Evaluation of the Use of TRIzol-Based Protein Extraction Approach for Gel-Based Proteomic Analysis of Dried Seafood Products and Chinese Tonic Foods
Abstract
1. Introduction
2. Results and Discussion
2.1. TRIzol Protein Extractions
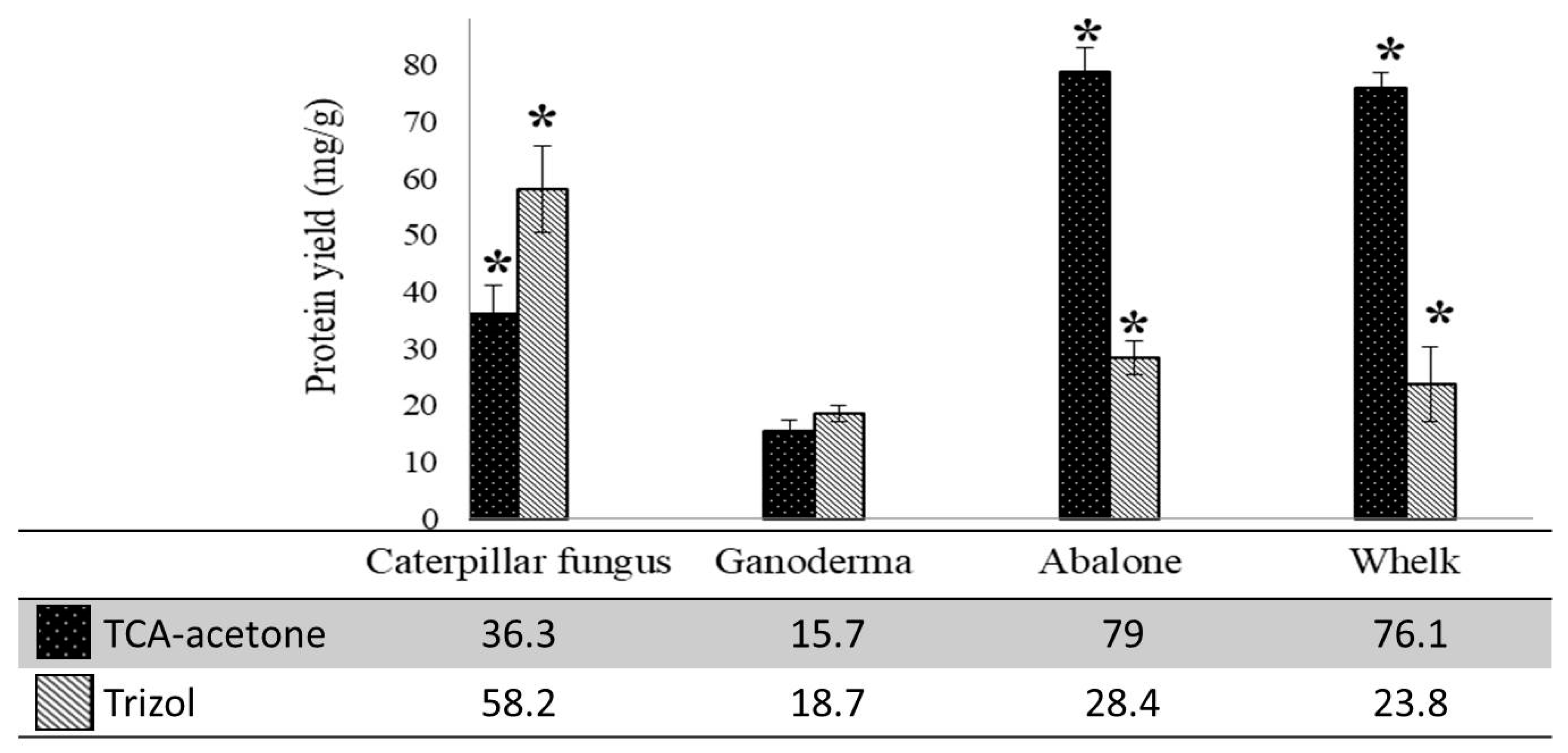
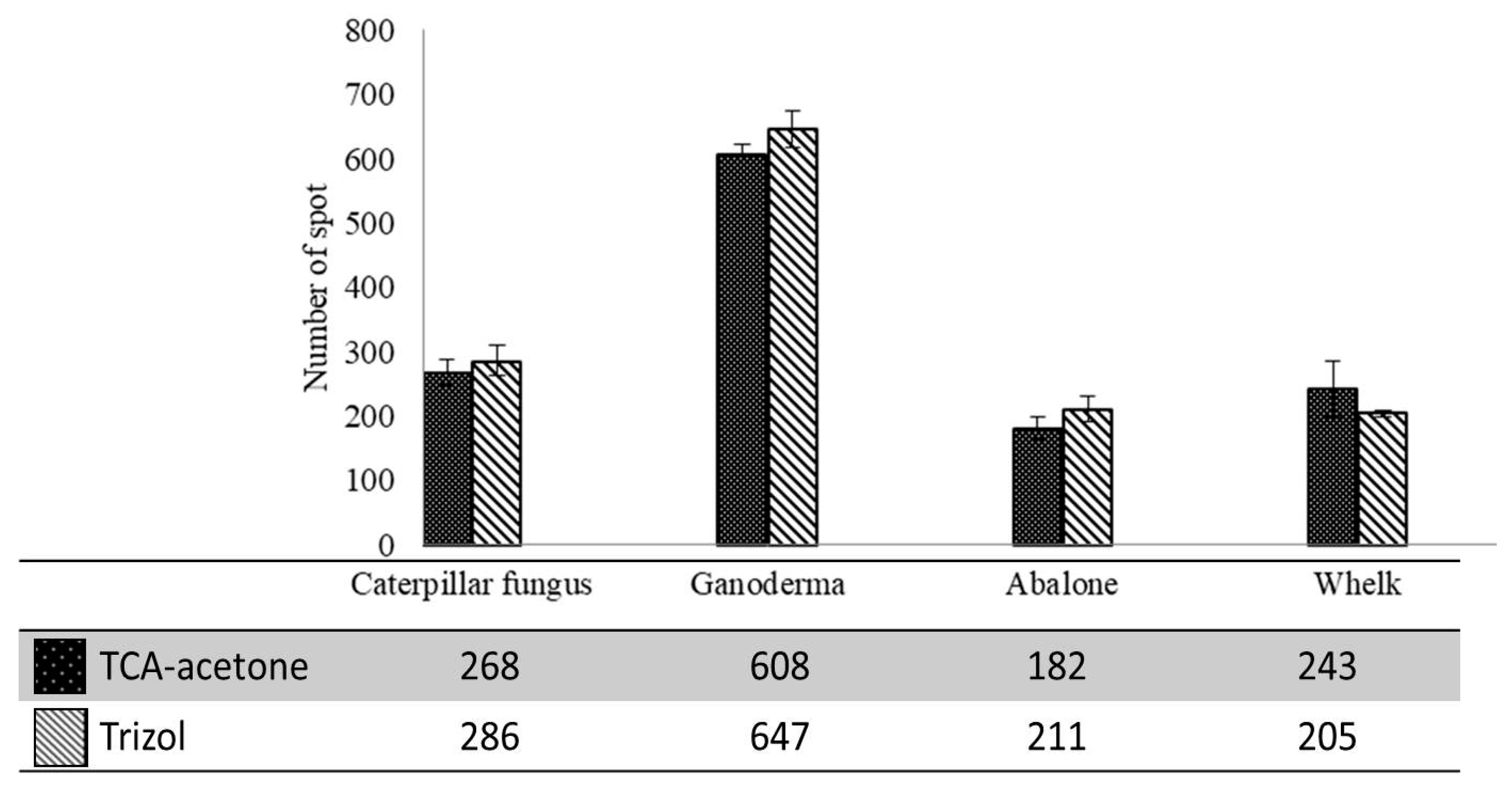
2.2. Protein Concentrations and Number of Spots
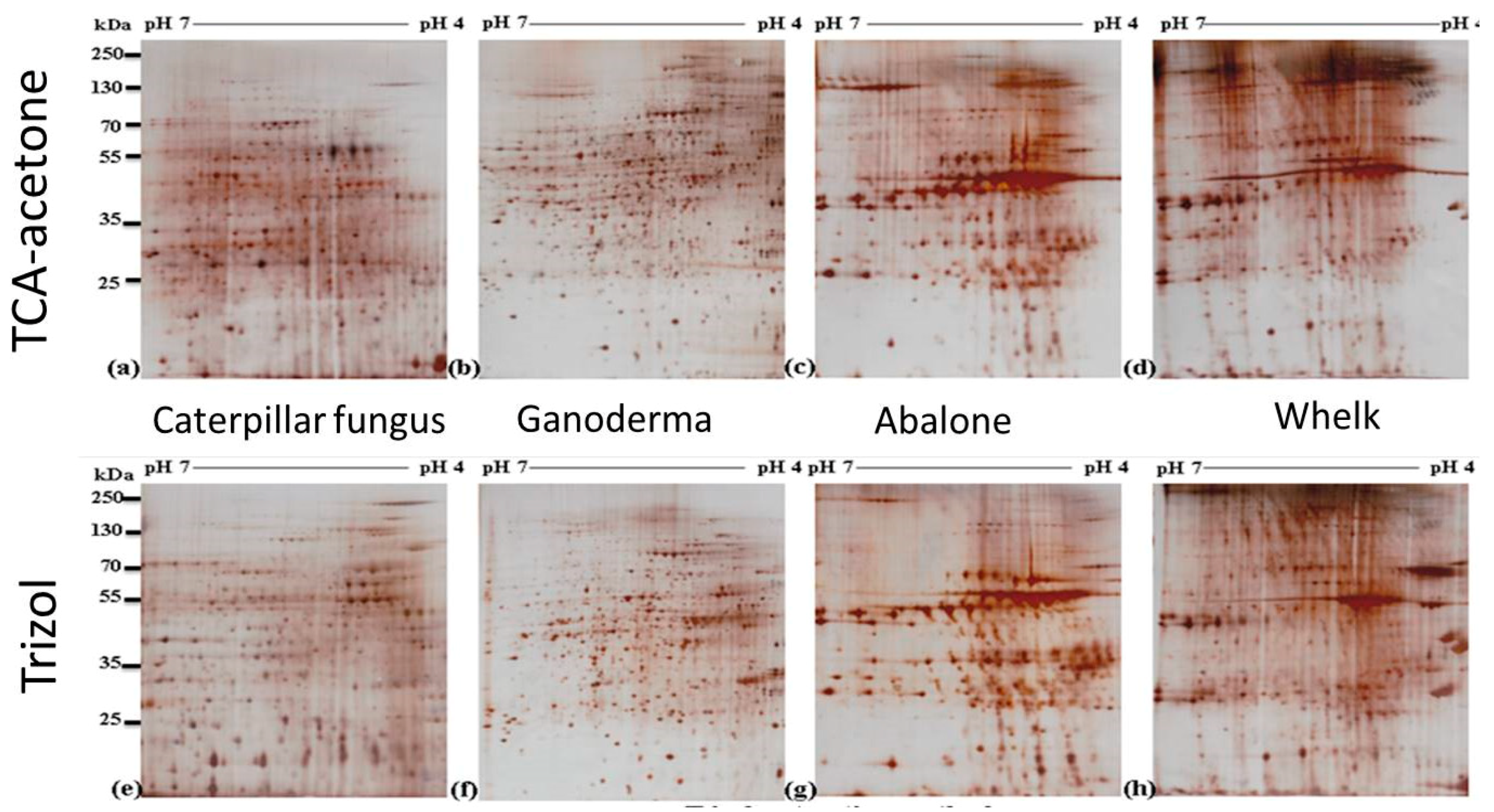
