Detection and Correlation of Single and Concomitant TP53, PTEN, and CDKN2A Alterations in Gliomas
Abstract
1. Introduction
2. Results
2.1. Clinical Parameters
2.2. Molecular Data
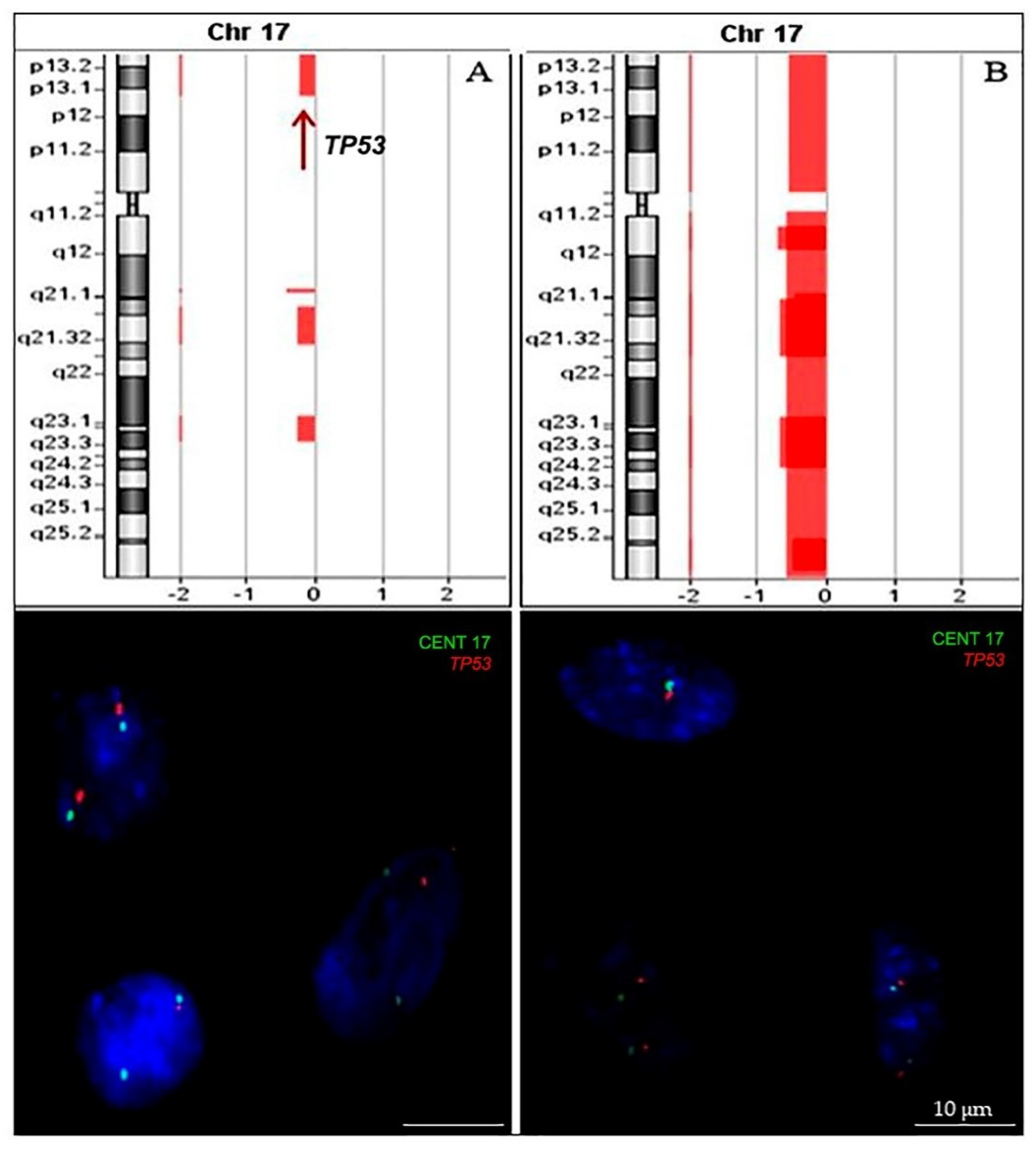
2.2.1. TP53 Status
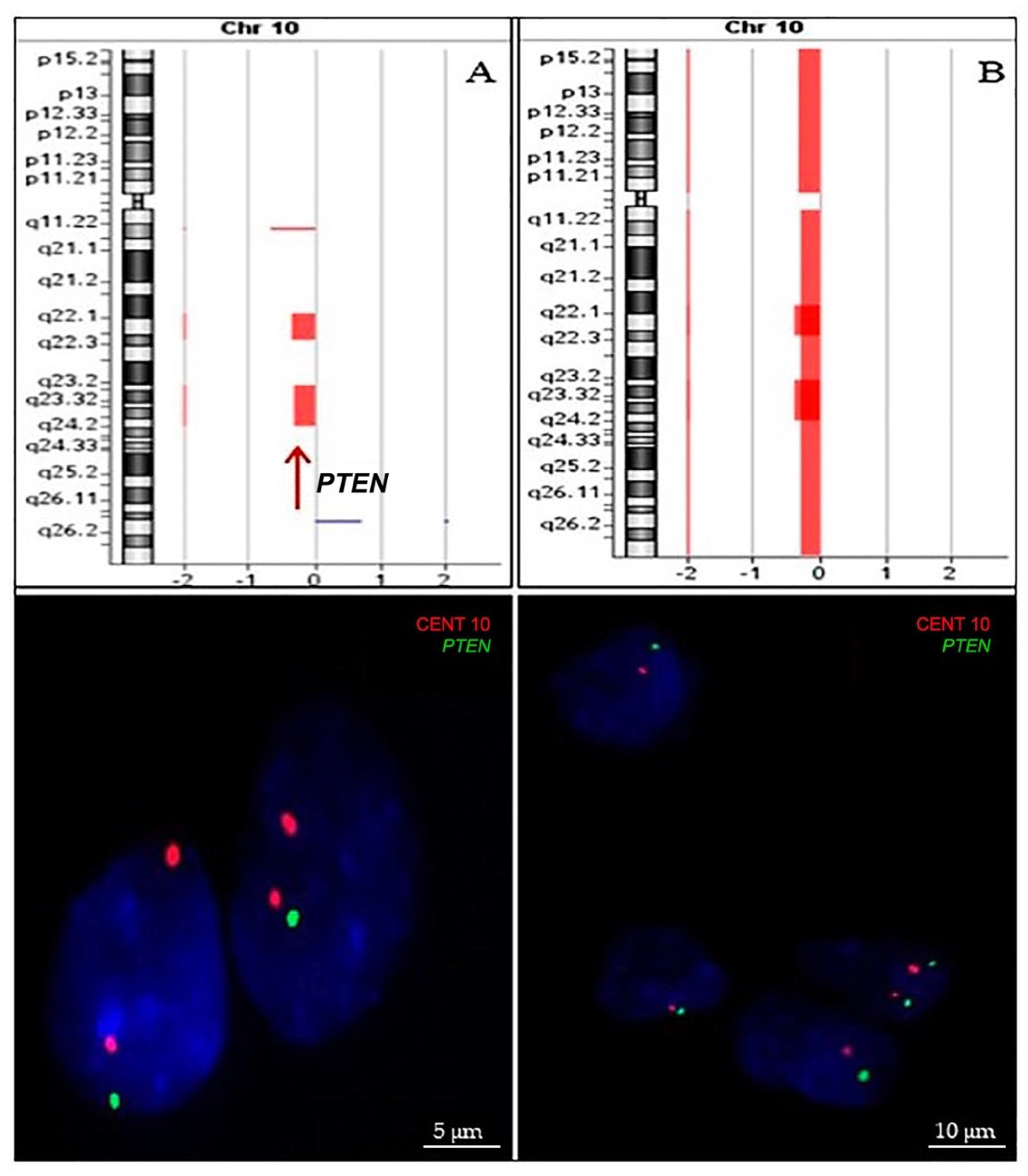
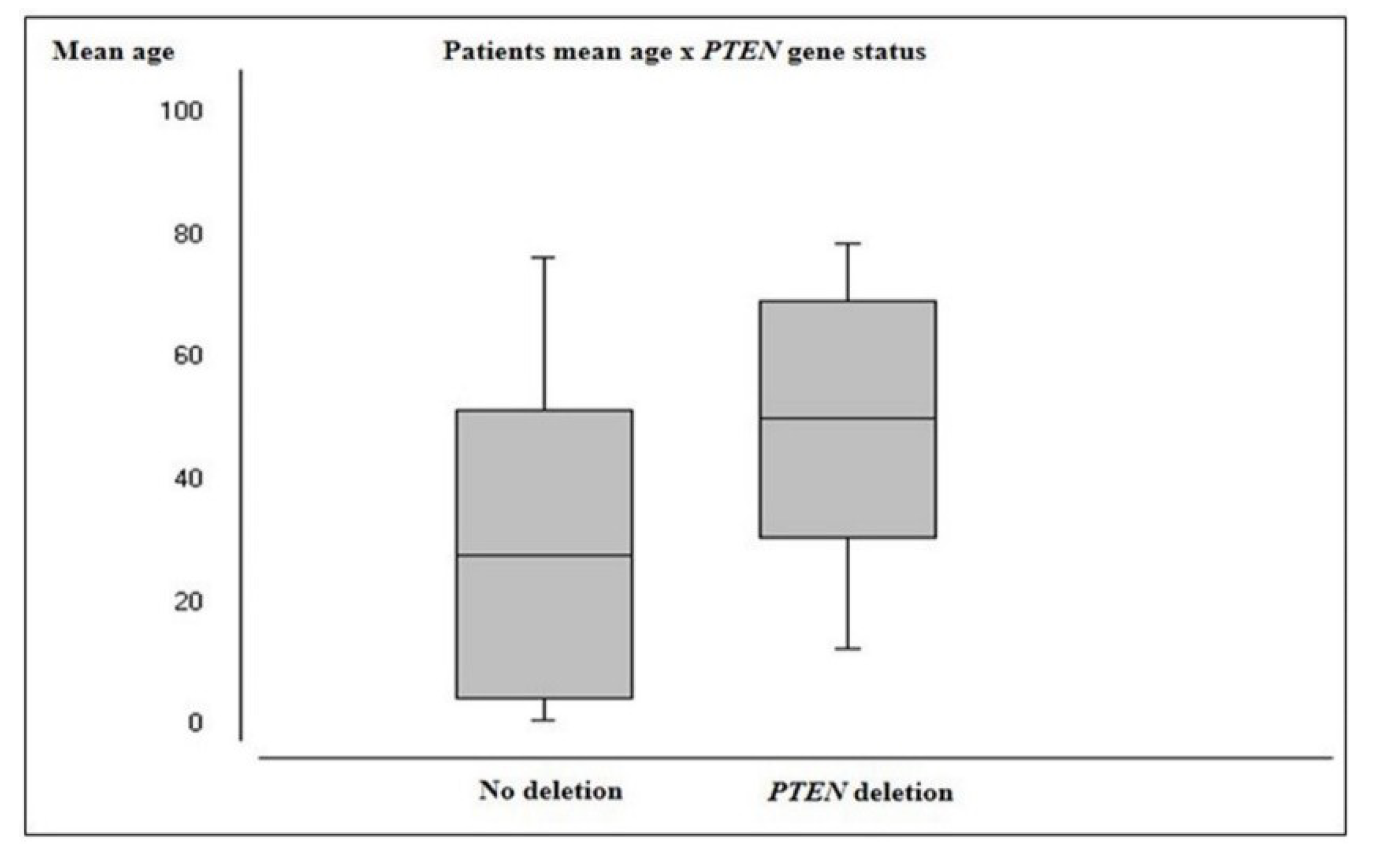
2.2.2. PTEN Status
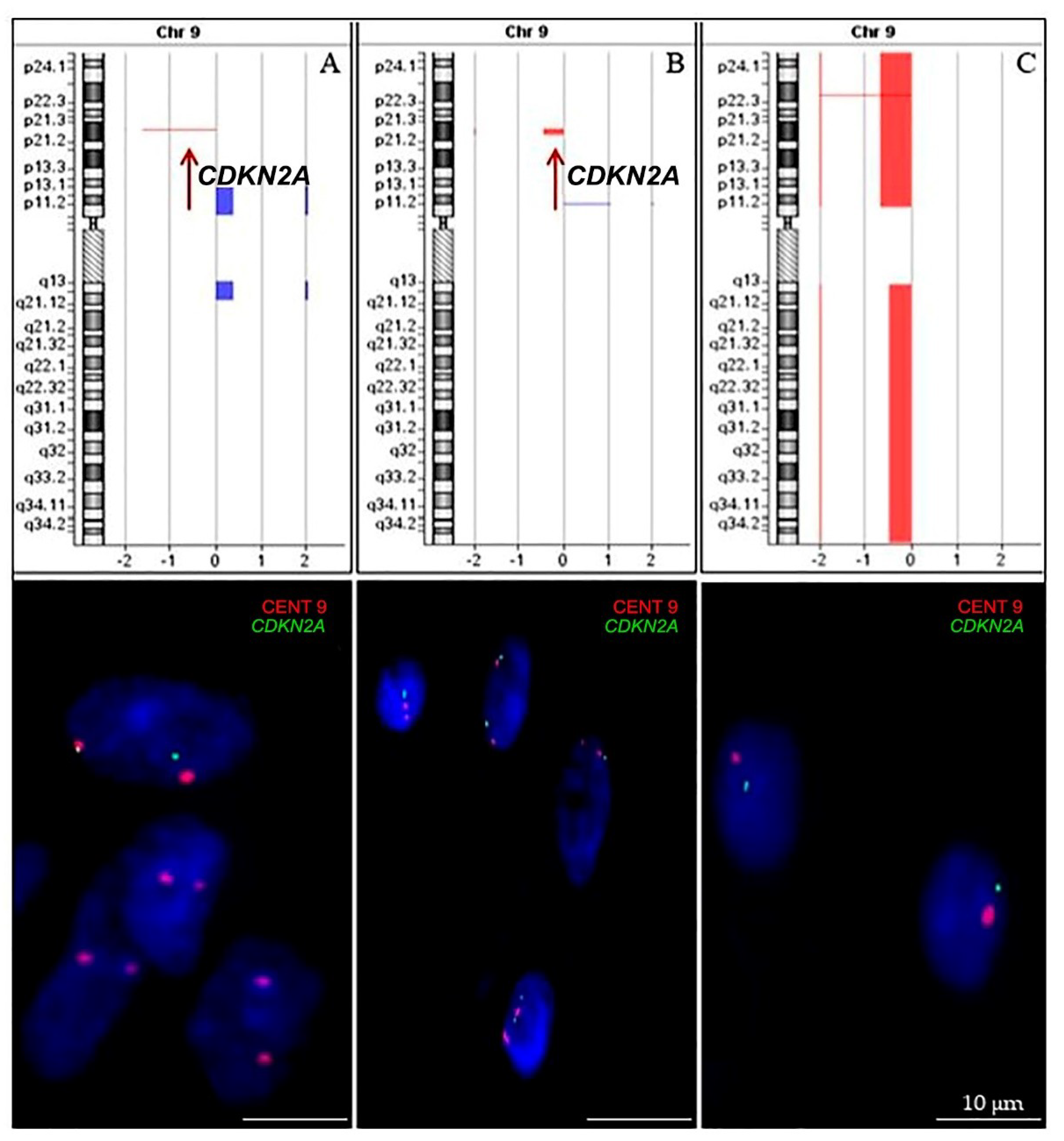
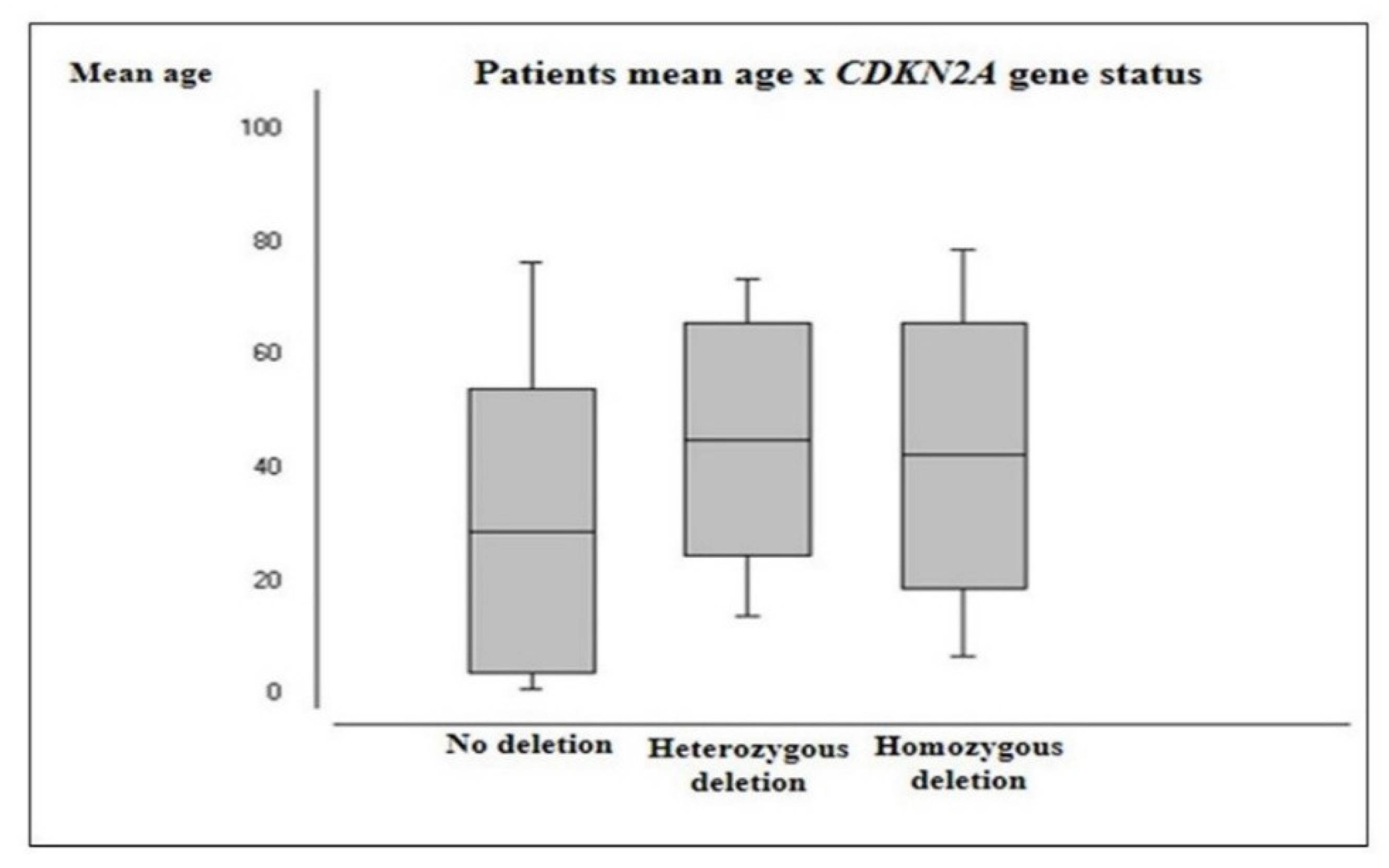
2.2.3. CDKN2A Status
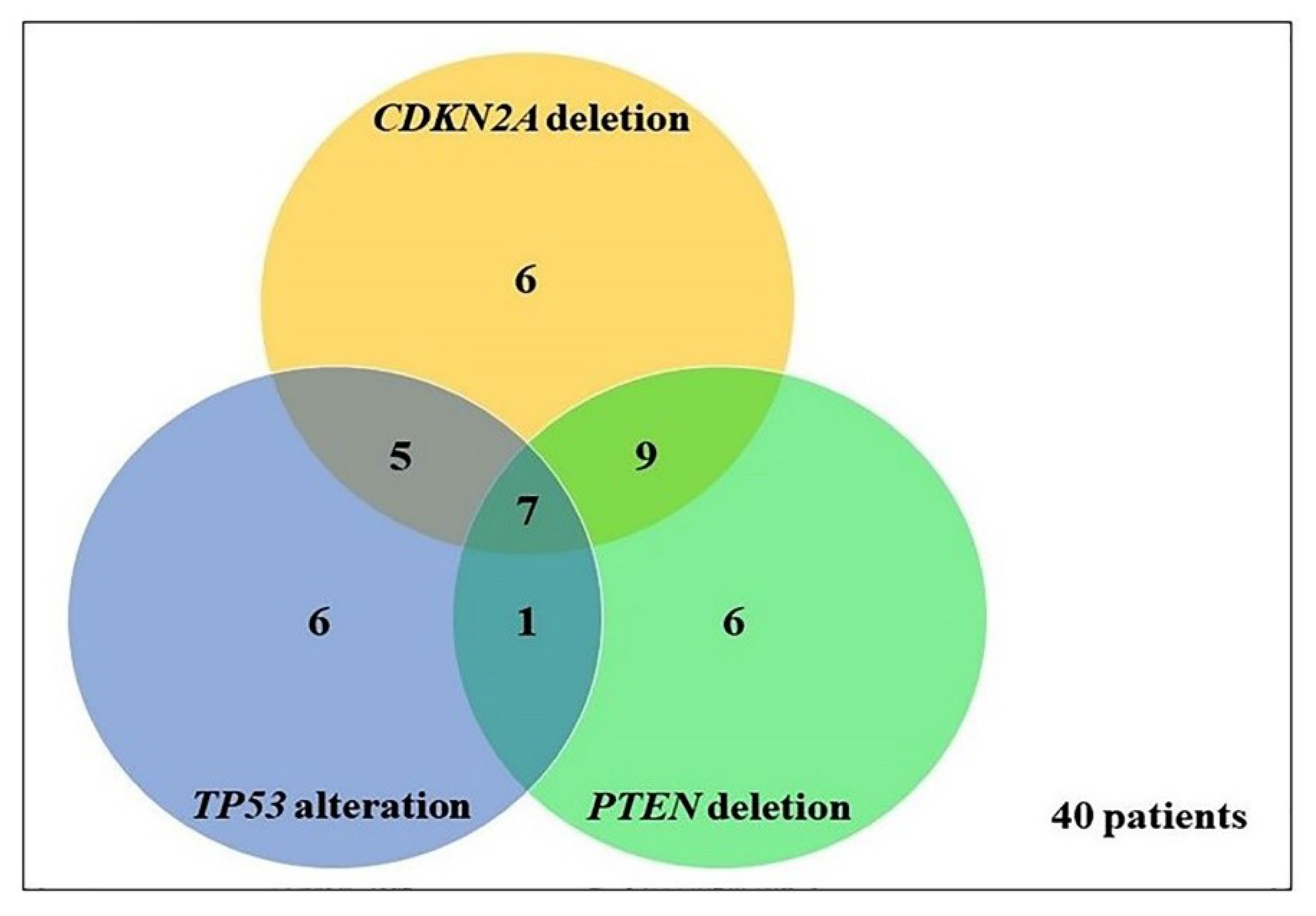
2.3. Frequency and Association between TP53, PTEN, and CDKN2A Genetic Alterations
3. Discussion
4. Materials and Methods
4.1. Patients and Tissue Samples
4.2. DNA Extraction
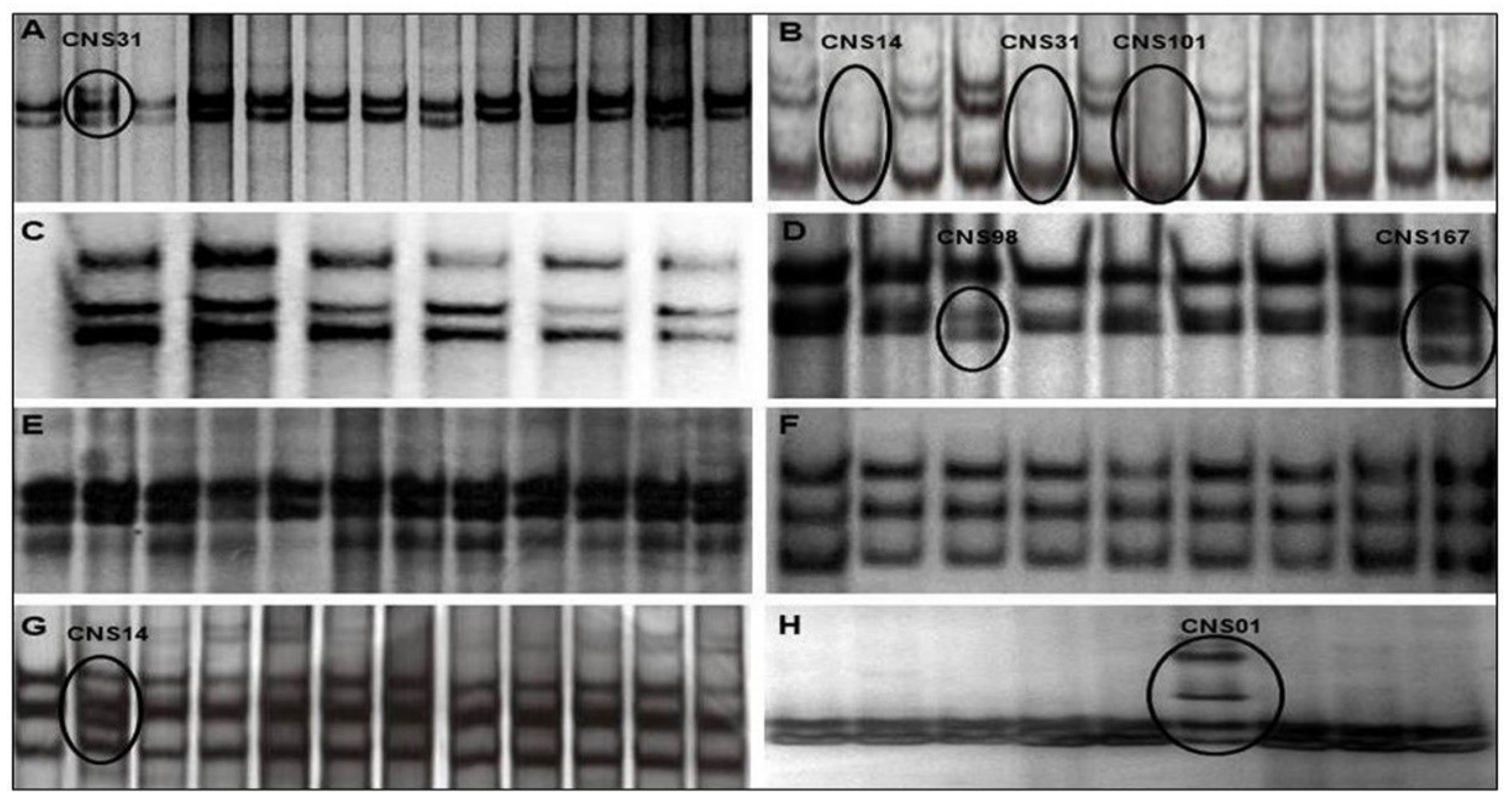
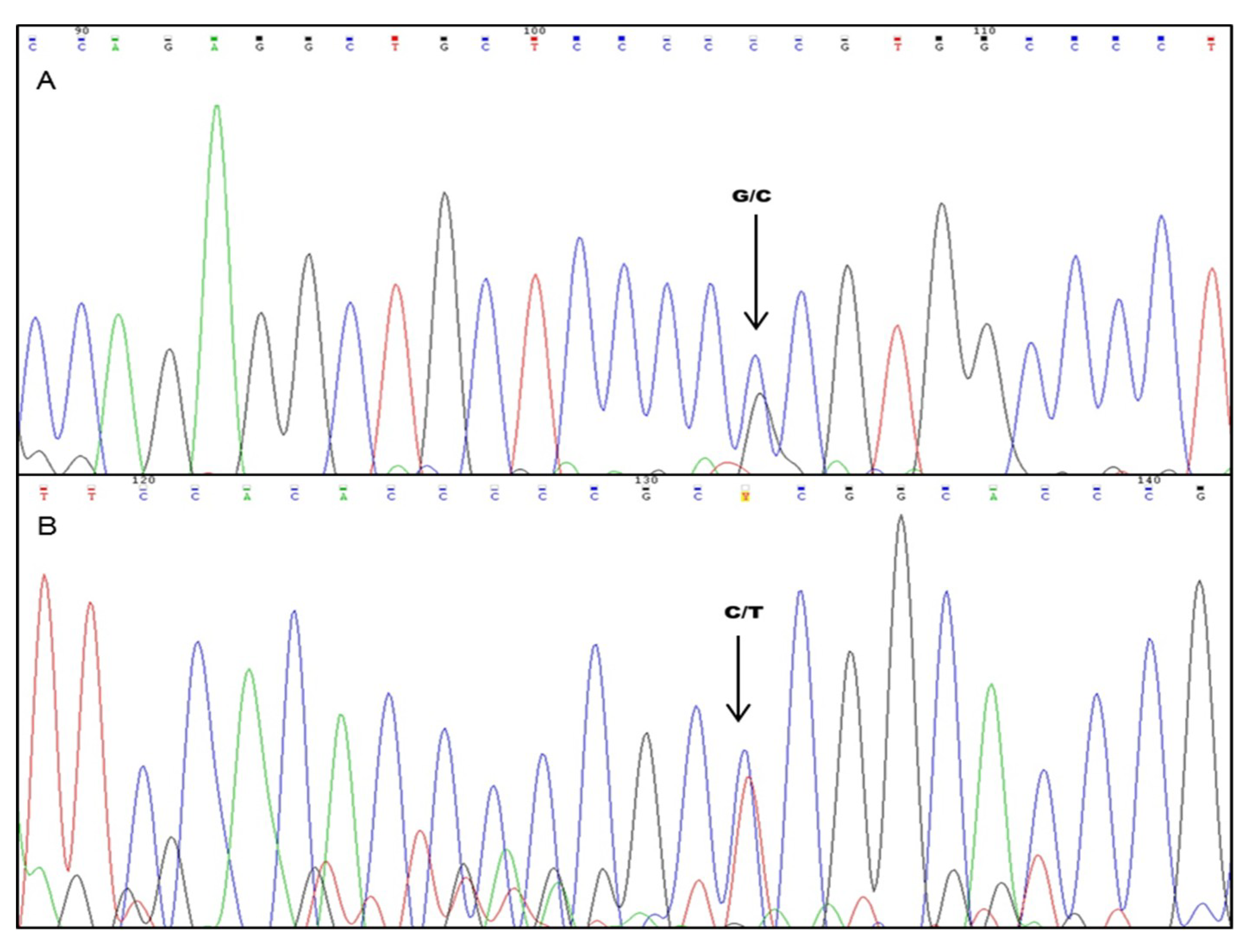
4.3. Single-Strand Conformational Polymorphism (SSCP) Analysis
4.4. Array-Comparative Genomic Hybridization (aCGH)
4.5. Fluorescence In Situ Hybridization (FISH)
4.6. Statistical Analysis
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Persaud-Sharma, D.; Burns, J.; Trangle, J.; Moulik, S. Disparities in Brain Cancer in the United States: A Literature Review of Gliomas. Med. Sci. 2017, 5, 16. [Google Scholar] [CrossRef]
- Alfonso, J.C.L.; Talkenberger, K.; Seifert, M.; Klink, B.; Hawkins-Daarud, A.; Swanson, K.R.; Hatzikirou, H.; Deutsch, A. The biology and mathematical modelling of glioma invasion: A review. J. R. Soc. Interface 2017, 14, 20170490. [Google Scholar] [CrossRef] [PubMed]
- Karsy, M.; Guan, J.; Cohen, A.L.; Jensen, R.L.; Colman, H. New Molecular Considerations for Glioma: IDH, ATRX, BRAF, TERT, H3 K27M. Curr. Neurol. Neurosci. Rep. 2017, 17, 19. [Google Scholar] [CrossRef] [PubMed]
- Massara, M.; Persico, P.; Bonavita, O.; Mollica, P.V.; Locati, M.; Simonelli, M.; Bonecchi, R. Neutrophils in Gliomas. Front. Immunol. 2017, 8, 1349. [Google Scholar] [CrossRef]
- Chen, R.; Smith-Cohn, M.; Cohen, A.L.; Colman, H. Glioma Subclassifications and Their Clinical Significance. Neurotherapeutics 2017, 14, 284–297. [Google Scholar] [CrossRef]
- Louis, D.N.; Perry, A.; Reifenberger, G.; Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A summary. Acta Neuropathol. 2016, 131, 803–820. [Google Scholar] [CrossRef]
- Stankovic, T.; Milinkovic, V.; Bankovic, J.; Dinic, J.; Tanic, N.; Dramicanin, T.; Tanic, N. Comparative analyses of individual and multiple alterations of p53, PTEN and p16 in non-small cell lung carcinoma, glioma and breast carcinoma samples. Biomed. Pharmacother. 2014, 68, 521–526. [Google Scholar] [CrossRef]
- Takami, H.; Yoshida, A.; Fukushima, S.; Arita, H.; Matsushita, Y.; Nakamura, T.; Ohno, M.; Miyakita, Y.; Shibui, S.; Narita, Y.; et al. Revisiting TP53 mutations and immunohistochemistry—A comparative study in 157 diffuse gliomas. Brain Pathol. 2015, 25, 256–265. [Google Scholar] [CrossRef] [PubMed]
- Han, F.; Hu, R.; Yang, H.; Liu, J.; Sui, J.; Xiang, X.; Wang, F.; Chu, L.; Song, S. PTEN gene mutations correlate to poor prognosis in glioma patients: A meta-analysis. Oncol. Targets Ther. 2016, 9, 3485–3492. [Google Scholar]
- Hill, V.K.; Kim, J.S.; James, C.D.; Waldman, T. Correction of PTEN mutations in glioblastoma cell lines via AAV-mediated gene editing. PLoS ONE 2017, 12, e0176683. [Google Scholar] [CrossRef] [PubMed]
