, S.; Dikic, I. Ubiquitination without E1 and E2 enzymes. Nature 2016, 533, 43. [Google Scholar] [CrossRef]
- Ciechanover, A.; Heller, H.; Katz-Etzion, R.; Hershko, A. Activation of the heat-stable polypeptide of the ATP-dependent proteolytic system. Proc. Natl. Acad. Sci. USA 1981, 78, 761–765. [Google Scholar] [CrossRef]
- Hershko, A.; Heller, H.; Elias, S.; Ciechanover, A. Components of ubiquitin-protein ligase system. Resolution, affinity purification, and role in protein breakdown. J. Biol. Chem. 1983, 258, 8206–8214. [Google Scholar]
- Hershko, A. Ubiquitin-mediated protein degradation. J. Biol. Chem. 1988, 263, 15237–15240. [Google Scholar]
- Ye, Y.; Rape, M. Building ubiquitin chains: E2 enzymes at work. Nat. Rev. Mol. Cell Biol. 2009, 10, 755–764. [Google Scholar] [CrossRef]
- Varshavsky, A. The ubiquitin system, an immense realm. Annu. Rev. Biochem. 2012, 81, 167–176. [Google Scholar] [CrossRef]
- Metzger, M.B.; Hristova, V.A.; Weissman, A.M. HECT and RING finger families of E3 ubiquitin ligases at a glance. J. Cell Sci. 2012, 125, 531–537. [Google Scholar] [CrossRef]
- Grossman, S.R.; Deato, M.E.; Brignone, C.; Chan, H.M.; Kung, A.L.; Tagami, H.; Nakatani, Y.; Livingston, D.M. Polyubiquitination of p53 by a ubiquitin ligase activity of p300. Science 2003, 300, 342–344. [Google Scholar] [CrossRef]
- Hoppe, T. Multiubiquitylation by E4 enzymes: ‘one size’ doesn't fit all. Trends Biochem. Sci. 2005, 30, 183–187. [Google Scholar] [CrossRef]
- Baranes-Bachar, K.; Levy-Barda, A.; Oehler, J.; Reid, D.A.; Soria-Bretones, I.; Voss, T.C.; Chung, D.; Park, Y.; Liu, C.; Yoon, J.-B.; et al. The Ubiquitin E3/E4 Ligase UBE4A Adjusts Protein Ubiquitylation and Accumulation at Sites of DNA Damage, Facilitating Double-Strand Break Repair. Mol. Cell 2018, 69, 866–878. [Google Scholar] [CrossRef]
- Liu, C.; Liu, W.; Ye, Y.; Li, W. Ufd2p synthesizes branched ubiquitin chains to promote the degradation of substrates modified with atypical chains. Nat. Commun. 2017, 8, 14274. [Google Scholar] [CrossRef]
- Wu, H.; Pomeroy, S.L.; Ferreira, M.; Teider, N.; Mariani, J.; Nakayama, K.I.; Hatakeyama, S.; Tron, V.A.; Saltibus, L.F.; Spyracopoulos, L.; et al. UBE4B promotes Hdm2-mediated degradation of the tumor suppressor p53. Nat. Med. 2011, 17, 347–355. [Google Scholar] [CrossRef]
- Wu, H.; Leng, R.P. UBE4B, a ubiquitin chain assembly factor, is required for MDM2-mediated p53 polyubiquitination and degradation. Cell Cycle 2011, 10, 1912–1915. [Google Scholar] [CrossRef]
- Ackermann, L.; Schell, M.; Pokrzywa, W.; Kevei, É.; Gartner, A.; Schumacher, B.; Hoppe, T. E4 ligase-specific ubiquitination hubs coordinate DNA double-strand-break repair and apoptosis. Nat. Struct. Mol. Biol. 2016, 23, 995–1002. [Google Scholar] [CrossRef]
- Koegl, M.; Hoppe, T.; Schlenker, S.; Ulrich, H.D.; Mayer, T.U.; Jentsch, S. A novel ubiquitination factor, E4, is involved in multiubiquitin chain assembly. Cell 1999, 96, 635–644. [Google Scholar] [CrossRef]
- Qiu, J.; Sheedlo, M.J.; Yu, K.; Tan, Y.; Nakayasu, E.S.; Das, C.; Liu, X.; Luo, Z.Q. Ubiquitination independent of E1 and E2 enzymes by bacterial effectors. Nature 2016, 533, 120–124. [Google Scholar] [CrossRef]
- Stewart, M.D.; Ritterhoff, T.; Klevit, R.E.; Brzovic, P.S. E2 enzymes: More than just middle men. Cell Res. 2016, 26, 423–440. [Google Scholar] [CrossRef]
- Plafker, S.M.; Plafker, K.S.; Weissman, A.M.; Macara, I.G. Ubiquitin charging of human class III ubiquitin-conjugating enzymes triggers their nuclear import. J. Cell Biol. 2004, 167, 649–659. [Google Scholar] [CrossRef]
- Morreale, F.E.; Walden, H. Types of Ubiquitin Ligases. Cell 2016, 165, 248. [Google Scholar] [CrossRef]
- Deshaies, R.J.; Joazeiro, C.A. RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 2009, 78, 399–434. [Google Scholar] [CrossRef] [PubMed]
- Chang, L.F.; Zhang, Z.; Yang, J.; McLaughlin, S.H.; Barford, D. Molecular architecture and mechanism of the anaphase-promoting complex. Nature 2014, 513, 388–393. [Google Scholar] [CrossRef]
- Min, M.; Mevissen, T.E.T.; De Luca, M.; Komander, D.; Lindon, C. Efficient APC/C substrate degradation in cells undergoing mitotic exit depends on K11 ubiquitin linkages. Mol. Biol. Cell 2015, 26, 4325–4332. [Google Scholar] [CrossRef] [PubMed]
- Kelly, A.; Wickliffe, K.E.; Song, L.; Fedrigo, I.; Rape, M. Ubiquitin Chain Elongation Requires E3-Dependent Tracking of the Emerging Conjugate. Mol. Cell 2014, 56, 232–245. [Google Scholar] [CrossRef]
- Scheffner, M.; Staub, O. HECT E3s and human disease. BMC. Biochem. 2007, 8 (Suppl. 1), S6. [Google Scholar] [CrossRef]
- Rotin, D.; Kumar, S. Physiological functions of the HECT family of ubiquitin ligases. Nat. Rev. Mol. Cell Biol. 2009, 10, 398–409. [Google Scholar] [CrossRef]
- Marin, I.; Lucas, J.I.; Gradilla, A.C.; Ferrus, A. Parkin and relatives: The RBR family of ubiquitin ligases. Physiol. Genomics 2004, 17, 253–263. [Google Scholar] [CrossRef]
- Dove, K.K.; Klevit, R.E. RING-Between-RING E3 Ligases: Emerging Themes amid the Variations. J. Mol. Biol. 2017, 429, 3363–3375. [Google Scholar] [CrossRef]
- Hochstrasser, M. Ubiquitin-dependent protein degradation. Ann. Rev. Genet. 1996, 30, 405–439. [Google Scholar] [CrossRef] [PubMed]
- Xu, P.; Peng, J. Characterization of Polyubiquitin Chain Structure by Middle-down Mass Spectrometry. Anal. Chem. 2008, 80, 3438–3444. [Google Scholar] [CrossRef]
- Swatek, K.N.; Komander, D. Ubiquitin modifications. Cell Res. 2016, 26, 399–422. [Google Scholar] [CrossRef] [PubMed]
- Yau, R.; Rape, M. The increasing complexity of the ubiquitin code. Nat. Cell Biol. 2016, 18, 579–586. [Google Scholar] [CrossRef] [PubMed]
- Xu, P.; Duong, D.M.; Seyfried, N.T.; Cheng, D.; Xie, Y.; Robert, J.; Rush, J.; Hochstrasser, M.; Finley, D.; Peng, J. Quantitative proteomics reveals the function of unconventional ubiquitin chains in proteasomal degradation. Cell 2009, 137, 133–145. [Google Scholar] [CrossRef] [PubMed]
- Pickart, C.M.; Fushman, D. Polyubiquitin chains: Polymeric protein signals. Curr. Opin. Chem. Biol. 2004, 8, 610–616. [Google Scholar] [CrossRef] [PubMed]
- Livneh, I.; Kravtsova-Ivantsiv, Y.; Braten, O.; Kwon, Y.T.; Ciechanover, A. Monoubiquitination joins polyubiquitination as an esteemed proteasomal targeting signal. Bioessays 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Ea, C.K.; Deng, L.; Xia, Z.P.; Pineda, G.; Chen, Z.J. Activation of IKK by TNFalpha requires site-specific ubiquitination of RIP1 and polyubiquitin binding by NEMO. Mol. Cell 2006, 22, 245–257. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Xin, H.; Xu, X.; Huang, M.; Zhang, X.; Chen, Y.; Zhang, S.; Fu, X.Y.; Chang, Z. CHIP mediates degradation of Smad proteins and potentially regulates Smad-induced transcription. Mol. Cell Biol. 2004, 24, 856–864. [Google Scholar] [CrossRef]
- Kanayama, A.; Seth, R.B.; Sun, L.; Ea, C.-K.; Hong, M.; Shaito, A.; Chiu, Y.-H.; Deng, L.; Chen, Z.J. TAB2 and TAB3 Activate the NF-κB Pathway through Binding to Polyubiquitin Chains. Mol. Cell 2004, 15, 535–548. [Google Scholar] [CrossRef]
- Skaug, B.; Jiang, X.; Chen, Z.J. The role of ubiquitin in NF-kappaB regulatory pathways. Annu. Rev. Biochem. 2009, 78, 769–796. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Ulrich, H.D. Distinct consequences of posttranslational modification by linear versus K63-linked polyubiquitin chains. Proc. Natl. Acad. Sci. USA 2010, 107, 7704–7709. [Google Scholar] [CrossRef]
- Liao, B.; Jin, Y. Wwp2 mediates Oct4 ubiquitination and its own auto-ubiquitination in a dosage-dependent manner. Cell Res. 2010, 20, 332–344. [Google Scholar] [CrossRef] [PubMed]
