Cytokinin at the Crossroads of Abiotic Stress Signalling Pathways
Abstract
1. Introduction
2. Nutrient Stress
2.1. Nitrogen
2.2. Phosphorus
2.3. Potassium
2.4. Sulphur
2.5. Boron
2.6. Iron
2.7. Silicon
2.8. Selenium
2.9. Xenobiotics
3. Cytokinin Roles in Drought and Salinity Tolerance
3.1. Cytokinin Modulates Photosynthesis under Water-Limiting Conditions and Salt Stress
3.2. Cytokinin Enhances Capacities of Antioxidant Systems
3.3. Cytokinin Influences Water Balance Regulation
3.4. Cytokinin Effects on Growth
3.5. Cytokinin Crosstalk with Stress-Related Phytohormones
3.5.1. Abscisic Acid
3.5.2. Jasmonates
4. Temperature and Cytokinin
4.1. Low Temperature Stress: Cold and Freezing
4.2. High Temperature and Heat Stress
5. Light Signalling and the Circadian Clock Interact with Cytokinin
6. Summary
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Suzuki, I.; Los, D.A.; Kanesaki, Y.; Mikami, K.; Murata, N. The pathway for perception and transduction of low-temperature signals in Synechocystis. EMBO J. 2000, 19, 1327–1334. [Google Scholar] [CrossRef] [PubMed]
- Hwang, I.; Chen, H.C.; Sheen, J. Two-component signal transduction pathways in Arabidopsis. Plant Physiol. 2002, 129, 500–515. [Google Scholar] [CrossRef] [PubMed]
- Wolanin, P.M.; Thomason, P.A.; Stock, J.B. Histidine protein kinases: Key signal transducers outside the animal kingdom. Genome Biol. 2002, 3. [Google Scholar] [CrossRef]
- O’Brien, J.A.; Benková, E. Cytokinin cross-talking during biotic and abiotic stress responses. Front. Plant Sci. 2013, 4, 451. [Google Scholar] [CrossRef] [PubMed]
- Kieber, J.J.; Schaller, G.E. Cytokinin signaling in plant development. Development 2018, 145, dev149344. [Google Scholar] [CrossRef] [PubMed]
- Krapp, A. Plant nitrogen assimilation and its regulation: A complex puzzle with missing pieces. Curr. Opin. Plant Biol. 2015, 25, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Gent, L.; Forde, B.G. How do plants sense their nitrogen status? J. Exp. Bot. 2017, 68, 2531–2539. [Google Scholar] [CrossRef] [PubMed]
- Bellegarde, F.; Gojon, A.; Martin, A. Signals and players in the transcriptional regulation of root responses by local and systemic N signaling in Arabidopsis thaliana. J. Exp. Bot. 2017, 68, 2553–2565. [Google Scholar] [CrossRef] [PubMed]
- Guan, P.; Ripoll, J.J.; Wang, R.; Vuong, L.; Bailey-Steinitz, L.J.; Ye, D.; Crawford, N.M. Interacting TCP and NLP transcription factors control plant responses to nitrate availability. Proc. Natl. Acad. Sci. USA 2017, 114, 2419–2424. [Google Scholar] [CrossRef] [PubMed]
- Kiba, T.; Takei, K.; Kojima, M.; Sakakibara, H. Side-chain modification of cytokinins controls shoot growth in Arabidopsis. Dev. Cell 2013, 27, 452–461. [Google Scholar] [CrossRef] [PubMed]
- Kieber, J.J.; Schaller, G.E. Cytokinins. Arabidopsis Book 2014, 12, e0168. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Tischner, R.; Gutiérrez, R.A.; Hoffman, M.; Xing, X.; Chen, M.; Coruzzi, G.; Crawford, N.M. Genomic analysis of the nitrate response using a nitrate reductase-null mutant of Arabidopsis. Plant Physiol. 2004, 136, 2512–2522. [Google Scholar] [CrossRef] [PubMed]
- Ramireddy, E.; Chang, L.; Schmülling, T. Cytokinin as a mediator for regulating root system architecture in response to environmental cues. Plant Signal. Behav. 2014, 9, e27771. [Google Scholar] [CrossRef] [PubMed]
- Menz, J.; Li, Z.; Schulze, W.X.; Ludewig, U. Early nitrogen-deprivation responses in Arabidopsis roots reveal distinct differences on transcriptome and (phospho-) proteome levels between nitrate and ammonium nutrition. Plant J. 2016, 88, 717–734. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.H.; Niu, Y.; Konishi, M.; Wu, Y.; Du, H.; Sun Chung, H.; Li, L.; Boudsocq, M.; McCormack, M.; Maekawa, S.; et al. Discovery of nitrate–CPK–NLP signalling in central nutrient–growth networks. Nature 2017, 545, 311–316. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Xing, X.; Wang, Y.; Tran, A.; Crawford, N.M. A Genetic screen for nitrate regulatory mutants captures the nitrate transporter gene NRT1.1. Plant Physiol. 2009, 151, 472–478. [Google Scholar] [CrossRef] [PubMed]
- Maeda, Y.; Konishi, M.; Kiba, T.; Sakuraba, Y.; Sawaki, N.; Kurai, T.; Ueda, Y.; Sakakibara, H.; Yanagisawa, S. A NIGT1-centred transcriptional cascade regulates nitrate signalling and incorporates phosphorus starvation signals in Arabidopsis. Nat. Commun. 2018, 9, 1376. [Google Scholar] [CrossRef] [PubMed]
- Ruffel, S.; Krouk, G.; Ristova, D.; Shasha, D.; Birnbaum, K.D.; Coruzzi, G.M. Nitrogen economics of root foraging: Transitive closure of the nitrate-cytokinin relay and distinct systemic signaling for N supply vs. demand. Proc. Natl. Acad. Sci. USA 2011, 108, 18524–18529. [Google Scholar] [CrossRef] [PubMed]
- Ruffel, S.; Poitout, A.; Krouk, G.; Coruzzi, G.M.; Lacombe, B. Long-distance nitrate signaling displays cytokinin dependent and independent branches. J. Integr. Plant Biol. 2016, 58, 226–229. [Google Scholar] [CrossRef] [PubMed]
- Poitout, A.; Crabos, A.; Petřík, I.; Novák, O.; Krouk, G.; Lacombe, B.; Ruffel, S. Responses to Systemic Nitrogen Signaling in Arabidopsis Roots Involve trans-Zeatin in Shoots. Plant Cell 2018, 30, 1243–1257. [Google Scholar] [CrossRef] [PubMed]
- Guan, P.; Wang, R.; Nacry, P.; Breton, G.; Kay, S.A.; Pruneda-Paz, J.L.; Davani, A.; Crawford, N.M. Nitrate foraging by Arabidopsis roots is mediated by the transcription factor TCP20 through the systemic signaling pathway. Proc. Natl. Acad. Sci. USA 2014, 111, 15267–15272. [Google Scholar] [CrossRef] [PubMed]
- Patterson, K.; Walters, L.A.; Cooper, A.M.; Olvera, J.G.; Rosas, M.A.; Rasmusson, A.G.; Escobar, M.A. Nitrate-Regulated Glutaredoxins Control Arabidopsis Primary Root Growth. Plant Physiol. 2016, 170, 989–999. [Google Scholar] [CrossRef] [PubMed]
- Walters, L.A.; Escobar, M.A. The AtGRXS3/4/5/7/8 glutaredoxin gene cluster on Arabidopsis thaliana chromosome 4 is coordinately regulated by nitrate and appears to control primary root growth. Plant Signal. Behav. 2016, 11, e1171450. [Google Scholar] [CrossRef] [PubMed]
- Walch-Liu, P.; Neumann, G.; Bangerth, F.; Engels, C. Rapid effects of nitrogen form on leaf morphogenesis in tobacco. J. Exp. Bot. 2000, 51, 227–237. [Google Scholar] [CrossRef] [PubMed]
- Rahayu, Y.S.; Walch-Liu, P.; Neumann, G.; Römheld, V.; von Wirén, N.; Bangerth, F. Root-derived cytokinins as long-distance signals for NO3-induced stimulation of leaf growth. J. Exp. Bot. 2005, 56, 1143–1152. [Google Scholar] [CrossRef] [PubMed]
- Müller, D.; Waldie, T.; Miyawaki, K.; To, J.P.; Melnyk, C.W.; Kieber, J.J.; Kakimoto, T.; Leyser, O. Cytokinin is required for escape but not release from auxin mediated apical dominance. Plant J. 2015, 82, 874–886. [Google Scholar] [CrossRef] [PubMed]
- Miyawaki, K.; Matsumoto-Kitano, M.; Kakimoto, T. Expression of cytokinin biosynthetic isopentenyltransferase genes in Arabidopsis: Tissue specificity and regulation by auxin, cytokinin, and nitrate. Plant J. 2004, 37, 128–138. [Google Scholar] [CrossRef] [PubMed]
- Takei, K.; Ueda, N.; Aoki, K.; Kuromori, T.; Hirayama, T.; Shinozaki, K.; Yamaya, T.; Sakakibara, H. AtIPT3 is a key determinant of nitrate-dependent cytokinin biosynthesis in Arabidopsis. Plant Cell Physiol. 2004, 45, 1053–1062. [Google Scholar] [CrossRef] [PubMed]
- Sakakibara, H.; Takei, K.; Hirose, N. Interactions between nitrogen and cytokinin in the regulation of metabolism and development. Trends Plant Sci. 2006, 11, 440–448. [Google Scholar] [CrossRef] [PubMed]
- Kiba, T.; Kudo, T.; Kojima, M.; Sakakibara, H. Hormonal control of nitrogen acquisition: Roles of auxin, abscisic acid, and cytokinin. J. Exp. Bot. 2011, 62, 1399–1409. [Google Scholar] [CrossRef] [PubMed]
- Osugi, A.; Kojima, M.; Takebayashi, Y.; Ueda, N.; Kiba, T.; Sakakibara, H. Systemic transport of trans-zeatin and its precursor have differing roles in Arabidopsis shoots. Nat. Plants 2017, 3, 17112. [Google Scholar] [CrossRef] [PubMed]
- Landrein, B.; Formosa-Jordan, P.; Malivert, A.; Schuster, C.; Melnyk, C.W.; Yang, W.; Turnbull, C.; Meyerowitz, E.M.; Locke, J.C.W.; Jönsson, H. Nitrate modulates stem cell dynamics in Arabidopsis shoot meristems through cytokinins. Proc. Natl. Acad. Sci. USA 2018, 115, 1382–1387. [Google Scholar] [CrossRef] [PubMed]
- Krishnakumar, V.; Hanlon, M.R.; Contrino, S.; Ferlanti, E.S.; Karamycheva, S.; Kim, M.; Rosen, B.D.; Cheng, C.Y.; Moreira, W.; Mock, S.A.; et al. Araport: The Arabidopsis Information Portal. Nucleic Acids Res. 2015, 43, D1003–D1009. [Google Scholar] [CrossRef] [PubMed]
- Rouached, H.; Arpat, A.B.; Poirier, Y. Regulation of Phosphate Starvation Responses in Plants: Signaling Players and Cross-Talks. Mol. Plant 2010, 3, 288–299. [Google Scholar] [CrossRef] [PubMed]
- Ham, B.K.; Chen, J.; Yan, Y.; Lucas, W.J. Insights into plant phosphate sensing and signaling. Curr. Opin. Biotechnol. 2018, 49, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Hirose, N.; Takei, K.; Kuroha, T.; Kamada-Nobusada, T.; Hayashi, H.; Sakakibara, H. Regulation of cytokinin biosynthesis, compartmentalization and translocation. J. Exp. Bot. 2008, 59, 75–83. [Google Scholar] [CrossRef] [PubMed]
- Franco-Zorrilla, J.M.; Martin, A.C.; Solano, R.; Rubio, V.; Leyva, A.; Paz-Ares, J. Mutations at CRE1 impair cytokinin-induced repression of phosphate starvation responses in Arabidopsis. Plant J. 2002, 32, 353–360. [Google Scholar] [CrossRef] [PubMed]
- Woo, J.; MacPherson, C.R.; Liu, J.; Wang, H.; Kiba, T.; Hannah, M.A.; Wang, X.J.; Bajic, V.B.; Chua, N.H. The response and recovery of the Arabidopsis thaliana transcriptome to phosphate starvation. BMC Plant Biol. 2012, 12, 62. [Google Scholar] [CrossRef] [PubMed]
- Werner, T.; Nehnevajova, E.; Köllmer, I.; Novák, O.; Strnad, M.; Krämer, U.; Schmülling, T. Root-Specific Reduction of Cytokinin Causes Enhanced Root Growth, Drought Tolerance, and Leaf Mineral Enrichment in Arabidopsis and Tobacco. Plant Cell 2010, 22, 3905–3920. [Google Scholar] [CrossRef] [PubMed]
- Nishiyama, R.; Le, D.T.; Watanabe, Y.; Matsui, A.; Tanaka, M.; Seki, M.; Yamaguchi-Shinozaki, K.; Shinozaki, K.; Tran, L.S. Transcriptome analyses of a salt-tolerant cytokinin-deficient mutant reveal differential regulation of salt stress response by cytokinin deficiency. PLoS ONE 2012, 7, e32124. [Google Scholar] [CrossRef] [PubMed]
- Mohan, T.C.; Castrillo, G.; Navarro, C.; Zarco-Fernández, S.; Ramireddy, E.; Mateo, C.; Zamarreño, A.M.; Paz-Ares, J.; Muñoz, R.; García-Mina, J.M.; et al. Cytokinin Determines Thiol-Mediated Arsenic Tolerance and Accumulation. Plant Physiol. 2016, 171, 1418–1426. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Cao, H.; Chen, Z.; Liu, C.; Cao, S.; Wei, Z.; Han, Y.; Gao, Q.; Wang, W. Cytokinin is involved in TPS22-mediated selenium tolerance in Arabidopsis thaliana. Ann. Bot. 2018. [Google Scholar] [CrossRef] [PubMed]