- Liggett, W.H.; Sidransky, D. Role of the p16 tumor suppressor gene in cancer. J. Clin. Oncol. 1998, 16, 1197–1206. [Google Scholar] [CrossRef]
- Ruas, M.; Peters, G. The p16INK4a/CDKN2A tumor suppressor and its relatives. Biochim. Biophys. Acta 1998, 1378, F115–F177. [Google Scholar] [CrossRef]
- Quelle, D.E.; Zindy, F.; Ashmun, R.A.; Sherr, C.J. Alternative reading frames of the INK4a tumor suppressor gene encode two unrelated proteins capable of inducing cell cycle arrest. Cell 1995, 83, 993–1000. [Google Scholar]
- Ivanchuk, S.M.; Mondal, S.; Dirks, P.B.; Rutka, J.T. The INK4A/ARF locus: Role in cell cycle control and apoptosis and implications for glioma growth. J. Neurooncol. 2001, 51, 219–229. [Google Scholar] [CrossRef]
- Purkait, S.; Jha, P.; Sharma, M.C.; Suri, V.; Sharma, M.; Kale, S.S.; Sarkar, C. CDKN2A deletion in pediatric versus adult glioblastomas and predictive value of p16 immunohistochemistry. Neuropathology 2013, 33, 405–412. [Google Scholar] [CrossRef]
- Jagannathan, J.; Oskouian, R.J.; Yeoh, H.K.; Saulle, D.; Dumont, A.S. Molecular biology of unreresectable meningiomas: Implications for new treatments and review of the literature. Skull Base 2008, 18, 173–187. [Google Scholar] [CrossRef]
- Nikiforova, M.N.; Wald, A.I.; Melan, M.A.; Roy, S.; Zhong, S.; Hamilton, R.L.; Lieberman, F.S.; Drappatz, J.; Amankulor, N.M.; Pollack, I.F.; et al. Targeted next-generation sequencing panel (GlioSeq) provides comprehensive genetic profiling of central nervous system tumors. Neuro Oncol. 2016, 18, 379–387. [Google Scholar] [CrossRef] [PubMed]
- Zacher, A.; Kaulich, K.; Stepanow, S.; Wolter, M.; Köhrer, K.; Felsberg, J.; Malzkorn, B.; Reifenberger, G. Molecular Diagnostics of Gliomas Using Next Generation Sequencing of a Glioma-Tailored Gene Panel. Brain Pathol. 2017, 27, 146–159. [Google Scholar] [CrossRef] [PubMed]
- Diamandis, P.; Aldape, K.D. Insights from Molecular Profiling of Adult Glioma. J. Clin. Oncol. 2017, 35, 2386–2393. [Google Scholar] [CrossRef]
- Ludwig, K.; Kornblum, H.I. Molecular markers in glioma. J. Neurooncol. 2017, 134, 505–512. [Google Scholar] [CrossRef] [PubMed]
- Jesionek-Kupnicka, D.; Szybka, M.; Malachowska, B.; Fendler, W.; Potemski, P.; Piaskowski, S.; Jaskolski, D.; Papierz, W.; Skowronski, W.; Och, W.; et al. TP53 Promoter Methylation in Primary Glioblastoma: Relationship with TP53 mRNA and Protein Expression and Mutation Status. DNA Cell Biol. 2014, 33, 217–226. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.Y.; Zhang, T.; Li, S.W.; Qian, T.Y.; Fan, X.; Peng, X.X.; Ma, J.; Wang, L.; Jiang, T. Mapping p53 Mutations in Low-Grade Glioma: A Voxel-Based Neuroimaging Analysis. AJNR Am. J. Neuroradiol. 2015, 36, 70–76. [Google Scholar] [CrossRef]
- Halldórsdóttir, A.M.; Lundin, A.; Murray, F.; Mansouri, L.; Knuutila, S.; Sundström, C.; Laurell, A.; Ehrencrona, H.; Sander, B.; Rosenquist, R. Impact of TP53 mutation and 17p deletion in mantle cell lymphoma. Leukemia 2011, 25, 1904–1908. [Google Scholar] [CrossRef]