- Garnett, M.J.; Mansfeld, J.; Godwin, C.; Matsusaka, T.; Wu, J.; Russell, P.; Pines, J.; Venkitaraman, A.R. UBE2S elongates ubiquitin chains on APC/C substrates to promote mitotic exit. Nat. Cell Biol. 2009, 11, 1363–1369. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Merbl, Y.; Huo, Y.; Gallop, J.L.; Tzur, A.; Kirschner, M.W. UBE2S drives elongation of K11-linked ubiquitin chains by the Anaphase-Promoting Complex. Proc. Natl. Acad. Sci. USA 2010, 107, 1355–1360. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhang, Y.; Hou, J.; Qian, X.; Zhang, H.; Zhang, Z.; Li, M.; Wang, R.; Liao, K.; Wang, Y.; et al. Ube2s regulates Sox2 stability and mouse ES cell maintenance. Cell Death Differ. 2016, 23, 393–404. [Google Scholar] [CrossRef]
- Li, Z.; Wang, Y.; Li, Y.; Yin, W.; Mo, L.; Qian, X.; Zhang, Y.; Wang, G.; Bu, F.; Zhang, Z.; et al. Ube2s stabilizes β-Catenin through K11-linked polyubiquitination to promote mesendoderm specification and colorectal cancer development. Cell Death Dis. 2018, 9, 456. [Google Scholar] [CrossRef]
- Zhang, W.; Qin, Z.; Zhang, X.; Xiao, W. Roles of sequential ubiquitination of PCNA in DNA-damage tolerance. FEBS Lett. 2011, 585, 2786–2794. [Google Scholar] [CrossRef]
- Hedglin, M.; Benkovic, S.J. Regulation of Rad6/Rad18 Activity During DNA Damage Tolerance. Annu. Rev. Biophys. 2015, 44, 207–228. [Google Scholar] [CrossRef]
- Leung, W.; Baxley, R.; Moldovan, G.-L.; Bielinsky, A. Mechanisms of DNA Damage Tolerance: Post-Translational Regulation of PCNA. Genes 2018, 10, 10. [Google Scholar] [CrossRef] [PubMed]
- Weissman, I.L. Stem cells: Units of development, units of regeneration, and units in evolution. Cell 2000, 100, 157–168. [Google Scholar] [CrossRef]
- Evans, M.J.; Kaufman, M.H. Establishment in culture of pluripotential cells from mouse embryos. Nature 1981, 292, 154–156. [Google Scholar] [CrossRef]
- Levine, M.; Tjian, R. Transcription regulation and animal diversity. Nature 2003, 424, 147–151. [Google Scholar] [CrossRef] [PubMed]
- Boyer, L.A.; Lee, T.I.; Cole, M.F.; Johnstone, S.E.; Levine, S.S.; Zucker, J.P.; Guenther, M.G.; Kumar, R.M.; Murray, H.L.; Jenner, R.G.; et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 2005, 122, 947–956. [Google Scholar] [CrossRef]
- Young, R.A. Control of the embryonic stem cell state. Cell 2011, 144, 940–954. [Google Scholar] [CrossRef]
- Silva, J.; Smith, A. Capturing pluripotency. Cell 2008, 132, 532–536. [Google Scholar] [CrossRef]
- Brehm, A.; Ohbo, K.; Scholer, H. The carboxy-terminal transactivation domain of Oct-4 acquires cell specificity through the POU domain. Mol. Cell Biol. 1997, 17, 154–162. [Google Scholar] [CrossRef]
- Zeineddine, D.; Hammoud, A.A.; Mortada, M.; Boeuf, H. The Oct4 protein: More than a magic stemness marker. Am. J. Stem Cells 2014, 3, 74–82. [Google Scholar]
- Niwa, H.; Miyazaki, J.; Smith, A.G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat. Genet. 2000, 24, 372–376. [Google Scholar] [CrossRef]
- Xu, H.; Wang, W.; Li, C.; Yu, H.; Yang, A.; Wang, B.; Jin, Y. WWP2 promotes degradation of transcription factor OCT4 in human embryonic stem cells. Cell Res. 2009, 19, 796. [Google Scholar] [CrossRef]
- Pirozzi, G.; McConnell, S.J.; Uveges, A.J.; Carter, J.M.; Sparks, A.B.; Kay, B.K.; Fowlkes, D.M. Identification of Novel Human WW Domain-containing Proteins by Cloning of Ligand Targets. J. Biol. Chem. 1997, 272, 14611–14616. [Google Scholar] [CrossRef]
- Xu, H.M.; Liao, B.; Zhang, Q.J.; Wang, B.B.; Li, H.; Zhong, X.M.; Sheng, H.Z.; Zhao, Y.X.; Zhao, Y.M.; Jin, Y. Wwp2, an E3 Ubiquitin Ligase That Targets Transcription Factor Oct-4 for Ubiquitination. J. Biol. Chem. 2004, 279, 23495–23503. [Google Scholar] [CrossRef]
- Li, S.; Xiao, F.; Zhang, J.; Sun, X.; Wang, H.; Zeng, Y.; Hu, J.; Tang, F.; Gu, J.; Zhao, Y.; et al. Disruption of OCT4 Ubiquitination Increases OCT4 Protein Stability and ASH2L-B-Mediated H3K4 Methylation Promoting Pluripotency Acquisition. Stem Cell Rep. 2018, 11, 973–987. [Google Scholar] [CrossRef] [PubMed]
- Liao, B.; Zhong, X.; Xu, H.; Xiao, F.; Fang, Z.; Gu, J.; Chen, Y.; Zhao, Y.; Jin, Y. Itch, an E3 ligase of Oct4, is required for embryonic stem cell self-renewal and pluripotency induction. J. Cell Physiol. 2013, 228, 1443–1451. [Google Scholar] [CrossRef] [PubMed]
- Bahnassawy, L.; Perumal, T.M.; Gonzalez-Cano, L.; Hillje, A.L.; Taher, L.; Makalowski, W.; Suzuki, Y.; Fuellen, G.; del Sol, A.; Schwamborn, J.C. TRIM32 modulates pluripotency entry and exit by directly regulating Oct4 stability. Sci. Rep. 2015, 5, 13456. [Google Scholar] [CrossRef] [PubMed]
- Novo, C.L.; Tang, C.; Ahmed, K.; Djuric, U.; Fussner, E.; Mullin, N.P.; Morgan, N.P.; Hayre, J.; Sienerth, A.R.; Elderkin, S.; et al. The pluripotency factor Nanog regulates pericentromeric heterochromatin organization in mouse embryonic stem cells. Genes Dev. 2016, 30, 1101–1115. [Google Scholar] [CrossRef] [PubMed]
- Chew, J.L.; Loh, Y.H.; Zhang, W.; Chen, X.; Tam, W.L.; Yeap, L.S.; Li, P.; Ang, Y.S.; Lim, B.; Robson, P.; et al. Reciprocal transcriptional regulation of Pou5f1 and Sox2 via the Oct4/Sox2 complex in embryonic stem cells. Mol. Cell Biol. 2005, 25, 6031–6046. [Google Scholar] [CrossRef]
- Kopp, J.L.; Ormsbee, B.D.; Desler, M.; Rizzino, A. Small increases in the level of Sox2 trigger the differentiation of mouse embryonic stem cells. Stem Cells 2008, 26, 903–911. [Google Scholar] [CrossRef]
- Wang, Z.; Oron, E.; Nelson, B.; Razis, S.; Ivanova, N. Distinct lineage specification roles for NANOG, OCT4, and SOX2 in human embryonic stem cells. Cell Stem Cell 2012, 10, 440–454. [Google Scholar] [CrossRef]
- Chen, X.; Xu, H.; Yuan, P.; Fang, F.; Huss, M.; Vega, V.B.; Wong, E.; Orlov, Y.L.; Zhang, W.; Jiang, J.; et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 2008, 133, 1106–1117. [Google Scholar] [CrossRef] [PubMed]
- Fang, L.; Zhang, L.; Wei, W.; Jin, X.; Wang, P.; Tong, Y.; Li, J.; Du, J.X.; Wong, J. A methylation-phosphorylation switch determines Sox2 stability and function in ESC maintenance or differentiation. Mol. Cell 2014, 55, 537–551. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.H.; Tsai, M.S.; Chiang, M.F.; Li, H. A novel NK-type homeobox gene, ENK (early embryo specific NK), preferentially expressed in embryonic stem cells. Gene Expr. Patterns 2003, 3, 99–103. [Google Scholar] [CrossRef]
- Chambers, I.; Colby, D.; Robertson, M.; Nichols, J.; Lee, S.; Tweedie, S.; Smith, A. Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell 2003, 113, 643–655. [Google Scholar] [CrossRef]
- Mishina, Y. Function of bone morphogenetic protein signaling during mouse development. Front Biosci. 2003, 8, 855–869. [Google Scholar] [CrossRef]
- Chambers, I.; Silva, J.; Colby, D.; Nichols, J.; Nijmeijer, B.; Robertson, M.; Vrana, J.; Jones, K.; Grotewold, L.; Smith, A. Nanog safeguards pluripotency and mediates germline development. Nature 2007, 450, 1230–1234. [Google Scholar] [CrossRef] [PubMed]
- Heurtier, V.; Owens, N.; Gonzalez, I.; Mueller, F.; Proux, C.; Mornico, D.; Clerc, P.; Dubois, A.; Navarro, P. The molecular logic of Nanog-induced self-renewal in mouse embryonic stem cells. Nat.Commun. 2019, 10, 1109. [Google Scholar] [CrossRef]
- Mitsui, K.; Tokuzawa, Y.; Itoh, H.; Segawa, K.; Murakami, M.; Takahashi, K.; Maruyama, M.; Maeda, M.; Yamanaka, S. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 2003, 113, 631–642. [Google Scholar] [CrossRef]
- Pei, D. Deubiquitylating Nanog: Novel role of USP21 in embryonic stem cell maintenance. Signal Transduct. Target Ther. 2017, 2, 17014. [Google Scholar] [CrossRef]
- Wang, J.; Qiao, M.; He, Q.; Shi, R.; Loh, S.J.; Stanton, L.W.; Wu, M. Pluripotency Activity of Nanog Requires Biochemical Stabilization by Variant Histone Protein H2A.Z. Stem Cells 2015, 33, 2126–2134. [Google Scholar] [CrossRef]