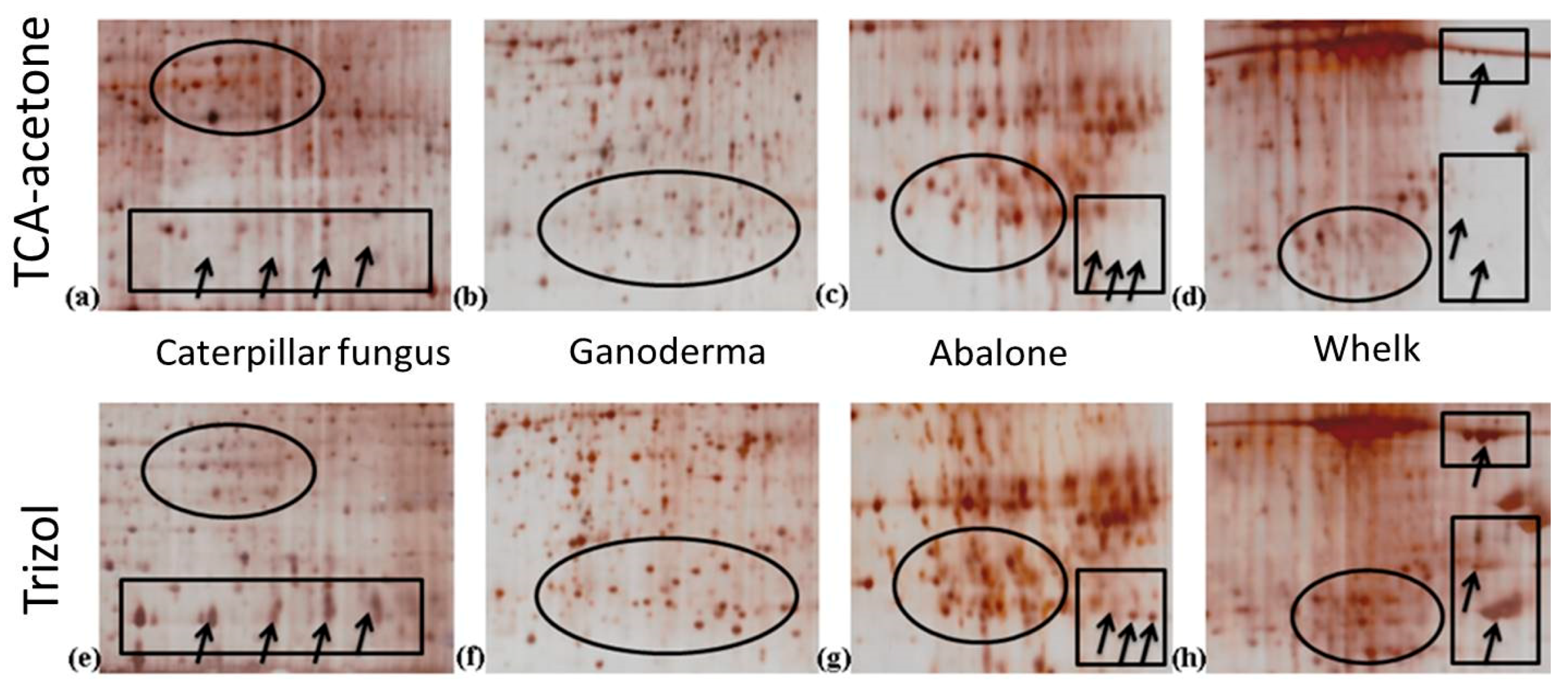
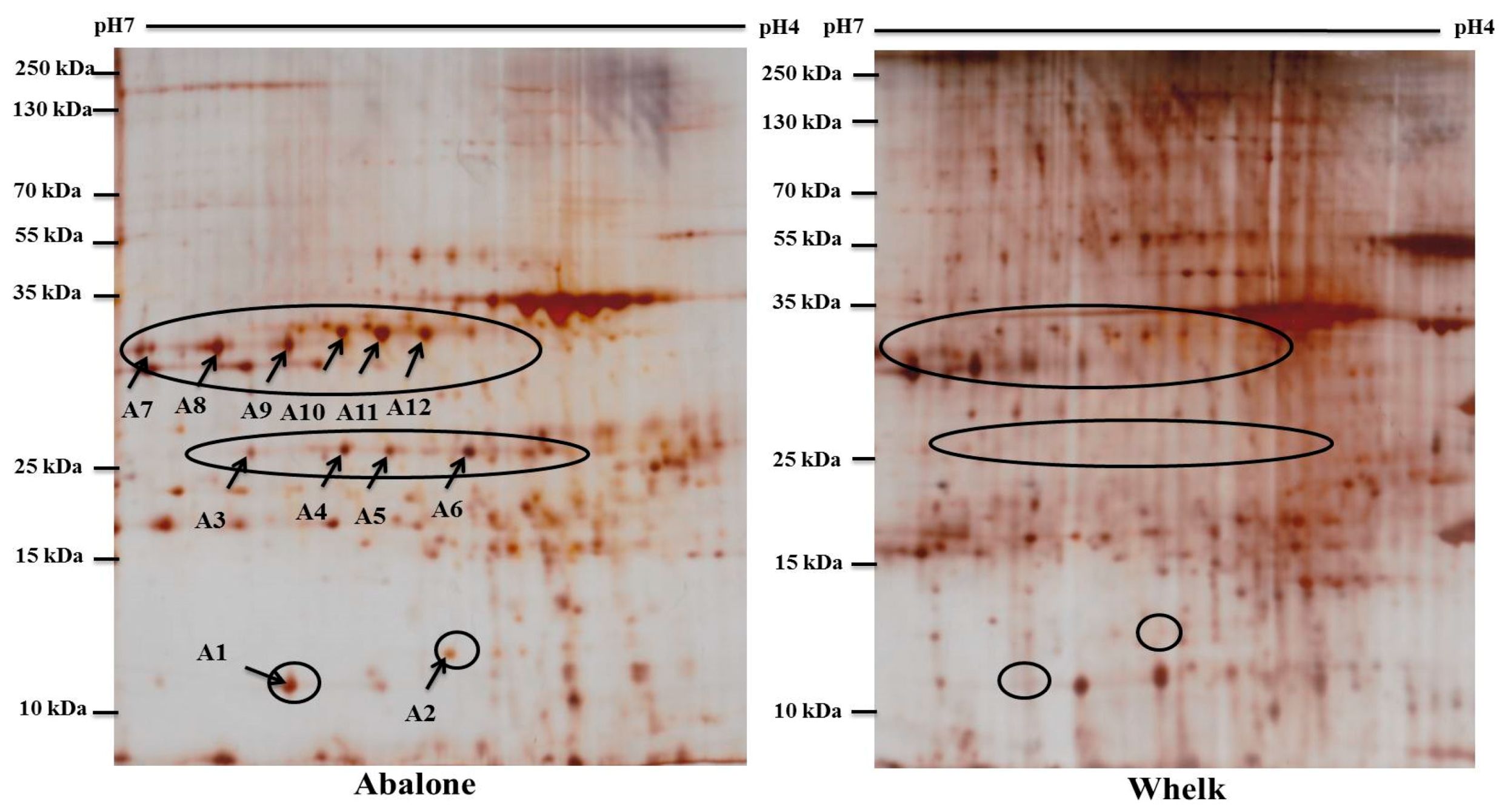
2.3. Two-Dimensional Gel Electrophoresis (2-DE) Images
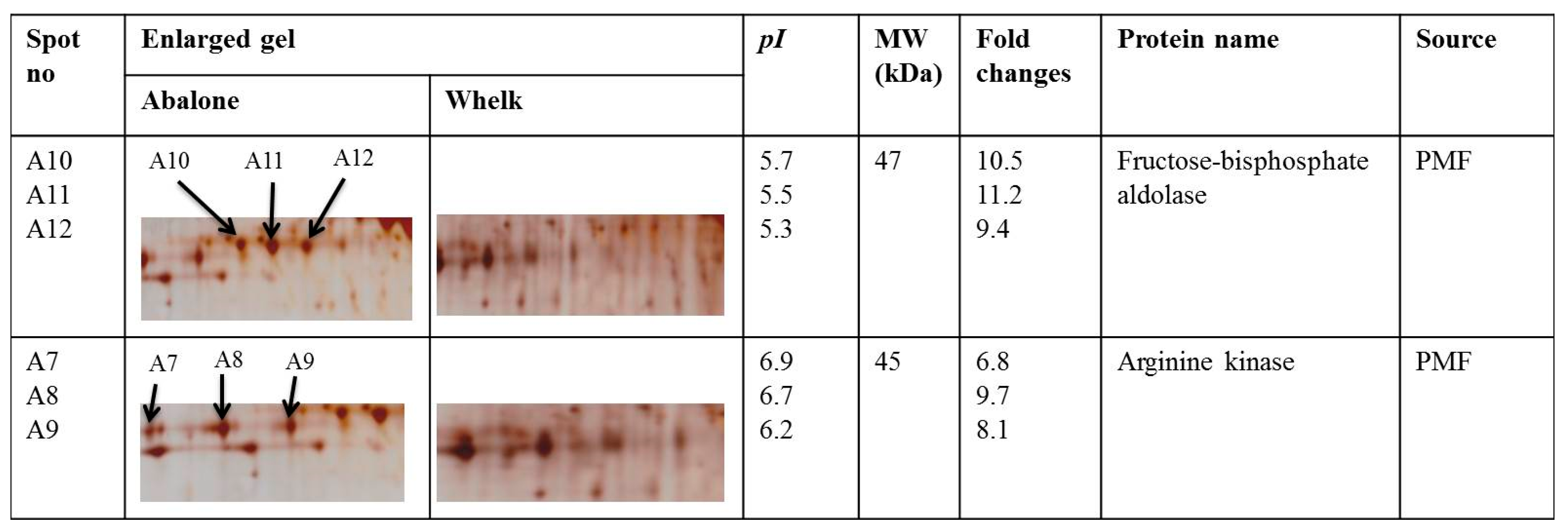
2.4. Identification of Differentially Expressed Proteins Between Abalone Slices and Whelk Slices through Matrix-Assisted Laser Desorption/Ionisation Time-of-Flight Mass Spectrometry (MALDI-TOF/TOF MS)
3. Materials and Methods
