TCO, a Putative Transcriptional Regulator in Arabidopsis, Is a Target of the Protein Kinase CK2
Abstract
1. Introduction
2. Results
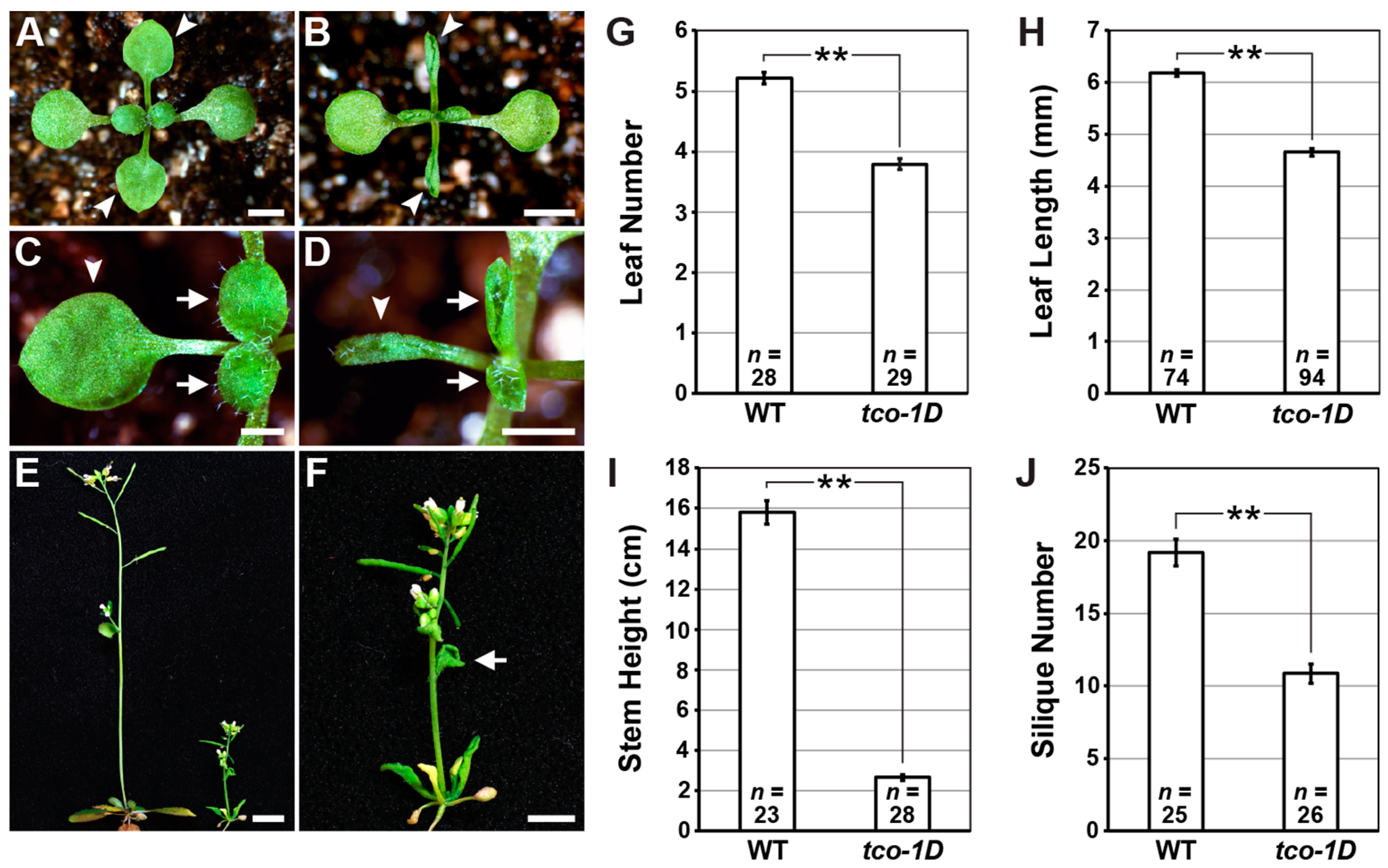
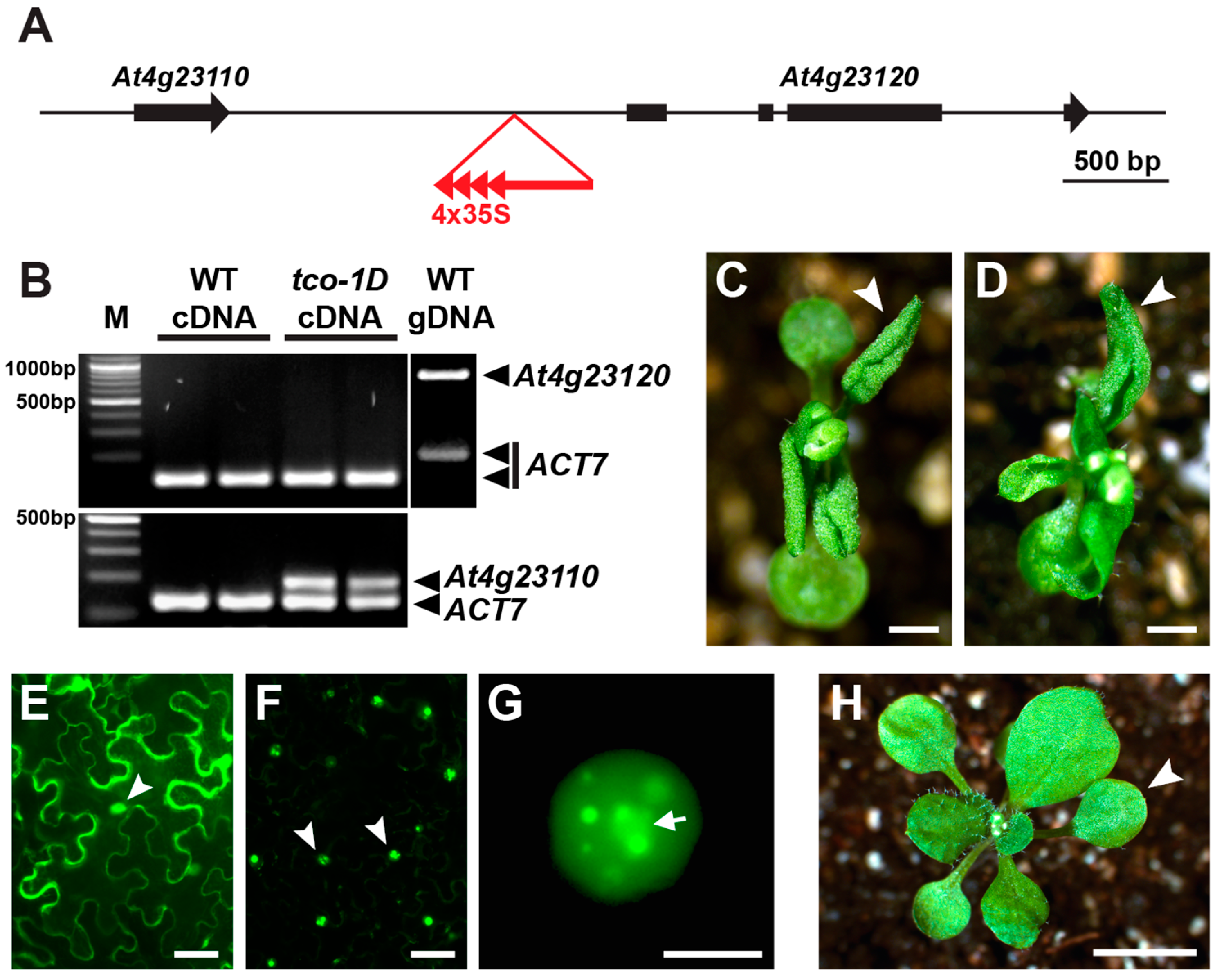
2.1. Identification and Characterization of the Leaf-Curling Mutant Taco Leaf-1D
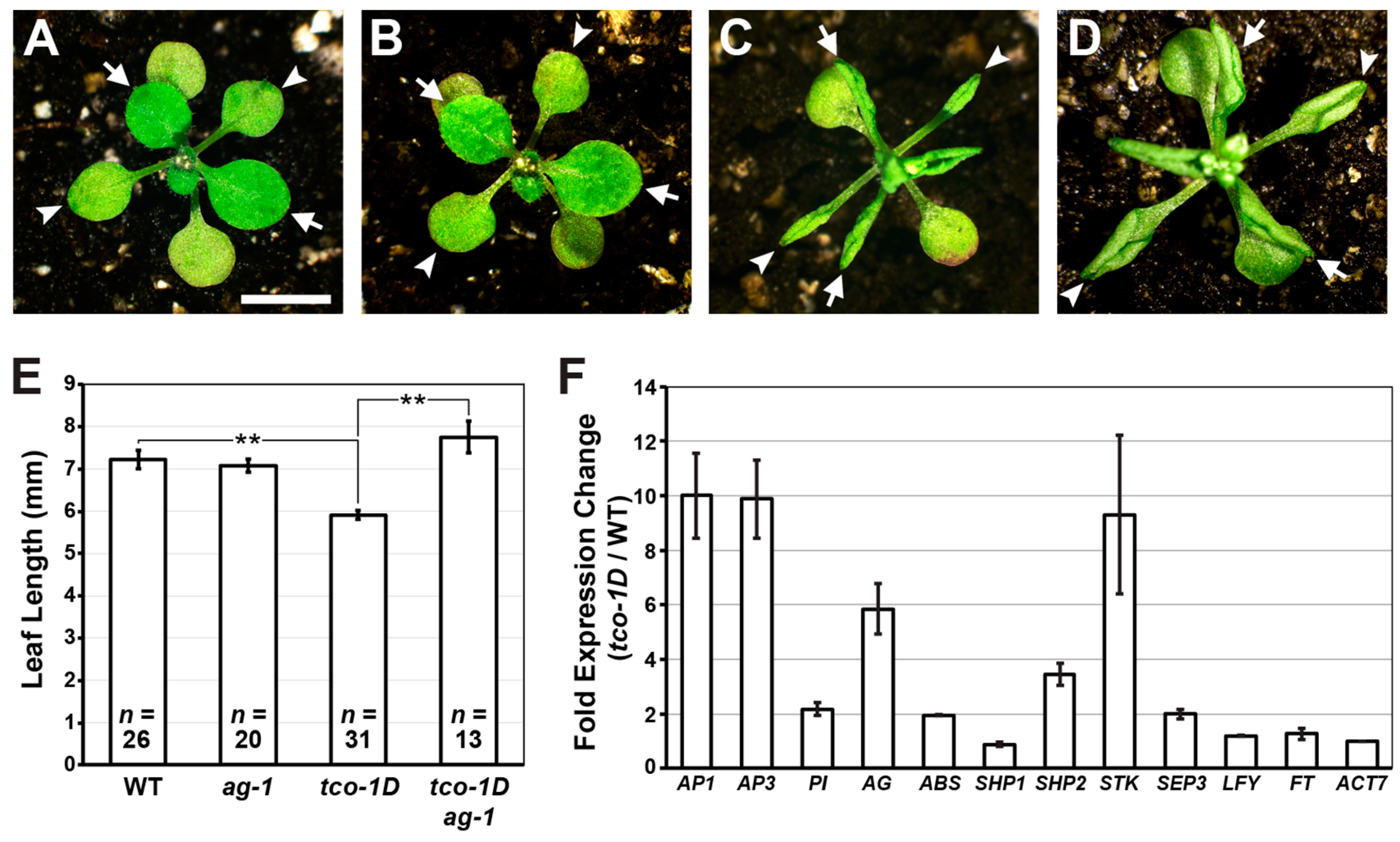
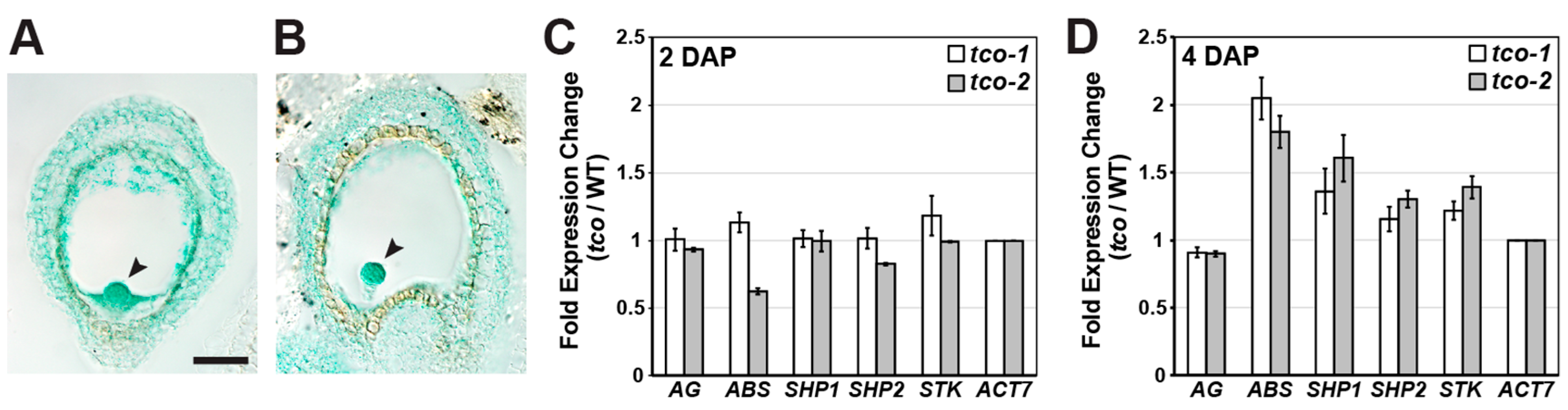
2.2. Reproductive Genes Are Ectopically Expressed in tco-1D Vegetative Tissues
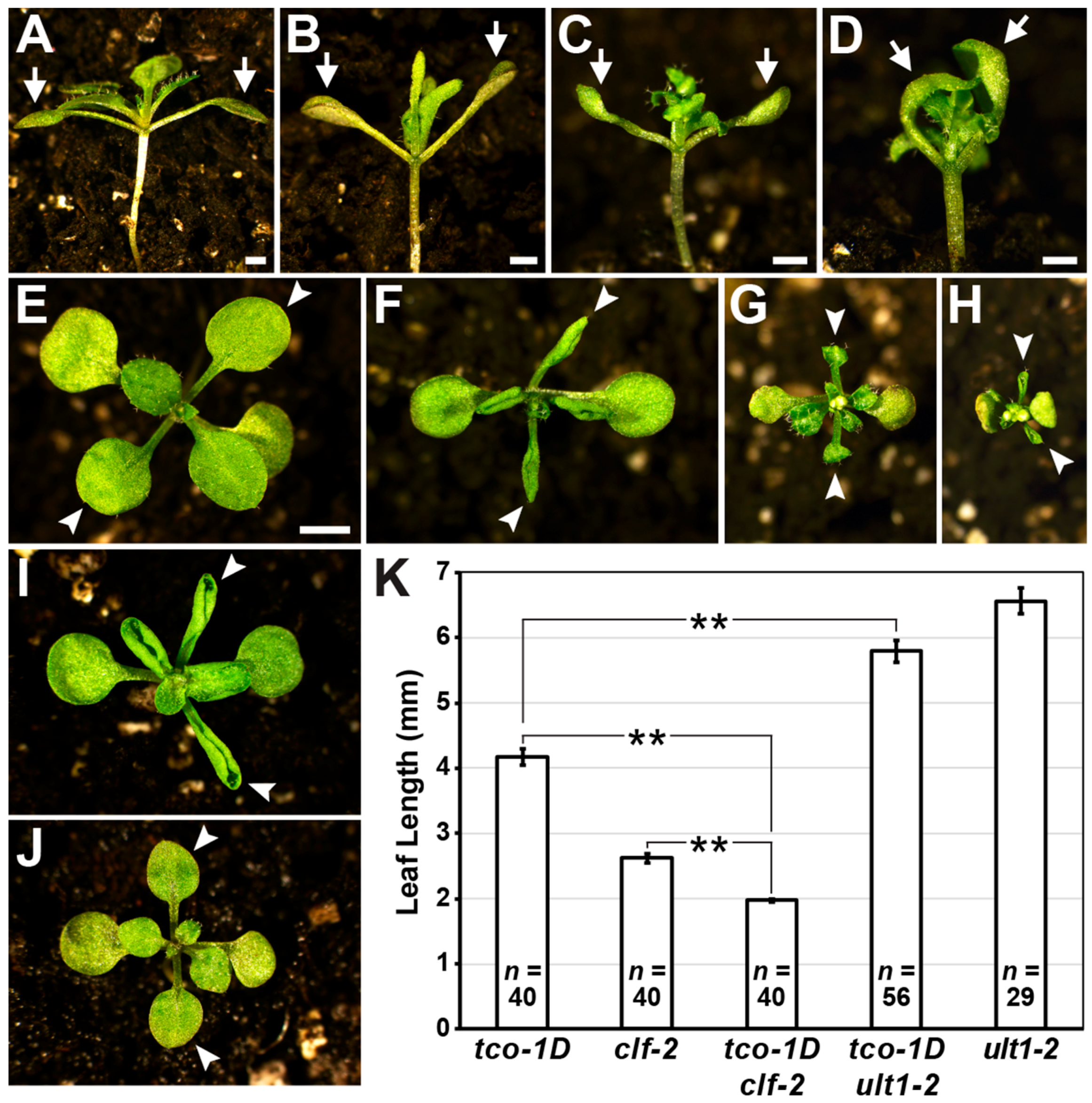
2.3. TCO Displays Characteristics of a Chromatin Regulator
2.4. TCO Exhibits Seed-Specific Expression
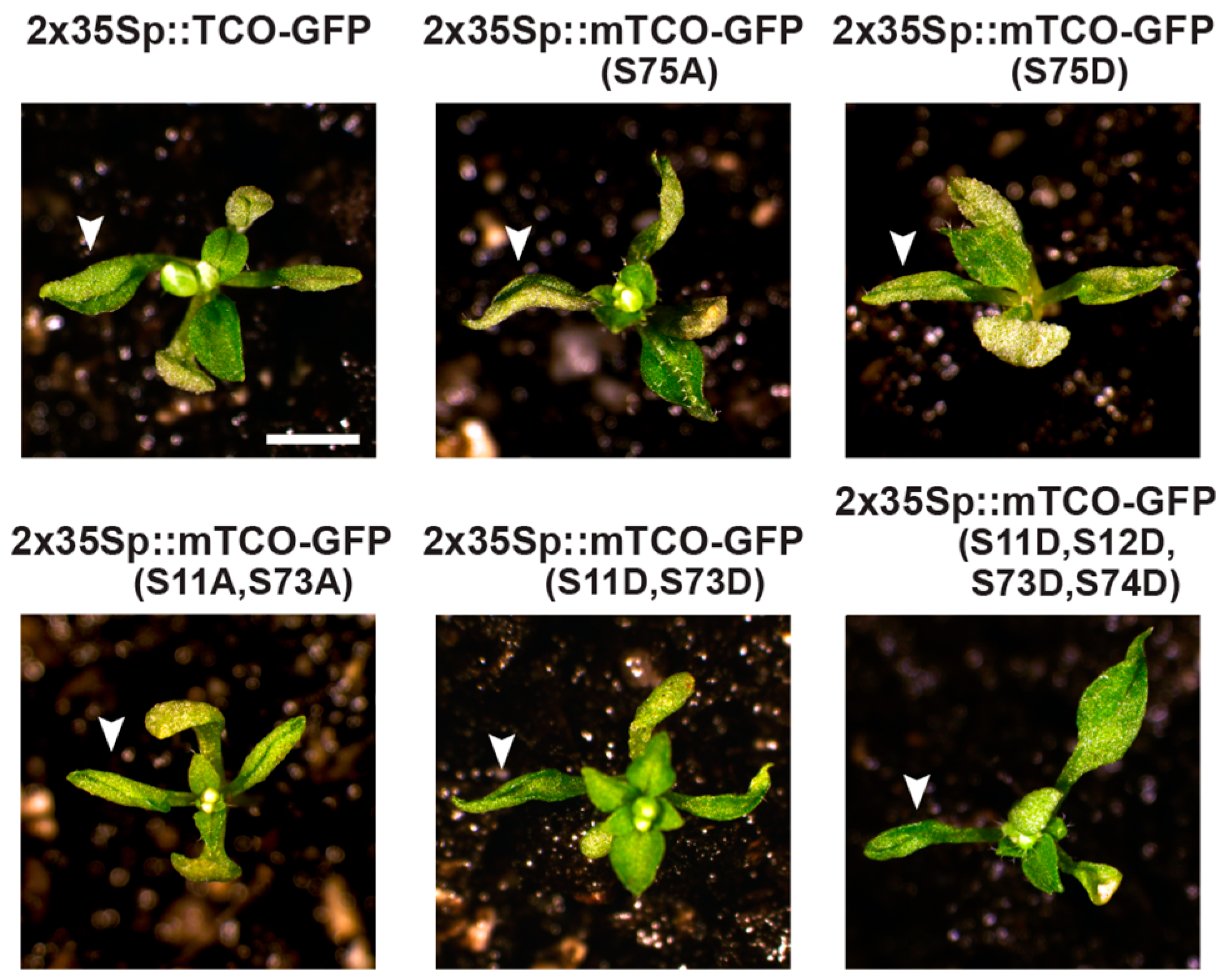
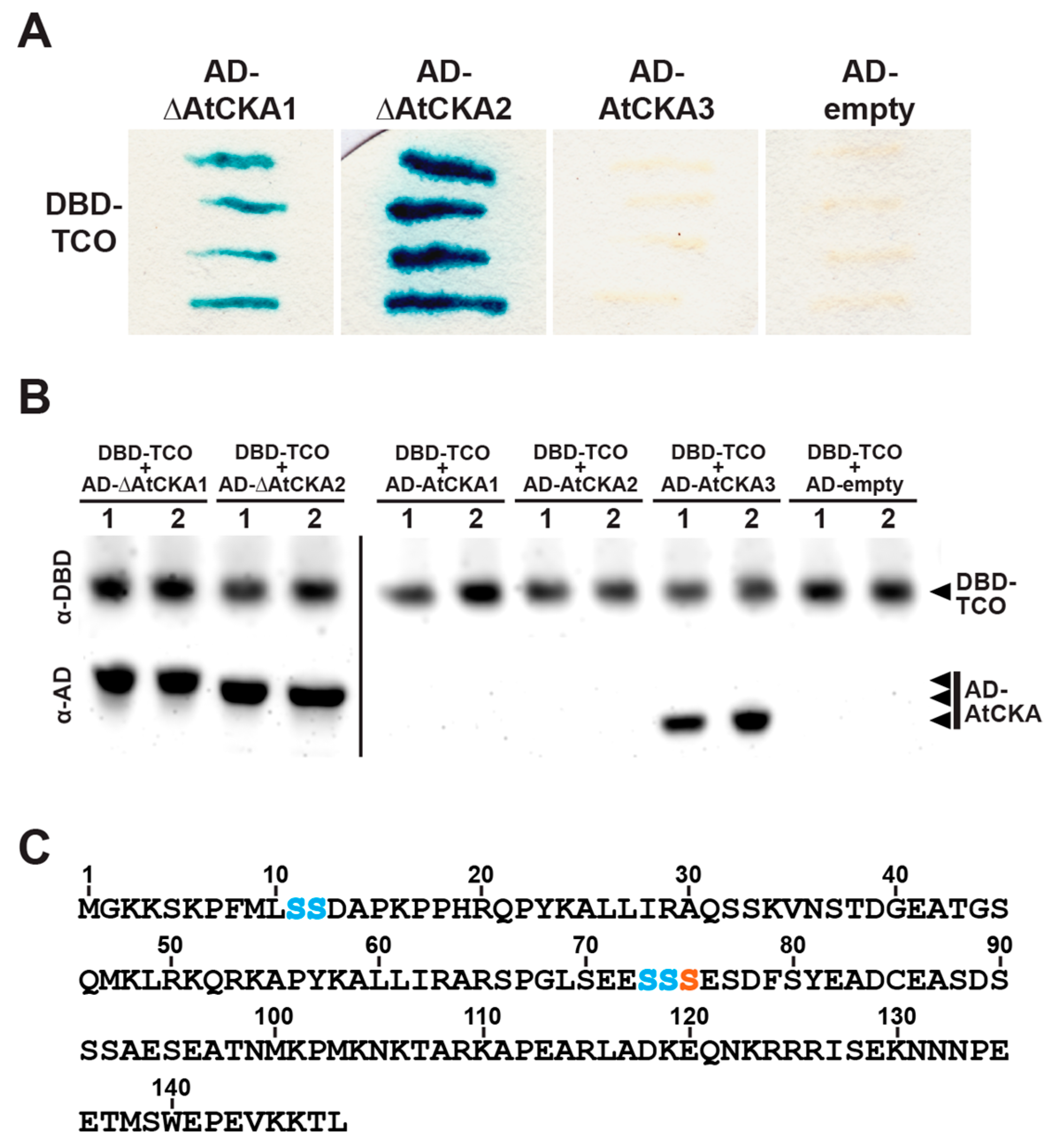
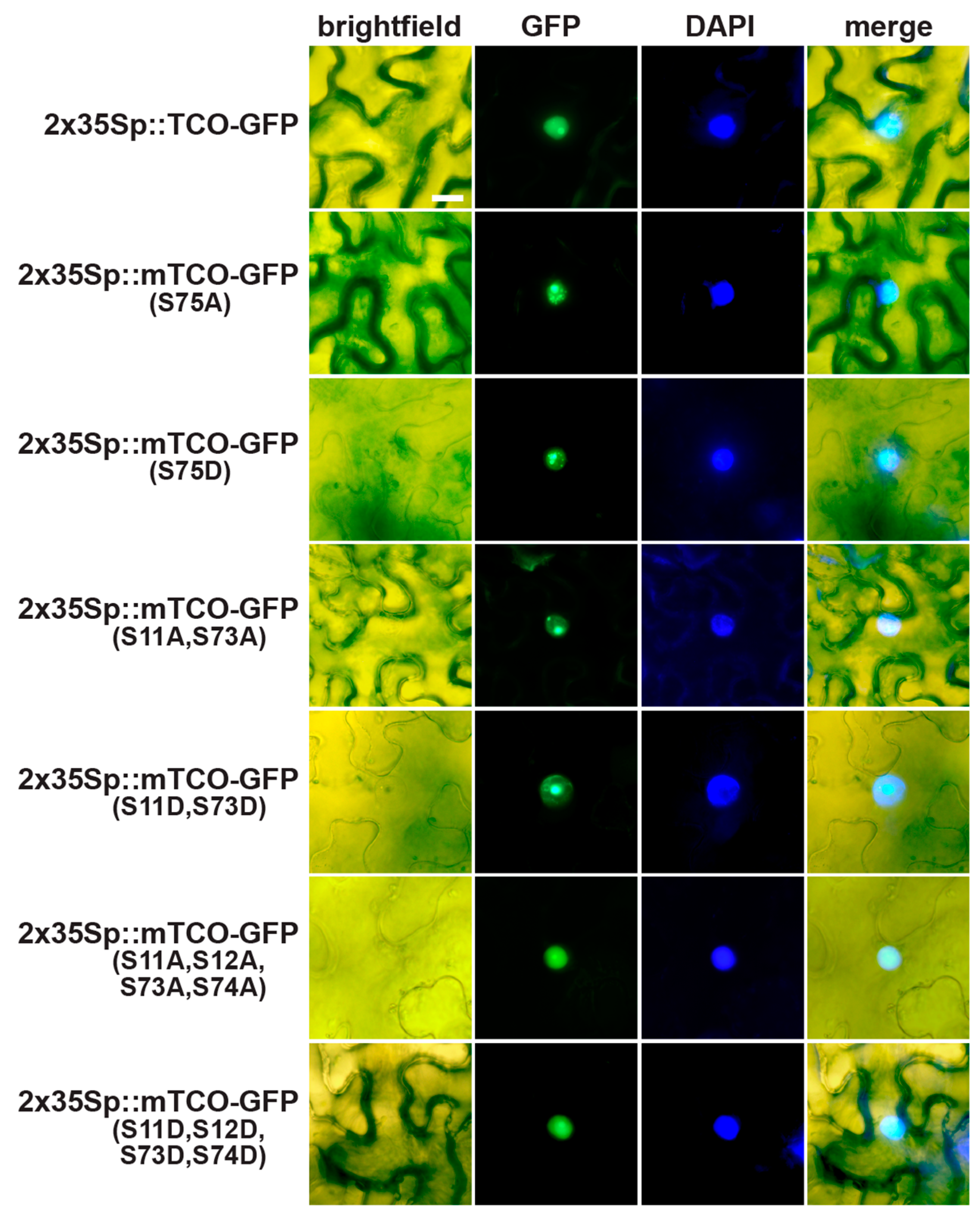
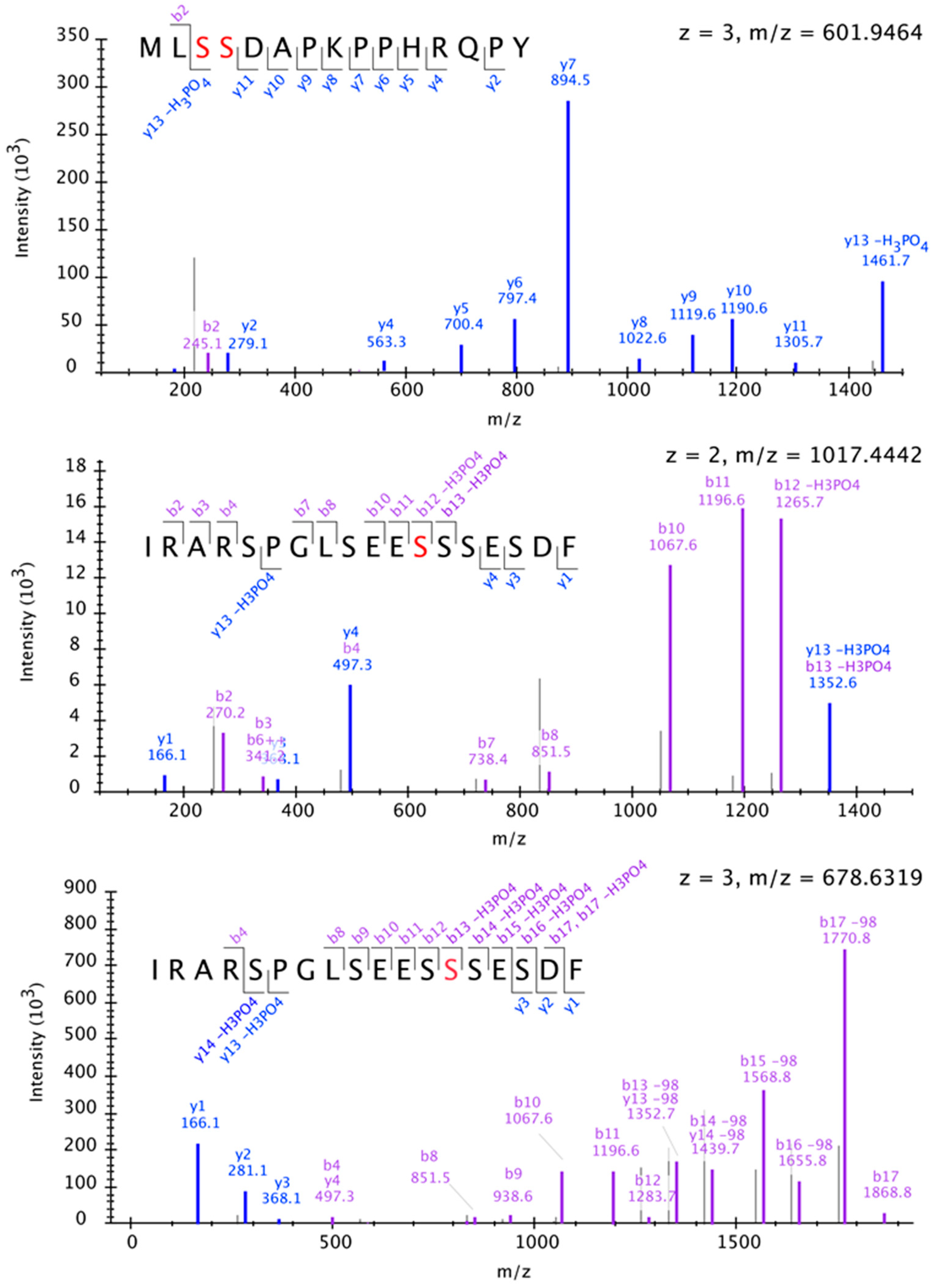
2.5. TCO Interacts with and Is Phosphorylated by the Protein Kinase CK2
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Reverse Transcription-Polymerase Chain Reaction (RT-PCR)
4.3. Transgenic Plant Lines
4.4. Histology and Microscopy
4.5. TCO Protein Expression and Analyses
4.6. Mass Spectrometry Sample Preparation, Acquisition and Data Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| DAG | days after germination |
| DAP | days after pollination |
| Y2H | yeast two-hybrid |
References
- Schuettengruber, B.; Bourbon, H.M.; Di Croce, L.; Cavalli, G. Genome regulation by Polycomb and Trithorax: 70 years and counting. Cell 2017, 171, 34–57. [Google Scholar] [CrossRef] [PubMed]
- Goodrich, J.; Puangsomlee, P.; Martin, M.; Long, D.; Meyerowitz, E.M.; Coupland, G. A Polycomb-group gene regulates homeotic gene expression in Arabidopsis. Nature 1997, 386, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Saleh, A.; Al-Abdallat, A.; Ndamukong, I.; Alvarez-Venegas, R.; Avramova, Z. The Arabidopsis homologs of trithorax (ATX1) and enhancer of zeste (CLF) establish ‘bivalent chromatin marks’ at the silent AGAMOUS locus. Nucleic Acids Res. 2007, 35, 6290–6296. [Google Scholar] [CrossRef] [PubMed]
- Carles, C.C.; Fletcher, J.C. The SAND domain protein ULTRAPETALA1 acts as a trithorax group factor to regulate cell fate in plants. Genes Dev. 2009, 23, 2723–2728. [Google Scholar] [CrossRef] [PubMed]