- Kwon, S.K.; Lee, D.H.; Kim, S.Y.; Park, J.H.; Choi, J.; Baek, K.H. Ubiquitin-specific protease 21 regulating the K48-linked polyubiquitination of NANOG. Biochem. Biophys. Res. Commun. 2017, 482, 1443–1448. [Google Scholar] [CrossRef]
- Jin, J.; Liu, J.; Chen, C.; Liu, Z.; Jiang, C.; Chu, H.; Pan, W.; Wang, X.; Zhang, L.; Li, B.; et al. The deubiquitinase USP21 maintains the stemness of mouse embryonic stem cells via stabilization of Nanog. Nat. Commun. 2016, 7, 13594. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yao, Y.; Ding, H.; Han, C.; Chen, Y.; Zhang, Y.; Wang, C.; Zhang, X.; Zhang, Y.; Zhai, Y.; et al. USP21 deubiquitylates Nanog to regulate protein stability and stem cell pluripotency. Signal Transduct. Target Ther. 2017, 2, 16046. [Google Scholar] [CrossRef]
- Pera, M.F.; Tam, P.P.L. Extrinsic regulation of pluripotent stem cells. Nature 2010, 465, 713. [Google Scholar] [CrossRef] [PubMed]
- Nichols, J.; Smith, A. The origin and identity of embryonic stem cells. Development 2011, 138, 3–8. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Thomson, J.A. Pluripotent stem cell lines. Genes Dev. 2008, 22, 1987–1997. [Google Scholar] [CrossRef]
- Niwa, H.; Burdon, T.; Chambers, I.; Smith, A. Self-renewal of pluripotent embryonic stem cells is mediated via activation of STAT3. Genes Dev. 1998, 12, 2048–2060. [Google Scholar] [CrossRef]
- Hirai, H.; Karian, P.; Kikyo, N. Regulation of embryonic stem cell self-renewal and pluripotency by leukaemia inhibitory factor. Biochem. J. 2011, 438, 11–23. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, T.; Nakamura, T.; Nakao, K.; Arai, T.; Katsuki, M.; Heike, T.; Yokota, T. STAT3 activation is sufficient to maintain an undifferentiated state of mouse embryonic stem cells. EMBO J. 1999, 18, 4261–4269. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, K.; Chambers, I.; Nichols, J.; Smith, A.; Saito, M.; Yasukawa, K.; Shoyab, M.; Taga, T.; Kishimoto, T. Maintenance of the pluripotential phenotype of embryonic stem cells through direct activation of gp130 signalling pathways. Mech. Dev. 1994, 45, 163–171. [Google Scholar] [CrossRef]
- Vallier, L.; Alexander, M.; Pedersen, R.A. Activin/Nodal and FGF pathways cooperate to maintain pluripotency of human embryonic stem cells. J. Cell Sci. 2005, 118, 4495–4509. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Rosler, E.; Jiang, J.; Lebkowski, J.S.; Gold, J.D.; O'Sullivan, C.; Delavan-Boorsma, K.; Mok, M.; Bronstein, A.; Carpenter, M.K. Basic fibroblast growth factor supports undifferentiated human embryonic stem cell growth without conditioned medium. Stem Cells 2005, 23, 315–323. [Google Scholar] [CrossRef]
- Massague, J. TGF-beta signal transduction. Annu. Rev. Biochem. 1998, 67, 753–791. [Google Scholar] [CrossRef] [PubMed]
- Derynck, R.; Zhang, Y.E. Smad-dependent and Smad-independent pathways in TGF-beta family signalling. Nature 2003, 425, 577–584. [Google Scholar] [CrossRef]
- Weissman, A.M. Themes and variations on ubiquitylation. Nat. Rev. Mol. Cell Biol. 2001, 2, 169–178. [Google Scholar] [CrossRef] [PubMed]
- Miyazono, K.; ten Dijke, P.; Heldin, C.H. TGF-beta signaling by Smad proteins. Adv. Immunol. 2000, 75, 115–157. [Google Scholar] [PubMed]
- Moustakas, A.; Souchelnytskyi, S.; Heldin, C.H. Smad regulation in TGF-beta signal transduction. J. Cell Sci. 2001, 114, 4359–4369. [Google Scholar] [PubMed]
- Hata, A.; Lagna, G.; Massagué, J.; Hemmati-Brivanlou, A. Smad6 inhibits BMP/Smad1 signaling by specifically competing with the Smad4 tumor suppressor. Genes Dev. 1998, 12, 186–197. [Google Scholar] [CrossRef] [PubMed]
- Hanyu, A.; Ishidou, Y.; Ebisawa, T.; Shimanuki, T.; Imamura, T.; Miyazono, K. The N domain of Smad7 is essential for specific inhibition of transforming growth factor-beta signaling. J. Cell Biol. 2001, 155, 1017–1027. [Google Scholar] [CrossRef]
- Ying, Q.L.; Nichols, J.; Chambers, I.; Smith, A. BMP induction of Id proteins suppresses differentiation and sustains embryonic stem cell self-renewal in collaboration with STAT3. Cell 2003, 115, 281–292. [Google Scholar] [CrossRef]
- Lee, T.H.; Shank, J.; Cusson, N.; Kelliher, M.A. The kinase activity of Rip1 is not required for tumor necrosis factor-alpha-induced IkappaB kinase or p38 MAP kinase activation or for the ubiquitination of Rip1 by Traf2. J. Biol. Chem. 2004, 279, 33185–33191. [Google Scholar] [CrossRef]
- Zhu, H.; Kavsak, P.; Abdollah, S.; Wrana, J.L.; Thomsen, G.H. A SMAD ubiquitin ligase targets the BMP pathway and affects embryonic pattern formation. Nature 1999, 400, 687–693. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Qiao, M.; Harris, S.E.; Oyajobi, B.O.; Mundy, G.R.; Chen, D. Smurf1 inhibits osteoblast differentiation and bone formation in vitro and in vivo. J. Biol. Chem. 2004, 279, 12854–12859. [Google Scholar] [CrossRef]
- Zhang, Y.; Chang, C.; Gehling, D.J.; Hemmati-Brivanlou, A.; Derynck, R. Regulation of Smad degradation and activity by Smurf2, an E3 ubiquitin ligase. Proc. Natl. Acad. Sci. USA 2001, 98, 974–979. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Liu, Y.T.; Hao, R.; Chen, L.; Chang, Z.; Wang, H.R.; Wang, Z.X.; Wu, J.W. Molecular mechanism of the negative regulation of Smad1/5 protein by carboxyl terminus of Hsc70-interacting protein (CHIP). J. Biol. Chem. 2011, 286, 15883–15894. [Google Scholar] [CrossRef]
- Moren, A.; Imamura, T.; Miyazono, K.; Heldin, C.H.; Moustakas, A. Degradation of the tumor suppressor Smad4 by WW and HECT domain ubiquitin ligases. J. Biol. Chem. 2005, 280, 22115–22123. [Google Scholar] [CrossRef]
- Zhang, L.; Huang, H.; Zhou, F.; Schimmel, J.; Pardo, C.G.; Zhang, T.; Barakat, T.S.; Sheppard, K.A.; Mickanin, C.; Porter, J.A.; et al. RNF12 Controls Embryonic Stem Cell Fate and Morphogenesis in Zebrafish Embryos by Targeting Smad7 for Degradation. Mol. Cell 2012, 46, 650–661. [Google Scholar]
- Kim, S.-E.; Huang, H.; Zhao, M.; Zhang, X.; Zhang, A.; Semonov, M.V.; MacDonald, B.T.; Zhang, X.; Abreu, J.G.; Peng, L.; et al. Wnt Stabilization of β-Catenin Reveals Principles for Morphogen Receptor-Scaffold Assemblies. Science 2013, 340, 867. [Google Scholar] [CrossRef] [PubMed]
- Moon, R.T.; Bowerman, B.; Boutros, M.; Perrimon, N. The Promise and Perils of Wnt Signaling Through β-Catenin. Science 2002, 296, 1644. [Google Scholar] [CrossRef]
- van Es, J.H.; Barker, N.; Clevers, H. You Wnt some, you lose some: Oncogenes in the Wnt signaling pathway. Curr. Opin. Genet. Dev. 2003, 13, 28–33. [Google Scholar] [CrossRef]
- Sato, N.; Meijer, L.; Skaltsounis, L.; Greengard, P.; Brivanlou, A.H. Maintenance of pluripotency in human and mouse embryonic stem cells through activation of Wnt signaling by a pharmacological GSK-3-specific inhibitor. Nat. Med. 2004, 10, 55–63. [Google Scholar] [CrossRef]
- Kielman, M.F.; Rindapaa, M.; Gaspar, C.; van Poppel, N.; Breukel, C.; van Leeuwen, S.; Taketo, M.M.; Roberts, S.; Smits, R.; Fodde, R. Apc modulates embryonic stem-cell differentiation by controlling the dosage of beta-catenin signaling. Nat. Genet. 2002, 32, 594–605. [Google Scholar] [CrossRef] [PubMed]
- Ying, Q.L.; Wray, J.; Nichols, J.; Batlle-Morera, L.; Doble, B.; Woodgett, J.; Cohen, P.; Smith, A. The ground state of embryonic stem cell self-renewal. Nature 2008, 453, 519–523. [Google Scholar] [CrossRef] [PubMed]
- Blauwkamp, T.A.; Nigam, S.; Ardehali, R.; Weissman, I.L.; Nusse, R. Endogenous Wnt signalling in human embryonic stem cells generates an equilibrium of distinct lineage-specified progenitors. Nat. Commun. 2012, 3, 1070. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Li, Y.; Semenov, M.; Han, C.; Baeg, G.H.; Tan, Y.; Zhang, Z.; Lin, X.; He, X. Control of beta-catenin phosphorylation/degradation by a dual-kinase mechanism. Cell 2002, 108, 837–847. [Google Scholar] [CrossRef]