- Liu, Y.; Chen, C.; Xu, Z.; Scuoppo, C.; Rillahan, C.D.; Gao, J.; Spitzer, B.; Bosbach, B.; Kastenhuber, E.R.; Baslan, T.; et al. Deletions linked to TP53 loss drive cancer through p53-independent mechanisms. Nature 2016, 531, 471–475. [Google Scholar] [CrossRef]
- Malcikova, J.; Smardova, J.; Rocnova, L.; Tichy, B.; Kuglik, P.; Vranova, V.; Cejkova, S.; Svitakova, M.; Skuhrova, F.H.; Brychtova, Y.; et al. Monoallelic and biallelic inactivation of TP53 gene in chronic lymphocytic leukemia: Selection, impact on survival, and response to DNA damage. Blood 2009, 114, 5307–5314. [Google Scholar] [CrossRef] [PubMed]
- Gillet, E.; Alentorn, A.; Doukouré, B.; Mundwiller, E.; van Thuijl, H.F.; Reijneveld, J.C.; Medina, J.A.; Liou, A.; Marie, Y.; Mokhtari, K.; et al. TP53 and p53 statuses and their clinical impact in diffuse low grade gliomas. J. Neurooncol. 2014, 118, 131–139. [Google Scholar] [PubMed]
- Wiencke, J.K.; Zheng, S.; Jelluma, N.; Tihan, T.; Vandenberg, S.; Tamgüney, T.; Baumber, R.; Parsons, R.; Lamborn, K.R.; Berger, M.S.; et al. Methylation of the PTEN promoter defines low-grade gliomas and secondary glioblastoma. Neuro Oncol. 2007, 9, 271–279. [Google Scholar] [CrossRef] [PubMed]
- Cryan, J.B.; Haidar, S.; Ramkissoon, L.A.; Bi, W.L.; Knoff, D.S.; Schultz, N.; Abedalthagafi, M.; Brown, L.; Wen, P.Y.; Reardon, D.A.; et al. Clinical multiplexed exome sequencing distinguishes adult oligodendroglial neoplasms from astrocytic and mixed lineage gliomas. Oncotarget 2014, 5, 8083–8092. [Google Scholar] [CrossRef]
- Yang, Y.; Shao, N.; Luo, G.; Li, L.; Zheng, L.; Nilsson-Ehle, P.; Xu, N. Mutations of PTEN Gene in Gliomas Correlate to Tumor Differentiation and Short-term Survival Rate. Anticancer Res. 2010, 30, 981–985. [Google Scholar]
- Bleeker, F.E.; Lamba, S.; Zanon, C.; Molenaar, R.J.; Hulsebos, T.J.; Troost, D.; van Tilborg, A.A.; Vandertop, W.P.; Leenstra, S.; van Noorden, C.J.; et al. Mutational profiling of kinases in glioblastoma. BMC Cancer 2014, 14, 718. [Google Scholar] [CrossRef]
- Tang, C.; Guo, J.; Chen, H.; Yao, C.J.; Zhuang, D.X.; Wang, Y.; Tang, W.J.; Ren, G.; Yao, Y.; Wu, J.S.; et al. Gene mutation profiling of primary glioblastoma through multiple tumor biopsy guided by 1H-magnetic resonance spectroscopy. Int. J. Clin. Exp. Pathol. 2015, 8, 5327–5335. [Google Scholar]
- Yakut, T.; Gutenberg, A.; Bekar, A.; Egeli, U.; Gunawan, B.; Ercan, I.; Tolunay, S.; Doygun, M.; Schulten, H.J. Correlation of chromosomal imbalances by comparative genomic hybridization and expression of EGFR, PTEN, p53, and MIB-1 in diffuse gliomas. Oncol. Rep. 2007, 17, 1037–1043. [Google Scholar] [CrossRef][Green Version]
- Jeuken, J.W.; Sijben, A.; Bleeker, F.E.; Boots-Sprenger, S.H.; Rijntjes, J.; Gijtenbeek, J.M.; Mueller, W.; Wesseling, P. The Nature and Timing of Specific Copy Number Changes in the Course of Molecular Progression in Diffuse Gliomas: Further Elucidation of Their Genetic “Life Story”. Brain Pathol. 2011, 21, 308–320. [Google Scholar] [CrossRef]
- Srividya, M.R.; Thota, B.; Shailaja, B.C.; Arivazhagan, A.; Thennarasu, K.; Chandramouli, B.A.; Hegde, A.S.; Santosh, V. Homozygous 10q23/PTEN deletion and its impact on outcome in glioblastoma: A prospective translational study on a uniformly treated cohort of adult patients. Neuropathology 2011, 31, 376–383. [Google Scholar] [CrossRef] [PubMed]