- Bemer, M.; Grossniklaus, U. Dynamic regulation of Polycomb group activity during plant development. Curr. Opin. Plant Biol. 2012, 15, 523–529. [Google Scholar] [CrossRef] [PubMed]
- De la Paz Sanchez, M.; Aceves-Garcia, P.; Petrone, E.; Steckenborn, S.; Vega-Leon, R.; Alvarez-Buylla, E.R.; Garay-Arroyo, A.; Garcia-Ponce, B. The impact of Polycomb group (PcG) and Trithorax group (TrxG) epigenetic factors in plant plasticity. New Phytol. 2015, 208, 684–694. [Google Scholar] [CrossRef] [PubMed]
- Perino, M.; Veenstra, G.J.C. Chromatin control of developmental dynamics and plasticity. Dev. Cell 2016, 38, 610–620. [Google Scholar] [CrossRef]
- Niessen, H.E.C.; Demmers, J.A.; Voncken, J.W. Talking to chromatin: Post-translational modulation of polycomb group function. Epigenet. Chromatin 2009, 2, 10. [Google Scholar] [CrossRef]
- Bigeard, J.; Rayapuram, N.; Pflieger, D.; Hirt, H. Phosphorylation-dependent regulation of plant chromatin and chromatin-associated proteins. Proteomics 2014, 14, 2127–2140. [Google Scholar] [CrossRef]
- Cha, T.L.; Zhou, B.P.; Xia, W.; Wu, Y.; Yang, C.C.; Chen, C.T.; Ping, B.; Otte, A.P.; Hung, M.C. Akt-mediated phosphorylation of EZH2 suppresses methylation of lysine 27 in histone H3. Science 2005, 310, 306–310. [Google Scholar] [CrossRef]
- Muchardt, C.; Reyes, J.C.; Bourachot, B.; Legouy, E.; Yaniv, M. The hbrm and BRG-1 proteins, components of the human SNF/SWI complex, are phosphorylated and excluded from the condensed chromosomes during mitosis. EMBO J. 1996, 15, 3394–3402. [Google Scholar] [CrossRef]
- Ng, J.; Hart, C.M.; Morgan, K.; Simon, J.A. A Drosophila ESC-E(Z) protein complex is distinct from other Polycomb group complexes and contains covalently modified ESC. Mol. Cell. Biol. 2000, 20, 3069–3078. [Google Scholar] [CrossRef] [PubMed]
- Tie, F.; Siebold, A.P.; Harte, P.J. The N-terminus of Drosophila ESC mediates its phosphorylation and dimerization. Biochem. Biophys. Res. Commun. 2005, 332, 622–632. [Google Scholar] [CrossRef] [PubMed]