- He, X.; Semenov, M.; Tamai, K.; Zeng, X. LDL receptor-related proteins 5 and 6 in Wnt/beta-catenin signaling: Arrows point the way. Development 2004, 131, 1663–1677. [Google Scholar] [CrossRef] [PubMed]
- Pez, F.; Lopez, A.; Kim, M.; Wands, J.R.; Caron de Fromentel, C.; Merle, P. Wnt signaling and hepatocarcinogenesis: Molecular targets for the development of innovative anticancer drugs. J. Hepatol. 2013, 59, 1107–1117. [Google Scholar] [CrossRef]
- Aberle, H.; Bauer, A.; Stappert, J.; Kispert, A.; Kemler, R. β-catenin is a target for the ubiquitin–proteasome pathway. EMBO J. 1997, 16, 3797–3804. [Google Scholar] [CrossRef]
- Ha, N.C.; Tonozuka, T.; Stamos, J.L.; Choi, H.J.; Weis, W.I. Mechanism of phosphorylation-dependent binding of APC to beta-catenin and its role in beta-catenin degradation. Mol. Cell 2004, 15, 511–521. [Google Scholar] [CrossRef]
- Su, Y.; Fu, C.; Ishikawa, S.; Stella, A.; Kojima, M.; Shitoh, K.; Schreiber, E.M.; Day, B.W.; Liu, B. APC is essential for targeting phosphorylated beta-catenin to the SCFbeta-TrCP ubiquitin ligase. Mol. Cell 2008, 32, 652–661. [Google Scholar] [CrossRef]
- Wu, G.; Xu, G.; Schulman, B.A.; Jeffrey, P.D.; Harper, J.W.; Pavletich, N.P. Structure of a β-TrCP1-Skp1-β-Catenin Complex: Destruction Motif Binding and Lysine Specificity of the SCFβ-TrCP1 Ubiquitin Ligase. Mol. Cell 2003, 11, 1445–1456. [Google Scholar] [CrossRef]
- Ji, L.; Jiang, B.; Jiang, X.; Charlat, O.; Chen, A.; Mickanin, C.; Bauer, A.; Xu, W.; Yan, X.; Cong, F. The SIAH E3 ubiquitin ligases promote Wnt/β-catenin signaling through mediating Wnt-induced Axin degradation. Genes Dev. 2017, 31, 904–915. [Google Scholar] [CrossRef] [PubMed]
- Meshorer, E.; Misteli, T. Chromatin in pluripotent embryonic stem cells and differentiation. Nat. Rev. Mol. Cell Biol. 2006, 7, 540–546. [Google Scholar] [CrossRef] [PubMed]
- Azuara, V.; Perry, P.; Sauer, S.; Spivakov, M.; Jorgensen, H.F.; John, R.M.; Gouti, M.; Casanova, M.; Warnes, G.; Merkenschlager, M.; et al. Chromatin signatures of pluripotent cell lines. Nat. Cell Biol. 2006, 8, 532–538. [Google Scholar] [CrossRef] [PubMed]
- Campos, E.I.; Reinberg, D. Histones: Annotating chromatin. Annu. Rev. Genet. 2009, 43, 559–599. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, B.E.; Mikkelsen, T.S.; Xie, X.; Kamal, M.; Huebert, D.J.; Cuff, J.; Fry, B.; Meissner, A.; Wernig, M.; Plath, K.; et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 2006, 125, 315–326. [Google Scholar] [CrossRef] [PubMed]
- Thorne, A.W.; Sautiere, P.; Briand, G.; Crane-Robinson, C. The structure of ubiquitinated histone H2B. EMBO J. 1987, 6, 1005–1010. [Google Scholar] [CrossRef]
- Minsky, N.; Shema, E.; Field, Y.; Schuster, M.; Segal, E.; Oren, M. Monoubiquitinated H2B is associated with the transcribed region of highly expressed genes in human cells. Nat. Cell Biol. 2008, 10, 483–488. [Google Scholar] [CrossRef]
- Kim, J.; Hake, S.B.; Roeder, R.G. The human homolog of yeast BRE1 functions as a transcriptional coactivator through direct activator interactions. Mol. Cell 2005, 20, 759–770. [Google Scholar] [CrossRef]
- Fuchs, G.; Shema, E.; Vesterman, R.; Kotler, E.; Wolchinsky, Z.; Wilder, S.; Golomb, L.; Pribluda, A.; Zhang, F.; Haj-Yahya, M.; et al. RNF20 and USP44 regulate stem cell differentiation by modulating H2B monoubiquitylation. Mol. Cell 2012, 46, 662–673. [Google Scholar] [CrossRef]
- de Napoles, M.; Mermoud, J.E.; Wakao, R.; Tang, Y.A.; Endoh, M.; Appanah, R.; Nesterova, T.B.; Silva, J.; Otte, A.P.; Vidal, M.; et al. Polycomb group proteins Ring1A/B link ubiquitylation of histone H2A to heritable gene silencing and X inactivation. Dev. Cell 2004, 7, 663–676. [Google Scholar] [CrossRef]
- Wang, H.; Wang, L.; Erdjument-Bromage, H.; Vidal, M.; Tempst, P.; Jones, R.S.; Zhang, Y. Role of histone H2A ubiquitination in Polycomb silencing. Nature 2004, 431, 873–878. [Google Scholar] [CrossRef]
- Stock, J.K.; Giadrossi, S.; Casanova, M.; Brookes, E.; Vidal, M.; Koseki, H.; Brockdorff, N.; Fisher, A.G.; Pombo, A. Ring1-mediated ubiquitination of H2A restrains poised RNA polymerase II at bivalent genes in mouse ES cells. Nat. Cell Biol. 2007, 9, 1428–1435. [Google Scholar] [CrossRef] [PubMed]
- van der Stoop, P.; Boutsma, E.A.; Hulsman, D.; Noback, S.; Heimerikx, M.; Kerkhoven, R.M.; Voncken, J.W.; Wessels, L.F.; van Lohuizen, M. Ubiquitin E3 ligase Ring1b/Rnf2 of polycomb repressive complex 1 contributes to stable maintenance of mouse embryonic stem cells. PloS ONE 2008, 3, 2235. [Google Scholar] [CrossRef]
- Endoh, M.; Endo, T.A.; Endoh, T.; Fujimura, Y.; Ohara, O.; Toyoda, T.; Otte, A.P.; Okano, M.; Brockdorff, N.; Vidal, M.; et al. Polycomb group proteins Ring1A/B are functionally linked to the core transcriptional regulatory circuitry to maintain ES cell identity. Development 2008, 135, 1513–1524. [Google Scholar] [CrossRef] [PubMed]
- Inoue, D.; Aihara, H.; Sato, T.; Mizusaki, H.; Doiguchi, M.; Higashi, M.; Imamura, Y.; Yoneda, M.; Miyanishi, T.; Fujii, S.; et al. Dzip3 regulates developmental genes in mouse embryonic stem cells by reorganizing 3D chromatin conformation. Sci. Rep. 2015, 5, 16567. [Google Scholar] [CrossRef] [PubMed]
- Singh, M.P.; Rai, S.; Suyal, S.; Singh, S.K.; Singh, N.K.; Agarwal, A.; Srivastava, S. Genetic and epigenetic markers in colorectal cancer screening: Recent advances. Expert Rev. Mol. Diagn. 2017, 17, 665–685. [Google Scholar] [CrossRef] [PubMed]
- Dagogo-Jack, I.; Shaw, A.T. Tumour heterogeneity and resistance to cancer therapies. Nat. Rev. Clin. Oncol. 2018, 15, 81–94. [Google Scholar] [CrossRef]
- Kinzler, K.W.; Vogelstein, B. Lessons from hereditary colorectal cancer. Cell 1996, 87, 159–170. [Google Scholar] [CrossRef]
- Markowitz, S.D.; Bertagnolli, M.M. Molecular origins of cancer: Molecular basis of colorectal cancer. N. Engl. J. Med. 2009, 361, 2449–2460. [Google Scholar] [CrossRef]
- Toyota, M.; Ahuja, N.; Ohe-Toyota, M.; Herman, J.G.; Baylin, S.B.; Issa, J.P. CpG island methylator phenotype in colorectal cancer. Proc. Natl. Acad. Sci. USA 1999, 96, 8681–8686. [Google Scholar] [CrossRef]
- Lynch, H.T.; Lynch, J.F. Genetics of colonic cancer. Digestion 1998, 59, 481–492. [Google Scholar] [CrossRef]
- Kuwai, T.; Kitadai, Y.; Tanaka, S.; Hiyama, T.; Tanimoto, K.; Chayama, K. Mutation of the von Hippel-Lindau (VHL) gene in human colorectal carcinoma: Association with cytoplasmic accumulation of hypoxia-inducible factor (HIF)-1alpha. Cancer Sci. 2004, 95, 149–153. [Google Scholar] [CrossRef] [PubMed]
- Lisztwan, J.; Imbert, G.; Wirbelauer, C.; Gstaiger, M.; Krek, W. The von Hippel-Lindau tumor suppressor protein is a component of an E3 ubiquitin-protein ligase activity. Genes Dev. 1999, 13, 1822–1833. [Google Scholar] [CrossRef] [PubMed]
- Iwai, K.; Yamanaka, K.; Kamura, T.; Minato, N.; Conaway, R.C.; Conaway, J.W.; Klausner, R.D.; Pause, A. Identification of the von Hippel-lindau tumor-suppressor protein as part of an active E3 ubiquitin ligase complex. Proc. Natl. Acad. Sci. USA 1999, 96, 12436–12441. [Google Scholar] [CrossRef]
- Ohh, M.; Park, C.W.; Ivan, M.; Hoffman, M.A.; Kim, T.Y.; Huang, L.E.; Pavletich, N.; Chau, V.; Kaelin, W.G. Ubiquitination of hypoxia-inducible factor requires direct binding to the beta-domain of the von Hippel-Lindau protein. Nat. Cell Biol. 2000, 2, 423–427. [Google Scholar] [CrossRef] [PubMed]
- Maxwell, P.H.; Wiesener, M.S.; Chang, G.W.; Clifford, S.C.; Vaux, E.C.; Cockman, M.E.; Wykoff, C.C.; Pugh, C.W.; Maher, E.R.; Ratcliffe, P.J. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 1999, 399, 271–2753. [Google Scholar] [CrossRef]
- Semenza, G.L. Regulation of Mammalian O2 Homeostasis by Hypoxia-Inducible Factor 1. Annu. Rev. Cell Dev. Biol. 1999, 15, 551–578. [Google Scholar] [CrossRef]