- Martín, A.C.; del Pozo, J.C.; Iglesias, J.; Rubio, V.; Solano, R.; de La Peña, A.; Leyva, A.; Paz-Ares, J. Influence of cytokinins on the expression of phosphate starvation responsive genes in Arabidopsis. Plant J. 2000, 24, 559–567. [Google Scholar] [CrossRef] [PubMed]
- Franco-Zorrilla, J.M.; Martín, A.C.; Leyva, A.; Paz-Ares, J. Interaction between phosphate-starvation, sugar, and cytokinin signaling in Arabidopsis and the roles of cytokinin receptors CRE1/AHK4 and AHK3. Plant Physiol. 2005, 138, 847–857. [Google Scholar] [CrossRef] [PubMed]
- Shin, H.; Shin, H.S.; Chen, R.; Harrison, M.J. Loss of At4 function impacts phosphate distribution between the roots and the shoots during phosphate starvation. Plant J. 2006, 45, 712–726. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yi, K.; Tao, Y.; Wang, F.; Wu, Z.; Jiang, D.; Chen, X.; Zhu, L.; Wu, P. Cytokinin represses phosphate-starvation response through increasing of intracellular phosphate level. Plant Cell Environ. 2006, 29, 1924–1935. [Google Scholar] [CrossRef] [PubMed]
- Shen, C.; Yue, R.; Yang, Y.; Zhang, L.; Sun, T.; Tie, S.; Wang, H. OsARF16 Is Involved in Cytokinin-Mediated Inhibition of Phosphate Transport and Phosphate Signaling in Rice (Oryza sativa L.). PLoS ONE 2014, 9, e112906. [Google Scholar] [CrossRef] [PubMed]
- Ribot, C.; Wang, Y.; Poirier, Y. Expression analyses of three members of the AtPHO1 family reveal differential interactions between signaling pathways involved in phosphate deficiency and the responses to auxin, cytokinin, and abscisic acid. Planta 2008, 227, 1025–1036. [Google Scholar] [CrossRef] [PubMed]
- Lai, F.; Thacker, J.; Li, Y.; Doerner, P. Cell division activity determines the magnitude of phosphate starvation responses in Arabidopsis. Plant J. 2007, 50, 545–556. [Google Scholar] [CrossRef] [PubMed]
- Schaller, G.E.; Bishopp, A.; Kieber, J.J. The Yin-Yang of Hormones: Cytokinin and Auxin Interactions in Plant Development. Plant Cell Online 2015, 27, 44–63. [Google Scholar] [CrossRef] [PubMed]
- Nam, Y.J.; Tran, L.S.P.; Kojima, M.; Sakakibara, H.; Nishiyama, R.; Shin, R. Regulatory roles of cytokinins and cytokinin signaling in response to potassium deficiency in Arabidopsis. PLoS ONE 2012, 7, e47797. [Google Scholar] [CrossRef] [PubMed]
- Schachtman, D.P. The Role of Ethylene in Plant Responses to K+ Deficiency. Front. Plant Sci. 2015, 6, 1153. [Google Scholar] [CrossRef] [PubMed]
- Rigas, S.; Ditengou, F.A.; Ljung, K.; Daras, G.; Tietz, O.; Palme, K.; Hatzopoulos, P. Root gravitropism and root hair development constitute coupled developmental responses regulated by auxin homeostasis in the Arabidopsis root apex. New Phytol. 2013, 197, 1130–1141. [Google Scholar] [CrossRef] [PubMed]
- Koprivova, A.; Kopriva, S. Sulfur metabolism and its manipulation in crops. J. Genet. Genom. 2016, 43, 623–629. [Google Scholar] [CrossRef] [PubMed]
- Honsel, A.; Kojima, M.; Haas, R.; Frank, W.; Sakakibara, H.; Herschbach, C.; Rennenberg, H. Sulphur limitation and early sulphur deficiency responses in poplar: Significance of gene expression, metabolites, and plant hormones. J. Exp. Bot. 2012, 63, 1873–1893. [Google Scholar] [CrossRef] [PubMed]
- Maruyama-Nakashita, A.; Nakamura, Y.; Yamaya, T.; Takahashi, H. A novel regulatory pathway of sulfate uptake in Arabidopsis roots: Implication of CRE1/WOL/AHK4-mediated cytokinin-dependent regulation. Plant J. 2004, 38, 779–789. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, K.H.; Ha, C. Van; Nishiyama, R.; Watanabe, Y.; Leyva-González, M.A.; Fujita, Y.; Tran, U.T.; Li, W.; Tanaka, M.; Seki, M.; Schaller, G.E.; et al. Arabidopsis type B cytokinin response regulators ARR1, ARR10, and ARR12 negatively regulate plant responses to drought. Proc. Natl. Acad. Sci. USA 2016, 113, 3090–3095. [Google Scholar] [CrossRef] [PubMed]
- Bhargava, A.; Clabaugh, I.; To, J.P.; Maxwell, B.B.; Chiang, Y.H.; Schaller, G.E.; Loraine, A.; Kieber, J.J. Identification of cytokinin-responsive genes using microarray meta-analysis and RNA-Seq in Arabidopsis. Plant Physiol. 2013, 162, 272–294. [Google Scholar] [CrossRef] [PubMed]
- Öztürk, S.E.; Göktay, M.; Has, C.; Babaoğlu, M.; Allmer, J.; Doğanlar, S.; Frary, A. Transcriptomic analysis of boron hyperaccumulation mechanisms in Puccinellia distans. Chemosphere 2018, 199, 390–401. [Google Scholar] [CrossRef] [PubMed]
- González-Fontes, A.; Herrera-Rodríguez, M.B.; Martín-Rejano, E.M.; Navarro-Gochicoa, M.T.; Rexach, J.; Camacho-Cristóbal, J.J. Root Responses to Boron Deficiency Mediated by Ethylene. Front. Plant Sci. 2015, 6, 1103. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.Q.; Liu, Y.Z.; An, J.C.; Li, S.; Jin, L.F.; Zhou, G.F.; Wei, Q.J.; Yan, H.Q.; Wang, N.N.; Fu, L.N.; et al. Digital gene expression analysis of corky split vein caused by boron deficiency in “Newhall” Navel Orange (Citrus sinensis Osbeck) for selecting differentially expressed genes related to vascular hypertrophy. PLoS ONE 2013, 8, e65737. [Google Scholar] [CrossRef] [PubMed]
- Abreu, I.; Poza, L.; Bonilla, I.; Bolaños, L. Boron deficiency results in early repression of a cytokinin receptor gene and abnormal cell differentiation in the apical root meristem of Arabidopsis thaliana. Plant Physiol. Biochem. 2014, 77, 117–121. [Google Scholar] [CrossRef] [PubMed]
- Eggert, K.; von Wirén, N. Response of the plant hormone network to boron deficiency. New Phytol. 2017, 216, 868–881. [Google Scholar] [CrossRef] [PubMed]
- Poza-Viejo, L.; Abreu, I.; González-García, M.P.; Allauca, P.; Bonilla, I.; Bolaños, L.; Reguera, M. Boron deficiency inhibits root growth by controlling meristem activity under cytokinin regulation. Plant Sci. 2018, 270, 176–189. [Google Scholar] [CrossRef] [PubMed]
- Séguéla, M.; Briat, J.F.; Vert, G.; Curie, C. Cytokinins negatively regulate the root iron uptake machinery in Arabidopsis through a growth-dependent pathway. Plant J. 2008, 55, 289–300. [Google Scholar] [CrossRef] [PubMed]
- Shen, C.; Yue, R.; Sun, T.; Zhang, L.; Yang, Y.; Wang, H. OsARF16, a transcription factor regulating auxin redistribution, is required for iron deficiency response in rice (Oryza sativa L.). Plant Sci. 2015, 231, 148–158. [Google Scholar] [CrossRef] [PubMed]
- Yin, L.; Wang, S.; Liu, P.; Wang, W.; Cao, D.; Deng, X.; Zhang, S. Silicon-mediated changes in polyamine and 1-aminocyclopropane-1-carboxylic acid are involved in silicon-induced drought resistance in Sorghum bicolor L. Plant Physiol. Biochem. 2014, 80, 268–277. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.H.; Khan, A.L.; Waqas, M.; Jeong, H.J.; Kim, D.H.; Shin, J.S.; Kim, J.G.; Yeon, M.H.; Lee, I.J. Regulation of jasmonic acid biosynthesis by silicon application during physical injury to Oryza sativa L. J. Plant Res. 2014, 127, 525–532. [Google Scholar] [CrossRef] [PubMed]
- Hosseini, S.A.; Maillard, A.; Hajirezaei, M.R.; Ali, N.; Schwarzenberg, A.; Jamois, F.; Yvin, J.C. Induction of Barley Silicon Transporter HvLsi1 and HvLsi2, increased silicon concentration in the shoot and regulated Starch and ABA Homeostasis under Osmotic stress and Concomitant Potassium Deficiency. Front. Plant Sci. 2017, 8, 1359. [Google Scholar] [CrossRef] [PubMed]
- Markovich, O.; Steiner, E.; Kouřil, Š.; Tarkowski, P.; Aharoni, A.; Elbaum, R. Silicon promotes cytokinin biosynthesis and delays senescence in Arabidopsis and Sorghum. Plant Cell Environ. 2017, 40, 1189–1196. [Google Scholar] [CrossRef] [PubMed]
- Hartikainen, H. Biogeochemistry of selenium and its impact on food chain quality and human health. J. Trace Elem. Med. Biol. 2005, 18, 309–318. [Google Scholar] [CrossRef] [PubMed]
- Pilon-Smits, E.A.; Quinn, C.F.; Tapken, W.; Malagoli, M.; Schiavon, M. Physiological functions of beneficial elements. Curr. Opin. Plant Biol. 2009, 12, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Schiavon, M.; Pilon-Smits, E.A. The fascinating facets of plant selenium accumulation—Biochemistry, physiology, evolution and ecology. New Phytol. 2017, 213, 1582–1596. [Google Scholar] [CrossRef] [PubMed]
- Shibagaki, N.; Rose, A.; McDermott, J.P.; Fujiwara, T.; Hayashi, H.; Yoneyama, T.; Davies, J.P. Selenate-resistant mutants of Arabidopsis thaliana identify Sultr1;2, a sulfate transporter required for efficient transport of sulfate into roots. Plant J. 2002, 29, 475–486. [Google Scholar] [CrossRef] [PubMed]
- Lehotai, N.; Kolbert, Z.; Peto, A.; Feigl, G.; Ördög, A.; Kumar, D.; Tari, I.; Erdei, L. Selenite-induced hormonal and signalling mechanisms during root growth of Arabidopsis thaliana L. J. Exp. Bot. 2012, 63, 5677–5687. [Google Scholar] [CrossRef] [PubMed]
- Kolbert, Z.; Lehotai, N.; Molnár, Á.; Feigl, G. “The roots” of selenium toxicity: A new concept. Plant Signal. Behav. 2016, 11, e1241935. [Google Scholar] [CrossRef] [PubMed]
- Lehotai, N.; Feigl, G.; Koós, Á.; Molnár, Á.; Ördög, A.; Pető, A.; Erdei, L.; Kolbert, Z. Nitric oxide–cytokinin interplay influences selenite sensitivity in Arabidopsis. Plant Cell Rep. 2016, 35, 2181–2195. [Google Scholar] [CrossRef] [PubMed]
- Bruno, L.; Pacenza, M.; Forgione, I.; Lamerton, L.R.; Greco, M.; Chiappetta, A.; Bitonti, M.B. In Arabidopsis thaliana Cadmium Impact on the Growth of Primary Root by Altering SCR Expression and Auxin-Cytokinin Cross-Talk. Front. Plant Sci. 2017, 8, 1323. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Liu, G.; Liu, J.; Zhang, B.; Meng, W.; Müller, B.; Hayashi, K.; Zhang, X.; Zhao, Z.; De Smet, I.; Ding, Z. Synergistic action of auxin and cytokinin mediates aluminum-induced root growth inhibition in Arabidopsis. EMBO Rep. 2017, 18, 1213–1230. [Google Scholar] [CrossRef] [PubMed]
- Gemrotová, M.; Kulkarni, M.G.; Stirk, W.A.; Strnad, M.; Van Staden, J.; Spíchal, L. Seedlings of medicinal plants treated with either a cytokinin antagonist (PI-55) or an inhibitor of cytokinin degradation (INCYDE) are protected against the negative effects of cadmium. Plant Growth Regul. 2013, 71, 137–145. [Google Scholar] [CrossRef]
- Fukudome, A.; Aksoy, E.; Wu, X.; Kumar, K.; Jeong, I.S.; May, K.; Russell, W.K.; Koiwa, H. Arabidopsis CPL4 is an essential C-terminal domain phosphatase that suppresses xenobiotic stress responses. Plant J. 2014, 80, 27–39. [Google Scholar] [CrossRef] [PubMed]
- Ramel, F.; Sulmon, C.; Cabello-Hurtado, F.; Taconnat, L.; Martin-Magniette, M.L.; Renou, J.P.; El Amrani, A.; Couée, I.; Gouesbet, G. Genome-wide interacting effects of sucrose and herbicide-mediated stress in Arabidopsis thaliana: Novel insights into atrazine toxicity and sucrose-induced tolerance. BMC Genom. 2007, 8, 450. [Google Scholar] [CrossRef] [PubMed]
- Ramel, F.; Sulmon, C.; Serra, A.A.; Gouesbet, G.; Couée, I. Xenobiotic sensing and signalling in higher plants. J. Exp. Bot. 2012, 63, 3999–4014. [Google Scholar] [CrossRef] [PubMed]
- Brenner, W.G.; Schmulling, T. Transcript profiling of cytokinin action in Arabidopsis roots and shoots discovers largely similar but also organ-specific responses. BMC Plant Biol. 2012, 12, 112. [Google Scholar] [CrossRef] [PubMed]