- Hatano, A.; Matsumoto, M.; Higashinakagawa, T.; Nakayama, K.I. Phosphorylation of the chromodomain changes the binding specificity of Cbx2 for methylated histone H3. Biochem. Biophys. Res. Commun. 2010, 397, 93–99. [Google Scholar] [CrossRef]
- Liu, Y.G.; Mitsukawa, N.; Oosumi, T.; Whittier, R.F. Efficient isolation and mapping of Arabidopsis thaliana T-DNA insert junctions by thermal asymmetric interlaced PCR. Plant J. 1995, 8, 457–463. [Google Scholar] [CrossRef]
- Berardini, T.Z.; Reiser, L.; Li, D.; Mezheritsky, Y.; Muller, R.; Strait, E.; Huala, E. The Arabidopsis Information Resource: Making and mining the “gold standard” annotated reference plant genome. Genesis 2015, 53, 474–485. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Bowman, J.L.; Smyth, D.R.; Meyerowitz, E.M. Genetic interactions among floral homeotic genes of Arabidopsis. Development 1991, 112, 1–20. [Google Scholar] [PubMed]
- Coen, E.S.; Meyerowitz, E.M. The war of the whorls: Genetic interactions controlling flower development. Nature 1991, 353, 31–37. [Google Scholar] [CrossRef] [PubMed]
- Kardailsky, I.; Shukla, V.K.; Ahn, J.H.; Dagenais, N.; Christensen, S.K.; Nguyen, J.T.; Chory, J.; Harrison, M.J.; Weigel, D. Activation tagging of the floral inducer FT. Science 1999, 286, 1962–1965. [Google Scholar] [CrossRef]
- Kobayashi, Y.; Kaya, H.; Goto, K.; Iwabuchi, M.; Araki, T. A pair of related genes with antagonistic roles in mediating flowering signals. Science 1999, 286, 1960–1962. [Google Scholar] [CrossRef] [PubMed]
- Mizukami, Y.; Ma, H. Ectopic expression of the floral homeotic gene AGAMOUS in transgenic Arabidopsis plants alters floral organ identity. Cell 1992, 71, 119–131. [Google Scholar] [CrossRef]
- Mandel, M.A.; Yanofsky, M.F. A gene triggering flower formation in Arabidopsis. Nature 1995, 377, 522–524. [Google Scholar] [CrossRef] [PubMed]
- Weigel, D.; Nilsson, O. A developmental switch sufficient for flower initiation in diverse plants. Nature 1995, 377, 495–500. [Google Scholar] [CrossRef] [PubMed]
- Krizek, B.A.; Meyerowitz, E.M. The Arabidopsis homeotic genes APETALA3 and PISTILLATA are sufficient to provide the B class organ identity function. Development 1996, 122, 11–22. [Google Scholar] [PubMed]