- Idbaih, A.; Dalmasso, C.; Kouwenhoven, M.; Jeuken, J.; Carpentier, C.; Gorlia, T.; Kros, J.M.; French, P.; Teepen, J.; Broët, P.; et al. Genomic aberrations associated with outcome in anaplastic oligodendroglial tumors treated within the EORTC phase III trial 26951. J. Neurooncol. 2011, 103, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Sabha, N.; Knobbe, C.B.; Maganti, M.; Omar, S.; Bernstein, M.; Cairns, R.; Çako, B.; von Deimling, A.; Capper, D.; Mak, T.W.; et al. Analysis of IDH mutation,1p/19q deletion, and PTEN loss delineates prognosis in clinical low-grade diffuse gliomas. Neuro Oncol. 2014, 16, 914–923. [Google Scholar] [CrossRef] [PubMed]
- Bidinotto, L.T.; Torrieri, R.; Mackay, A.; Almeida, G.C.; Viana-Pereira, M.; Cruvinel-Carloni, A.; Spina, M.L.; Campanella, N.C.; Pereira de Menezes, W.; Clara, C.A.; et al. Copy Number Profiling of Brazilian Astrocytomas. G3 2016, 6, 1867–1878. [Google Scholar] [CrossRef]
- Smith, J.S.; Tachibana, I.; Passe, S.M.; Huntley, B.K.; Borell, T.J.; Iturria, N.; O’Fallon, J.R.; Schaefer, P.L.; Scheithauer, B.W.; James, C.D.; et al. PTEN Mutation, EGFR Amplification, and Outcome in Patients with Anaplastic Astrocytoma and Glioblastoma Multiforme. J. Natl. Cancer Inst. 2001, 93, 1246–1256. [Google Scholar] [CrossRef]
- Dubbink, H.J.; Atmodimedjo, P.N.; Kros, J.M.; French, P.J.; Sanson, M.; Idbaih, A.; Wesseling, P.; Enting, R.; Spliet, W.; Tijssen, C.; et al. Molecular classification of anaplastic oligodendroglioma using next-generation sequencing: A report of the prospective randomized EORTC Brain Tumor Group 26951 phase III trial. Neuro Oncol. 2016, 18, 388–400. [Google Scholar] [CrossRef]
- Pollack, I.F.; Hamilton, R.L.; James, C.D.; Finkelstein, S.D.; Burnham, J.; Yates, A.J.; Holmes, E.J.; Zhou, T.; Finlay, J.L.; Children’s Oncology Group. Rarity of PTEN deletions and EGFR amplification in malignant gliomas of childhood: Results from the Children’s Cancer Group 945 cohort. J. Neurosurg. 2006, 105, 418–424. [Google Scholar]
- Purkait, S.; Sharma, V.; Jha, P.; Sharma, M.C.; Suri, V.; Suri, A.; Sharma, B.S.; Sarkar, C. EZH2 expression in gliomas: Correlation with CDKN2A gene deletion/ p16 loss and MIB-1 proliferation index. Neuropathology 2015, 35, 421–431. [Google Scholar] [CrossRef]
- Reis, G.F.; Pekmezci, M.; Hansen, H.M.; Rice, T.; Marshall, R.E.; Molinaro, A.M.; Phillips, J.J.; Vogel, H.; Wiencke, J.K.; Wrensch, M.R.; et al. CDKN2A Loss Is Associated with Shortened Overall Survival in Lower Grade (World Health Organization II-III) Astrocytomas. J. Neuropathol. Exp. Neurol. 2015, 74, 442–452. [Google Scholar] [CrossRef]
- Kogiso, M.; Qi, L.; Lindsay, H.; Huang, Y.; Zhao, X.; Liu, Z.; Braun, F.K.; Du, Y.; Zhang, H.; Bae, G.; et al. Xenotransplantation of pediatric low grade gliomas confirms the enrichment of BRAF V600E mutation and preservation of CDKN2A deletion in a novel orthotopic xenograft mouse model of progressive pleomorphic xanthoastrocytoma. Oncotarget 2017, 8, 87455–87471. [Google Scholar] [CrossRef]