- Miyoshi, Y.; Nagase, H.; Ando, H.; Horii, A.; Ichii, S.; Nakatsuru, S.; Aoki, T.; Miki, Y.; Mori, T.; Nakamura, Y. Somatic mutations of the APC gene in colorectal tumors: Mutation cluster region in the APC gene. Hum. Mol. Genet. 1992, 1, 229–233. [Google Scholar]
- Morin, P.J.; Sparks, A.B.; Korinek, V.; Barker, N.; Clevers, H.; Vogelstein, B.; Kinzler, K.W. Activation of beta-catenin-Tcf signaling in colon cancer by mutations in beta-catenin or APC. Science 1997, 275, 1787–1790. [Google Scholar] [CrossRef]
- Wu, H.; Lu, X.X.; Wang, J.R.; Yang, T.Y.; Li, X.M.; He, X.S.; Li, Y.; Ye, W.L.; Wu, Y.; Gan, W.J.; et al. TRAF6 inhibits colorectal cancer metastasis through regulating selective autophagic CTNNB1/beta-catenin degradation and is targeted for GSK3B/GSK3beta-mediated phosphorylation and degradation. Autophagy 2019, 26, 1554–8627. [Google Scholar]
- Liu, L.; Zhang, Y.; Wong, C.C.; Zhang, J.; Dong, Y.; Li, X.; Kang, W.; Chan, F.K.L.; Sung, J.J.Y.; Yu, J. RNF6 Promotes Colorectal Cancer by Activating the Wnt/beta-Catenin Pathway via Ubiquitination of TLE3. Cancer Res. 2018, 78, 1958–1971. [Google Scholar] [CrossRef] [PubMed]
- Daniels, D.L.; Weis, W.I. Beta-catenin directly displaces Groucho/TLE repressors from Tcf/Lef in Wnt-mediated transcription activation. Nat. Struct. Mol. Biol. 2005, 12, 364–371. [Google Scholar] [CrossRef]
- Barker, N.; van Es, J.H.; Kuipers, J.; Kujala, P.; van den Born, M.; Cozijnsen, M.; Haegebarth, A.; Korving, J.; Begthel, H.; Peters, P.J.; et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 2007, 449, 1003. [Google Scholar] [CrossRef] [PubMed]
- Barker, N.; van de Wetering, M.; Clevers, H. The intestinal stem cell. Genes Dev. 2008, 22, 1856–1864. [Google Scholar] [CrossRef]
- Datta, K.; Suman, S.; Kumar, S.; Fornace, A.J., Jr. Colorectal Carcinogenesis, Radiation Quality, and the Ubiquitin-Proteasome Pathway. J. Cancer 2016, 7, 174–183. [Google Scholar] [CrossRef] [PubMed]
- Sancho, R.; Jandke, A.; Davis, H.; Diefenbacher, M.E.; Tomlinson, I.; Behrens, A. F-box and WD repeat domain-containing 7 regulates intestinal cell lineage commitment and is a haploinsufficient tumor suppressor. Gastroenterology 2010, 139, 929–941. [Google Scholar] [CrossRef] [PubMed]
- Babaei-Jadidi, R.; Li, N.; Saadeddin, A.; Spencer-Dene, B.; Jandke, A.; Muhammad, B.; Ibrahim, E.E.; Muraleedharan, R.; Abuzinadah, M.; Davis, H.; et al. FBXW7 influences murine intestinal homeostasis and cancer, targeting Notch, Jun, and DEK for degradation. J. Exp. Med. 2011, 208, 295–312. [Google Scholar] [CrossRef]
- Li, N.; Babaei-Jadidi, R.; Lorenzi, F.; Spencer-Dene, B.; Clarke, P.; Domingo, E.; Tulchinsky, E.; Vries, R.G.J.; Kerr, D.; Pan, Y.; et al. An FBXW7-ZEB2 axis links EMT and tumour microenvironment to promote colorectal cancer stem cells and chemoresistance. Oncogenesis 2019, 8, 13. [Google Scholar] [CrossRef]
- Kemp, Z.; Rowan, A.; Chambers, W.; Wortham, N.; Halford, S.; Sieber, O.; Mortensen, N.; von Herbay, A.; Gunther, T.; Ilyas, M.; et al. CDC4 mutations occur in a subset of colorectal cancers but are not predicted to cause loss of function and are not associated with chromosomal instability. Cancer Res. 2005, 65, 11361–11366. [Google Scholar] [CrossRef] [PubMed]
- Miyaki, M.; Yamaguchi, T.; Iijima, T.; Takahashi, K.; Matsumoto, H.; Mori, T. Somatic mutations of the CDC4 (FBXW7) gene in hereditary colorectal tumors. Oncology 2009, 76, 430–434. [Google Scholar] [CrossRef] [PubMed]
- Iwatsuki, M.; Mimori, K.; Ishii, H.; Yokobori, T.; Takatsuno, Y.; Sato, T.; Toh, H.; Onoyama, I.; Nakayama, K.I.; Baba, H.; et al. Loss of FBXW7, a cell cycle regulating gene, in colorectal cancer: Clinical significance. Int. J. Cancer 2010, 126, 1828–1837. [Google Scholar]
- Inuzuka, H.; Shaik, S.; Onoyama, I.; Gao, D.; Tseng, A.; Maser, R.S.; Zhai, B.; Wan, L.; Gutierrez, A.; Lau, A.W.; et al. SCF(FBW7) regulates cellular apoptosis by targeting MCL1 for ubiquitylation and destruction. Nature 2011, 471, 104–109. [Google Scholar] [CrossRef] [PubMed]
- Wertz, I.E.; Kusam, S.; Lam, C.; Okamoto, T.; Sandoval, W.; Anderson, D.J.; Helgason, E.; Ernst, J.A.; Eby, M.; Liu, J.; et al. Sensitivity to antitubulin chemotherapeutics is regulated by MCL1 and FBW7. Nature 2011, 471, 110–114. [Google Scholar] [CrossRef]
- Bhola, P.D.; Letai, A. Mitochondria-Judges and Executioners of Cell Death Sentences. Mol. Cell 2016, 61, 695–704. [Google Scholar] [CrossRef] [PubMed]
- Tong, J.; Tan, S.; Zou, F.; Yu, J.; Zhang, L. FBW7 mutations mediate resistance of colorectal cancer to targeted therapies by blocking Mcl-1 degradation. Oncogene 2017, 36, 787–796. [Google Scholar] [CrossRef]
- Adhikary, S.; Marinoni, F.; Hock, A.; Hulleman, E.; Popov, N.; Beier, R.; Bernard, S.; Quarto, M.; Capra, M.; Goettig, S.; et al. The ubiquitin ligase HectH9 regulates transcriptional activation by Myc and is essential for tumor cell proliferation. Cell 2005, 123, 409–421. [Google Scholar] [CrossRef]
- Myant, K.B.; Cammareri, P.; Hodder, M.C.; Wills, J.; Von Kriegsheim, A.; Gyorffy, B.; Rashid, M.; Polo, S.; Maspero, E.; Vaughan, L.; et al. HUWE1 is a critical colonic tumour suppressor gene that prevents MYC signalling, DNA damage accumulation and tumour initiation. EMBO Mol. Med. 2017, 9, 181–197. [Google Scholar] [CrossRef]
- Dominguez-Brauer, C.; Khatun, R.; Elia, A.J.; Thu, K.L.; Ramachandran, P.; Baniasadi, S.P.; Hao, Z.; Jones, L.D.; Haight, J.; Sheng, Y.; et al. E3 ubiquitin ligase Mule targets beta-catenin under conditions of hyperactive Wnt signaling. Proc. Natl. Acad. Sci. USA 2017, 114, 1148–1157. [Google Scholar] [CrossRef]
- Tarcic, O.; Pateras, I.S.; Cooks, T.; Shema, E.; Kanterman, J.; Ashkenazi, H.; Boocholez, H.; Hubert, A.; Rotkopf, R.; Baniyash, M.; et al. RNF20 Links Histone H2B Ubiquitylation with Inflammation and Inflammation-Associated Cancer. Cell Rep. 2016, 14, 1462–1476. [Google Scholar] [CrossRef]
- Wang, Z.; Zhu, L.; Guo, T.; Wang, Y.; Yang, J. Decreased H2B monoubiquitination and overexpression of ubiquitin-specific protease enzyme 22 in malignant colon carcinoma. Hum. Pathol. 2015, 46, 1006–1014. [Google Scholar] [CrossRef] [PubMed]
- Espinosa, J.M. Histone H2B ubiquitination: The cancer connection. Genes Dev. 2008, 22, 2743–2749. [Google Scholar] [CrossRef][Green Version]
- Chernikova, S.B.; Razorenova, O.V.; Higgins, J.P.; Sishc, B.J.; Nicolau, M.; Dorth, J.A.; Chernikova, D.A.; Kwok, S.; Brooks, J.D.; Bailey, S.M.; et al. Deficiency in mammalian histone H2B ubiquitin ligase Bre1 (Rnf20/Rnf40) leads to replication stress and chromosomal instability. Cancer Res. 2012, 72, 2111–2119. [Google Scholar] [CrossRef] [PubMed]
- Carter, S.L.; Eklund, A.C.; Kohane, I.S.; Harris, L.N.; Szallasi, Z. A signature of chromosomal instability inferred from gene expression profiles predicts clinical outcome in multiple human cancers. Nat. Genet. 2006, 38, 1043–1048. [Google Scholar] [CrossRef]
- Thompson, L.L.; Jeusset, L.M.; Lepage, C.C.; McManus, K.J. Evolving Therapeutic Strategies to Exploit Chromosome Instability in Cancer. Cancers 2017, 9, 151. [Google Scholar] [CrossRef] [PubMed]
- Brocato, J.; Costa, M. SATB1 and 2 in colorectal cancer. Carcinogenesis 2014, 36, 186–191. [Google Scholar] [CrossRef]
- Mir, R.; Pradhan, S.J.; Patil, P.; Mulherkar, R.; Galande, S. Wnt/beta-catenin signaling regulated SATB1 promotes colorectal cancer tumorigenesis and progression. Oncogene 2016, 35, 1679–1691. [Google Scholar] [CrossRef]
- Yu, L.; Dong, L.; Wang, Y.; Liu, L.; Long, H.; Li, H.; Li, J.; Yang, X.; Liu, Z.; Duan, G.; et al. Reversible regulation of SATB1 ubiquitination by USP47 and SMURF2 mediates colon cancer cell proliferation and tumor progression. Cancer Lett. 2019, 448, 40–51. [Google Scholar] [CrossRef] [PubMed]