- Pelaz, S.; Gustafson-Brown, C.; Kohalmi, S.E.; Crosby, W.L.; Yanofsky, M.F. APETALA1 and SEPALLATA3 interact to promote flower development. Plant J. 2001, 26, 385–394. [Google Scholar] [CrossRef] [PubMed]
- Nesi, N.; Debeaujon, I.; Jond, C.; Stewart, A.J.; Jenkins, G.I.; Caboche, M.; Lepiniec, L. The TRANSPARENT TESTA16 locus encodes the ARABIDOPSIS BSISTER MADS domain protein and is required for proper development and pigmentation of the seed coat. Plant Cell 2002, 14, 2463–2479. [Google Scholar] [CrossRef] [PubMed]
- Favaro, R.; Pinyopich, A.; Battaglia, R.; Kooiker, M.; Borghi, L.; Ditta, G.; Yanofsky, M.F.; Kater, M.M.; Colombo, L. MADS-box protein complexes control carpel and ovule development in Arabidopsis. Plant Cell 2003, 15, 2603–2611. [Google Scholar] [CrossRef]
- Pinyopich, A.; Ditta, G.S.; Savidge, B.; Liljegren, S.J.; Baumann, E.; Wisman, E.; Yanofsky, M.F. Assessing the redundancy of MADS-box genes during carpel and ovule development. Nature 2003, 424, 85–88. [Google Scholar] [CrossRef]
- Dietzel, S.; Niemann, H.; Bruckner, B.; Maurange, C.; Paro, R. The nuclear distribution of Polycomb during Drosophila melanogaster development shown with a GFP fusion protein. Chromosoma 1999, 108, 83–94. [Google Scholar] [CrossRef]
- Gaudin, V.; Libault, M.; Pouteau, S.; Juul, T.; Zhao, G.; Lefebvre, D.; Grandjean, O. Mutations in LIKE HETEROCHROMATIN PROTEIN 1 affect flowering time and plant architecture in Arabidopsis. Development 2001, 128, 4847–4858. [Google Scholar] [PubMed]
- Calonje, M.; Sanchez, R.; Chen, L.; Sung, Z.R. EMBRYONIC FLOWER1 participates in Polycomb group-mediated AG gene silencing in Arabidopsis. Plant Cell 2008, 20, 277–291. [Google Scholar] [CrossRef] [PubMed]
- Hernandez-Munoz, I.; Lund, A.H.; van der Stoop, P.; Boutsma, E.; Muijrers, I.; Verhoeven, E.; Nusinow, D.A.; Panning, B.; Marahrens, Y.; van Lohuizen, M. Stable X chromosome inactivation involves the PRC1 Polycomb complex and requires histone MACROH2A1 and the CULLIN3/SPOP ubiquitin E3 ligase. Proc. Natl. Acad. Sci. USA 2005, 102, 7635–7640. [Google Scholar] [CrossRef] [PubMed]