- Fan, Y.; Yue, J.; Xiao, M.; Han-Zhang, H.; Wang, Y.V.; Ma, C.; Deng, Z.; Li, Y.; Yu, Y.; Wang, X.; et al. FXR1 regulates transcription and is required for growth of human cancer cells with TP53/FXR2 homozygous deletion. Elife 2017, 6, e26129. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, N.; Nobusawa, S.; Nakata, S.; Nakada, M.; Yamazaki, T.; Matsumura, N.; Harada, K.; Matsuda, H.; Funata, N.; Nagai, S.; et al. BRAF V600E, TERT promoter mutations and CDKN2A/B homozygous deletions are frequent in epithelioid glioblastomas: A histological and molecular analysis focusing on intratumoral heterogeneity. Brain Pathol. 2018, 28, 663–673. [Google Scholar] [CrossRef] [PubMed]
- Knudson, A.G. Two genetic hits (more or less) to cancer). Nat. Rev. Cancer 2001, 1, 157–162. [Google Scholar] [CrossRef]
- Picanço-Albuquerque, C.G.; Morais, C.L.; Carvalho, F.L.; Peskoe, S.B.; Hicks, J.L.; Ludkovski, O.; Vidotto, T.; Fedor, H.; Humphreys, E.; Han, M.; et al. In prostate cancer needle biopsies, detections of PTEN loss by fluorescence in situ hybridization (FISH) and by immunohistochemistry (IHC) are concordant and show consistent association with upgrading. Virchows Arch. 2016, 468, 607–617. [Google Scholar] [CrossRef]
- Vidotto, T.; Tiezzi, D.G.; Squire, J.A. Distinct subtypes of genomic PTEN deletion size influence the landscape of aneuploidy and outcome in prostate cancer. Mol. Cytogenet. 2018, 11, 1. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Huang, C.; He, J.; Lamb, K.L.; Kang, X.; Gu, T.; Shen, W.H.; Yin, Y. PTEN C-terminal deletion causes genomic instability and tumor development. Cell Rep. 2014, 6, 844–854. [Google Scholar] [CrossRef] [PubMed]
- Kluth, M.; Harasimowicz, S.; Burkhardt, L.; Grupp, K.; Krohn, A.; Prien, K.; Gjoni, J.; Haß, T.; Galal, R.; Graefen, M.; et al. Clinical significance of different types of p53 gene alteration in surgically treated prostate cancer. Int. J. Cancer. 2014, 135, 1369–1380. [Google Scholar] [CrossRef]
- Mistry, M.; Zhukova, N.; Merico, D.; Rakopoulos, P.; Krishnatry, R.; Stavropoulos, M.S.; Alon, N.; Pole, J.D.; Ray, P.N.; Navickiene, V.; et al. BRAF Mutation and CDKN2A Deletion Define a Clinically Distinct Subgroup of Childhood Secondary High-Grade Glioma. J. Clin. Oncol. 2015, 9, 1015–1022. [Google Scholar] [CrossRef] [PubMed]
- Kato, H.; Kato, S.; Kumabe, T.; Sonoda, Y.; Yoshimoto, T.; Kato, S.; Han, S.Y.; Suzuki, T.; Shibata, H.; Kanamaru, R.; et al. Functional evaluation of p53 and pten gene mutation in gliomas. Clin. Cancer Res. 2000, 6, 3937–3943. [Google Scholar] [PubMed]
- Riemenschneider, M.J.; Jeuken, J.W.; Wesseling, P.; Reifenberger, G. Molecular diagnostics of gliomas: State of the art. Acta Neuropathol. 2010, 120, 567–584. [Google Scholar] [CrossRef] [PubMed]
| WHO Histological Grading | Mean Age | Significance (p Value) |
|---|---|---|
| I | 11.8 | Median I and II (p > 0.05) Median I and III (p < 0.05) Median I and IV (p < 0.05) |
| II | 23.6 | Median II and III (p < 0.05) Median II and IV (p < 0.05) |
| III | 42.8 | Median III and IV (p > 0.05) |
| IV | 51.7 | - |
| Sample | Tumor | WHO Grading | Gender | Age | Exon |
|---|---|---|---|---|---|