- Brenner, W.G.; Schmülling, T. Summarizing and exploring data of a decade of cytokinin-related transcriptomics. Front. Plant Sci. 2015, 6, 29. [Google Scholar] [CrossRef] [PubMed]
- Daryanto, S.; Wang, L.; Jacinthe, P.A. Global Synthesis of Drought Effects on Maize and Wheat Production. PLoS ONE 2016, 11, e0156362. [Google Scholar] [CrossRef] [PubMed]
- Clauw, P.; Coppens, F.; De Beuf, K.; Dhondt, S.; Van Daele, T.; Maleux, K.; Storme, V.; Clement, L.; Gonzalez, N.; Inzé, D. Leaf Responses to Mild Drought Stress in Natural Variants of Arabidopsis. Plant Physiol. 2015, 167, 800–816. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Huang, B. Effects of foliar-applied ethylene inhibitor and synthetic cytokinin on creeping bentgrass to enhance heat tolerance. Crop Sci. 2009, 49, 1876–1884. [Google Scholar] [CrossRef]
- Xu, S.; Brockmöller, T.; Navarro-Quezada, A.; Kuhl, H.; Gase, K.; Ling, Z.; Zhou, W.; Kreitzer, C.; Stanke, M.; Tang, H.; et al. Wild tobacco genomes reveal the evolution of nicotine biosynthesis. Proc. Natl. Acad. Sci. USA 2017, 114, 6133–6138. [Google Scholar] [CrossRef] [PubMed]
- Le, D.T.; Nishiyama, R.; Watanabe, Y.; Vankova, R.; Tanaka, M.; Seki, M.; Ham, L.H.; Yamaguchi-Shinozaki, K.; Shinozaki, K.; Tran, L.S.P. Identification and expression analysis of cytokinin metabolic genes in soybean under normal and drought conditions in relation to cytokinin levels. PLoS ONE 2012, 7, e42411. [Google Scholar] [CrossRef] [PubMed]
- Dobra, J.; Motyka, V.; Dobrev, P.; Malbeck, J.; Prasil, I.T.; Haisel, D.; Gaudinova, A.; Havlova, M.; Gubis, J.; Vankova, R. Comparison of hormonal responses to heat, drought and combined stress in tobacco plants with elevated proline content. J. Plant Physiol. 2010, 167, 1360–1370. [Google Scholar] [CrossRef] [PubMed]
- Nishiyama, R.; Watanabe, Y.; Fujita, Y.; Le, D.T.; Kojima, M.; Werner, T.; Vankova, R.; Yamaguchi-Shinozaki, K.; Shinozaki, K.; Kakimoto, T.; et al. Analysis of Cytokinin Mutants and Regulation of Cytokinin Metabolic Genes Reveals Important Regulatory Roles of Cytokinins in Drought, Salt and Abscisic Acid Responses, and Abscisic Acid Biosynthesis. Plant Cell 2011, 23, 2169–2183. [Google Scholar] [CrossRef] [PubMed]
- Bano, A.; Dorffling, K.; Bettin, D.; Hahn, H. Abscisic acid and cytokinins as possible root-to-shoot signals in xylem sap of rice plants in drying soils. Funct. Plant Biol. 1993, 20, 109–115. [Google Scholar] [CrossRef]
- Argueso, C.T.; Ferreira, F.J.; Kieber, J.J. Environmental perception avenues: The interaction of cytokinin and environmental response pathways. Plant Cell Environ. 2009, 32, 1147–1160. [Google Scholar] [CrossRef] [PubMed]
- Tran, L.S.; Urao, T.; Qin, F.; Maruyama, K.; Kakimoto, T.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Functional analysis of AHK1/ATHK1 and cytokinin receptor histidine kinases in response to abscisic acid, drought, and salt stress in Arabidopsis. Proc. Natl. Acad. Sci. USA 2007, 104, 20623–20628. [Google Scholar] [CrossRef] [PubMed]
- Jang, G.; Chang, S.H.; Um, T.Y.; Lee, S.; Kim, J.K.; Choi, Y.D. Antagonistic interaction between jasmonic acid and cytokinin in xylem development. Sci. Rep. 2017, 7, 10212. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Hou, L.; Meng, J.; You, H.; Li, Z.; Gong, Z.; Yang, S.; Shi, Y. The Antagonistic Action of Abscisic Acid and Cytokinin Signaling Mediates Drought Stress Response in Arabidopsis. Mol. Plant 2018, 11, 970–982. [Google Scholar] [CrossRef] [PubMed]
- Rivero, R.M.; Ruiz, J.M.; García, P.C.; López-Lefebre, L.R.; Sánchez, E.; Romero, L. Resistance to cold and heat stress: Accumulation of phenolic compounds in tomato and watermelon plants. Plant Sci. 2001, 160, 315–321. [Google Scholar] [CrossRef]
- Ghanem, M.E.; Albacete, A.; Smigocki, A.C.; Frébort, I.; Pospísilová, H.; Martínez-Andújar, C.; Acosta, M.; Sánchez-Bravo, J.; Lutts, S.; Dodd, I.C.; Pérez-Alfocea, F. Root-synthesized cytokinins improve shoot growth and fruit yield in salinized tomato (Solanum lycopersicum L.) plants. J. Exp. Bot. 2011, 62, 125–140. [Google Scholar] [CrossRef] [PubMed]
- Rivero, R.M.; Kojima, M.; Gepstein, A.; Sakakibara, H.; Mittler, R.; Gepstein, S.; Blumwald, E. Delayed leaf senescence induces extreme drought tolerance in a flowering plant. Proc. Natl. Acad. Sci. USA 2007, 104, 19631–19636. [Google Scholar] [CrossRef] [PubMed]
- Rivero, R.M.; Gimeno, J.; Van Deynze, A.; Walia, H.; Blumwald, E. Enhanced Cytokinin Synthesis in Tobacco Plants Expressing PSARK::IPT Prevents the Degradation of Photosynthetic Protein Complexes During Drought. Plant Cell Physiol. 2010, 51, 1929–1941. [Google Scholar] [CrossRef] [PubMed]
- Reguera, M.; Peleg, Z.; Abdel-Tawab, Y.M.; Tumimbang, E.B.; Delatorre, C.A.; Blumwald, E. Stress-Induced Cytokinin Synthesis Increases Drought Tolerance through the Coordinated Regulation of Carbon and Nitrogen Assimilation in Rice. Plant Physiol. 2013, 163, 1609–1622. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Zhang, J.; Huang, B. Cytokinin-mitigation of salt-induced leaf senescence in perennial ryegrass involving the activation of antioxidant systems and ionic balance. Environ. Exp. Bot. 2016, 125, 1–11. [Google Scholar] [CrossRef]
- Décima Oneto, C.; Otegui, M.E.; Baroli, I.; Beznec, A.; Faccio, P.; Bossio, E.; Blumwald, E.; Lewi, D. Water deficit stress tolerance in maize conferred by expression of an isopentenyltransferase (IPT) gene driven by a stress- and maturation-induced promoter. J. Biotechnol. 2016, 220, 66–77. [Google Scholar] [CrossRef] [PubMed]
- Vojta, P.; Kokáš, F.; Husičková, A.; Grúz, J.; Bergougnoux, V.; Marchetti, C.F.; Jiskrová, E.; Ježilová, E.; Mik, V.; Ikeda, Y.; Galuszka, P. Whole transcriptome analysis of transgenic barley with altered cytokinin homeostasis and increased tolerance to drought stress. New Biotechnol. 2016, 33, 676–691. [Google Scholar] [CrossRef] [PubMed]
- Ramireddy, E.; Hosseini, S.A.; Eggert, K.; Gillandt, S.; Gnad, H.; von Wirén, N.; Schmülling, T. Root Engineering in Barley: Increasing Cytokinin Degradation Produces a Larger Root System, Mineral Enrichment in the Shoot and Improved Drought Tolerance. Plant Physiol. 2018, 177, 1078–1095. [Google Scholar] [CrossRef] [PubMed]
- Černý, M.; Kuklová, A.; Hoehenwarter, W.; Fragner, L.; Novák, O.; Rotková, G.; Jedelský, P.L.P. L.; Žáková, K. K.; Šmehilová, M.; Strnad, M.; et al. Proteome and metabolome profiling of cytokinin action in Arabidopsis identifying both distinct and similar responses to cytokinin down- and up-regulation. J. Exp. Bot. 2013, 64, 4193–4206. [Google Scholar] [CrossRef] [PubMed]
- Zavaleta-Mancera, H.A.; López-Delgado, H.; Loza-Tavera, H.; Mora-Herrera, M.; Trevilla-García, C.; Vargas-Suárez, M.; Ougham, H. Cytokinin promotes catalase and ascorbate peroxidase activities and preserves the chloroplast integrity during dark-senescence. J. Plant Physiol. 2007, 164, 1572–1582. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Zhang, J.; Burgess, P.; Rossi, S.; Huang, B. Interactive effects of melatonin and cytokinin on alleviating drought-induced leaf senescence in creeping bentgrass (Agrostis stolonifera). Environ. Exp. Bot. 2018, 145, 1–11. [Google Scholar] [CrossRef]
- Liao, X.; Guo, X.; Wang, Q.; Wang, Y.; Zhao, D.; Yao, L.; Wang, S.; Liu, G.; Li, T. Overexpression of MsDREB6.2 results in cytokinin-deficient developmental phenotypes and enhances drought tolerance in transgenic apple plants. Plant J. 2017, 89, 510–526. [Google Scholar] [CrossRef] [PubMed]
- Nakabayashi, R.; Yonekura-Sakakibara, K.; Urano, K.; Suzuki, M.; Yamada, Y.; Nishizawa, T.; Matsuda, F.; Kojima, M.; Sakakibara, H.; Shinozaki, K.; et al. Enhancement of oxidative and drought tolerance in Arabidopsis by overaccumulation of antioxidant flavonoids. Plant J. 2014, 77, 367–379. [Google Scholar] [CrossRef] [PubMed]
- Novák, J.; Pavlů, J.; Novák, O.; Nožková-Hlaváčková, V.; Špundová, M.; Hlavinka, J.; Koukalová, Š.; Skalák, J.; Černý, M.; Brzobohatý, B. High cytokinin levels induce a hypersensitive-like response in tobacco. Ann. Bot. 2013, 112, 41–55. [Google Scholar] [CrossRef] [PubMed]
- White, R.G.; Kirkegaard, J.A. The distribution and abundance of wheat roots in a dense, structured subsoil—Implications for water uptake. Plant Cell Environ. 2010, 33, 133–148. [Google Scholar] [CrossRef] [PubMed]
- Novák, J.; Černý, M.; Pavlů, J.; Zemánková, J.; Skalák, J.; Plačková, L.; Brzobohatý, B. Roles of proteome dynamics and cytokinin signaling in root to hypocotyl ratio changes induced by shading roots of Arabidopsis seedlings. Plant Cell Physiol. 2015, 56, 1006–1018. [Google Scholar] [CrossRef] [PubMed]
- Laplaze, L.; Benkova, E.; Casimiro, I.; Maes, L.; Vanneste, S.; Swarup, R.; Weijers, D.; Calvo, V.; Parizot, B.; Herrera-Rodriguez, M.B.; et al. Cytokinins act directly on lateral root founder cells to inhibit root initiation. Plant Cell 2007, 19, 3889–3900. [Google Scholar] [CrossRef] [PubMed]
- Pospíšilová, H.; Jiskrová, E.; Vojta, P.; Mrízová, K.; Kokáš, F.; Čudejková, M.M.; Bergougnoux, V.; Plíhal, O.; Klimešová, J.; Novák, O.; et al. Transgenic barley overexpressing a cytokinin dehydrogenase gene shows greater tolerance to drought stress. New Biotechnol. 2016, 33, 692–705. [Google Scholar] [CrossRef] [PubMed]
- Jang, G.; Choi, Y.D. Drought stress promotes xylem differentiation by modulating the interaction between cytokinin and jasmonic acid. Plant Signal. Behav. 2018, 13, e1451707. [Google Scholar] [CrossRef] [PubMed]
- Guan, C.; Wang, X.; Feng, J.; Hong, S.; Liang, Y.; Ren, B.; Zuo, J. Cytokinin Antagonizes Abscisic Acid-Mediated Inhibition of Cotyledon Greening by Promoting the Degradation of ABSCISIC ACID INSENSITIVE5 Protein in Arabidopsis. Plant Physiol. 2014, 164, 1515–1526. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Sun, M.M.; Ye, Q.; Wu, X.Q.; Wu, W.H.; Chen, Y.F. Abscisic Acid Modulates Seed Germination via ABA INSENSITIVE5-Mediated PHOSPHATE1. Plant Physiol. 2017, 175, 1661–1668. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.Y.; Huang, Y.C.; Ou, S.L. ERF73/HRE1 is involved in H2O2 production via hypoxia-inducible Rboh gene expression in hypoxia signaling. Protoplasma 2017, 254, 1705–1714. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.Q.; Emery, R.J.N. Is ABA the earliest upstream inhibitor of apical dominance? J. Exp. Bot. 2017, 68, 881–884. [Google Scholar] [CrossRef]
- Prerostova, S.; Dobrev, P.I.; Gaudinova, A.; Knirsch, V.; Körber, N.; Pieruschka, R.; Fiorani, F.; Brzobohatý, B.; Černý, M.; Spichal, L.; et al. Cytokinins: Their Impact on Molecular and Growth Responses to Drought Stress and Recovery in Arabidopsis. Front. Plant Sci. 2018, 9, 655. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Wu, H.; Ma, S.; Xiang, D.; Liu, R.; Xiong, L. OsJAZ1 Attenuates Drought Resistance by Regulating JA and ABA Signaling in Rice. Front. Plant Sci. 2017, 8, 2108. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, P.; Rasool, S.; Gul, A.; Sheikh, S.A.; Akram, N.A.; Ashraf, M.; Kazi, A.M.; Gucel, S. Jasmonates: Multifunctional Roles in Stress Tolerance. Front. Plant Sci. 2016, 7, 813. [Google Scholar] [CrossRef] [PubMed]