- Carles, C.C.; Choffnes-Inada, D.; Reville, K.; Lertpiriyapong, K.; Fletcher, J.C. ULTRAPETALA1 encodes a SAND domain putative transcriptional regulator that controls shoot and floral meristem activity in Arabidopsis. Development 2005, 132, 897–911. [Google Scholar] [CrossRef] [PubMed]
- Klepikova, A.V.; Kasianov, A.S.; Gerasimov, E.S.; Logacheva, M.D.; Penin, A.A. A high resolution map of the Arabidopsis thaliana developmental transcriptome based on RNA-seq profiling. Plant J. 2016, 88, 1058–1070. [Google Scholar] [CrossRef] [PubMed]
- Salinas, P.; Fuentes, D.; Vidal, E.; Jordana, X.; Echeverria, M.; Holuigue, L. An extensive survey of CK2 α and β subunits in Arabidopsis: Multiple isoforms exhibit differential subcellular localization. Plant Cell Physiol. 2006, 47, 1295–1308. [Google Scholar] [CrossRef] [PubMed]
- Meggio, F.; Pinna, L.A. One-thousand-and-one substrates of protein kinase CK2? FASEB J. 2003, 17, 349–368. [Google Scholar] [CrossRef] [PubMed]
- Chanvivattana, Y.; Bishopp, A.; Schubert, D.; Stock, C.; Moon, Y.H.; Sung, Z.R.; Goodrich, J. Interaction of Polycomb-group proteins controlling flowering in Arabidopsis. Development 2004, 131, 5263–5276. [Google Scholar] [CrossRef]
- Spillane, C.; Schmid, K.J.; Laoueille-Duprat, S.; Pien, S.; Escobar-Restrepo, J.M.; Baroux, C.; Gagliardini, V.; Page, D.R.; Wolfe, K.H.; Grossniklaus, U. Positive darwinian selection at the imprinted MEDEA locus in plants. Nature 2007, 448, 349–352. [Google Scholar] [CrossRef]
- Delgerekh, N. Study on the Role of EARLY FLOWERING AND CURLY LEAVES (EFC) in the FIS2-PRC2 Complex of Arabidopsis thaliana. Master’s Thesis, Seoul National University, Seoul, Korea, 2016. [Google Scholar]
- Kang, H.G.; Klessig, D.F. Salicylic acid-inducible Arabidopsis CK2-like activity phosphorylates TGA2. Plant Mol. Biol. 2005, 57, 541–557. [Google Scholar] [CrossRef]
- Allende-Vega, N.; McKenzie, L.; Meek, D. Transcription factor TAFII250 phosphorylates the acidic domain of Mdm2 through recruitment of protein kinase CK2. Mol. Cell. Biochem. 2008, 316, 99–106. [Google Scholar] [CrossRef] [PubMed]