| CNS 1 | Subependymal giant cell astrocytoma | I | M | 23 | 11 |
| CNS 14 | Glioblastoma | IV | F | 68 | 5, 10 |
| CNS 31 | Analastic astrocytoma | III | F | 65 | 4, 5 |
| CNS 98 | Diffuse astrocytoma | II | M | 26 | 7 |
| CNS 101 | Glioblastoma | IV | M | 79 | 5 |
| CNS 167 | Diffuse astrocytoma | II | M | 14 | 7 |
| TP53 | All Cases | A-I | A-II | A-III | A-IV | OD-II | OD-III | OA-II | OA-III | ED-II | ED-III | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| N | % | N | % | N | % | N | % | N | % | N | % | N | % | N | % | N | % | N | % | N | % | |
| Intact | 46 | 70.8 | 5 | 55.6 | 9 | 90 | 4 | 66.7 | 16 | 66.7 | 3 | 100 | 0 | 0 | 2 | 100 | 1 | 50 | 3 | 75 | 3 | 75 |
| Hom. Del | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Het. Del | 19 | 29.2 | 4 | 44.4 | 1 | 10 | 2 | 33.3 | 8 | 33.3 | 0 | 0 | 1 | 100 | 0 | 0 | 1 | 50 | 1 | 25 | 1 | 25 |
| Total | 65 | 9 | 10 | 6 | 24 | 3 | 1 | 2 | 2 | 4 | 4 | |||||||||||
| PTEN | All Cases | A-I | A-II | A-III | A-IV | OD-II | OD-III | OA-II | OA-III | ED-II | ED-III | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| N | % | N | % | N | % | N | % | N | % | N | % | N | % | N | % | N | % | N | % | N | % | |
| Intact | 39 | 62.9 | 8 | 100 | 7 | 77.8 | 5 | 83.3 | 6 | 26.1 | 2 | 66.7 | 0 | 0 | 2 | 100 | 1 | 50 | 4 | 100 | 4 | 100 |
| Hom. Del | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Het. Del | 23 | 37.1 | 0 | 0 | 2 | 22.2 | 1 | 16.7 | 17 | 73.9 | 1 | 33.3 | 1 | 100 | 0 | 0 | 1 | 50 | 0 | 0 | 0 | 0 |
| Total | 62 | 8 | 9 | 6 | 23 | 3 | 1 | 2 | 2 | 4 | 4 | |||||||||||
| CDKN2A | All Cases | A-I | A-II | A-III | A-IV | OD-II | OD-III | OA-II | OA-III | ED-II | ED-III | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| N | % | N | % | N | % | N | % | N | % | N | % | N | % | N | % | N | % | N | % | N | % | |
| Intact | 35 | 55.6 | 6 | 75 | 4 | 44.4 | 3 | 50 | 10 | 41.7 | 2 | 66.7 | 0 | 0 | 2 | 100 | 1 | 50 | 4 | 100 | 3 | 75 |
| Hom. Del | 8 | 12.7 | 0 | 0 | 2 | 22.2 | 1 | 16.7 | 5 | 20.8 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Het. Del | 20 | 31.7 | 2 | 25 | 3 | 33.3 | 2 | 33.3 | 9 | 37.5 | 1 | 33.3 | 1 | 100 | 0 | 0 | 1 | 50 | 0 | 0 | 1 | 25 |
| Total | 63 | 8 | 9 | 6 | 24 | 3 | 1 | 2 | 2 | 4 | 4 | |||||||||||
| Gene Alterations | WHO Grading | |||
|---|---|---|---|---|
| Grade I | Grade II | Grade III | Grade IV | |
| One Gene | ||||
| TP53 */PTEN/CDKN2A | 1/8 (12.5%) | 1/18 (5.6%) | 2/13 (15.4%) | 2/23 (8.7%) |
| TP53/PTEN */CDKN2A | - | 1/18 (5.6%) | - | 5/23 (21.7%) |
| TP53/PTEN/CDKN2A* | - | 4/18 (22.2%) | 1/13 (7.7%) | 1/23 (4.3%) |
| Two Genes | ||||
| TP53*/PTEN */CDKN2A | - | - | - | 1/23 (4.3%) |
| TP53*/PTEN/CDKN2A* | 2/8 (25%) | - | 2/13 (15.4%) | 1/23 (4.3%) |
| TP53/PTEN */CDKN2A* | - | 1/18 (5.6%) | 2/13 (15.4%) | 6/23 (26.1%) |
| Three Genes | ||||
| TP53*/PTEN */CDKN2A* | - | 1/18 (5.6%) | 1/13 (7.7%) | 5/23 (21.7%) |
| Patient | Histopathologic | WHO | Sex | Age | SSCP | aCGH + iFISH | |||
|---|---|---|---|---|---|---|---|---|---|
| TP53 | PTEN | TP53 | PTEN | CDKN2A | |||||
| 1 | SGCA | 1 | M | 23 | mut | wt | x | x | x |
| 2 | DA | 2 | F | 12 | wt | wt | - | x | x |
| 3 | PA | 1 | M | 13 | wt | x | x | x | x |