- Veselova, S.V.; Farhutdinov, R.G.; Veselov, S.Y.; Kudoyarova, G.R.; Veselov, D.S.; Hartung, W. The effect of root cooling on hormone content, leaf conductance and root hydraulic conductivity of durum wheat seedlings (Triticum durum L.). J. Plant Physiol. 2005, 162, 21–26. [Google Scholar] [CrossRef] [PubMed]
- Ntatsi, G.; Savvas, D.; Papasotiropoulos, V.; Katsileros, A.; Zrenner, R.M.; Hincha, D.K.; Zuther, E.; Schwarz, D. Rootstock Sub-Optimal Temperature Tolerance Determines Transcriptomic Responses after Long-Term Root Cooling in Rootstocks and Scions of Grafted Tomato Plants. Front. Plant Sci. 2017, 8, 911. [Google Scholar] [CrossRef] [PubMed]
- Maruyama, K.; Urano, K.; Yoshiwara, K.; Morishita, Y.; Sakurai, N.; Suzuki, H.; Kojima, M.; Sakakibara, H.; Shibata, D.; Saito, K.; et al. Integrated Analysis of the Effects of Cold and Dehydration on Rice Metabolites, Phytohormones, and Gene Transcripts. Plant Physiol. 2014, 164, 1759–1771. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Yang, Y.; Zhang, Q.; Liu, N.; Xu, Q.; Hu, L. Differential physiological and metabolic response to low temperature in two zoysiagrass genotypes native to high and low latitude. PLoS ONE 2018, 13, e0198885. [Google Scholar] [CrossRef] [PubMed]
- Jeon, J.; Kim, N.Y.; Kim, S.; Kang, N.Y.; Novák, O.; Ku, S.J.; Cho, C.; Lee, D.J.; Lee, E.J.; Strnad, M.; Kim, J. A subset of cytokinin two-component signaling system plays a role in cold temperature stress response in Arabidopsis. J. Biol. Chem. 2010, 285, 23371–23386. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Tian, S.; Hou, L.; Huang, X.; Zhang, X.; Guo, H.; Yang, S. Ethylene signaling negatively regulates freezing tolerance by repressing expression of CBF and type-A ARR genes in Arabidopsis. Plant Cell 2012, 24, 2578–2595. [Google Scholar] [CrossRef] [PubMed]
- Kang, N.Y.; Cho, C.; Kim, J. Inducible Expression of Arabidopsis Response Regulator 22 (ARR22), a Type-C ARR, in Transgenic Arabidopsis Enhances Drought and Freezing Tolerance. PLoS ONE 2013, 8, e79248. [Google Scholar] [CrossRef] [PubMed]
- Zwack, P.J.; Compton, M.A.; Adams, C.I.; Rashotte, A.M. Cytokinin response factor 4 (CRF4) is induced by cold and involved in freezing tolerance. Plant Cell Rep. 2016, 35, 573–584. [Google Scholar] [CrossRef] [PubMed]
- Jeon, J.; Cho, C.; Lee, M.R.; Van Binh, N.; Kim, J. CYTOKININ RESPONSE FACTOR2 (CRF2) and CRF3 Regulate Lateral Root Development in Response to Cold Stress in Arabidopsis. Plant Cell 2016, 28, 1828–1843. [Google Scholar] [CrossRef] [PubMed]
- Dobrá, J.; Černý, M.; Štorchová, H.; Dobrev, P.; Skalák, J.; Jedelský, P.L.; Lukšanová, H.; Gaudinová, A.; Pešek, B.; Malbecka, J.; et al. The impact of heat stress targeting on the hormonal and transcriptomic response in Arabidopsis. Plant Sci. 2015, 231, 52–61. [Google Scholar] [CrossRef] [PubMed]
- Skalák, J.; Černý, M.; Jedelský, P.; Dobrá, J.; Ge, E.; Novák, J.; Hronková, M.; Dobrev, P.; Vanková, R.; Brzobohatý, B. Stimulation of ipt overexpression as a tool to elucidate the role of cytokinins in high temperature responses of Arabidopsis thaliana. J. Exp. Bot. 2016, 67, 2861–2873. [Google Scholar] [CrossRef] [PubMed]
- Černý, M.; Jedelský, P.L.; Novák, J.; Schlosser, A.; Brzobohatý, B. Cytokinin modulates proteomic, transcriptomic and growth responses to temperature shocks in Arabidopsis. Plant Cell Environ. 2014, 37, 1641–1655. [Google Scholar] [CrossRef] [PubMed]
- Escandón, M.; Cañal, M.J.; Pascual, J.; Pinto, G.; Correia, B.; Amaral, J.; Meijón, M. Integrated physiological and hormonal profile of heat-induced thermotolerance in Pinus radiata. Tree Physiol. 2016, 36, 63–77. [Google Scholar] [CrossRef] [PubMed]
- Escandón, M.; Meijón, M.; Valledor, L.; Pascual, J.; Pinto, G.; Cañal, M.J. Metabolome Integrated Analysis of High-Temperature Response in Pinus radiata. Front. Plant Sci. 2018, 9, 485. [Google Scholar] [CrossRef] [PubMed]
- Lochmanová, G.; Zdráhal, Z.; Konečná, H.; Koukalová, Š.; Malbeck, J.; Souček, P.; Válková, M.; Kiran, N.S.; Brzobohatý, B. Cytokinin-induced photomorphogenesis in dark-grown Arabidopsis: A proteomic analysis. J. Exp. Bot. 2008, 59, 3705–3719. [Google Scholar] [CrossRef] [PubMed]
- Černý, M.; Dyčka, F.; Bobáľová, J.; Brzobohatý, B. Early cytokinin response proteins and phosphoproteins of Arabidopsis thaliana identified by proteome and phosphoproteome profiling. J. Exp. Bot. 2011, 62, 921–937. [Google Scholar] [CrossRef] [PubMed]
- Macková, H.; Hronková, M.; Dobrá, J.; Turečková, V.; Novák, O.; Lubovská, Z.; Motyka, V.; Haisel, D.; Hájek, T.; Prášil, I.T.; et al. Enhanced drought and heat stress tolerance of tobacco plants with ectopically enhanced cytokinin oxidase/dehydrogenase gene expression. J. Exp. Bot. 2013, 64, 2805–2815. [Google Scholar] [CrossRef] [PubMed]
- Danilova, M.N.; Kudryakova, N.V.; Doroshenko, A.S.; Zabrodin, D.A.; Vinogradov, N.S.; Kuznetsov, V.V. Molecular and physiological responses of Arabidopsis thaliana plants deficient in the genes responsible for ABA and cytokinin reception and metabolism to heat shock. Russ. J. Plant Physiol. 2016, 63, 308–318. [Google Scholar] [CrossRef]
- Yang, Y.; Jiang, Y.; Mi, X.; Gan, L.; Gu, T.; Ding, J.; Li, Y. Identification and expression analysis of cytokinin response regulators in Fragaria vesca. Acta Physiol. Plant. 2016, 38, 198. [Google Scholar] [CrossRef]
- Mi, X.; Wang, X.; Wu, H.; Gan, L.; Ding, J.; Li, Y. Characterization and expression analysis of cytokinin biosynthesis genes in Fragaria vesca. Plant Growth Regul. 2017, 82, 139–149. [Google Scholar] [CrossRef]
- Cortleven, A.; Schmülling, T. Regulation of chloroplast development and function by cytokinin. J. Exp. Bot. 2015, 66, 4999–5013. [Google Scholar] [CrossRef] [PubMed]
- Sweere, U.; Eichenberg, K.; Lohrmann, J.; Mira-Rodado, V.; Bäurle, I.; Kudla, J.; Nagy, F.; Schafer, E.; Harter, K. Interaction of the response regulator ARR4 with phytochrome B in modulating red light signaling. Science 2001, 294, 1108–1111. [Google Scholar] [CrossRef] [PubMed]
- Chi, W.; Li, J.; He, B.; Chai, X.; Xu, X.; Sun, X.; Jiang, J.; Feng, P.; Zuo, J.; Lin, R.; et al. DEG9, a serine protease, modulates cytokinin and light signaling by regulating the level of ARABIDOPSIS RESPONSE REGULATOR 4. Proc. Natl. Acad. Sci. USA 2016, 113, E3568–E3576. [Google Scholar] [CrossRef] [PubMed]
- Dobisova, T.; Hrdinova, V.; Cuesta, C.; Michlickova, S.; Urbankova, I.; Hejatkova, R.; Zadnikova, P.; Pernisova, M.; Benkova, E.; Hejatko, J. Light Controls Cytokinin Signaling via Transcriptional Regulation of Constitutively Active Sensor Histidine Kinase CKI1. Plant Physiol. 2017, 174, 387–404. [Google Scholar] [CrossRef] [PubMed]
- Nováková, M.; Motyka, V.; Dobrev, P.I.; Malbeck, J.; Gaudinová, A.; Vanková, R. Diurnal variation of cytokinin, auxin and abscisic acid levels in tobacco leaves. J. Exp. Bot. 2005, 56, 2877–2883. [Google Scholar] [CrossRef] [PubMed]
- Edwards, K.D.; Takata, N.; Johansson, M.; Jurca, M.; Novák, O.; Hényková, E.; Liverani, S.; Kozarewa, I.; Strnad, M.; Millar, A.J.; et al. Circadian clock components control daily growth activities by modulating cytokinin levels and cell division-associated gene expression in Populus trees. Plant Cell Environ. 2018, 41, 1468–1482. [Google Scholar] [CrossRef] [PubMed]
- Nitschke, S.; Cortleven, A.; Iven, T.; Feussner, I.; Havaux, M.; Riefler, M.; Schmülling, T. Circadian stress regimes affect the circadian clock and cause jasmonic acid-dependent cell death in cytokinin-deficient Arabidopsis plants. Plant Cell 2016, 28, 1616–1639. [Google Scholar] [CrossRef] [PubMed]
- Janečková, H.; Husičková, A.; Ferretti, U.; Prčina, M.; Pilařová, E.; Plačková, L.; Pospíšil, P.; Doležal, K.; Špundová, M. The interplay between cytokinins and light during senescence in detached Arabidopsis leaves. Plant Cell Environ. 2018, 41, 1870–1885. [Google Scholar] [CrossRef] [PubMed]
- Vandenbussche, F.; Habricot, Y.; Condiff, A.S.; Maldiney, R.; Van der Straeten, D.; Ahmad, M. HY5 is a point of convergence between cryptochrome and cytokinin signalling pathways in Arabidopsis thaliana. Plant J. 2007, 49, 428–441. [Google Scholar] [CrossRef] [PubMed]
- Cortleven, A.; Nitschke, S.; Klaumunzer, M.; AbdElgawad, H.; Asard, H.; Grimm, B.; Riefler, M.; Schmulling, T. A Novel Protective Function for Cytokinin in the Light Stress Response Is Mediated by the ARABIDOPSIS HISTIDINE KINASE2 and ARABIDOPSIS HISTIDINE KINASE3 Receptors. Plant Physiol. 2014, 164, 1470–1483. [Google Scholar] [CrossRef] [PubMed]
- Danilova, M.N.; Kudryakova, N.V.; Voronin, P.Y.; Oelmüller, R.; Kusnetsov, V.V.; Kulaeva, O.N. Membrane receptors of cytokinin and their regulatory role in Arabidopsis thaliana plant response to photooxidative stress under conditions of water deficit. Russ. J. Plant Physiol. 2014, 61, 434–442. [Google Scholar] [CrossRef]
- Bashri, G.; Singh, M.; Mishra, R.K.; Kumar, J.; Singh, V.P.; Prasad, S.M. Kinetin Regulates UV-B-Induced Damage to Growth, Photosystem II Photochemistry, and Nitrogen Metabolism in Tomato Seedlings. J. Plant Growth Regul. 2018, 37, 233–245. [Google Scholar] [CrossRef]
- Patterson, K.; Cakmak, T.; Cooper, A.; Lager, I.; Rasmusson, A.G.; Escobar, M.A. Distinct signalling pathways and transcriptome response signatures differentiate ammonium- and nitrate-supplied plants. Plant Cell Environ. 2010, 33, 1486–14501. [Google Scholar] [CrossRef] [PubMed]
- Canales, J.; Moyano, T.C.; Villarroel, E.; Gutiérrez, R.A. Systems analysis of transcriptome data provides new hypotheses about Arabidopsis root response to nitrate treatments. Front. Plant Sci. 2014, 5, 22. [Google Scholar] [CrossRef] [PubMed]
- Krapp, A.; Berthome, R.; Orsel, M.; Mercey-Boutet, S.; Yu, A.; Castaings, L.; Elftieh, S.; Major, H.; Renou, J.-P.; Daniel-Vedele, F. Arabidopsis Roots and Shoots Show Distinct Temporal Adaptation Patterns toward Nitrogen Starvation. Plant Physiol. 2011, 157, 1255–1282. [Google Scholar] [CrossRef] [PubMed]
- Kiba, T.; Inaba, J.; Kudo, T.; Ueda, N.; Konishi, M.; Mitsuda, N.; Takiguchi, Y.; Kondou, Y.; Yoshizumi, T.; Ohme-Takagi, M.; et al. Repression of Nitrogen Starvation Responses by Members of the Arabidopsis GARP-Type Transcription Factor NIGT1/HRS1 Subfamily. Plant Cell 2018, 30, 925–945. [Google Scholar] [CrossRef] [PubMed]
- Pant, B.D.; Musialak-Lange, M.; Nuc, P.; May, P.; Buhtz, A.; Kehr, J.; Walther, D.; Scheible, W.-R. Identification of Nutrient-Responsive Arabidopsis and Rapeseed MicroRNAs by Comprehensive Real-Time Polymerase Chain Reaction Profiling and Small RNA Sequencing. Plant Physiol. 2009, 150, 1541–1555. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Ding, H.; Zhu, J.K.; Zhang, F.; Li, W.X. Involvement of miR169 in the nitrogen-starvation responses in Arabidopsis. New Phytol. 2011, 190, 906–915. [Google Scholar] [CrossRef] [PubMed]