- Litchfield, D.W. Protein kinase CK2: Structure, regulation and role in cellular decisions of life and death. Biochem. J. 2003, 369, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; Lee, P.; Stafford, J.M.; von Schimmelmann, M.; Schaefer, A.; Reinberg, D. An AUTS2-Polycomb complex activates gene expression in the CNS. Nature 2014, 516, 349–354. [Google Scholar] [CrossRef] [PubMed]
- Prelich, G. Gene overexpression: Uses, mechanisms, and interpretation. Genetics 2012, 190, 841–854. [Google Scholar] [CrossRef] [PubMed]
- Turck, F.; Roudier, F.; Farrona, S.; Martin-Magniette, M.L.; Guillaume, E.; Buisine, N.; Gagnot, S.; Martienssen, R.A.; Coupland, G.; Colot, V. Arabidopsis TFL2/LHP1 specifically associates with genes marked by trimethylation of histone H3 lysine 27. PLoS Genet. 2007, 3, e86. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Germann, S.; Blus, B.J.; Khorasanizadeh, S.; Gaudin, V.; Jacobsen, S.E. The Arabidopsis LHP1 protein colocalizes with histone H3 Lys27 trimethylation. Nat. Struct. Mol. Biol. 2007, 14, 869–871. [Google Scholar] [CrossRef] [PubMed]
- Larsson, A.S.; Landberg, K.; Meeks-Wagner, D.R. The TERMINAL FLOWER2 (TFL2) gene controls the reproductive transition and meristem identity in Arabidopsis thaliana. Genetics 1998, 149, 597–605. [Google Scholar] [PubMed]
- Derkacheva, M.; Steinbach, Y.; Wildhaber, T.; Mozgova, I.; Mahrez, W.; Nanni, P.; Bischof, S.; Gruissem, W.; Hennig, L. Arabidopsis MSI1 connects LHP1 to PRC2 complexes. EMBO J. 2013, 32, 2073–2085. [Google Scholar] [CrossRef] [PubMed]
- Gehring, M.; Bubb, K.L.; Henikoff, S. Extensive demethylation of repetitive elements during seed development underlies gene imprinting. Science 2009, 324, 1447–1451. [Google Scholar] [CrossRef]
- Gehring, M.; Satyaki, P.R. Endosperm and imprinting, inextricably linked. Plant Physiol. 2017, 173, 143–154. [Google Scholar] [CrossRef]
- Xiao, J.; Wagner, D. Polycomb repression in the regulation of growth and development in Arabidopsis. Curr. Opin. Plant Biol. 2015, 23, 15–24. [Google Scholar] [CrossRef] [PubMed]