- Klupp, F.; Giese, C.; Halama, N.; Franz, C.; Lasitschka, F.; Warth, A.; Schmidt, T.; Kloor, M.; Ulrich, A.; Schneider, M. E3 ubiquitin ligase Smurf2: A prognostic factor in microsatellite stable colorectal cancer. Cancer Manag. Res. 2019, 11, 1795–1803. [Google Scholar] [CrossRef] [PubMed]
- Ougolkov, A.; Zhang, B.; Yamashita, K.; Bilim, V.; Mai, M.; Fuchs, S.Y.; Minamoto, T. Associations among beta-TrCP, an E3 ubiquitin ligase receptor, beta-catenin, and NF-kappaB in colorectal cancer. J. Natl. Cancer Inst. 2004, 96, 1161–1170. [Google Scholar] [CrossRef]
- Kitagawa, M.; Hatakeyama, S.; Shirane, M.; Matsumoto, M.; Ishida, N.; Hattori, K.; Nakamichi, I.; Kikuchi, A.; Nakayama, K.; Nakayama, K. An F-box protein, FWD1, mediates ubiquitin-dependent proteolysis of beta-catenin. EMBO J. 1999, 18, 2401–2410. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Kato, Y.; Zhang, Z.; Do, V.M.; Yankner, B.A.; He, X. beta-Trcp couples beta-catenin phosphorylation-degradation and regulates Xenopus axis formation. Proc. Natl. Acad. Sci. USA 1999, 96, 6273–6278. [Google Scholar] [CrossRef] [PubMed]
- Hadjihannas, M.V.; Bernkopf, D.B.; Bruckner, M.; Behrens, J. Cell cycle control of Wnt/beta-catenin signalling by conductin/axin2 through CDC20. EMBO Rep. 2012, 13, 347–354. [Google Scholar] [CrossRef] [PubMed]
- Uddin, S.; Ahmed, M.; Bavi, P.; El-Sayed, R.; Al-Sanea, N.; AbdulJabbar, A.; Ashari, L.H.; Alhomoud, S.; Al-Dayel, F.; Hussain, A.R.; et al. Bortezomib (Velcade) induces p27Kip1 expression through S-phase kinase protein 2 degradation in colorectal cancer. Cancer Res. 2008, 68, 3379–3388. [Google Scholar] [CrossRef] [PubMed]
- Shapira, M.; Ben-Izhak, O.; Linn, S.; Futerman, B.; Minkov, I.; Hershko, D.D. The prognostic impact of the ubiquitin ligase subunits Skp2 and Cks1 in colorectal carcinoma. Cancer 2005, 103, 1336–1346. [Google Scholar] [CrossRef]
- Bochis, O.V.; Irimie, A.; Pichler, M.; Berindan-Neagoe, I. The role of Skp2 and its substrate CDKN1B (p27) in colorectal cancer. J. Gastrointestin Liver Dis. 2015, 24, 225–234. [Google Scholar] [PubMed]
- Hao, B.; Zheng, N.; Schulman, B.A.; Wu, G.; Miller, J.J.; Pagano, M.; Pavletich, N.P. Structural Basis of the Cks1-Dependent Recognition of p27Kip1 by the SCFSkp2 Ubiquitin Ligase. Mol. Cell 2005, 20, 9–19. [Google Scholar] [CrossRef]
- Okuda, H.; Saitoh, K.; Hirai, S.; Iwai, K.; Takaki, Y.; Baba, M.; Minato, N.; Ohno, S.; Shuin, T. The von Hippel-Lindau tumor suppressor protein mediates ubiquitination of activated atypical protein kinase C. J. Biol. Chem. 2001, 276, 43611–43617. [Google Scholar] [CrossRef]
- Cockman, M.E.; Masson, N.; Mole, D.R.; Jaakkola, P.; Chang, G.-W.; Clifford, S.C.; Maher, E.R.; Pugh, C.W.; Ratcliffe, P.J.; Maxwell, P.H. Hypoxia Inducible Factor-α Binding and Ubiquitylation by the von Hippel-Lindau Tumor Suppressor Protein. J. Biol. Chem. 2000, 275, 25733–25741. [Google Scholar] [CrossRef]
- Jung, C.R.; Hwang, K.S.; Yoo, J.; Cho, W.K.; Kim, J.M.; Kim, W.H.; Im, D.S. E2-EPF UCP targets pVHL for degradation and associates with tumor growth and metastasis. Nat. Med. 2006, 12, 809–816. [Google Scholar] [CrossRef]
- Suzuki, Y.; Nakabayashi, Y.; Takahashi, R. Ubiquitin-protein ligase activity of X-linked inhibitor of apoptosis protein promotes proteasomal degradation of caspase-3 and enhances its anti-apoptotic effect in Fas-induced cell death. Proc. Natl. Acad. Sci. USA 2001, 98, 8662–8667. [Google Scholar] [CrossRef]
- Ehrenschwender, M.; Bittner, S.; Seibold, K.; Wajant, H. XIAP-targeting drugs re-sensitize PIK3CA-mutated colorectal cancer cells for death receptor-induced apoptosis. Cell Death Dis. 2014, 5, e1570. [Google Scholar] [CrossRef]
- Hanson, A.J.; Wallace, H.A.; Freeman, T.J.; Beauchamp, R.D.; Lee, L.A.; Lee, E. XIAP monoubiquitylates Groucho/TLE to promote canonical Wnt signaling. Mol. Cell 2012, 45, 619–628. [Google Scholar] [CrossRef]
- Zhu, B.; Zheng, Y.; Pham, A.D.; Mandal, S.S.; Erdjument-Bromage, H.; Tempst, P.; Reinberg, D. Monoubiquitination of human histone H2B: The factors involved and their roles in HOX gene regulation. Mol. Cell 2005, 20, 601–611. [Google Scholar] [CrossRef]
- Hattori, T.; Kishino, T.; Stephen, S.; Eberspaecher, H.; Maki, S.; Takigawa, M.; de Crombrugghe, B.; Yasuda, H. E6-AP/UBE3A protein acts as a ubiquitin ligase toward SOX9 protein. J. Biol. Chem. 2013, 288, 35138–35148. [Google Scholar] [CrossRef] [PubMed]
- Heng, S.; Xue-Bing, L.; Ya, M.; Li, F.; Min, L.; Jing, F.J.C.R. TRAF6 upregulates expression of HIF-1α and promotes tumor angiogenesis. Cancer Res. 2013, 73, 4950–4959. [Google Scholar]
- Zhang, T.; Wang, H.; Han, L. Expression and Clinical Significance of Tumor Necrosis Factor Receptor-Associated Factor 6 in Patients with Colon Cancer. Iran. Red. Crescent Med. J. 2016, 18, e23931. [Google Scholar] [CrossRef][Green Version]
- Liu, L.; Wong, C.C.; Gong, B.; Yu, J. Functional significance and therapeutic implication of ring-type E3 ligases in colorectal cancer. Oncogene 2017, 37, 148. [Google Scholar] [CrossRef] [PubMed]
- Deng, L.; Wang, C.; Spencer, E.; Yang, L.; Braun, A.; You, J.; Slaughter, C.; Pickart, C.; Chen, Z.J. Activation of the IkappaB Kinase Complex by TRAF6 Requires a Dimeric Ubiquitin-Conjugating Enzyme Complex and a Unique Polyubiquitin Chain. Cell 2000, 103, 351–361. [Google Scholar] [CrossRef]
- Krajewska, M.; Kim, H.; Kim, C.; Kang, H.; Welsh, K.; Matsuzawa, S.; Tsukamoto, M.; Thomas, R.G.; Assa-Munt, N.; Piao, Z.; et al. Analysis of apoptosis protein expression in early-stage colorectal cancer suggests opportunities for new prognostic biomarkers. Clin. Cancer Res. 2005, 11, 5451–5461. [Google Scholar] [CrossRef] [PubMed]
- Welcker, M.; Clurman, B.E. FBW7 ubiquitin ligase: A tumour suppressor at the crossroads of cell division, growth and differentiation. Nat. Rev. Cancer 2008, 8, 83. [Google Scholar] [CrossRef] [PubMed]
- Nateri, A.S.; Riera-Sans, L.; Da Costa, C.; Behrens, A. The ubiquitin ligase SCFFbw7 antagonizes apoptotic JNK signaling. Science 2004, 303, 1374–1378. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.Y.; Lee, Y.; Kim, J.H.; Chung, A.S.; Joo, J.H.; Kim, C.N.; Kim, N.S.; Choe, I.S.; Kim, J.W. Over-expression of human UREB1 in colorectal cancer: HECT domain of human UREB1 inhibits the activity of tumor suppressor p53 protein. Biochem. Biophys. Res. Commun. 2005, 326, 7–17. [Google Scholar] [CrossRef]
- Kubbutat, M.H.; Jones, S.N.; Vousden, K.H. Regulation of p53 stability by Mdm2. Nature 1997, 387, 299–303. [Google Scholar] [CrossRef]
- Abdel-Fattah, G.; Yoffe, B.; Krishnan, B.; Khaoustov, V.; Ltani, K. MDM2/p53 protein expression in the development of colorectal adenocarcinoma. J. Gastrointest Surg. 2000, 4, 109–114. [Google Scholar] [CrossRef]
- Thomas, J.J.; Abed, M.; Heuberger, J.; Novak, R.; Zohar, Y.; Beltran Lopez, A.P.; Trausch-Azar, J.S.; Ilagan, M.X.G.; Benhamou, D.; Dittmar, G.; et al. RNF4-Dependent Oncogene Activation by Protein Stabilization. Cell Rep. 2016, 16, 3388–3400. [Google Scholar] [CrossRef]
- Gonzalez-Prieto, R.; Cuijpers, S.A.; Kumar, R.; Hendriks, I.A.; Vertegaal, A.C. c-Myc is targeted to the proteasome for degradation in a SUMOylation-dependent manner, regulated by PIAS1, SENP7 and RNF4. Cell Cycle 2015, 14, 1859–1872. [Google Scholar] [CrossRef] [PubMed]
- Hakli, M.; Lorick, K.L.; Weissman, A.M.; Janne, O.A.; Palvimo, J.J. Transcriptional coregulator SNURF (RNF4) possesses ubiquitin E3 ligase activity. FEBS Lett. 2004, 560, 56–62. [Google Scholar] [CrossRef]
- Wu, B.; Piloto, S.; Zeng, W.; Hoverter, N.P.; Schilling, T.F.; Waterman, M.L. Ring Finger Protein 14 is a new regulator of TCF/β-catenin-mediated transcription and colon cancer cell survival. EMBO Rep. 2013, 14, 347–355. [Google Scholar] [CrossRef]