- Konishi, N.; Ishiyama, K.; Matsuoka, K.; Maru, I.; Hayakawa, T.; Yamaya, T.; Kojima, S. NADH-dependent glutamate synthase plays a crucial role in assimilating ammonium in the Arabidopsis root. Physiol. Plant. 2014, 152, 138–151. [Google Scholar] [CrossRef] [PubMed]
- Bi, Y.M.; Wang, R.L.; Zhu, T.; Rothstein, S.J. Global transcription profiling reveals differential responses to chronic nitrogen stress and putative nitrogen regulatory components in Arabidopsis. BMC Genom. 2007, 8, 281. [Google Scholar] [CrossRef] [PubMed]
- Canales, J.; Rueda-López, M.; Craven-Bartle, B.; Avila, C.; Cánovas, F.M. Novel Insights into Regulation of Asparagine Synthetase in Conifers. Front. Plant Sci. 2012, 3, 100. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, J.M.; Riveras, E.; Vidal, E.A.; Gras, D.E.; Contreras-López, O.; Tamayo, K.P.; Aceituno, F.; Gómez, I.; Ruffel, S.; Lejay, L.; et al. Systems approach identifies TGA1 and TGA4 transcription factors as important regulatory components of the nitrate response of Arabidopsis thaliana roots. Plant J. 2014, 80, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Konishi, M.; Yanagisawa, S. Arabidopsis NIN-like transcription factors have a central role in nitrate signalling. Nat. Commun. 2013, 4, 1617. [Google Scholar] [CrossRef] [PubMed]
- Rubin, G.; Tohge, T.; Matsuda, F.; Saito, K.; Scheible, W.R. Members of the LBD Family of Transcription Factors Repress Anthocyanin Synthesis and Affect Additional Nitrogen Responses in Arabidopsis. Plant Cell 2009, 21, 3567–3584. [Google Scholar] [CrossRef] [PubMed]
- Jost, R.; Pharmawati, M.; Lapis-Gaza, H.R.; Rossig, C.; Berkowitz, O.; Lambers, H.; Finnegan, P.M. Differentiating phosphate-dependent and phosphate-independent systemic phosphate-starvation response networks in Arabidopsis thaliana through the application of phosphite. J. Exp. Bot. 2015, 66, 2501–2514. [Google Scholar] [CrossRef] [PubMed]
- Hammond, J.P.; Bennett, M.J.; Bowen, H.C.; Broadley, M.R.; Eastwood, D.C.; May, S.T.; Rahn, C.; Swarup, R.; Woolaway, K.E.; White, P.J. Changes in Gene Expression in Arabidopsis Shoots during Phosphate Starvation and the Potential for Developing Smart Plants. Plant Physiol. 2003, 132, 578–596. [Google Scholar] [CrossRef] [PubMed]
- Ayadi, A.; David, P.; Arrighi, J.F.; Chiarenza, S.; Thibaud, M.C.; Nussaume, L.; Marin, E. Reducing the Genetic Redundancy of Arabidopsis PHOSPHATE TRANSPORTER1 Transporters to Study Phosphate Uptake and Signaling. Plant Physiol. 2015, 167, 1511–1526. [Google Scholar] [CrossRef] [PubMed]
- Gu, M.; Chen, A.; Sun, S.; Xu, G. Complex Regulation of Plant Phosphate Transporters and the Gap between Molecular Mechanisms and Practical Application: What Is Missing? Mol. Plant 2016, 9, 396–416. [Google Scholar] [CrossRef] [PubMed]
- Lapis-Gaza, H.R.; Jost, R.; Finnegan, P.M. Arabidopsis PHOSPHATE TRANSPORTER1 genes PHT1;8 and PHT1;9 are involved in root-to-shoot translocation of orthophosphate. BMC Plant Biol. 2014, 14, 334. [Google Scholar] [CrossRef] [PubMed]
- Puga, M.I.; Mateos, I.; Charukesi, R.; Wang, Z.; Franco-Zorrilla, J.M.; de Lorenzo, L.; Irigoyen, M.L.; Masiero, S.; Bustos, R.; Rodriguez, J.; et al. SPX1 is a phosphate-dependent inhibitor of PHOSPHATE STARVATION RESPONSE 1 in Arabidopsis. Proc. Natl. Acad. Sci. USA 2014, 111, 14947–14952. [Google Scholar] [CrossRef] [PubMed]
- Maruyama-Nakashita, A.; Nakamura, Y.; Tohge, T.; Saito, K.; Takahashi, H. Arabidopsis SLIM1 is a central transcriptional regulator of plant sulfur response and metabolism. Plant Cell 2006, 18, 3235–3251. [Google Scholar] [CrossRef] [PubMed]
- Bielecka, M.; Watanabe, M.; Morcuende, R.; Scheible, W.R.; Hawkesford, M.J.; Hesse, H.; Hoefgen, R. Transcriptome and metabolome analysis of plant sulfate starvation and resupply provides novel information on transcriptional regulation of metabolism associated with sulfur, nitrogen and phosphorus nutritional responses in Arabidopsis. Front. Plant Sci. 2014, 5, 805. [Google Scholar] [CrossRef] [PubMed]
- Nikiforova, V.; Freitag, J.; Kempa, S.; Adamik, M.; Hesse, H.; Hoefgen, R. Transcriptome analysis of sulfur depletion in Arabidopsis thaliana: Interlacing of biosynthetic pathways provides response specificity. Plant J. 2003, 33, 633–650. [Google Scholar] [CrossRef] [PubMed]
- Kopriva, S.; Calderwood, A.; Weckopp, S.C.; Koprivova, A. Plant sulfur and Big Data. Plant Sci. 2015, 241, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Pasini, R.; Dan, H.; Joshi, N.; Zhao, Y.; Leustek, T.; Zheng, Z.L. Aberrant gene expression in the Arabidopsis SULTR1;2 mutants suggests a possible regulatory role for this sulfate transporter in response to sulfur nutrient status. Plant J. 2014, 77, 185–197. [Google Scholar] [CrossRef] [PubMed]
- Henríquez-Valencia, C.; Arenas-M, A.; Medina, J.; Canales, J. Integrative Transcriptomic Analysis Uncovers Novel Gene Modules That Underlie the Sulfate Response in Arabidopsis thaliana. Front. Plant Sci. 2018, 9, 470. [Google Scholar] [CrossRef] [PubMed]
- Kawashima, C.G.; Berkowitz, O.; Hell, R.; Noji, M.; Saito, K. Characterization and expression analysis of a serine acetyltransferase gene family involved in a key step of the sulfur assimilation pathway in Arabidopsis. Plant Physiol. 2005, 137, 220–230. [Google Scholar] [CrossRef] [PubMed]
- Forieri, I.; Sticht, C.; Reichelt, M.; Gretz, N.; Hawkesford, M.J.; Malagoli, M.; Wirtz, M.; Hell, R. System analysis of metabolism and the transcriptome in Arabidopsis thaliana roots reveals differential co-regulation upon iron, sulfur and potassium deficiency. Plant Cell Environ. 2017, 40, 95–107. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Cao, B.; Wang, S.; Liu, P.; Deng, X.; Yin, L.; Zhang, S. Silicon moderated the K deficiency by improving the plant-water status in sorghum. Sci. Rep. 2016, 6, 22882. [Google Scholar] [CrossRef] [PubMed]
- Gierth, M.; Mäser, P.; Schroeder, J.I. The Potassium Transporter AtHAK5 Functions in K+ Deprivation-Induced High-Affinity K+ Uptake and AKT1 K+ Channel Contribution to K+ Uptake Kinetics in Arabidopsis Roots. Plant Physiol. 2005, 137, 1105–1114. [Google Scholar] [CrossRef] [PubMed]
- Pyo, Y.J.; Gierth, M.; Schroeder, J.I.; Cho, M.H. High-Affinity K+ Transport in Arabidopsis: AtHAK5 and AKT1 Are Vital for Seedling Establishment and Postgermination Growth under Low-Potassium Conditions. Plant Physiol. 2010, 153, 863–875. [Google Scholar] [CrossRef] [PubMed]
- Ragel, P.; Ródenas, R.; García-Martín, E.; Andrés, Z.; Villalta, I.; Nieves-Cordones, M.; Rivero, R.M.; Martínez, V.; Pardo, J.M.; Quintero, F.J.; Rubio, F. CIPK23 regulates HAK5-mediated high-affinity K+ uptake in Arabidopsis roots. Plant Physiol. 2015, 169, 01401.2015. [Google Scholar] [CrossRef] [PubMed]
- Osakabe, Y.; Arinaga, N.; Umezawa, T.; Katsura, S.; Nagamachi, K.; Tanaka, H.; Ohiraki, H.; Yamada, K.; Seo, S.-U.; Abo, M.; et al. Osmotic Stress Responses and Plant Growth Controlled by Potassium Transporters in Arabidopsis. Plant Cell 2013, 25, 609–624. [Google Scholar] [CrossRef] [PubMed]
- Pilot, G.; Gaymard, F.; Mouline, K.; Chérel, I.; Sentenac, H. Regulated expression of Arabidopsis shaker K+ channel genes involved in K+ uptake and distribution in the plant. Plant Mol. Biol. 2003, 51, 773–787. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wu, W.H. Regulation of potassium transport and signaling in plants. Curr. Opin. Plant Biol. 2017, 39, 123–128. [Google Scholar] [CrossRef] [PubMed]
- Rigas, S.; Debrosses, G.; Haralampidis, K.; Vicente-Agullo, F.; Feldmann, K.A.; Grabov, A.; Dolan, L.; Hatzopoulos, P. TRH1 encodes a potassium transporter required for tip growth in Arabidopsis root hairs. Plant Cell 2001, 13, 139–151. [Google Scholar] [CrossRef] [PubMed]
- Han, M.; Wu, W.; Wu, W.-H.; Wang, Y. Potassium Transporter KUP7 Is Involved in K+ Acquisition and Translocation in Arabidopsis Root under K+ -Limited Conditions. Mol. Plant 2016, 9, 437–446. [Google Scholar] [CrossRef] [PubMed]
- Armengaud, P.; Breitling, R.; Amtmann, A. The Potassium-Dependent Transcriptome of Arabidopsis Reveals a Prominent Role of Jasmonic Acid in Nutrient Signaling. Plant Physiol. 2004, 136, 2556–2576. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Lan, P. The Understanding of the Plant Iron Deficiency Responses in Strategy I Plants and the Role of Ethylene in This Process by Omic Approaches. Front. Plant Sci. 2017, 8, 40. [Google Scholar] [CrossRef] [PubMed]
- Mai, H.J.; Pateyron, S.; Bauer, P. Iron homeostasis in Arabidopsis thaliana: Transcriptomic analyses reveal novel FIT-regulated genes, iron deficiency marker genes and functional gene networks. BMC Plant Biol. 2016, 16, 211. [Google Scholar] [CrossRef] [PubMed]
- Stein, R.J.; Waters, B.M. Use of natural variation reveals core genes in the transcriptome of iron-deficient Arabidopsis thaliana roots. J. Exp. Bot. 2012, 63, 1039–1055. [Google Scholar] [CrossRef] [PubMed]
- Mendoza-Cózatl, D.G.; Xie, Q.; Akmakjian, G.Z.; Jobe, T.O.; Patel, A.; Stacey, M.G.; Song, L.; Demoin, D.W.; Jurisson, S.S.; Stacey, G.; Schroeder, J.I. OPT3 Is a Component of the Iron-Signaling Network between Leaves and Roots and Misregulation of OPT3 Leads to an Over-Accumulation of Cadmium in Seeds. Mol. Plant 2014, 7, 1455–1469. [Google Scholar] [CrossRef] [PubMed]
- Takano, J.; Noguchi, K.; Yasumori, M.; Kobayashi, M.; Gajdos, Z.; Miwa, K.; Hayashi, H.; Yoneyama, T.; Fujiwara, T. Arabidopsis boron transporter for xylem loading. Nature 2002, 420, 337–340. [Google Scholar] [CrossRef] [PubMed]
- Miwa, K.; Aibara, I.; Fujiwara, T. Arabidopsis thaliana BOR4 is upregulated under high boron conditions and confers tolerance to high boron. Soil Sci. Plant Nutr. 2014, 60, 349–355. [Google Scholar] [CrossRef]
- Msanne, J.; Lin, J.; Stone, J.M.; Awada, T. Characterization of abiotic stress-responsive Arabidopsis thaliana RD29A and RD29B genes and evaluation of transgenes. Planta 2011, 234, 97–107. [Google Scholar] [CrossRef] [PubMed]
- Huang, K.C.; Lin, W.C.; Cheng, W.H. Salt hypersensitive mutant 9, a nucleolar APUM23 protein, is essential for salt sensitivity in association with the ABA signaling pathway in Arabidopsis. BMC Plant Biol. 2018, 18, 40. [Google Scholar] [CrossRef] [PubMed]
- Vogel, J.T.; Zarka, D.G.; Van Buskirk, H.A.; Fowler, S.G.; Thomashow, M.F. Roles of the CBF2 and ZAT12 transcription factors in configuring the low temperature transcriptome of Arabidopsis. Plant J. 2005, 41, 195–211. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.; Henderson, D.A.; Zhu, J.K. The Arabidopsis cold-responsive transcriptome and its regulation by ICE1. Plant Cell 2005, 17, 3155–3175. [Google Scholar] [CrossRef] [PubMed]
- Maruyama, K.; Sakuma, Y.; Kasuga, M.; Ito, Y.; Seki, M.; Goda, H.; Shimada, Y.; Yoshida, S.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Identification of cold-inducible downstream genes of the Arabidopsis DREB1A/CBF3 transcriptional factor using two microarray systems. Plant J. 2004, 38, 982–993. [Google Scholar] [CrossRef] [PubMed]