- Bowman, J.L.; Smyth, D.R.; Meyerowitz, E.M. Genes directing flower development in Arabidopsis. Plant Cell 1989, 1, 37–52. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, J.C. The ULTRAPETALA gene controls shoot and floral meristem size in Arabidopsis. Development 2001, 128, 1323–1333. [Google Scholar] [PubMed]
- Hacisalihoglu, G.; Hilgert, U.; Nash, E.B.; Micklos, D.A. An innovative plant genomics and gene annotation program for high school, community college, and university faculty. CBE Life Sci. Educ. 2008, 7, 310–316. [Google Scholar] [CrossRef] [PubMed]
- Weigel, D.; Ahn, J.H.; Blazquez, M.A.; Borevitz, J.O.; Christensen, S.K.; Frankhauser, C.; Ferrandiz, C.; Kardailsky, I.; Malancharuvil, E.J.; Neff, M.M.; et al. Activation tagging in Arabidopsis. Plant Physiol. 2000, 122, 1003–1013. [Google Scholar] [CrossRef] [PubMed]
- Alonso, J.M.; Stepanova, A.N.; Leisse, T.J.; Kim, C.J.; Chen, H.; Shinn, P.; Stevenson, D.K.; Zimmerman, J.; Barajas, P.; Cheuk, R.; et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 2003, 301, 653–657. [Google Scholar] [CrossRef] [PubMed]
- Krogan, N.T.; Yin, X.; Ckurshumova, W.; Berleth, T. Distinct subclades of Aux/IAA genes are direct targets of ARF5/MP transcriptional regulation. New Phytol. 2014, 204, 474–483. [Google Scholar] [CrossRef]
- Eshed, Y.; Baum, S.F.; Perea, J.V.; Bowman, J.L. Establishment of polarity in lateral organs of plants. Curr. Biol. 2001, 11, 1251–1260. [Google Scholar] [CrossRef]
- Gleave, A.P. A versatile binary vector system with a T-DNA organisational structure conducive to efficient integration of cloned DNA into the plant genome. Plant Mol. Biol. 1992, 20, 1203–1207. [Google Scholar] [CrossRef]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef]
- Szemenyei, H.; Hannon, M.; Long, J.A. TOPLESS mediates auxin-dependent transcriptional repression during Arabidopsis embryogenesis. Science 2008, 319, 1384–1386. [Google Scholar] [CrossRef] [PubMed]
- Warner, C.A.; Biedrzycki, M.L.; Jacobs, S.S.; Wisser, R.J.; Caplan, J.L.; Sherrier, D.J. An optical clearing technique for plant tissues allowing deep imaging and compatible with fluorescence microscopy. Plant Physiol. 2014, 166, 1684–1687. [Google Scholar] [CrossRef] [PubMed]