3.1. Dried Seafood Products and Chinese Tonic Foods
3.2. Trichloroacetic Acid (TCA)-Acetone Protein Precipitation Method
3.3. TRIzol Protein Extraction Method
3.4. Protein Quantification
3.5. Two-Dimensional Polyacrylamide Gel Electrophoresis
3.6. In-Gel Protein Digestion and MALDI-TOF/TOF Analysis
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| 2-DE | Two Dimensional Gel Electrophoresis |
| AC | Acetonitrile |
| CHAPS | 3-[(3-cholamidopropyl)dimethylammonio]-1-propanesulfonate |
| DDT | Dithiothreitol |
| HCCA | α-cyano-4-hydroxycinnamic Acid |
| IAA | Iodoacetamine |
| IEF | Isoelectric Focusing |
| MALDI-TOF | Matrix-Assisted Laser Desorption/Ionization Mass Spectrometry |
| PMF | Peptide Mass Fingerprint |
| SDS-PAGE | Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis |
| TCA | Trichloroacetic Acid |
| TFA | Trifluoroacetic Acid |
References
- Zhang, S.; Lai, X.; Li, B.; Wu, C.; Wang, S.; Chen, X.; Huang, J.; Yang, G. Application of Differential Proteomic Analysis to Authenticate Ophiocordyceps sinensis. Curr. Microbiol. 2016, 72, 337–343. [Google Scholar] [CrossRef] [PubMed]
- Customs and Excise Department. Customs Seizes Fake Abalone Slices; Customs and Excise Department: Hong Kong, China, 2010.