| 4 | GBM | 4 | F | 68 | mut | x | no del | no del | no del |
| 5 | GBM | 4 | F | 64 | wt | x | - | - | - - |
| 6 | PMA | 2 | M | 3 | wt | wt | no del | no del | no del |
| 7 | GBM | 4 | M | 65 | wt | x | - | - | no del |
| 8 | OA | 2 | M | 2 | wt | wt | no del | no del | no del |
| 9 | AA | 3 | F | 55 | mut | wt | - | no del | - |
| 10 | GBM | 4 | F | 7 | wt | x | no del | x | - - |
| 11 | ED | 2 | F | 2 | x | x | - | no del | no del |
| 12 | AA | 3 | M | 60 | wt | wt | no del | no del | - |
| 13 | GSA | 4 | M | 64 | wt | wt | no del | - | no del |
| 14 | DA | 2 | M | 37 | wt | x | no del | no del | - |
| 15 | GBM | 4 | M | 43 | wt | wt | no del | no del | no del |
| 16 | ED | 2 | F | 1 | wt | wt | no del | no del | no del |
| 17 | DA | 2 | F | 52 | wt | x | no del | no del | no del |
| 18 | AOA | 3 | F | 63 | wt | wt | - | no del | no del |
| 19 | ED | 2 | M | 8 | wt | wt | no del | no del | no del |
| 20 | OD | 2 | M | 19 | wt | wt | no del | no del | no del |
| 21 | OD | 2 | F | 43 | wt | wt | no del | - | no del |
| 22 | DA | 2 | M | 26 | mut | x | x | x | x |
| 23 | GBM | 4 | M | 79 | mut | x | - | - | - - |
| 24 | AED | 3 | F | 54 | wt | wt | - | no del | no del |
| 25 | GBM | 4 | M | 15 | wt | x | - | - | - |
| 26 | SGCA | 1 | M | 20 | wt | x | - | no del | - |
| 27 | AA | 3 | F | 23 | x | x | no del | no del | no del |
| 28 | PA | 1 | F | 16 | wt | wt | - | no del | - |
| 29 | AOD | 3 | M | 61 | x | x | - | - | - |
| 30 | PXA | 2 | M | 26 | wt | wt | no del | no del | - - |
| 31 | PA | 1 | F | 3 | wt | x | - | x | x |
| 32 | PMA | 2 | F | 2 | x | x | no del | no del | no del |
| 33 | GBM | 4 | F | 70 | wt | wt | no del | - | - |
| 34 | DA | 2 | F | 34 | wt | wt | no del | no del | no del |
| 35 | GBM | 4 | M | 51 | wt | wt | no del | - | - |
| 36 | DA | 2 | M | 64 | wt | x | x | x | x |
| 37 | ED | 2 | F | 8 | wt | wt | no del | no del | no del |
| 38 | AED | 3 | M | 62 | wt | wt | no del | no del | - |
| 39 | PA | 1 | F | 27 | wt | wt | no del | no del | no del |
| 40 | AA | 3 | F | 31 | wt | wt | - | no del | - - |
| 41 | GBM | 4 | M | 60 | wt | wt | no del | - | - - |
| 42 | GBM | 4 | M | 43 | wt | wt | - | no del | - - |
| 43 | GBM | 4 | F | 13 | x | x | no del | - | no del |
| 44 | DA | 2 | M | 14 | mut | x | no del | - | - |
| 45 | GBM | 4 | M | 46 | x | x | no del | - | - |
| 46 | GBM | 4 | F | 49 | x | x | no del | - | no del |
| 47 | OD | 2 | F | 23 | x | x | no del | no del | - |
| 48 | AA | 3 | M | 59 | x | x | no del | - | - |
| 49 | GBM | 4 | M | 56 | x | x | - | - | - |
| 50 | GBM | 4 | M | 74 | wt | wt | no del | no del | - |
| 51 | GBM | 4 | M | 48 | wt | wt | no del | - | no del |
| 52 | GBM | 4 | M | 32 | x | x | - | - | - |
| 53 | PA | 1 | M | 7 | wt | wt | no del | no del | no del |
| 54 | AED | 3 | M | 19 | wt | wt | no del | no del | no del |
| 55 | PA | 1 | F | 9 | x | x | no del | no del | no del |
| 56 | GSA | 4 | M | 77 | wt | wt | - | no del | no del |
| 57 | PA | 1 | M | 2 | wt | wt | no del | no del | no del |
| 58 | DA | 2 | M | 65 | wt | wt | no del | - | - |
| 59 | AOA | 3 | M | 24 | wt | wt | no del | - | - |
| 60 | AA | 3 | M | 58 | x | x | no del | no del | no del |