- Krogan, N.T.; Hogan, K.; Long, J.A. APETALA2 negatively regulates multiple floral organ identity genes in Arabidopsis by recruiting the co-repressor TOPLESS and the histone deacetylase HDA19. Development 2012, 139, 4180–4190. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 2008, 26, 1367–1372. [Google Scholar] [CrossRef] [PubMed]
- MacLean, B.; Tomazela, D.M.; Shulman, N.; Chambers, M.; Finney, G.L.; Frewen, B.; Kern, R.; Tabb, D.L.; Liebler, D.C.; MacCoss, M.J. Skyline: An open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 2010, 26, 966–968. [Google Scholar] [CrossRef] [PubMed]
- Vizcaino, J.A.; Csordas, A.; del-Toro, N.; Dianes, J.A.; Griss, J.; Lavidas, I.; Mayer, G.; Perez-Riverol, Y.; Reisinger, F.; Ternent, T.; et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 2016, 44, D447–D456. [Google Scholar] [CrossRef] [PubMed]

© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Weinman, L.M.; Running, K.L.D.; Carey, N.S.; Stevenson, E.J.; Swaney, D.L.; Chow, B.Y.; Krogan, N.J.; Krogan, N.T. TCO, a Putative Transcriptional Regulator in Arabidopsis, Is a Target of the Protein Kinase CK2. Int. J. Mol. Sci. 2019, 20, 99. https://doi.org/10.3390/ijms20010099
Weinman LM, Running KLD, Carey NS, Stevenson EJ, Swaney DL, Chow BY, Krogan NJ, Krogan NT. TCO, a Putative Transcriptional Regulator in Arabidopsis, Is a Target of the Protein Kinase CK2. International Journal of Molecular Sciences. 2019; 20(1):99. https://doi.org/10.3390/ijms20010099
Chicago/Turabian StyleWeinman, Laina M., Katherine L. D. Running, Nicholas S. Carey, Erica J. Stevenson, Danielle L. Swaney, Brenda Y. Chow, Nevan J. Krogan, and Naden T. Krogan. 2019. "TCO, a Putative Transcriptional Regulator in Arabidopsis, Is a Target of the Protein Kinase CK2" International Journal of Molecular Sciences 20, no. 1: 99. https://doi.org/10.3390/ijms20010099
APA StyleWeinman, L. M., Running, K. L. D., Carey, N. S., Stevenson, E. J., Swaney, D. L., Chow, B. Y., Krogan, N. J., & Krogan, N. T. (2019). TCO, a Putative Transcriptional Regulator in Arabidopsis, Is a Target of the Protein Kinase CK2. International Journal of Molecular Sciences, 20(1), 99. https://doi.org/10.3390/ijms20010099