- Anderson, N.; Anderson, G.N. Proteome and proteomics: New technologies, new concepts, and new words. Electrophoresis 1998, 19, 1853–1861. [Google Scholar] [CrossRef] [PubMed]
- Lima, T.B.; Pinto, M.F.S.; Ribeiro, S.M.; de Lima, L.A.; Viana, J.C.; Gomes Júnior, N.; de Souza Cândido, E.; Dias, S.C.; Franco, O.L. Bacterial resistance mechanism: What proteomics can elucidate. FASEB J. 2013, 27, 1291–1303. [Google Scholar] [CrossRef] [PubMed]
- Sheehan, D. Environmental proteomics and biomarker discovery. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 2010, 157, S42. [Google Scholar] [CrossRef]
- Barry, R.; Soloviev, M. Quantitative protein profiling using antibody arrays. Proteomics 2004, 4, 3717–3726. [Google Scholar] [CrossRef] [PubMed]
- Roe, M.R. Gel-free mass spectrometry-based high throughput proteomics: Tools for studying biological response of proteins and proteomes. Proteomics 2006, 6, 4678–4687. [Google Scholar] [CrossRef] [PubMed]
- Rabilloud, T. Two-dimensional gel electrophoresis in proteomics: Past, present and future. J. Proteom. 2010, 73, 2064–2077. [Google Scholar] [CrossRef] [PubMed]
- Gallardo, J.M.; Ortea, I.; Carrera, M. Proteomics and its applications for food authentication and food-technology research. Trends Anal. Chem. 2013, 52, 135–141. [Google Scholar] [CrossRef]
- Ortea, I.; O’Connor, G.; Maquet, A. Review on proteomics for food authentication. J. Proteom. 2016, 147, 212–225. [Google Scholar] [CrossRef] [PubMed]
- Li, C.-H.; Zuo, H.-L.; Chen, C.; Hu, Y.-J.; Qian, Z.-M.; Li, W.-J.; Xia, Z.-N.; Yang, F.-Q. SDS-PAGE and 2-DE protein profiles of Ganoderma lucidum from different origins. Pak. J. Pharm. Sci. 2018, 31, 447–454. [Google Scholar] [PubMed]
- Tong, X.; Wang, Y.; Xue, Z.; Chen, L.; Qiu, Y.; Cao, J.; Peng, C.; Guo, J. Proteomic identification of marker proteins ans its application to authenticate Ophiocordyceps sinensis. 3 Biotech 2018, 8, 246. [Google Scholar] [CrossRef] [PubMed]
- Bodzon-Kulakowska, A.; Bierczynska-Krzysik, A.; Dylag, T.; Drabik, A.; Suder, P.; Noga, M.; Jarzebinska, J.; Silberring, J. Methods for samples preparation in proteomic research. J. Chromatogr. B 2007, 849, 1–31. [Google Scholar] [CrossRef] [PubMed]
- Ruan, Y.; Wan, M. An optimized procedure for solubilization, reduction, and transfer of human breast cancer membrane-enriched fraction by 2-DE. Electrophoresis 2007, 28, 3333–3340. [Google Scholar] [CrossRef] [PubMed]
- Westermeier, R. Looking at proteins from two dimensions: A review on five decades of 2D electrophoresis. Arch. Physiol. Biochem. 2014, 120, 168–172. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Zhou, J.; Zhang, X.; Zheng, X.; Jiang, X.; Shi, L.; Yin, W.; Wang, J. Optimized sample preparation for two-dimensional gel electrophoresis of soluble proteins from chicken bursa of Fabricius. Proteome Sci. 2009, 7, 38. [Google Scholar] [CrossRef] [PubMed]
- Hao, R.; Adoligbe, C.; Jiang, B.; Zhao, X.; Gui, L.; Qu, K.; Wu, S.; Zan, L.; Mazzuca, S. An Optimized Trichloroacetic Acid/Acetone Precipitation Method for Two-Dimensional Gel Electrophoresis Analysis of Qinchuan Cattle Longissimus Dorsi Muscle Containing High Proportion of Marbling. PLoS ONE 2015, 10, e0124723. [Google Scholar] [CrossRef] [PubMed]
- Tal, I.; Cynthia, M.B.D.; Ramu, S.S.; Yonghua, H.; Carmen, C.; Montserrat, S.; Jocelyn, K.C.R. Sample extraction techniques for enhanced proteomic analysis of plant tissues. Nat. Protoc. 2006, 1, 769. [Google Scholar]
- Kirkland, P.A.; Busby, J.; Stevens, S.; Maupin-Furlow, J.A. Trizol-based method for sample preparation and isoelectric focusing of halophilic proteins. Anal. Biochem. 2006, 351, 254–259. [Google Scholar] [CrossRef] [PubMed]
- Butt, R.; Pfeifer, T.; Delaney, A.; Grigliatti, T.; Tetzlaff, W.; Coorssen, J. Enabling Coupled Quantitative Genomics and Proteomics Analyses from Rat Spinal Cord Samples. Mol. Cell. Proteom. 2007, 6, 1574–1588. [Google Scholar] [CrossRef] [PubMed]
- Lasserre, J.P. Effects of the endocrine disruptors atrazine and PCB 153 on the protein expression of MCF-7 human cells. J. Proteome Res. 2009, 8, 5485–5496. [Google Scholar] [CrossRef] [PubMed]
- Rajagopalan, S.; Nicole, K.; Peicheng, D.; Edward, N.; Thomas, J.B.; Abdissa, N.; Ruth Hogue, A.; Michael, B.P. Global proteomic analysis distinguishes biologic differences in head and neck squamous carcinoma. Lab. Investig. 2007, 87, 755. [Google Scholar]