- Cantù, C.; Valenta, T.; Basler, K. A RING finger to wed TCF and β-catenin. EMBO Rep. 2013, 14, 295–296. [Google Scholar] [CrossRef]
- Jiang, X.; Charlat, O.; Zamponi, R.; Yang, Y.; Cong, F. Dishevelled promotes Wnt receptor degradation through recruitment of ZNRF3/RNF43 E3 ubiquitin ligases. Mol. Cell 2015, 58, 522–533. [Google Scholar] [CrossRef]
- Koo, B.K.; Spit, M.; Jordens, I.; Low, T.Y.; Stange, D.E.; van de Wetering, M.; van Es, J.H.; Mohammed, S.; Heck, A.J.; Maurice, M.M.; et al. Tumour suppressor RNF43 is a stem-cell E3 ligase that induces endocytosis of Wnt receptors. Nature 2012, 488, 665–669. [Google Scholar] [CrossRef] [PubMed]
- Giannakis, M.; Hodis, E.; Jasmine Mu, X.; Yamauchi, M.; Rosenbluh, J.; Cibulskis, K.; Saksena, G.; Lawrence, M.S.; Qian, Z.R.; Nishihara, R.; et al. RNF43 is frequently mutated in colorectal and endometrial cancers. Nat. Genet. 2014, 46, 1264–1266. [Google Scholar] [CrossRef]
- Piao, M.Y.; Cao, H.L.; He, N.N.; Xu, M.Q.; Dong, W.X.; Wang, W.Q.; Wang, B.M.; Zhou, B. Potential role of TRIM3 as a novel tumour suppressor in colorectal cancer (CRC) development. Scand. J. Gastroenterol. 2016, 51, 572–582. [Google Scholar] [CrossRef]
- Lee, O.H.; Lee, J.; Lee, K.H.; Woo, Y.M.; Kang, J.H.; Yoon, H.G.; Bae, S.K.; Songyang, Z.; Oh, S.H.; Choi, Y. Role of the focal adhesion protein TRIM15 in colon cancer development. Biochim. Biophys. Acta. 2015, 1853, 409–421. [Google Scholar] [CrossRef] [PubMed]
- Glebov, O.K.; Rodriguez, L.M.; Soballe, P.; DeNobile, J.; Cliatt, J.; Nakahara, K.; Kirsch, I.R. Gene expression patterns distinguish colonoscopically isolated human aberrant crypt foci from normal colonic mucosa. Cancer Epidemiol. Biomarkers Prev. 2006, 15, 2253–2262. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Coulie, P.G.; Van den Eynde, B.J.; van der Bruggen, P.; Boon, T. Tumour antigens recognized by T lymphocytes: At the core of cancer immunotherapy. Nat. Rev. Cancer 2014, 14, 135–146. [Google Scholar] [CrossRef]
- Botter, S.M.; Neri, D.; Fuchs, B. Recent advances in osteosarcoma. Curr. Opin. Pharmacol. 2014, 16, 15–23. [Google Scholar] [CrossRef] [PubMed]
- Kaste, S.C.; Pratt, C.B.; Cain, A.M.; Jones-Wallace, D.J.; Rao, B.N. Metastases detected at the time of diagnosis of primary pediatric extremity osteosarcoma at diagnosis: Imaging features. Cancer 1999, 86, 1602–1608. [Google Scholar] [CrossRef]
- Haydon, R.C.; Luu, H.H.; He, T.C. Osteosarcoma and osteoblastic differentiation: A new perspective on oncogenesis. Clin. Orthop. Relat. Res. 2007, 454, 237–246. [Google Scholar] [CrossRef]
- Thomas, D.M.; Johnson, S.A.; Sims, N.A.; Trivett, M.K.; Slavin, J.L.; Rubin, B.P.; Waring, P.; McArthur, G.A.; Walkley, C.R.; Holloway, A.J.; et al. Terminal osteoblast differentiation, mediated by runx2 and p27KIP1, is disrupted in osteosarcoma. J. Cell Biol. 2004, 167, 925–934. [Google Scholar] [CrossRef]
- Luo, X.; Chen, J.; Song, W.X.; Tang, N.; Luo, J.; Deng, Z.L.; Sharff, K.A.; He, G.; Bi, Y.; He, B.C.; et al. Osteogenic BMPs promote tumor growth of human osteosarcomas that harbor differentiation defects. Lab. Invest. 2008, 88, 1264–1277. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Kang, Q.; Cheng, H.; Li, X.; Sun, M.H.; Jiang, W.; Luu, H.H.; Park, J.Y.; Haydon, R.C.; He, T.-C. Transcriptional Characterization of Bone Morphogenetic Proteins (BMPs)-Mediated Osteogenic Signaling. J. Cell Biochem. 2003, 90, 1149–1165. [Google Scholar] [CrossRef]
- Peng, Y.; Kang, Q.; Luo, Q.; Jiang, W.; Si, W.; Liu, B.A.; Luu, H.H.; Park, J.K.; Li, X.; Luo, J.; et al. Inhibitor of DNA binding/differentiation helix-loop-helix proteins mediate bone morphogenetic protein-induced osteoblast differentiation of mesenchymal stem cells. J. Biol. Chem. 2004, 279, 32941–32949. [Google Scholar] [CrossRef] [PubMed]
- Ducy, P.; Schinke, T.; Karsenty, G. The osteoblast: A sophisticated fibroblast under central surveillance. Science 2000, 289, 1501–1504. [Google Scholar] [CrossRef]
- Wang, L.; Park, P.; Lin, C.Y. Characterization of stem cell attributes in human osteosarcoma cell lines. Cancer Biol. Ther. 2009, 8, 543–552. [Google Scholar] [CrossRef]
- Levings, P.P.; McGarry, S.V.; Currie, T.P.; Nickerson, D.M.; McClellan, S.; Ghivizzani, S.C.; Steindler, D.A.; Gibbs, C.P. Expression of an exogenous human Oct-4 promoter identifies tumor-initiating cells in osteosarcoma. Cancer Res. 2009, 69, 5648–5655. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Zhuang, Y.; Zhang, Y.; Yang, X.; Zhang, H.; Wang, G.; Yin, W.; Wang, R.; Zhang, Z.; Xiao, W. Uev1A facilitates osteosarcoma differentiation by promoting Smurf1-mediated Smad1 ubiquitination and degradation. Cell Death Dis. 2017, 8, 2974. [Google Scholar] [CrossRef]
- Jones, D.C.; Wein, M.N.; Oukka, M.; Hofstaetter, J.G.; Glimcher, M.J.; Glimcher, L.H. Regulation of Adult Bone Mass by the Zinc Finger Adapter Protein Schnurri-3. Science 2006, 312, 1223–1227. [Google Scholar] [CrossRef]
- Ying, M.; Zhang, L.; Zhou, Q.; Shao, X.; Cao, J.; Zhang, N.; Li, W.; Zhu, H.; Yang, B.; He, Q. The E3 ubiquitin protein ligase MDM2 dictates all-trans retinoic acid-induced osteoblastic differentiation of osteosarcoma cells by modulating the degradation of RARα. Oncogene 2016, 35, 4358. [Google Scholar] [CrossRef]
- Chen, Z.; Shen, B.L.; Fu, Q.G.; Wang, F.; Tang, Y.X.; Hou, C.L.; Chen, L. CUL4B promotes proliferation and inhibits apoptosis of human osteosarcoma cells. Oncol. Rep. 2014, 32, 2047–2053. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Zhang, W.; Jiang, K.; Chen, B.; Wang, K.; Lao, L.; Hou, C.; Wang, F.; Zhang, C.; Shen, H. MicroRNA-300 Regulates the Ubiquitination of PTEN through the CRL4B(DCAF13) E3 Ligase in Osteosarcoma Cells. Mol. Ther. Nucleic Acids 2018, 10, 254–268. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Chen, B.; Jiang, K.; Lao, L.; Shen, H.; Chen, Z. Activation of TNF-α/NF-κB axis enhances CRL4B(DCAF)(11) E3 ligase activity and regulates cell cycle progression in human osteosarcoma cells. Mol. Oncol. 2018, 12, 476–494. [Google Scholar] [CrossRef] [PubMed]
- Severe, N.; Dieudonne, F.X.; Marty, C.; Modrowski, D.; Patino-Garcia, A.; Lecanda, F.; Fromigue, O.; Marie, P.J. Targeting the E3 ubiquitin casitas B-lineage lymphoma decreases osteosarcoma cell growth and survival and reduces tumorigenesis. J. Bone. Miner. Res. 2012, 27, 2108–2117. [Google Scholar] [CrossRef]
- Archange, C.; Nowak, J.; Garcia, S.; Moutardier, V.; Calvo, E.L.; Dagorn, J.C.; Iovanna, J.L. The WSB1 gene is involved in pancreatic cancer progression. PLoS ONE 2008, 3, 2475. [Google Scholar] [CrossRef]
- Dentice, M.; Bandyopadhyay, A.; Gereben, B.; Callebaut, I.; Christoffolete, M.A.; Kim, B.W.; Nissim, S.; Mornon, J.P.; Zavacki, A.M.; Zeold, A.; et al. The Hedgehog-inducible ubiquitin ligase subunit WSB-1 modulates thyroid hormone activation and PTHrP secretion in the developing growth plate. Nat. Cell Biol. 2005, 7, 698–705. [Google Scholar] [CrossRef]
- Choi, D.W.; Seo, Y.M.; Kim, E.A.; Sung, K.S.; Ahn, J.W.; Park, S.J.; Lee, S.R.; Choi, C.Y. Ubiquitination and degradation of homeodomain-interacting protein kinase 2 by WD40 repeat/SOCS box protein WSB-1. J. Biol. Chem. 2008, 283, 4682–4689. [Google Scholar] [CrossRef]
- Cao, J.; Wang, Y.; Dong, R.; Lin, G.; Zhang, N.; Wang, J.; Lin, N.; Gu, Y.; Ding, L.; Ying, M.; et al. Hypoxia-Induced WSB1 Promotes the Metastatic Potential of Osteosarcoma Cells. Cancer Res. 2015, 75, 4839–4851. [Google Scholar] [CrossRef]
- El-Naggar, A.M.; Clarkson, P.W.; Negri, G.L.; Turgu, B.; Zhang, F.; Anglesio, M.S.; Sorensen, P.H. HACE1 is a potential tumor suppressor in osteosarcoma. Cell Death Dis. 2019, 10, 21. [Google Scholar] [CrossRef]
- Chen, L.; Pei, H.; Lu, S.J.; Liu, Z.J.; Yan, L.; Zhao, X.M.; Hu, B.; Lu, H.G. SPOP suppresses osteosarcoma invasion via PI3K/AKT/NF-kappaB signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 609–615. [Google Scholar]