| 61 | OA | 2 | M | 3 | wt | wt | no del | no del | no del |
| 62 | DA | 2 | M | 31 | x | x | no del | no del | - - |
| 63 | GBM | 4 | M | 33 | x | x | no del | no del | no del |
| 64 | PA | 1 | F | 4 | x | x | no del | no del | no del |
| 65 | AED | 3 | M | 2 | x | x | no del | no del | no del |
| 66 | PA | 1 | F | 6 | x | x | - | no del | no del |
| 67 | GBM | 4 | F | 66 | x | x | no del | - | - |
| 68 | GBM | 4 | M | 43 | x | x | no del | - | no del |
| 69 | GBM | 4 | F | 74 | x | x | no del | - | no del |
| Exon | Primer | Primer Lenght (bp) | Exon Lenght (pb) | Annealing Temperature (°C) |
|---|---|---|---|---|
| Exon 4 | F-5′ TTTCACCCATCTACAGTCCC 3′ R-5′ CATTGAAGTCTCATGGAAGC 3′ | 20 20 | 279 | 54 |
| Exon 5 | F-5′ TTCACTTGTGCCCTGACTT 3′ R-5′ AACCAGCCCTGTCCGTCTC 3′ | 19 19 | 184 | 57 |
| Exon 6 | F-5′ CAGGGCTGGTTGCCCAGGGTCCCCC 3′ R-5′ ACTGACAACCACCCTTAACCCCTCC 3′ | 25 25 | 113 | 59 |
| Exon 7 | F-5′ TGCTTGCCACAGGTCT 3′ R-5′ ACAGCAGGCAGTGT 3′ | 16 14 | 110 | 54 |
| Exon 8 | F-5′ CCACCGCTTCTTGTCCTGC 3′ R-5′ CCTACTGCCTCTTTGCTTC 3′ | 19 19 | 137 | 58 |
| Exon 9 | F-5′ AGGGTGCAGTTATGCCTCAG 3′ R-5′ ACTTGATAAGAGGTCC 3′ | 20 16 | 74 | 52 |
| Exon 10 | F-5′ CTGAGGCACAAGAATCAC 3′ R-5′ TCCTATGGCTTTCCAACC 3′ | 18 18 | 107 | 59 |
| Exon 11 | F-5′ GGAAGGGTCAACATCTTTTACA 3′ R–5′ TAAAAAGGGAGAAGGAGGGG 3′ | 22 20 | 1289 | 59 |
| Exon | Primer | Primer Lenght (bp) | Exon Lenght (pb) | Annealing Temperature (°C) |
|---|---|---|---|---|
| Exon 5 | F-5′ ACCTGTTAAGTTTGTATGCAAC 3′ R-5′ TCTGTTTTCCAATAAATTCTC 3′ | 22 21 | 239 | 51 |
| Exon 6 | F-5′ CTACGACCCAGTTACCATAGCA 3′ R-5′ GGCTTCTTTAGCCCAATGAGTTG 3′ | 22 23 | 142 | 59 |
| Exon 7 | F-5′ TGACAGTTTGACAGTTAAAGG 3′ R-5′ CTCCCAATGAAAGTAAAGTACA 3′ | 21 22 | 167 | 53 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pessôa, I.A.; Amorim, C.K.; Ferreira, W.A.S.; Sagica, F.; Brito, J.R.; Othman, M.; Meyer, B.; Liehr, T.; de Oliveira, E.H.C. Detection and Correlation of Single and Concomitant TP53, PTEN, and CDKN2A Alterations in Gliomas. Int. J. Mol. Sci. 2019, 20, 2658. https://doi.org/10.3390/ijms20112658
Pessôa IA, Amorim CK, Ferreira WAS, Sagica F, Brito JR, Othman M, Meyer B, Liehr T, de Oliveira EHC. Detection and Correlation of Single and Concomitant TP53, PTEN, and CDKN2A Alterations in Gliomas. International Journal of Molecular Sciences. 2019; 20(11):2658. https://doi.org/10.3390/ijms20112658
Chicago/Turabian StylePessôa, Igor Andrade, Carolina Koury Amorim, Wallax Augusto Silva Ferreira, Fernanda Sagica, José Reginaldo Brito, Moneeb Othman, Britta Meyer, Thomas Liehr, and Edivaldo Herculano C. de Oliveira. 2019. "Detection and Correlation of Single and Concomitant TP53, PTEN, and CDKN2A Alterations in Gliomas" International Journal of Molecular Sciences 20, no. 11: 2658. https://doi.org/10.3390/ijms20112658
APA StylePessôa, I. A., Amorim, C. K., Ferreira, W. A. S., Sagica, F., Brito, J. R., Othman, M., Meyer, B., Liehr, T., & de Oliveira, E. H. C. (2019). Detection and Correlation of Single and Concomitant TP53, PTEN, and CDKN2A Alterations in Gliomas. International Journal of Molecular Sciences, 20(11), 2658. https://doi.org/10.3390/ijms20112658