- Young, C. Proteins isolated with TRIzol are compatible with two-dimensional electrophoresis and mass spectrometry analyses. Anal. Biochem. 2012, 421, 330–332. [Google Scholar] [CrossRef] [PubMed]
- Lee, F.W.-F.; Lo, S.C.-L. The use of Trizol reagent (phenol/guanidine isothiocyanate) for producing high quality two-dimensional gel electrophoretograms (2-DE) of dinoflagellates. J. Microbiol. Methods 2008, 73, 26–32. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Wang, D.; Dong, H.; Xie, Z.; Hong, H. Proteomics of a toxic dinoflagellate Alexandrium catenella DHOI: Detection and identification of cell surface proteins using fluorescent labeling. Chin. Sci. Bull. 2012, 57, 3320–3327. [Google Scholar] [CrossRef]
- Liu, B.; Lo, S.C.-L.; Matton, D.P.; Lang, B.F.; Morse, D. Daily Changes in the Phosphoproteome of the Dinoflagellate Lingulodinium. Protist 2012, 163, 746–754. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.-Z.; Li, C.; Xie, Z.-X.; Dong, H.-P.; Lin, L.; Hong, H.-S. Homology-Driven Proteomics of Dinoflagellates with Unsequenced Genomes Using MALDI-TOF/TOF and Automated De Novo Sequencing. Evid.-Based Complement. Altern. Med. 2011, 2011, 471020. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.-Z.; Zhang, Y.-J.; Zhang, S.-F.; Lin, L.; Hong, H.-S.; Lin, S. Quantitative Proteomic Analysis of Cell Cycle of the Dinoflagellate Prorocentrum donghaiense (Dinophyceae). PLoS ONE 2013, 8, e63659. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.-L.; Zhang, H.-Y.; Guo, Z.-Y.; Ying, W.-T.; Qian, X.-H.; Wang, J.-L. Comparison of Three Methods of Protein Extraction from Dermatophagoides Pteronyssinus for Two-dimensional Electrophoresis. Chin. Med. Sci. J. 2009, 24, 64–68. [Google Scholar] [CrossRef]
- Xiong, J.; Yang, Q.; Kang, J.; Sun, Y.; Zhang, T.; Margaret, G.; Ding, W. Simultaneous isolation of DNA, RNA, and protein from Medicago truncatula L. Electrophoresis 2011, 32, 321–330. [Google Scholar] [CrossRef] [PubMed]
- Li, S.F.; Wang, F.; Zhang, G.H.; Xu, H.N. Establishment of two-dimensional electrophoresis system of caudal gland. China J. Chin. Mater. Med. 2012, 37, 2542–2547. [Google Scholar]
- Pop, C.; Ameling, S.; Empen, K.; Rüdebusch, J.; Dhople, V.M.; Felix, S.B.; Loghin, F.; Völker, U.; Hammer, E. Proteome analysis of heart biopsies using a TRIzol-based protein extraction. Clin. Chim. Acta 2015, 438, 246–247. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, H.; Hasegawa, K.; Esumi, M. Protein from the fraction remaining after RNA extraction is useful for proteomics but care must be exercised in its application. Exp. Mol. Pathol. 2013, 95, 46–50. [Google Scholar] [CrossRef] [PubMed]
- Di, G.; You, W.; Yu, J.; Wang, D.; Ke, C. Genetic changes in muscle protein following hybridization between Haliotis diversicolor reeve Japan and Taiwan populations revealed using a proteomic approach. Proteomics 2013, 13, 845–859. [Google Scholar] [CrossRef] [PubMed]
- Di, G.; Luo, X.; You, W.; Zhao, J.; Kong, X.; Ke, C. Proteomic analysis of muscle between hybrid abalone and parental lines Haliotis gigantea Reeve and Haliotis discus hannai Ino. Heredity 2015, 114, 564–575. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Ji, C.; Wei, L.; Zhao, J. Evaluation of protein extraction protocols for 2 DE in marine ecotoxicoproteomics. Proteomics 2013, 13, 3205–3210. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Zhao, H.; Yang, T.; Ruan, S.; Wang, H.; Xiang, N.; Zhou, H.; Li, Q.X.; Diao, X. Comparative evaluation of five protocols for protein extraction from stony corals (Scleractinia) for proteomics. Electrophoresis 2018, 39, 1062–1070. [Google Scholar] [CrossRef] [PubMed]
- Jiang, X.-W.; Wang, J.; Chan, L.; Lam, P.; Gu, J.-D. Comparison of three protein extraction procedures from toxic and non-toxic dinoflagellates for proteomics analysis. Ecotoxicology 2015, 24, 1395–1406. [Google Scholar] [CrossRef] [PubMed]
- Kline, K.G.; Frewen, B.; Bristow, M.R.; Maccoss, M.J.; Wu, C.C. High quality catalog of proteotypic peptides from human heart. J. Proteome Res. 2008, 7, 5055–5061. [Google Scholar] [CrossRef] [PubMed]