- Zhao, M.; Su, Z.; Zhang, S.; Zhuang, L.; Xie, Y.; Li, X. Suppressive Role of MicroRNA-148a in Cell Proliferation and Invasion in Ovarian Cancer Through Targeting Transforming Growth Factor-β-Induced 2. Oncol. Res. 2016, 24, 353–360. [Google Scholar] [CrossRef]
- Moon, S.; Lee, B.H. Chemically Induced Cellular Proteolysis: An Emerging Therapeutic Strategy for Undruggable Targets. Mol. Cells 2018, 41, 933–942. [Google Scholar]
- Zeng, X.; Sigoillot, F.; Gaur, S.; Choi, S.; Pfaff, K.L.; Oh, D.C.; Hathaway, N.; Dimova, N.; Cuny, G.D.; King, R.W. Pharmacologic inhibition of the anaphase-promoting complex induces a spindle checkpoint-dependent mitotic arrest in the absence of spindle damage. Cancer cell 2010, 18, 382–395. [Google Scholar] [CrossRef] [PubMed]
- Sackton, K.L.; Dimova, N.; Zeng, X.; Tian, W.; Zhang, M.; Sackton, T.B.; Meaders, J.; Pfaff, K.L.; Sigoillot, F.; Yu, H.; et al. Synergistic blockade of mitotic exit by two chemical inhibitors of the APC/C. Nature 2014, 514, 646–649. [Google Scholar] [CrossRef]
- Zhang, B.; Golding, B.T.; Hardcastle, I.R. Small-molecule MDM2-p53 inhibitors: Recent advances. Future Med. Chem. 2015, 7, 631–645. [Google Scholar] [CrossRef]
- Zhao, Y.; Aguilar, A.; Bernard, D.; Wang, S. Small-molecule inhibitors of the MDM2-p53 protein-protein interaction (MDM2 Inhibitors) in clinical trials for cancer treatment. J. Med. Chem. 2015, 58, 1038–1052. [Google Scholar] [CrossRef] [PubMed]
- Chan, C.-H.; Morrow, J.K.; Li, C.-F.; Gao, Y.; Jin, G.; Moten, A.; Stagg, L.J.; Ladbury, J.E.; Cai, Z.; Xu, D.; et al. Pharmacological inactivation of Skp2 SCF ubiquitin ligase restricts cancer stem cell traits and cancer progression. Cell 2013, 154, 556–568. [Google Scholar] [CrossRef]
- Wu, L.; Grigoryan, A.V.; Li, Y.; Hao, B.; Pagano, M.; Cardozo, T.J. Specific small molecule inhibitors of Skp2-mediated p27 degradation. Chem. Biol. 2012, 19, 1515–1524. [Google Scholar] [CrossRef] [PubMed]
- Veggiani, G.; Gerpe, M.C.R.; Sidhu, S.S.; Zhang, W. Emerging drug development technologies targeting ubiquitination for cancer therapeutics. Pharmacol. Ther. 2019, 19. [Google Scholar] [CrossRef] [PubMed]
| E2 | E3 | Function | Reference |
|---|---|---|---|
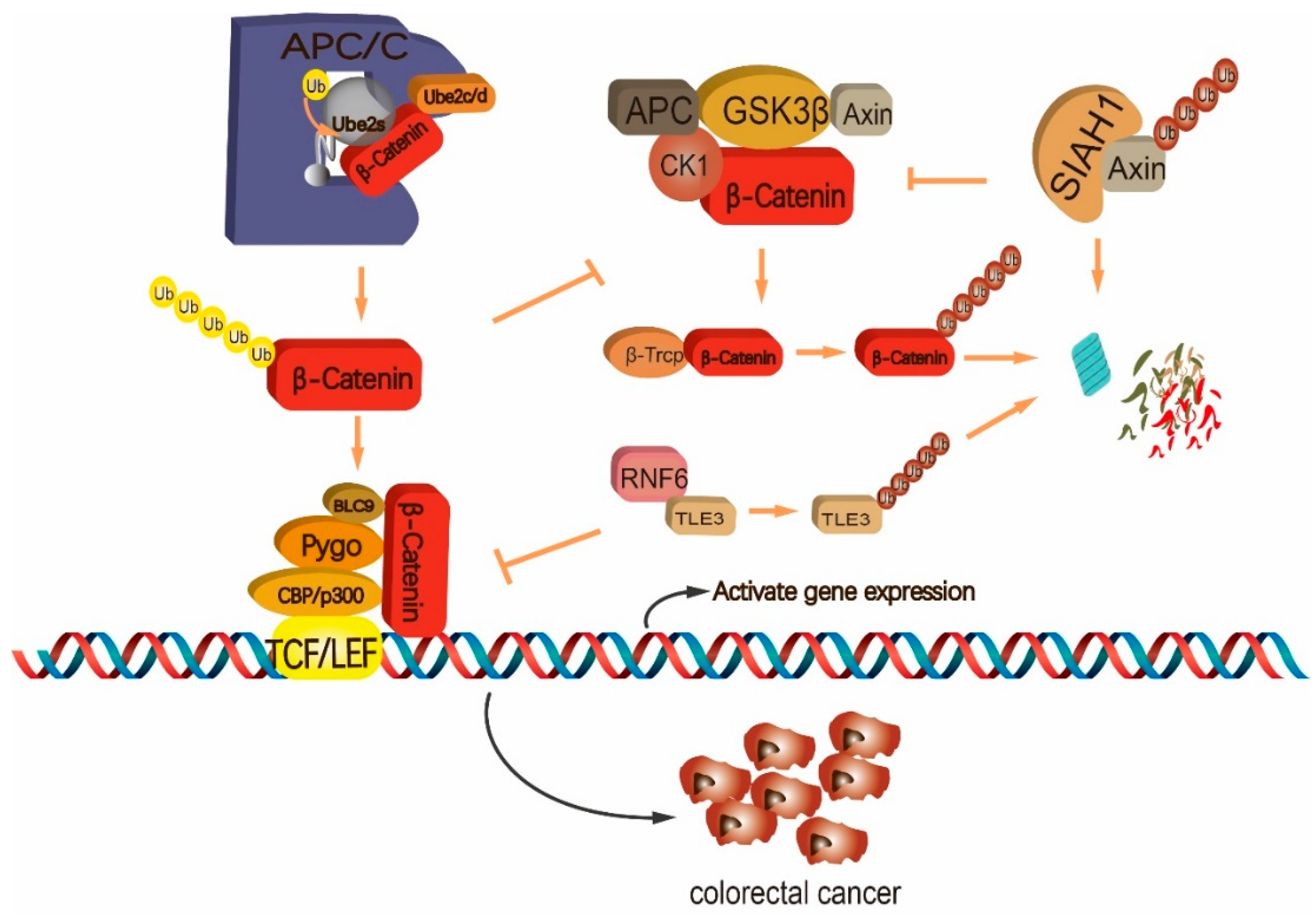
| UBC3 /UBC4 | β-TrCP | 1. β-TrCP ubiquitinates phosphorylated IκB for degradation, which enhances the NF-κB signaling. Increased β-TrCP is associated with an enhanced NF-κB signaling in CRC. 2. β-TrCP ubiquitinates β-Catenin via K48-linked poly-Ub chain for proteasomal degradation, which suppresses CRC progression. | [190,191,192] |
| UBCH5 /UBCH10/UBE2S | APC/C | 1. APC/Ccdc20 ubiquitinates Conductin for degradation during mitotic exit, which regulates the Wnt/β-catenin signaling and CRC cell growth. 2. UBE2S collaborates with the APC/C complex to stablize β-Catenin via K11-linked polyubiqutination. This activity enhances CRC proliferation and metastasis. | [58,193] |
| UBCH5 | HECTH9 | HECTH9 modifies C-MYC with K63-linked poly-Ub chain to promote CRC cell proliferation. The expression of HECTH9 is increased in the cancer tissues of CRC patients. | [177] |
| UBCH5 | S-phase kinase protein 2 (SKP2) | SKP2 ubiquitinates p27Kip1 for degradation in CRC cells. Elevated expression of SKP2 and reduced expression of p27Kip1 is associated with poor prognosis and decreased survival of CRC patients. | [194,195,196,197] |
| UBCH5B/ UBE2S | von Hippel-Lindau protein (pVHL) | VCB-Cul2 ubiquitinates HIF-1α for degradation under hypoxic conditions, which suppresses CRC malignancy. | [156,198,199,200] |
| UBCH5B | X-chromosome-linked IAP (XIAP), | 1. XIAP ubiquitinates active caspase-3 for degradation to suppress apoptosis. Inhibition of XIAP increases the sensitivity of PIK3CA-mutated CRC cells for induced cell death. 2. XIAP monoubiquitinates TLE, which promotes β-catenin-TCF association and enhances activation of the Wnt pathway in CRC cells. | [201,202,203] |
| UBCH6 | RNF20/RNF40 | RNF20/RNF40 monoubiquitinates H2B-K120, which is required by transcription regulation. Loss of H2BK120ub1 is associated with poor therapeutic outcome in CRC. | [181,182,204] |
| UBC9 | E6-AP | E6-AP coordinates with UBC9 to ubiquitinate SOX9 for degradation, which may repress Sox9-enhanced CRC malignancy. | [205] |
| UBC13/ UEV1A | Tumor necrosis factor receptor-associated factor 6 (TARF6) | 1. TARF6 ubiquitinates IKK via K63-linked poly-Ub chain to promote the NF-κB signaling pathway. High expression of TRAF6 is associated with a decreased survival of CRC patients. 2. TARF6 stabilizes hypoxia-inducible factor (HIF)–1a through K63-linked polyubiquitination, which promotes angiogenesis and growth of CRC. 3. TRAF6 ubiquitinates LC3B via K63 linkage, which allows LC3B to recognize β-Catenin and drive it for autophagic degradation. This activity is involved in inhibiting the metastasis of CRC cells. | [161,206,207,208,209] |
| c-IAP | c-IAP is upregulated in CRC patients with a reduced survival. | [210] | |
| FBXW7 | FBXW7 ubiquitinates ZEB2 and MCL1 for degradation, which is involved in regulating the malignancy and therapy resistance of CRC cells. Somatic mutations in FBXW7 is detected in CRC patients. | [169,171,172,176,211,212] | |
| Human upstream regulatory element binding protein 1 (hUREB1) | hUREB1 down-regulates p53 through ubiquitination in CRC cells. Increased expression of hUREB1 is correlated with p53 destabilization in CRC patients. | [213] | |
| MDM2 | MDM2 ubiquitinates p53 for degradation. Increased expression of MDM2 is correlated with negative expression of p53 in CRC patients. | [214,215] | |
| RNF4 | RNF4 ubiquitinates and stabilizes multiple oncoproteins, such as c-Myc and β-Catenin. Elevated expression of RNF4 is correlated with CRC tumorigenesis. | [216,217,218] | |
| RNF6 | RNF6 enhances the interaction between β-Catenin and TCF4/LEF through ubiquitinating transducin-like enhancer of split 3 (TLE3) for degradation, which promotes CRC cell growth and metastasis. | [162] | |
| RNF14 | RNF14 activates the Wnt pathway through interacting with TCFs to promote β-Catenin recruitment, which promotes CRC cell growth. | [219,220] | |
| Ubiquitin E3 ligase ring finger 43 (RNF43) | RNF43 ubiquitinates Frizzled (FZD) and LRP6 for degradation. About 18% of CRC patients bear RNF43 truncating mutation. | [221,222,223] | |
| Tripartite motif (TRIM3) | TRIM3 enhances the stability of p53 and suppresses CRC development. | [224] | |
| TRIM15 | TRIM15 serves as a putative CRC suppressor, which inhibits CRC cell growth and metastasis. | [225] | |
| TRIM29 | TRIM29 is upregulated in aberrant crypt foci in human colon and serves as a putative biomarker for CRC diagnosis. | [226] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, D.; Bu, F.; Zhang, W. The Role of Ubiquitination in Regulating Embryonic Stem Cell Maintenance and Cancer Development. Int. J. Mol. Sci. 2019, 20, 2667. https://doi.org/10.3390/ijms20112667
Wang D, Bu F, Zhang W. The Role of Ubiquitination in Regulating Embryonic Stem Cell Maintenance and Cancer Development. International Journal of Molecular Sciences. 2019; 20(11):2667. https://doi.org/10.3390/ijms20112667
Chicago/Turabian StyleWang, Dian, Fan Bu, and Weiwei Zhang. 2019. "The Role of Ubiquitination in Regulating Embryonic Stem Cell Maintenance and Cancer Development" International Journal of Molecular Sciences 20, no. 11: 2667. https://doi.org/10.3390/ijms20112667
APA StyleWang, D., Bu, F., & Zhang, W. (2019). The Role of Ubiquitination in Regulating Embryonic Stem Cell Maintenance and Cancer Development. International Journal of Molecular Sciences, 20(11), 2667. https://doi.org/10.3390/ijms20112667