- Nakaminami, K.; Matsui, A.; Nakagami, H.; Minami, A.; Nomura, Y.; Tanaka, M.; Morosawa, T.; Ishida, J.; Takahashi, S.; Uemura, M.; et al. Analysis of differential expression patterns of mRNA and protein during cold-acclimation and de-acclimation in Arabidopsis. Mol. Cell. Proteom. 2014, 13, 3602–3611. [Google Scholar] [CrossRef] [PubMed]
- Janská, A.; Aprile, A.; Zámečník, J.; Cattivelli, L.; Ovesná, J. Transcriptional responses of winter barley to cold indicate nucleosome remodelling as a specific feature of crown tissues. Funct. Integr. Genom. 2011, 11, 307–325. [Google Scholar] [CrossRef] [PubMed]
- Teige, M.; Scheikl, E.; Eulgem, T.; Dóczi, R.; Ichimura, K.; Shinozaki, K.; Dangl, J.L.; Hirt, H. The MKK2 Pathway Mediates Cold and Salt Stress Signaling in Arabidopsis. Mol. Cell 2004, 15, 141–152. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.L.; Chen, R.-G.; Du, X.H.; Zhang, Z.; Yin, Y.X.; Gong, Z.H.; Wang, G.Y. Reduced tolerance to abiotic stress in transgenic Arabidopsis overexpressing a Capsicum annuum multiprotein bridging factor 1. BMC Plant Biol. 2014, 14, 138. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.M.; Kang, B.R.; Kim, Y.C. Transcriptome Analysis of Induced Systemic Drought Tolerance Elicited by Pseudomonas chlororaphis O6 in Arabidopsis thaliana. Plant Pathol. J. 2013, 29, 209–220. [Google Scholar] [CrossRef] [PubMed]
- Rest, J.S.; Wilkins, O.; Yuan, W.; Purugganan, M.D.; Gurevitch, J. Meta-analysis and meta-regression of transcriptomic responses to water stress in Arabidopsis. Plant J. 2016, 85, 548–560. [Google Scholar] [CrossRef] [PubMed]
- MacGregor, D.R.; Penfield, S. Exploring the pleiotropy of hos1. J. Exp. Bot. 2015, 66, 1661–1671. [Google Scholar] [CrossRef] [PubMed]
- Simpson, S.D.; Nakashima, K.; Narusaka, Y.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Two different novel cis-acting elements of erd1, a clpA homologous Arabidopsis gene function in induction by dehydration stress and dark-induced senescence. Plant J. 2003, 33, 259–270. [Google Scholar] [CrossRef] [PubMed]
- Sakuma, Y.; Maruyama, K.; Qin, F.; Osakabe, Y.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Dual function of an Arabidopsis transcription factor DREB2A in water-stress-responsive and heat-stress-responsive gene expression. Proc. Natl. Acad. Sci. USA 2006, 103, 18822–18827. [Google Scholar] [CrossRef] [PubMed]
- Baek, D.; Chun, H.J.; Kang, S.; Shin, G.; Park, S.J.; Hong, H.; Kim, C.; Kim, D.H.; Lee, S.Y.; Kim, M.C.; et al. A Role for Arabidopsis miR399f in Salt, Drought, and ABA Signaling. Mol. Cells 2016, 39, 111–118. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, J.M.; Boisson-Dernier, A.; Dizon, M.B.; Maktabi, M.H.; Schroeder, J.I. The protein phosphatase AtPP2CA negatively regulates abscisic acid signal transduction in Arabidopsis, and effects of abh1 on AtPP2CA mRNA. Plant Physiol. 2006, 140, 127–139. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhang, X.; Fan, S. Meta-analysis of salt-related gene expression profiles identifies common signatures of salt stress responses in Arabidopsis. Plant Syst. Evol. 2017, 303, 757–774. [Google Scholar] [CrossRef]
- Yamada, K.; Fukao, Y.; Hayashi, M.; Fukazawa, M.; Suzuki, I.; Nishimura, M. Cytosolic HSP90 Regulates the Heat Shock Response That Is Responsible for Heat Acclimation in Arabidopsis thaliana. J. Biol. Chem. 2007, 282, 37794–37804. [Google Scholar] [CrossRef] [PubMed]
- Larkindale, J.; Vierling, E. Core genome responses involved in acclimation to high temperature. Plant Physiol. 2008, 146, 748–761. [Google Scholar] [CrossRef] [PubMed]
- Charng, Y.Y.; Liu, H.C.; Liu, N.Y.; Chi, W.T.; Wang, C.N.; Chang, S.H.; Wang, T.T. A Heat-Inducible Transcription Factor, HsfA2, Is Required for Extension of Acquired Thermotolerance in Arabidopsis. Plant Physiol. 2007, 143, 251–262. [Google Scholar] [CrossRef] [PubMed]
- Lin, K.F.; Tsai, M.Y.; Lu, C.A.; Wu, S.J.; Yeh, C.H. The roles of Arabidopsis HSFA2, HSFA4a, and HSFA7a in the heat shock response and cytosolic protein response. Bot. Stud. 2018, 59, 15. [Google Scholar] [CrossRef] [PubMed]
- Sung, D.Y.; Vierling, E.; Guy, C.L. Comprehensive expression profile analysis of the Arabidopsis Hsp70 gene family. Plant Physiol. 2001, 126, 789–800. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Dai, X.; Xu, W.; Ma, M. Overexpressing GSH1 and AsPCS1 simultaneously increases the tolerance and accumulation of cadmium and arsenic in Arabidopsis thaliana. Chemosphere 2008, 72, 1020–1026. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Feng, S.J.; Chen, J.; Zhao, W.T.; Yang, Z.M. A cadmium stress-responsive gene AtFC1 confers plant tolerance to cadmium toxicity. BMC Plant Biol. 2017, 17, 187. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yang, L.; Yan, X.; Liu, Y.; Wang, R.; Fan, T.; Ren, Y.; Tang, X.; Xiao, F.; Liu, Y.; et al. Zinc-Finger Transcription Factor ZAT6 Positively Regulates Cadmium Tolerance through the Glutathione-Dependent Pathway in Arabidopsis. Plant Physiol. 2016, 171, 707–719. [Google Scholar] [CrossRef] [PubMed]
- Song, W.Y.; Martinoia, E.; Lee, J.; Kim, D.; Kim, D.Y.; Vogt, E.; Shim, D.; Choi, K.S.; Hwang, I.; Lee, Y. A Novel Family of Cys-Rich Membrane Proteins Mediates Cadmium Resistance in Arabidopsis. Plant Physiol. 2004, 135, 1027–1039. [Google Scholar] [CrossRef] [PubMed]
- Morel, M.; Crouzet, J.; Gravot, A.; Auroy, P.; Leonhardt, N.; Vavasseur, A.; Richaud, P. AtHMA3, a P1B-ATPase Allowing Cd/Zn/Co/Pb Vacuolar Storage in Arabidopsis. Plant Physiol. 2008, 149, 894–904. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Martinoia, E.; Lee, Y. Vacuolar Transporters for Cadmium and Arsenic in Plants and their Applications in Phytoremediation and Crop Development. Plant Cell Physiol. 2018, 59, 1317–1325. [Google Scholar] [CrossRef] [PubMed]
- Mills, R.F.; Krijger, G.C.; Baccarini, P.J.; Hall, J.L.; Williams, L.E. Functional expression of AtHMA4, a P1B-type ATPase of the Zn/Co/Cd/Pb subclass. Plant J. 2003, 35, 164–176. [Google Scholar] [CrossRef] [PubMed]
- Van Hoewyk, D.; Takahashi, H.; Inoue, E.; Hess, A.; Tamaoki, M.; Pilon-Smits, E.A.H. Transcriptome analyses give insights into selenium-stress responses and selenium tolerance mechanisms in Arabidopsis. Physiol. Plant. 2008, 132, 236–253. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Zhang, Y.; Peng, J.S.; Zhong, C.; Yi, H.Y.; Ow, D.W.; Gong, J.M. Fission Yeast HMT1 Lowers Seed Cadmium through Phytochelatin-Dependent Vacuolar Sequestration in Arabidopsis. Plant Physiol. 2012, 158, 1779–1788. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.O.; Kang, H. Comparative expression analysis of genes encoding metallothioneins in response to heavy metals and abiotic stresses in rice (Oryza sativa) and Arabidopsis thaliana. Biosci. Biotechnol. Biochem. 2018, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Sanz-Fernández, M.; Rodríguez-Serrano, M.; Sevilla-Perea, A.; Pena, L.; Mingorance, M.D.; Sandalio, L.M.; Romero-Puertas, M.C. Screening Arabidopsis mutants in genes useful for phytoremediation. J. Hazard. Mater. 2017, 335, 143–151. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.Y.; Bovet, L.; Maeshima, M.; Martinoia, E.; Lee, Y. The ABC transporter AtPDR8 is a cadmium extrusion pump conferring heavy metal resistance. Plant J. 2007, 50, 207–218. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.Y.; Bovet, L.; Kushnir, S.; Noh, E.W.; Martinoia, E.; Lee, Y. AtATM3 Is Involved in Heavy Metal Resistance in Arabidopsis. Plant Physiol. 2006, 140, 922–932. [Google Scholar] [CrossRef] [PubMed]
- Park, J.; Song, W.Y.; Ko, D.; Eom, Y.; Hansen, T.H.; Schiller, M.; Lee, T.G.; Martinoia, E.; Lee, Y. The phytochelatin transporters AtABCC1 and AtABCC2 mediate tolerance to cadmium and mercury. Plant J. 2012, 69, 278–288. [Google Scholar] [CrossRef] [PubMed]
- Song, W.Y.; Park, J.; Mendoza-Cozatl, D.G.; Suter-Grotemeyer, M.; Shim, D.; Hortensteiner, S.; Geisler, M.; Weder, B.; Rea, P.A.; Rentsch, D.; et al. Arsenic tolerance in Arabidopsis is mediated by two ABCC-type phytochelatin transporters. Proc. Natl. Acad. Sci. USA 2010, 107, 21187–21192. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Bermejo, E.; Castrillo, G.; del Llano, B.; Navarro, C.; Zarco-Fernández, S.; Martinez-Herrera, D.J.; Leo-del Puerto, Y.; Muñoz, R.; Cámara, C.; Paz-Ares, J.; et al. Natural variation in arsenate tolerance identifies an arsenate reductase in Arabidopsis thaliana. Nat. Commun. 2014, 5, 4617. [Google Scholar] [CrossRef] [PubMed]
- Sawaki, Y.; Iuchi, S.; Kobayashi, Y.; Kobayashi, Y.; Ikka, T.; Sakurai, N.; Fujita, M.; Shinozaki, K.; Shibata, D.; Kobayashi, M.; et al. STOP1 regulates multiple genes that protect arabidopsis from proton and aluminum toxicities. Plant Physiol. 2009, 150, 281–294. [Google Scholar] [CrossRef] [PubMed]
- Iuchi, S.; Koyama, H.; Iuchi, A.; Kobayashi, Y.; Kitabayashi, S.; Kobayashi, Y.; Ikka, T.; Hirayama, T.; Shinozaki, K.; Kobayashi, M. Zinc finger protein STOP1 is critical for proton tolerance in Arabidopsis and coregulates a key gene in aluminum tolerance. Proc. Natl. Acad. Sci. USA 2007, 104, 9900–9905. [Google Scholar] [CrossRef] [PubMed]
- Larsen, P.B.; Geisler, M.J.B.; Jones, C.A.; Williams, K.M.; Cancel, J.D. ALS3 encodes a phloem-localized ABC transporter-like protein that is required for aluminum tolerance in Arabidopsis. Plant J. 2004, 41, 353–363. [Google Scholar] [CrossRef] [PubMed]
- Skipsey, M.; Knight, K.M.; Brazier-Hicks, M.; Dixon, D.P.; Steel, P.G.; Edwards, R. Xenobiotic Responsiveness of Arabidopsis thaliana to a Chemical Series Derived from a Herbicide Safener. J. Biol. Chem. 2011, 286, 32268–32276. [Google Scholar] [CrossRef] [PubMed]
- Gandia-Herrero, F.; Lorenz, A.; Larson, T.; Graham, I.A.; Bowles, D.J.; Rylott, E.L.; Bruce, N.C. Detoxification of the explosive 2,4,6-trinitrotoluene in Arabidopsis: Discovery of bifunctional O- and C-glucosyltransferases. Plant J. 2008, 56, 963–974. [Google Scholar] [CrossRef] [PubMed]
- Landa, P.; Prerostova, S.; Langhansova, L.; Marsik, P.; Vanek, T. Transcriptomic response of Arabidopsis thaliana (L.) Heynh. roots to ibuprofen. Int. J. Phytoremediation 2017, 19, 695–700. [Google Scholar] [CrossRef] [PubMed]
- Weisman, D.; Alkio, M.; Colón-Carmona, A. Transcriptional responses to polycyclic aromatic hydrocarbon-induced stress in Arabidopsis thaliana reveal the involvement of hormone and defense signaling pathways. BMC Plant Biol. 2010, 10, 59. [Google Scholar] [CrossRef] [PubMed]
- Behringer, C.; Bartsch, K.; Schaller, A. Safeners recruit multiple signalling pathways for the orchestrated induction of the cellular xenobiotic detoxification machinery in Arabidopsis. Plant Cell Environ. 2011, 34, 1970–1985. [Google Scholar] [CrossRef] [PubMed]
- Manabe, Y.; Tinker, N.; Colville, A.; Miki, B. CSR1, the Sole Target of Imidazolinone Herbicide in Arabidopsis thaliana. Plant Cell Physiol. 2007, 48, 1340–1358. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, S.; Schnoor, J.L.; Van Aken, B. Effects of Polychlorinated Biphenyls (PCBs) and Their Hydroxylated Metabolites (OH-PCBs) on Arabidopsis thaliana. Environ. Sci. Technol. 2017, 51, 7263–7270. [Google Scholar] [CrossRef] [PubMed]