- Hsu, T.-Y.; Yang, K.D.; Tsai, C.-C.; Ou, C.-Y.; Cheng, B.-H.; Wong, Y.-H.; Hung, H.-N.; Chou, A.-K.; Hsiao, C.-C.; Lin, H. Proteomic profiling reveals α1-antitrypsin, α1-microglobulin, and clusterin as preeclampsia-related serum proteins in pregnant women. Taiwan. J. Obstet. Gynecol. 2015, 54, 499–504. [Google Scholar] [CrossRef] [PubMed]
- Pappa, I.K.; Lygirou, V.; Kontostathi, G.; Zoidakis, J.; Makridakis, M.; Vougas, K.; Daskalakis, G.; Polyzos, A.; Anagnou, P.N. Proteomic Analysis of Normal and Cancer Cervical Cell Lines Reveals Deregulation of Cytoskeleton-associated Proteins. Cancer Genom. Proteom. 2017, 14, 253–266. [Google Scholar]
- Knigge, T.; Letendre, J.; Monsinjon, T. Sample preparation for two-dimensional gel electrophoresis: Considering the composition of biological material. Proteomics 2013, 13, 3106–2108. [Google Scholar] [CrossRef] [PubMed]
- Malafaia, C.B. Selection of a protein solubilization method suitable for phytopathogenic bacteria: A proteomics approach. Proteome Sci. 2015, 13, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.-Z. Comparative studies of four protein preparation methods for proteomic study of the dinoflagellate Alexandrium sp. using two-dimensional electrophoresis. Harmful Algae 2009, 8, 685–692. [Google Scholar] [CrossRef]
- Ramagli, L.S.; Rodriguez, L.V. Quantitation of microgram amounts of protein in two-dimensional polyacrylamide gel electrophoresis sample buffer. Eelectrophoresis 1985, 6, 559–563. [Google Scholar] [CrossRef]
| Study Organism/Sample Type | Selected Sample(s) | Year | Reference |
|---|---|---|---|
| Halophile microorganisms | Haloferax volcanii | 2006 | [19] |
| Rat | Spinal cords segment | 2007 | [20] |
| Cell lines | Neck squamous cell carcinoma cell lines (SCC-25; FaDu) | 2007 | [22] |
| MCF-7 cells | 2009 | [21] | |
| Human neuroblastoma cells (SH-SY5Y cells) | 2012 | [23] | |
| Dinoflagellates | Alexandrium sp.; Scrippsiella sp. | 2008 | [24] |
| Alexandriumcatenella | 2011 | [27] | |
| Alexandriumcatenella (DH01) | 2012 | [25] | |
| Lingulodinium sp. | 2012 | [26] | |
| Prorocentrum donghaiense | 2013 | [28] | |
| Alexandrium minutum (AMTK4; AL1TAB); Alexandrium tamarense (ATCI01; SP3B8-3); Gymnodinium catenatum (GCHK; GCEO) | 2015 | [38] | |
| Mites | Dermatophagoides pteronyssinus | 2009 | [29] |
| Plants | Medicago truncatula tissues | 2011 | [30] |
| Clinical samples | Human heart tissue | 2008 | [39] |
| Caudal gland | 2012 | [31] | |
| Non-cancerous liver tissues | 2013 | [33] | |
| Heart biopsies | 2015 | [32] | |
| Marine animals | Mytilus galloprovincialis (mussel); Paralichthys olivaceus (flounder); Nereis diversicolor (Polychaete) | 2013 | [36] |
| Haliotis diversicolor reeve | 2013 | [34] | |
| Haliotis gigantean reeve; Haliotis discus hannai lno | 2015 | [35] | |
| Reef corals | Acropora hyacinthus; Acropora humilis; Acropora muricata | 2018 | [37] |
| Dried seafood and dried tonic foods | Dried Abalone slices; Dried Whelk slices; Dried Ganoderma; Dried Caterpillar fungus | 2018 | This study |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chan, K.-K.; Kwok, C.S.-N.; Sze, E.T.-P.; Lee, F.W.-F. Evaluation of the Use of TRIzol-Based Protein Extraction Approach for Gel-Based Proteomic Analysis of Dried Seafood Products and Chinese Tonic Foods. Int. J. Mol. Sci. 2018, 19, 1998. https://doi.org/10.3390/ijms19071998
Chan K-K, Kwok CS-N, Sze ET-P, Lee FW-F. Evaluation of the Use of TRIzol-Based Protein Extraction Approach for Gel-Based Proteomic Analysis of Dried Seafood Products and Chinese Tonic Foods. International Journal of Molecular Sciences. 2018; 19(7):1998. https://doi.org/10.3390/ijms19071998
Chicago/Turabian StyleChan, Kin-Ka, Celia Sze-Nga Kwok, Eric Tung-Po Sze, and Fred Wang-Fat Lee. 2018. "Evaluation of the Use of TRIzol-Based Protein Extraction Approach for Gel-Based Proteomic Analysis of Dried Seafood Products and Chinese Tonic Foods" International Journal of Molecular Sciences 19, no. 7: 1998. https://doi.org/10.3390/ijms19071998
APA StyleChan, K.-K., Kwok, C. S.-N., Sze, E. T.-P., & Lee, F. W.-F. (2018). Evaluation of the Use of TRIzol-Based Protein Extraction Approach for Gel-Based Proteomic Analysis of Dried Seafood Products and Chinese Tonic Foods. International Journal of Molecular Sciences, 19(7), 1998. https://doi.org/10.3390/ijms19071998