- DeRidder, B.P.; Dixon, D.P.; Beussman, D.J.; Edwards, R.; Goldsbrough, P.B. Induction of Glutathione S-Transferases in Arabidopsis by Herbicide Safeners. Plant Physiol. 2002, 130, 1497–1505. [Google Scholar] [CrossRef] [PubMed]
- Nutricati, E.; Miceli, A.; Blando, F.; De Bellis, L. Characterization of two Arabidopsis thaliana glutathione S-transferases. Plant Cell Rep. 2006, 25, 997–1005. [Google Scholar] [CrossRef] [PubMed]
- Grzam, A.; Martin, M.N.; Hell, R.; Meyer, A.J. γ-Glutamyl transpeptidase GGT4 initiates vacuolar degradation of glutathione S -conjugates in Arabidopsis. FEBS Lett. 2007, 581, 3131–3138. [Google Scholar] [CrossRef] [PubMed]
- Merewitz, E. Chemical Priming-Induced Drought Stress Tolerance in Plants. In Drought Stress Tolerance in Plants; Springer International Publishing: Cham, Switzerland, 2016; Volume 1, pp. 77–103. ISBN 978-3-319-28899-4. [Google Scholar]
- Černý, M.; Novák, J.; Habánová, H.; Cerna, H.; Brzobohatý, B. Role of the proteome in phytohormonal signaling. Biochim. Biophys. Acta Proteins Proteom. 2016, 1864, 1003–1015. [Google Scholar] [CrossRef] [PubMed]
- Šimura, J.; Antoniadi, I.; Široká, J.; Tarkowská, D.; Strnad, M.; Ljung, K.; Novák, O. Plant Hormonomics: Multiple Phytohormone Profiling by Targeted Metabolomics. Plant Physiol. 2018, 177, 476–489. [Google Scholar] [CrossRef] [PubMed]
- Černý, M.; Skalák, J.; Cerna, H.; Brzobohatý, B. Advances in purification and separation of posttranslationally modified proteins. J. Proteom. 2013, 92, 2–27. [Google Scholar] [CrossRef] [PubMed]
- Hsu, C.C.; Zhu, Y.; Arrington, J. V; Paez, J.S.; Wang, P.; Zhu, P.; Chen, I.H.; Zhu, J.K.; Tao, W.A. Universal plant phosphoproteomics workflow and its application to tomato signaling in response to cold stress. Mol. Cell. Proteom. 2018, mcp.TIR118.000702. [Google Scholar] [CrossRef] [PubMed]
- Vandereyken, K.; Van Leene, J.; De Coninck, B.; Cammue, B.P.A. Hub Protein Controversy: Taking a Closer Look at Plant Stress Response Hubs. Front. Plant Sci. 2018, 9, 694. [Google Scholar] [CrossRef] [PubMed]
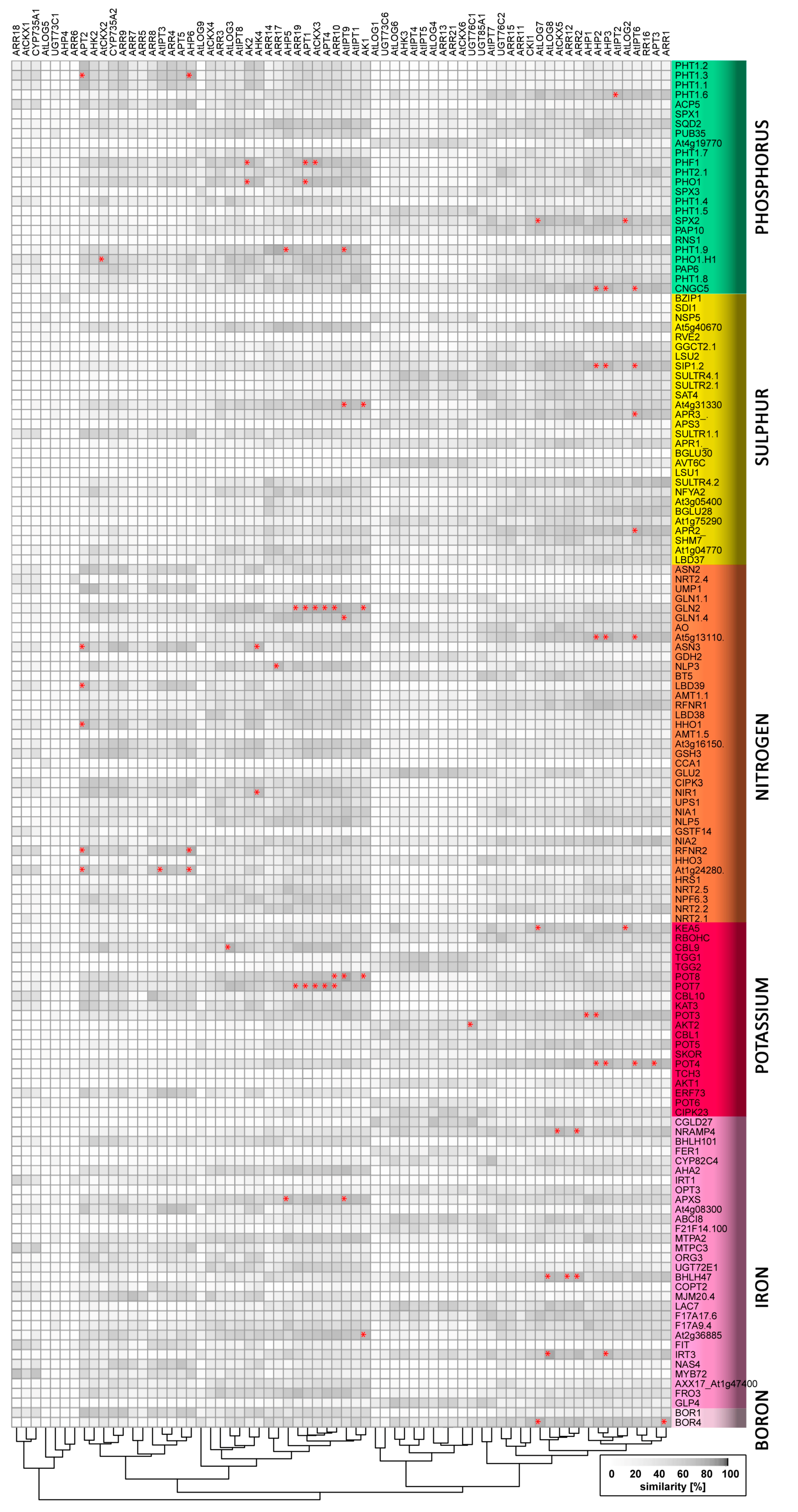

| Gene Name | AGI Code | UniProt Protein Name | Significant Co-Expression with Cytokinin Signalling/Metabolism Genes | References |
|---|---|---|---|---|
| NITROGEN | ||||
| NPF6.3 | AT1G12110 | Protein NRT1/PTR FAMILY 6.3 | [157] | |
| NRT2.1 | AT1G08090 | High-affinity nitrate transporter 2.1 | [158] | |
| NRT2.2 | AT1G08100 | High-affinity nitrate transporter 2.2 | [157] | |
| NRT2.4 | AT5G60770 | High affinity nitrate transporter 2.4 | [159,160] | |
| NRT2.5 | AT1G12940 | High affinity nitrate transporter 2.5 | [13,157,161] | |
| AMT1-1 | AT4G13510 | Ammonium transporter 1 member 1 | [157] | |
| AMT1-5 | AT3G24290 | Putative ammonium transporter 1 member 5 | [161,162] | |
| GDH2 | AT5G07440 | Glutamate dehydrogenase 2 | [157] | |
| GSH3 | AT3G03910 | Probable glutamate dehydrogenase 3 | [157] | |
| GLN2 | AT5G35630 | Glutamine synthetase | 2/4 | [157,158] |
| GLU2 | AT2G41220 | Ferredoxin-dependent glutamate synthase 2 | [163] | |
| NIA1 | AT1G77760 | Nitrate reductase [NADH] 1 | [13,158] | |
| NIA2 | AT1G37130 | Nitrate reductase [NADH] 2 | [157] | |
| NIR1 | AT2G15620 | Ferredoxin–nitrite reductase | 1/0 | [13,158] |
| UMP1 | AT5G40850 | Urophorphyrin methylase 1 | [13,157,158] | |
| GLN1-1 | AT5G37600 | Glutamine synthetase cytosolic isozyme 1-1 | [159,164] | |
| GLN1-4 | AT5G16570 | Glutamine synthetase cytosolic isozyme 1-4 | 0/1 | [157,164] |
| At3g16150 | AT3G16150 | Probable isoaspartyl peptidase/l-asparaginase 2 | [157,159] | |
| ASN2 | AT5G65010 | Asparagine synthetase [glutamine-hydrolyzing] 2 | [22,157] | |
| ASN3 | AT5G10240 | Asparagine synthetase [glutamine-hydrolyzing] 3 | 1/1 | [165] |
| At5g13110 | AT5G13110 | Glucose-6-phosphate 1-dehydrogenase 2 | 2/1 | [157,158] |
| At1g24280 | AT1G24280 | Glucose-6-phosphate 1-dehydrogenase 3 | 1/2 | [22,158] |
| UPS1 | AT2G03590 | Ureide permease 1 | [159] | |
| AT4G39795 | AT4G39795 | Uncharacterized protein | [159] | |
| RFNR1 | AT4G05390 | Ferredoxin–NADP reductase | [14,158] | |
| RFNR2 | AT1G30510 | Ferredoxin–NADP reductase | 1/1 | [158] |
| GSTF14 | AT1G49860 | Glutathione S-transferase F14 | [158] | |
| BT5 | AT4G37610 | BTB/POZ and TAZ domain-containing protein 5 | [158] | |
| CCA1 | AT2G46830 | Protein CCA1 | [157] | |
| TGA1 | AT5G65210 | Transcription factor TGA1 | [166] | |
| TGA4 | AT5G10030 | Transcription factor TGA4 | [166] | |
| NLP3 | AT4G38340 | Protein NLP3 | 1/0 | [167] |
| NLP5 | AT1G76350 | Protein NLP5 | [167] | |
| NLP7 | AT4G24020 | Protein NLP7 | [167] | |
| HHO1 | AT3G25790 | Transcription factor HHO1 | 0/1 | [158,160] |
| HRS1 | AT1G13300 | Transcription factor HRS1 | [158,160] | |
| HHO3 | AT1G25550 | Transcription factor HHO3 | [22,158,160] | |
| LBD37 | AT5G67420 | LOB domain-containing protein 37 | [14,158,168] | |
| LBD38 | AT3G49940 | LOB domain-containing protein 38 | [14,158,168] | |
| LBD39 | AT4G37540 | LOB domain-containing protein 39 | 0/1 | [158,168] |
| CIPK3 | AT2G26980 | CBL-interacting serine/threonine-protein kinase 3 | [158] | |
| CIPK13 | AT2G34180 | CBL-interacting serine/threonine-protein kinase 13 | [157] | |
| AO | AT5G14760 | l-aspartate oxidase | [158] | |
| PHOSPHORUS | ||||
| PHO1 | AT3G23430 | Phosphate transporter PHO1 | 0/2 | [38] |
| PHO1-H1 | AT1G68740 | Phosphate transporter PHO1 homolog 1 | 0/1 | [169] |
| PHF1 | AT3G52190 | SEC12-like protein 1 | 0/3 | [169] |
| PHT1-1 | AT5G43350 | Inorganic phosphate transporter 1-1 | [170,171] | |
| PHT1-2 | AT5G43370 | Probable inorganic phosphate transporter 1-2 | [171,172] | |
| PHT1-3 | AT5G43360 | Probable inorganic phosphate transporter 1-3 | 1/1 | [38,171] |
| PHT1-4 | AT2G38940 | Inorganic phosphate transporter 1-4 | 1/0 | [171,172] |
| PHT1-5 | AT2G32830 | Probable inorganic phosphate transporter 1-5 | 1/0 | [170,171] |
| PHT1-6 | AT5G43340 | Probable inorganic phosphate transporter 1-6 | 0/1 | [172] |
| PHT1-7 | AT3G54700 | Probable inorganic phosphate transporter 1-7 | [173] | |
| PHT1-8 | AT1G20860 | Probable inorganic phosphate transporter 1-8 | [172,173] | |
| PHT1-9 | AT1G76430 | Probable inorganic phosphate transporter 1-9 | 1/1 | [172,173] |
| PHT2-1 | AT3G26570 | Inorganic phosphate transporter 2-1 | [170,172] | |
| SPX1 | AT5G20150 | SPX domain-containing protein 1 | [169,174] | |
| SPX2 | AT2G26660 | SPX domain-containing protein 2 | 0/2 | [174] |
| SPX3 | AT2G45130 | SPX domain-containing protein 3 | [38] | |
| IPS1 | AT3G09922 | INDUCED BY PHOSPHATE STARVATION1 | [169,174] | |
| F12E4_330 | AT5G03545 | At5g03545 | [169] | |
| ACP5 | AT5G27200 | Acyl carrier protein 5 | [38] | |
| RNS1 | AT2G02990 | Ribonuclease 1 | [169,170] | |
| SQD2 | AT5G01220 | Sulfoquinovosyl transferase SQD2 | [169] | |
| PAP10 | AT2G16430 | Purple acid phosphatase 10 | [38] | |
| PAP6 | AT1G56360 | Purple acid phosphatase 6 | [38] | |
| At4g19770 | AT4G19770 | Glycosyl hydrolase family protein with chitinase insertion domain-containing protein | [38] | |
| PUB35 | AT4G25160 | U-box domain-containing protein 35 | [38] | |
| GDPD3 | AT5G43300 | Glycerophosphodiester phosphodiesterase GDPD3 | [38] | |
| ETC3 | AT4G01060 | MYB-like transcription factor ETC3 | [38] | |
| SULPHUR | ||||
| SULTR1;1 | AT4G08620 | Sulfate transporter 1.1 | [175,176] | |
| SULTR2;1 | AT5G10180 | Sulfate transporter 2.1 | [175,176] | |
| SULTR4;1 | AT5G13550 | Sulfate transporter 4.1 | [177,178] | |
| SULTR4;2 | AT3G12520 | Probable sulfate transporter 4.2 | [178,179,180,181] | |
| APS3 | AT4G14680 | ATP-sulfurylase 3 | [176,178] | |
| APR1 | AT4G04610 | 5′-adenylylsulfate reductase 1 | [176,178] | |
| APR2 | AT1G62180 | 5′-adenylylsulfate reductase 2 | 0/1 | [178] |
| APR3 | AT4G21990 | 5′-adenylylsulfate reductase 3 | 0/1 | [178,180] |
| SAT4 | AT4G35640 | Serine acetyltransferase 4 | [181,182] | |
| BGLU28 | AT2G44460 | Beta-glucosidase 28 | [179,180] | |
| BGLU30 | AT3G60140 | Beta-glucosidase 30 | [180,182] | |
| SDI1 | AT5G48850 | Protein SULFUR DEFICIENCY-INDUCED 1 | [178,179] | |
| SDI2 | AT1G04770 | Protein SULFUR DEFICIENCY-INDUCED 2 | [176,178] | |
| SHM7 | AT1G36370 | Serine hydroxymethyltransferase 7 | [178,180] | |
| GGCT2;1 | AT5G26220 | Gamma-glutamylcyclotransferase 2-1 | [178,180,182] | |
| LSU1 | AT3G49580 | Protein RESPONSE TO LOW SULFUR 1 | [178,180] | |
| LSU2 | AT5G24660 | Protein RESPONSE TO LOW SULFUR 2 | [178,180,182] | |
| At3g05400 | AT3G05400 | Sugar transporter ERD6-like 12 | [180,182] | |
| At4g31330 | AT4G31330 | Protein of unknown function | 0/2 | [180,182] |
| SIP1-2 | AT5G18290 | Probable aquaporin SIP1-2 | 2/1 | [180] |
| At5g40670 | AT5G40670 | Cystinosin homolog | [180] | |
| At1g75290 | AT1G75290 | NAD | [180] | |
| NSP5 | AT5G48180 | Nitrile-specifier protein 5 | [180] | |
| AVT6C | AT3G56200 | Amino acid transporter AVT6C | [178,180] | |
| NFYA2 | AT3G05690 | Nuclear transcription factor Y subunit A-2 | [180] | |
| BZIP1 | AT5G49450 | Basic leucine zipper 1 | [180] | |
| RVE2 | AT5G37260 | Homeodomain-like superfamily protein | [180] | |
| POTASSIUM | ||||
| POT5 | AT4G13420 | Potassium transporter 5 | [182,183] | |
| POT4 | AT3G02050 | Potassium transporter 4 | 2/2 | [183] |
| AKT1 | AT2G26650 | Potassium channel AKT1 | [184,185] | |
| RBOHC | AT5G51060 | Respiratory burst oxidase homolog protein C | [183] | |
| CIPK23 | AT1G30270 | CBL-interacting serine/threonine-protein kinase 23 | [183,186] | |
| TGG1 | AT5G26000 | Myrosinase 1 | [182] | |
| TGG2 | AT5G25980 | Myrosinase 2 | [182] | |
| POT6 | AT1G70300 | Potassium transporter 6 | [187] | |
| POT8 | AT5G14880 | Potassium transporter 8 | 1/2 | [184,187] |
| KEA5 | AT5G51710 | K+ efflux antiporter 5 | 0/2 | [183] |
| KAT3 | AT4G32650 | Potassium channel KAT3 | [184] | |
| SKOR | AT3G02850 | Potassium channel SKOR | [184,188,189] | |
| AKT2 | AT4G22200 | Potassium channel AKT2/3 | 0/1 | [189] |
| POT3 | AT4G23640 | Potassium transporter 3 | 2/0 | [190] |
| POT7 | AT5G09400 | Potassium transporter 7 | 2/3 | [191] |
| CBL1 | AT4G17615 | Calcineurin B-like protein 1 | [189] | |
| CBL9 | AT5G47100 | Calcineurin B-like protein 9 | 0/1 | [189] |
| CBL10 | AT4G33000 | Calcineurin B-like protein 10 | [183] | |
| TCH3 | AT2G41100 | Calcium-binding EF hand family protein | [192] | |
| ERF73 | AT1G72360 | Integrase-type DNA-binding superfamily protein | [183] | |
| IRON | ||||
| AXX17_At1g47400 | AT1G47400 | Uncharacterized protein | [182,193] | |
| At1g47395 | AT1G47395 | At1g47390 | [182] | |
| AT2G14247 | AT2G14247 | Expressed protein | [182] | |
| At1g13609 | AT1G13609 | Defensin-like | [182] | |
| IRT1 | AT4G19690 | Fe2+ transport protein 1 | [182,193,194] | |
| F17A17.6 | AT3G07720 | AT3g07720/F17A17_6 | [182,193,194] | |
| MTPA2 | AT3G58810 | Metal tolerance protein A2 | [193,194] | |
| MJM20.4 | AT3G12900 | 2-oxoglutarate | [193,194] | |
| F21F14.100 | AT3G61930 | Uncharacterized protein At3g61930/F21F14_100 | [193,194] | |
| COPT2 | AT3G46900 | Copper transporter 2 | [193,194] | |
| CYP82C4 | AT4G31940 | Cytochrome P450 82C4 | [193] | |
| GLP4 | AT1G09560 | Germin-like protein subfamily 2 member 1 | [193,194] | |
| F17A9.4 | AT3G06890 | At3g06890 | [193,194] | |
| UGT72E1 | AT3G50740 | UDP-glycosyltransferase 72E1 | [193,194] | |
| ORG3 | AT3G56980 | Transcription factor ORG3 | [182,193,194] | |
| MYB72 | AT1G56160 | Transcription factor MYB72 | [193,194] | |
| MTPC3 | AT3G58060 | Putative metal tolerance protein C3 | [193,194] | |
| FIT | AT2G28160 | Transcription factor FER-LIKE IRON DEFICIENCY-INDUCED TRANSCRIPTION FACTOR | [194,195] | |
| BHLH47 | AT3G47640 | Transcription factor bHLH47 | 2/1 | [193,195] |
| BHLH101 | AT5G04150 | Transcription factor bHLH101 | [182,193] | |
| NAS4 | AT1G56430 | Probable nicotianamine synthase 4 | [182,193] | |
| OPT3 | AT4G16370 | Oligopeptide transporter 3 | [195,196] | |
| CGLD27 | AT5G67370 | Protein CONSERVED IN THE GREEN LINEAGE AND DIATOMS 27 | [182] | |
| FRO2 | AT1G01580 | Ferric reduction oxidase 2 | [182] | |
| FRO3 | AT1G23020 | Ferric reduction oxidase 3 | [193,195] | |
| AHA2 | AT4G30190 | Plasma membrane ATPase | [195] | |
| NRAMP4 | AT5G67330 | Metal transporter Nramp4 | 1/1 | [193,195] |
| FER1 | AT5G01600 | Ferritin-1 | [193,194] | |
| ABCI8 | AT4G04770 | UPF0051 protein ABCI8 | [193,194] | |
| At2g36885 | AT2G36885 | Translation initiation factor | 0/1 | [193,194,195] |
| APXS | AT4G08390 | Stromal ascorbate peroxidase | 1/1 | [193,194] |
| LAC7 | AT3G09220 | Laccase-7 | [193,194,195] | |
| IRT3 | AT1G60960 | Fe2+ transport protein 3, chloroplastic | 1/1 | [193] |
| At4g08300 | AT4G08300 | WAT1-related protein At4g08300 | [182] | |
| FRO4 | AT5G23980 | Ferric reduction oxidase 4 | [185] | |
| BORON | ||||
| BOR1 | AT2G47160 | Boron transporter 1 | [197] | |
| BOR4 | AT1G15460 | Boron transporter 4 | 1/1 | [198] |
| TEMPERATURE/DROUGHT | ||||
| RD29A | AT5G52310 | Low-temperature-induced 78 kDa protein | [141,199,200] | |
| KIN1 | AT5G15960 | Stress-induced protein KIN1 | [201,202] | |
| KIN2 | AT5G15970 | Stress-induced protein KIN2 | [203,204] | |
| COR15A | AT2G42540 | Protein COLD-REGULATED 15A | [200,201,202] | |
| COR47 | AT1G20440 | Dehydrin COR47 | [201,202] | |
| ERD10 | AT1G20450 | Dehydrin ERD10 | [201,202] | |
| ERD7 | AT2G17840 | Protein EARLY-RESPONSIVE TO DEHYDRATION 7 | [201,202,205] | |
| At1g30790 | AT1G30790 | F-box protein At1g30790 | 0/2 | [205] |
| MKK2 | AT4G29810 | Mitogen-activated protein kinase kinase 2 | [206] | |
| RAB18 | AT5G66400 | Dehydrin Rab18 | [207,208,209] | |
| LTI65/RD29B | AT5G52300 | Low-temperature-induced 65 kDa protein | [129,199,200] | |
| RD22 | AT5G25610 | BURP domain protein RD22 | [207,209] | |
| HOS1 | AT2G39810 | E3 ubiquitin-protein ligase HOS1 | 4/3 | [203,210] |
| DREB1B | AT4G25490 | Dehydration-responsive element-binding protein 1B | [201,202] | |
| DREB1C | AT4G25470 | Dehydration-responsive element-binding protein 1C | [201,202,209] | |
| DREB1A | AT4G25480 | Dehydration-responsive element-binding protein 1A | [201,202] | |
| RABC1 | AT1G43890 | Ras-related protein RABC1 | 0/2 | [202] |
| CLPD | AT5G51070 | Chaperone protein ClpD | [209,211] | |
| SWEET15 | AT5G13170 | Bidirectional sugar transporter SWEET15 | [40,209] | |
| P5CSA | AT2G39800 | Delta-1-pyrroline-5-carboxylate synthase A | [40,209] | |
| ABI1 | AT4G26080 | Protein phosphatase 2C 56 | [209] | |
| DREB2A | AT5G05410 | Dehydration-responsive element-binding protein 2A | [209,212] | |
| NCED3 | AT3G14440 | 9-cis-epoxycarotenoid dioxygenase NCED3 | [200,209] | |
| ABF3 | AT4G34000 | ABSCISIC ACID-INSENSITIVE 5-like protein 6 | [209,213] | |
| PP2CA | AT3G11410 | Protein phosphatase 2C 37 | 2/2 | [200,209,214] |
| PXG3/RD20 | AT2G33380 | Probable peroxygenase 3 | [200,209,211] | |
| LEA7 | AT1G52690 | Late embryogenesis abundant protein 7 | [209,215] | |
| LEA29 | AT3G15670 | Late embryogenesis abundant protein 29 | [40] | |
| At3g17520 | AT3G17520 | Late embryogenesis abundant protein | [208,209] | |
| NAC072 | AT4G27410 | NAC domain-containing protein 72 | [208,209] | |
| MBF1C | AT3G24500 | Multiprotein-bridging factor 1c | [216,217] | |
| HSFA2 | AT2G26150 | Heat stress transcription factor A-2 | [217,218,219] | |
| HSA32 | AT4G21320 | Protein HEAT-STRESS-ASSOCIATED 32 | [216,217,218] | |
| CLPB1 | AT1G74310 | Chaperone protein ClpB1 | [216,217,218] | |
| CLPB3 | AT5G15450 | Chaperone protein ClpB3 | [216,218] | |
| HSFB2A | AT5G62020 | Heat stress transcription factor B-2a | [216,219] | |
| HSFA7A | AT3G51910 | Heat stress transcription factor A-7a | [217,218,219] | |
| HSP90-1 | AT5G52640 | Heat shock protein 90-1 | [216,217,218] | |
| HSP90-2 | AT5G56030 | Heat shock protein 90-2 | [216,217] | |
| At2g20560 | AT2G20560 | At2g20560/T13C7.15 | [216,217,218] | |
| HSFB1 | AT4G36990 | Heat stress transcription factor B-1 | [216,217,218] | |
| HSP23.6 | AT4G25200 | 23.6 kDa heat shock protein | [217,218] | |
| HSP18.1 | AT5G59720 | 18.1 kDa class I heat shock protein | [217,218] | |
| HSP17.4B | AT1G54050 | 17.4 kDa class III heat shock protein | [216,217] | |
| MED37C | AT3G12580 | Probable mediator of RNA polymerase II transcription subunit 37c | [217,220] | |
| HSP70-5 | AT1G16030 | Heat shock 70 kDa protein 5 | [217,218,220] | |
| HSP70-10 | AT5G09590 | Heat shock 70 kDa protein 10 | [216,217] | |
| GOLS1 | AT2G47180 | Galactinol synthase 1 | [216,217,218] | |
| APX2 | AT3G09640 | L-ascorbate peroxidase 2 | [217,218] | |
| ERDJ3A | AT3G08970 | DnaJ protein ERDJ3A | [216,217] | |
| HSP90-6 | AT3G07770 | Heat shock protein 90-6 | 2/0 | [216,217] |
| HSP90-4 | AT5G56000 | Heat shock protein 90-4 | [216,217] | |
| HSP70-8 | AT2G32120 | Heat shock 70 kDa protein 8 | [216,217] | |
| MED37D | AT5G02490 | Probable mediator of RNA polymerase II transcription subunit 37c | [216,217] | |
| HSP70-3 | AT3G09440 | Heat shock 70 kDa protein 3 | [216,217] | |
| HSP70-15 | AT1G79920 | Heat shock 70 kDa protein 15 | 0/2 | [216,217] |
| HSP90-5 | AT2G04030 | Heat shock protein 90-5 | 0/1 | [216,217] |
| XENOBIOTIC STRESS | ||||
| GSH1 | AT4G23100 | Glutamate–cysteine ligase | [221,222] | |
| GSH2 | AT5G27380 | Glutathione synthetase | [221,222] | |
| PCS1 | AT5G44070 | Glutathione gamma-glutamylcysteinyltransferase 1 | [221,222] | |
| MAN3 | AT3G10890 | Mannan endo-1 | [223] | |
| ZAT6 | AT5G04340 | Zinc finger protein ZAT6 | [223] | |
| PCR1 | AT1G14880 | Protein PLANT CADMIUM RESISTANCE 1 | [224] | |
| HMA3 | AT4G30120 | Putative inactive cadmium/zinc-transporting ATPase HMA3 | [225,226] | |
| HMA4 | AT2G19110 | Putative cadmium/zinc-transporting ATPase HMA4 | 2/1 | [227] |
| HSFA4A | AT4G18880 | Heat stress transcription factor A-4a | [228] | |
| FC1 | AT5G26030 | Ferrochelatase-1 | [222] | |
| HMT-1 | AT3G25900 | Homocysteine S-methyltransferase 1 | 2/2 | [229] |
| MT1A | AT1G07600 | Metallothionein-like protein 1A | 1/4 | [230] |
| NRAMP5 | AT4G18790 | Metal transporter Nramp5 | 2/2 | [226,231] |
| ABCG36 | AT1G59870 | ABC transporter G family member 36 | [232] | |
| ABCB25 | AT5G58270 | ABC transporter B family member 25 | 4/3 | [233] |
| ABCC1 | AT1G30400 | ABC transporter C family member 1 | 4/2 | [226,234,235] |
| ABCC2 | AT2G34660 | ABC transporter C family member 2 | 10/7 | [226,234,235] |
| HAC1 | AT2G21045 | Protein HIGH ARSENIC CONTENT 1 | 1/1 | [236] |
| ALMT1 | AT1G08430 | Aluminum-activated malate transporter 1 | [237,238] | |
| ALS3 | AT2G37330 | Protein ALUMINUM SENSITIVE 3 | 2/4 | [237,239] |
| STOP1 | AT1G34370 | Protein SENSITIVE TO PROTON RHIZOTOXICITY 1 | 0/1 | [237,239] |
| CYP81D11 | AT3G28740 | Cytochrome P450 81D11 | [240,241,242,243] | |
| CYP710A1 | AT2G34500 | Cytochrome P450 710A1 | [82,244,245] | |
| CYP81D8 | AT4G37370 | Cytochrome P450 | [81,243,244,245] | |
| UGT73B2 | AT4G34135 | UDP-glucosyl transferase 73B2 | [241,242,244] | |
| UGT73B3 | AT4G34131 | UDP-glycosyltransferase 73B3 | [242,244] | |
| UGT73B4 | AT2G15490 | UDP-glycosyltransferase 73B4 | [241,242,244] | |
| UGT73C1 | AT2G36750 | UDP-glycosyltransferase 73C1 | [241] | |
| GSTU3 | AT2G29470 | Glutathione S-transferase U3 | [241,244] | |
| GSTU10 | AT1G74590 | Glutathione S-transferase U10 | [82,243,246] | |
| GSTU19 | AT1G78380 | Glutathione S-transferase U19 | 2/3 | [243,247] |
| GSTU24 | AT1G17170 | Glutathione S-transferase U24 | [240,241,242,243] | |
| GSTU25 | AT1G17180 | Glutathione S-transferase U25 | [241,242,243,244] | |
| GSTU26 | AT1G17190 | Glutathione S-transferase U26 | [248] | |
| GGT4 | AT4G29210 | Glutathione hydrolase 3 | 1/0 | [249] |
| ABCC3 | AT3G13080 | ABC transporter C family member 3 | [244] | |
| ABCI21 | AT5G44110 | ABC transporter I family member 21 | [242,243] | |
| DTX1 | AT2G04040 | Protein DETOXIFICATION 1 | [243,245] | |
| DTX3 | AT2G04050 | Protein DETOXIFICATION 3 | [243,245] | |
| DTX4 | AT2G04070 | Protein DETOXIFICATION 4 | [241,245] | |
| CYP710A2 | AT2G34490 | Cytochrome P450 710A2 | [82] | |
| DHAR2 | AT1G75270 | Glutathione S-transferase DHAR2 | 1/0 | [243,244] |
| DHAR3 | AT5G16710 | Glutathione S-transferase DHAR3 | 2/0 | [243] |
| GSTU4 | AT2G29460 | Glutathione S-transferase U4 | [241,244] | |
| UGT74F2 | AT2G43820 | UDP-glycosyltransferase 74F2 | [244,246] | |
| UGT73C6 | AT2G36790 | UDP-glycosyltransferase 73C6 | 0/1 | [241,245] |
| UGT74E2 | AT1G05680 | UDP-glycosyltransferase 74E2 | [241,245] | |
| UGT73B5 | AT2G15480 | UDP-glycosyltransferase 73B5 | [241,244] | |
| UGT75B1 | AT1G05560 | UDP-glycosyltransferase 75B1 | [241,244] | |
| CYP81F2 | AT5G57220 | Cytochrome P450 81F2 | [241] | |
| CYP87A2 | AT1G12740 | Photosynthetic NDH subunit of lumenal location 5 | 1/1 | [240] |
| GSTF7 | AT1G02920 | Glutathione S-transferase F7 | [243] | |
| GSTF6 | AT1G02930 | Glutathione S-transferase F6 | [243,246] | |
| ABCB15 | AT3G28345 | ABC transporter B family member 15 | [242] | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pavlů, J.; Novák, J.; Koukalová, V.; Luklová, M.; Brzobohatý, B.; Černý, M. Cytokinin at the Crossroads of Abiotic Stress Signalling Pathways. Int. J. Mol. Sci. 2018, 19, 2450. https://doi.org/10.3390/ijms19082450
Pavlů J, Novák J, Koukalová V, Luklová M, Brzobohatý B, Černý M. Cytokinin at the Crossroads of Abiotic Stress Signalling Pathways. International Journal of Molecular Sciences. 2018; 19(8):2450. https://doi.org/10.3390/ijms19082450
Chicago/Turabian StylePavlů, Jaroslav, Jan Novák, Vladěna Koukalová, Markéta Luklová, Břetislav Brzobohatý, and Martin Černý. 2018. "Cytokinin at the Crossroads of Abiotic Stress Signalling Pathways" International Journal of Molecular Sciences 19, no. 8: 2450. https://doi.org/10.3390/ijms19082450
APA StylePavlů, J., Novák, J., Koukalová, V., Luklová, M., Brzobohatý, B., & Černý, M. (2018). Cytokinin at the Crossroads of Abiotic Stress Signalling Pathways. International Journal of Molecular Sciences, 19(8), 2450. https://doi.org/10.3390/ijms19082450







